Meaty Aroma Compounds Enhance MSG Umami Perception Through Allosteric Modulation of T1R1/T1R3 Receptor: Evidence from Nasal Clip Sensory Evaluation and Molecular Dynamics Simulation
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials and Chemicals
2.2. Taste Evaluation
2.2.1. Panelist Training
2.2.2. Sensory Evaluation
2.2.3. Nasal Clip Sensory Evaluation of Umami-Enhancing Meaty Flavorings on MSG
2.2.4. Evaluation of Umami-Enhancing Aroma Compounds on MSG
2.3. Determination of Aroma Compounds in Meaty Flavorings
2.4. Homology Modeling Method of Umami Receptor T1R1/T1R3
2.5. Molecular Dynamics (MD)
2.6. Statistical Analysis
3. Results and Discussions
3.1. Effects of Meaty Flavorings on Umami Characteristics of MSG
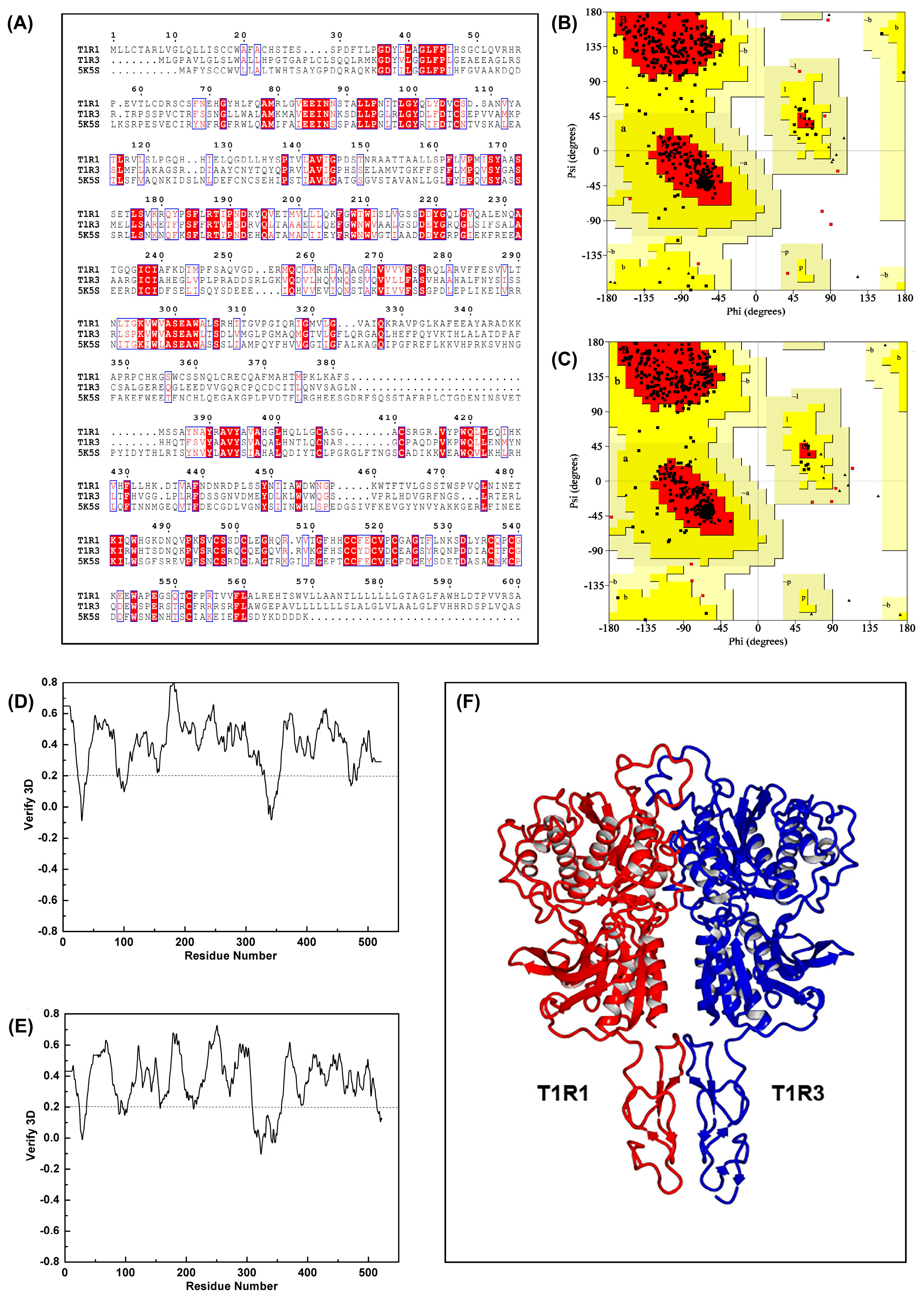
3.2. Homology Modeling of Umami Receptor T1R1/T1R3
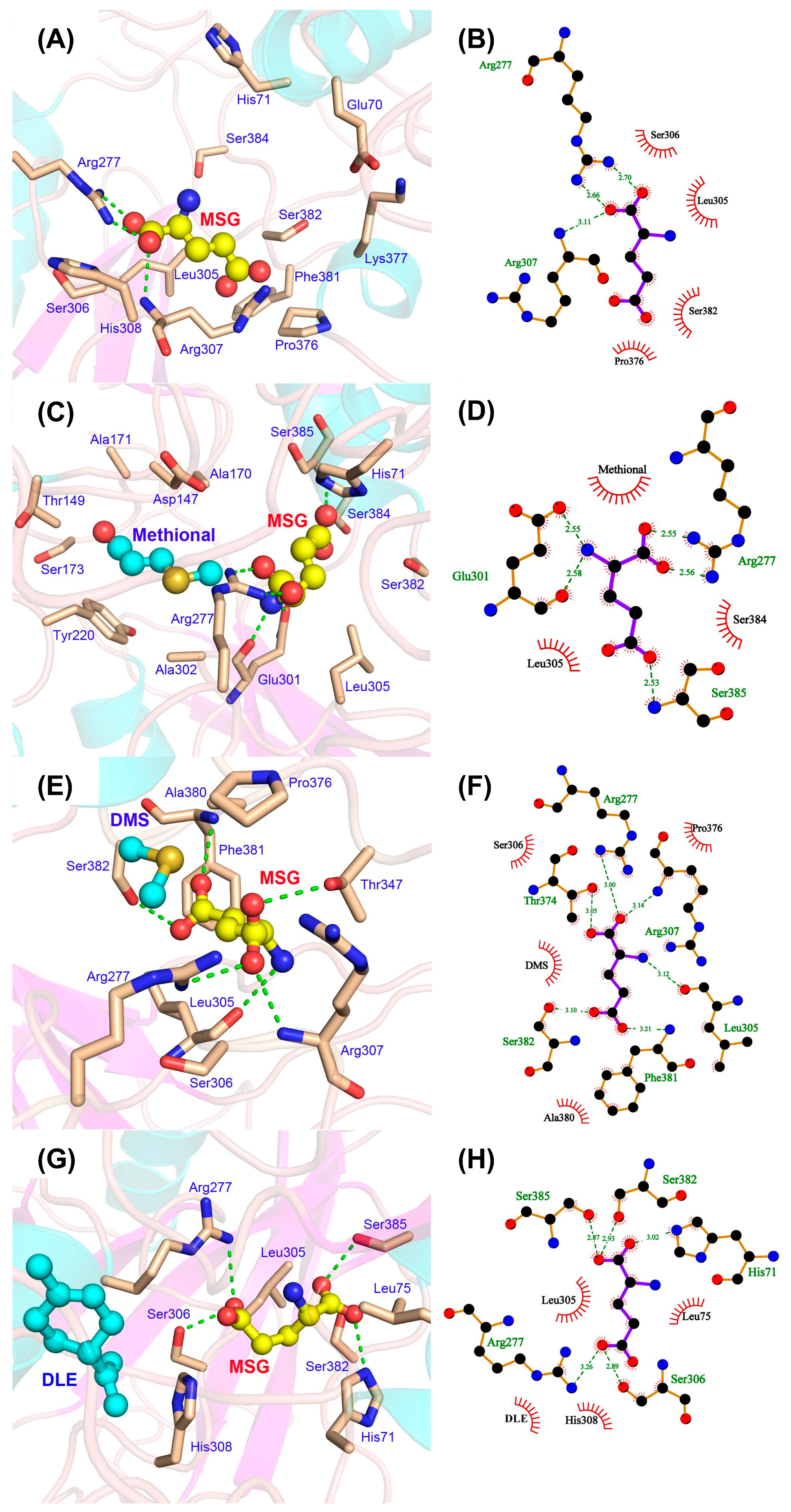
3.3. Modulatory Effects of Aroma Compounds on MSG Binding to the Umami Receptor T1R1/T1R3
3.3.1. Molecular Insights into Umami Enhancement Mechanisms by Aroma Compounds via MD Simulations
3.3.2. Aroma Compound-Induced Stabilization of Key Domains in the Umami Receptor T1R1/T1R3
3.3.3. Allosteric Enhancement of MSG–Receptor Interactions by Aroma Compounds
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Le Quéré, J.-L.; Schoumacker, R. Dynamic instrumental and sensory methods used to link aroma release and aroma perception: A review. Molecules 2023, 28, 6380. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Badar, I.H.; Zhang, L.; Yang, L.; Xu, B. Odor and taste characteristics, transduction mechanism, and perceptual interaction in fermented foods: A review. Crit. Rev. Food Sci. Nutr. 2025, 65, 3947–3965. [Google Scholar] [CrossRef]
- Wang, X.; Cui, B.; Lin, H.; Pan, R.; Zeng, J.; Fang, X.; Liu, Y.; Chen, Z.-Y.; Chen, Y.; Zhu, H. Research progress in saltiness perception and salty substitutes. J. Agric. Food Chem. 2025, 73, 2745–2759. [Google Scholar] [CrossRef] [PubMed]
- Zhou, R.; Zhu, J.; Niu, Y.; Zhang, J.; Xiao, Z.; Zhao, L. Identification of characteristic compounds of sweet orange oil and their sweetening effects on the sucrose solution with sweetness meter, sensory analysis, electronic tongue, and molecular dynamics simulation. Food Chem. 2024, 461, 140815. [Google Scholar] [CrossRef]
- Horie, F.; Sanematsu, K.; Yasumatsu, K.; Hirokawa, T.; Shigemura, N.; Yamashita, A.; Kusakabe, Y. Receptor mechanism producing a sweet taste from plant aroma compounds. Sci. Rep. 2025, 15, 6795. [Google Scholar] [CrossRef]
- Gao, Y.; Wang, W.; Zhang, H.; Chen, H.; Wang, S.; Sun, B. The enhancement of the perception of saltiness by odorants selected from chinese douchi in salt solution. Foods 2022, 11, 2260. [Google Scholar] [CrossRef]
- Lopez, J.; Kerley, T.; Jenkinson, L.; Luckett, C.R.; Munafo, J.P., Jr. Odorants from the thermal treatment of hydrolyzed mushroom protein and cysteine enhance saltiness perception. J. Agric. Food Chem. 2019, 67, 11444–11453. [Google Scholar] [CrossRef]
- Lawrence, G.; Salles, C.; Palicki, O.; Septier, C.; Busch, J.; Thomas-Danguin, T. Using cross-modal interactions to counterbalance salt reduction in solid foods. Int. Dairy J. 2011, 21, 103–110. [Google Scholar] [CrossRef]
- Lawrence, G.; Salles, C.; Septier, C.; Busch, J.; Thomas-Danguin, T. Odour-taste interactions: A way to enhance saltiness in low-salt content solutions. Food Qual. Prefer. 2009, 20, 241–248. [Google Scholar] [CrossRef]
- Djordjevic, J.; Zatorre, R.J.; Jones-gotman, M. Odor-induced changes in taste perception. Exp. Brain Res. 2004, 159, 405–408. [Google Scholar] [CrossRef]
- Mccabe, C.; Rolls, E.T. Umami: A delicious flavor formed by convergence of taste and olfactory pathways in the human brain. Eur. J. Neurosci. 2007, 25, 1855–1864. [Google Scholar] [CrossRef]
- Niimi, J.; Eddy, A.I.; Overington, A.R.; Heenan, S.P.; Silcock, P.; Bremer, P.J.; Delahunty, C.M. Aroma–taste interactions between a model cheese aroma and five basic tastes in solution. Food Qual. Prefer. 2014, 31, 1–9. [Google Scholar] [CrossRef]
- Kitajima, S.; Maruyama, Y.; Kuroda, M. Volatile short-chain aliphatic aldehydes act as taste modulators through the orally expressed calcium-sensing receptor casr. Molecules 2023, 28, 4585. [Google Scholar] [CrossRef]
- Sun-Waterhouse, D.; Wadhwa, S.S. Industry-relevant approaches for minimising the bitterness of bioactive compounds in functional foods: A review. Food Bioprocess Technol. 2013, 6, 607–627. [Google Scholar] [CrossRef]
- Toda, Y.; Nakagita, T.; Hirokawa, T.; Yamashita, Y.; Nakajima, A.; Narukawa, M.; Ishimaru, Y.; Uchida, R.; Misaka, T. Positive/negative allosteric modulation switching in an umami taste receptor (t1r1/t1r3) by a natural flavor compound, methional. Sci. Rep. 2018, 8, 11796. [Google Scholar] [CrossRef] [PubMed]
- Malik, B.; Elkaddi, N.; Turkistani, J.; Spielman, A.I.; Ozdener, M.H. Mammalian taste cells express functional olfactory receptors. Chem. Senses 2019, 44, 289–301. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, M.; Canon, F.; Feron, G.; Neiers, F.; Gamero, A. Impact of oral microbiota on flavor perception: From food processing to in-mouth metabolization. Foods 2021, 10, 2006. [Google Scholar] [CrossRef]
- Cao, C.; Waterhouse, G.I.N.; Sun, W.; Zhao, M.; Sun-Waterhouse, D.; Su, G. Effects of fermentation with tetragenococcus halophilus and zygosaccharomyces rouxii on the volatile profiles of soybean protein hydrolysates. Foods 2023, 12, 4513. [Google Scholar] [CrossRef]
- Boratyn, G.M.; Schäffer, A.A.; Agarwala, R.; Altschul, S.F.; Lipman, D.J.; Madden, T.L. Domain enhanced lookup time accelerated blast. Biol. Direct 2012, 7, 12. [Google Scholar] [CrossRef]
- Šali, A.; Blundell, T.L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 1993, 234, 270–277. [Google Scholar] [CrossRef]
- Maier, J.A.; Martinez, C.; Kasavajhala, K.; Wickstrom, L.; Hauser, K.E.; Simmerling, C. Ff14sb improving the accuracy of protein side chain and backbone parameters from ff99sb. J. Chem. Theory Comput. 2015, 11, 3696–3713. [Google Scholar] [CrossRef]
- Van der spoel, D.; Lindahl, E.; Hess, B.; Groenhof, G.; Mark, A.E.; Berendsen, H.J.C. Gromacs: Fast, flexible, and free. J. Comput. Chem. 2005, 26, 1701–1718. [Google Scholar] [CrossRef]
- Jorgensen, W.L.; Chandrasekhar, J.; Madura, J.D.; Impey, R.W.; Klein, M.L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 1998, 79, 926–935. [Google Scholar] [CrossRef]
- Hess, B.; Bekker, H.; Berendsen, H.J.C.; Fraaije, J.G.E.M. Lincs: A linear constraint solver for molecular simulations. J. Comput. Chem. 1997, 18, 1463–1472. [Google Scholar] [CrossRef]
- Darden, T.; York, D.; Pedersen, L. Particle mesh Ewald: An N log(N) method for Ewald sums in large systems. J. Chem. Phys. 1993, 98, 10089–10092. [Google Scholar] [CrossRef]
- Berendsen, H.J.C.; Postma, J.P.M.; Van gunsteren, W.F.; DiNola, A.; Haak, J.R. Molecular dynamics with coupling to an external bath. J. Chem. Phys. 1984, 81, 3684–3690. [Google Scholar] [CrossRef]
- Martoňák, R.; Laio, A.; Parrinello, M. Predicting crystal structures: The parrinello-rahman method revisited. Phys. Rev. Lett. 2003, 90, 075503. [Google Scholar] [CrossRef]
- Nasri, N.; Beno, N.; Septier, C.; Salles, C.; Thomas-Danguin, T. Cross-modal interactions between taste and smell: Odour-induced saltiness enhancement depends on salt level. Food Qual. Prefer. 2011, 22, 678–682. [Google Scholar] [CrossRef]
- Nasri, N.; Septier, C.; Beno, N.; Salles, C.; Thomas-Danguin, T. Enhancing salty taste through odour–taste–taste interactions: Influence of odour intensity and salty tastants’ nature. Food Qual. Prefer. 2013, 28, 134–140. [Google Scholar] [CrossRef]
- Ogasawara, Y.; Mochimaru, S.; Ueda, R.; Ban, M.; Kabuto, S.; Abe, K. Preparation of an aroma fraction from dried bonito by steam distillation and its effect on modification of salty and umami taste qualities. J. Food Sci. 2016, 81, C308–C316. [Google Scholar] [CrossRef] [PubMed]
- Wise, P.M.; Ledyard, A. Human oral sensitivity to and taste modulation by 3-mercapto-2-methylpentan-1-ol. Chemosens. Percept. 2022, 15, 70–86. [Google Scholar] [CrossRef]
- Geng, Y.; Mosyak, L.; Kurinov, I.; Zuo, H.; Sturchler, E.; Cheng, T.C.; Subramanyam, P.; Brown, A.P.; Brennan, S.C.; Mun, H.-C.; et al. Structural mechanism of ligand activation in human calcium-sensing receptor. eLife 2015, 5, e13662. [Google Scholar] [CrossRef]
- Singh, K.D.; Muthusamy, K. Molecular modeling, quantum polarized ligand docking and structure-based 3d-qsar analysis of the imidazole series as dual at1 and eta receptor antagonists. Acta Pharmacol. Sin. 2013, 34, 1592–1606. [Google Scholar] [CrossRef] [PubMed]
- Zhang, N.; Cui, Z.; Li, M.; Fan, Y.; Liu, J.; Wang, W.; Zhang, Y.; Liu, Y. Typical umami ligand-induced binding interaction and conformational change of t1r1-vft. J. Agric. Food Chem. 2022, 70, 11652–11666. [Google Scholar] [CrossRef] [PubMed]
- Diepeveen, J.; Moerdijk-poortvliet, T.C.W.; van der Leij, F.R. Molecular insights into human taste perception and umami tastants: A review. J. Food Sci. 2022, 87, 1449–1465. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Sun-waterhouse, D.; Su, G.; Zhao, M. New insight into umami receptor, umami/umami-enhancing peptides and their derivatives: A review. Trends Food Sci. Technol. 2019, 88, 429–438. [Google Scholar] [CrossRef]
- Liu, H.; Da, L.-T.; Liu, Y. Understanding the molecular mechanism of umami recognition by t1r1-t1r3 using molecular dynamics simulations. Biochem. Biophys. Res. Commun. 2019, 514, 967–973. [Google Scholar] [CrossRef]
- Pallante, L.; Malavolta, M.; Grasso, G.; Korfiati, A.; Mavroudi, S.; Mavkov, B.; Kalogeras, A.; Alexakos, C.; Martos, V.; Amoroso, D.; et al. On the human taste perception: Molecular-level understanding empowered by computational methods. Trends Food Sci. Technol. 2021, 116, 445–459. [Google Scholar] [CrossRef]





| No. | CAS | Compound | RI Calc | RI Ref | m/z | Integral Fragment | ID * | M | B | S |
|---|---|---|---|---|---|---|---|---|---|---|
| Sulfur-containing compounds | ||||||||||
| 1 | 110-66-7 | 1-Pentanethiol | 818 | 817 | 104, 42 | 104 | MS, RI | -- | 2.2639 | -- |
| 2 | 513-44-0 | 1-Propanethiol, 2-methyl- | 685 | 685 | 41, 90 | 41 | MS, RI | 2.0815 | -- | -- |
| 3 | 24295-03-2 | 2-Acetylthiazole | 1023 | 1023 | 43, 99, 127 | 43 | MS, RI | 2.7729 | -- | -- |
| 4 | NA | 2-Furfuryl-2-methyl-3-furyl disulfide | 1654 | 1655 | 81 | 81 | MS, RI | 64.3061 | -- | -- |
| 5 | 98-02-2 | 2-Furfurylthiol | 913 | 913 | 81, 53, 114 | 81 | MS, RI | 290.5882 | -- | -- |
| 6 | 28588-75-2 | 2-Methyl-3-[(2-methylfuran-3-yl)disulfanyl]furan | 1553 | 1547 | 113, 226 | 113 | MS, RI | 607.1735 | -- | -- |
| 7 | 28588-74-1 | 2-Methyl-3-furanthiol | 870 | 870 | 114 | 114 | MS, RI | 99.5886 | 15.1273 | -- |
| 8 | 5271-38-5 | 2-Methylthioethanol | 850 | 850 | 61, 92 | 61 | MS, RI | -- | -- | 0.0123 |
| 9 | 1618-26-4 | Bis(methylthio)methane | 892 | 892 | 61, 108 | 61 | MS, RI | -- | -- | 0.0585 |
| 10 | 624-92-0 | Dimethyl disulfide | 744 | 744 | 94, 79 | 94 | MS, RI | 0.2441 | -- | 29.6650 |
| 11 | 75-18-3 | Dimethyl sulfide | -- | 521 | 62, 47 | 62 | MS | -- | -- | 9.9339 |
| 12 | 5756-24-1 | Dimethyl tetrasulfide | 1215 | 1215 | 79, 158 | 79 | MS, RI | -- | -- | 0.0117 |
| 13 | 3658-80-8 | Dimethyl trisulfide | 975 | 975 | 126 | 126 | MS, RI | 0.5471 | -- | 1.7146 |
| 14 | 7783-06-4 | Hydrogen sulfide | -- | -- | 34 | 34 | MS | 0.9203 | -- | -- |
| 15 | 74-93-1 | Methanethiol | -- | 464 | 47 | 47 | MS | -- | -- | 0.1530 |
| 16 | 57500-00-2 | Methyl furfuryl disulfide | 1223 | 1226 | 81 | 81 | MS, RI | 6.6786 | -- | -- |
| 17 | 4861-58-9 | Thiophene, 2-pentyl- | 1168 | 1170 | 97, 154 | 97 | MS, RI | 434.6640 | 8.1651 | 0.1374 |
| 18 | 616-44-4 | Thiophene, 3-methyl | 772 | 772 | 97 | 97 | MS, RI | 0.4060 | -- | -- |
| Pyrazines | ||||||||||
| 19 | 14667-55-1 | 2,3,5-Trimethylpyrazine | 1006 | 1006 | 122, 42 | 122 | MS, RI | 3.7262 | -- | 2.2367 |
| 20 | 15707-24-1 | 2,3-Diethylpyrazine | 1081 | 1080 | 121, 136 | 121 | MS, RI | -- | -- | 1.3320 |
| 21 | 5910-89-4 | 2,3-Dimethylpyrazine | 919 | 919 | 108, 67 | 108 | MS, RI | -- | -- | 3.3991 |
| 22 | 109-08-0 | 2-Methylpyrazine | 830 | 830 | 94, 67 | 94 | MS, RI | -- | -- | 1.1609 |
| Alkenes | ||||||||||
| 23 | 18172-67-3 | (−)-β-Pinene | 976 | -- | 93, 41 | 93 | MS | -- | -- | 0.1352 |
| 24 | 7785-70-8 | (+)-α-Pinene | 938 | 937 | 93 | 93 | MS, RI | -- | -- | 0.0819 |
| 25 | 18368-95-1 | 1,3,8-p-Menthatriene | 1117 | 1119 | 119, 134 | 119 | MS, RI | -- | -- | 0.0000 |
| 26 | 586-62-9 | 1-Methyl-4-(1-methylethylidene) cyclohexene | 1093 | 1094 | 93, 121, 136 | 93 | MS, RI | -- | 1.2262 | -- |
| 27 | 28634-89-1 | 2-Thujene | 978 | 978 | 93 | 93 | MS, RI | -- | 0.7750 | -- |
| 28 | 13466-78-9 | 3-Carene | 1015 | 1015 | 93 | 93 | MS, RI | -- | 5.7671 | -- |
| 29 | 104-46-1 | Anethole | 1294 | 1289 | 148 | 148 | MS, RI | -- | 6550.5112 | -- |
| 30 | 79-92-5 | Camphene | 954 | 954 | 93, 121 | 93 | MS, RI | -- | 0.3364 | -- |
| 31 | 87-44-5 | Caryophyllene | 1441 | 1441 | 93, 133 | 93 | MS, RI | 0.3445 | 128.6968 | -- |
| 32 | 1139-30-6 | Caryophyllene oxide | 1614 | 1613 | 93 | 93 | MS, RI | -- | 0.5720 | -- |
| 33 | 18252-46-5 | Cis-α-bergamotene | 1428 | 1428 | 119, 93 | 119 | MS, RI | -- | 103.4251 | -- |
| 34 | 3856-25-5 | Copaene | 1391 | 1391 | 119, 105, 161 | 119 | MS, RI | -- | 63.5100 | -- |
| 35 | 5989-27-5 | D-Limonene | 1034 | -- | 68, 93 | 68 | MS | 0.8658 | 27.2089 | 5.6548 |
| 36 | 6753-98-6 | Humulene | 1476 | 1477 | 93 | 93 | MS, RI | -- | 13.0814 | -- |
| 37 | 78-70-6 | Linalool | 1102 | 1102 | 71, 93 | 71 | MS, RI | -- | 4.2015 | -- |
| 38 | 58319-06-5 | Sesquithujene | 1414 | 1417 | 119, 93 | 119 | MS, RI | -- | 3.2553 | -- |
| 39 | 562-74-3 | Terpinen-4-ol | 1186 | 1186 | 71, 111 | 71 | MS, RI | -- | 0.8337 | -- |
| 40 | 546-80-5 | Thujone | 1112 | 1112 | 81, 110 | 81 | MS, RI | -- | 0.7441 | -- |
| 41 | 13474-59-4 | Trans-α-bergamotene | 1449 | 1450 | 119, 93 | 119 | MS, RI | -- | 103.8404 | -- |
| 42 | 495-60-3 | Zingiberene | 1506 | 1506 | 119, 93 | 119 | MS, RI | -- | 53.5610 | -- |
| 43 | 469-61-4 | α-Cedrene | 1436 | 1436 | 119 | 119 | MS, RI | -- | 4.6260 | -- |
| 44 | 502-61-4 | α-Farnesene | 1513 | 1513 | 93 | 93 | MS, RI | -- | 3.2607 | -- |
| 45 | 495-61-4 | β-Bisabolene | 1520 | 1520 | 69, 93 | 69 | MS, RI | -- | 20.1507 | -- |
| 46 | 515-13-9 | β-Elemene | 1405 | 1405 | 93, 81, 107 | 93 | MS, RI | -- | 13.1540 | -- |
| 47 | 127-91-3 | β-Pinene | 982 | 982 | 93 | 93 | MS, RI | -- | 2.3582 | -- |
| 48 | 17066-67-0 | β-Selinene | 1510 | 1509 | 105 | 105 | MS, RI | -- | 1.1195 | -- |
| 49 | 20307-83-9 | β-Sesquiphellandrene | 1538 | 1536 | 69, 93 | 69 | MS, RI | -- | 14.9319 | -- |
| 50 | 99-85-4 | γ-Terpinene | 1064 | 1064 | 93, 136 | 93 | MS, RI | -- | 3.0459 | 0.0401 |
| 51 | 20307-84-0 | δ-Elemene | 1350 | 1351 | 121, 93, 136, 161 | 121 | MS, RI | -- | 23.6534 | -- |
| 52 | 106-99-0 | 1,3-Butadiene | -- | 400 | 54, 39 | 54 | MS | -- | -- | 0.0988 |
| 53 | 872-05-9 | 1-Decene | 992 | 992 | 43, 56, 70 | 43 | MS, RI | -- | -- | 0.0933 |
| 54 | 112-41-4 | 1-Dodecene | 1196 | 1195 | 69, 43 | 69 | MS, RI | -- | -- | 0.0746 |
| 55 | 124-11-8 | 1-Nonene | 892 | 892 | 43, 56 | 43 | MS, RI | 0.8351 | 1.7049 | -- |
| 56 | 627-20-3 | 2-Pentene, (Z)- | -- | 510 | 55, 70 | 55 | MS | -- | -- | 0.9789 |
| 57 | 691-37-2 | 4-Methyl-1-Pentene | -- | 553 | 43, 56 | 43 | MS | -- | -- | 0.2348 |
| 58 | 100-42-5 | Styrene | 894 | 894 | 104, 78 | 104 | MS, RI | -- | 0.7356 | -- |
| 59 | 17699-05-7 | α-Bergamotene | 1428 | 1428 | 93, 119 | 93 | MS, RI | 0.0505 | -- | -- |
| Alcohols | ||||||||||
| 60 | 928-96-1 | (Z)-hex-3-en-1-ol | 855 | 855 | 67, 82, 41, 55 | 67 | MS, RI | -- | -- | 2.2041 |
| 61 | 1565-80-6 | 1-Butanol, 2-methyl-, (S)- | -- | -- | 56, 41 | 56 | MS | -- | -- | 0.3792 |
| 62 | 15250-22-3 | 1-Octanol, 2,7-dimethyl- | -- | 144 | 43, 58 | 43 | MS | -- | -- | 0.1527 |
| 63 | 3391-86-4 | 1-Octen-3-ol | 982 | 982 | 57 | 57 | MS, RI | 5.0144 | -- | 10.8963 |
| 64 | 928-97-2 | 3-Hexen-1-ol, (E)- | 855 | 855 | 67, 41 | 67 | MS, RI | -- | -- | 0.0231 |
| 65 | 123-51-3 | Isoamyl alcohol | -- | 730 | 55, 70 | 55 | MS | -- | -- | 2.9690 |
| Ketones | ||||||||||
| 66 | 600-14-6 | 2,3-Pentanedione | -- | 698 | 43, 57, 100 | 43 | MS | 0.6208 | -- | -- |
| 67 | 78-93-3 | 2-Butantone | -- | 609 | 43, 72 | 43 | MS | 1.4412 | 5.7114 | -- |
| 68 | 693-54-9 | 2-Decanone | 1195 | 1195 | 58, 43, 71 | 58 | MS, RI | 0.4498 | -- | -- |
| 69 | 110-43-0 | 2-Heptanon | 889 | 889 | 43, 58 | 43 | MS, RI | -- | -- | 0.1451 |
| 70 | 821-55-6 | 2-Nonanone | 1093 | 1094 | 58 | 58 | MS, RI | -- | 0.4057 | 0.0206 |
| 71 | 111-13-7 | 2-Octanone | 992 | 992 | 43, 58 | 43 | MS, RI | 1.0648 | -- | -- |
| 72 | 107-87-9 | 2-Pentanone | -- | 697 | 43, 86 | 43 | MS | 1.1319 | 4.2671 | -- |
| 73 | 563-80-4 | 3-Methyl-2-butanone | -- | 653 | 43, 86 | 43 | MS | -- | -- | 0.2921 |
| 74 | 96-22-0 | 3-Pentanone | 700 | 700 | 57, 86 | 57 | MS, RI | 0.8016 | 3.3342 | -- |
| 75 | 2216-87-7 | 3-Undecanone | 1290 | 1283 | 141, 57, 72 | 141 | MS, RI | -- | 2.1575 | -- |
| 76 | 30086-02-3 | Trans,trans-3,5-octadien-2-one | 1097 | 1098 | 95, 43 | 95 | MS, RI | -- | -- | 0.0048 |
| Furan(one)s | ||||||||||
| 77 | 104-61-0 | 2(3H)-Furanone, dihydro-5-pentyl- | 1363 | 1363 | 85, 44 | 85 | MS, RI | -- | -- | 0.0115 |
| 78 | 3188-00-9 | Dihydro-2-methyl-3-furanone | 808 | 808 | 43, 72, 100 | 43 | MS, RI | 12.5517 | -- | 0.1365 |
| 79 | 1197-40-6 | Furan, 2,2′-methylenebis- | 1088 | 1089 | 91, 148 | 91 | MS, RI | 0.7623 | -- | -- |
| 80 | 930-27-8 | Furan, 3-methyl- | -- | 614 | 82, 53 | 82 | MS | 2.6717 | -- | -- |
| 81 | 3777-69-3 | Furan, 2-pentyl | 994 | 994 | 81 | 81 | MS, RI | 0.6586 | -- | -- |
| Aldehydes | ||||||||||
| 82 | 25152-84-5 | (E,E)-2,4-Decadienal | 1324 | 1324 | 81 | 81 | MS, RI | -- | -- | 0.0143 |
| 83 | 557-48-2 | 2,6-Nonadienal, (E,Z) | 1156 | 1156 | 41, 70 | 41 | MS, RI | -- | -- | 0.3315 |
| 84 | 3913-81-3 | 2-Decenal, (E)- | 1268 | 1268 | 70, 55 | 70 | MS, RI | 0.0544 | 1.1817 | -- |
| 85 | 13019-16-4 | 2-Octenal, 2-butyl- | 1379 | 1378 | 55 | 55 | MS, RI | -- | 0.6627 | -- |
| 86 | 590-86-3 | 3-Methylbutanal | -- | 655 | 44, 58, 71 | 44 | MS | -- | -- | 0.0901 |
| 87 | 6728-31-0 | 4-Heptenal, (Z)- | 902 | 902 | 41, 55, 68, 84 | 41 | MS, RI | -- | -- | 0.2251 |
| 88 | 122-03-2 | 4-Isopropylbenzaldehyde | 1252 | 1252 | 133, 105 | 133 | MS, RI | -- | 3.8862 | -- |
| 89 | 100-52-7 | Benzaldehyde | 966 | 965 | 105, 77, 51 | 105 | MS, RI | -- | 6.8549 | 0.3813 |
| 90 | 112-31-2 | Decanal | 1208 | 1208 | 57 | 57 | MS, RI | 0.1426 | 3.0725 | -- |
| 91 | 56-82-6 | DL-Glyceraldehyde | -- | -- | 91, 61 | 91 | MS | -- | -- | 0.0174 |
| 92 | 98-01-1 | Furfural | 835 | 835 | 95, 39 | 95 | MS, RI | -- | -- | 0.2524 |
| 93 | 111-71-7 | Heptanal | 903 | 903 | 57 | 57 | MS, RI | 0.1651 | 1.6242 | -- |
| 94 | 66-25-1 | Hexanal | 801 | 801 | 44, 56 | 44 | MS, RI | 0.2643 | 1.2348 | 0.0580 |
| 95 | 3268-49-3 | Methional | 909 | 909 | 48, 104 | 48 | MS, RI | -- | -- | 8.5728 |
| 96 | 124-19-6 | Nonanal | 1105 | 1105 | 57 | 57 | MS, RI | 1.5959 | 9.2310 | 0.0256 |
| 97 | 124-13-0 | Octanal | 1004 | 1004 | 43, 57, 84 | 43 | MS, RI | 0.1715 | 0.8794 | -- |
| 98 | 124-25-4 | Tetradecanal | 1579 | 1579 | 82 | 82 | MS, RI | -- | 39.9319 | -- |
| Esters | ||||||||||
| 99 | 123-92-2 | 1-Butanol, 3-methyl-, acetate | 878 | 878 | 43, 55, 70 | 43 | MS, RI | -- | -- | 0.0938 |
| 100 | 110-45-2 | 1-Butanol, 3-methyl-, formate | 792 | 792 | 55, 70 | 55 | MS, RI | -- | -- | 0.0827 |
| 101 | 868-57-5 | 2-Methylbutanoic acid methyl ester | 771 | 771 | 88, 57 | 88 | MS, RI | -- | -- | 0.0693 |
| 102 | 3681-82-1 | 3-Hexen-1-ol,1-acetate, (3E)- | 1008 | 1008 | 67, 82 | 67 | MS, RI | -- | 6.7108 | -- |
| 103 | 2035-99-6 | 3-Methylbutyl octanoate | 1453 | 1452 | 70 | 70 | MS, RI | -- | -- | 0.0077 |
| 104 | 140-11-4 | Acetic acid benzyl ester | 1170 | 1170 | 108, 91, 150 | 108 | MS, RI | -- | 1.4869 | -- |
| 105 | 112-14-1 | Acetic acid octyl ester | 1212 | 1213 | 43, 56, 70 | 43 | MS, RI | -- | 46.0491 | -- |
| 106 | 123-86-4 | Acetic acid, butyl ester | 816 | 816 | 43, 56 | 43 | MS, RI | 0.8136 | -- | 0.0524 |
| 107 | 105-54-4 | Butanoic acid, ethyl ester | 803 | 803 | 71, 43, 88 | 71 | MS, RI | 0.3658 | 0.9472 | 0.5675 |
| 108 | 623-42-7 | Butanoic acid, methyl ester | 723 | 723 | 74, 43 | 74 | MS, RI | 1.1396 | 3.4593 | 0.2654 |
| 109 | 589-75-3 | Butyl caprylate | 1388 | 1388 | 56, 145 | 56 | MS, RI | 0.4217 | -- | -- |
| 110 | 110-38-3 | Decanoic acid, ethyl ester | 1395 | 1395 | 88, 101 | 88 | MS, RI | 6.4838 | 5.0649 | -- |
| 111 | 110-42-9 | Decanoic acid, methyl ester | 1327 | 1327 | 74, 87, 143 | 74 | MS, RI | -- | -- | 0.0141 |
| 112 | 616-38-6 | Dimethyl carbonate | -- | 620 | 45, 59 | 45 | MS | -- | -- | -- |
| 113 | 106-33-2 | Dodecanoic acid, ethyl ester | 1594 | 1594 | 88, 101 | 88 | MS, RI | -- | 1.4096 | -- |
| 114 | 141-78-6 | Ethyl acetate | -- | 615 | 43 | 43 | MS | 4.3639 | -- | 0.2620 |
| 115 | 93-89-0 | Ethyl benzoate | 1178 | 1177 | 105, 77, 122 | 105 | MS, RI | -- | 1.3972 | -- |
| 116 | 105-37-3 | Ethyl propionate | 726 | 726 | 57, 77, 43 | 57 | MS, RI | -- | -- | 0.0315 |
| 117 | 142-92-7 | Hexyl acetate | 1014 | 1014 | 43, 56 | 43 | MS, RI | -- | 11.8858 | -- |
| 118 | 554-12-1 | Methyl propionate | -- | 622 | 57, 88 | 57 | MS | -- | 2.3802 | 0.1936 |
| 119 | 106-32-1 | Octanoic acid, ethyl ester | 1197 | 1197 | 88, 101 | 88 | MS, RI | 43.0167 | 2.2618 | 0.0115 |
| 120 | 111-11-5 | Octanoic acid, methyl ester | 1125 | 1125 | 74, 87 | 74 | MS, RI | 3.0726 | 0.3832 | 0.2696 |
| 121 | 80-26-2 | Terpinyl acetate | 1359 | 1359 | 121, 93, 136 | 121 | MS, RI | -- | 10.8402 | -- |
| 122 | 124-06-1 | Tetradecanoic acid, ethyl ester | 1793 | 1793 | 88, 101 | 88 | MS, RI | -- | 5.6830 | -- |
| 123 | 102-76-1 | Triacetin | 1349 | 1350 | 43, 103, 145 | 43 | MS, RI | -- | -- | 0.0132 |
| ethers | ||||||||||
| 124 | 140-67-0 | Estragole | 1205 | 1204 | 148 | 148 | MS, RI | -- | 109.1040 | -- |
| Acids | ||||||||||
| 125 | 334-48-5 | Decanoic acid | 1367 | 1367 | 60, 73, 129 | 60 | MS, RI | 0.7768 | -- | -- |
| 126 | 143-07-7 | Dodecanoic acid | 1563 | 1562 | 73 | 73 | MS, RI | -- | 1.8547 | -- |
| 127 | 124-07-2 | Octanoic acid | 1175 | 1175 | 60, 73 | 60 | MS, RI | 2.3087 | 1.4629 | -- |
| Others | ||||||||||
| 128 | 95-47-6 | 1,2-Xylene | 896 | 896 | 91, 106 | 91 | MS, RI | 0.7234 | -- | 0.0595 |
| 129 | 91-57-6 | 2-Methylnaphthalene | 1308 | 1308 | 142 | 142 | MS, RI | -- | 2.4538 | -- |
| 130 | 100-41-4 | Ethylbenzene | 864 | 864 | 91, 106 | 91 | MS, RI | 0.8535 | 0.9918 | -- |
| 131 | 527-84-4 | o-Cymene | 1029 | 1029 | 119 | 119 | MS, RI | -- | 8.9806 | 0.1382 |
| 132 | 108-88-3 | Toluene | 767 | 767 | 91 | 91 | MS, RI | 1.3531 | 2.4152 | -- |
| Crystal Structure | T1R1 | T1R3 | ||||||
|---|---|---|---|---|---|---|---|---|
| Score | E Value | Identity | Cover | Score | E Value | Identity | Cover | |
| 5K5S | 299 | 2 × 10−90 | 31.59% | 64% | 283 | 2 × 10−84 | 31.05% | 64% |
| 5X2M | 267 | 4 × 10−80 | 34.75% | 55% | 204 | 5 × 10−57 | 30.17% | 54% |
| 6N52 | 266 | 4 × 10−76 | 26.85% | 95% | 124 | 1 × 10−28 | 25.71% | 75% |
| 5FBH | 239 | 8 × 10−69 | 31.02% | 54% | 228 | 1 × 10−64 | 30.11% | 55% |
| 2E4U | 164 | 2 × 10−42 | 26.22% | 62% | 166 | 6 × 10−43 | 26.00% | 63% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, Y.; Zhang, J.; Zhao, M.; Zhao, X.; Su, G. Meaty Aroma Compounds Enhance MSG Umami Perception Through Allosteric Modulation of T1R1/T1R3 Receptor: Evidence from Nasal Clip Sensory Evaluation and Molecular Dynamics Simulation. Foods 2025, 14, 3041. https://doi.org/10.3390/foods14173041
Zhao Y, Zhang J, Zhao M, Zhao X, Su G. Meaty Aroma Compounds Enhance MSG Umami Perception Through Allosteric Modulation of T1R1/T1R3 Receptor: Evidence from Nasal Clip Sensory Evaluation and Molecular Dynamics Simulation. Foods. 2025; 14(17):3041. https://doi.org/10.3390/foods14173041
Chicago/Turabian StyleZhao, Yaqi, Jianan Zhang, Mouming Zhao, Xuan Zhao, and Guowan Su. 2025. "Meaty Aroma Compounds Enhance MSG Umami Perception Through Allosteric Modulation of T1R1/T1R3 Receptor: Evidence from Nasal Clip Sensory Evaluation and Molecular Dynamics Simulation" Foods 14, no. 17: 3041. https://doi.org/10.3390/foods14173041
APA StyleZhao, Y., Zhang, J., Zhao, M., Zhao, X., & Su, G. (2025). Meaty Aroma Compounds Enhance MSG Umami Perception Through Allosteric Modulation of T1R1/T1R3 Receptor: Evidence from Nasal Clip Sensory Evaluation and Molecular Dynamics Simulation. Foods, 14(17), 3041. https://doi.org/10.3390/foods14173041








