Emerging Understanding of Gut Microbiome in Colorectal Cancer and Food-Related Intervention Strategies
Abstract
1. Introduction
2. Associations Between Gut Microbiome and CRC
2.1. Fusobacterium Nucleatum Is a Major Risk Factor of CRC
| Microbes | Organisms | CRC-Modulating Effects | Proposed Mechanisms | References | Correlations | Models |
|---|---|---|---|---|---|---|
| F. nucleatum | Frozen tumor specimens from humans | Positive correlation with lymph node metastasis | ND | [28] | Association | Human |
| Human samples and Apc (Min/+) mouse | Potentiates the intestinal tumorigenesis | Recruitment of tumor-infiltrating myeloid cells | [35] | Association and interventional | Human and animal | |
| Chinese cohorts | Enriched in CRC tissues and associated with CRC development and metastasis | ND | [29] | Association | Human | |
| A large cohort of 616 participants | Elevated abundance from intramucosal carcinoma to more advanced stage | ND | [30] | Association | Human | |
| Cell line and humans | Stimulates growth of CRC cells | FadA binds to E-cadherin, activates β-catenin signaling, and differentially regulates the inflammatory and oncogenic responses | [31] | Interventional and association | In Vitro and human | |
| 598 rectal and colon carcinoma cases | Overrepresented in tumor samples | F. nucleatum is inversely associated with CD3+ T-cell density in colorectal carcinoma tissue | [32] | Association | Human | |
| A Mexican cohort | Enriched in CRC tissues | ND | [33] | Association | Human | |
| Enterotoxigenic Bacteroides fragilis | Colonic mucosa of patients with familial adenomatous polyposis | Associated with faster tumor onset and greater mortality | Increased interleukin-17 in the colon and DNA damage in colonic epithelium | [36] | Association | Human |
| Nude mice | Promoted colorectal carcinogenesis | Upregulated JMJD2B levels in a TLR4-NFAT5-dependent pathway | [37] | Interventional | Animal | |
| 40 CRC patients | KRAS mutations positively correlate with the abundance of this bacterium | Existence of this bacterium affected by miR3655/SURF6/IRF7/IFNβ axis | [38] | Association | Human | |
| ETBF-treated cells and patients | Promotes intestinal inflammation and malignancy | Inhibiting exosome-packaged miR-149-3p | [39] | Interventional and association | In vitro and human | |
| Mice | Induces colitis, colonic hyperplasia, and tumor formation | Activation of T helper type 17 T-cell responses | [40] | Interventional | Animal | |
| ApcMin mice | Induces onset of distal colon tumorigenesis | IL-17-dependent NF-κB activation, which relays CXCR2-expressing polymorphonuclear immature myeloid cells in a mucosal Th17 response | [41] | Interventional | Animal | |
| pks+ E. coli | 147 Caucasian cohort | Escherichia coli was enriched in carcinoma samples compared with both healthy and advanced adenoma sample | ND | [42] | Association | Human |
| 3741 stool metagenomes from 18 cohorts for cross-stage | Although not significant, the carriage of colibactin-producing genes by E. coli and Klebsiella spp. was increased in CRC; however, no correct time point for an impact of pks+ E coli was captured on CRC progression | ND | [43] | Association | Human | |
| Human intestinal organoids and two separate cohorts | Mutational signature in colorectal cancer was detected via colonization of pks+ E. coli | Presumptively, alkylate DNA on adenine residues induced double-strand breaks in cultured cells | [44] | Association | Human | |
| Streptococcus | Human epithelial colonic Caco-2 cells and rat colonic mucosa | S. bovis proteins showed procarcinogenic properties | Promoted release of CXC chemokines and prostaglandin E2, correlated with in vitro overexpression of COX-2 | [45] | Interventional | In vitro and animal |
| 50 colorectal cancer, 14 colorectal adenoma patients, and controls | Higher levels of serum S. gallolyticus IgG antibodies were associated with adenoma patients | NF-kappa B and IL-8 mRNAs more highly expressed in tumorous sections | [46] | Association | Human | |
| Cancer cell line and mouse models | Proliferation-promoting strains can promote carcinogenesis, while others cannot | ND | [47] | Interventional | In vitro and animal | |
| Peptostreptococcus anaerobius | Mice and patients with CRC in Hong Kong | Contributes to colon cancer formation | P. anaerobius interacted with TLR2 and TLR4 to increase intracellular levels of reactive oxidative species, promoting cholesterol synthesis and cell proliferation | [48] | Interventional and association | Animal and human |
| ApcMin/+ mice and cell line | Adhere to the CRC mucosa and accelerate CRC development | Via a PCWBR2-integrin α2/β1-PI3K-Akt-NF-κB signaling axis | [49] | Interventional | Animal and in vitro | |
| Salmonella | Animal model | Enhances colonic tumorigenesis | Salmonella protein AvrA upregulated transcriptional activity of STAT3 and its target genes | [50] | Interventional | Animal |
| 2D and organotypic 3D cultures | Promoting a microenvironment conducive to malignant transformation together with the loss of APC (adenomatous polyposis coli) | Reduced DNA repair capacity and inability to activate adequate checkpoint responses, as well as increased genomic instability | [51] | Interventional | In vitro | |
| Mouse models | Disrupts tumors | This therapy was accompanied by a compromised activation of tumor infiltrating lymphocytes | [52] | Interventional | Animal | |
| Fungi and virus | Cohort patients from Hong Kong | Abundance of 14 fungal biomarkers distinguished CRC from controls, characterized by a higher Basidiomycota–Ascomycota ratio | ND | [53] | Association | Human |
| 74 patients with CRC in Hong Kong | 22 viral taxa discriminate controls from patients | ND | [54] | Association | Human | |
| 12 CRC patients before and after surgery | Elevated viral correlations and network connectivity were observed in CRC, exhibiting cross-kingdom correlation | ND | [55] | Association | Human | |
| One patient with 313-day prolonged response and three non-responders | A rapid decrease in circulating DNA with virotherapy and immunotherapy | A progressive increase in CD4+ T cells, CD8+ T cells, and B cells, along with upregulated transcriptional factors for T-cell activation | [56] | Association | Human |
2.2. Enterotoxigenic Bacteroides fragilis and CRC
2.3. Enterococcus faecalis Strains Are Potential Biomarkers of CRC
2.4. pks+ E. coli Contributes to the Development of CRC
2.5. Streptococcus Is a Potential Contributor to CRC
2.6. Peptostreptococcus Anaerobius Is a Potential Contributor to CRC
2.7. Salmonella Has Relevance to CRC
2.8. Fungi and Viruses Are Emerging Contributors to CRC
2.9. Intratumoral Microbiota Is an Emerging Field to Impact CRC Pathogenesis
2.10. Microbe-Derived Metabolites Mediate the Microbiota-Related Effects on CRC
2.11. Inflammation Is Suggested to Underlie the Microbial Effects on CRC
3. Emerging Intervention Strategies for CRC
3.1. Antibiotics
3.2. FMT (Fecal Microbiota Transplantation)
3.3. Probiotics
 ) indicates suppression.
) indicates suppression.
 ) indicates suppression.
) indicates suppression.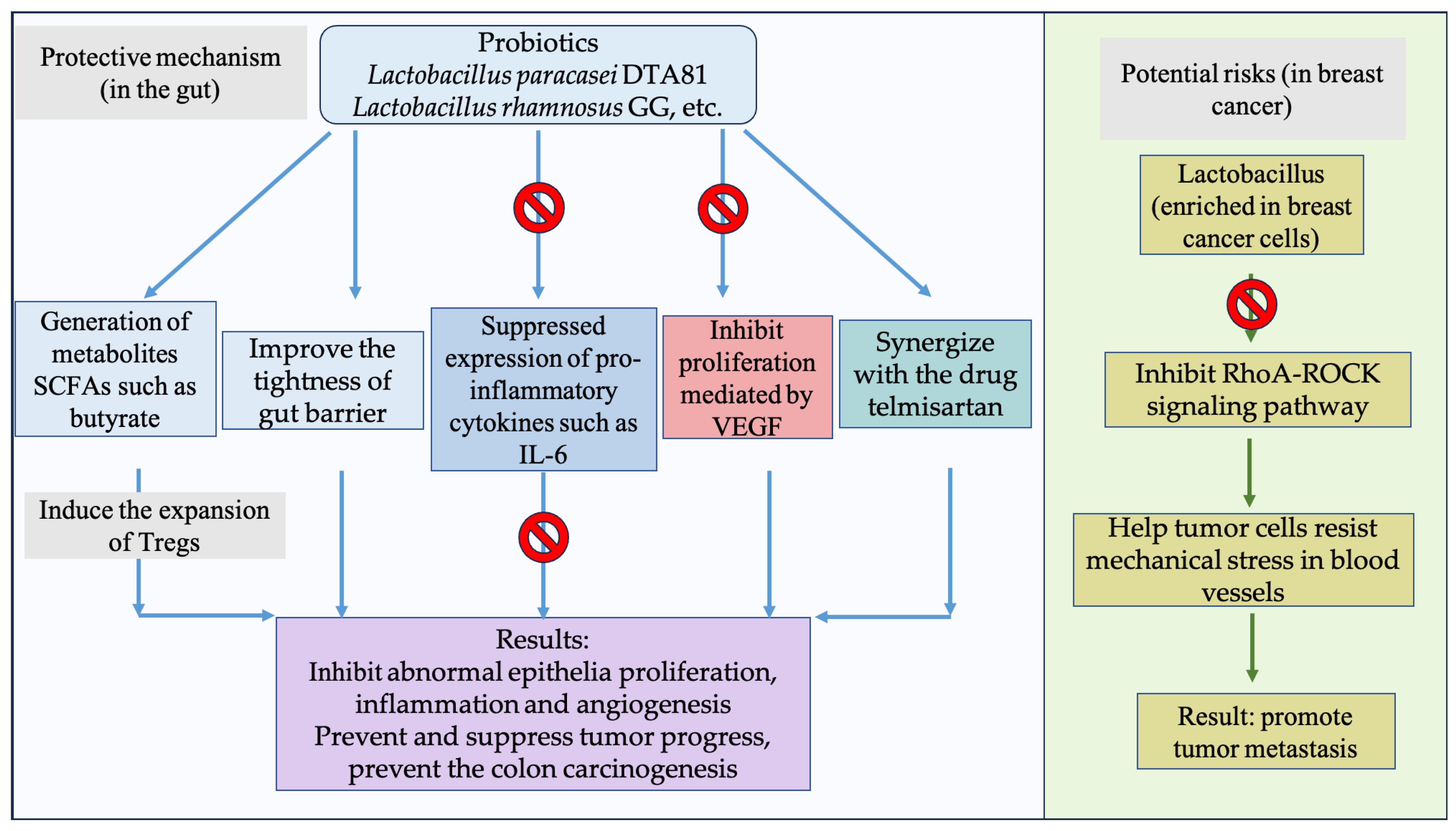
3.4. Dietary Intervention and CRC
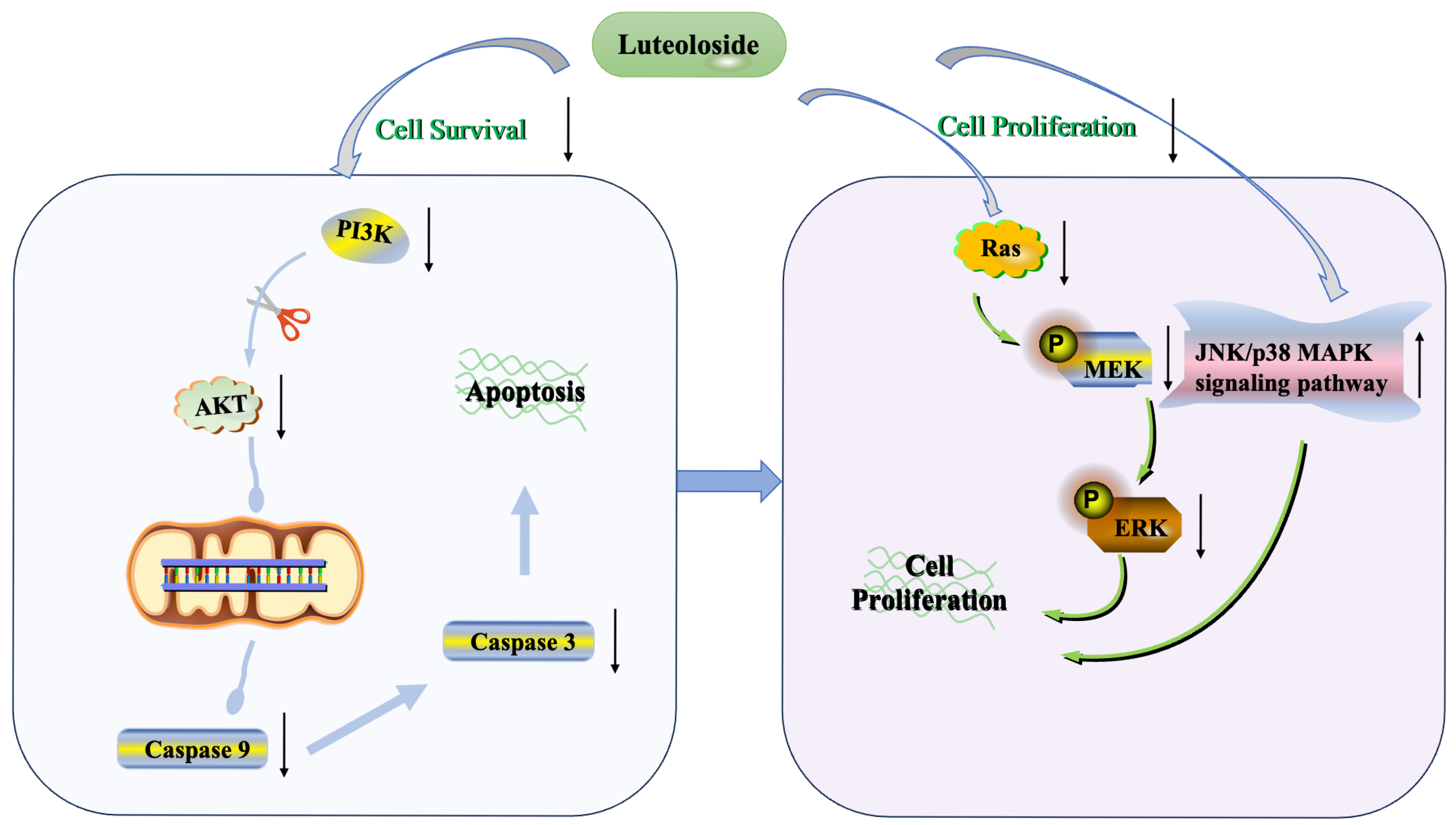
3.5. Natural Compounds and CRC
4. Future Perspectives
5. Methodology
- “gut microbiota” OR the concrete terms of specific bacterium.
- “CRC” OR “colorectal cancer.”
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Garrett, W.S. The gut microbiota and colon cancer. Science 2019, 364, 1133–1135. [Google Scholar] [CrossRef]
- Kim, J.; Lee, H.K. Potential Role of the Gut Microbiome in Colorectal Cancer Progression. Front. Immunol. 2021, 12, 807648. [Google Scholar] [CrossRef]
- Kamath, H.S.; Shukla, R.; Shah, U.; Patel, S.; Das, S.; Chordia, A.; Satish, P.; Ghosh, D. Role of Gut Microbiota in Predisposition to Colon Cancer: A Narrative Review. Indian J. Microbiol. 2024, 64, 1–13. [Google Scholar] [CrossRef]
- Cuisiniere, T.; Hajjar, R.; Oliero, M.; Calve, A.; Fragoso, G.; Rendos, H.V.; Gerkins, C.; Taleb, N.; Gagnon-Konamna, M.; Dagbert, F.; et al. Initial gut microbiota composition is a determining factor in the promotion of colorectal cancer by oral iron supplementation: Evidence from a murine model. Microbiome 2025, 13, 100. [Google Scholar] [CrossRef] [PubMed]
- Erfanian, N.; Tavakoli, T.; Mahdiabadi, M.A.; Nasseri, S.; Safarpour, H.; Fakharian, T.; Namaei, M.H. Gut microbiota dysbiosis and the anti-inflammatory effects of probiotic-derived cell-free supernatants in HT-29 cells: Insights into early stage colorectal cancer. Lett. Appl. Microbiol. 2025, 78, ovaf060. [Google Scholar] [CrossRef] [PubMed]
- Darnindro, N.; Abdullah, M.; Sukartini, N.; Rumende, C.M.; Pitarini, A.; Nursyirwan, S.A.; Fauzi, A.; Makmun, D.; Nelwan, E.J.; Shatri, H.; et al. Differences in diversity and composition of mucosa-associated colonic microbiota in colorectal cancer and non-colorectal cancer in Indonesia. World J. Gastroenterol. 2025, 31, 100051. [Google Scholar] [CrossRef] [PubMed]
- Akimoto, N.; Ugai, T.; Zhong, R.; Hamada, T.; Fujiyoshi, K.; Giannakis, M.; Wu, K.; Cao, Y.; Ng, K.; Ogino, S. Rising incidence of early-onset colorectal cancer—A call to action. Nat. Rev. Clin. Oncol. 2021, 18, 230–243. [Google Scholar] [CrossRef]
- Siegel, R.L.; Torre, L.A.; Soerjomataram, I.; Hayes, R.B.; Bray, F.; Weber, T.K.; Jemal, A. Global patterns and trends in colorectal cancer incidence in young adults. Gut 2019, 68, 2179–2185. [Google Scholar] [CrossRef]
- Vuik, F.E.; Nieuwenburg, S.A.; Bardou, M.; Lansdorp-Vogelaar, I.; Dinis-Ribeiro, M.; Bento, M.J.; Zadnik, V.; Pellise, M.; Esteban, L.; Kaminski, M.F.; et al. Increasing incidence of colorectal cancer in young adults in Europe over the last 25 years. Gut 2019, 68, 1820–1826. [Google Scholar] [CrossRef]
- Wong, S.H.; Yu, J. Gut microbiota in colorectal cancer: Mechanisms of action and clinical applications. Nat. Rev. Gastroenterol. Hepatol. 2019, 16, 690–704. [Google Scholar] [CrossRef]
- Lichtenstein, P.; Holm, N.V.; Verkasalo, P.K.; Iliadou, A.; Kaprio, J.; Koskenvuo, M.; Pukkala, E.; Skytthe, A.; Hemminki, K. Environmental and heritable factors in the causation of cancer--analyses of cohorts of twins from Sweden, Denmark, and Finland. N. Engl. J. Med. 2000, 343, 78–85. [Google Scholar] [CrossRef]
- Mucci, L.A.; Hjelmborg, J.B.; Harris, J.R.; Czene, K.; Havelick, D.J.; Scheike, T.; Graff, R.E.; Holst, K.; Möller, S.; Unger, R.H.; et al. Familial Risk and Heritability of Cancer Among Twins in Nordic Countries. JAMA 2016, 315, 68–76. [Google Scholar] [CrossRef] [PubMed]
- Wu, G.D.; Chen, J.; Hoffmann, C.; Bittinger, K.; Chen, Y.-Y.; Keilbaugh, S.A.; Bewtra, M.; Knights, D.; Walters, W.A.; Knight, R.; et al. Linking long-term dietary patterns with gut microbial enterotypes. Science 2011, 334, 105–108. [Google Scholar] [CrossRef] [PubMed]
- Mehta, R.S.; Song, M.; Nishihara, R.; Drew, D.A.; Wu, K.; Qian, Z.R.; Fung, T.T.; Hamada, T.; Masugi, Y.; da Silva, A.; et al. Dietary Patterns and Risk of Colorectal Cancer: Analysis by Tumor Location and Molecular Subtypes. Gastroenterology 2017, 152, 1944–1953. [Google Scholar] [CrossRef]
- Qu, R.; Zhang, Y.; Ma, Y.; Zhou, X.; Sun, L.; Jiang, C.; Zhang, Z.; Fu, W. Role of the Gut Microbiota and Its Metabolites in Tumorigenesis or Development of Colorectal Cancer. Adv. Sci. 2023, 10, e2205563. [Google Scholar] [CrossRef]
- Gacesa, R.; Kurilshikov, A.; Vich Vila, A.; Sinha, T.; Klaassen, M.A.Y.; Bolte, L.A.; Andreu-Sanchez, S.; Chen, L.; Collij, V.; Hu, S.; et al. Environmental factors shaping the gut microbiome in a Dutch population. Nature 2022, 604, 732–739. [Google Scholar] [CrossRef]
- Xiao, J.; Wang, T.; Xu, Y.; Gu, X.; Li, D.; Niu, K.; Wang, T.; Zhao, J.; Zhou, R.; Wang, H.L. Long-term probiotic intervention mitigates memory dysfunction through a novel H3K27me3-based mechanism in lead-exposed rats. Transl. Psychiatry 2020, 10, 25. [Google Scholar] [CrossRef]
- Robinson, W.; Gertz, E.M.; Greten, T.F.; Ruppin, E. Medication effects on the gut microbiome in allo-HCT. Cell 2023, 186, 2520–2523. [Google Scholar] [CrossRef]
- Yang, R.; Xu, Y.; Zhu, F.; Ma, X.; Fan, T.; Wang, H.-L. Gut microbiome, a potential modulator of neuroepigenome. J. Nutr. Biochem. 2025, 144, 109961. [Google Scholar] [CrossRef]
- Gomaa, E.Z. Human gut microbiota/microbiome in health and diseases: A review. Antonie Van Leeuwenhoek 2020, 113, 2019–2040. [Google Scholar] [CrossRef]
- Li, J.; Hou, W.; Lin, S.; Wang, L.; Pan, C.; Wu, F.; Liu, J. Polydopamine nanoparticle-mediated dopaminergic immunoregulation in colitis. Adv. Sci. 2022, 9, 2104006. [Google Scholar] [CrossRef]
- Wang, Z.; Dan, W.; Zhang, N.; Fang, J.; Yang, Y. Colorectal cancer and gut microbiota studies in China. Gut Microbes 2023, 15, 2236364. [Google Scholar] [CrossRef]
- Tsenkova, M.; Brauer, M.; Pozdeev, V.I.; Kasakin, M.; Busi, S.B.; Schmoetten, M.; Cheung, D.; Meyers, M.; Rodriguez, F.; Gaigneaux, A.; et al. Ketogenic diet suppresses colorectal cancer through the gut microbiome long chain fatty acid stearate. Nat. Commun. 2025, 16, 1792. [Google Scholar] [CrossRef]
- Qin, Y.; Tong, X.; Mei, W.J.; Cheng, Y.; Zou, Y.; Han, K.; Yu, J.; Jie, Z.; Zhang, T.; Zhu, S.; et al. Consistent signatures in the human gut microbiome of old- and young-onset colorectal cancer. Nat. Commun. 2024, 15, 3396. [Google Scholar] [CrossRef]
- Qin, Y.; Wang, Q.; Lin, Q.; Liu, F.; Pan, X.; Wei, C.; Chen, J.; Huang, T.; Fang, M.; Yang, W.; et al. Multi-omics analysis reveals associations between gut microbiota and host transcriptome in colon cancer patients. mSystems 2025, 10, e0080524. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Zhang, X.; Hang, D.; Lau, H.C.-H.; Du, J.; Liu, C.; Xie, M.; Pan, Y.; Wang, L.; Liang, C.; et al. Integrative plasma and fecal metabolomics identify functional metabolites in adenoma-colorectal cancer progression and as early diagnostic biomarkers. Cancer Cell 2024, 42, 1386–1400. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.W. Fusobacterium nucleatum: A commensal-turned pathogen. Curr. Opin. Microbiol. 2015, 23, 141–147. [Google Scholar] [CrossRef] [PubMed]
- Castellarin, M.; Warren, R.L.; Freeman, J.D.; Dreolini, L.; Krzywinski, M.; Strauss, J.; Barnes, R.; Watson, P.; Allen-Vercoe, E.; Moore, R.A.; et al. Fusobacterium nucleatum infection is prevalent in human colorectal carcinoma. Genome Res. 2012, 22, 299–306. [Google Scholar] [CrossRef]
- Li, Y.Y.; Ge, Q.X.; Cao, J.; Zhou, Y.J.; Du, Y.L.; Shen, B.; Wan, Y.J.Y.; Nie, Y.Q. Association of Fusobacterium nucleatum infection with colorectal cancer in Chinese patients. World J. Gastroenterol. 2016, 22, 3227. [Google Scholar] [CrossRef]
- Yachida, S.; Mizutani, S.; Shiroma, H.; Shiba, S.; Nakajima, T.; Sakamoto, T.; Watanabe, H.; Masuda, K.; Nishimoto, Y.; Kubo, M.; et al. Metagenomic and metabolomic analyses reveal distinct stage-specific phenotypes of the gut microbiota in colorectal cancer. Nat. Med. 2019, 25, 968–976. [Google Scholar] [CrossRef]
- Rubinstein, M.R.; Wang, X.; Liu, W.; Hao, Y.; Cai, G.; Han, Y.W. Fusobacterium nucleatum promotes colorectal carcinogenesis by modulating E-cadherin/β-catenin signaling via its FadA adhesin. Cell Host Microbe 2013, 14, 195–206. [Google Scholar] [CrossRef]
- Mima, K.; Sukawa, Y.; Nishihara, R.; Qian, Z.R.; Yamauchi, M.; Inamura, K.; Kim, S.A.; Masuda, A.; Nowak, J.A.; Nosho, K.; et al. Fusobacterium nucleatum and T cells in colorectal carcinoma. JAMA Oncol. 2015, 1, 653–661. [Google Scholar] [CrossRef]
- Cuellar-Gómez, H.; Ocharán-Hernández, M.E.; Calzada-Mendoza, C.C.; Comoto-Santacruz, D.A. Association of Fusobacterium nucleatum infection and colorectal cancer: A Mexican study. Rev. Gastroenterol. Méx. (Engl. Ed.) 2022, 87, 277–284. [Google Scholar] [CrossRef]
- Brennan, C.A.; Garrett, W.S. Fusobacterium nucleatum—Symbiont, opportunist and oncobacterium. Microbiology 2019, 17, 156–166. [Google Scholar] [CrossRef] [PubMed]
- Kostic, A.D.; Chun, E.; Robertson, L.; Glickman, J.N.; Gallini, C.A.; Michaud, M.; Clancy, T.E.; Chung, D.C.; Lochhead, P.; Hold, G.L.; et al. Fusobacterium nucleatum potentiates intestinal tumorigenesis and modulates the tumor-immune microenvironment. Cell Host Microbe 2013, 14, 207–215. [Google Scholar] [CrossRef] [PubMed]
- Dejea, C.M.; Fathi, P.; Craig, J.M.; Boleij, A.; Taddese, R.; Geis, A.L.; Wu, X.; DeStefano Shields, C.E.; Hechenbleikner, E.M.; Huso, D.L.; et al. Patients with familial adenomatous polyposis harbor colonic biofilms containing tumorigenic bacteria. Science 2018, 359, 592–597. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.-Q.; Li, C.-M.; Fu, L.-N.; Wang, H.-L.; Tan, J.; Wang, Y.-Q.; Sun, D.-F.; Gao, Q.-Y.; Chen, Y.-X.; Fang, J.-Y. Enterotoxigenic Bacteroides fragilis induces the stemness in colorectal cancer via upregulating histone demethylase JMJD2B. Gut Microbes 2020, 12, 1788900. [Google Scholar] [CrossRef]
- Chen, Y.; Liu, S.; Tan, S.; Zheng, Y.; Chen, Y.; Yang, C.; Lin, S.; Mi, Y.; Li, W. KRAS mutations promote the intratumoral colonization of enterotoxigenic bacteroides fragilis in colorectal cancer through the regulation of the miRNA3655/SURF6/IRF7/IFNβ axis. Gut Microbes 2024, 16, 2423043. [Google Scholar] [CrossRef]
- Cao, Y.; Wang, Z.; Yan, Y.; Ji, L.; He, J.; Xuan, B.; Shen, C.; Ma, Y.; Jiang, S.; Ma, D.; et al. Enterotoxigenic Bacteroidesfragilis Promotes Intestinal Inflammation and Malignancy by Inhibiting Exosome-Packaged miR-149-3p. Gastroenterology 2021, 161, 1552–1566. [Google Scholar] [CrossRef]
- Wu, S.; Rhee, K.-J.; Albesiano, E.; Rabizadeh, S.; Wu, X.; Yen, H.-R.; Huso, D.L.; Brancati, F.L.; Wick, E.; McAllister, F.; et al. A human colonic commensal promotes colon tumorigenesis via activation of T helper type 17 T cell responses. Nat. Med. 2009, 15, 1016–1022. [Google Scholar] [CrossRef]
- Chung, L.; Orberg, E.T.; Geis, A.L.; Chan, J.L.; Fu, K.; Shields, C.E.D.; Dejea, C.M.; Fathi, P.; Chen, J.; Finard, B.B.; et al. Bacteroides fragilis toxin coordinates a pro-carcinogenic inflammatory cascade via targeting of colonic epithelial cells. Cell Host Microbe 2018, 23, 203–214.e5. [Google Scholar] [CrossRef]
- Feng, Q.; Liang, S.; Jia, H.; Stadlmayr, A.; Tang, L.; Lan, Z.; Zhang, D.; Xia, H.; Xu, X.; Jie, Z.; et al. Gut microbiome development along the colorectal adenoma–carcinoma sequence. Nat. Commun. 2015, 6, 6528. [Google Scholar] [CrossRef] [PubMed]
- Piccinno, G.; Thompson, K.N.; Manghi, P.; Ghazi, A.R.; Thomas, A.M.; Blanco-Miguez, A.; Asnicar, F.; Mladenovic, K.; Pinto, F.; Armanini, F.; et al. Pooled analysis of 3,741 stool metagenomes from 18 cohorts for cross-stage and strain-level reproducible microbial biomarkers of colorectal cancer. Nat. Med. 2025, 31, 2416–2429. [Google Scholar] [CrossRef] [PubMed]
- Pleguezuelos-Manzano, C.; Puschhof, J.; Rosendahl Huber, A.; van Hoeck, A.; Wood, H.M.; Nomburg, J.; Gurjao, C.; Manders, F.; Dalmasso, G.; Stege, P.B.; et al. Mutational signature in colorectal cancer caused by genotoxic pks+ E. coli. Nature 2020, 580, 269–273. [Google Scholar] [CrossRef] [PubMed]
- Biarc, J.; Nguyen, I.S.; Pini, A.; Gosse, F.; Richert, S.; Thierse, D.; Van Dorsselaer, A.; Leize-Wagner, E.; Raul, F.; Klein, J.-P.; et al. Carcinogenic properties of proteins with pro-inflammatory activity from Streptococcus infantarius (formerly S. bovis). Carcinogenesis 2004, 25, 1477–1484. [Google Scholar] [CrossRef]
- Abdulamir, A.S.; Hafidh, R.R.; Mahdi, L.K.; Al-jeboori, T.; Abubaker, F. Investigation into the controversial association of Streptococcus gallolyticus with colorectal cancer and adenoma. BMC Cancer 2009, 9, 403. [Google Scholar] [CrossRef]
- Kumar, R.; Herold, J.L.; Taylor, J.; Xu, J.; Xu, Y. Variations among Streptococcus gallolyticus subsp. gallolyticus strains in connection with colorectal cancer. Sci. Rep. 2018, 8, 1514. [Google Scholar] [CrossRef]
- Tsoi, H.; Chu, E.S.; Zhang, X.; Sheng, J.; Nakatsu, G.; Ng, S.C.; Chan, A.W.; Chan, F.K.; Sung, J.J.; Yu, J. Peptostreptococcus anaerobius induces intracellular cholesterol biosynthesis in colon cells to induce proliferation and causes dysplasia in mice. Gastroenterology 2017, 152, 1419–1433.e5. [Google Scholar] [CrossRef]
- Long, X.; Wong, C.C.; Tong, L.; Chu, E.S.; Ho Szeto, C.; Go, M.Y.; Coker, O.O.; Chan, A.W.; Chan, F.K.; Sung, J.J.; et al. Peptostreptococcus anaerobius promotes colorectal carcinogenesis and modulates tumour immunity. Nat. Microbiol. 2019, 4, 2319–2330. [Google Scholar] [CrossRef]
- Lu, R.; Wu, S.; Zhang, Y.-g.; Xia, Y.; Zhou, Z.; Kato, I.; Dong, H.; Bissonnette, M.; Sun, J. Salmonella protein AvrA activates the STAT3 signaling pathway in colon cancer. Neoplasia 2016, 18, 307–316. [Google Scholar] [CrossRef]
- Martin, O.C.; Bergonzini, A.; d’Amico, F.; Chen, P.; Shay, J.W.; Dupuy, J.; Svensson, M.; Masucci, M.G.; Frisan, T. Infection with genotoxin-producing Salmonella enterica synergises with loss of the tumour suppressor APC in promoting genomic instability via the PI3K pathway in colonic epithelial cells. Cell. Microbiol. 2019, 21, e13099. [Google Scholar] [CrossRef]
- Copland, A.; Mackie, G.M.; Scarfe, L.; Jinks, E.; Lecky, D.A.J.; Gudgeon, N.; McQuade, R.; Ono, M.; Barthel, M.; Hardt, W.-D.; et al. Salmonella cancer therapy metabolically disrupts tumours at the collateral cost of T cell immunity. EMBO Mol. Med. 2024, 16, 3057–3088. [Google Scholar] [CrossRef]
- Coker, O.O.; Nakatsu, G.; Dai, R.Z.; Wu, W.K.K.; Wong, S.H.; Ng, S.C.; Chan, F.K.L.; Sung, J.J.Y.; Yu, J. Enteric fungal microbiota dysbiosis and ecological alterations in colorectal cancer. Gut 2019, 68, 654–662. [Google Scholar] [CrossRef]
- Nakatsu, G.; Zhou, H.; Wu, W.K.K.; Wong, S.H.; Coker, O.O.; Dai, Z.; Li, X.; Szeto, C.-H.; Sugimura, N.; Lam, T.Y.-T.; et al. Alterations in enteric virome are associated with colorectal cancer and survival outcomes. Gastroenterology 2018, 155, 529–541.e5. [Google Scholar] [CrossRef] [PubMed]
- Ho, S.X.; Law, J.-H.; Png, C.-W.; Alberts, R.; Zhang, Y.; Chu, J.J.H.; Tan, K.-K. Alterations in colorectal cancer virome and its persistence after surgery. Sci. Rep. 2024, 14, 2819. [Google Scholar] [CrossRef]
- Zhang, H.; Ren, Y.; Wang, F.; Tu, X.; Tong, Z.; Liu, L.; Zheng, Y.; Zhao, P.; Cheng, J.; Li, J.; et al. The long-term effectiveness and mechanism of oncolytic virotherapy combined with anti-PD-L1 antibody in colorectal cancer patient. Cancer Gene Ther. 2024, 31, 1412–1426. [Google Scholar] [CrossRef] [PubMed]
- Miyazaki, K.; Sakuma, K.; Kawamura, Y.I.; Izawa, M.; Ohmori, K.; Mitsuki, M.; Yamaji, T.; Hashimoto, Y.; Suzuki, A.; Saito, Y.; et al. Colonic epithelial cells express specific ligands for mucosal macrophage immunosuppressive receptors siglec-7 and-9. The J. Immunol. 2012, 188, 4690–4700. [Google Scholar] [CrossRef] [PubMed]
- Xing, J.; Fang, Y.; Zhang, W.; Zhang, H.; Tang, D.; Wang, D. Bacterial driver–passenger model in biofilms: A new mechanism in the development of colorectal cancer. Clin. Transl. Oncol. 2022, 24, 784–795. [Google Scholar] [CrossRef]
- Kwong, T.N.; Wang, X.; Nakatsu, G.; Chow, T.C.; Tipoe, T.; Dai, R.Z.; Tsoi, K.K.; Wong, M.C.; Tse, G.; Chan, M.T.; et al. Association between bacteremia from specific microbes and subsequent diagnosis of colorectal cancer. Gastroenterology 2018, 155, 383–390.e8. [Google Scholar] [CrossRef]
- Gong, D.; Adomako-Bonsu, A.G.; Wang, M.; Li, J. Three specific gut bacteria in the occurrence and development of colorectal cancer: A concerted effort. PeerJ 2023, 11, e15777. [Google Scholar] [CrossRef]
- Ternes, D.; Karta, J.; Tsenkova, M.; Wilmes, P.; Haan, S.; Letellier, E. Microbiome in Colorectal Cancer: How to Get from Meta-omics to Mechanism? Trends Microbiol. 2020, 28, 401–423. [Google Scholar] [CrossRef]
- Daca, A.; Jarzembowski, T. From the friend to the foe—Enterococcus faecalis diverse impact on the human immune system. Int. J. Mol. Sci. 2024, 25, 2422. [Google Scholar] [CrossRef] [PubMed]
- Winters, M.D.; Schlinke, T.L.; Joyce, W.A.; Glore, S.R.; Huycke, M.M. Prospective case-cohort study of intestinal colonization with enterococci that produce extracellular superoxide and the risk for colorectal adenomas or cancer. Off. J. Am. Coll. Gastroenterol. ACG 1998, 93, 2491–2500. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Allen, T.D.; May, R.J.; Lightfoot, S.; Houchen, C.W.; Huycke, M.M. Enterococcus faecalis induces aneuploidy and tetraploidy in colonic epithelial cells through a bystander effect. Cancer Res. 2008, 68, 9909–9917. [Google Scholar] [CrossRef] [PubMed]
- Evans, M.D.; Dizdaroglu, M.; Cooke, M.S. Oxidative DNA damage and disease: Induction, repair and significance. Mutat. Res./Rev. Mutat. Res. 2004, 567, 1–61. [Google Scholar] [CrossRef]
- Zhang, L.; Liu, J.; Deng, M.; Chen, X.; Jiang, L.; Zhang, J.; Tao, L.; Yu, W.; Qiu, Y. Enterococcus faecalis promotes the progression of colorectal cancer via its metabolite: Biliverdin. J. Transl. Med. 2023, 21, 72. [Google Scholar] [CrossRef]
- de Almeida, C.V.; Taddei, A.; Amedei, A. The controversial role of Enterococcus faecalis in colorectal cancer. Ther. Adv. Gastroenterol. 2018, 11, 1756284818783606. [Google Scholar] [CrossRef]
- Wilson, M.R.; Jiang, Y.; Villalta, P.W.; Stornetta, A.; Boudreau, P.D.; Carrá, A.; Brennan, C.A.; Chun, E.; Ngo, L.; Samson, L.D.; et al. The human gut bacterial genotoxin colibactin alkylates DNA. Science 2019, 363, eaar7785. [Google Scholar] [CrossRef]
- Zhang, H.; Jin, K.; Xiong, K.; Jing, W.; Pang, Z.; Feng, M.; Cheng, X. Disease-associated gut microbiome and critical metabolomic alterations in patients with colorectal cancer. Cancer Med. 2023, 12, 15720–15735. [Google Scholar] [CrossRef]
- Gupta, A.; Madani, R.; Mukhtar, H. Streptococcus bovis endocarditis, a silent sign for colonic tumour. Color. Dis. 2010, 12, 164–171. [Google Scholar] [CrossRef]
- Boleij, A.; van Gelder, M.M.; Swinkels, D.W.; Tjalsma, H. Clinical Importance of Streptococcus gallolyticus infection among colorectal cancer patients: Systematic review and meta-analysis. Clin. Infect. Dis. 2011, 53, 870–878. [Google Scholar] [CrossRef]
- Liu, M.; Wei, F.; Wang, J.; Yu, W.; Shen, M.; Liu, T.; Zhang, D.; Wang, Y.; Ren, X.; Sun, Q. Myeloid-derived suppressor cells regulate the immunosuppressive functions of PD-1− PD-L1+ Bregs through PD-L1/PI3K/AKT/NF-κB axis in breast cancer. Cell Death Dis. 2021, 12, 465. [Google Scholar] [CrossRef]
- Yu, L.; Zhao, G.; Wang, L.; Zhou, X.; Sun, J.; Li, X.; Zhu, Y.; He, Y.; Kofonikolas, K.; Bogaert, D.; et al. A systematic review of microbial markers for risk prediction of colorectal neoplasia. Br. J. Cancer 2022, 126, 1318–1328. [Google Scholar] [CrossRef] [PubMed]
- Starnes, C.O. Coley’s toxins in perspective. Nature 1992, 357, 11–12. [Google Scholar] [CrossRef] [PubMed]
- Cani, P.D. Human gut microbiome: Hopes, threats and promises. Gut 2018, 67, 1716–1725. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Wang, Y.; Huang, X.; Yu, X. Unveiling the overlooked fungi: The vital of gut fungi in inflammatory bowel disease and colorectal cancer. Gut Pathog. 2024, 16, 59. [Google Scholar] [CrossRef]
- Hannigan, G.D.; Duhaime, M.B.; Ruffin IV, M.T.; Koumpouras, C.C.; Schloss, P.D. Diagnostic potential and interactive dynamics of the colorectal cancer virome. mBio 2018, 9, 10–1128. [Google Scholar] [CrossRef]
- Yang, L.; Li, A.; Wang, Y.; Zhang, Y. Intratumoral microbiota: Roles in cancer initiation, development and therapeutic efficacy. Signal Transduct. Target. Ther. 2023, 8, 35. [Google Scholar] [CrossRef]
- Lin, D.; Shen, Y.; Liang, T. Oncolytic virotherapy: Basic principles, recent advances and future directions. Signal Transduct. Target. Ther. 2023, 8, 156. [Google Scholar] [CrossRef]
- Cao, Y.; Xia, H.; Tan, X.; Shi, C.; Ma, Y.; Meng, D.; Zhou, M.; Lv, Z.; Wang, S.; Jin, Y.J.S.T.; et al. Intratumoural microbiota: A new frontier in cancer development and therapy. Signal Transduct. Target. Ther. 2024, 9, 15. [Google Scholar] [CrossRef]
- Fu, A.; Yao, B.; Dong, T.; Chen, Y.; Yao, J.; Liu, Y.; Li, H.; Bai, H.; Liu, X.; Zhang, Y.; et al. Tumor-resident intracellular microbiota promotes metastatic colonization in breast cancer. Cell 2022, 185, 1356–1372.e26. [Google Scholar] [CrossRef] [PubMed]
- Galeano Niño, J.L.; Wu, H.; LaCourse, K.D.; Kempchinsky, A.G.; Baryiames, A.; Barber, B.; Futran, N.; Houlton, J.; Sather, C.; Sicinska, E.; et al. Effect of the intratumoral microbiota on spatial and cellular heterogeneity in cancer. Nature 2022, 611, 810–817. [Google Scholar] [CrossRef] [PubMed]
- Riquelme, E.; Zhang, Y.; Zhang, L.; Montiel, M.; Zoltan, M.; Dong, W.; Quesada, P.; Sahin, I.; Chandra, V.; San Lucas, A.; et al. Tumor microbiome diversity and composition influence pancreatic cancer outcomes. Cell 2019, 178, 795–806.e12. [Google Scholar] [CrossRef]
- Narunsky-Haziza, L.; Sepich-Poore, G.D.; Livyatan, I.; Asraf, O.; Martino, C.; Nejman, D.; Gavert, N.; Stajich, J.E.; Amit, G.; González, A.; et al. Pan-cancer analyses reveal cancer-type-specific fungal ecologies and bacteriome interactions. Cell 2022, 185, 3789–3806.e17. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Veledo, S.; Vendrell, J. Gut microbiota-derived succinate: Friend or foe in human metabolic diseases? Rev. Endocr. Metab. Disord. 2019, 20, 439–447. [Google Scholar] [CrossRef]
- Fu, T.; Huan, T.; Rahman, G.; Zhi, H.; Xu, Z.; Oh, T.G.; Guo, J.; Coulter, S.; Tripathi, A.; Martino, C.; et al. Paired microbiome and metabolome analyses associate bile acid changes with colorectal cancer progression. Cell Rep. 2023, 42, 112997. [Google Scholar] [CrossRef]
- Yoshimoto, S.; Loo, T.M.; Atarashi, K.; Kanda, H.; Sato, S.; Oyadomari, S.; Iwakura, Y.; Oshima, K.; Morita, H.; Hattori, M.; et al. Obesity-induced gut microbial metabolite promotes liver cancer through senescence secretome. Nature 2013, 499, 97–101. [Google Scholar] [CrossRef]
- Ma, R.; Sun, E.D.; Zou, J. A spectral method for assessing and combining multiple data visualizations. Nat. Commun. 2023, 14, 780. [Google Scholar] [CrossRef]
- Stidham, R.W.; Higgins, P.D.R. Colorectal Cancer in Inflammatory Bowel Disease. Clin. Colon Rectal Surg. 2018, 31, 168–178. [Google Scholar] [CrossRef]
- van Staa, T.P.; Card, T.; Logan, R.; Leufkens, H.G.M. 5-Aminosalicylate use and colorectal cancer risk in inflammatory bowel disease: A large epidemiological study. Gut 2005, 54, 1573–1578. [Google Scholar] [CrossRef]
- Wang, T.; Xu, J.; Xu, Y.; Xiao, J.; Bi, N.; Gu, X.; Wang, H.-L. Gut microbiota shapes social dominance through modulating HDAC2 in the medial prefrontal cortex. Cell Rep. 2022, 38, 110478. [Google Scholar] [CrossRef]
- Quaglio, A.E.V.; Grillo, T.G.; De Oliveira, E.C.S.; Di Stasi, L.C.; Sassaki, L.Y. Gut microbiota, inflammatory bowel disease and colorectal cancer. World J. Gastroenterol. 2022, 28, 4053. [Google Scholar] [CrossRef]
- Jochems, C.; Schlom, J. Tumor-infiltrating immune cells and prognosis: The potential link between conventional cancer therapy and immunity. Exp. Biol. Med. 2011, 236, 567–579. [Google Scholar] [CrossRef]
- Wang, Z.-T.; Deng, Z.-M.; Dai, F.-F.; Yuan, M.-Q.; Liu, S.-Y.; Li, B.-S.; Cheng, Y.-X. Tumor immunity: A brief overview of tumor-infiltrating immune cells and research advances into tumor-infiltrating lymphocytes in gynecological malignancies. Exp. Ther. Med. 2024, 27, 166. [Google Scholar] [CrossRef]
- Kikuchi, T.; Mimura, K.; Ashizawa, M.; Okayama, H.; Endo, E.; Saito, K.; Sakamoto, W.; Fujita, S.; Endo, H.; Saito, M.; et al. Immunotherapy, Characterization of tumor-infiltrating immune cells in relation to microbiota in colorectal cancers. Cancer Immunol. Immunother. 2020, 69, 23–32. [Google Scholar] [CrossRef]
- Cao, M.; Deng, Y.; Hao, Q.; Yan, H.; Wang, Q.-L.; Dong, C.; Wu, J.; He, Y.; Huang, L.-B.; Xia, X.; et al. Single-cell transcriptomic analysis reveals gut microbiota-immunotherapy synergy through modulating tumor microenvironment. Signal Transduct. Target. Ther. 2025, 10, 140. [Google Scholar] [CrossRef]
- Bullman, S.; Pedamallu, C.S.; Sicinska, E.; Clancy, T.E.; Zhang, X.; Cai, D.; Neuberg, D.; Huang, K.; Guevara, F.; Nelson, T.; et al. Analysis of Fusobacterium persistence and antibiotic response in colorectal cancer. Science 2017, 358, 1443–1448. [Google Scholar] [CrossRef] [PubMed]
- Simin, J.; Fornes, R.; Liu, Q.; Olsen, R.S.; Callens, S.; Engstrand, L.; Brusselaers, N. Antibiotic use and risk of colorectal cancer: A systematic review and dose–response meta-analysis. Br. J. Cancer 2020, 123, 1825–1832. [Google Scholar] [CrossRef] [PubMed]
- Bakken, J.S.; Borody, T.; Brandt, L.J.; Brill, J.V.; Demarco, D.C.; Franzos, M.A.; Kelly, C.; Khoruts, A.; Louie, T.; Martinelli, L.P.; et al. Treating Clostridium difficile infection with fecal microbiota transplantation. Clin. Gastroenterol. Hepatol. Off. Clin. Pract. J. Am. Gastroenterol. Assoc. 2011, 9, 1044–1049. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wiesnoski, D.H.; Helmink, B.A.; Gopalakrishnan, V.; Choi, K.; DuPont, H.L.; Jiang, Z.-D.; Abu-Sbeih, H.; Sanchez, C.A.; Chang, C.-C.; et al. Fecal microbiota transplantation for refractory immune checkpoint inhibitor-associated colitis. Nat. Med. 2018, 24, 1804–1808. [Google Scholar] [CrossRef]
- Davar, D.; Dzutsev, A.K.; McCulloch, J.A.; Rodrigues, R.R.; Chauvin, J.-M.; Morrison, R.M.; Deblasio, R.N.; Menna, C.; Ding, Q.; Pagliano, O.; et al. Fecal microbiota transplant overcomes resistance to anti–PD-1 therapy in melanoma patients. Science 2021, 371, 595–602. [Google Scholar] [CrossRef]
- Baruch, E.N.; Youngster, I.; Ben-Betzalel, G.; Ortenberg, R.; Lahat, A.; Katz, L.; Adler, K.; Dick-Necula, D.; Raskin, S.; Bloch, N.; et al. Fecal microbiota transplant promotes response in immunotherapy-refractory melanoma patients. Science 2021, 371, 602–609. [Google Scholar] [CrossRef]
- da Silva Duarte, V.; dos Santos Cruz, B.C.; Tarrah, A.; Sousa Dias, R.; de Paula Dias Moreira, L.; Lemos Junior, W.J.F.; Fidelis Silva, L.C.; Rocha Santana, G.; Licursi de Oliveira, L.; Gouveia Peluzio, M.D.C.; et al. Chemoprevention of DMH-induced early colon carcinogenesis in male BALB/c mice by administration of lactobacillus paracasei DTA81. Microorganisms 2020, 8, 1994. [Google Scholar] [CrossRef] [PubMed]
- Rafter, J.; Bennett, M.; Caderni, G.; Clune, Y.; Hughes, R.; Karlsson, P.C.; Klinder, A.; O’Riordan, M.; O’Sullivan, G.C.; Pool-Zobel, B.; et al. Dietary synbiotics reduce cancer risk factors in polypectomized and colon cancer patients23. Am. J. Clin. Nutr. 2007, 85, 488–496. [Google Scholar] [CrossRef]
- Lee, H.A.; Kim, H.; Lee, K.-W.; Park, K.-Y. Dead nano-sized Lactobacillus plantarum inhibits azoxymethane/dextran sulfate sodium-induced colon cancer in Balb/c mice. J. Med. Food 2015, 18, 1400–1405. [Google Scholar] [CrossRef] [PubMed]
- El-Baz, A.M.; El-Mahmoudy, A.A.; Saber, S.; ElRakaiby, M.T. The coadministration of Lactobacillus probiotic augments the antitumor effect of telmisartan in rats. AMB Express 2025, 15, 38. [Google Scholar] [CrossRef] [PubMed]
- Frankel, T.L.; di Magliano, M.P. Immune sensing of microbial metabolites: Action at the tumor. Immunity 2022, 55, 192–194. [Google Scholar] [CrossRef]
- Yang, J.; McDowell, A.; Kim, E.K.; Seo, H.; Lee, W.H.; Moon, C.M.; Kym, S.M.; Lee, D.H.; Park, Y.S.; Jee, Y.K.; et al. Development of a colorectal cancer diagnostic model and dietary risk assessment through gut microbiome analysis. Exp. Mol. Med. 2019, 51, 1–15. [Google Scholar] [CrossRef]
- Rodriguez, D.M.; Hintze, K.J.; Rompato, G.; Wettere, A.J.V.; Ward, R.E.; Phatak, S.; Neal, C.; Armbrust, T.; Stewart, E.C.; Thomas, A.J.; et al. Dietary Supplementation with Black Raspberries Altered the Gut Microbiome Composition in a Mouse Model of Colitis-Associated Colorectal Cancer, although with Differing Effects for a Healthy versus a Western Basal Diet. Nutrients 2022, 14, 5270. [Google Scholar] [CrossRef]
- Papier, K.; Bradbury, K.E.; Balkwill, A.; Barnes, I.; Smith-Byrne, K.; Gunter, M.J.; Berndt, S.I.; Le Marchand, L.; Wu, A.H.; Peters, U.; et al. Diet-wide analyses for risk of colorectal cancer: Prospective study of 12,251 incident cases among 542,778 women in the UK. Nat. Commun. 2025, 16, 375. [Google Scholar] [CrossRef]
- Dehelean, C.A.; Marcovici, I.; Soica, C.; Mioc, M.; Coricovac, D.; Iurciuc, S.; Cretu, O.M.; Pinzaru, I. Plant-derived anticancer compounds as new perspectives in drug discovery and alternative therapy. Molecules 2021, 26, 1109. [Google Scholar] [CrossRef]
- Brockmueller, A.; Ruiz de Porras, V.; Shakibaei, M. Curcumin and its anti-colorectal cancer potential: From mechanisms of action to autophagy. Phytother. Res. 2024, 38, 3525–3551. [Google Scholar] [CrossRef]
- Bouyahya, A.; Taha, D.; Benali, T.; Zengin, G.; El Omari, N.; El Hachlafi, N.; Khalid, A.; Abdalla, A.N.; Ardianto, C.; Tan, C.S.; et al. Natural sources, biological effects, and pharmacological properties of cynaroside. Biomed. Pharmacother. 2023, 161, 114337. [Google Scholar] [CrossRef]
- Zhang, J.; Zhang, X.-X.; Zhang, W.-W.; Thakur, K.; Zhang, J.-G.; Fan, G. Luteoloside induces apoptosis in human HCT116 colorectal cancer cells via mitochondrial and PI3K/AKT signaling pathways. Food Biosci. 2024, 61, 104738. [Google Scholar] [CrossRef]
- Zhang, J.; Zhang, W.-W.; Zhang, X.-X.; Zhong, Q.; Thakur, K.; Zhang, J.-G.; Fan, G. Luteoloside inhibits cell migration and invasion in human HCT116 colorectal cancer via Ras/ERK/MAPK pathway. Food Biosci. 2025, 64, 105915. [Google Scholar] [CrossRef]
- Ionescu, V.A.; Diaconu, C.C.; Gheorghe, G.; Mihai, M.M.; Diaconu, C.C.; Bostan, M.; Bleotu, C. Gut Microbiota and Colorectal Cancer: A Balance Between Risk and Protection. Int. J. Mol. Sci. 2025, 26, 3733. [Google Scholar] [CrossRef]
- Bai, X.; Liu, B.; Fan, D.; Lu, Y.; Zhao, X. Modulating the gut microbiota: A novel perspective in colorectal cancer treatment. Cancer Lett. 2025, 612, 217459. [Google Scholar] [CrossRef] [PubMed]
- Byrd, D.A.; Damerell, V.; Gomez Morales, M.F.; Hogue, S.R.; Lin, T.; Ose, J.; Himbert, C.; Ilozumba, M.N.; Kahlert, C.; Shibata, D.; et al. The gut microbiome is associated with disease-free survival in stage I-III colorectal cancer patients. Int. J. Cancer 2025, 157, 64–73. [Google Scholar] [CrossRef] [PubMed]
- Tito, R.Y.; Verbandt, S.; Aguirre Vazquez, M.; Lahti, L.; Verspecht, C.; Llorens-Rico, V.; Vieira-Silva, S.; Arts, J.; Falony, G.; Dekker, E.; et al. Microbiome confounders and quantitative profiling challenge predicted microbial targets in colorectal cancer development. Nat. Med. 2024, 30, 1339–1348. [Google Scholar] [CrossRef] [PubMed]
- Forster, S.C.; Clare, S.; Beresford-Jones, B.S.; Harcourt, K.; Notley, G.; Stares, M.D.; Kumar, N.; Soderholm, A.T.; Adoum, A.; Wong, H.; et al. Identification of gut microbial species linked with disease variability in a widely used mouse model of colitis. Nat. Microbiol. 2022, 7, 590–599. [Google Scholar] [CrossRef]
- Piawah, S.; Kyaw, T.S.; Trepka, K.; Stewart, A.L.; Mora, R.V.; Stanfield, D.; Levine, K.; Van Blarigan, E.L.; Venook, A.; Turnbaugh, P.J.; et al. Associations between the Gut Microbiota, Race, and Ethnicity of Patients with Colorectal Cancer: A Pilot and Feasibility Study. Cancers 2023, 15, 4546. [Google Scholar] [CrossRef]
- Soderholm, A.T.; Pedicord, V.A. Intestinal epithelial cells: At the interface of the microbiota and mucosal immunity. Immunology 2019, 158, 267–280. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, J.; Wei, Z.-J.; Fan, G. Emerging Understanding of Gut Microbiome in Colorectal Cancer and Food-Related Intervention Strategies. Foods 2025, 14, 3040. https://doi.org/10.3390/foods14173040
Zhang J, Wei Z-J, Fan G. Emerging Understanding of Gut Microbiome in Colorectal Cancer and Food-Related Intervention Strategies. Foods. 2025; 14(17):3040. https://doi.org/10.3390/foods14173040
Chicago/Turabian StyleZhang, Jie, Zhao-Jun Wei, and Guangsen Fan. 2025. "Emerging Understanding of Gut Microbiome in Colorectal Cancer and Food-Related Intervention Strategies" Foods 14, no. 17: 3040. https://doi.org/10.3390/foods14173040
APA StyleZhang, J., Wei, Z.-J., & Fan, G. (2025). Emerging Understanding of Gut Microbiome in Colorectal Cancer and Food-Related Intervention Strategies. Foods, 14(17), 3040. https://doi.org/10.3390/foods14173040







