Abstract
Fermentation is one of the most ancient strategies to improve safety and extend shelf-life of the products. Starter cultures are mainly represented by lactic acid bacteria (LAB), which may also be bioprotective agents controlling the fermentation process, the native microbiota and pathogen outgrowth. This work aimed to select new LAB strains from spontaneously fermented sausages produced in different areas of Italy, which can be effective as starter cultures and bioprotective agents in fermented salami. The strains, mainly belonging to the Latilactobacillus sakei species, were characterized for their ability to inhibit major meat pathogens, the presence of antibiotic resistances and amine production. Moreover, technological performances, such as growth and acidification kinetics at increasing NaCl concentrations, were studied. As a result, new autochthonous Lat. sakei strains were obtained, lacking antibiotic resistance, possessing antimicrobial activity against Clostridium sporogenes, Listeria monocytogenes, Salmonella and Escherichia coli and with high growth performance under osmotic pressure. These strains have the potential for future application to improve the safety of fermented meats, even under conditions in which chemical preservatives are reduced or eliminated. Moreover, studies on autochthonous cultures are pivotal for guaranteeing specific characteristics of traditional products that represent an important cultural heritage.
1. Introduction
Fermented meat products, such as salamis, represent a cultural fingerprint dating back millennia. Traditional salami production in Italy is sometimes carried out on small-scale family-based farms, with in-house production of all raw materials [1]. The artisanal products bring the “taste of tradition” where indigenous microbiota, raw materials and production conditions are the main protagonists [2]. Spontaneous fermentation in artisanal small-scale salami production is led by environmental and meat-indigenous biota, whereas industrial-scale fermentation is standardized with the employment of starter cultures [3]. Fermented salamis are restricted ecological niches, harbouring lactic acid bacteria (LAB), Staphylococcus, Debaryomyces and Penicillium as the main bacterial and fungi genera. LAB are the most employed starters for several matrices, including meat, due to their technological, functional and safety features [3]. The limited carbon availability and the rich amino acid and lipid content in the meat matrix are the shaping parameters of the LAB metabolic behaviour during the fermentation and late ripening process. The initial sugar fermentation and the successive amino acid catabolism have a significant impact on the hygienic and sensory quality of long-ripened fermented meats [4].
Fast LAB growth and the consequent rapid pH drop are the most desirable technological and safety characteristic of meat starter cultures. The fast acidification in salami manufacturing mainly inhibits spoilage and pathogenic bacteria and contributes to desirable textural properties and sensory profile, due to the coagulation of muscle proteins [5]. Moreover, to ensure the safety of fermented dry-cured salamis, nitrate and nitrite are usually added to the meat before the fermentation process. However, this is raising concern among consumers because of the adverse effects of nitrite on health [6].
Fermentation and the consequent biopreservation is one of the most ancient microbial-based strategies to improve product safety and extend shelf-life [7]. Protective cultures used in fermented foods are mainly represented by LAB, which might also be added as starters aimed at controlling both the native microbiota and the pathogen outgrowth [8]. The production of antimicrobial compounds such as organic acids, hydrogen peroxide and lytic agents are some of the antagonistic mechanisms employed by bioprotective bacteria [9], but also the production of bacteriocins, known for their bactericidal effects at a specific concentration, may act against mainly gram-positive bacteria [10].
The employment of food grade cultures in Europe is regulated by the General EU Food Law (EU Regulation No. 178/2002). It relies on the obligations of the culture supplier for a careful safety assessment of the newly released products [11]. The safety of new microbial candidate strains should be guaranteed by the study of potentially transmissible antibiotic resistance traits and biogenic amine production [12], although antibiotic resistant strains and biogenic amine producers are often naturally present in foods [13,14]. Therefore, the safety characterization of strains to be employed as starter cultures is a prerequisite for their use.
In this work, indigenous LAB strains were isolated from spontaneously fermented sausages produced in different areas of Italy and characterized for their bioprotective function, safety assessment and technological performances. The aim was to obtain and characterise LAB strains that could be proposed as starter cultures for fermented sausage production and have, at the same time, antimicrobial activity against potential food pathogens.
2. Materials and Methods
2.1. Traditional Spontaneously Fermented Sausages Characterization
Three artisanal, homemade, starter-free salami produced in different Italian regions were considered for isolation of new LAB strains: Salame Bresciano (BRE), from the Lombardy region; Salame Romagnolo (ROM), from the Emilia Romagna Region; and Salame Basilicata (BAS), from the Basilicata Region. The sausages from Basilicata differed from the others in their traditional preservation. In fact, after ripening, they were stored in pots and covered with pork fat. For this reason, they are called “Salsiccia sotto sugna” [15]. The samples analysed in this work remained under fat for one month at room temperature before sampling. The main characteristics of these fermented sausages, obtained by the producers, are reported in Table 1.

Table 1.
Main characteristics of the fermented sausages used for the isolation of LAB strains.
All sausages contained pepper and no sugar was added. After aseptically removing the casing, 10 g of fermented sausages were 10-fold diluted with 90 mL of 0.9% (w/v) NaCl and homogenized in a Lab Blender Stomacher (Seward Medical, London, UK) for 2 min. Decimal dilutions were performed and plated onto selective media. For lactobacilli count, MRS agar (Oxoid, Milan, Italy) plates were used and incubated at 30 °C for 48 h in anaerobic conditions. Staphylococci were counted by surface-plating on Baird-Parker (Oxoid), added with egg yolk tellurite emulsion, with incubation at 30 °C for 48 h. For enterococci counts, Slanetz and Bartley medium (Oxoid) incubated at 42 °C for 48 h was used. Finally, Enterobacteriaceae were enumerated by plating in violet red bile glucose agar (Oxoid) at 37 °C for 24 h. The analyses were performed in triplicate.
The determination of water activity (aw) and pH was performed in triplicate by using an Aqualab CX3-TE (Labo-Scientifica, Parma, Italy) and a pH meter Basic 20 (Crison Instruments, Barcelona, Spain), respectively.
2.2. Isolation of LAB Strains and DNA Extraction
Colonies grown onto MRS plates were randomly picked with sterile loop and single colony lines were streaked onto new MRS agar plates. Each selected colony was then inoculated in 10 mL MRS broth and incubated at 30 °C for 24 h. Next, 1 mL overnight culture was submitted to DNA extraction with the Wizard® Genomic DNA Purification Kit (Promega Italia S.r.l., Milan, Italy). The remaining bacterial culture was centrifuged and from the obtained pellet single strain skim milk stocks were prepared and kept at −80 °C. Manufacturer’s instructions for isolating genomic DNA from gram-positive bacteria were followed, except for an additional lysis step (100 mg/mL of lysozyme, followed by an overnight incubation at 37 °C).
2.3. Fingerprinting-Based Clustering and 16S rRNA Identification of Biotypes
Clustering of LAB isolates was performed by Random Amplification of Polymorphic DNA (RAPD)-PCR as described by Di Gioia et al. [16]. Cluster analysis of obtained RAPD profiles was carried out with GelCompar II, 6.6 (Applied Maths, Sint-Martens-Latem, Belgium) using Dice’s coefficient of similarity with the un-weighted pair group method arithmetic averages clustering algorithm (UPGMA). Based on the genotypic clustering, amplification of 16S rRNA gene region of representative isolates was performed, according to Gaggia et al. [17] and then sequenced (MWG, Eurofins genomics). The obtained forward and reverse sequences were edited, and consensus sequences were built using the BioEdit software package. Sequences assignment to species or genera was achieved with the genomic data available on NCBI by BLASTn procedure (https://blast.ncbi.nlm.nih.gov/Blast.cgi?PAGE_TYPE=BlastSearch, accessed on 3 June 2021).
2.4. Determination of Antibiotic Susceptibility
Antibiotic susceptibility profiles of LAB were phenotypically determined, using Lact-1 and Lact-2 VetMIC microplates, purchased from the National Veterinary Institute (SVA, Uppsala, Sweden). Briefly, individual colonies were suspended in a sterile glass tube containing 4 mL maximum recovery diluent (Biolife, Milan, Italy) to a turbidity of 1 on the McFarland scale (~1 × 108 CFU/mL). Next, 20 μL from the bacterial suspension was diluted in 10 mL ISO-MRS broth (90% Iso-sensitest IST broth, Oxoid + 10% MRS broth) to obtain a final inoculum of ~5 × 105 CFU/mL. After filling with 100 μL of the final suspension (5 × 105 CFU/mL), VetMIC plates were sealed with provided clear film and incubated at 30 °C for 24–48 h, depending on the growth of the strain in the control wells.
An inverted light microscope was used for results interpretation. MIC was considered as the lowest concentration completely inhibiting visible growth. The resistance of LAB to antibiotics was determined according to the cut-off values reported by the FEEDAP Panel and adopted by EFSA [18] for gentamicin (16 mg/L), streptomycin (64 mg/L), tetracycline (8 mg/L), erythromycin (1 mg/L), clindamycin (1 mg/L), chloramphenicol (4 mg/L), kanamycin (16 mg/L for Leuconostoc sp. and 64 mg/L for lactobacilli facultative heterofermentative) and ampicillin (2 mg/L for Leuconostoc sp. and 4 mg/L for lactobacilli facultative heterofermentative).
2.5. Biogenic Amine Production
The identification of biogenic amines production was performed with the use of the medium proposed by Bover-Cid and Holzapfel [19] and further confirmed by HPLC analysis as described by Tabanelli et al. [20].
2.6. Antimicrobial Activity Assay
The potential antagonistic activity of LAB strains was evaluated using different indicator strains (listed in Table 2). The activity of all the cells was tested with a spot-on-the-lawn assay, whereas the potential production of antimicrobial compounds present in the cell-free supernatant was tested with the well-diffusion assay (WDA). For the direct antagonistic activity, 10 µL of fresh cell pellet, from LAB strains previously grown overnight, were spotted onto MRS agar plates and incubated for 24 h at 30 °C. Then, plates were overlaid with 10 mL of 0.8% BHI or RCM soft agar (Oxoid, Milan, Italy) containing 105 CFU/mL Listeria spp., E. coli, Salmonella spp. or Clostridium spp., respectively, and incubated as indicated in Table 2. Supernatants from meat-borne LAB strains, grown in MRS broth for 24 h at 30 °C, were obtained by centrifugation at 5000× g at 4 °C for 10 min. Unmodified and neutralized supernatants at pH 6.8–7.2 with 0.1–1 M NaOH were used for the WDA. After solidification of MRS, BHA (BHI+1.5% agar) and RCA (RCM+1.5% agar) agar plates inoculated with 105 CFU/mL of the indicator strain, 50 µL of supernatant were inoculated into preformed 6-mm diameter wells. Plates were initially placed for 2 h at 4 °C and further incubated as indicated in Table 2. Both assays were performed in duplicate and the presence of inhibition zones around the spotted cells or around the wells was analysed.

Table 2.
Indicator strains used for the in vitro antimicrobial activity.
2.7. Kinetic Modelling of Microbial Growth and Acidification
Growth and acidification kinetics of LAB were tested in MRS broth at 30 °C. Frozen stock strains were streaked on MRS agar plates, incubated for 48 h and colonies propagated overnight in MRS broth at 30 °C. Then, cells were inoculated at a concentration of about 105 CFU/mL in MRS containing 0%, 2%, 4%, 6% and 8% (w/v) of NaCl and incubated at 30 °C. Cell turbidity was recorded for 72 h with DEN-1B McFarland densitometer (Biosan, Riga, Latvia) and pH was monitored for the same time with pH meter Basic20 (Crison Instruments, Barcelona, Spain).
Data were modelled using the Gompertz equation as modified by Zwietering et al. [21].
When the growth was measured in McFarland units (MF), y is the MF value at time t; A represents maximum growth expressed as MF value, µmax is the maximum increase in the growth rate (MF values/h) and λ is the time (h) after which MF significantly changes from its baseline. In the case of the pH measurement, the meaning of the parameters was as follows: y is the pH decrease at time t; A represents the maximum pH decrease; µmax is the maximum pH decrease rate (ΔpH/h) and λ is the lag (h) to detect a measurable pH drop.
2.8. Statistical Analysis
The analyses regarding spontaneous fermented sausages characterization were performed in triplicate and the data were statistically analysed using the two-way ANOVA. The Tukey critical difference test was performed to determine differences between samples (p < 0.05).
3. Results
3.1. Spontaneous Fermented Sausages Characterization
The three spontaneous fermented sausages, produced without the use of starter cultures or the addition of sugars to meat batter, were characterized and used as a source of isolation of LAB. Table 3 shows the microbiological parameters, pH and aw of the sausages at the end of ripening.

Table 3.
Microbiological counts and chemico-physical features of the spontaneously fermented sausages at the end of ripening.
The products showed high pH values, because of low initial pH decrease due to slow and weak fermentation for the absence of added sugars and starter cultures. Differences among samples can be attributed to the absence, in BAS, of mould growth on the casing, and therefore not contributing to a rise in pH during ripening. The aw was particularly low in the BAS sample (0.831), while it was higher (0.908) in BRE, characterized by a larger diameter.
The microbiological analyses showed a high concentration of LAB in BRE and ROM, with a concentration of about 9 and 8 log CFU/g, respectively. On the contrary, this microbial population was present at a very low concentration (2.54 log CFU/g) in BAS, probably due to the low aw and storage conditions. In BAS, staphylococci and enterococci also showed low concentrations, being enterococci under the detection limit. In BRE and ROM, staphylococci were present at similar levels (about 7 log CFU/g) while enterococci were found in high concentration exclusively in the BRE sample (4.23 log CFU/g). This high number of enterococci could depend on several factors, such as the microbial quality of raw materials, the environment of production or the ripening conditions. The differences in the viable population in BAS can have many reasons. The BAS sample remained one month after ripening under fat, undergoing much longer osmotic stress than other samples that reached final aw values progressively over time during ripening. These stresses can have great influences on microbial populations and their cultivability. In addition, the absence of oxygen in BAS can affect the stress response mechanisms of several bacteria, in particular staphylococci.
3.2. Clustering of LAB Isolates and 16S rRNA Sequencing
Representative colonies were picked up from MRS plates deriving from ROM (97 colonies), BRE (50) and BAS (62). After purification, DNA was extracted from each isolate. RAPD fingerprinting and clustering analysis allowed the obtainment of a total of 41 biotypes, as 12 clusters and 10 single strains from ROM, 6 clusters and 6 single strains from BRE and 5 clusters and 5 single strains from BAS (results are shown in the Supplementary Material Figure S1). From this clustering, different scenarios in the spontaneously fermented salami could be observed: (i) dominance of a specific strain in BAS, represented by the cluster containing more than 80% of all isolates; (ii) about 50% single isolate in BRE, with no clear dominance of a particular isolate in the remaining 50%; and (iii) co-dominance of two clusters that comprise more than 50% of the isolates in ROM.
A total of 41 biotypes, comprising one representative strain for each cluster and the single strains, showing good growth after sub-culturing, were planned for 16S rRNA sequencing and subsequent analyses (as schematized in Figure 1). Three single isolates present in the clustering profiles (one from BRE and two form ROM) did not show growth after sub-culturing and were discarded. Therefore, the molecular identification was performed on 11 strains from BRE, 20 strains from ROM and 10 strains from BAS (Table 4). Results showed that Latilactobacillus sakei was the dominant species (Table 4) in 80.5% of all strains. This confirmed its strong adaptation to the meat matrix and, in particular, to fermented sausages [22,23]. However, within Lat. sakei, a great strain biodiversity could be appreciated in all sausages. Latilactobacillus curvatus was the second most abundant species among the biotypes, representing 9.1% in BRE, 15.0% in ROM and 20.0% in BAS, whereas Leuconostoc mesenteroides was detected only in BAS, representing 20.0% of all isolated strains.
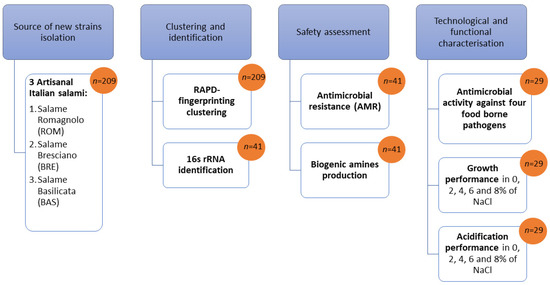
Figure 1.
Summary of the screening procedure for isolation and characterization of new potential bioprotective starter lactic acid bacteria (n = number of isolates).

Table 4.
Identification of lactobacilli isolated from three spontaneously fermented sausages.
Several authors reported that LAB species diversity in fermented sausages is limited. In fact, Lat. sakei predominates during the ripening process, given its competitiveness and assertiveness in the meat matrix, and it was evidenced as the dominant species in some European spontaneously fermented sausages, together with Lat. curvatus [24,25]. This predominance is mainly due to its physiological and technological characteristics and to its peculiar metabolic pathways for the meat ecological niche, including the arginine deiminase pathway and the utilization of nucleosides [26,27,28]. Lat. sakei can show a high biodiversity degree and, interestingly, strain-specific differences in species performance in meat environments were demonstrated [25].
The present work highlighted the presence of different biotypes in the same sample, but it is known that the co-dominance of these strains can confer competitive exclusion of other strains in the sausage environment and can lead to the peculiar characteristic of the products [29].
The scheme in Figure 1 was used both for the screening procedure and for safety and technological characterization of the studied biotypes.
3.3. Determination of Antibiotic Susceptibility (MIC)
The minimal inhibition concentration of eight antibiotics was examined in the 41 LAB biotypes previously identified, following the indications given by EFSA [18]. Table 5 shows the results obtained.

Table 5.
Phenotypic characterization of the antibiotic resistance in LAB meat-borne strains previously identified.
No strains were resistant to gentamycin, streptomycin, erythromycin and clindamycin, while the resistance against the other antibiotics was 14.6% for tetracycline and chloramphenicol, 7.3% for ampicillin and 2.4% for kanamycin.
Resistances to tetracycline and erythromycin are the most widely observed and studied among lactobacilli from fermented meats. Tetracycline is a commonly used antibiotic in pig farming, explaining the abundance of tet resistance genes in the pig microbiome and, consequently, in the raw meat used for fermented sausages [30]. In many cases, this specific resistance involved the lactic acid bacteria responsible for meat fermentation; however, the prevalence of resistant strains varied highly in products from different geographical areas [31,32]. In this study, all strains were susceptible to erythromycin and 6 out of 41 (14.6%) were tetracycline resistant. Three of these latter strains were isolated from ROM while the other three were from BAS. According to Fontana et al. [33], 26.4% of Lat. sakei and Lactiplantibacillus plantarum strains from fermented sausages differing in meat and geographical origin were resistant to tetracycline while the percentage was reduced at 10.7% for erythromycin. Although, according to Abriouel et al. [34], many lactobacilli are intrinsically resistant to kanamycin, streptomycin and gentamicin, in this study, no strains were resistant to streptomycin and gentamicin, and only one strain, isolated from BAS and belonging to Leuc. mesenteroides species, was resistant to kanamycin. Danielsen and Wind [35] found higher resistance for Lat. sakei/curvatus to kanamycin (66.7%), gentamicin (50%) and streptomycin (100%). Resistant strains were also not found for clindamycin, confirming the data reported by Danielsen and Wind [35] and Fontana et al. [27]. Three ampicillin-resistant strains were found, two from ROM and one from BAS. A higher percentage of resistant strains to this antibiotic were found by Aymerich et al. [36] and by Federici et al. [37].
A higher resistance frequency was observed for chloramphenicol. Five strains out of twenty isolated from ROM were resistant, while only one strain from BRE and none from BAS showed a MIC higher than the cut-off value. Aymerich et al. [36] described a low percentage (1.2%) of Lat. sakei resistant to this antibiotic, while Danielsen and Wind [35] found all the strains of Lat. sakei/curvatus susceptible.
Interestingly, three strains from ROM showed multiple resistances: Lat. sakei C12G (tetracycline and chloramphenicol), C22G (tetracycline and ampicillin) and E3G (tetracycline, chloramphenicol and ampicillin). It was demonstrated that genes responsible for tetracycline resistance (tetM and tetS, which encode ribosomal protection proteins) can mediate the resistance to other antimicrobials such as erythromycin, clindamycin and chloramphenicol [34].
3.4. Determination of Biogenic Amines
The 41 identified strains were tested for their ability to decarboxylate tyrosine, histidine, ornithine and lysine. Lat. curvatus C26G was the only biogenic amine producer strain. The HPLC analysis of C26G supernatant confirmed the production of putrescine (600 mg/L in Bover–Cid and Holzapfel medium). The production of putrescine by Lat. curvatus isolated from dairy products, meat and sausages, ranged from 10 to 10,000 mg/kg as reviewed in the work of Wunderlichova et al. [38]. Putrescine production derives from the decarboxylation of ornithine, an amino acid that can be produced through the arginine deiminase pathway [39]. No strains produced tyramine or histamine, the most dangerous biogenic amine [40,41]. This was expected for Lat. sakei strains, while those belonging to Lat. curvatus are often involved in tyramine production [42].
3.5. Antimicrobial Activity Assay
Twenty-nine strains lacking antimicrobial resistance traits and amino-biogenic potential were included in further screening steps (Figure 1). The LAB culture antagonistic activity against target foodborne pathogens showed that all strains were able to inhibit at least one pathogen to a different extent. Some strains, namely Lat. sakei E13G, E15G and C10B, were the most performant antagonistic strains against all four screened pathogens (Table 6). Conversely, non-neutralized and neutralized LAB supernatants did not exert any antimicrobial activity against the same pathogens.

Table 6.
Spot-test assay: direct antagonistic activity of meat-borne LAB against foodborne pathogens.
However, two Lat. sakei strains, namely C21B and E23B, showed an anti-LAB activity against Lat. curvatus DSM 20019T and Lat. sakei subsp. sakei LMG 13558T in the well diffusion assay (data not shown). Although strain-specific monitoring systems are rarely investigated, a recent study demonstrated how the bacteriogenic strain Lat. curvatus TMW 1.624 could exclude some of the Lat. sakei strains during sausage fermentation competition, while coexisting in the same environment with bacteriocin resistant Lat. sakei 1.417 [43]. The innate resistance against bacteriocin produced by closely related competitors was hypothesised as useful competitive fitness in Lat. sakei 23K [26]. Further investigation of bacteriocins in the Lat. sakei C21B and Lat. sakei E23B genome would elucidate the nature of this anti-LAB activity.
3.6. Growth and Acidification Kinetics
The growth and acidification behaviour of 29 safe LAB strains not presenting antibiotic resistance or amino biogenic potential and studied for their antimicrobial activity, were monitored through the turbidity increase (measured on the McFarland scale) and pH variation under different salt concentrations. The growth data for each strain were modelled with the Gompertz equation [21]. Table S1 and Figure 2 show the biotype growth performances at different NaCl concentrations (0, 2, 4, 6 and 8%). The strains were grouped according to their origin and, for each condition, the estimated parameters of the Gompertz equation (A, λ and μmax) were reported as mean (and standard deviation).
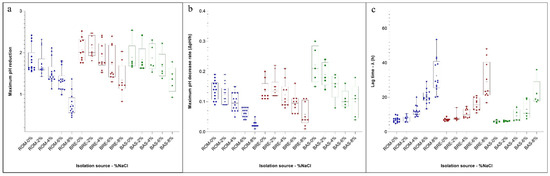
Figure 2.
Box and whiskers plots of the parameter estimated for the Gompertz equation of the pH drop curves of strains in relation to the isolation source and NaCl concentration. Boxes represent the limit of the 25th and 75th percentile of the distribution, the line inside the boxes represents the median and the whiskers represent the minimum and maximum values. (a) Maximum pH reduction estimates; (b) Maximum pH decrease rate estimates (ΔpH/h); (c) lag (h) to detect a measurable pH drop. Blue = Salame Romagnolo, Red = Salame Bresciano, Green = Salame Basilicata.
The maximum cell density of the McFarland scale (A) reached after the incubation was lower, as expected, in the presence of higher salt concentrations. No significant differences were observed in relation to the isolation source in cells grown at 0 and 2% of NaCl. In general, the strains isolated from BRE and BAS showed higher A estimates if compared with the strains isolated from ROM. It is noteworthy that comparable standard deviations characterize these data. However, this is not true for the means concerning the λ values. In fact, higher NaCl concentration not only determined longer λ time but also greater standard deviation of the means, indicating a wide variability in the responses of strains subjected to more severe stress conditions. Analogously to what was observed for A, the values of λ did not present significant differences in the samples added with 0 and 2% salt. Shorter λ characterized the strains deriving from BRE and BAS at a higher concentration than those isolated from ROM. The estimates of λ followed the same behaviour of A, showing higher values for BRE and BAS strains, and were characterized by homogeneous standard deviations. In fermented sausage production, the rate at which starter cultures decrease the pH is crucial for safety and technological aspects [44].
In the same samples analysed for the determination of the McFarland values, pH was also periodically monitored. In fact, the LAB metabolism increased the acidity, and the data were expressed as differences of pH with respect to the initial value. The curve obtained showed a behaviour similar to those obtained measuring the change of the turbidity on the McFarland scale. In this case, the results for the three parameters estimated with the Gompertz equation and obtained in relation to the isolation source and NaCl concentration are presented as a box and whiskers plot (Figure 2a–c).
Regarding the maximum pH decrease (parameter A of the model), data showed a great variability within the strains in relation to the source of isolation. In particular, biotypes isolated from ROM were generally characterized by lower pH decreases if compared with BRE and BAS, independently of the NaCl concentration. Even if the extent of the acid production was affected by the NaCl concentration, it is interesting to observe that the variability of the results within each group was of the same order.
Regarding the rate of acidification (the parameter µmax of the Gompertz equation) (Figure 2b), the strains isolated from BAS presented median values generally higher with respect to the other isolation sources, while the ROM isolates showed lower performances at higher salt concentrations.
Another important characteristic of the starter culture is the time needed to modify the initial pH of the medium (the λ value). Also, in this case, the strains isolated from BAS were the more competitive, especially at increasing NaCl concentrations. As observed for the λ value for the McFarland scale, the variability within each group markedly increased when the higher content of salts was considered. Highly performant strains could be observed at 6% of NaCl with a short lag phase of 7.5 h. A high heterogeneity among strains can be observed when higher osmotic stress was applied, with a lag phase ranging from 16.92 to 53.55 h. This again confirms the high physiological heterogeneity and potential technological traits of the indigenous meat-borne strains.
The growth and acidification performances of starter cultures are important parameters for ensuring the safety and quality of fermented meats. The rapidity with which the cells can multiply in defined conditions, colonizing the environment and producing organic acids, results in a more efficient inhibition of pathogens or undesirable microbial outgrowth [45]. In addition, in the case of fermented meats, a fast acidification performance represents a crucial technological point allowing the correct initial dehydration process of sausages [46].
The different behaviour shown in the metabolic activity in relation to different NaCl concentrations is often linked to the isolation source. It is noteworthy that the strains isolated from BAS, which were the most performant also at high NaCl concentrations, were those isolated from the harsher environment, whose final aw was 0.831. The great diversity within the species Lat. sakei is well known and justifies the differences in adaptation and competition in natural systems, such as fermented sausages [47]. In addition, there can be at least two relationships during meat colonization by members of this species, according to Janßen et al. (2018) [25]. In the first case, a strain outcompetes the other members of microbiota (assertiveness), based on peculiar metabolic characteristics (lag phase, growth rate, antimicrobials, stress responses). Alternatively, a co-dominance or a cooperation between different strains is established, with the consequent creation of a colonization resistance. According to the data reported here, the second strategy characterized the ROM sausages, while BAS and BRE presented a selection of a reduced number of strains at the end of production.
4. Conclusions
The present study allowed the selection of autochthonous meat-borne lactic acid bacteria that are safe for their utilization in fermented meat production, effective as starter cultures and promising as bioprotective agents. Some strains isolated from BAS samples showed high growth performances under osmotic pressure and strains C21G, E13G, E15G and C10B showed the greatest antimicrobial activity against C. sporogenes, L. monocytogenes, Salmonella spp. and E. coli. These natural isolates have the highest potential as starter cultures and as biopreservative agents in fermented sausage production. The real challenge for future application studies is the use of these microbial cultures to improve the safety of fermented meats, even in situations in which antimicrobial factors, such as the addition of nitrates and nitrites, are reduced or eliminated, as requested by consumers for their potential carcinogenic activity. At the same time, studies concerning the use of autochthonous cultures are pivotal for guaranteeing the specific characteristics and the recognizability of traditional and typical products representing an important cultural heritage.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/foods12040727/s1, Figure S1: RAPD-profiles of 209 autochthonous lactic acid bacteria, isolated from three artisanal Italian salami; Table S1: Mean values (±standard deviation) of the parameters A (Mc Farland value), λ (h) and μmax (h−1) of the growth curves obtained by the modelization with the Gompertz equation, for strains isolated from the different salami at different NaCl concentrations.
Author Contributions
Conceptualization, D.D.G.; Data curation, I.N. and F.B.; Formal analysis, I.N., G.T. and F.B.; Funding acquisition, D.D.G.; Investigation, I.N.; Methodology, I.N., G.T., L.B., F.G. (Fausto Gardini) and F.G. (Francesca Gaggìa); Supervision, F.G. (Fausto Gardini) and D.D.G.; Writing—original draft, I.N.; Writing—review and editing, G.T., L.B., F.G. (Fausto Gardini) and F.G. (Francesca Gaggìa). All authors have read and agreed to the published version of the manuscript.
Funding
The research described in this paper was funded by the project LONGLIFE, Joint Action Food Processing for Health of the Joint Programming Initiative a Healthy Diet for a Healthy Life (JPI HDHL) (https://www.healthydietforhealthylife.eu/index.php/joint-activities-2/re-port/183?p=1).
Data Availability Statement
The datasets generated during and/or analysed during the current study are available from the corresponding author upon reasonable request.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Roccato, A.; Uyttendaele, M.; Barrucci, F.; Cibin, V.; Favretti, M.; Cereser, A.; Dal Cin, M.; Pezzuto, A.; Piovesana, A.; Longo, A.; et al. Artisanal Italian salami and soppresse: Identification of control strategies to manage microbiological hazards. Food Microbiol. 2017, 61, 5–13. [Google Scholar] [CrossRef] [PubMed]
- Tabanelli, G.; Coloretti, F.; Chiavari, C.; Grazia, L.; Lanciotti, R.; Gardini, F. Effects of starter cultures and fermentation climate on the properties of two types of typical Italian dry fermented sausages produced under industrial conditions. Food Control 2012, 26, 416–426. [Google Scholar] [CrossRef]
- Franciosa, I.; Alessandria, V.; Dolci, P.; Rantsiou, K.; Cocolin, L. Sausage fermentation and starter cultures in the era of molecular biology methods. Int. J. Food Microbiol. 2018, 279, 26–32. [Google Scholar] [CrossRef] [PubMed]
- Gänzle, M.G. Lactic metabolism revisited: Metabolism of lactic acid bacteria in food fermentations and food spoilage. Curr. Opin. Food Sci. 2015, 2, 106–117. [Google Scholar] [CrossRef]
- Lücke, F.K. Utilization of microbes to process and preserve meat. Meat Sci. 2000, 56, 105–115. [Google Scholar] [CrossRef] [PubMed]
- IARC. IARC Monographs on the Evaluation of Carcinogenic Risks to Humans. 2010. Available online: http://monographs.iarc.fr/ENG/Monographs/vol96/index.php (accessed on 1 February 2020).
- Stiles, M.E. Biopreservation by lactic acid bacteria. Antonie Van Leeuwenhoek 1996, 70, 331–345. [Google Scholar] [CrossRef] [PubMed]
- Zagorec, M.; Champomier-Vergès, M.C. Lactobacillus sakei: A starter for sausage fermentation, a protective culture for meat products. Microorganisms 2017, 5, 56. [Google Scholar] [CrossRef]
- Gálvez, A.; Abriouel, H.; Benomar, N.; Lucas, R. Microbial antagonists to food-borne pathogens and biocontrol. Curr. Opin. Biotechnol. 2010, 21, 142–148. [Google Scholar] [CrossRef]
- Chikindas, M.L.; Weeks, R.; Drider, D.; Chistyakov, V.A.; Dicks, L.M.T. Functions and emerging applications of bacteriocins. Curr. Opin. Biotechnol. 2018, 49, 23–28. [Google Scholar] [CrossRef]
- Laulund, S.; Wind, A.; Derkx, P.M.F.; Zuliani, V. Regulatory and safety requirements for food cultures. Microorganisms 2017, 5, 28. [Google Scholar] [CrossRef]
- Herman, L.; Chemaly, M.; Cocconcelli, P.S.; Fernandez, P.; Klein, G.; Peixe, L.; Prieto, M.; Querol, A.; Suarez, J.E.; Sundh, I.; et al. The qualified presumption of safety assessment and its role in EFSA risk evaluations: 15 years past. FEMS Microbiol. Lett. 2019, 366, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Jaimee, G.; Halami, P.M. High level aminoglycoside resistance in Enterococcus, Pediococcus and Lactobacillus species from farm animals and commercial meat products. Ann. Microbiol. 2016, 66, 101–110. [Google Scholar] [CrossRef]
- Coton, M.; Lebreton, M.; Leyva Salas, M.; Garnier, L.; Navarri, M.; Pawtowski, A.; Le Blay, G.; Valence, F.; Coton, E.; Mounier, J. Biogenic amine and antibiotic resistance profiles determined for lactic acid bacteria and a Propionibacterium prior to use as antifungal bioprotective cultures. Int. Dairy J. 2018, 85, 21–26. [Google Scholar] [CrossRef]
- Gardini, F.; Suzzi, S.; Lombardi, A.; Galgano, F.; Crudele, M.A.; Andrighetto, C.; Schirone, M.; Tofalo, R. A survey of yeasts in traditional sausages of southern Italy. FEMS Yeast Res. 2001, 1, 161–167. [Google Scholar] [CrossRef] [PubMed]
- Di Gioia, D.; Mazzola, G.; Nikodinoska, I.; Aloisio, I.; Langerholc, T.; Rossi, M.; Raiomondi, S.; Melero, B.; Rovira, J. Lactic acid bacteria as protective cultures in fermented pork meat to prevent Clostridium spp. growth. Int. J. Food Microbiol. 2016, 235, 53–59. [Google Scholar] [CrossRef]
- Gaggìa, F.; Baffoni, L.; Galiano, M.; Nielsen, D.S.; Jakobsen, R.R.; Castro-Mejía, J.L.; Bosi, S.; Truzzi, F.; Musumeci, F.; Dinelli, G.; et al. Kombucha beverage from green, black and rooibos teas: A comparative study looking at microbiology, chemistry and antioxidant activity. Nutrients 2018, 11, 1. [Google Scholar] [CrossRef]
- European Food Safety Authority (EFSA)-FEEDAP. Guidance on the assessment of bacterial susceptibility to antimicrobials of human and veterinary importance. EFSA J. 2012, 10, 2740–2749. [Google Scholar] [CrossRef]
- Bover-Cid, S.; Holzapfel, W.H. Improved screening procedure for biogenic amine production by lactic acid bacteria. Int. J. Food Microbiol. 1999, 53, 33–41. [Google Scholar] [CrossRef]
- Tabanelli, G.; Montanari, C.; Bargossi, E.; Lanciotti, R.; Gatto, V.; Felis, G.; Torriani, S.; Gardini, F. Control of tyramine and histamine accumulation by lactic acid bacteria using bacteriocin forming lactococci. Int. J. Food Microbiol. 2014, 190, 14–23. [Google Scholar] [CrossRef]
- Zwietering, M.H.; Jongenburger, I.; Rombouts, F.M.; Van’t Riet, K. Modeling of the bacterial growth curve. Appl. Environ. Microbiol. 1990, 56, 1875–1881. [Google Scholar] [CrossRef]
- Tremonte, P.; Sorrentino, E.; Pannella, G.; Tipaldia, L.; Sturchio, M.; Masucci, A.; Maiuro, L.; Coppola, R.; Succia, M. Detection of different microenvironments and Lactobacillus sakei biotypes in Ventricina, a traditional fermented sausage from central Italy. Int. J. Food Microbiol. 2017, 242, 132–140. [Google Scholar] [CrossRef] [PubMed]
- Guilbaud, M.; Zagorec, M.; Chaillou, S.; Champomier-Vergès, M.C. Intraspecies diversity of Lactobacillus sakei response to oxidative stress and variability of strain performance in mixed strains challenges. Food Microbiol. 2012, 29, 197–204. [Google Scholar] [CrossRef] [PubMed]
- Barbieri, F.; Tabanelli, G.; Montanari, C.; Dall’Osso, N.; Šimat, V.; Smole Možina, S.; Baños, A.; Özogul, F.; Bassi, D.; Fontana, C.; et al. Mediterranean spontaneously fermented sausages: Spotlight on microbiological and quality features to exploit their bacterial biodiversity. Foods 2021, 10, 2691. [Google Scholar] [CrossRef]
- Janßen, D.; Eisenbach, L.; Ehrmann, M.A.; Vogel, R.F. Assertiveness of Lactobacillus sakei and Lactobacillus curvatus in a fermented sausage model. Int. J. Food Microbiol. 2018, 285, 188–197. [Google Scholar] [CrossRef] [PubMed]
- Chaillou, S.; Champomier-Vergès, M.C.; Cornet, M. The complete genome sequence of the meat-borne lactic acid bacterium Lactobacillus sakei 23K. Nat. Biotechnol. 2005, 23, 1527–1533. [Google Scholar] [CrossRef] [PubMed]
- Fontana, C.; Bassi, D.; López, C.; Pisacane, V.; Otero, M.C.; Puglisi, E.; Rebecchi, A.; Cocconcelli, P.S.; Vignolo, G. Microbial ecology involved in the ripening of naturally fermented llama meat sausages. A focus on lactobacilli diversity. Int. J. Food Microbiol. 2016, 236, 17–25. [Google Scholar] [CrossRef] [PubMed]
- Montanari, C.; Barbieri, F.; Magnani, M.; Grazia, L.; Gardini, F.; Tabanelli, G. Phenotypic diversity of Lactobacillus sakei strains. Front. Microbiol. 2003, 9, 2009. [Google Scholar] [CrossRef]
- Widenmann, A.W.; Schiffer, C.J.; Ehrmann, M.A.; Vogel, R.F. Impact of different sugars and glycosyltransferases on the assertiveness of Latilactobacillus sakei in raw sausage fermentations. Int. J. Food Microbiol. 2022, 366, 109575. [Google Scholar] [CrossRef]
- Monger, X.C.; Gilbert, A.A.; Saucier, L.; Vincent, A.T. Antibiotic resistance: From pig to meat. Antibiotics 2021, 10, 1209. [Google Scholar] [CrossRef]
- Gevers, D.; Danielsen, M.; Huys, G.; Swings, J. Molecular characterization of tet (M) genes in Lactobacillus isolates from different types of fermented dry sausage. Appl. Environ. Microb. 2003, 69, 1270–1275. [Google Scholar] [CrossRef]
- Zonenschain, D.; Rebecchi, A.; Morelli, L. Erythromycin- and tetracycline-resistant lactobacilli in Italian fermented dry sausages. J. Appl. Microbiol. 2009, 107, 1559–1568. [Google Scholar] [CrossRef] [PubMed]
- Fontana, C.; Patrone, V.; Lopez, C.M.; Morelli, L.; Rebecchi, A. Incidence of tetracycline and erythromycin resistance in meat-associated bacteria: Impact of different livestock management strategies. Microorganisms 2021, 9, 2111. [Google Scholar] [CrossRef] [PubMed]
- Abriouel, H.; Muñoz, M.D.C.C.; Lerma, L.L.; Montoro, B.P.; Bockelmann, W.; Pichner, R.; Kabisch, J.; Cho, G.-S.; Franz, C.M.A.P.; Gálvez, A.; et al. New insights in antibiotic resistance of Lactobacillus species from fermented foods. Food Res. Int. 2015, 78, 465–481. [Google Scholar] [CrossRef] [PubMed]
- Danielsen, M.; Wind, A. Susceptibility of Lactobacillus spp. to antimicrobial agents. Int. J. Food Microbiol. 2003, 82, 1–11. [Google Scholar] [CrossRef]
- Aymerich, T.; Martin, B.; Garriga, M.; Vidal-Carou, M.C.; Bover-Cid, S.; Hugas, M. Safety properties and molecular strain typing of lactic acid bacteria from slightly fermented sausages. J. Appl. Microbiol. 2006, 100, 40–49. [Google Scholar] [CrossRef] [PubMed]
- Federici, S.; Ciarrocchi, F.; Campana, R.; Ciandrini, E.; Blasi, G.; Baffone, W. Identification and functional traits of lactic acid bacteria isolated from Ciauscolo salami produced in Central Italy. Meat Sci. 2014, 98, 575–584. [Google Scholar] [CrossRef]
- Wunderlichova, L.; Bunkova, L.; Koutry, M.; Jancova, P.; Bunka, F. Formation, degradation, and detoxification of putrescine by foodborne bacteria: A review. Compr. Rev. Food Sci. Food Saf. 2014, 13, 1012–1030. [Google Scholar] [CrossRef]
- Rimaux, T.; Rivière, A.; Illeghems, K.; Weckx, S.; De Vuyst, L.; Leroy, F. Expression of the arginine deiminase pathway genes in Lactobacillus sakei is strain dependent and is affected by the environmental pH. Appl. Environ. Microbiol. 2012, 78, 4874–4883. [Google Scholar] [CrossRef]
- Omer, A.K.; Mohammed, R.R.; Ameen, P.S.M.; Abas, Z.A.; Ekici, K. Presence of biogenic amines in food and their public health implications: A review. J. Food Protect. 2021, 84, 1539–1548. [Google Scholar] [CrossRef]
- EFSA Panel on Biological Hazards (BIOHAZ). Scientific opinion on risk based control of biogenic amine formation in fermented foods. EFSA J. 2011, 9, 2393. [Google Scholar] [CrossRef]
- Marcobal, A.; De Las Rivas, B.; Landete, J.M.; Tabera, L.; Muñoz, R. Tyramine and phenylethylamine biosynthesis by food bacteria. Crit. Rev. Food Sci. 2012, 52, 448–467. [Google Scholar] [CrossRef] [PubMed]
- Janßen, D.; Dworschak, L.; Ludwig, C.; Ehrmann, M.A.; Vogel, R.F. Interspecies assertiveness of Lactobacillus curvatus and Lactobacillus sakei in sausage fermentations. Int. J. Food Microbiol. 2020, 331, 108689. [Google Scholar] [CrossRef] [PubMed]
- Fontana, C.; Cocconcelli, P.S.; Vignolo, G.; Saavedra, L. Occurrence of antilisterial structural bacteriocins genes in meat borne lactic acid bacteria. Food Control 2015, 47, 53–59. [Google Scholar] [CrossRef]
- Swinnen, I.A.M.; Bernaerts, K.; Dens, E.J.J.; Geeraerd, A.H.; Impe, V.J.F. Predictive modelling of the microbial lag phase: A review. Int. J. Food Microbiol. 2004, 94, 137–159. [Google Scholar] [CrossRef]
- Ammor, S.; Dufour, E.; Zagorec, M.; Chaillou, S.; Chevallier, I. Characterization and selection of Lactobacillus sakei strains isolated from traditional dry sausage for their potential use as starter cultures. Food Microbiol. 2005, 22, 529–538. [Google Scholar] [CrossRef]
- Chaillou, S.; Lucquin, I.; Najjari, A.; Zagorec, M.; Champomier-Vergès, M.C. Population genetics of Lactobacillus sakei reveals three lineages with distinct evolutionary histories. PLoS ONE 2013, 8, e73253. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).