Associations between the Bacterial Composition of Farm Bulk Milk and the Microbiota in the Resulting Swedish Long-Ripened Cheese
Abstract
:1. Introduction
2. Materials and Methods
2.1. Participating Farms
2.2. Cheese Production Practice
2.3. Sampling of Farm Bulk Milk, Dairy Silo Milk and the Resulting Cheese
2.4. Total Bacteria Count in Farm Bulk Milk, Dairy Silo Milk and Cheese
2.5. DNA Extraction from Bacteria in Milk and Cheese
2.6. Illumina Amplicon Library Construction, Sequencing and Bioinformatic Analysis
2.7. Statistical Analysis
3. Results
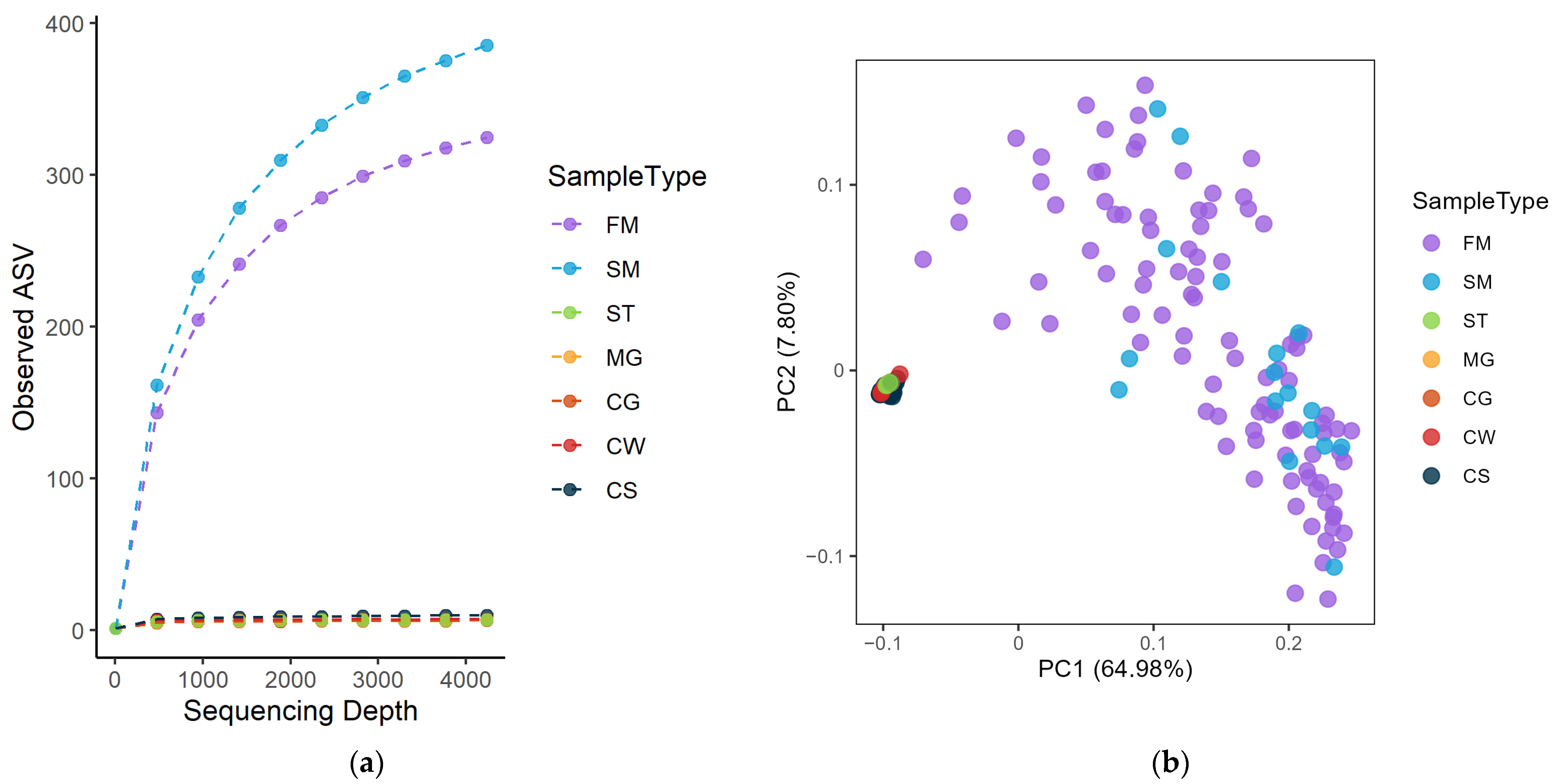
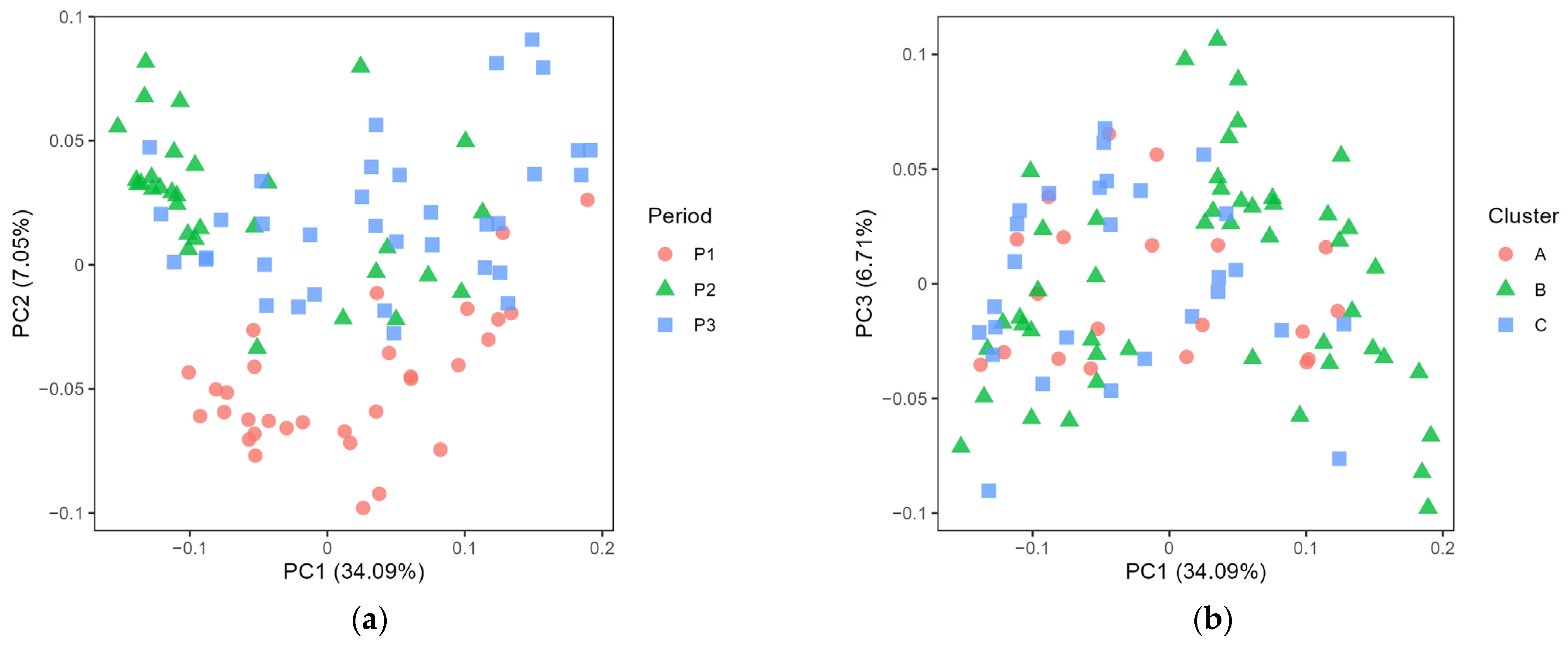
3.1. Exploring the Microbiota Present in Farm Bulk and Dairy Silo Milk
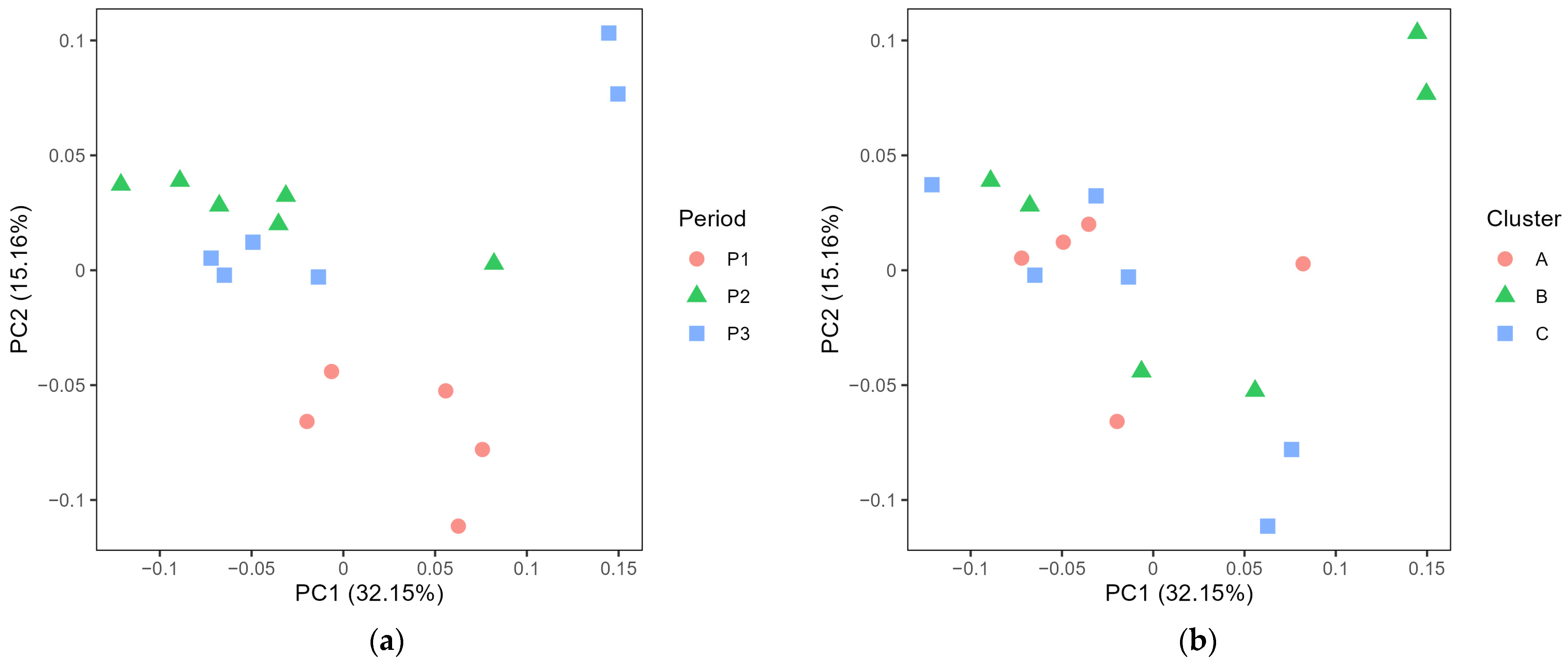
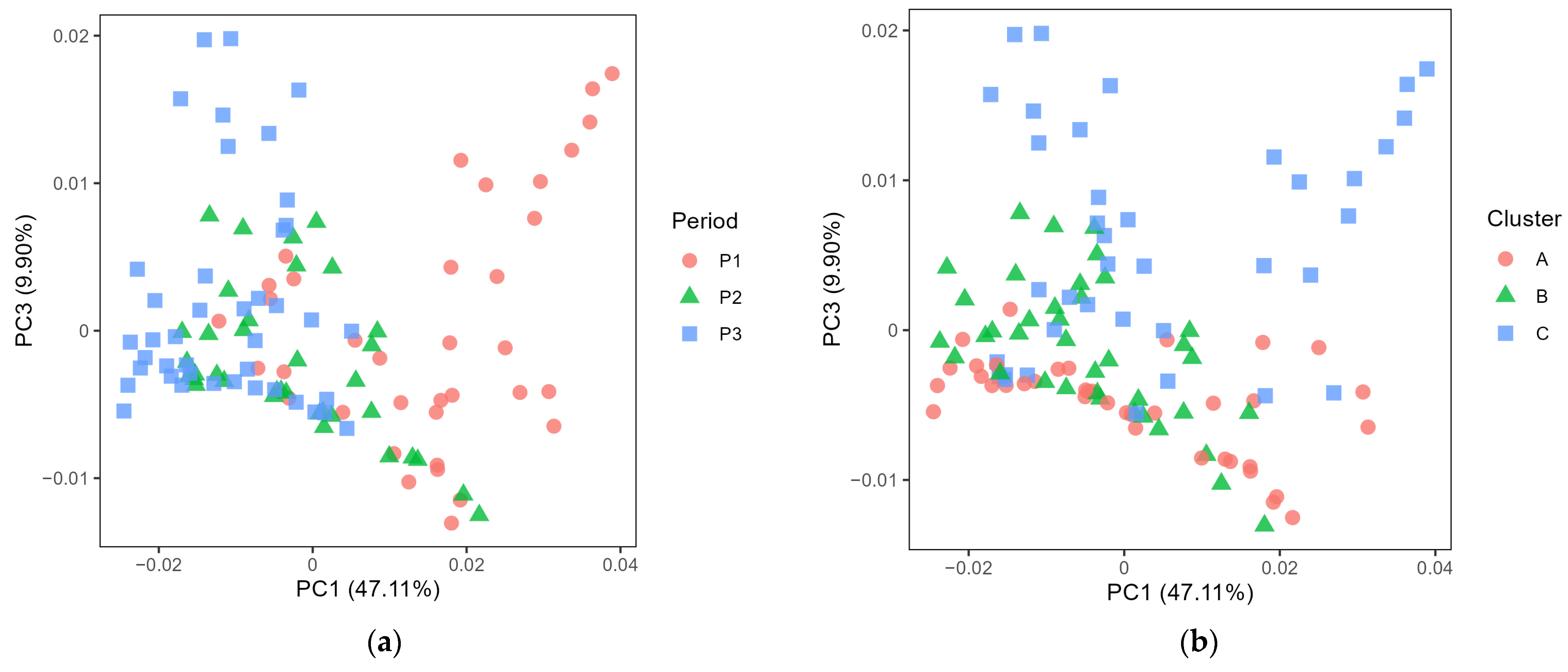
3.2. Exploring the Microbiota in Cheese
3.3. Common ASVs Found in Different Sample Types
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- McSweeney, P.L.H. Biochemistry of cheese ripening. Int. J. Dairy Technol. 2004, 57, 127–144. [Google Scholar] [CrossRef]
- Ardö, Y. Flavour formation by amino acid catabolism. Biotechnol. Adv. 2006, 24, 238–242. [Google Scholar] [CrossRef]
- Bokulich, N.A.; Mills, D.A. Facility-specific “house” microbiome drives microbial landscapes of artisan cheesemaking plants. Appl. Environ. Microbiol. 2013, 79, 5214–5223. [Google Scholar] [CrossRef]
- Broadbent, J.R.; Houck, K.; Johnson, M.; Oberg, C. Influence of adjunct use and cheese microenvironment on nonstarter bacteria in reduced-fat Cheddar-type cheese. J. Dairy Sci. 2003, 86, 2773–2782. [Google Scholar] [CrossRef]
- Levante, A.; De Filippis, F.; La Storia, A.; Gatti, M.; Neviani, E.; Ercolini, D.; Lazzi, C. Metabolic gene-targeted monitoring of non-starter lactic acid bacteria during cheese ripening. Int. J. Food Microbiol. 2017, 257, 276–284. [Google Scholar] [CrossRef]
- Beresford, T.P.; Fitzsimons, N.A.; Brennan, N.L.; Cogan, T.M. Recent advances in cheese microbiology. Int. Dairy J. 2001, 11, 259–274. [Google Scholar] [CrossRef]
- Williams, A.G.; Choi, S.-C.; Banks, J.M. Variability of the species and strain phenotype composition of the non-starter lactic acid bacterial population of cheddar cheese manufactured in a commercial creamery. Food Res. Int. 2002, 35, 483–493. [Google Scholar] [CrossRef]
- Sun, L.; Lundh, Å.; Höjer, A.; Bernes, G.; Nilsson, D.; Johansson, M.; Hetta, M.; Gustafsson, A.H.; Saedén, K.H.; Dicksved, J. Milking system and premilking routines have a strong effect on the microbial community in bulk tank milk. J. Dairy Sci. 2022, 105, 123–139. [Google Scholar] [CrossRef] [PubMed]
- Doyle, C.J.; Gleeson, D.; O’Toole, P.W.; Cotter, P.D. Impacts of seasonal housing and teat preparation on raw milk microbiota: A high-throughput sequencing study. Appl. Environ. Microbiol. 2017, 83, e02694-16. [Google Scholar] [CrossRef] [PubMed]
- Gagnon, M.; Ouamba, A.J.; LaPointe, G.; Chouinard, P.Y.; Roy, D. Prevalence and abundance of lactic acid bacteria in raw milk associated with forage types in dairy cow feeding. J. Dairy Sci. 2020, 103, 5931–5946. [Google Scholar] [CrossRef]
- Priyashantha, H.; Lundh, Å.; Höjer, A.; Bernes, G.; Nilsson, D.; Hetta, M.; Saedén, K.H.; Gustafsson, A.H.; Johansson, M. Composition and properties of bovine milk: A study from dairy farms in northern Sweden; Part I. Eff. Dairy Farming System. J. Dairy Sci. 2021, 104, 8582–8594. [Google Scholar]
- Hugerth, L.W.; Wefer, H.A.; Lundin, S.; Jakobsson, H.E.; Lindberg, M.; Rodin, S.; Engstrand, L.; Andersson, A.F. DegePrime, a Program for Degenerate Primer Design for Broad-Taxonomic-Range PCR in Microbial Ecology Studies. Appl. Environ. Microbiol. 2014, 80, 5116–5123. [Google Scholar] [CrossRef]
- Sun, L.; Dicksved, J.; Priyashantha, H.; Lundh, Å.; Johansson, M. Distribution of bacteria between different milk fractions, investigated using culture-dependent methods and molecular-based and fluorescent microscopy approaches. J. Appl. Microbiol. 2019, 127, 1028–1037. [Google Scholar] [CrossRef] [PubMed]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 2019, 37, 852–857. [Google Scholar] [CrossRef] [PubMed]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef]
- Katoh, K.; Misawa, K.; Kuma, K.-i.; Miyata, T. MAFFT: A novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 2002, 30, 3059–3066. [Google Scholar] [CrossRef]
- Price, M.N.; Dehal, P.S.; Arkin, A.P. FastTree 2—Approximately Maximum-Likelihood Trees for Large Alignments. PLoS ONE 2010, 5, e9490. [Google Scholar] [CrossRef]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V. Scikit-learn: Machine learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2013, 41, D590–D596. [Google Scholar] [CrossRef]
- Bokulich, N.A.; Kaehler, B.D.; Rideout, J.R.; Dillon, M.; Bolyen, E.; Knight, R.; Huttley, G.A.; Gregory Caporaso, J. Optimizing taxonomic classification of marker-gene amplicon sequences with QIIME 2’s q2-feature-classifier plugin. Microbiome 2018, 6, 90. [Google Scholar] [CrossRef]
- Kruskal, W.H.; Wallis, W.A. Use of ranks in one-criterion variance analysis. J. Am. Stat. Assoc. 1952, 47, 583–621. [Google Scholar] [CrossRef]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Soc. Ser. B Stat. Methodol. 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Anderson, M.J. A new method for non-parametric multivariate analysis of variance. Austral Ecol. 2001, 26, 32–46. [Google Scholar]
- Choi, J.; In Lee, S.; Rackerby, B.; Frojen, R.; Goddik, L.; Ha, S.-D.; Park, S.H. Assessment of overall microbial community shift during Cheddar cheese production from raw milk to aging. Appl. Microbiol. Biotechnol. 2020, 104, 6249–6260. [Google Scholar] [CrossRef] [PubMed]
- Bottari, B.; Levante, A.; Bancalari, E.; Sforza, S.; Bottesini, C.; Prandi, B.; De Filippis, F.; Ercolini, D.; Nocetti, M.; Gatti, M. The Interrelationship between Microbiota and Peptides during Ripening as a Driver for Parmigiano Reggiano Cheese Quality. Front. Microbiol. 2020, 11, 581658. [Google Scholar] [CrossRef]
- McHugh, A.J.; Feehily, C.; Fenelon, M.A.; Gleeson, D.; Hill, C.; Cotter, P.D. Tracking the Dairy Microbiota from Farm Bulk Tank to Skimmed Milk Powder. mSystems 2020, 5, e00226-20. [Google Scholar] [CrossRef] [PubMed]
- Skeie, S.B.; Håland, M.; Thorsen, I.M.; Narvhus, J.; Porcellato, D. Bulk tank raw milk microbiota differs within and between farms: A moving goalpost challenging quality control. J. Dairy Sci. 2019, 102, 1959–1971. [Google Scholar] [CrossRef]
- Porcellato, D.; Smistad, M.; Bombelli, A.; Abdelghani, A.; Jørgensen, H.J.; Skeie, S.B. Longitudinal study of the bulk tank milk microbiota reveals major temporal shifts in composition. Front. Microbiol. 2021, 12, 616429. [Google Scholar] [CrossRef]
- Porcellato, D.; Skeie, S.B. Bacterial dynamics and functional analysis of microbial metagenomes during ripening of Dutch-type cheese. Int. Dairy J. 2016, 61, 182–188. [Google Scholar] [CrossRef]
- de Vos, W.M. Systems solutions by lactic acid bacteria: From paradigms to practice. Microb. Cell Factories 2011, 10, S2. [Google Scholar] [CrossRef] [PubMed]
- Somers, E.; Johnson, M.; Wong, A. Biofilm formation and contamination of cheese by nonstarter lactic acid bacteria in the dairy environment. J. Dairy Sci. 2001, 84, 1926–1936. [Google Scholar] [CrossRef]
- Rehn, U.; Petersen, M.A.; Saedén, K.H.; Ardö, Y. Ripening of extra-hard cheese made with mesophilic DL-starter. Int. Dairy J. 2010, 20, 844–851. [Google Scholar] [CrossRef]
- Dasen, A.; Berthier, F.; Grappin, R.; Williams, A.G.; Banks, J. Genotypic and phenotypic characterization of the dynamics of the lactic acid bacterial population of adjunct-containing Cheddar cheese manufactured from raw and microfiltered pasteurised milk. J. Appl. Microbiol. 2003, 94, 595–607. [Google Scholar] [CrossRef] [PubMed]
- Ruggirello, M.; Cocolin, L.; Dolci, P. Fate of Lactococcus lactis starter cultures during late ripening in cheese models. Food Microbiol. 2016, 59, 112–118. [Google Scholar] [CrossRef]
- Desfossés-Foucault, É.; LaPointe, G.; Roy, D. Dynamics and rRNA transcriptional activity of lactococci and lactobacilli during Cheddar cheese ripening. Int. J. Food Microbiol. 2013, 166, 117–124. [Google Scholar] [CrossRef]
- van Mastrigt, O.; Gallegos Tejeda, D.; Kristensen, M.N.; Abee, T.; Smid, E.J. Aroma formation during cheese ripening is best resembled by Lactococcus lactis retentostat cultures. Microb. Cell Factories 2018, 17, 104. [Google Scholar] [CrossRef]
- van Mastrigt, O.; Abee, T.; Lillevang, S.K.; Smid, E.J. Quantitative physiology and aroma formation of a dairy Lactococcus lactis at near-zero growth rates. Food Microbiol. 2018, 73, 216–226. [Google Scholar] [CrossRef]
- Barzideh, Z.; Siddiqi, M.; Mohamed, H.M.; LaPointe, G. Dynamics of Starter and Non-Starter Lactic Acid Bacteria Populations in Long-Ripened Cheddar Cheese Using Propidium Monoazide (PMA) Treatment. Microorganisms 2022, 10, 1669. [Google Scholar] [CrossRef]
- Hols, P.; Hancy, F.; Fontaine, L.; Grossiord, B.; Prozzi, D.; Leblond-Bourget, N.; Decaris, B.; Bolotin, A.; Delorme, C.; Dusko Ehrlich, S. New insights in the molecular biology and physiology of Streptococcus thermophilus revealed by comparative genomics. FEMS Microbiol. Rev. 2005, 29, 435–463. [Google Scholar]
- Iyer, R.; Tomar, S.; Maheswari, T.U.; Singh, R. Streptococcus thermophilus strains: Multifunctional lactic acid bacteria. Int. Dairy J. 2010, 20, 133–141. [Google Scholar] [CrossRef]
- Johnson, J.; Selover, B.; Curtin, C.; Waite-Cusic, J. Nonstarter Bacterial Communities in Aged Cheddar Cheese: Patterns on Two Timescales. Appl. Environ. Microbiol. 2022, 88, e01939-21. [Google Scholar] [CrossRef] [PubMed]



| Farm Cluster and Selection Criteria | Number of Farms | Average Number of Cows per Herd (Min–Max) 1 | Dominant Cow Breed 2 on the Farm (Number of Farms) | Milking System (Number of Farms) | Proportion of Silage Fed as Round Bales 1 |
|---|---|---|---|---|---|
| A: Average milk quality | 4 | 110 a (68–180) | SH (3) Mixed (1) | AMS (3) Milking parlour (1) | 29% b |
| B: Higher fat and protein content, fewer clostridia in milk | 9 | 49 b (17–90) | SH (1) SRB (2) SKB (1) SJB (1) Mixed (4) | Tie-stall (6) AMS (2) Milking parlour (1) | 73% a |
| C: Higher total bacteria number, higher abundance of lactic acid bacteria in milk | 5 | 84 a (41–127) | SH (4) SKB (1) | AMS (3) Milking parlour (2) | 60% ab |
| Batches with ASV Found in Cheese and in Samples from Early Stages in the Process | Average RA (%) in Cheese Sampled after Ageing for 7–20 Months | |||
|---|---|---|---|---|
| ASV | FM | SM 1 | EPS 2 | |
| Lactococcus b39c | 17 | 17 | 17 | 42.1 |
| Leuconostoc 7f62 | 10 | 6 | 17 | 41.7 |
| Leuconostoc 24cd | 11 | 2 | 15 | 6.1 |
| Lactococcus d90e | 16 | 8 | 17 | 3.2 |
| Streptococcus bd2e | 18 | 16 | 9 | 1.8 |
| Lactobacillus c982 | 17 | 12 | 11 | 1.3 |
| Lactobacillus 2653 | 0 | 0 | 0 | 0.75 |
| Lactobacillus 3486 | 6 | 4 | 2 | 0.61 |
| Lactococcus 072f | 0 | 0 | 14 | 0.52 |
| Lactococcus bb15 | 0 | 0 | 11 | 0.51 |
| Lactococcus cc1f | 8 | 5 | 11 | 0.45 |
| Lactobacillus 0b36 | 1 | 0 | 0 | 0.34 |
| Acinetobacter 6933 | 0 | 0 | 1 | 0.17 |
| Streptococcus 0c46 | 2 | 1 | 1 | 0.15 |
| Lactobacillus bf19 | 0 | 0 | 1 | 0.13 |
| Lactobacillus 57c1 | 1 | 0 | 0 | 0.064 |
| Lactococcus 6fa5 | 0 | 0 | 0 | 0.041 |
| Romboutsia 464a | 15 | 14 | 0 | 0.013 |
| Tetragenococcus 49a8 | 1 | 0 | 0 | 0.011 |
| Turicibacter dacd | 12 | 11 | 0 | 0.011 |
| Paeniclostridium 69a9 | 14 | 13 | 0 | 0.010 |
| Streptococcus 84e3 | 1 | 1 | 0 | 0.010 |
| Lactobacillus 61ee | 2 | 2 | 1 | 0.008 |
| Staphylococcus 1355 | 10 | 9 | 0 | 0.005 |
| Clostridium ddf8 | 0 | 0 | 1 | 0.005 |
| Trueperella b116 | 4 | 3 | 0 | 0.005 |
| Enterococcus 028e | 5 | 3 | 0 | 0.004 |
| Lactococcus ffd9 | 0 | 0 | 0 | 0.003 |
| Unclassified Enterobacteriaceae 0315 | 2 | 1 | 2 | 0.002 |
| Unclassified Enterobacteriaceae 0942 | 0 | 0 | 1 | 0.0004 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sun, L.; Höjer, A.; Johansson, M.; Saedén, K.H.; Bernes, G.; Hetta, M.; Gustafsson, A.H.; Dicksved, J.; Lundh, Å. Associations between the Bacterial Composition of Farm Bulk Milk and the Microbiota in the Resulting Swedish Long-Ripened Cheese. Foods 2023, 12, 3796. https://doi.org/10.3390/foods12203796
Sun L, Höjer A, Johansson M, Saedén KH, Bernes G, Hetta M, Gustafsson AH, Dicksved J, Lundh Å. Associations between the Bacterial Composition of Farm Bulk Milk and the Microbiota in the Resulting Swedish Long-Ripened Cheese. Foods. 2023; 12(20):3796. https://doi.org/10.3390/foods12203796
Chicago/Turabian StyleSun, Li, Annika Höjer, Monika Johansson, Karin Hallin Saedén, Gun Bernes, Mårten Hetta, Anders H. Gustafsson, Johan Dicksved, and Åse Lundh. 2023. "Associations between the Bacterial Composition of Farm Bulk Milk and the Microbiota in the Resulting Swedish Long-Ripened Cheese" Foods 12, no. 20: 3796. https://doi.org/10.3390/foods12203796
APA StyleSun, L., Höjer, A., Johansson, M., Saedén, K. H., Bernes, G., Hetta, M., Gustafsson, A. H., Dicksved, J., & Lundh, Å. (2023). Associations between the Bacterial Composition of Farm Bulk Milk and the Microbiota in the Resulting Swedish Long-Ripened Cheese. Foods, 12(20), 3796. https://doi.org/10.3390/foods12203796






