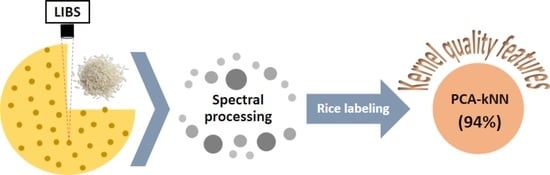
Rice Labeling according to Grain Quality Features Using Laser-Induced Breakdown Spectroscopy
Abstract
1. Introduction
2. Materials and Methods
2.1. Samples
2.2. LIBS Instrumentation and Measurements
2.3. Chemometric Data Analysis
2.3.1. Preprocessing
2.3.2. Variable Selection
2.3.3. Model Optimization and Validation
2.3.4. Classification Metrics and Software
3. Results and Discussion
4. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Qiu, Z.; Chen, J.; Zhao, Y.; Zhu, S.; He, Y.; Zhang, C. Variety identification of single rice seed using hyperspectral imaging combined with convolutional neural network. Appl. Sci. 2018, 8, 212. [Google Scholar] [CrossRef]
- Olivares Díaz, E.; Kawamura, S.; Matsuo, M.; Kato, M.; Koseki, S. Combined analysis of near-infrared spectra, colour, and physicochemical information of brown rice to develop accurate calibration models for determining amylose content. Food Chem. 2019, 286, 297–306. [Google Scholar] [CrossRef] [PubMed]
- Custodio, M.C.; Cuevas, R.P.; Ynion, J.; Laborte, A.G.; Velasco, M.L.; Demont, M. Rice quality: How is it defined by consumers, industry, food scientists, and geneticists? Trends Food Sci. Technol. 2019, 92, 122–137. [Google Scholar] [CrossRef] [PubMed]
- Bao, J. Genes and QTLs for Rice Grain Quality Improvement. In Rice–Germplasm, Genetics and Improvement; InTech–Open Science Open Mind: London, UK, 2014; pp. 239–278. [Google Scholar]
- Biselli, C.; Bagnaresi, P.; Cavalluzzo, D.; Urso, S.; Desiderio, F.; Orasen, G.; Gianinetti, A.; Righettini, F.; Gennaro, M.; Perrini, R.; et al. Deep sequencing transcriptional fingerprinting of rice kernels for dissecting grain quality traits. BMC Genom. 2015, 16, 1–28. [Google Scholar] [CrossRef] [PubMed]
- BRASIL, Ministério da Agricultura, Pecuária e Abastecimento (MAPA). Instrução Normativa no 06, de 16 de fevereiro de 2009; Diário Oficial [da] República Federativa do Brasil: Brasília, DF, 2009; 17 fev. Seção 1. [Google Scholar]
- Joint FAO/WHO Codex Alimentarius Commission; World Health Organization; Joint FAO/WHO Food Standards Programme. Cereals, Pulses, Legumes and Vegetable Proteins; Food & Agriculture Organization: Rome, Italy, 2007. [Google Scholar]
- Elmasry, G.; Mandour, N.; Al-Rejaie, S.; Belin, E.; Rousseau, D. Recent applications of multispectral imaging in seed phenotyping and quality monitoring—An overview. Sensors 2019, 19, 1090. [Google Scholar] [CrossRef]
- Marschalek, R.; Silva, M.C.; dos Santos, S.B.; Manke, J.R.; Bieging, C.; Porto, G.; Wickert, E.; de Andrade, A. Image–Rice Grain Scanner: A three-dimensional fully automated assessment of grain size and quality traits. Crop Breed. Appl. Biotechnol. 2017, 17, 89–97. [Google Scholar] [CrossRef]
- Courtois, F.; Faessel, M.; Bonazzi, C. Assessing breakage and cracks of parboiled rice kernels by image analysis techniques. Food Control. 2010, 21, 567–572. [Google Scholar] [CrossRef]
- Cheng, F.; Ying, Y.B.; Li, Y.B. Detection of defects in rice seeds using machine vision. Trans ASABE 2006, 49, 1929–1934. [Google Scholar] [CrossRef]
- Miziolek, A.W.; Palleschi, V.; Schechter, I. Laser-Induced Breakdown Spectroscopy (LIBS)–Fundamentals and Applications; Cambridge University Press: Cambridge, UK, 2006; pp. 1–5. [Google Scholar]
- Cremers, D.A.; Radziemski, L.J. Handbook of Laser-Induced Breakdown Spectroscopy; John Wiley & Sons: Hoboken, NJ, USA, 2006; pp. 1–283. [Google Scholar]
- Sezer, B.; Durna, S.; Bilge, G.; Berkkan, A.; Yetisemiyen, A. Identi fi cation of milk fraud using laser-induced breakdown spectroscopy (LIBS). Int. Dairy J. 2018, 81, 1–7. [Google Scholar] [CrossRef]
- Yang, P.; Zhou, R.; Zhang, W.; Tang, S.; Hao, Z.; Li, X.; Lu, Y.; Zeng, X. Laser-induced breakdown spectroscopy assisted chemometric methods for rice geographic origin classification. Appl. Opt. 2018, 57, 8297. [Google Scholar] [CrossRef]
- Zhu, L.; Sun, J.; Wu, G.; Wang, Y.; Zhang, H.; Wang, L. Identi fi cation of rice varieties and determination of their geographical origin in China using Raman spectroscopy. J. Cereal Sci. 2018, 82, 175–182. [Google Scholar] [CrossRef]
- Larios, G.S.; Nicolodelli, G.; Senesi, G.S.; Ribeiro, M.C.S.; Xavier, A.A.P.; Milori, D.M.B.P.; Alves, C.Z.; Marangoni, B.S.; Cena, C. Laser-Induced Breakdown Spectroscopy as a Powerful Tool for Distinguishing High- and Low-Vigor Soybean Seed Lots. Food Anal. Methods 2020, 13, 1691–1698. [Google Scholar] [CrossRef]
- Silva, T.V.; Hubinger, S.Z.; Gomes Neto, J.A.; Milori, D.M.B.P.; Ferreira, E.J.; Ferreira, E.C. Potential of Laser Induced Breakdown Spectroscopy for analyzing the quality of unroasted and ground coffee. Spectrochim. Acta–Part B At. Spectrosc. 2017, 135, 29–33. [Google Scholar] [CrossRef]
- Chu, Y.W.; Tang, S.S.; Ma, S.X.; Ma, Y.Y.; Hao, Z.Q.; Guo, Y.M.; Guo, L.B.; Lu, Y.F.; Zeng, X.Y. Accuracy and stability improvement for meat species identification using multiplicative scatter correction and laser-induced breakdown spectroscopy. Opt. Express 2018, 26, 10119. [Google Scholar] [CrossRef]
- Nespeca, M.G.; Vieira, A.L.; Júnior, D.S.; Neto, J.A.G.; Ferreira, E.C. Detection and quantification of adulterants in honey by LIBS. Food Chem. 2020, 311, 125886. [Google Scholar] [CrossRef]
- Yang, P.; Zhu, Y.; Yang, X.; Li, J.; Tang, S.; Hao, Z.; Guo, L.; Li, X.; Zeng, X.; Lu, Y. Evaluation of sample preparation methods for rice geographic origin classification using laser-induced breakdown spectroscopy. J. Cereal Sci. 2018, 80, 111–118. [Google Scholar] [CrossRef]
- Pérez-Rodríguez, M.; Dirchwolf, P.M.; Silva, T.V.; Villafañe, R.N.; Neto, J.A.G.; Pellerano, R.G.; Ferreira, E.C. Brown rice authenticity evaluation by spark discharge-laser-induced breakdown spectroscopy. Food Chem. 2019, 297, 124960. [Google Scholar] [CrossRef]
- Luo, Z.; Zang, L.; Chen, T.; Liu, M.; Chen, J.; Hou, H.; Yao, M. Rapid identification of rice species by laser-induced breakdown spectroscopy combined with pattern recognition. Appl. Opt. 2019, 58, 1631–1638. [Google Scholar] [CrossRef]
- Pérez-Rodríguez, M.; Dirchwolf, P.M.; Silva, T.V.; Vieira, A.L.; Neto, J.A.G.; Pellerano, R.G.; Ferreira, E.C. Fast spark discharge-laser-induced breakdown spectroscopy method for rice botanic origin determination. Food Chem. 2020, 331, 127051. [Google Scholar] [CrossRef]
- Vieira, A.L.; Silva, T.V.; de Sousa, F.S.I.; Senesi, G.S.; Júnior, D.S.; Ferreira, E.C.; Neto, J.A.G. Determinations of phosphorus in fertilizers by spark discharge-assisted laser-induced breakdown spectroscopy. Microchem. J. 2018, 139, 322–326. [Google Scholar] [CrossRef]
- Bro, R.; Smilde, A.K. Principal component analysis. Anal. Methods 2014, 6, 2812–2831. [Google Scholar] [CrossRef]
- Lantz, B. Machine Learning with R; Packt Publishing: Birmingham, UK, 2015. [Google Scholar]
- Mchugh, M.L. Lessons in biostatistics Interrater reliability: The kappa statistic. Biochem. Medica 2012, 22, 276–282. [Google Scholar] [CrossRef]
- Team, R.C.; R: A Language and Environment for Statistical Computing. The R Foundation - The R Project for Statistical Computing. 2020. Available online: http://www.R-project.org/ (accessed on 25 August 2022).
- Hastie, T.; Tibshirani, R.; Friedman, J. The Elements of Statistical Learning: Data Mining, Inference, and Prediction; Springer: New York, NY, USA, 2009; Volume 2. [Google Scholar]
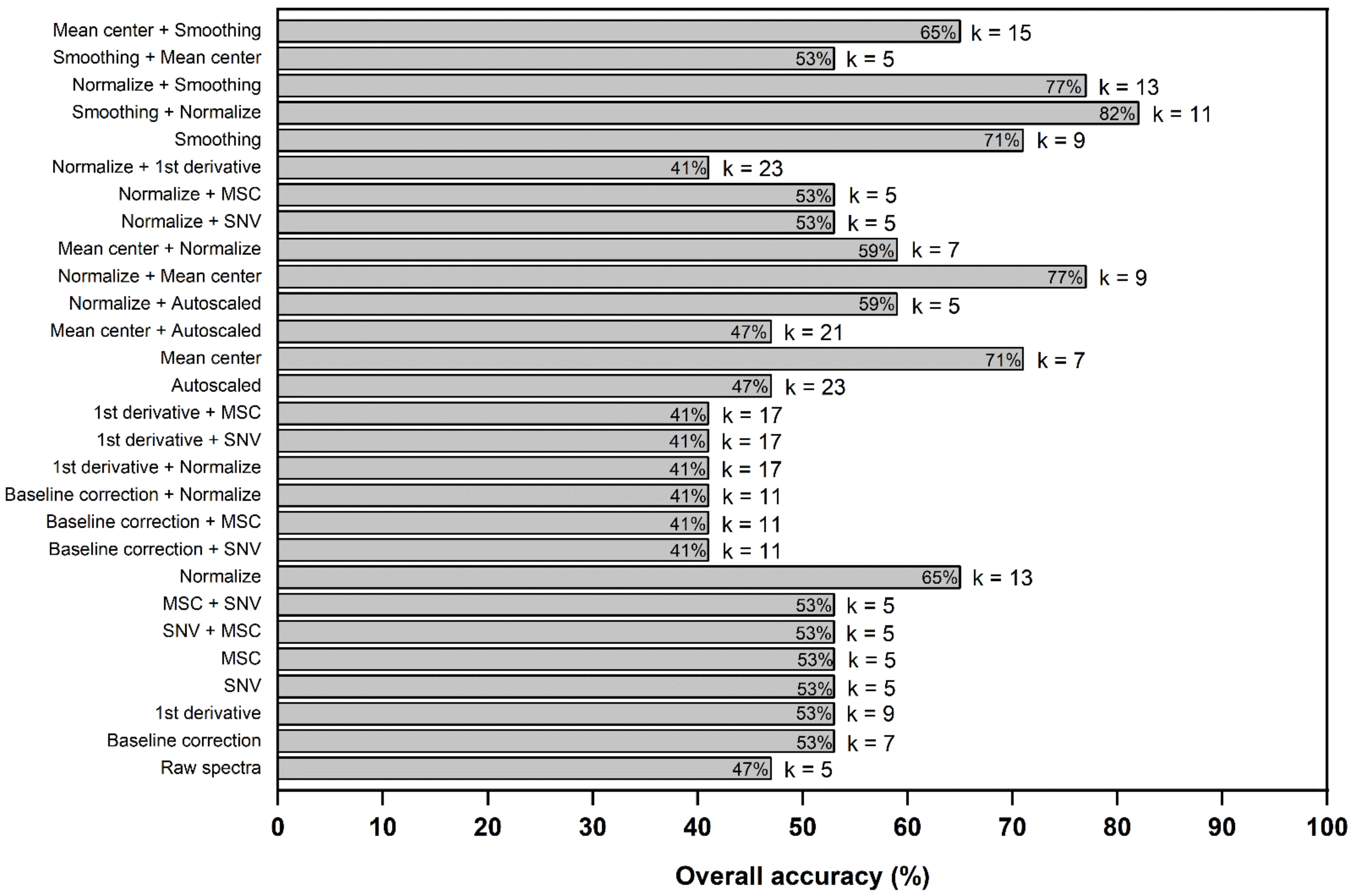

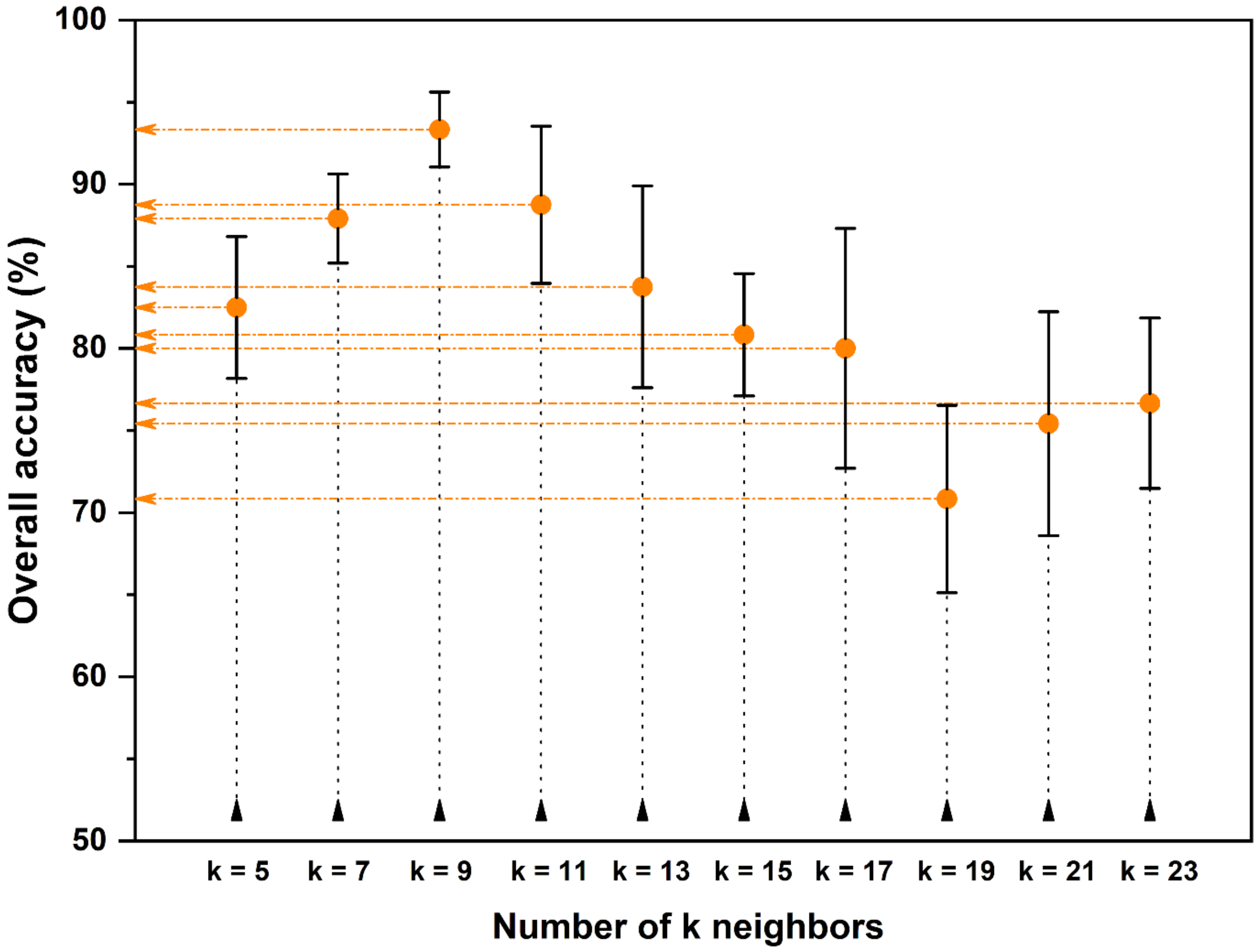
| Quality Type | General Defects (%) * | |||||
|---|---|---|---|---|---|---|
| Moldy and Sour | Chopped or Stained | Plastered or Green | Brindle | Yellow | Broken | |
| 1 | 0.15 | 1.75 | 2.00 | 1.00 | 0.50 | 7.00 |
| 2 | 0.30 | 3.00 | 4.00 | 1.50 | 1.00 | 14.00 |
| 3 | 0.50 | 4.50 | 6.00 | 2.00 | 2.00 | 23.00 |
| Element * | Registered Spectral Lines (nm) |
|---|---|
| C II | 426.7 |
| Ca I | 397.4 |
| Ca II | 393.4 |
| CN | 387.1, 388.3 |
| Fe I | 302.3, 304.6, 305.9, 326.0, 326.5, 331.9, 335.3, 336.7, 337.4, 339.0, 340.7, 343.7, 347.1, 371.2, 372.7, 375.4, 375.9, 377.0, 383.9, 395.6, 396.9, 397.4, 407.6, 409.8, 410.4, 426.7, 435.0, 444.7, 459.2, 459.6, 460.2, 464.3, 466.2, 470.0, 470.6, 478.9, 553.4, 568.6, 571.0 |
| Fe II | 371.2, 375.9, 441.6 |
| Mg I | 383.2, 383.9, 518.3 |
| N II | 332.9, 399.6, 460.8, 461.5, 462.2, 463.1, 464.3, 478.1, 480.4, 499.5, 504.6, 549.5, 553.4, 566.6, 567.9, 574.7, 593.2, 594.1 |
| Na I | 589.0, 589.5 |
| O II | 313.5, 374.9, 412.0 |
| Grain Quality | Sensitivity (%) | Specificity (%) | Accuracy (%) |
|---|---|---|---|
| Type 1 | 100 | 100 | 100 |
| Type 2 | 83 | 100 | 92 |
| Type 3 | 100 | 90 | 95 |
| Overall | 94 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pérez-Rodríguez, M.; Mendoza, A.; González, L.T.; Lima Vieira, A.; Pellerano, R.G.; Gomes Neto, J.A.; Ferreira, E.C. Rice Labeling according to Grain Quality Features Using Laser-Induced Breakdown Spectroscopy. Foods 2023, 12, 365. https://doi.org/10.3390/foods12020365
Pérez-Rodríguez M, Mendoza A, González LT, Lima Vieira A, Pellerano RG, Gomes Neto JA, Ferreira EC. Rice Labeling according to Grain Quality Features Using Laser-Induced Breakdown Spectroscopy. Foods. 2023; 12(2):365. https://doi.org/10.3390/foods12020365
Chicago/Turabian StylePérez-Rodríguez, Michael, Alberto Mendoza, Lucy T. González, Alan Lima Vieira, Roberto Gerardo Pellerano, José Anchieta Gomes Neto, and Edilene Cristina Ferreira. 2023. "Rice Labeling according to Grain Quality Features Using Laser-Induced Breakdown Spectroscopy" Foods 12, no. 2: 365. https://doi.org/10.3390/foods12020365
APA StylePérez-Rodríguez, M., Mendoza, A., González, L. T., Lima Vieira, A., Pellerano, R. G., Gomes Neto, J. A., & Ferreira, E. C. (2023). Rice Labeling according to Grain Quality Features Using Laser-Induced Breakdown Spectroscopy. Foods, 12(2), 365. https://doi.org/10.3390/foods12020365