Abstract
In recent years, several alimentary diseases have been connected with the consumption or tasting of raw flour and dough. Microbiological quality concern is also raising due to increased consumer demand for plant powders, while some of them can be consumed without prior thermal processing. In this study, we have focused on the occurrence of antibiotic-resistant coliform bacteria and enterococci in flour, plant powder and dough from Slovak retail. Our results indicated the presence of both total and antibiotic-resistant coliform bacteria and enterococci in the flour and powder samples. Lower numbers of the total, as well as resistant bacteria, were detected in flours compared to plant powders. Coliform bacteria isolates were predominantly identified as Klebsiella spp. and Enterobacter spp. Ampicillin resistance appeared in 97% of isolates followed by chloramphenicol resistance (22%) and tetracycline resistance (17%). The presence of the blaSHV gene was confirmed in 13% of isolates. The tetA and tetE genes were present in 25% of isolates of coliform bacteria. The presence of enterococci was detected only in plant powders. Antibiotic-resistant strains were identified as the following: Enterococcus casseliflavus, E. gallinarium and E. faecium. Despite the isolates showing resistance to vancomycin, the presence of the vanA gene was not detected. The majority of antibiotic-resistant isolates belonged to the group of medium biofilm producers. None of these isolates showed efflux pump overproduction. Antibiotic-resistant coliform bacteria and enterococci were not detected in the processed doughs.
1. Introduction
Flour is classified as a food with low water activity, which ensures the growth inhibition of most microorganisms [1,2]. Nevertheless, it can be contaminated with pathogens, some of which are able to maintain viability long enough to cause disease [1,3]. For example, studies have shown that Escherichia coli and Salmonella sp. can survive 12 to 16 weeks in flour stored at room temperature [1]. The flour quality and safety are greatly affected by the initial microbiological quality of the cereal grains. A comparison of the final milling products of high and low microbiological quality wheat confirmed that flour with a higher number of microorganisms is obtained from lower microbiological quality wheat [4]. Conditioning is the process of adding water to wheat grains before milling to strengthen the bran and soften the endosperm. However, a long time (6–48 h), appropriate temperature (24–25 °C) and humidity during this process promote the growth and reproduction of the microorganisms’ present [5]. No significant increase in the total number of aerobic microorganisms was observed in the experimental system, but after conditioning, the presence of bacteria that were not previously detectable was detected, and a significant increase in E. coli was also demonstrated. This suggests selective enrichment with bacteria that were below the detection limit prior to conditioning or their recovery from a non-culturable but non-infectious state [6]. At the same time, there is a risk of cross-contamination by microorganisms potentially inhabiting the grinding equipment. Equipment used to purify and condition wheat has been shown to contain populations of aerobic mesophilic bacteria, Enterobacterales, and micromycetes [7]. As none of the operations in the flour recovery process involve chemical or thermal treatment to ensure safety, it can be expected that microorganisms originally present on cereal grains, including the possible presence of pathogenic microorganisms, will also be present in reduced quantities in ground cereal grains [4].
The Lower Saxon State Office for Consumer Protection and Food Safety carries out regular monitoring of the microbiological quality of flour. In 2017, shiga-toxigenic E. coli (STEC) was isolated from five retail flour samples [8]. As part of the Federal Monitoring Plan, the presence of STEC was confirmed in Germany in 2018 in 50 of the 328 samples of wheat, spelled and rye flour tested. A wide variety of STEC serotypes were detected in the samples, including human pathogenic types with different combinations of pathogenicity factors [6]. The presence of STEC was detected in 10.8% of 93 flour samples of different cereal varieties (wheat and mixtures of wheat with other cereal flour) purchased at retail markets in Switzerland [6,9].
In 2008, in New Zealand, 66 people suffered from salmonellosis due to consumption of an uncooked baking mixture containing flour contaminated with S. Typhimurium (STM42). This was the first time that contaminated flour had been identified by laboratory and epidemiological research as a source of Salmonella [10,11,12]. The first outbreak of flour-related STEC occurred in the United States in 2016. In 24 states, 63 patients were infected with STEC O121 or STEC O26 strains [3,13]. In 2017, there were 29 cases of the STEC disease in six provinces in Canada. This was the first nationwide E. coli epidemic associated with flour consumption in Canada [11,14]. These flour related epidemics point to the consequences of risky consumer behavior, in particular to the health risks associated with the consumption of raw products that are intended for heat treatment before consumption [12].
Lifestyle changes, together with the increasing incidence of celiac disease, have caused increased consumer demand for gluten-free products [15]. The market for gluten-free products is growing significantly as better diagnostic procedures make it possible to identify an increasing number of people with celiac disease and other disorders related to gluten sensitivity, such as dermatitis herpetiformis, gluten ataxia, wheat allergy and non-celiac gluten sensitivity [16]. In 2019, 22,735 patients with celiac disease were registered in Slovakia [17]. The basic treatment for these types of disorders is to follow a strict and permanent gluten-free diet. While patients suffering from gluten-related diseases rely on a gluten-free diet, other consumers choose it as a lifestyle because it is of cultural, environmental, civic, historical, ethnic or health concern. All these factors support the gluten-free market and its continued growth [16]. The current market provides several alternatives to flour (such as plant or insect powders) that can replace traditional wheat flour. Some of these flours can be consumed without previous heat treatment, for example, by adding to smoothie drinks, yogurts, cocktails, etc. [18]. However, if these products are contaminated with pathogenic microorganisms, they may pose a serious risk to human health.
According to these facts, we have decided to evaluate the microbiological quality of flours and powders as well as processed doughs available in Slovak retail. Since antibiotic resistance is a global emerging problem, we have focused on the detection and characterization of antibiotic-resistant coliform bacteria and enterococci in these foodstuffs used in common cuisine.
2. Materials and Methods
2.1. Sample Collection
In total, 60 samples of flours and powders, 29 of cereal flours and 31 of plant and insect powders were collected during 2019 and 2021 in Slovakia, predominantly Bratislava and subjected to microbial analyses. The abbreviation of each sample used in the paper and details of analyzed flours or powders are summarized in Table 1. Samples of 13 cooled doughs were also subjected to microbial analyses, which were obtained from markets in Bratislava, Slovakia. The abbreviation of each sample and details of analyzed doughs are summarized in Table 2. All tested samples were subjected to microbial analyses immediately after arriving at the laboratory. Samples of dough were retained at a cooling temperature of 8 °C during the time of transport to the laboratory.

Table 1.
Details and types of flours and powders subjected for microbial analyses.

Table 2.
Details and types of dough subjected for microbial analyses.
2.2. Microbiological Analysis
Ninety mL of sterile peptone water solution (0.85% NaCl and 0.1% peptone) was homogenized with 10 g of sample of powder or flour or cooled dough [19,20,21]. Total aerobic bacteria, total and antibiotic-resistant coliform bacteria and enterococci were enumerated using the plate count agar (Biolife Italiana srl., Milan, Italy), Chromocult-coliform agar (VWR, Darmstadt, Germany) and Slanetz-Bartley agar (BioLife Italiana srl., Milan, Italy), respectively. Antibiotic-resistant colonies of coliform bacteria and enterococci were monitored by adding specific concentrations to the agar. Different concentrations were chosen according to resistance limits established by the European Committee on Antimicrobial Susceptibility Testing and in the case of antibiotic tetracycline according to Clinical and Laboratory Standard Institute [22,23]. To detect antibiotic-resistant coliform bacteria, antibiotics were used: ampicillin, gentamicin, chloramphenicol, ciprofloxacin, and tetracycline (Sigma-Aldrich, St. Louis, MO, USA). To detect antibiotic-resistant enterococci, antibiotics were used: ampicillin, gentamicin, ciprofloxacin, and vancomycin (Sigma-Aldrich, St. Louis, MO, USA). Agar plates with plate count agar were cultivated aerobically at 30 °C for 72 h. Chromocult coliform agars were cultivated aerobically at 37 °C for 24 h. Agar plates with Slanetz-Bartley agar were cultivated aerobically at 40 °C for 48 h. The results were expressed as the mean of all the repetitions together with the standard deviation (SD).
2.3. Identification of Antibiotic-Resistant Strains
The identification of antibiotic-resistant bacterial strains was performed by matrix-assisted laser desorption ionization-time of flight (MALDI-TOF) mass spectrometry (Bruker, Germany). After the monitoring of resistant coliform bacteria, colonies were randomly picked and isolated by the streak plate method on Mueller Hinton agar (Biolife, Italy) plates for 24 h at 37 °C (enterococci for 48 h at 40 °C). A sample of each strain (one colony) was applied on a steel target plate and covered with 1 μL of the matrix (saturated solution of α-cyano-4-hydroxycinnamic acid in 2.5% trifluoroacetic acid and 50% acetonitrile) and left to air-dry. The target plate was inserted into MALDI-TOF mass spectrometry, and analysis was performed via an AutoFlex I TOF-TOF apparatus (Bruker Daltonics Inc., Billerica, MA, US) in linear positive-ion mode (m/z range of 2000 to 20,000 with gating of ions below m/z 400 and a delayed extraction time of 450 ns). Gaining spectra were analyzed using MALDI BioTyper software (v 2.0) based on an algorithm for matching spectral patterns in logarithmic scores 0–3 (BioTyper Library v 3.0; Bruker Daltonics s.r.o., Brno, Czech Republic). A score above 1.9 ensured bacterial species identification by comparison of the obtained bacterial fingerprints with the existing database [24,25].
2.4. Susceptibility Testing of Resistant Isolates
The susceptibility of bacterial isolates to different antibiotics was detected with the macro-dilution drop method. Several ATBs of different classes were used for testing coliform bacteria (ampicillin, ceftazidime, ciprofloxacin, tetracycline, gentamicin, chloramphenicol, and meropenem). For enterococci, antibiotics ampicillin, gentamicin, ciprofloxacin, and vancomycin were used. The concentration of each ATB is listed in Table 3. Antibiotic concentrations were defined by resistant breakpoints according to EUCAST (R1) and according to CLSI (R2) [22,23]. The third concentration was selected to determine the highest level of resistance in coliform bacteria isolates as described previously by Krahulcová et al. [25]. The experiment was performed using Mueller-Hinton agar (Biolife, Italy). Incubation of plates was at 37 °C for 24 h for coliforms and 40 °C for 48 h for enterococci. The susceptibility profiles after incubation were evaluated visually [24,25]. Each experiment ran in triplicates and was repeated three times. For statistical analysis, the Student’s t-test was applied.

Table 3.
Resistant breakpoints of ATBs used in susceptibility testing.
2.5. Biofilm Production Testing
Biofilm production assessment was performed according to Beenken et al. (2003) [26]. A sterile 96-well microtiter plate was inoculated with diluted overnight cultures of tested isolates (1:200 in tryptic soy broth). Incubation was static at 37 °C, 24 h for coliforms, and 40 °C, 48 h for enterococci. After incubation the overnight cultures were removed, and the wells of the microtiter plate were washed twice with 200 μL of PBS. The forming biofilm was fixed with 200 μL of 96% ethanol for few seconds and the microtiter plate was left to air-dry. Bacterial cells forming biofilm in the wells were stained with crystal violet solution (0.41% in 12% ethanol) for 3 min. After staining, wells were washed twice with PBS. As the final step, 200 µL of 96% ethanol was added to each well to dissolve the absorbed crystal violet in biofilm. The absorbance of crystal violet in each well was measured at 570 nm using a plate reader device (BioTek Inc., Seattle, WA, USA). Each experiment was repeated three times and ran in six parallels. For statistical analysis, the Student’s t-test was applied. The positive control strain was Pseudomonas aeruginosa (CCM 3955) considered strong producer of biofilm. Biofilm producers were then divided into 4 groups according to the values of measured absorbances: weak (<0.2), medium (0.2–0.3), strong (0.3–0.9), and very strong (>1.0) biofilm producers as described in Taniguchi et al. (2009) [24,25,27].
2.6. Efflux Pumps Overproduction Testing with Ethidium Bromide (EtBr)
The detection of efflux pump overproduction was evaluated via the EtBr-agar Carthweel method [28]. Detection was performed on Mueller-Hinton agar plates supplemented with ethidium bromide (2.5 mg/L). Inoculums in a density of 0.5 McFarland standard were prepared from overnight cultures and applied on marked EtBr plates with a swab. Incubation of inoculated EtBr-plates was at 37 °C for 16 h (coliforms) or 40 °C for 48 h (enterococci). The experiment was based on visual evaluation with UV irradiation due to fluorescence active compound EtBr (Sigma-Aldrich, St. Louis, MO, USA) [25,29]. The reference strain used as a comparative negative control for coliforms was E. coli (CCM 3988) and for enterococci Staphylococcus epidermidis (CCM 3953). Each experiment ran in triplicates and was repeated three times. For statistical analysis Student’s t-test was applied.
2.7. Detection of Antibiotic Resistance Genes
Resistance genes were detected via single and multiplex polymerase chain reaction (PCR). The presence of several resistance genes was detected in the bacterial isolates: β-lactamases TEM, SHV, OXA [30], tetracycline resistance genes Group II: tetA, tetE in coliforms [31] and vancomycin resistance gene vanA in enterococci [32]. The reaction mixture for multiplex PCR for detection of β-lactamases and tet genes consisted of 0,5 μL of each primer (0.25 μL for tet genes) and PCR water in a total volume of 25 μL. The reaction mixture for vanA PCR consisted of 0.25 μL of both primers, 10 μL of PCR buffer, 1 μL of deoxyribonucleotides mixture, 2 μL of 50 mmol MgCl2 solution and PCR water in total volume of 50 μL. A single colony was used as a DNA sample of tested isolates. Primers used during each PCR are listed in Table 4. PCR was carried out in a thermocycler (Mastercycler personal Eppendorf, Hamburg, Germany) using the following conditions for amplifying specific sections in β-lactamases: initial denaturation at 94 °C for 20 min; 30 cycles of denaturation at 94 °C for 40 s, annealing temperature 60.5°C for 1 min, 72 °C for 1 min and 30 s, and the final elongation step at 72 °C for 10 min. The same protocol was applied for the detection of vanA gene, except for an annealing temperature of 65 °C. The conditions used to determine tetracycline genes were initial denaturation at 94 °C for 20 min, 35 cycles of denaturation at 94 °C for 40 s, annealing temperature at 55 °C for 1 min, 72 °C for 1 min 30 s, and the final elongation step at 72 °C for 10 min. After initial denaturation, 25 μL of Multiplex PCR Master mix (Biotechrabbit, Berlin, Germany) for multiplex PCR or 0.5 μL of Hot Start Taq DNA Polymerase (Biotechrabbit, Berlin, Germany) for single PCR was added to the mixture [25,29].

Table 4.
Resistant genes detected during PCR in resistant coliform isolates.
PCR products were dyed with Loading Dye (Biotechrabbit, Berlin, Germany) and separated by gel electrophoresis (1.5% agarose gel in TAE buffer) set up at 100 V for 1 h and 40 min. Bands were visualized by subsequent staining in Gel Red (Biotium, Fremont, CA, USA) solution in TAE for 30 min and 100 bp DNA Ladder was used as a size marker. Positive controls used during the PCR reaction were subjected to sequence analyses to prove the presence of specific resistance genes.
3. Results and Discussion
3.1. Total Coliform Bacteria and Enterococci in Flour, Powder and Dough
Twenty-nine flour samples and 31 samples of plant or insect powders and 13 samples of dough were subjected to microbiological analysis. In 87% of samples of flour and powder were determined the total number of aerobic bacteria (Table 5). Fecal indicators (coliform bacteria or enterococci) were detected in 53% of samples of flour and powder. A total of 85% of samples of dough were determined total aerobic (Table 6). Coliform bacteria and enterococci were presented in 38% of all samples of dough, where total enterococci were observed only in one sample, namely sample of strudel dough in number of 4.1 log CFU/g.

Table 5.
Number of total aerobic and indicator bacteria in samples of flour and powder.

Table 6.
Number of total aerobic bacteria, coliform bacteria, and enterococci in samples of dough.
The majority of detected numbers of total coliform bacteria (TCB) has ranged between 2.3 and 4.1 log CFU/g. These counts are comparable to Batool et al. (2012), who determined coliform bacteria counts in Pakistani flour in the range of 3.0–4.0 log CFU/g [33]. Twelve powder samples contained coliforms ranging from 2.3 to 5.3 log CFU/g. We recorded the most numerous growths in the case of chestnut powder (M3) originating from Italy. The microbiological quality of flour and other mill products is defined in the second part of the Codex Alimentarius of the Slovak Republic, which regulates the microbial requirements for food and packaging. This section sets out the requirements for coliform bacteria, fungi, and Bacillus cereus. As for coliform bacteria, these can be found in flours in the number of 103, the limit value being 104 in two samples out of five examined samples [34]. Microbiological criteria for powders are not currently available. Four samples of pizza and puff dough contained coliform bacteria (Table 6). In March 2022, the consumption of frozen pizza was connected to the E. coli outbreak in France [35]. Escherichia coli was detected only in one sample of pumpkin powder.
Compared to coliform bacteria, the number of samples with enterococci was much lower. We detected enterococci only in six samples of powder and one sample of strudel pastry. In cricket powder, we observed the highest count of enterococci. The genus Enterococcus spp. is part of the healthy intestinal microbiota of humans and animals, and some strains are used as starter cultures or probiotics. Due to their preferred intestinal habitat, their wide occurrence and ease of cultivation serve as indicators of fecal contamination and are part of hygienic standards for water and food products [36,37].
3.2. Antibiotic Resistant Coliform Bacteria and Enterococci in Flour, Powder and Dough
Besides TCB and TEC (total enterococci), we also detected their possible antibiotic resistance. When quantifying resistant coliform bacteria and enterococci, we applied the concentrations of individual antibiotics in accordance with EUCAST guidelines. In case of coliform bacteria, we applied the antibiotics ampicillin (penicillin), gentamicin (aminoglycoside), chloramphenicol (amphenicol), ciprofloxacin (fluoroquinolone), and tetracycline (tetracycline), which represent different classes of antibiotics. For tetracycline, we used the concentration specified by CLSI guidelines, while there are no breakpoints given by EUCAST. The antibiotics ampicillin (penicillin), gentamicin (aminoglycoside), ciprofloxacin (fluoroquinolone), and vancomycin (glycopeptide) were applied in the determination of resistant enterococci.
In 45% of flour samples, we detected antibiotic-resistant coliforms (Table 7). The majority was observed only in the case of ampicillin. Only in one sample of rye flour did we also observe gentamicin-resistant coliforms. In 39% of powder samples, we observed ampicillin resistance. We recorded the highest number in a sample of chestnut powder (M3). Significantly higher resistance to ampicillin was expected because most coliforms have intrinsic resistance to this antibiotic, except for the genus Escherichia. In five powder samples, we detected gentamicin resistance in the range of 2.7–4.3 log CFU/g. We recorded chloramphenicol and tetracycline resistance only in chestnut powder (M3). We did not detect resistance to ciprofloxacin in any sample. We have observed ampicillin-resistant enterococci only in six powder samples. In samples of chestnut (M3) and coconut powder (M19, M30), we also detected vancomycin-resistant enterococci. The only sample of insect flour-cricket flour (M49) contained ampicillin (penicillins) and gentamicin (aminoglycosides)-resistant coliform bacteria. Ampicillin-resistant enterococci were also present in this flour sample. In dough samples, we did not register any antibiotic-resistant bacteria.

Table 7.
Occurrence of antibiotic-resistant coliform bacteria and enterococci in samples of flour and powder.
The presence of total and antibiotic-resistant coliforms and enterococci in powders from chickpeas, rice, banana, almond, and coconut is of considerable concern, while these powders can be used in cold cuisine as an addition to drinks, spreads, salads, yogurt, smoothies, etc. [18].
In the United States in 2018, voluntary recalls of approximately 6300 packages of organic coconut powder were reported, due to the possible presence of Salmonella, which was detected in one package during routine testing. In the same year, Salmonella was the cause of the further withdrawal of flour from the market, namely 2099 packages of organic amaranth powder. Fortunately, none of these withdrawals have reported cases of disease due to flour consumption [38,39].
3.3. Identification of Antibiotic-Resistant Strains
Isolated colonies of antibiotic-resistant coliforms and enterococci were identified using a MALDI-TOF mass spectrometer, which can be used to analyze biological macromolecules by determining the weight of molecules after mixing with a matrix and laser beam ionization. Identification of microorganisms is performed by comparing the obtained mass spectrum with the spectra of reference strains in the relevant database. This method provides fast, well reproducible and accurate identification of microorganisms at the genus and species level [40].
In total, we identified 32 resistant coliform isolates belonging to six genera (Figure 1).
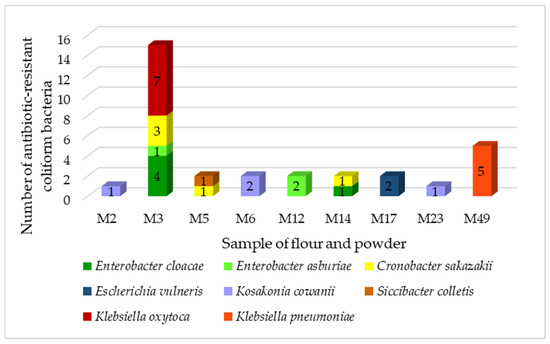
Figure 1.
Antibiotic-resistant coliform bacteria isolated from flour, powder. M2—Teff, M3—Chestnut, M5—Amaranth, M6—Rice wholegrain, M12—Banana, M14—Chestnut, M17—Sorghum, M23—Rye, M49—Cricket.
The majority was from the Klebsiella family (37%). Bacteria of the genus Klebsiella are widespread in nature. They are associated with several pathological processes in humans and animals but can also enter the food chain. Klebsiella is not considered to be a major intestinal or foodborne pathogen, but its participation in intestinal diseases cannot be ruled out. Multidrug-resistant Klebsiella spp. producing broad-spectrum β-lactamases that can be transported through food [41]. We identified two species of the genus Klebsiella, K. oxytoca and K. pneumoniae. These two species are the most common pathogens causing nosocomial infections in humans and are of great concern regarding the development of resistance to multiple antibiotics [42]. Klebsiella pneumoniae is a common cause of community-acquired and nosocomial infections, such as urinary tract infections, lower respiratory tract infections, and liver abscesses. Although not traditionally considered a foodborne pathogen, there have been reports of K. pneumoniae infections preceded by intestinal colonization, supporting the theory of food as a possible vector for the transmission of these pathogens [43]. All strains of K. pneumoniae were isolated from cricket powder.
Strains of Cronobacter sakazakii were isolated from chestnut and amaranth powder. Cronobacter sakazakii is an opportunistic pathogen associated with life-threatening neonatal infections that can lead to severe manifestations such as brain abscesses, meningitis, necrotizing enterocolitis, and systemic sepsis. Most cases are associated with the consumption of powdered infant formula contaminated with these microorganisms [44].
Kosakonia cowanii was isolated from teff and rye flour as well as from rice powder. Kosakonua cowanii is a recently reclassified bacterial species formerly known as Enterobacter cowanii. The bacterium is thought to be primarily a plant pathogen, but cases of human infections have been reported in which K. cowanii has been identified as the cause of rhabdomyolysis and bacteremia associated with rose thorns and as the cause of acute cholecystitis [45,46]. Up to 63% of Enterobacter were isolated from chestnut powder. These bacteria are ubiquitous and have been found in a wide variety of foods, including fruits and vegetables, meat, fish, eggs, tea, herbs, legumes, spices, dry animal feed, dairy products, powdered baby food, cereals, nuts, seeds, flour, pasta, chocolate, drinks, and water. Enterobacter cloacae is the most isolated species of the genus Enterobacter from humans and animals. It is referred to as an opportunistic pathogen identified in nosocomial infections [47]. Escherichia vulneris and Siccibacter colletis were isolated from chickpea powder. Enterobacter vulneris is an opportunistic human pathogen that has been identified in clinical cases of urosepsis, osteomyelitis, intravenous catheter-related bacteremia, meningitis, and dialysis-related peritonitis [48].
We have isolated and identified 10 antibiotic-resistant enterococci. The majority belonged to E. casseliflavus (70%), followed by E.gallinarium (20%) and E. faecium (10%) (Figure 2). Enterococcus faecium is the second most common species of Enterococcus, which is the cause of nosocomial diseases. It often possesses vancomycin resistance genes, making it a priority pathogen for the development of new antibiotics [49]. Most enterococci were isolated from coconut powder, only one isolate was derived from amaranth powder.
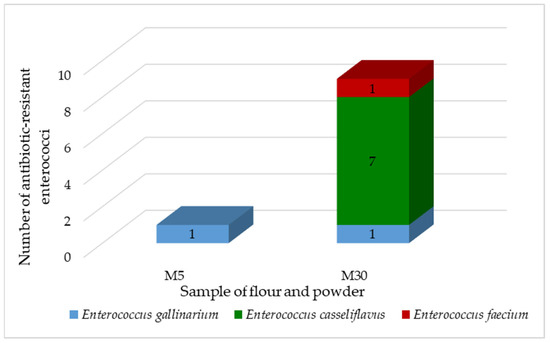
Figure 2.
Antibiotic-resistant enterococci isolated from flour, powder. M5—Amarant, M30—Coconut.
3.4. Characterization of Antibiotic-Resistant Strains
In almost all isolates we have detected ampicillin resistance (Table 8). Only one isolate of K. cowani from rye flour showed susceptibility to this antibiotic. The majority of coliform bacteria are intrinsically resistant to ampicillin, in the case of the genus Escherichia it is an acquired mechanism of resistance [24]. We did not observe resistance to gentamicin, ciprofloxacin and meropenem in any of the isolates. In seven isolates, we detected chloramphenicol resistance. In five K. oxytoca isolates from chestnut powder, we observed tetracycline resistance.

Table 8.
Antibiotic-resistant profiles of isolated resistant coliform bacteria against selected types of antibiotics.
Enterococci have intrinsic and acquired resistance to most antibiotics used in humans, leading to treatment complications. Enterococci are naturally resistant to β-lactams, carbapenems and cephalosporins. They may also acquire resistance to tetracyclines, quinolones, macrolides, erythromycin, streptogramin, polymixins, clindamycin, and glycopeptides (vancomycin) [50]. Of the ten isolates tested, only three showed resistance to ampicillin at CLSI concentrations (Table 9). Ampicillin and penicillin, belonging to β-lactam ATBs, inhibit the synthesis of peptidoglycan, a critical component required for bacterial viability [50]. Higher levels of ampicillin resistance in enterococci are associated with the presence of the species-specific chromosomal gene pbp5, which encodes class B penicillin-binding proteins (PBPs) with binding affinity for ampicillin and cephalosporins [51]. All isolates were vancomycin-resistant. Vancomycin-resistant enterococci found in food can temporarily colonize the human gut and confer multidrug resistance to human E. faecalis via a conjugation plasmid [52].

Table 9.
Antibiotic-resistant profiles of isolated resistant enterococci against selected types of antibiotics.
Multidrug resistance, defined as resistance to three or more antimicrobials, was not detected in any of the isolates. The high level of antibiotic resistance is due to the widespread use of antibiotics in medical practice, especially in developing countries [42].
Efflux causes a decrease in intracellular drug concentrations, leading to subtoxic levels, which may ultimately induce phenotypic resistance [53]. All isolates of coliform bacteria and enterococci showed normal efflux. In contrast to our results, Krahulcova et al. recorded in their work the overproduction of efflux pumps in up to 19% of coliforms obtained from smoothie drinks [25].
About 1300 types of β-lactamases are currently known, of which 200 are types of broad-spectrum β-lactamases (ESBLs) [54]. ESBLs are commonly present in Klebsiella and E. coli species but have also been detected in other members of the Enterobacterales family, such as Salmonella, Enterobacter, Citrobacter, Serratia and Proteus. These microorganisms are the cause of septicemia, pneumonia, intra-abdominal abscess as well as various infections, e.g., urinary tract infections [55]. ESBL-related infections are on the rise and have been observed worldwide in hospitals and communities [56].
Coliform isolates, in which we confirmed ampicillin resistance were subjected to the detection of the blaTEM, blaSHV and blaOXA genes. We did not detect the presence of the blaTEM and blaOXA genes in any of the isolates. We confirmed the presence of the blaSHV-1 gene in 13% of ampicillin-resistant coliform bacteria, of which all isolates belonged to the K. oxytoca species (Table 10). Klebsiella species exhibit resistance to a broad spectrum of antibiotics, with resistance to penicillins, particularly ampicillin and carbenicillin, being mediated by the enzyme SHV-1 [57]. In the remaining 87% of isolates, ampicillin resistance may be due to other mechanisms or genes.

Table 10.
Detected resistance genes in resistant coliform bacteria.
We also determined the presence of tetA and tetE genes in coliform isolates (Table 10). These genes mediate tetracycline resistance through membrane-associated proteins that export the drug from the cell and thereby reduce its intracellular concentration [58]. Although we did not detect excessive production of efflux pumps in any of the isolates, we recorded the presence of the tetA and tetE genes in eight isolates. It follows that the isolates have genes encoding specific efflux pumps, but they are not expressed. In addition to overproduction of efflux pumps, resistance to tetracycline antibiotics is mediated by two other mechanisms, namely the protection of ribosomes by proteins that prevent the efficient binding of tetracycline or the production of enzymes that inhibit the ATB. Each of these mechanisms is encoded by a wide variety of genes [58].
In the case of enterococci, we focused on the detection of the vanA gene, which is the most common gene encoding glycopeptide resistance [59]. Worldwide, the vanA gene is frequently detected in E. faecium and E. faecalis strains isolated from humans and animals but is less common in other species, such as E. gallinarum, E. hirae, and E. durans [60]. Although all isolates showed vancomycin resistance, we did not detect the vanA gene in any of the isolates. However, vancomycin resistance is mediated by several genes referred to as vanA -B, -C, -D, -E, -G. Of these, the vanA and vanB genes located on plasmids are most frequently detected, followed by chromosomally localized vanD and vanC genes. These genes encode changes in the amino acid sequence of the peptidoglycan precursor protein, thereby preventing vancomycin binding [61]. The relationship of food enterococci with clinical infections has not yet been clearly elucidated [62]. Foodborne enterococci are not known to be a direct source of resistant enterococci in humans but could pose a risk of transmitting resistance determinants to strains present in the human intestinal microbiota [60].
Biofilms can be found in food, medical and natural environments. Pathogenic bacteria that enter food production areas can remain there in the form of a biofilm covering the surfaces of machinery and equipment. Biofilms formed on these surfaces are a major cause of contamination of the final product, which ultimately leads to disease transmission, reduced food shelf life and quality, and thus huge economic losses [63]. Studies suggest that the biofilm is important for the transfer of conjugation plasmids due to the high proximity of cells in this structure. The resistance of a biofilm to ATB depends on various factors, such as physical, physiological, and genetic factors [63]. In a mature biofilm, bacterial cells can tolerate antibiotics at concentrations 10–1000-fold higher than those required to kill planktonic cells [64]. The majority of isolated coliforms (72%) were strong producers of biofilm (Figure 3), which correlates with the results of Krahulcová, who evaluated 73% of coliform isolates from raw milk and smoothie as strong biofilm producers [25]. The remaining 28% of isolates were medium-strength biofilm producers. We identified 86% of K. oxytoca as a strong biofilm producer. All K. pneumoniae isolates were classified as strong biofilm producers. Biofilm-forming K. pneumoniae is a major source of nosocomial infections associated with high morbidity and mortality due to limited treatment options [63]. Two isolates of C. sakazakii showed strong production, while the remaining three isolates were intermediate biofilm producers. The survival and persistence of C. sakazakii in powdered infant formula require the body’s ability to adapt to harsh osmotic and dry conditions. Studies have shown that C. sakazakii cells in biofilms were more persistent compared to plankton cells when exposed to low humidity conditions [65].
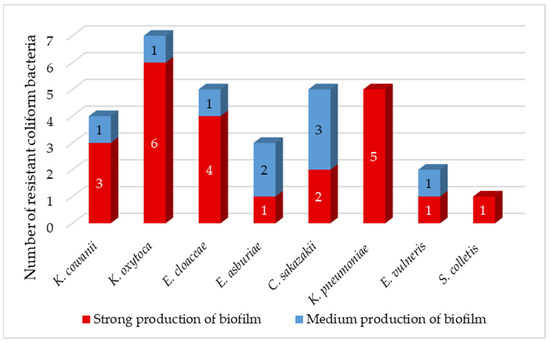
Figure 3.
Production of biofilm between isolated resistant coliform bacteria.
Enterococci are known for their ability to form biofilms [66]. Enterococcal biofilms have been reported in many infections, including urinary tract, wounds, gastrointestinal tract, and endocarditis. Biofilm-associated enterococcal infections are not only difficult to eradicate but also serve as a reservoir for antibiotic resistance genes. Low penetration of antibiotics through the biofilm matrix and the presence of persistent cells contribute to antibiotic tolerance of biofilms, leading to persistent infections [49]. Approximately 80% of persistent bacterial infections in the United States have been found to be associated with biofilms [63]. We observed medium biofilm production in isolates of E. casseliflavus and E. gallinarium (Figure 4). In the remaining 40% of isolates, including E. faecium isolate, we have detected strong production of biofilm. We did not observe very strong biofilm production in any of the coliform bacteria nor enterococci isolates (Figure 4).
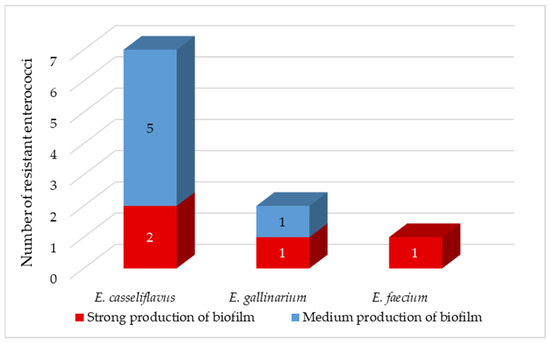
Figure 4.
Production of biofilm between isolated resistant enterococci.
4. Conclusions
The contamination of food with antibiotic-resistant pathogens poses a serious risk to public health. In recent years, flour has been recorded as an important source of food-borne diseases, while outbreaks were due to the consumption of raw dough. The new nutrition trends together with food supply globalization and increased civilizational disease incidence led to a change in consumers’ diet habits. For these reasons, we can observe an increased consumer demand for alternative flours or powders. Based on previous results, samples of flours and powders contain antibiotic-resistant bacteria where the majority were identified as Klebsiella spp. and Enterobacter spp. Although none of the resistant isolates harbored multidrug resistance and all showed normal efflux, we detected resistance genes such as blaSHV, tetA and tetE. Enterococci were less detected compared to coliform bacteria and were present only in non-cereal flours (powders). Identified strains were E. casseliflavus, E. gallinarium a E. faecium. Samples of dough contained only total coliform bacteria and enterococci. The presence of potentially pathogenic bacteria resistant to antibiotics in tested samples confirms the role of the food chain in the spread of resistance in the environment. Therefore, it is necessary to inform consumers about the correct handling of flour and dough and the importance of heat treatment. At the same time, we would like to suggest to legislatively define microbiological criteria for alternative plant-based flours, especially those which can be used without previous heat treatment.
Author Contributions
Conceptualization, K.Č. and K.C.; Formal analysis, L.B.; Investigation, J.K., M.K. and K.C.; Methodology, K.C.; Supervision, L.B.; Writing—original draft, J.K. and L.B.; Writing—review & editing, M.K. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Scientific Grant Agency VEGA under the contract 1/0464/21, by the Slovak Research and Development Agency under the contract APVV-16-0171 and by the Operational Program Integrated Infrastructure for the project: “Strategic research in the field of SMART monitoring, treatment and preventive protection against coronavirus (SARS-CoV-2)”, Project no. 313011ASS8, co-financed by the European Regional Development Fund.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Data of the current study are available from the corresponding author. The data are not publicly available due to privacy.
Conflicts of Interest
The authors declare that there is no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
References
- Feng, Y.; Archila-Godínez, J.C. Consumer Knowledge and Behaviors Regarding Food Safety Risks Associated with Wheat Flour. J. Food Prot. 2021, 84, 628–638. [Google Scholar] [CrossRef] [PubMed]
- Magallanes López, A.M.; Simsek, S. Pathogens control on wheat and wheat flour: A review. Cereal Chem. 2021, 98, 17–30. [Google Scholar] [CrossRef]
- Crowe, S.J.; Bottichio, L.; Shade, L.N.; Whitney, B.M.; Corral, N.; Melius, B.; Arends, K.D.; Donovan, D.; Stone, J.; Allen, K.; et al. Shiga Toxin–Producing E. coli Infections Associated with Flour. N. Engl. J. Med. 2017, 377, 2036–2043. [Google Scholar] [CrossRef] [PubMed]
- Sabillón, L.; Bianchini, A. From Field to Table: A Review on the Microbiological Quality and Safety of Wheat-Based Products. Cereal Chem. J. 2016, 93, 105–115. [Google Scholar] [CrossRef]
- Chen, Y.-X.; Guo, X.-N.; Xing, J.-J.; Zhu, K.-X. Effects of tempering with steam on the water distribution of wheat grains and quality properties of wheat flour. Food Chem. 2020, 323, 126842. [Google Scholar] [CrossRef] [PubMed]
- Bundesinstitut für Risikobewertung (BfR). Escherichia coli in Flour—Sources, Risks and Prevention; BfR-Stellungnahmen: Berlin, Germany, 2020; Volume 2020, pp. 1–28. [Google Scholar] [CrossRef]
- Sabillón, L.; Stratton, J.; Rose, D.; Bianchini, A. Microbiological survey of equipment and wheat-milled fractions of a milling operation. Cereal Chem. 2021, 98, 44–51. [Google Scholar] [CrossRef]
- LAVES. Roher Teig birgt Risiken. Available online: https://www.laves.niedersachsen.de/startseite/lebensmittel/lebensmittelgruppen/getreide_getreideerzeugnisse/roher-kuchenteig-birgt-risiken-160553.html (accessed on 8 August 2022).
- Boss, R.; Hummerjohann, J. Whole Genome Sequencing Characterization of Shiga Toxin–Producing Escherichia coli Isolated from Flour from Swiss Retail Markets. J. Food Prot. 2019, 82, 1398–1404. [Google Scholar] [CrossRef]
- McCallum, L.; Paine, S.; Sexton, K.; Dufour, M.; Dyet, K.; Wilson, M.; Campbell, D.; Bandaranayake, D.; Hope, V. An Outbreak of Salmonella Typhimurium Phage Type 42 Associated with the Consumption of Raw Flour. Foodborne Pathog. Dis. 2013, 10, 159–164. [Google Scholar] [CrossRef]
- Gill, A.; Carrillo, C.; Hadley, M.; Kenwell, R.; Chui, L. Bacteriological analysis of wheat flour associated with an outbreak of Shiga toxin-producing Escherichia coli O121. Food Microbiol. 2019, 82, 474–481. [Google Scholar] [CrossRef]
- Myoda, S.P.; Gilbreth, S.; Akins-Lewenthal, D.; Davidson, S.K.; Samadpour, M. Occurrence and Levels of Salmonella, Enterohemorrhagic Escherichia coli, and Listeria in Raw Wheat. J. Food Prot. 2019, 82, 1022–1027. [Google Scholar] [CrossRef]
- FDA. FDA Investigated Multistate Outbreak of Shiga Toxin-Producing E. coli Infections Linked to Flour. Available online: https://www.fda.gov/food/outbreaks-foodborne-illness/fda-investigation-multistate-outbreak-shiga-toxin-producing-e-coli-infections-linked-flour#Problem (accessed on 8 August 2022).
- Morton, V.; Cheng, J.; Sharma, D.; Kearney, A. An outbreak of Shiga toxin-producing Escherichia coli O121 infections associated with flour—Canada, 2016–2017. Can. Commun. Dis. Rep. 2017, 43, 154–155. [Google Scholar] [CrossRef] [PubMed]
- Losio, M.-N.; Dalzini, E.; Pavoni, E.; Merigo, D.; Finazzi, G.; Daminelli, P. A survey study on safety and microbial quality of “gluten-free” products made in Italian pasta factories. Food Control 2017, 73, 316–322. [Google Scholar] [CrossRef]
- Foschia, M.; Horstmann, S.; Arendt, E.K.; Zannini, E. Nutritional therapy—Facing the gap between coeliac disease and gluten-free food. Int. J. Food Microbiol. 2016, 239, 113–124. [Google Scholar] [CrossRef] [PubMed]
- Pekárková, B. Celiakia na Slovensku—čo o nej Možno Neviete—BEZlepku. Available online: https://www.bezlepku.sk/2020/05/28/celiakia-na-slovensku-co-o-nej-mozno-neviete/ (accessed on 8 August 2022).
- Muky.sk. Available online: https://www.muky.sk/ (accessed on 8 August 2022).
- Eglezos, S. Microbiological Quality of Wheat Grain and Flour from Two Mills in Queensland, Australia. J. Food Prot. 2010, 73, 1533–1536. [Google Scholar] [CrossRef]
- Li, Z.; Li, H.; Bian, K. Microbiological characterization of traditional dough fermentation starter (Jiaozi) for steamed bread making by culture-dependent and culture-independent methods. Int. J. Food Microbiol. 2016, 234, 9–14. [Google Scholar] [CrossRef]
- Formato, A.; Pepe, O. Pizza Dough Differentiation by Principal Component Analysis of Alveographic, Microbiological, and Chemical Parameters. Cereal Chem. J. 2005, 82, 356–360. [Google Scholar] [CrossRef]
- The European Committee on Antimicrobial Susceptibility Testing (EUCAST). Breakpoint Tables for Interpretation of MICs, Version 12.0. Available online: http://www.eucast.org (accessed on 8 August 2022).
- CLSI. M100-ED32:2022 Performance Standards for Antimicrobial Susceptibility Testing, 32nd ed.; Available online: http://em100.edaptivedocs.net/GetDoc.aspx?doc=CLSIM100ED32:2022&scope=user (accessed on 8 August 2022).
- Lépesová, K.; Olejníková, P.; Mackuľak, T.; Tichý, J.; Birošová, L. Annual changes in the occurrence of antibiotic-resistant coliform bacteria and enterococci in municipal wastewater. Environ. Sci. Pollut. Res. 2019, 26, 18470–18483. [Google Scholar] [CrossRef]
- Krahulcová, M.; Cverenkárová, K.; Olejníková, P.; Micajová, B.; Koreneková, J.; Bírošová, L. Characterization of Antibiotic Resistant Coliform Bacteria and Resistance Genes Isolated from Samples of Smoothie Drinks and Raw Milk. Foods 2022, 11, 1324. [Google Scholar] [CrossRef]
- Beenken, K.E.; Blevins, J.S.; Smeltzer, M.S. Mutation of sarA in Staphylococcus aureus Limits Biofilm Formation. Infect. Immun. 2003, 71, 4206–4211. [Google Scholar] [CrossRef]
- Taniguchi, L.; de Fátima Faria, B.; Rosa, R.T.; de Paula e Carvalho, A.; Gursky, L.C.; Elifio-Esposito, S.L.; Parahitiyawa, N.; Samaranayake, L.P.; Rosa, E.A.R. Proposal of a low-cost protocol for colorimetric semi-quantification of secretory phospholipase by Candida albicans grown in planktonic and biofilm phases. J. Microbiol. Methods 2009, 78, 171–174. [Google Scholar] [CrossRef]
- Martins, M.; McCusker, M.P.; Viveiros, M.; Couto, I.; Fanning, S.; Pagès, J.-M.; Amaral, L. A Simple Method for Assessment of MDR Bacteria for Over-Expressed Efflux Pumps. Open Microbiol. J. 2013, 7, 72–82. [Google Scholar] [CrossRef]
- Lépesová, K.; Olejníková, P.; Mackuľak, T.; Cverenkárová, K.; Krahulcová, M.; Bírošová, L. Hospital Wastewater—Important Source of Multidrug Resistant Coliform Bacteria with ESBL-Production. Int. J. Environ. Res. Public Health 2020, 17, 7827. [Google Scholar] [CrossRef]
- Dallenne, C.; Da Costa, A.; Decré, D.; Favier, C.; Arlet, G. Development of a set of multiplex PCR assays for the detection of genes encoding important β-lactamases in Enterobacteriaceae. J. Antimicrob. Chemother. 2010, 65, 490–495. [Google Scholar] [CrossRef] [PubMed]
- Ng, L.-K.; Martin, I.; Alfa, M.; Mulvey, M. Multiplex PCR for the detection of tetracycline resistant genes. Mol. Cell. Probes 2001, 15, 209–215. [Google Scholar] [CrossRef] [PubMed]
- Depardieu, F.; Perichon, B.; Courvalin, P. Detection of the van Alphabet and Identification of Enterococci and Staphylococci at the Species Level by Multiplex PCR. J. Clin. Microbiol. 2004, 42, 5857–5860. [Google Scholar] [CrossRef] [PubMed]
- Batool, S.A.; Rauf, N.; Kalsoom, R. Microbial and Physico-chemical contamination in the wheat flour of the twin cities of Pakistan. Int. J. Food Saf. 2012, 14, 75–82. [Google Scholar]
- Ministry of Agriculture and Rural Develompment of the Slovak Republic. MPSR Decree No. 06267/2006-SL. Available online: https://www.svps.sk/legislativa/legislativa_kodex.asp (accessed on 8 August 2022).
- Food Safety News. French Officials Confirm Some E. coli Cases Linked to Nestlé Pizza. Available online: https://www.foodsafetynews.com/2022/03/french-officials-confirm-some-e-coli-cases-linked-to-nestle-pizza/ (accessed on 8 August 2022).
- Ghosh, A.; Zurek, L. Antibiotic Resistance in Enterococci. In Antimicrobial Resistance and Food Safety; Chen, C.-Y., Yan, X., Jackson, C.R., Eds.; Elsevier: San Diego, CA, USA, 2015; pp. 155–180. ISBN 978-0-12-801214-7. [Google Scholar]
- Ramos, S.; Silva, V.; Dapkevicius, M.; Igrejas, G.; Poeta, P. Enterococci, from Harmless Bacteria to a Pathogen. Microorganisms 2020, 8, 1118. [Google Scholar] [CrossRef]
- FDA; King Arthur Flour Company, Inc. Voluntarily Recalls Organic Coconut Flour (16 Oz.) Because of Possible Health Risk. Available online: https://www.fda.gov/safety/recalls-market-withdrawals-safety-alerts/king-arthur-flour-company-inc-voluntarily-recalls-organic-coconut-flour-16-oz-because-possible#recall-announcement (accessed on 8 August 2022).
- FDA; Bob’ s Red Mill Natural Foods, Inc. Voluntarily Recalls Organic Amaranth Flour (22 Oz.) Because of Possible Health Risk. Available online: https://www.fda.gov/safety/recalls-market-withdrawals-safety-alerts/bobs-red-mill-natural-foods-inc-voluntarily-recalls-organic-amaranth-flour-22-oz-because-possible#recall-announcement (accessed on 8 August 2022).
- Beňo, P.; Kaiglová, A.; Blažíčková, S.; Dovinová, I. Vybrané Laboratórne Vyšetrovacie Metódy v Zdravotníctve; Typi Universitatis Tyrnaviensis: Trnava, Slovakia, 2018; ISBN 9788056800768. [Google Scholar]
- Gundogan, N. Klebsiella. In Encyclopedia of Food Microbiology; Batt, C.A., Tortorello, M.L., Eds.; Elsevier: Oxford, UK, 2014; pp. 383–388. ISBN 978-0-12-384733-1. [Google Scholar]
- Chakraborty, S. Prevalence, antibiotic susceptibility profiles and ESBL production in Klebsiella pneumoniae and Klebsiella oxytoca among hospitalized patients. Period. Biol. 2016, 118, 53–58. [Google Scholar] [CrossRef]
- Theocharidi, N.A.; Balta, I.; Houhoula, D.; Tsantes, A.G.; Lalliotis, G.P.; Polydera, A.C.; Stamatis, H.; Halvatsiotis, P. High Prevalence of Klebsiella pneumoniae in Greek Meat Products: Detection of Virulence and Antimicrobial Resistance Genes by Molecular Techniques. Foods 2022, 11, 708. [Google Scholar] [CrossRef]
- Abdesselam, K.; Pagotto, F. Bacteria: Cronobacter (Enterobacter) sakazakii and Other Cronobacter spp. In Encyclopedia of Food Safety, History, Science and Methods; Motarjemi, Y., Moy, G., Todd, E., Eds.; Academic Press: London, UK, 2014; Volume 1, pp. 424–432. [Google Scholar]
- Berinson, B.; Bellon, E.; Christner, M.; Both, A.; Aepfelbacher, M.; Rohde, H. Identification of Kosakonia cowanii as a rare cause of acute cholecystitis: Case report and review of the literature. BMC Infect. Dis. 2020, 20, 366. [Google Scholar] [CrossRef]
- Petrzik, K.; Brázdová, S.; Krawczyk, K. Novel Viruses That Lyse Plant and Human Strains of Kosakonia cowanii. Viruses 2021, 13, 1418. [Google Scholar] [CrossRef] [PubMed]
- Guglielmetti, P.; Bartoloni, A. Escherichia coli|Occurrence and Epidemiology of Species other than Escherichia coli. In Encyclopedia of Food Sciences and Nutrition; Academic Press: London, UK, 2003; pp. 2162–2166. ISBN 978-0-12-227055-0. [Google Scholar]
- Jain, S.; Nagarjuna, D.; Gaind, R.; Chopra, S.; Debata, P.K.; Dawar, R.; Sardana, R.; Yadav, M. Escherichia vulneris: An unusual cause of complicated diarrhoea and sepsis in an infant. A case report and review of literature. New Microbes New Infect. 2016, 13, 83–86. [Google Scholar] [CrossRef] [PubMed]
- Ch’ng, J.-H.; Chong, K.K.L.; Lam, L.N.; Wong, J.J.; Kline, K.A. Biofilm-associated infection by enterococci. Nat. Rev. Microbiol. 2019, 17, 82–94. [Google Scholar] [CrossRef] [PubMed]
- Sanlibaba, P.; Senturk, E. Prevalence, characterization and antibiotic resistance of enterococci from traditional cheeses in Turkey. Int. J. Food Prop. 2018, 21, 1955–1963. [Google Scholar] [CrossRef]
- Miller, W.R.; Munita, J.M.; Arias, C.A. Mechanisms of antibiotic resistance in enterococci. Expert Rev. Anti. Infect. Ther. 2014, 12, 1221–1236. [Google Scholar] [CrossRef] [PubMed]
- Lim, S.-K.; Tanimoto, K.; Tomita, H.; Ike, Y. Pheromone-Responsive Conjugative Vancomycin Resistance Plasmids in Enterococcus faecalis Isolates from Humans and Chicken Feces. Appl. Environ. Microbiol. 2006, 72, 6544–6553. [Google Scholar] [CrossRef] [PubMed]
- Lamut, A.; Peterlin Mašič, L.; Kikelj, D.; Tomašič, T. Efflux pump inhibitors of clinically relevant multidrug resistant bacteria. Med. Res. Rev. 2019, 39, 2460–2504. [Google Scholar] [CrossRef]
- Roopashree, S.; Kaup, S. Prevalence of various Beta-lactamases in Enterobacteriaceae in a tertiary care hospital in South India: A Cross-sectional study. IP Int. J. Med. Microbiol. Trop. Dis. 2021, 2021, 186–191. [Google Scholar] [CrossRef]
- Dirar, M.H.; Bilal, N.E.; Ibrahim, M.E.; Hamid, M.E. Prevalence of extended-spectrum β-lactamase (ESBL) and molecular detection of bla TEM, bla SHV and bla CTX-M genotypes among Enterobacteriaceae isolates from patients in Khartoum, Sudan. Pan Afr. Med. J. 2020, 37, 213. [Google Scholar] [CrossRef]
- Malik, I.A.; Elhag, K.M. Characterisation of Extended-Spectrum β-Lactamases among Multidrug Resistant Enterobacteriaceae from Sudan. J. Pure Appl. Microbiol. 2019, 13, 61–68. [Google Scholar] [CrossRef]
- Cooney, S.; O’Brien, S.; Iversen, C.; Fanning, S. Bacteria: Other pathogenic Enterobacteriaceae—Enterobacter and other genera. In Encyclopedia of Food Safety; Motarjemi, Y., Moy, G., Todd, E., Eds.; Academic Press: London, UK, 2014; Volume 1, pp. 433–441. ISBN 978-0-12-378613-5. [Google Scholar]
- Pazda, M.; Kumirska, J.; Stepnowski, P.; Mulkiewicz, E. Antibiotic resistance genes identified in wastewater treatment plant systems—A review. Sci. Total Environ. 2019, 697, 134023. [Google Scholar] [CrossRef] [PubMed]
- Vehreschild, M.J.G.T.; Haverkamp, M.; Biehl, L.M.; Lemmen, S.; Fätkenheuer, G. Vancomycin-resistant enterococci (VRE): A reason to isolate? Infection 2019, 47, 7–11. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, M.O.; Baptiste, K.E. Vancomycin-Resistant Enterococci: A Review of Antimicrobial Resistance Mechanisms and Perspectives of Human and Animal Health. Microb. Drug Resist. 2018, 24, 590–606. [Google Scholar] [CrossRef] [PubMed]
- Levitus, M.; Rewane, A.; Perera, T.B. Vancomycin-Resistant Enterococci. In StatPearls (Internet); StatPearls: Treasure Island, FL, USA, 2022. [Google Scholar]
- Terkuran, M.; Turhan, E.Ü.; Erginkaya, Z. The Risk of Vancomycin Resistant Enterococci Infections from Food Industry. In Health and Safety Aspects of Food Processing Technologies; Malik, A., Erginkaya, Z., Erten, H., Eds.; Springer International Publishing: Cham, Switzerland, 2019; pp. 513–535. ISBN 978-3-030-24903-8. [Google Scholar]
- Abebe, G.M. The Role of Bacterial Biofilm in Antibiotic Resistance and Food Contamination. Int. J. Microbiol. 2020, 2020, 1705814. [Google Scholar] [CrossRef]
- Zurob, E.; Dennett, G.; Gentil, D.; Montero-Silva, F.; Gerber, U.; Naulín, P.; Gómez, A.; Fuentes, R.; Lascano, S.; Rodrigues da Cunha, T.H.; et al. Inhibition of Wild Enterobacter cloacae Biofilm Formation by Nanostructured Graphene- and Hexagonal Boron Nitride-Coated Surfaces. Nanomaterials 2019, 9, 49. [Google Scholar] [CrossRef]
- Ling, N.; Forsythe, S.; Wu, Q.; Ding, Y.; Zhang, J.; Zeng, H. Insights into Cronobacter sakazakii Biofilm Formation and Control Strategies in the Food Industry. Engineering 2020, 6, 393–405. [Google Scholar] [CrossRef]
- Hashem, Y.A.; Amin, H.M.; Essam, T.M.; Yassin, A.S.; Aziz, R.K. Biofilm formation in enterococci: Genotype-phenotype correlations and inhibition by vancomycin. Sci. Rep. 2017, 7, 5733. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).