Bacteriocin Production by Escherichia coli during Biofilm Development
Abstract
:1. Introduction
2. Materials and Methods
2.1. Bacterial Strains and Culture Conditions
2.2. Biofilm Formation Assay
2.3. Antimicrobial Activity Assay
2.4. Culturability Assessment of Dual-Species Biofilms
2.5. Bacterial Growth Inhibition Measurements
2.6. Determination of Bacteriocin Concentration
2.7. Screening for Siderophore Production
2.8. PCR Amplifications
2.9. RNA Extraction
2.10. Real-Time PCR (qPCR)
2.11. Statistical Analysis
3. Results
3.1. E. coli Biofilm Formation Ability, Bacteriocin Synthesis, and Siderophore Production
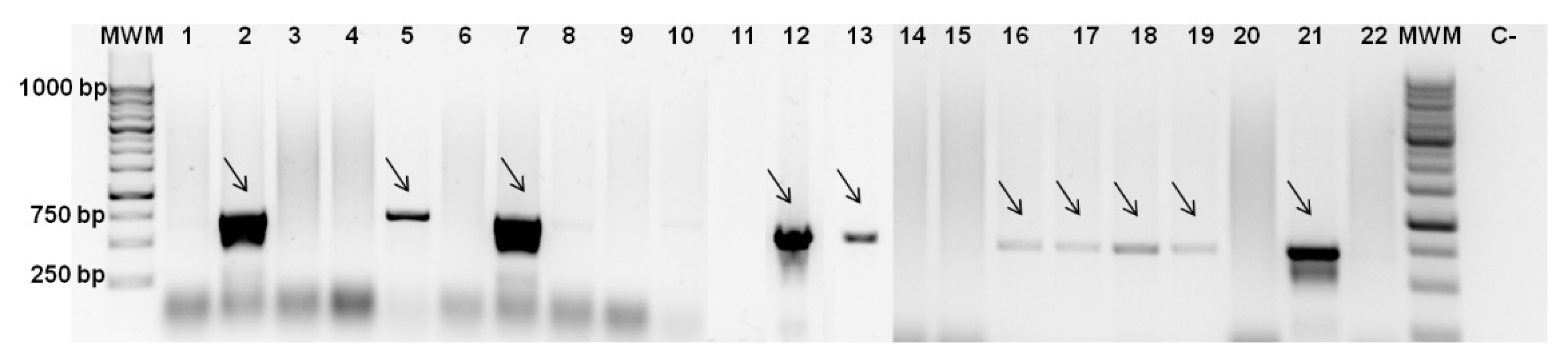
3.2. Screening for Colicin Genes and Antimicrobial Spectrum
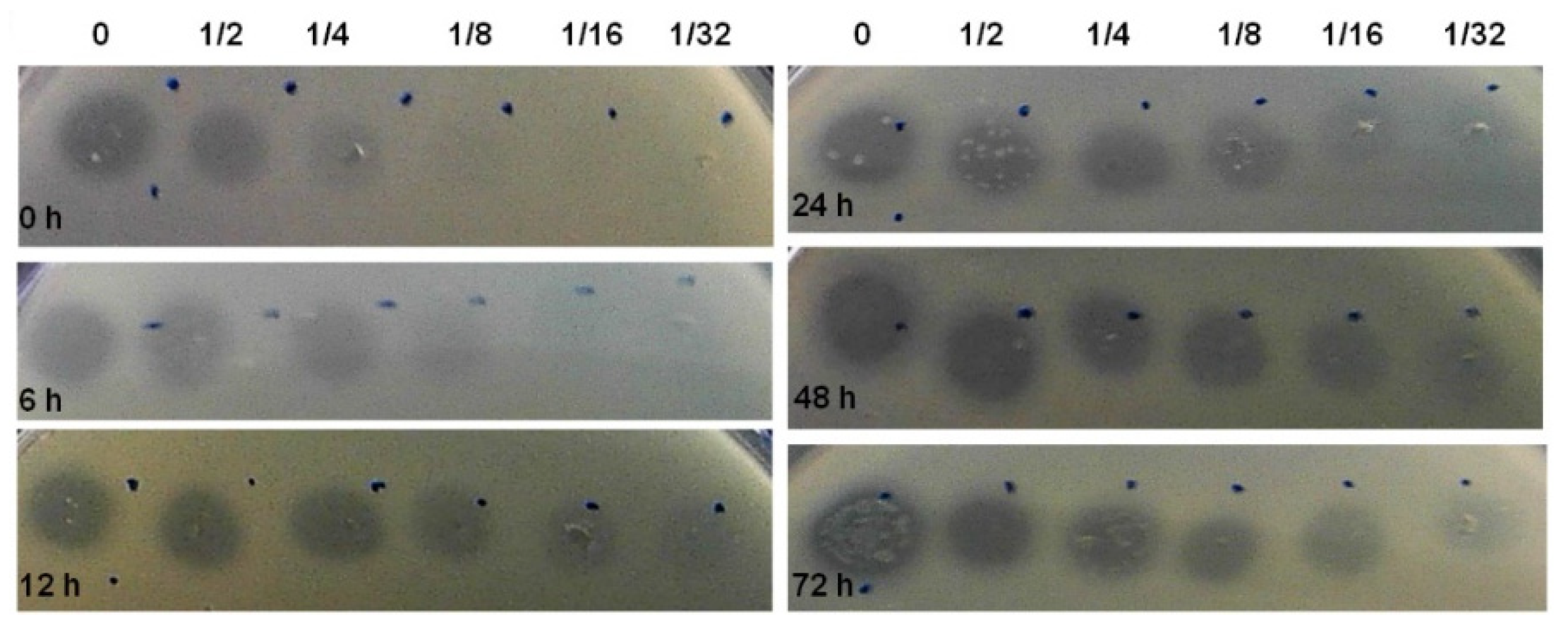
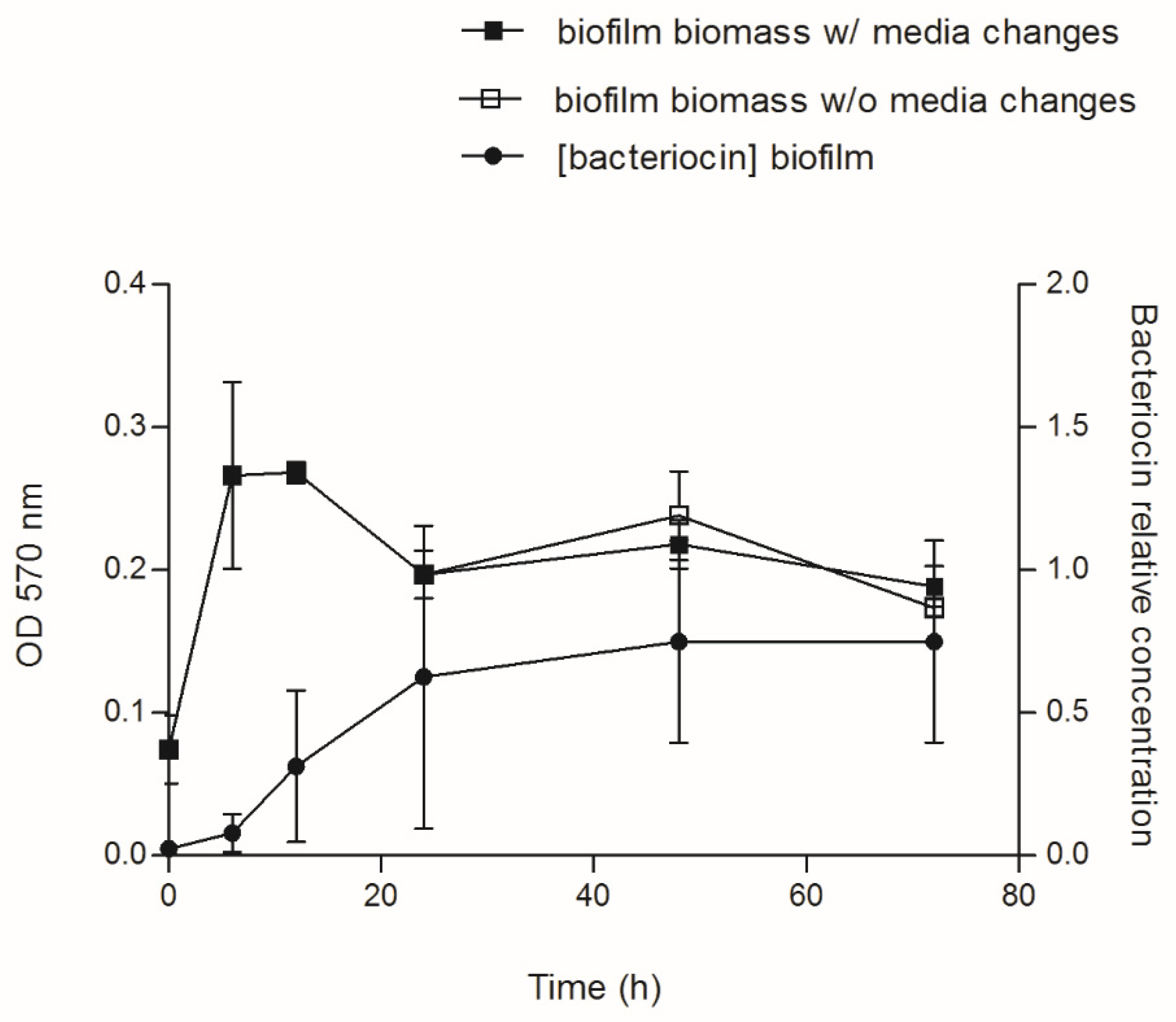
3.3. Bacteriocin Production during Biofilm Development
3.4. Dual-Species Biofilms
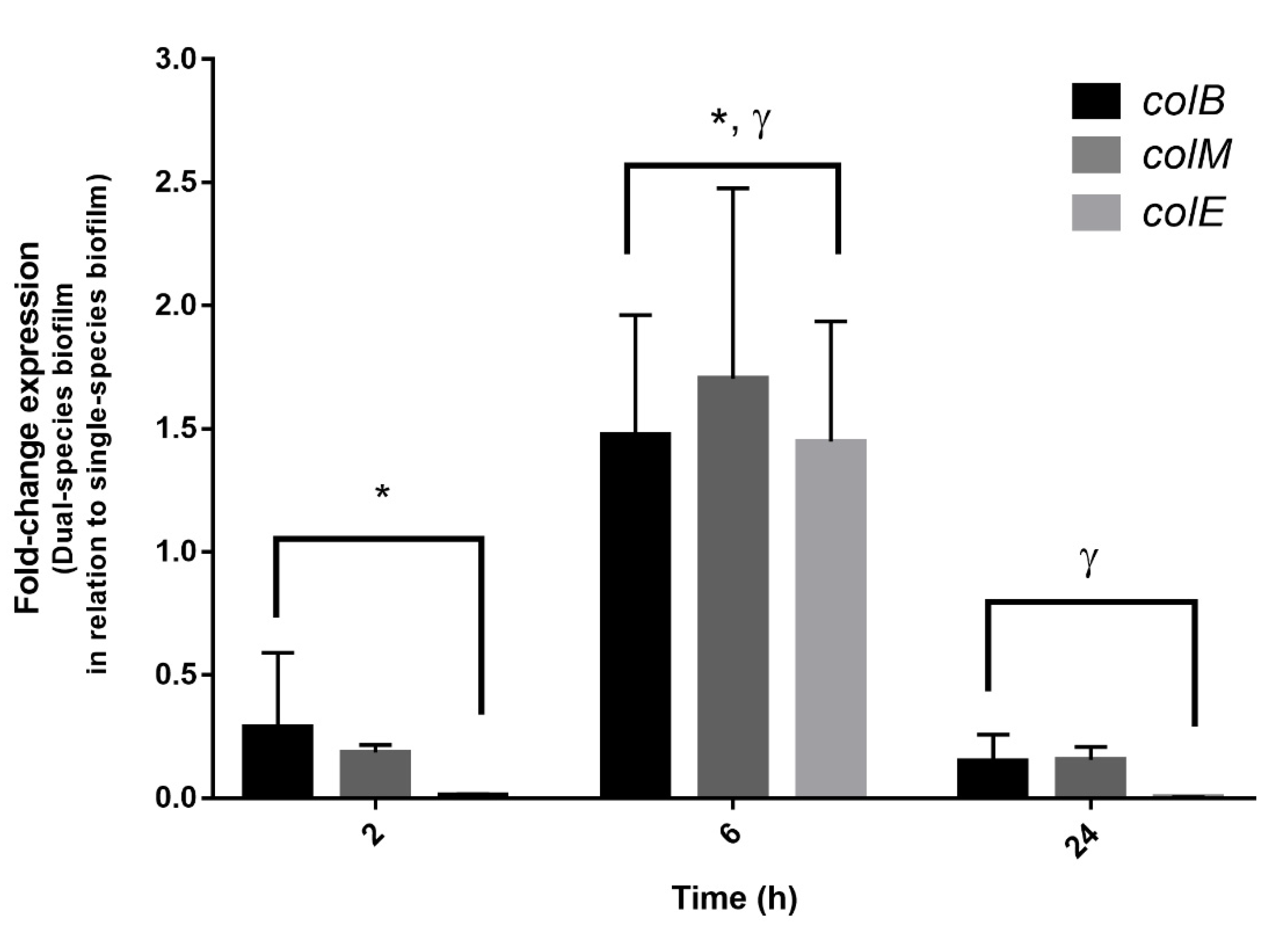
3.5. Expression of Colicins in Dual-Species Biofilms
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Yang, S.C.; Lin, C.H.; Aljuffali, I.A.; Fang, J.Y. Current Pathogenic Escherichia Coli Foodborne Outbreak Cases and Therapy Development. Arch. Microbiol. 2017, 199, 811–825. [Google Scholar] [CrossRef] [PubMed]
- Wasiński, B. Extra-Intestinal Pathogenic Escherichia coli—Threat Connected with Food-Borne Infections. Ann. Agric. Environ. Med. 2019, 26, 532–537. [Google Scholar] [CrossRef] [PubMed]
- Niamah, A.K.; Thyab, S.; Al-Sahlany, G. Detecting for E. coli O157:H7 in dairy products which were locally processed and found in Basra city markets. Basrah J. Agric. Sci. 2011, 24, 290–299. [Google Scholar]
- Mulder, C.J.J.; Zanen, H.C. A study of 280 cases of neonatal meningitis in The Netherlands. J. Infect. 1984, 9, 77–84. [Google Scholar] [CrossRef]
- Kaper, J.B.; Nataro, J.P.; Mobley, H.L.T. Pathogenic Escherichia coli. Nat. Rev. Microbiol. 2004, 2, 123–140. [Google Scholar] [CrossRef]
- Robbins, J.B.; McCracken, G.H.; Gotschlich, E.C.; Ørskov, I.; Hanson, L.A. Escherichia coli K1 Capsular Polysaccharide Associated with Neonatal Meningitis. N. Engl. J. Med. 1974, 30, 1216–1220. [Google Scholar] [CrossRef]
- Stamm, W.E.; Counts, G.W.; Running, K.R.; Fihn, S.; Turck, M.; Holmes, K.K. Diagnosis of Coliform Infection in Acutely Dysuric Women. N. Engl. J. Med. 2014, 307, 463–468. [Google Scholar] [CrossRef] [PubMed]
- Post, R.E.; Hainer, B.L. Gastrointestinal Tract Infections. In Management of Antimicrobials in Infectious Diseases: Impact of Antibiotic Resistance; MainousIII, A.G., Pomeroy, C., Eds.; Springer Science + Business Media: New York, NY, USA; Elsevier BV: Amsterdam, The Netherlands, 2010; pp. 231–265. [Google Scholar]
- Probert, H.M.; Gibson, G.R. Bacterial Biofilms in the Human Gastrointestinal Tract. Curr. Issues Intest. Microbiol. 2002, 3, 23–27. [Google Scholar]
- Sharma, G.; Sharma, S.; Sharma, P.; Chandola, D.; Dang, S.; Gupta, S.; Gabrani, R. Escherichia coli Biofilm: Development and Therapeutic Strategies. J. Appl. Microbiol. 2016, 121, 309–319. [Google Scholar] [CrossRef]
- Costerton, J.W.; Stewart, P.S.; Greenberg, E.P. Bacterial Biofilms: A Common Cause of Persistent Infections. Science 1999, 284, 1318–1322. [Google Scholar] [CrossRef]
- Eick, S. Biofilms. Monogr. Oral Sci. 2021, 29, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Høiby, N.; Bjarnsholt, T.; Givskov, M.; Molin, S.; Ciofu, O. Antibiotic Resistance of Bacterial Biofilms. Int. J. Antimicrob. Agents 2010, 35, 322–332. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Castro, J.; França, A.; Bradwell, K.R.; Serrano, M.G.; Jefferson, K.K.; Cerca, N. Comparative transcriptomic analysis of Gardnerella vaginalis biofilms vs. planktonic cultures using RNA-Seq. NPJ Biofilms Microbiomes 2017, 3. [Google Scholar] [CrossRef] [PubMed]
- Douglas, L.J. Candida biofilms and their role in infection. Trends Microbiol. 2003, 11, 30–36. [Google Scholar] [CrossRef]
- Hall-Stoodley, L.; Stoodley, P. Biofilm Formation and Dispersal and the Transmission of Human Pathogens. Trends Microbiol. 2005, 13, 7–10. [Google Scholar] [CrossRef]
- Parsek, M.R.; Fuqua, C. Biofilms 2003: Emerging Themes and Challenges in Studies of Surface-Associated Microbial Life. J. Bacteriol. 2004, 186, 4427–4440. [Google Scholar] [CrossRef]
- Dykes, G.A. Bacteriocins: Ecological and Evolutionary Significance. Tree 1995, 10, 186–189. [Google Scholar] [CrossRef]
- Riley, M.A.; Gordon, D.M. The Ecology and Evolution of Bacteriocins. J. Ind. Microbiol. 1996, 17, 151–158. [Google Scholar] [CrossRef]
- Meade, E.; Slattery, M.A.; Garvey, M. Bacteriocins, Potent Antimicrobial Peptides and the Fight against Multi Drug Resistant Species: Resistance Is Futile? Antibiotics 2020, 9, 32. [Google Scholar] [CrossRef]
- Hassan, M.; Kjos, M.; Nes, I.F.; Diep, D.B.; Lotfipour, F. Natural Antimicrobial Peptides from Bacteria: Characteristics and Potential Applications to Fight against Antibiotic Resistance. J. Appl. Microbiol. 2012, 113, 723–736. [Google Scholar] [CrossRef]
- Riley, M.A. Molecular mechanisms of bacteriocin evolution. Annu. Rev. Genet 1998, 32, 255–278. [Google Scholar] [CrossRef] [PubMed]
- Riley, M.A.; Wertz, J.E. Bacteriocin Diversity: Ecological and Evolutionary Perspectives. Biochimie 2002, 84, 357–364. [Google Scholar] [CrossRef]
- Verma, D.K.; Thakur, M.; Singh, S.; Tripathy, S.; Gupta, A.K.; Baranwal, D.; Patel, A.R.; Shah, N.; Utama, G.L.; Niamah, A.K.; et al. Bacteriocins as Antimicrobial and Preservative Agents in Food: Biosynthesis, Separation and Application. Food Biosci. 2022, 46, 101594. [Google Scholar] [CrossRef]
- Cascales, E.; Buchanan, S.K.; Duché, D.; Kleanthous, C.; Lloubès, R.; Postle, K.; Riley, M.; Slatin, S.; Cavard, D. Colicin Biology. Microbiol. Mol. Biol. Rev. 2007, 71, 158–229. [Google Scholar] [CrossRef] [PubMed]
- Gillor, O.; Kirkup, B.C.; Riley, M.A. Colicins and Microcins: The next Generation Antimicrobials. Adv. Appl. Microbiol. 2004, 54, 129–146. [Google Scholar] [CrossRef]
- Noinaj, N.; Guillier, M.; Barnard, T.J.; Buchanan, S.K. TonB-Dependent Transporters: Regulation, Structure, and Function. Annu. Rev. Microbiol. 2010, 64, 43–60. [Google Scholar] [CrossRef]
- Kim, Y.C.; Tarr, A.W.; Penfold, C.N. Colicin Import into E. coli Cells: A Model System for Insights into the Import Mechanisms of Bacteriocins. Biochim. Biophys. Acta Mol. Cell Res. 2014, 1843, 1717–1731. [Google Scholar] [CrossRef]
- Jin, X.; Kightlinger, W.; Kwon, Y.C.; Hong, S.H. Rapid Production and Characterization of Antimicrobial Colicins Using Escherichia coli-Based Cell-Free Protein Synthesis. Synth. Biol. 2018, 3, ysy004. [Google Scholar] [CrossRef]
- Andrews, S.C. Iron Storage in Bacteria. Adv. Microb. Physiol. 1998, 40, 281–351. [Google Scholar] [CrossRef]
- Pons, A.-M.; Lanneluc, I.; Cottenceau, G.; Sable, S. New Developments in Non-Post Translationally Modified Microcins. Biochimie 2002, 84, 531–537. [Google Scholar] [CrossRef]
- Li, C.; Bai, J.; Cai, Z.; Ouyang, F. Optimization of a Cultural Medium for Bacteriocin Production by Lactococcus lactis Using Response Surface Methodology. J. Biotechnol. 2002, 93, 27–34. [Google Scholar] [CrossRef]
- Salomón, R.A.; Farías, R.N. Influence of Iron on Microcin 25 Production. FEMS Microbiol. Lett. 1994, 121, 275–279. [Google Scholar] [CrossRef]
- Parente, E.; Ricciardi, A. Influence of PH on the Production of Enterocin 1146 during Batch Fermentation. Lett. Appl. Microbiol. 1994, 19, 12–15. [Google Scholar] [CrossRef]
- Martínez-Cardeñas, J.A.; de La Fuente-Salcido, N.M.; Salcedo-Hernández, R.; Bideshi, D.K.; Barboza-Corona, J.E. Effects of Physical Culture Parameters on Bacteriocin Production by Mexican Strains of Bacillus thuringiensis after Cellular Induction. J. Ind. Microbiol. Biotechnol. 2012, 39, 183–189. [Google Scholar] [CrossRef] [PubMed]
- Mintz, K.P.; Fives-Taylor, P.M. ImpA, a Gene Coding for an Inner Membrane Protein, Influences Colonial Morphology of Actinobacillus Actinomycetemcomitans. Infect. Immun. 2000, 68, 6580–6586. [Google Scholar] [CrossRef]
- Roche, E.D.; Walsh, C.T. Dissection of the EntF Condensation Domain Boundary and Active Site Residues in Nonribosomal Peptide Synthesis. Biochemistry 2003, 42, 1334–1344. [Google Scholar] [CrossRef] [PubMed]
- Sezonov, G.; le Joseleau-Petit, D.; Ari, R.D.’. Escherichia coli Physiology in Luria-Bertani Broth. J. Bacteriol. 2007, 189, 8746–8749. [Google Scholar] [CrossRef]
- Almeida, C.; Azevedo, N.F.; Santos, S.; Keevil, C.W.; Vieira, M.J. Discriminating Multi-Species Populations in Biofilms with Peptide Nucleic Acid Fluorescence in Situ Hybridization (PNA FISH). PLoS ONE 2011, 6, e14786. [Google Scholar] [CrossRef]
- O’toole, G.A.; Kolter, R. Initiation of Biofilm Formation in Pseudomonas fluorescens WCS365 Proceeds via Multiple, Convergent Signalling Pathways: A Genetic Analysis. Mol. Microbiol. 1998, 28, 449–461. [Google Scholar] [CrossRef]
- Hockett, K.L.; Baltrus, D.A. Use of the Soft-Agar Overlay Technique to Screen for Bacterially Produced Inhibitory Compounds. J. Vis. Exp. 2017, 119, e55064. [Google Scholar] [CrossRef]
- Alsop, G.M.; Waggy, G.T.; Conway, R.A. Bacterial Growth Inhibition Test. J. Water Pollut. Control. Fed. 1980, 25, 2452–2456. [Google Scholar]
- Goebel, W.F.; Barry, G.T.; Shedlovsky, T.; Colicine, K.I. The Production of Colicine K in Media Maintained at Constant pH. J. Exp. Med. 1956, 103, 577–588. [Google Scholar] [CrossRef] [PubMed]
- Neilands, B.B. Universal Chemical Assay for the Detection and Determination of Siderophores. Anal. Biochem. 1987, 160, 47–56. [Google Scholar]
- Noriega, N.C.; Kohama, S.G.; Urbanski, H.F. Microarray Analysis of Relative Gene Expression Stability for Selection of Internal Reference Genes in the Rhesus Macaque Brain. BMC Mol. Biol. 2010, 11, 47. [Google Scholar] [CrossRef] [Green Version]
- Stepanovic, S.; Vukovic, D.; Dakic, I.; Savic, B.; Svabic-Vlahovic, M. A Modified Microtiter-Plate Test for Quantification of Staphylococcal Biofilm Formation. J. Microbiol. Methods 2000, 40, 175–179. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2-ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Donlan, R.M. Biofilms: Microbial Life on Surfaces. Emerg. Infect. Dis. 2002, 8, 881–890. [Google Scholar] [CrossRef]
- Xavier, J.B.; Foster, K.R. Cooperation and Conflict in Microbial Biofilms. Proc. Natl. Acad. Sci. USA 2007, 104, 876–881. [Google Scholar] [CrossRef]
- Smarda, J.; Smajs, D. Colicins Exocellular Lethal Proteins of Escherichia coli. Folia Microbiol. Praha 1998, 43, 563–582. [Google Scholar] [CrossRef]
- Schaible, U.E.; Kaufmann, S.H.E. Iron and Microbial Infection. Nat. Rev. Microbiol. 2004, 2, 946–953. [Google Scholar] [CrossRef]
- Schalk, I.J. Innovation and Originality in the Strategies Developed by Bacteria To Get Access to Iron. ChemBioChem 2013, 14, 293–294. [Google Scholar] [CrossRef] [PubMed]
- Drider, D.; Rebuffat, S. Prokaryotic Antimicrobial Peptides. In Genes to Application; Springer: Berlin/Heidelberg, Germany, 2011. [Google Scholar]
- Budič, M.; Rijavec, M.; Petkovšek, Ž.; Žgur-Bertok, D. Escherichia coli Bacteriocins: Antimicrobial Efficacy and Prevalence among Isolates from Patients with Bacteraemia. PLoS ONE 2011, 6, e28769. [Google Scholar] [CrossRef] [PubMed]
- Tait, K.; Sutherland, I.W. Antagonistic Interactions amongst Bacteriocin-Producing Enteric Bacteria in Dual Species Biofilms. Appl. Microbiol. 2002, 93, 345–352. [Google Scholar] [CrossRef]
- Aoki, S.K.; Pamma, R.; Hernday, A.D.; Bickham, J.E.; Braaten, B.A.; Low, D.A. Microbiology: Contact-Dependent Inhibition of Growth in Escherichia coli. Science 2005, 309, 1245–1248. [Google Scholar] [CrossRef] [Green Version]
- Cowan, S.E.; Gilbert, E.; Liepmann, D.; Keasling, J.D.; Bioengineering Graduate Program, J.; Francisco, S. Commensal Interactions in a Dual-Species Biofilm Exposed to Mixed Organic Compounds. Appl. Environ. Microbiol. 2000, 66, 4481–4485. [Google Scholar] [CrossRef] [PubMed]
- Okabe, S.; Ito, T.; Sugita, K.; Satoh, H. Succession of Internal Sulfur Cycles and Sulfur-Oxidizing Bacterial Communities in Microaerophilic Wastewater Biofilms. Appl. Environ. Microbiol. 2005, 71, 2520–2529. [Google Scholar] [CrossRef]
- Werner, E.; Roe, F.; Bugnicourt, A.; Franklin, M.J.; Heydorn, A.; Molin, S.; Pitts, B.; Stewart, P.S. Stratified Growth in Pseudomonas aeruginosa Biofilms. Appl. Environ. Microbiol. 2004, 70, 6188–6196. [Google Scholar] [CrossRef]
- Leriche, V.; Briandet, R.; Carpentier, B. Ecology of Mixed Biofilms Subjected Daily to a Chlorinated Alkaline Solution: Spatial Distribution of Bacterial Species Suggests a Protective Effect of One Species to Another. Environ. Microbiol. 2003, 5, 64–71. [Google Scholar] [CrossRef]
- Rao, D.; Webb, J.S.; Kjelleberg, S. Competitive Interactions in Mixed-Species Biofilms Containing the Marine Bacterium Pseudoalteromonas tunicata. Appl. Environ. Microbiol. 2005, 71, 1729–1736. [Google Scholar] [CrossRef] [Green Version]

| Biofilm Forming Capacity a | Bacteriocin Production | Colicin Genes | Siderophore Synthesis c | ||
|---|---|---|---|---|---|
| Strain | Planktonic Culture b | Biofilm b | |||
| NCTC 12900 | - | - | ND | Ia; Ib | + |
| CECT 4267 | - | - | ND | Ia; Ib | + |
| CECT 4782 | - | - | ND | B; M | + |
| CECT 4783 | + | - | - | Ia; Ib; B; M | + |
| CECT 5947 | - | - | ND | Ia; Ib; B; M | + |
| K12 | - | - | ND | - | - |
| CECT 352 | + | - | - | - | + |
| CECT 434 | ++ | + | - | - | + |
| CECT 504 | ++ | + | + | E1-3; E6; E8; E9; B; M | - |
| CECT 515T | - | - | ND | Ia; Ib | + |
| CECT 533 | + | - | - | - | + |
| CECT 727 | - | + | - | E1-3; E6; E8; E9; B; M | + |
| CECT 730 | + | + | - | - | + |
| CECT 736 | - | - | ND | - | - |
| CECT 740 | + | - | - | Ia; Ib; B; M | + |
| CECT 744 | ++ | - | - | - | + |
| CECT 832 | - | + | ND | Ia; Ib; B; M | - |
| CECT 4537 | - | - | ND | Ia; Ib; B; M | + |
| CECT 4555 | - | + | ND | 5; 10; B; M | + |
| N5 | + | + | - | Ia; Ib; 5; 10; B; M | + |
| Growth Inhibition Zone Diameter (mm) | |||||||
|---|---|---|---|---|---|---|---|
| Bacteria Lawns | |||||||
| Spent Media | E. coli K12 | E. coli CECT 352 | E. coli CECT 736 | E. coli CECT 744 | E. coli NCTC 12900 | S. dysenteriae ATCC 11335 | |
| 24 h-biofilm | CECT 504 | 10.40 ± 2.26 | 6.00 ± 0.00 | 5.33 ± 0.47 | 7.00 ± 0.00 | 6.00 ± 0.00 | * |
| CECT 434 | - | - | - | - | - | - | |
| 24 h-planktonic culture | CECT 504 | 9.30 ± 1.06 | 6.33 ± 0.29 | - | - | - | * |
| CECT 434 | - | 4.00 ± 0.00 | 4.83 ± 0.29 | 4.17 ± 0.29 | 4.83 ± 0.29 | - | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fokt, H.; Cleto, S.; Oliveira, H.; Araújo, D.; Castro, J.; Cerca, N.; Vieira, M.J.; Almeida, C. Bacteriocin Production by Escherichia coli during Biofilm Development. Foods 2022, 11, 2652. https://doi.org/10.3390/foods11172652
Fokt H, Cleto S, Oliveira H, Araújo D, Castro J, Cerca N, Vieira MJ, Almeida C. Bacteriocin Production by Escherichia coli during Biofilm Development. Foods. 2022; 11(17):2652. https://doi.org/10.3390/foods11172652
Chicago/Turabian StyleFokt, Hanna, Sara Cleto, Hugo Oliveira, Daniela Araújo, Joana Castro, Nuno Cerca, Maria João Vieira, and Carina Almeida. 2022. "Bacteriocin Production by Escherichia coli during Biofilm Development" Foods 11, no. 17: 2652. https://doi.org/10.3390/foods11172652
APA StyleFokt, H., Cleto, S., Oliveira, H., Araújo, D., Castro, J., Cerca, N., Vieira, M. J., & Almeida, C. (2022). Bacteriocin Production by Escherichia coli during Biofilm Development. Foods, 11(17), 2652. https://doi.org/10.3390/foods11172652










