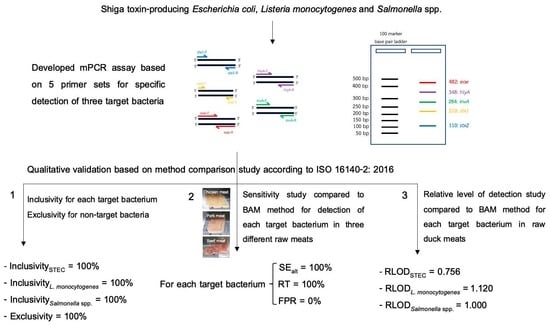
In-House Validation of Multiplex PCR for Simultaneous Detection of Shiga Toxin-Producing Escherichia coli, Listeria monocytogenes and Salmonella spp. in Raw Meats
Abstract
:1. Introduction
2. Materials and Methods
2.1. Bacterial Strains and Culture Conditions
2.2. DNA Extraction and the Optimized Multiplex PCR Conditions
2.3. Qualitative Validation Study
2.3.1. Inclusivity and Exclusivity Studies
2.3.2. Sensitivity Study
2.3.3. Relative Level of Detection Study
3. Results
3.1. Inclusivity and Exclusivity
3.2. Sensitivity Study
3.3. Relative Level of Detection Study
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Naditz, A.L.; Dzieciol, M.; Wagner, M.; Schmitz-Esser, S. Plasmids contribute to food processing environment—associated stress survival in three Listeria monocytogenes ST121, ST8, and ST5 strains. Int. J. Food Microbiol. 2019, 299, 39–46. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Costa, M.; Sucari, A.; Epszteyn, S.; Oteiza, J.; Gentiluomo, J.; Melamed, C.; Figueroa, Y.; Mingorance, S.; Grisaro, A.; Spioussas, S.; et al. Comparison of six commercial systems for the detection of non-O157 STEC in meat and vegetables. Food Microbiol. 2019, 84, 103273. [Google Scholar] [CrossRef] [PubMed]
- Trudelle, D.M.; Bryan, D.W.; Hudson, L.K.; Denes, T.G. ross-resistance to phage infection in Listeria monocytogenes serotype 1/2a mutants. Food Microbiol. 2019, 84, 103239. [Google Scholar] [CrossRef] [PubMed]
- Vico, J.P.; Lorenzutti, A.M.; Zogbi, A.P.; Aleu, G.; Sánchez, I.C.; Caffer, M.I.; Rosmini, M.R.; Mainar-Jaimed, R.C. Prevalence, associated risk factors, and antimicrobial resistance profiles of non-typhoidal Salmonella in large scale swine production in Córdoba. Vet. Sci. Res. J. 2020, 130, 161–169. [Google Scholar] [CrossRef]
- Xu, Y.G.; Liu, Z.M.; Zhang, B.Q.; Qu, M.; Mo, C.S.; Luo, J.; Li, S.L. Development of a novel target-enriched multiplex PCR (Tem-PCR) assay for simultaneous detection of five foodborne pathogens. Food Control. 2016, 64, 54–59. [Google Scholar] [CrossRef]
- Hu, A.; Cancan, G.; Zhaoxin, L.; Fengxia, L.; Liangyu, K.; Xiaomei, B. Detection of Exiguobacterium spp. and E. acetylicum on fresh-cut leafy vegetables by a multiplex PCR assay. J. Microbiol. Methods 2021, 180, 106100. [Google Scholar] [CrossRef]
- Elizaquível, P.; Maryam, A.; Gloria, S.; Rosa, A. Evaluation of Zataria multiflora Boiss. essential oil activity against Escherichia coli O157:H7, Salmonella enterica and Listeria monocytogenes by propidium monoazide quantitative PCR in vegetables. Food Control. 2013, 34, 770–776. [Google Scholar] [CrossRef]
- Garrido, A.; María-José, C.; Belén, R.; Paula, F.; Jorge, L.; Juan, M.V.; Ana, G.C. A new multiplex real-time PCR developed method for Salmonella spp. and Listeria monocytogenes detection in food and environmental samples. Food control. 2013, 30, 76–85. [Google Scholar] [CrossRef]
- Hasapa, L.; Thanakiatkraia, P.; Linacreb, A.; Kitpipit, T. Heptaplex-direct PCR assay for simultaneous detection of foodborne pathogens. Forensic Sci. Int. Genet. Suppl. Ser. 2017, 6, 173–174. [Google Scholar] [CrossRef] [Green Version]
- Kotzekidou, P. Survey of Listeria monocytogenes, Salmonella spp. and Escherichia coli O157:H7 in raw ingredients and ready-to-eat products by commercial real-time PCR kits. Food Microbiol. 2013, 35, 86–91. [Google Scholar] [CrossRef]
- Yang, Y.; Feng, X.; Hengyi, X.; Zoraida, P.; Aguilar, C.; Ruijiang, N.; Yong, Y.; Jichang, S.; Xingyong, Y.; Weihua, L.; et al. Magnetic nano-beads based separation combined with propidium monoazide treatment and multiplex PCR assay for simultaneous detection of viable Salmonella Typhimurium, Escherichia coli O157: H7 and Listeria monocytogenes in food products. Food Microbiol. 2013, 34, 418–424. [Google Scholar] [CrossRef]
- Damkerng, B.; Supawasit, W.; Trevanich, S. A new single-tube platform of melting temperature curve analysis based on multiplex real-time PCR using EvaGreen for simultaneous screening detection of Shiga toxin-producing Escherichia coli, Salmonella spp. and Listeria monocytogenes in food. Food Control 2018, 94, 195–204. [Google Scholar]
- International Organization for Standardization. ISO 16140-2, Microbiology of the Food Chain-Method Validation-Part 2: Protocol for the Validation of Alternative (Proprietary) Methods Against a Reference Method; ISO: Geneva, Switzerland, 2016. [Google Scholar]
- Official Methods of Analysis, 20th ed.; Appendix J [Online]; AOAC International: Rockville, MD, USA, 2016; Available online: http://www.eoma.aoac.org/app_j.pdf (accessed on 28 June 2020).
- Cetin, E.; Temelli, S.; Eyigor, A. Salmonella prevalence and serovar distribution in healthy slaughter sheep and cattle determined by ISO 6579 and VIDAS UP Salmonella methods. J. Food Sci. Technol. 2019, 56, 5317–5325. [Google Scholar] [CrossRef] [PubMed]
- D’Agostino, M.; Robles, S.; Hansen, F.; Ntafis, V.; Ikonomopoulos, J.; Kokkinos, P.; Alvarez-Ordonez, A.; Jordan, K.; Delibato, E.; Kukier, E.; et al. Validation of a loop-mediated amplification/ISO 6579-based method for analysing soya meal for the presence of Salmonella enterica. Food Anal. Methods. 2016, 9, 2979–2985. [Google Scholar] [CrossRef]
- Labrador, M.; Rota, M.C.; Perez-Arquillue, C.; Herrera, A.; Bayarri, S. Comparative evaluation of impedanciometry combined with chromogenic agars or RNA hybridization and real-time PCR methods for the detection of L. monocytogenes in dry-cured ham. Food Control 2018, 94, 108–115. [Google Scholar] [CrossRef] [Green Version]
- Supawasit, W.; Bundidamorn, D.; Trevanich, S. Effect of K2HPO4-modified buffer in simultaneous enrichment broth for improvement of multiplex PCR for detection of Shiga toxin-producing Escherichia coli, Salmonella spp. and Listeria monocytogenes. In Proceedings of the 6th International conference on creative technology, Pattaya, Thailand, 24–26 July 2018. [Google Scholar]
- Meng, J.; Zhao, S.; Doyle, P.M.; Mitchell, E.S.; Kresovich, S. A multiplex PCR for identifying Shiga like toxin-producing Escherichia coli O157:H7. Lett. Appl. 1997, 24, 172–176. [Google Scholar] [CrossRef]
- Pal, A.; Ghosh, S.; Ramamurthy, T.; Yamasaki, S.; Tsukamoto, T.; Bhattacharya, S.K.; Balakrish Nair, G.; Takeda, Y. Shiga toxin-producing Escherichia coli from healthy cattle in a semi-urban community in Calcutta, India. Indian J. Med. Res. 1999, 110, 83–85. [Google Scholar]
- Vidal, R.; Vidal, M.; Lagos, R.; Levine, M.; Prado, V. Multiplex PCR for Diagnosis of Enteric Infections Associated with Diarrheagenic Escherichia coli. J. Clin. Microbiol. 2004, 42, 1787–1789. [Google Scholar] [CrossRef] [Green Version]
- Liu, P.; Mizue, H.; Fujihara, K.; Kobayashi, H.; Kamikado, H.; Tanaka, T.; Honjoh, K.; Miyamoto, T. A new rapid real-time PCR method for detection of Listeria monocytogenes targeting the hlyA gene. Food Sci. Technol. Res. 2012, 18, 47–57. [Google Scholar] [CrossRef] [Green Version]
- Rahn, K.; Grandis, S.A.; Clarke, R.C.; McEwen, S.A.; Galin, J.E.; Ginocchio, C.; Curtiss, R.; Gyles, C.L. Amplification of an invA gene sequence of Salmonella Typhimurium by polymerase chain reaction as a specific method of Salmonella. Mol. Cell. Probes. 1992, 6, 271–279. [Google Scholar] [CrossRef]
- Li, B.; Chen, J.Q. Real-time PCR methodology for selective detection of viable Escherichia coli O157:H7 cells by targeting Z3276 as a genetic marker. Appl. Environ. 2012, 78, 5297–5304. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Detection and Enumeration of Listeria monocytogene. In Bacteriological Analytical Manual; U.S. Food and Drug Administration: Silver Spring, MD, USA, 2017. Available online: https://www.fda.gov/food/laboratory-methods-food/bam-detection-and-enumeration-listeria-monocytogenes (accessed on 28 May 2020).
- Enumeration of Escherichia coli and the Coliform Bacteria. In Bacteriological Analytical Manual; U.S; Food and Drug Administration: Silver Spring, MD, USA, 2017. Available online: https://www.fda.gov/food/laboratory-methods-food/bam-4-enumeration-escherichia-coli-and-coliform-bacteria (accessed on 28 May 2020).
- Salmonella . In Bacteriological Analytical Manual; U.S. Food and Drug Administration: Silver Spring, MD, USA, 2018. Available online: https://www.fda.gov/food/laboratory-methods-food/bacteriological-analytical-manual-bam-chapter-5-salmonella (accessed on 28 May 2020).
- Department of Medical Sciences Thailand (Thailand Ministry of Public Health, Nonthaburi, Thailand). DMST Bacteriology Culture Guide. 2016; Unpublished work. (In Thai)
- Benie, C.K.D.; Dadié, A.; Guessennd, N.; N’gbesso-Kouadio1, N.A.; Kouame, N.D.; N’golo, D.C.; Aka, S.; Dak, E.; Dje, K.M.; Dosso, M. Characterization of virulence potential of Pseudomonas aeruginosa isolated from bovine meat, fresh fish, and smoked fish. Eur. J. Microbiol. Immunol. 2017, 7, 55–64. [Google Scholar] [CrossRef] [Green Version]
- Doulgeraki, A.I.; Ercolini, D.; Villani, F.; Nychas, G.J.E. Spoilage microbiota associated to the storage of raw meat in different conditions. Int. J. Food Microbiol. 2012, 157, 130–141. [Google Scholar] [CrossRef] [PubMed]
- Gu, X.; Sun, Y.; Tu, K.; Dong, Q.; Pan, L. Predicting the growth situation of Pseudomonas aeruginosa on agar plates and meat stuffs using gas sensors. Sci. Rep. 2016, 6, 38721. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, G.; Ma, F.; Zeng, L.; Bai, Y.; Wang, H.; Xu, X.; Zhou., G. Modified Atmosphere Packaging Decreased Pseudomonas Fragi Cell T Metabolism and Extracellular Proteolytic Activities on Meat. Food Microbiol. 2018, 76, 443–449. [Google Scholar] [CrossRef]
- Aerobic Plate Count. In Bacteriological Analytical Manual; U.S. Food and Drug Administration: Silver Spring, MD, USA, 2001. Available online: https://www.fda.gov/food/laboratory-methods-food/bam-chapter-3-aerobic-plate-count (accessed on 28 May 2020).
- Delibato, E.; Rodriguez-Lazaro, D.; Gianfranceschi, M.; De Cesare, A.; Comin, D.; Gattuso, A.; Hernandez, M.; Sonnessa, M.; Pasquali, F.; Sreter-Lancz, Z.; et al. European validation of Real-Time PCR method for detection of Salmonella spp. in pork meat. Int. J. Food Microbiol. 2014, 184, 134–138. [Google Scholar] [CrossRef]
- Maurischat, S.; Baumann, B.; Martin, A.; Malorn, B. Rapid detection and specific differentiation of Salmonella enterica subsp. Enteritidis, Typhimurium and its monophasic variant 4,[5],12:i:- by real-time multiplex PCR. Int. J. Food Sci. 2015, 193, 8–14. [Google Scholar] [CrossRef]
- Balière, C.; Rincé, A.; Delannoy, S.; Fach, P.; Gourmelon, M. Molecular profiling of Shiga toxin-producing Escherichia coli and enteropathogenic E. coli strains isolated from french coastal environments. Appl. Environ. Microbiol. 2016, 82, 3913–3927. [Google Scholar] [CrossRef] [Green Version]
- Brusa, V.; Galli, L.; Linares, L.H.; Ortega, E.E.; Lirón, J.P.; Leotta, G. Development and validation of two SYBR green PCR assays and a multiplex real-time PCR for the detection of Shiga toxin-producing Escherichia coli in meat. J. Microbiol. Methods. 2015, 119, 10–17. [Google Scholar] [CrossRef]
- Chiang, Y.U.; Wang, H.H.; Ramireddy, L.; Chen, H.Y.; Shih, C.M.; Lin, C.K.; Tsen, H.Y. Designing a biochip following multiplex polymerase chain reaction for the detection of Salmonella serovars Typhimurium, Enteritidis, Infantis, Hadar, and Virchow in poultry products. J. Food Drug Anal. 2018, 26, 58–66. [Google Scholar] [CrossRef]
- Hu, Q.; Coburn, B.; Deng, W.; Li, Y.; Shi, X.; Lan, Q.; Wang, B.; Coombes, B.K.; Finlay, B.B. Salmonella enterica serovar Senftenberg human clinical isolates lacking SPI-1. J. Clin. Microbiol. 2008, 46, 1330–1336. [Google Scholar] [CrossRef] [PubMed] [Green Version]
| Number of Strains | Target Bacterial Strains |
|---|---|
| 55 | Shiga toxin-producing Escherichia coli (STEC a) DMST b 48719; STEC DMST 50662; STEC PSU c 5023; STEC DMST 19341; STEC DMST 50661; STEC PSU 38; STEC PSU 5030; STEC DMST 50660; STEC PSU 4189; STEC PSU 4159; STEC DMST 30538; STEC DMST 30536; STEC DMST 30537; STEC PSU 149; STEC PSU 150; STEC DMST 19340; STEC DMST 19342; STEC DMST 30539; STEC DMST 50659; STEC PSU 133; STEC PSU 135; STEC PSU 148; STEC PSU 4153; STEC PSU 4154; STEC PSU 4155; STEC PSU 4156; STEC PSU 4157; STEC PSU 4158; STEC PSU 4160; STEC PSU 4161; STEC PSU 4162; STEC PSU 4163; STEC PSU 4164; STEC PSU 4165; STEC PSU 4166; STEC PSU 4167; STEC PSU 4169; STEC PSU 4170; STEC PSU 4171; STEC PSU 4172; STEC PSU 4173; STEC PSU 4191; STEC PSU 4192; STEC PSU 4193; STEC PSU 4196; STEC PSU 4197; STEC PSU 5026; STEC PSU 5027; STEC PSU 5028; STEC PSU 5029; STEC PSU 3802; STEC PSU 4190; STEC PSU 4195; STEC PSU 4198 and STEC CDC d 03-3014 |
| 50 | L. monocytogenes ATCC e 7644; L. monocytogenes Li 23 ATCC 19114 L. monocytogenes Li 2107 ATCC 19116; L. monocytogenes DMST 41455; L. monocytogenes DMST 17303; L. monocytogenes DMST 20093; L. monocytogenes DMST 20422; L. monocytogenes DMST 20423; L. monocytogenes DMST 20425; L. monocytogenes DMST 21164; L. monocytogenes DMST 21165; L. monocytogenes DMST 23136; L. monocytogenes DMST 23146; L. monocytogenes DMST 23150; L. monocytogenes DMST 23151; L. monocytogenes DMST 23710; L. monocytogenes DMST 27738; L. monocytogenes DMST 31799; L. monocytogenes DMST 31800; L. monocytogenes DMST 31801; L. monocytogenes DMST 31802; L. monocytogenes DMST 31804; L. monocytogenes DMST 32862; L. monocytogenes DMST 33253; L. monocytogenes DMST 36156; L. monocytogenes DMST 37884; L. monocytogenes DMST 37885; L. monocytogenes DMST 41456; L. monocytogenes DMST 41457; L. monocytogenes DMST 41458; L. monocytogenes DMST 41642; L. monocytogenes DMST 41647; L. monocytogenes DMST 44932; L. monocytogenes DMST 44933; L. monocytogenes DMST 44934; L. monocytogenes DMST 44935; L. monocytogenes DMST 44936; L. monocytogenes DMST 45433; L. monocytogenes DMST 45683; L. monocytogenes DMST 45984; L. monocytogenes DMST 46456; L. monocytogenes DMST 47501; L. monocytogenes DMST 47502; L. monocytogenes DMST 47503; L. monocytogenes DMST 50339; L. monocytogenes 100; L. monocytogenes 101; L. monocytogenes 108; L. monocytogenes 310 and L. monocytogenes Scott A |
| 102 | S. Aberdeen DMST 19198; S. Abony PVKU 1 f; S. Agona DMST 23970; S. Alachua DMST 19203; S. Albany DMST 50696; S. Altona DMST 62226; S. Amsterdam WPKU 1 g; S. Anatum DMST 50705; S. Apeyeme PVKU 2; S. Augustenborg DMST 50631; S. Bangkok DMST 50834; S. Bareilly DMST 62231; S. Bergen DMST 19206; S. Blockley DMST 16821; S. Bongori ATCC 43975; S. Bovismorbificans DMST 17379; S. Braenderup DMST 62234; S. Bredeney PVKU 3; S. Canstatt PVKU 4; S. Cerro DMST 19200; S. Chester WPKU 2; S. Chicago KU 1 h; S. Corvalis KU 2; S. Derby DMST 16880; S. Dublin WPKU 3; S. Eastbourne WPKU 4; S. Emek WPKU 5; S. enterica subsp. Salamae ser, 17:gt: DMST 19207; S. Adelaide ATCC 10718; S. Bispebjerg ATCC 9842; S. Choleraesuis ATCC 6958; S. Gaminara ATCC 8324; S. Heerlen ATCC 15792; S. Hillingdon ATCC 9184; S. Illinois ATCC 11646; S. Inverness ATCC 10720; S. Kirkee ATCC 8322; S. Oranienburg ATCC 9239; S. Pullorum ATCC 9120; S. Simsbury ATCC 12004; S. Vellore ATCC 15611; S. Zwickau ATCC 15805; S. Dar-es-salaam ATCC 6959; S. Hooggraven ATCC 15786; S. Enteritidis DMST 15676; S. Falkensee DMST 50716; S. Fresno DMST 19197; S. Give DMST 50827; S. Hadar DMST 32769; S. Havana DMST 50710; S. Heidelberg PVKU 5; S. Hvittingfoss DMST 62220; S. Indiana PVKU 6; S. Infatis PVKU 7; S. I4,12:i:- PVKU 8; S. Johnnesburg DMST 50835; S. Kedougou DMST 33890; S. Kentucky DMST 50701; S. KiamBu PVKU 9; S. Krefeld DMST 62227; S. Lexington DMST 50707; S. Liverpool PVKU 10; S. Livingstone DMST 50633; S. London DMST 62232; S. Manhattan PVKU 11; S. Matopeni DMST 62218; S. Mbandaka DMST 62238; S. Minnesota KU 3; S. Molade PVKU 12; S. Montevideo PVKU 13; S. Moscow PVKU 14; S. Muenchen PVKU 15; S. Muenster DMST 62235; S. Newport DMST 15675; S. Orion WPKU 6; S. Oslo WPKU 7; S. Ouakam DMST 50824; S. Panama DMST 50703; S. Paratyphi A DMST 15673; S. Paratyphi B WPKU 8; S. Paratyphi C WPKU 9; S. Poona KU 4; S. Ramat-gan WPKU 10; S. Rissen DMST 16876; S. Rubislaw DMST 62223; S. Saintpaul DMST 62225; S. Schwarzengrund WPKU 11; S. Typhi WPKU 12; S. Singapore DMST 50636; S. Stanley DMST 33894; S. Tennessee WPKU 13; S. Thempson PVKU 16; S. Typhimurium DMST 562; S. Brunei KU 5; S. Virchow DMST 16857; S. Wandsworth DMST 19204; S. Warthington DMST 33889; S. Waycross DMST 19205; S. Weltevreden DMST 16820; S. Urbana PVKU 17; S. Soerenga PVKU 18 and S. Weston PVKU 19 |
| Number of Strains | Non-Target Bacterial Strains |
|---|---|
| 30 | Aeromonas hydrophila DMST a 21250; Bacillus cereus DMST 5040; Campylobacter coli DMST 18034; Campylobacter jejuni DMST 15190; Citrobacter freundii DMST 16368; Cronobacter sakazakii DMST 17894; Enterobacter cloacae DMST 434; Enterococcus faecalis DMST 4736; Klebsiella pneumonia DMST 8216; Lactococcus lactis KU 6 b; Lactobacillus brevis KU 7; Listeria innocua DMST 9011; Listeria ivanovii DMST 9012; Pediococcus pentosaceus DMST 18752; Proteus vulgaris DMST 557; Pseudomonas aeruginosa DMST 4739; Pseudomonas fluorescens KU 8; Shigella dysenteriae DMST 15111; Shigella flexneri DMST 4423; Shigella sonnei DMST 561; Staphylococcus aureus DMST 8840; Streptococcus pyogenes DMST 30653; Streptococcus suis serotype II DMST 18783; Vibrio cholerae nonO1/nonO139 DMST 2873; Vibrio parahaemolyticus DMST 21243; Vibrio vulnificus DMST 21245; Yersinia enterocolitica DMST 8012; Yersinia pseudotubercolosis DMST 16385; Bacillus subtilis (Ehrenberg) Cohn BCC c 6327 and Staphylococcus epidermidis KU 9 |
| Primer | Primer Sequence (5′–3′) | Target Gene | Target | Product Size (bp) | Reference |
|---|---|---|---|---|---|
| stx1-F stx1-R | TGTAACTGGAAAGGTGGAGTATACA GCTATTCTGAGTCAACGAAAAATAAC | stx1 | STEC | 210 | [19] |
| stx2-F stx2-R | ATCAGTCGTCACTCACTGGT CTGCTGTCACAGTGACAAA | stx2 | STEC | 110 | [20] |
| eae-F eae-R | TCAATGCAGTTCCGTTATCAGTT GTAAAGTCCGTTACCCCAACCTG | eae | STEC | 482 | [21] |
| hlyA-F hlyA-R | AAATCATCGACGGCAACCT GGACGATGTGAAATGAGC | hlyA | L. monocytogenes | 348 | [22] |
| invA-F invA-R | GTGAAATTATCGCCACGTTCGGGCAA TCATCGCACCGTCAAAGGAACC | invA | Salmonella spp. | 284 | [23] |
| IAC-F IAC-R | CAGGATTGACAGAGCGAGGTATG CGTAGTTAGGCCACCACTTCAAG | ori | pUC19 | 65 | [24] |
| Number of Strains | Bacterial Strains | Test Result | |
|---|---|---|---|
| Target bacterial strains | 3 | Shiga toxin-producing Escherichia coli (STEC a) DMST b 48719; STEC DMST 50662 and STEC PSU c 5023 | Positive for stx1 gene e (product size 210 bp) |
| 4 | STEC DMST 19341; STEC DMST 50661; STEC PSU 38 and STEC PSU 5030 | Positive for stx2 gene f (product size 110 bp) | |
| 3 | STEC DMST 50660; STEC PSU 4189 and STEC PSU 4159 | Positive for stx1 and stx2 genes | |
| 5 | STEC DMST 30538; STEC DMST 30536; STEC DMST 30537; STEC PSU 149 and STEC PSU 150 | Positive for stx1 and eae g genes (product size 482 bp) | |
| 35 | STEC DMST 19340; STEC DMST 19342; STEC DMST 30539; STEC DMST 50659; STEC PSU 133; STEC PSU 135; STEC PSU 148; STEC PSU 4153; STEC PSU 4154; STEC PSU 4155; STEC PSU 4156; STEC PSU 4157; STEC PSU 4158; STEC PSU 4160; STEC PSU 4161; STEC PSU 4162; STEC PSU 4163; STEC PSU 4164; STEC PSU 4165; STEC PSU 4166; STEC PSU 4167; STEC PSU 4169; STEC PSU 4170; STEC PSU 4171; STEC PSU 4172; STEC PSU 4173; STEC PSU 4191; STEC PSU 4192; STEC PSU 4193; STEC PSU 4196; STEC PSU 4197; STEC PSU 5026; STEC PSU 5027; STEC PSU 5028 and STEC PSU 5029 | Positive for stx2 and eae genes | |
| 5 | STEC PSU 3802; STEC PSU 4190; STEC PSU 4195; STEC PSU 4198 and STEC CDC d 03-3014 | Positive for stx1, stx2 and eae genes | |
| 50 | L. monocytogenes strains in Table 1 | Positive for hlyA gene h (product size 348 bp) | |
| 102 | Salmonella spp. serotypes in Table 1 | Positive for invA gene i (product size 284 bp) | |
| Non-target bacterial strains | 30 | Non-target bacterial strains in Table 2 | Negative |
| Bacteria | Total Number of Isolates Tested | Total Number of Positive Results by the Developed mPCR Assay | Inclusivity/Exclusivity (%) |
|---|---|---|---|
| Shiga toxin-producing E. coli | 55 | 55 | 100 |
| L. monocytogenes | 50 | 50 | 100 |
| Salmonella spp. | 102 | 102 | 100 |
| non-target bacteria a | 30 | 30 | 100 |
| Parameter | Value (%) |
|---|---|
| Sensitivity for developed mPCR assay (SEalt) | 100 |
| Sensitivity for BAM method (SEref) | 100 |
| Relative trueness (RT) | 100 |
| False positive ratio (FPR) | 0 |
| Method | Positive Results of Sample Contaminated with Different Levels of Contamination | RLOD g | ||
|---|---|---|---|---|
| 0 a | 0.5 b | 1.5 c | ||
| mPCR for STEC | 0/5 d | 12/20 e | 5/5 f | 0.756 |
| BAM h for STEC | 0/5 | 10/20 | 5/5 | |
| mPCR for L. monocytogenes | 0/5 | 8/20 | 5/5 | 1.170 |
| BAM for L. monocytogenes | 0/5 | 9/20 | 5/5 | |
| mPCR for Salmonella spp. | 0/5 | 11/20 | 5/5 | 1.000 |
| BAM for Salmonella spp. | 0/5 | 11/20 | 5/5 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jaroenporn, C.; Supawasit, W.; Bundidamorn, D.; Udompijitkul, P.; Assawamakin, A.; Trevanich, S. In-House Validation of Multiplex PCR for Simultaneous Detection of Shiga Toxin-Producing Escherichia coli, Listeria monocytogenes and Salmonella spp. in Raw Meats. Foods 2022, 11, 1557. https://doi.org/10.3390/foods11111557
Jaroenporn C, Supawasit W, Bundidamorn D, Udompijitkul P, Assawamakin A, Trevanich S. In-House Validation of Multiplex PCR for Simultaneous Detection of Shiga Toxin-Producing Escherichia coli, Listeria monocytogenes and Salmonella spp. in Raw Meats. Foods. 2022; 11(11):1557. https://doi.org/10.3390/foods11111557
Chicago/Turabian StyleJaroenporn, Chanokchon, Wannakarn Supawasit, Damkerng Bundidamorn, Pathima Udompijitkul, Anunchai Assawamakin, and Sudsai Trevanich. 2022. "In-House Validation of Multiplex PCR for Simultaneous Detection of Shiga Toxin-Producing Escherichia coli, Listeria monocytogenes and Salmonella spp. in Raw Meats" Foods 11, no. 11: 1557. https://doi.org/10.3390/foods11111557
APA StyleJaroenporn, C., Supawasit, W., Bundidamorn, D., Udompijitkul, P., Assawamakin, A., & Trevanich, S. (2022). In-House Validation of Multiplex PCR for Simultaneous Detection of Shiga Toxin-Producing Escherichia coli, Listeria monocytogenes and Salmonella spp. in Raw Meats. Foods, 11(11), 1557. https://doi.org/10.3390/foods11111557