Autochthonous Enterococcus durans PFMI565 and Lactococcus lactis subsp. lactis BGBU1–4 in Bio-Control of Listeria monocytogenes in Ultrafiltered Cheese
Abstract
1. Introduction
2. Materials and Methods
2.1. Cheese Samples, Isolation of Lactic Acid Bacteria with Antilisterial Activity
2.2. Proteinaceous Nature of Antilisterial Molecule
2.3. Identification of Lactic Acid Bacteria with Antilisterial Activity
2.4. Detection of Hydrogen Peroxide
2.5. Detection of Diacetyl
2.6. Antibiotic Resistance
2.7. Acidifying Activity
2.8. UF Cheese Making Procedure
2.9. Sampling and Microbiological Analysis of Experimental UF Cheeses
2.10. Statistical Analysis
3. Results and Discussion
3.1. Isolation and Identification of LAB with Antilisterial Activity
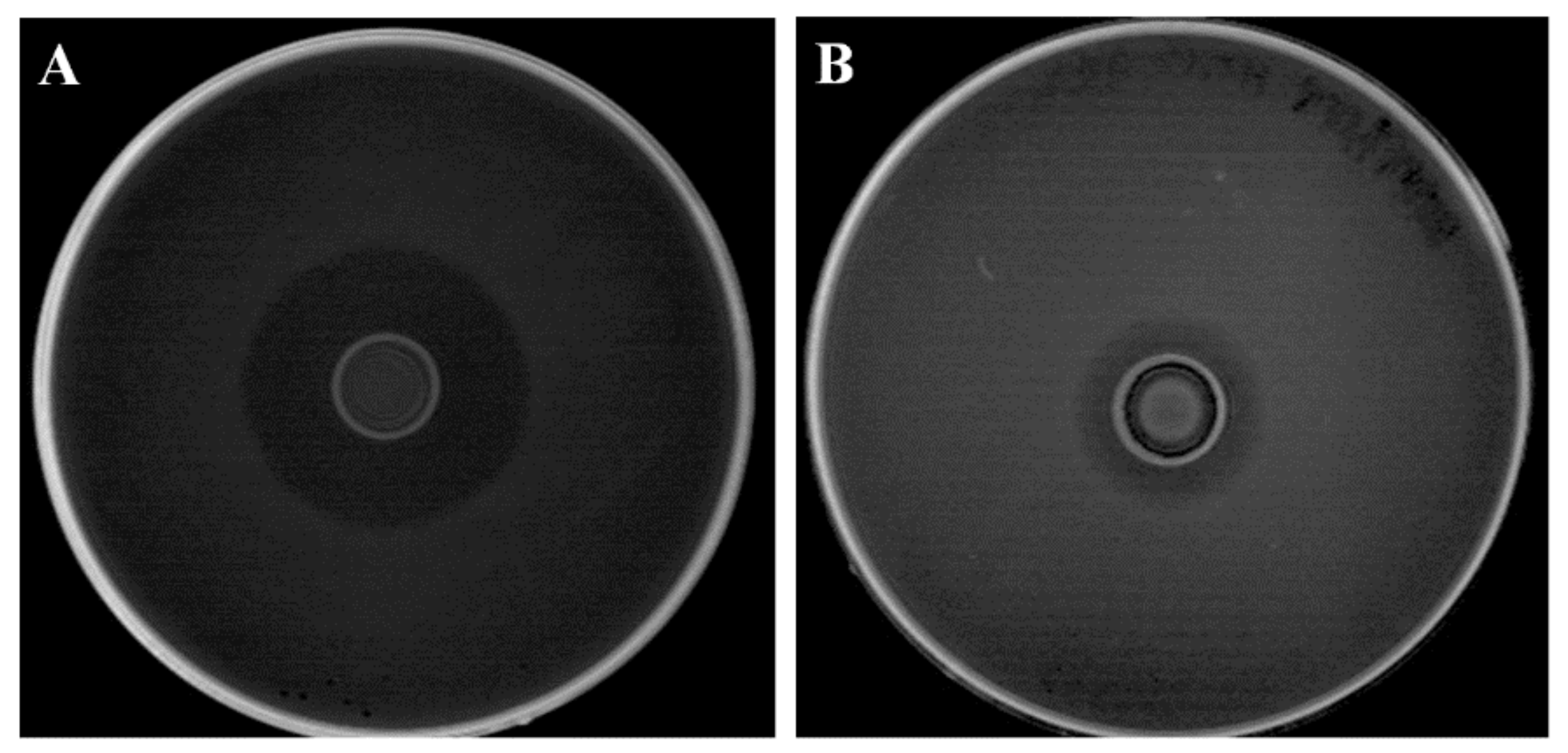
3.2. Proteinaceous Nature of Antilisterial Compounds
3.3. Hydrogen Peroxide and Diacetyl Production
3.4. Antibiotic Resistance
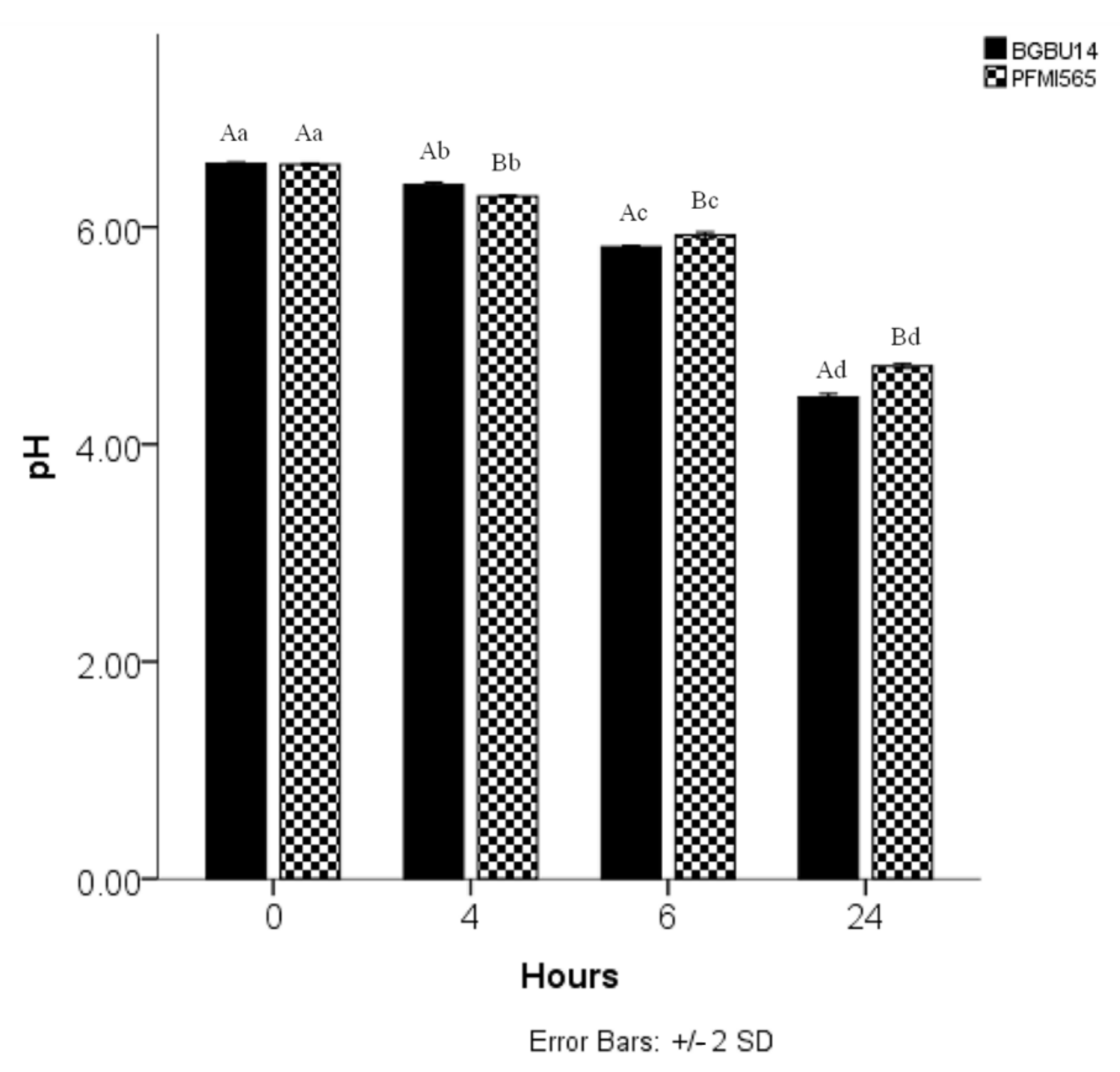
3.5. Acidifying Activity
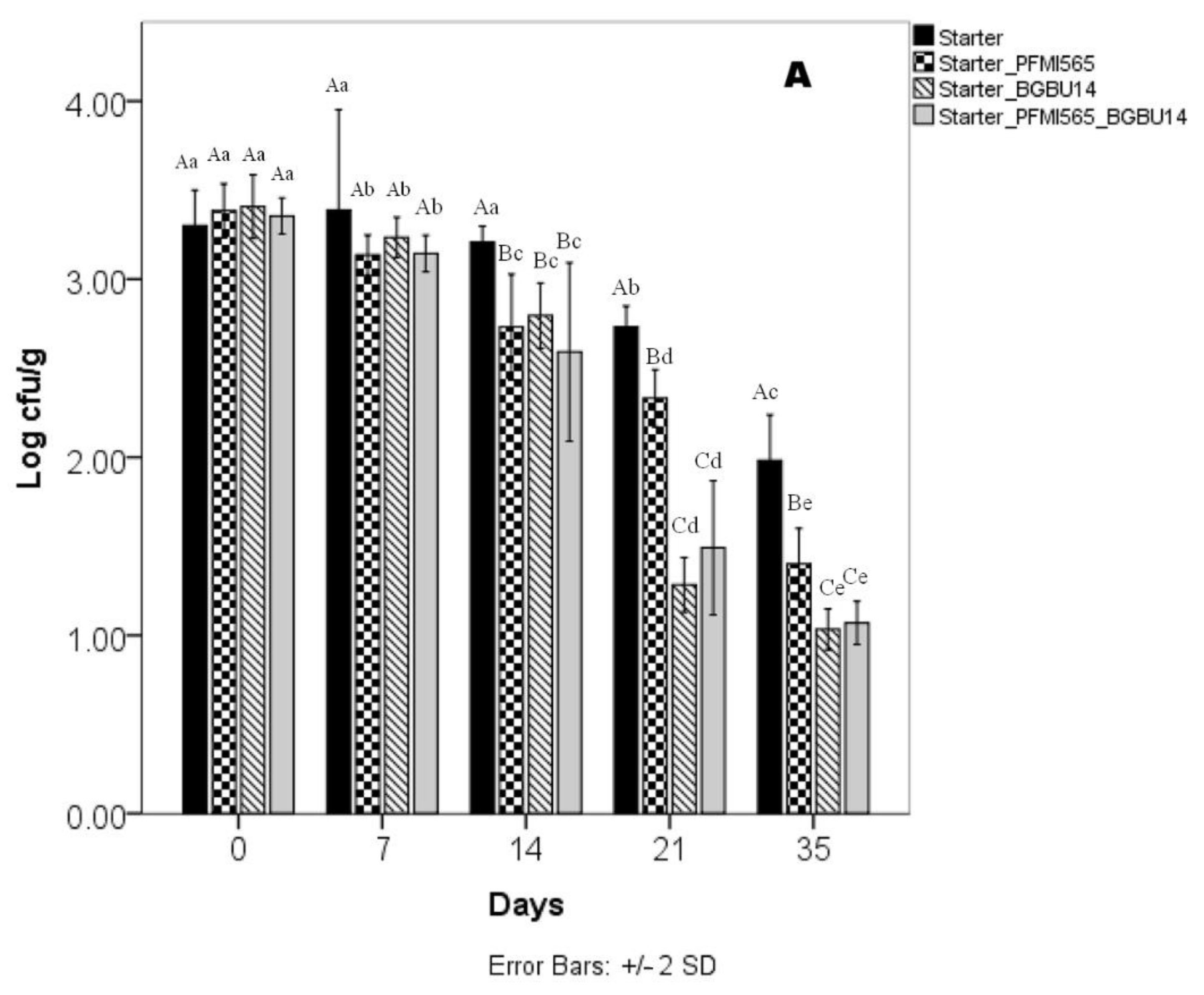
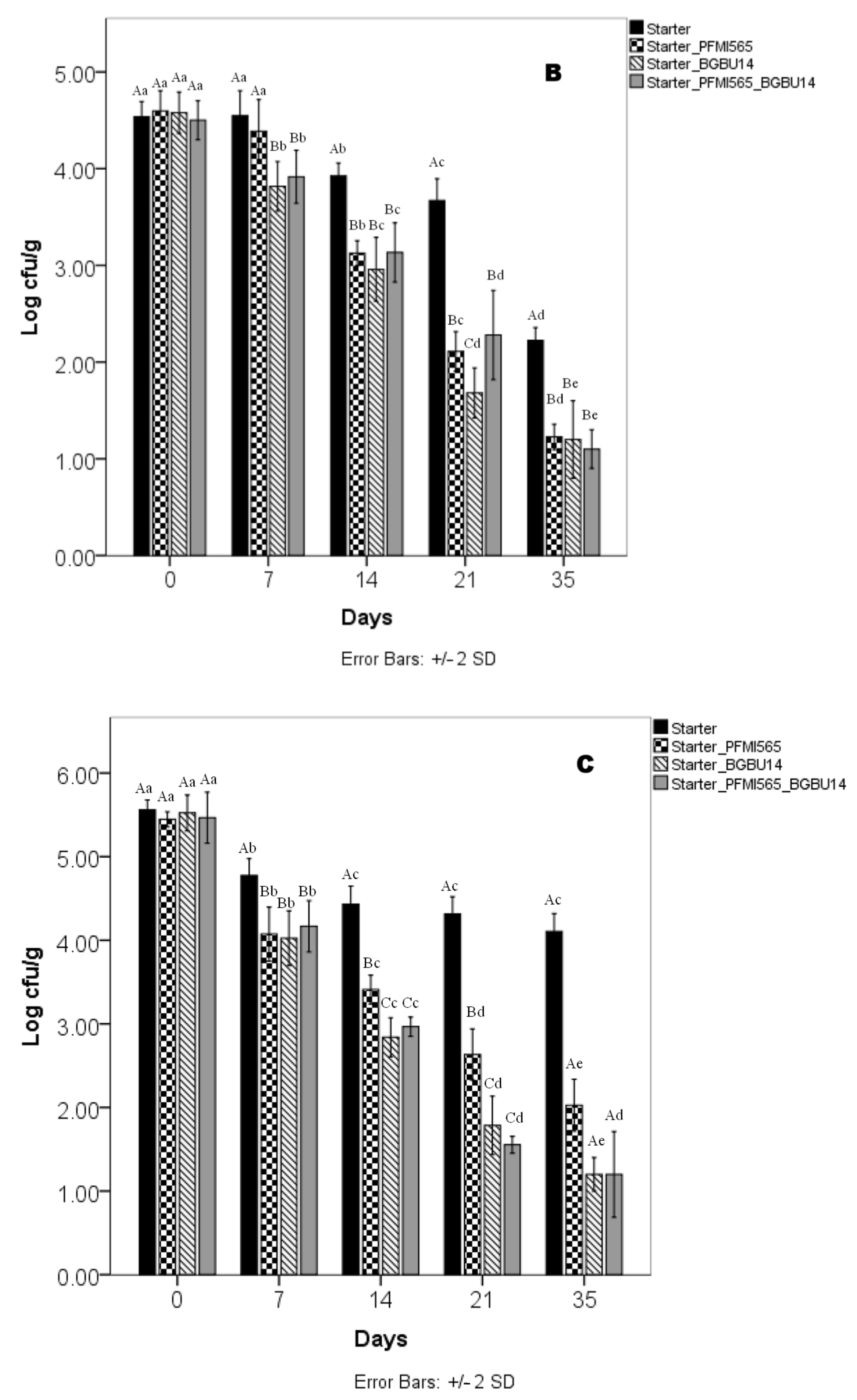
3.6. Microbiological Analysis of UF Cheeses
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Walther, B.; Schmid, A.; Sieber, R.; Wehrmüller, K. Cheese in nutrition and health. Dairy Sci. Technol. 2008, 88, 389–405. [Google Scholar] [CrossRef]
- Santiago-López, L.; Aguilar-Toalá, J.E.; Hernández-Mendoza, A.; Vallejo-Cordoba, B.; Liceaga, A.M.; González-Córdova, A.F. Invited review: Bioactive compounds produced during cheese ripening and health effects associated with aged cheese consumption. J. Dairy Sci. 2018, 101, 3742–3757. [Google Scholar] [CrossRef] [PubMed]
- Miočinović, J.; Puđa, P.; Radulović, Z.; Pavlović, V.; Miloradović, Z.; Radovanović, M.; Paunović, D. Development of low fat UF cheese technology. Hrvatska Mljekarska Udruga 2011, 61, 33–44. [Google Scholar]
- Miloradovic, Z.; Miocinovic, J.; Kljajevic, N.; Tomasevic, I.; Pudja, P. The influence of milk heat treatment on composition, texture, colour and sensory characteristics of cows’ and goats’ Quark-type cheeses. Small Rumin. Res. 2018, 169, 154–159. [Google Scholar] [CrossRef]
- Melo, J.; Andrew, P.W.; Faleiro, M.L. Listeria monocytogenes in cheese and the dairy environment remains a food safety challenge: The role of stress responses. Food Res. Int. 2015, 67, 75–90. [Google Scholar] [CrossRef]
- Thakur, M.; Asrani, R.K.; Patial, V. Listeria monocytogenes: A food-borne pathogen. In Foodborne Diseases; Elsevier: Amsterdam, The Netherlands, 2018; Volume 15, pp. 157–192. [Google Scholar]
- McLauchlin, J.; Mitchell, R.T.; Smerdon, W.J.; Jewell, K. Listeria monocytogenes and listeriosis: A review of hazard characterisation for use in microbiological risk assessment of foods. Int. J. Food Microbiol. 2004, 92, 15–33. [Google Scholar] [CrossRef]
- Vázquez-Boland, J.A.; Kuhn, M.; Berche, P.; Chakraborty, T.; Domínguez-Bernal, G.; Goebel, W.; González-Zorn, B.; Wehland, J.; Kreft, J. Listeria pathogenesis and molecular virulence determinants. Clin. Microbiol. Rev. 2001, 14, 584–640. [Google Scholar] [CrossRef]
- The European Union summary report on trends and sources of zoonoses, zoonotic agents and food-borne outbreaks in 2013. EFSA J. 2015, 13. [CrossRef]
- Regulation on the General and Special Conditions of Food Hygiene at any Stage of Production, Processing and Marketing” 33/2010. Official Bulletin of the Republic of Serbia. 2010. Available online: https://www.vet.minpolj.gov.rs/legislativa/pravilnici/Pravilnik_mikrobioloski_kriterijumi.pdf (accessed on 20 June 2021).
- Schlech, W.F. Foodborne listeriosis. Clin. Infect. Dis. 2000, 31, 770–775. [Google Scholar] [CrossRef]
- Oh, H.; Kim, S.; Lee, S.; Lee, H.; Ha, J.; Lee, J.; Choi, Y.; Choi, K.H.; Yoon, Y. Prevalence and genetic characteristics of meatborne listeria monocytogenes isolates from livestock farms in Korea. Korean J. Food Sci. Anim. Resour. 2016, 36, 779–786. [Google Scholar] [CrossRef]
- Jung, D.S.; Bodyfelt, F.W.; Daeschel, M.A. Influence of Fat and Emulsifiers on the Efficacy of Nisin in Inhibiting Listeria monocytogenes in Fluid Milk. J. Dairy Sci. 1992, 75, 387–393. [Google Scholar] [CrossRef]
- Reis, J.A.; Paula, A.T.; Casarotti, S.N.; Penna, A.L.B. Lactic Acid Bacteria Antimicrobial Compounds: Characteristics and Applications. Food Eng. Rev. 2012, 4, 124–140. [Google Scholar] [CrossRef]
- Guillier, L.; Stahl, V.; Hezard, B.; Notz, E.; Briandet, R. Modelling the competitive growth between Listeria monocytogenes and biofilm microflora of smear cheese wooden shelves. Int. J. Food Microbiol. 2008, 128, 51–57. [Google Scholar] [CrossRef]
- Rodríguez, E.; Calzada, J.; Arqués, J.L.; Rodríguez, J.M.; Nuñez, M.; Medina, M. Antimicrobial activity of pediocin-producing Lactococcus lactis on Listeria monocytogenes, Staphylococcus aureus and Escherichia coli O157:H7 in cheese. Int. Dairy J. 2005, 15, 51–57. [Google Scholar] [CrossRef]
- Gálvez, A.; López, R.L.; Abriouel, H.; Valdivia, E.; Omar, N. Ben Application of bacteriocins in the control of foodborne pathogenic and spoilage bacteria. Crit. Rev. Biotechnol. 2008, 28, 125–152. [Google Scholar] [CrossRef] [PubMed]
- Radulović, Z.; Miočinović, J.; Mirković, N.; Mirković, M.; Paunović, D.; Ivanović, M.; Seratlić, S. Survival of spray-dried and free-cells of potential probiotic Lactobacillus plantarum 564 in soft goat cheese. Anim. Sci. J. 2017, 88, 1849–1854. [Google Scholar] [CrossRef] [PubMed]
- Topisirovic, L.; Kojic, M.; Fira, D.; Golic, N.; Strahinic, I.; Lozo, J. Potential of lactic acid bacteria isolated from specific natural niches in food production and preservation. Int. J. Food Microbiol. 2006, 112, 230–235. [Google Scholar] [CrossRef] [PubMed]
- Castro, J.M.; Tornadijo, M.E.; Fresno, J.M.; Sandoval, H. Biocheese: A food probiotic carrier. Biomed. Res. Int. 2015. [Google Scholar] [CrossRef]
- Vera Pingitore, E.; Todorov, S.D.; Sesma, F.; Gombossy de Melo Franco, B.D. Application of bacteriocinogenic Enterococcus mundtii CRL35 and Enterococcus faecium ST88Ch in the control of Listeria monocytogenes in fresh Minas cheese. Food Microbiol. 2012, 32, 38–47. [Google Scholar] [CrossRef] [PubMed]
- Aspri, M.; Field, D.; Cotter, P.D.; Ross, P.; Hill, C.; Papademas, P. Application of bacteriocin-producing Enterococcus faecium isolated from donkey milk, in the bio-control of Listeria monocytogenes in fresh whey cheese. Int. Dairy J. 2017, 73, 1–9. [Google Scholar] [CrossRef]
- Dal Bello, B.; Cocolin, L.; Zeppa, G.; Field, D.; Cotter, P.D.; Hill, C. Technological characterization of bacteriocin producing Lactococcus lactis strains employed to control Listeria monocytogenes in Cottage cheese. Int. J. Food Microbiol. 2012, 153, 58–65. [Google Scholar] [CrossRef]
- Delves-Broughton, J. Nisin and its application as a food preservative. Int. J. Dairy Technol. 1990, 43, 73–76. [Google Scholar] [CrossRef]
- Rodríguez, J.M.; Martínez, M.I.; Kok, J. Pediocin PA-1, a wide-spectrum bacteriocin from lactic acid bacteria. Crit. Rev. Food Sci. Nutr. 2002, 42, 91–121. [Google Scholar] [CrossRef]
- Lozo, J.; Mirkovic, N.; O’Connor, P.M.; Malesevic, M.; Miljkovic, M.; Polovic, N.; Jovcic, B.; Cotter, P.D.; Kojic, M. Lactolisterin BU, a novel class II broad-spectrum bacteriocin from Lactococcus lactis subsp. lactis bv. diacetylactis BGBU1–4. Appl. Environ. Microbiol. 2017, 83. [Google Scholar] [CrossRef]
- Golić, N.; Čadež, N.; Terzić-Vidojević, A.; Šuranská, H.; Beganović, J.; Lozo, J.; Kos, B.; Šušković, J.; Raspor, P.; Topisirović, L. Evaluation of lactic acid bacteria and yeast diversity in traditional white pickled and fresh soft cheeses from the mountain regions of Serbia and lowland regions of Croatia. Int. J. Food Microbiol. 2013, 166, 294–300. [Google Scholar] [CrossRef] [PubMed]
- Mirkovic, N.; Kulas, J.; Miloradovic, Z.; Miljkovic, M.; Tucovic, D.; Miocionovic, J.; Jovcic, B.; Mirkov, I.; Kojic, M. Lactolisterin BU-producer Lactococcus lactis subsp. lactis BGBU1–4: Bio-control of Listeria monocytogenes and Staphylocococcus aureus in fresh soft cheese and effect on immunological response of rats. Food Control 2020, 111, 107076. [Google Scholar] [CrossRef]
- Kojic, M.; Svircevic, J.; Banina, A.; Topisirovic, L. Bacteriocin-Producing Strain of Lactococcus lactis subsp. diacitilactis S50. Appl. Environ. Microbiol. 1991, 57, 1835–1837. [Google Scholar] [CrossRef]
- Hopwood, D.A.; Bibb, M.J.; Chater, K.F.; Kjeser, T.; Bruton, C.J.; Kieser, H.M.; Lydiate, D.J.; Smith, C.P.; Ward, J.M. Genetic Manipulation of Streptomyces: A Laboratory Manual; The John Innes Foundation: Norwich, UK, 1985. [Google Scholar]
- Jovčić, B.; Begović, J.; Lozo, J.; Topisirović, L.; Kojić, M. Dynamics of sodium dodecyl sulfate utilization and antibiotic susceptibility of strain Pseudomonas sp. ATCC19151. Arch. Biol. Sci. 2009, 61, 159–164. [Google Scholar] [CrossRef]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed]
- Tomás, M.S.J.; Claudia Otero, M.; Ocaña, V.; Elena Nader-Macías, M. Production of antimicrobial substances by lactic acid bacteria I: Determination of hydrogen peroxide. Methods Mol. Biol. 2004, 268, 337–346. [Google Scholar] [CrossRef]
- Richelieu, M.; Houlberg, U.; Nielsen, J.C. Determination of α-Acetolactic Acid and Volatile Compounds by Headspace Gas Chromatography. J. Dairy Sci. 1997, 80, 1918–1925. [Google Scholar] [CrossRef]
- Clinical & Laboratory Standards Institute. CLSI Guidelines. Available online: https://clsi.org/ (accessed on 24 March 2021).
- International Dairy Federation. IDF guideline-determination of acidifying activity of dairy cultures. Bull. Int. Dairy Fed. 1995, 306, 34–36. [Google Scholar]
- Mazinani, S.; Fadaei, V.; Khosravi-Darani, K. Impact of Spirulina platensis on Physicochemical Properties and Viability of Lactobacillus acidophilus of Probiotic UF Feta Cheese. J. Food Process. Preserv. 2016, 40, 1318–1324. [Google Scholar] [CrossRef]
- Frece, J.; Kos, B.; Beganović, J.; Vuković, S.; Šušković, J. In vivo testing of functional properties of three selected probiotic strains. World J. Microbiol. Biotechnol. 2005, 21, 1401–1408. [Google Scholar] [CrossRef]
- Cintas, L.M.; Casaus, P.; Håvarstein, S.; Hernandez, P.E.; Hernandez, H.; Nes, I.F. Biochemical and genetic characterization of enterocin P, a novel sec-dependent bacteriocin from enterococcus faecium P13 with a broad antimicrobial spectrum. Appl. Environ. Microbio. 1997, 63, 4321–4330. [Google Scholar] [CrossRef] [PubMed]
- Campagnollo, F.B.; Margalho, L.P.; Kamimura, B.A.; Feliciano, M.D.; Freire, L.; Lopes, L.S.; Alvarenga, V.O.; Cadavez, V.A.P.; Gonzales-Barron, U.; Schaffner, D.W.; et al. Selection of indigenous lactic acid bacteria presenting anti-listerial activity, and their role in reducing the maturation period and assuring the safety of traditional Brazilian cheeses. Food Microbiol. 2018, 73, 288–297. [Google Scholar] [CrossRef] [PubMed]
- Ahmed Sheikh, H.M. Antimicrobial activity of certain bacteria and fungi isolated from soil mixed with human saliva against pathogenic microbes causing dermatological diseases. Saudi J. Biol. Sci. 2010, 17, 331–339. [Google Scholar] [CrossRef]
- Foulquié Moreno, M.R.; Callewaert, R.; Devreese, B.; Van Beeumen, J.; De Vuyst, L. Isolation and biochemical characterisation of enterocins produced by enterococci from different sources. J. Appl. Microbiol. 2003, 94, 214–229. [Google Scholar] [CrossRef]
- Zheng, W.; Zhang, Y.; Lu, H.M.; Li, D.T.; Zhang, Z.L.; Tang, Z.X.; Shi, L.E. Antimicrobial activity and safety evaluation of Enterococcus faecium KQ 2.6 isolated from peacock feces. BMC Biotechnol. 2015, 15, 30. [Google Scholar] [CrossRef] [PubMed]
- Daeschel, M.A. Antimicrobial substances from lactic acid bacteria for use as food preservatives. Food Technol. 1989, 49, 164–1677. [Google Scholar]
- Ghrairi, T.; Manai, M.; Berjeaud, J.M.; Frère, J. Antilisterial activity of lactic acid bacteria isolated from rigouta, a traditional Tunisian cheese. J. Appl. Microbiol. 2004, 97, 621–628. [Google Scholar] [CrossRef]
- Ribeiro, S.C.; Coelho, M.C.; Todorov, S.D.; Franco, B.D.G.M.; Dapkevicius, M.L.E.; Silva, C.C.G. Technological properties of bacteriocin-producing lactic acid bacteria isolated from Pico cheese an artisanal cow’s milk cheese. J. Appl. Microbiol. 2014, 116, 573–585. [Google Scholar] [CrossRef]
- Rivas, F.P.; Castro, M.P.; Vallejo, M.; Marguet, E.; Campos, C.A. Antibacterial potential of Enterococcus faecium strains isolated from ewes’ milk and cheese. LWT-Food Sci. Technol. 2012, 46, 428–436. [Google Scholar] [CrossRef]
- Ness, I.F.; Diep, D.B.; Ike, Y. Enterococcal Bacteriocins and Antimicrobial Proteins that Contribute to Niche Control; Massachusetts Eye and Ear Infirmary: Boston, MA, USA, 2014. [Google Scholar]
- Moy, T.I.; Mylonakis, E.; Calderwood, S.B.; Ausubel, F.M. Cytotoxicity of hydrogen peroxide produced by Enterococcus faecium. Infect. Immun. 2004, 72, 4512–4520. [Google Scholar] [CrossRef]
- Ito, A.; Sato, Y.; Kudo, S.; Sato, S.; Nakajima, H.; Toba, T. The screening of hydrogen peroxide-producing lactic acid bacteria and their application to inactivating psychrotrophic food-borne pathogens. Curr. Microbiol. 2003, 47, 231–236. [Google Scholar] [CrossRef] [PubMed]
- St. Amant, D.C.; Valentin-Bon, I.E.; Jerse, A.E. Inhibition of Neisseria gonorrhoeae by Lactobacillus species that are commonly isolated from the female genital tract. Infect. Immun. 2002, 70, 7169–7171. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ocaña, V.S.; Pesce de Ruiz Holgado, A.A.; Nader-Macà as, M.E. Growth inhibition of Staphylococcus aureus by H 2 O 2-producing Lactobacillus paracasei subsp. paracasei isolated from the human vagina. FEMS Immunol. Med. Microbiol. 1999, 23, 87–92. [Google Scholar] [CrossRef] [PubMed]
- Hawes, S.E.; Hillier, S.L.; Benedetti, J.; Stevens, C.E.; Koutsky, L.A.; Wølner-Hanssen, P.; Holmes, K.K. Hydrogen peroxide-producing lactobacilli and acquisition of vaginal infections. J. Infect. Dis. 1996, 174, 1058–1063. [Google Scholar] [CrossRef] [PubMed]
- Lee, W. Production of Diacetyl (2,3, butanediol) by Continuous Fermentation with Simultaneous Product Separation; Purdue University: West Lafayette, IN, USA, 1991. [Google Scholar]
- Lanciotti, R.; Patrignani, F.; Bagnolini, F.; Guerzoni, M.E.; Gardini, F. Evaluation of diacetyl antimicrobial activity against Escherichia coli, Listeria monocytogenes and Staphylococcus aureus. Food Microbiol. 2003, 20, 537–543. [Google Scholar] [CrossRef]
- Williams-Campbell, A.M.; Jay, J.M. Effects of diacetyl and carbon dioxide on spoilage microflora in ground beef. J. Food Prot. 2002, 65, 523–527. [Google Scholar] [CrossRef] [PubMed]
- Amaral, D.M.F.; Silva, L.F.; Casarotti, S.N.; Nascimento, L.C.S.; Penna, A.L.B. Enterococcus faecium and Enterococcus durans isolated from cheese: Survival in the presence of medications under simulated gastrointestinal conditions and adhesion properties. J. Dairy Sci. 2017, 100, 933–949. [Google Scholar] [CrossRef]
- Werner, G.; Coque, T.M.; Franz, C.M.A.P.; Grohmann, E.; Hegstad, K.; Jensen, L.; van Schaik, W.; Weaver, K. Antibiotic resistant enterococci-Tales of a drug resistance gene trafficker. Int. J. Med. Microbiol. 2013, 303, 360–379. [Google Scholar] [CrossRef]
- Morandi, S.; Brasca, M.; Andrighetto, C.; Lombardi, A.; Lodi, R. Technological and molecular characterisation of enterococci isolated from north-west Italian dairy products. Int. Dairy J. 2006, 16, 867–875. [Google Scholar] [CrossRef]
- Miller, W.R.; Munita, J.M.; Arias, C.A. Mechanisms of antibiotic resistance in enterococci. Expert Rev. Anti. Infect. Ther. 2014, 12, 1221–1236. [Google Scholar] [CrossRef] [PubMed]
- Herreros, M.A.; Fresno, J.M.; González Prieto, M.J.; Tornadijo, M.E. Technological characterization of lactic acid bacteria isolated from Armada cheese (a Spanish goats’ milk cheese). Int. Dairy J. 2003, 13, 469–479. [Google Scholar] [CrossRef]
- Jamaly, N.; Benjouad, A.; Comunian, R.; Daga, E.; Bouksaim, M. Characterization of Enterococci isolated from Moroccan dairy products. Afr. J. Microbiol. Res. 2010, 4, 1768–1774. [Google Scholar]
- Cogan, T.M.; Barbosa, M.; Beuvier, E.; Bianchi-Salvadori, B.; Cocconcelli, P.S.; Fernandes, I.; Gomez, J.; Gomez, R.; Kalantzopoulos, G.; Ledda, A.; et al. Characterization of the lactic acid bacteria in artisanal dairy products. J. Dairy Res. 1997, 64, 409–421. [Google Scholar] [CrossRef]
- Iskandar, C.F.; Cailliez-Grimal, C.; Borges, F.; Revol-Junelles, A.M. Review of lactose and galactose metabolism in Lactic Acid Bacteria dedicated to expert genomic annotation. Trends Food Sci. Technol. 2019, 88, 121–132. [Google Scholar] [CrossRef]
- Radulović, Z.; Miočinović, J.; Petrović, T.; Dimitrijević-Branković, S.; Nedović, V. Traditional and emerging technologies for autochthonous lactic acid bacteria application. In Emerging and Traditional Technologies for Safe, Healthy and Quality Food; Springer: Cham, Switzerland, 2016; pp. 237–256. [Google Scholar]
- Beresford, T.P.; Fitzsimons, N.A.; Brennan, N.L.; Cogan, T.M. Recent advances in cheese microbiology. Int. Dairy J. 2001, 11, 259–274. [Google Scholar] [CrossRef]
- Manolopoulou, E.; Sarantinopoulos, P.; Zoidou, E.; Aktypis, A.; Moschopoulou, E.; Kandarakis, I.G.; Anifantakis, E.M. Evolution of microbial populations during traditional Feta cheese manufacture and ripening. Int. J. Food Microbiol. 2003, 82, 153–161. [Google Scholar] [CrossRef]
- Papageorgiou, D.K.; Marth, E.H. Fate of Listeria monocytogenes during the manufacture, ripening and storage of feta cheese. J. Food Prot. 1989, 52, 82–87. [Google Scholar] [CrossRef] [PubMed]
- Yang, E.; Fan, L.; Yan, J.; Jiang, Y.; Doucette, C.; Fillmore, S.; Walker, B. Influence of culture media, pH and temperature on growth and bacteriocin production of bacteriocinogenic lactic acid bacteria. AMB Express 2018, 8, 10. [Google Scholar] [CrossRef] [PubMed]
- Maisnier-Patin, S.; Deschamps, N.; Tatini, S.R.; Richard, J. Inhibition of Listeria monocytogenes in Camembert cheese made with a nisin-producing starter. Lait 1992, 72, 249–263. [Google Scholar] [CrossRef]
- Rai, M.; Pandit, R.; Gaikwad, S.; Kövics, G. Antimicrobial peptides as natural bio-preservative to enhance the shelf-life of food. J. Food Sci. Technol. 2016, 53, 3381–3394. [Google Scholar] [CrossRef]
- Izquierdo, E.; Marchioni, E.; Aoude-Werner, D.; Hasselmann, C.; Ennahar, S. Smearing of soft cheese with Enterococcus faecium WHE 81, a multi-bacteriocin producer, against Listeria monocytogenes. Food Microbiol. 2009, 26, 16–20. [Google Scholar] [CrossRef] [PubMed]
| Cheese Designation | Bacterial Cultures and Level of Contamination |
|---|---|
| C | CHN 11 |
| CB | CHN11, BGBU1–4 |
| CBL3 | CHN11, BGBU1–4, L.monocyt. 3 log cfu g−1 |
| CBL4 | CHN11, BGBU1–4, L.monocyt. 4 log cfu g−1 |
| CBL5 | CHN11, BGBU1–4, L.monocyt. 5 log cfu g−1 |
| C565 | CHN11, isolate-PFMIX* |
| C565L3 | CHN11, isolate-PFMIX*, L.monocyt. 3log cfu g−1 |
| C565L4 | CHN11, isolate-PFMIX*, L.monocyt. 4 log cfu g−1 |
| C565L5 | CHN11, isolate-PFMIX*, L.monocyt. 5 log cfu g−1 |
| B565C | CHN11, BGBU1–4, isolate-PFMIX* |
| B565CL3 | CHN11, BGBU1–4, isolate-PFMIX*, L.monocyt. 3 log cfu g−1 |
| B565CL4 | CHN11, BGBU1–4, isolate-PFMIX*, L.monocyt. 4 log cfu g−1 |
| B565CL5 | CHN11, BGBU1–4, isolate-PFMIX*, L.monocyt. 5 log cfu g−1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ivanovic, M.; Mirkovic, N.; Mirkovic, M.; Miocinovic, J.; Radulovic, A.; Solevic Knudsen, T.; Radulovic, Z. Autochthonous Enterococcus durans PFMI565 and Lactococcus lactis subsp. lactis BGBU1–4 in Bio-Control of Listeria monocytogenes in Ultrafiltered Cheese. Foods 2021, 10, 1448. https://doi.org/10.3390/foods10071448
Ivanovic M, Mirkovic N, Mirkovic M, Miocinovic J, Radulovic A, Solevic Knudsen T, Radulovic Z. Autochthonous Enterococcus durans PFMI565 and Lactococcus lactis subsp. lactis BGBU1–4 in Bio-Control of Listeria monocytogenes in Ultrafiltered Cheese. Foods. 2021; 10(7):1448. https://doi.org/10.3390/foods10071448
Chicago/Turabian StyleIvanovic, Marina, Nemanja Mirkovic, Milica Mirkovic, Jelena Miocinovic, Ana Radulovic, Tatjana Solevic Knudsen, and Zorica Radulovic. 2021. "Autochthonous Enterococcus durans PFMI565 and Lactococcus lactis subsp. lactis BGBU1–4 in Bio-Control of Listeria monocytogenes in Ultrafiltered Cheese" Foods 10, no. 7: 1448. https://doi.org/10.3390/foods10071448
APA StyleIvanovic, M., Mirkovic, N., Mirkovic, M., Miocinovic, J., Radulovic, A., Solevic Knudsen, T., & Radulovic, Z. (2021). Autochthonous Enterococcus durans PFMI565 and Lactococcus lactis subsp. lactis BGBU1–4 in Bio-Control of Listeria monocytogenes in Ultrafiltered Cheese. Foods, 10(7), 1448. https://doi.org/10.3390/foods10071448







