High-Pressure-Induced Sublethal Injuries of Food Pathogens—Microscopic Assessment
Abstract
:1. Introduction
2. Materials and Methods
2.1. Preparation of Bacterial Suspensions and Culture Conditions
2.2. HHP Device and Parameters
2.3. Scanning Electron Microscope Protocol (SEM)
2.4. Transmission Electron Microscopy Protocol (TEM)
2.5. Epifluorescent Microscopy (EFM)
2.6. Statistical Analysis
3. Results and Discussion
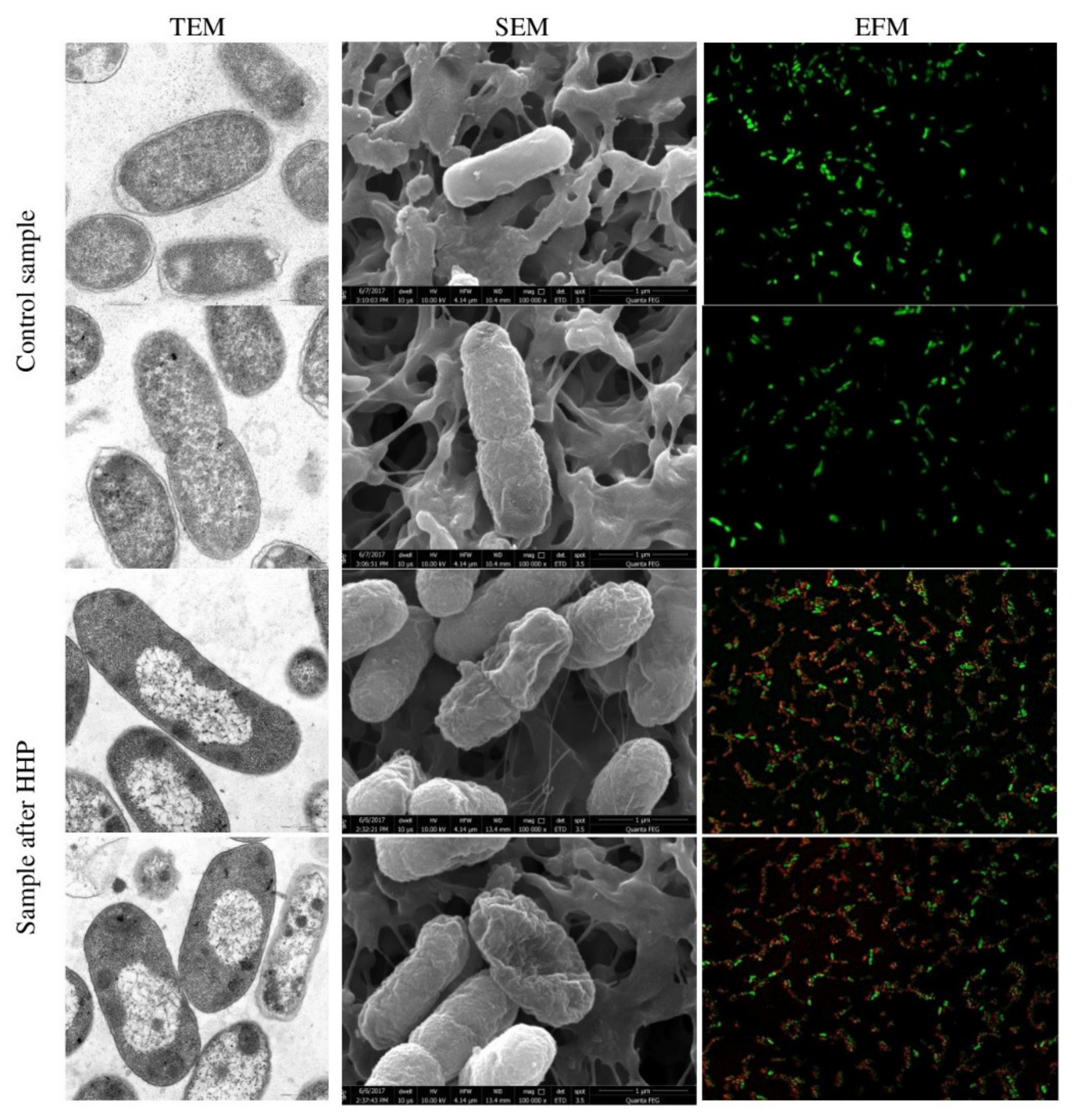
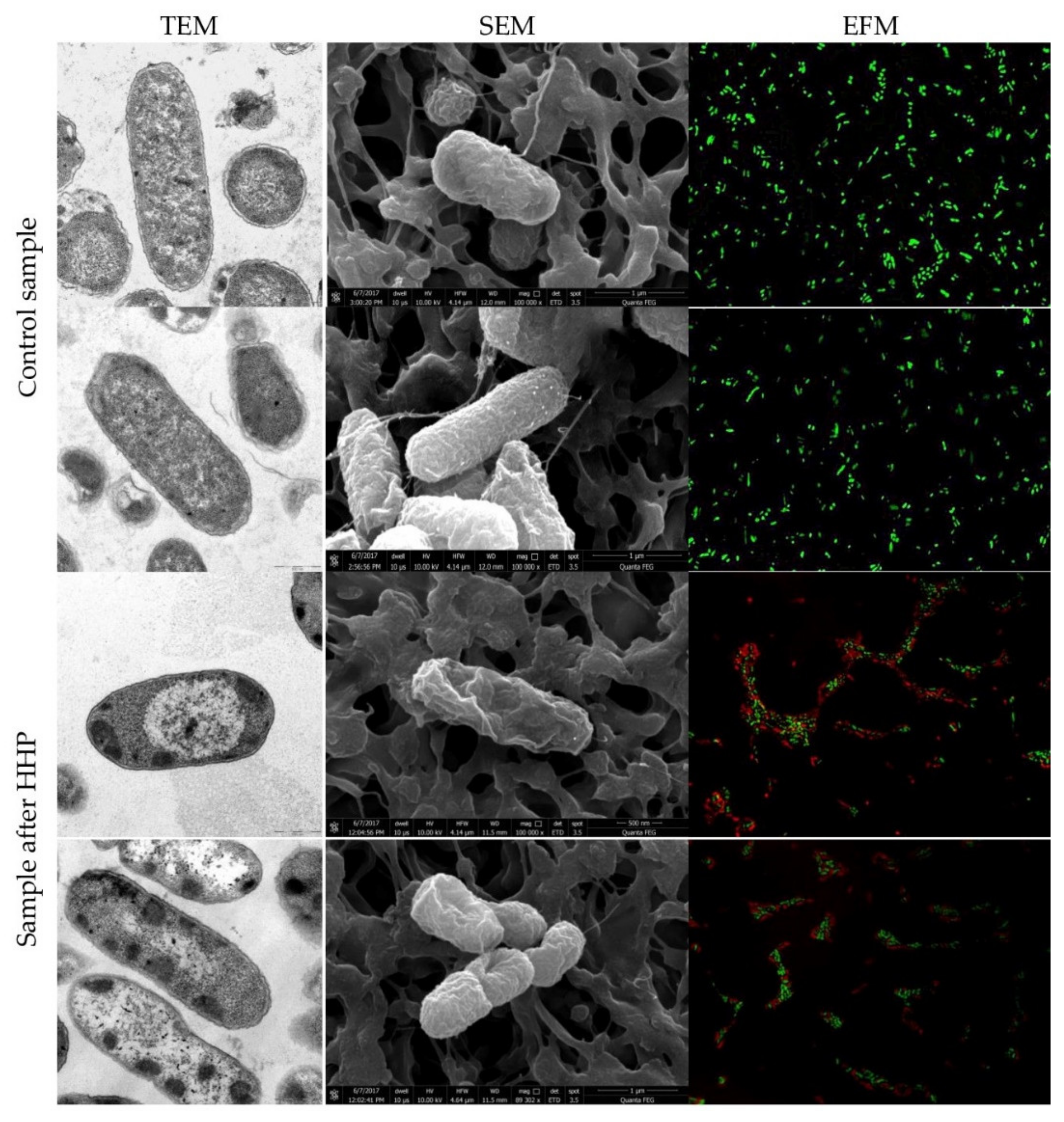
3.1. Escherichia coli EMs Observations
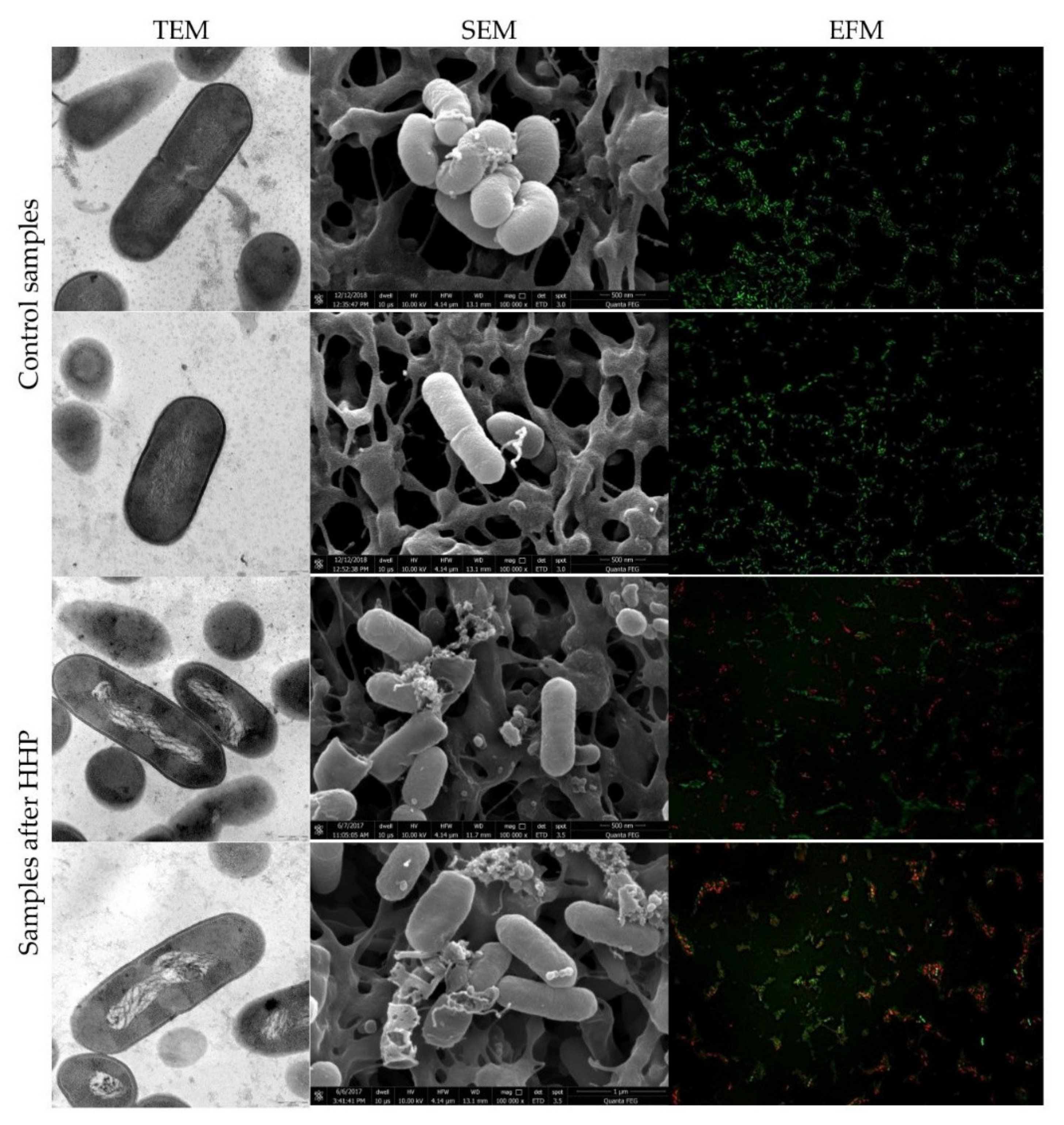
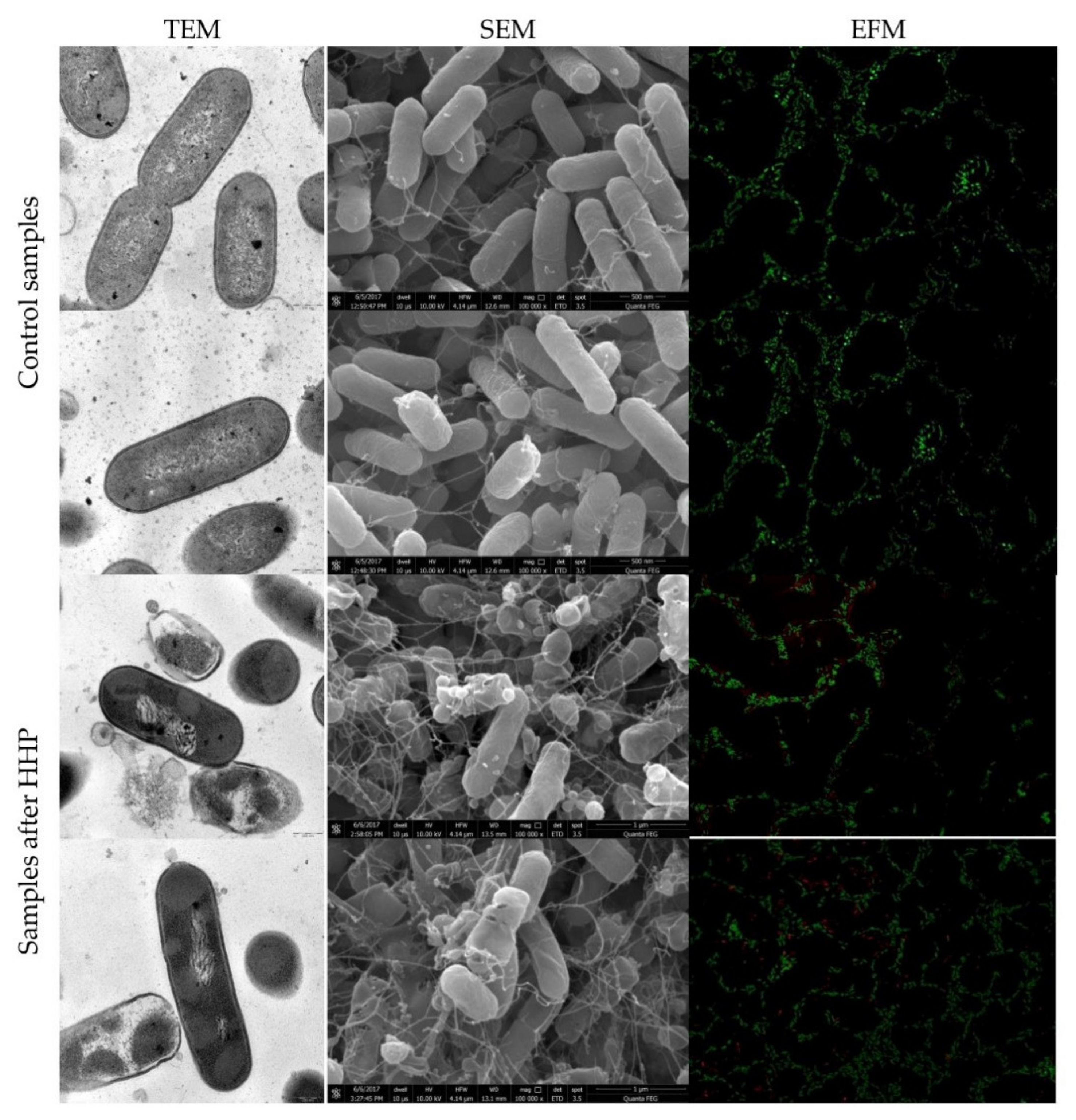
3.2. Listeria innocua EMs Observations
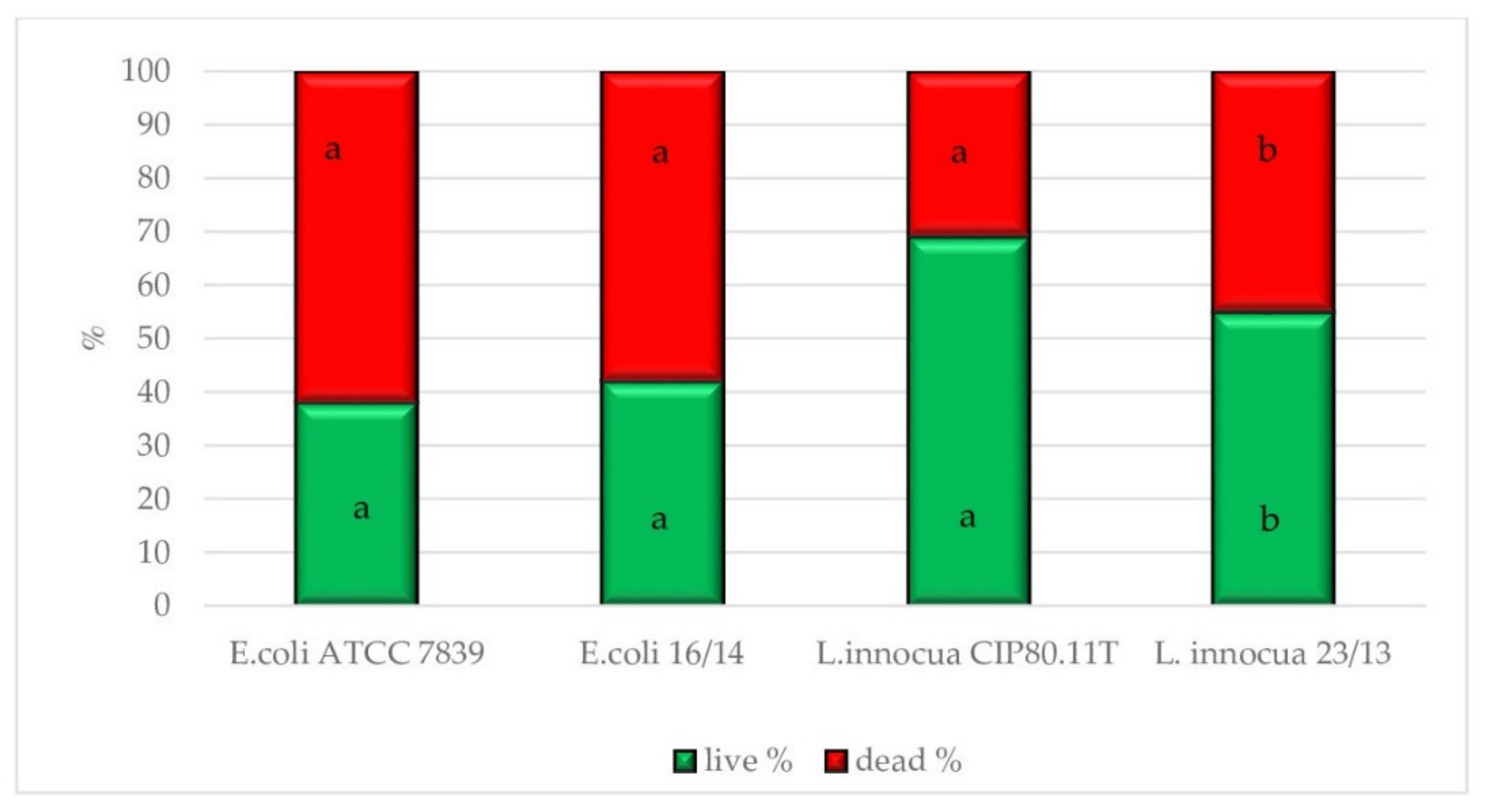
3.3. Physiological State of Cells Assessed by EFM
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Gelderblom, H.R.; Hazelton, P.R. Specimen collection for electron microscopy. Emerg. Infect. Dis. 2000, 6, 433–434. [Google Scholar] [CrossRef] [PubMed]
- Golding, C.G.; Lamboo, L.L.; Beniac, D.R.; Booth, T.F. The scanning electron microscope in microbiology and diagnosis of infectious disease. Sci. Rep. 2016, 6, 26516. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hameed, S.; Xie, L.; Ying, Y. Conventional and emerging detection techniques for pathogenic bacteria in food science: A review. Trends Food Sci. Technol. 2018, 81, 61–73. [Google Scholar] [CrossRef]
- Perfumo, A.; Elsaesser, A.; Littmann, S.; Foste, R.A.; Kuypers, M.M.M.; Cockell, C.S.; Kminek, G. Epifluorescence, SEM, TEM and nanoSIMS image analysis of the cold phenotype of Clostridium psychrophilum at subzero temperatures. FEMS Microbiol. Ecol. 2014, 90, 869–882. [Google Scholar] [CrossRef] [Green Version]
- Ruiz, L.; Aertsen, A.; Nguyen-The, C.; Gänzle, M.G.; Alvarez-Ordóñez, A. Editorial: Industrial and Host Associated Stress Responses in Food Microbes. Implications for Food Technology and Food Safety. Front. Microbiol. 2017, 8, 1522. [Google Scholar] [CrossRef]
- Donczew, M.; Ginda, K.; Zakrzewska-Czerwińska, J.; Jakimowicz, D. Solving the mysteries of the bacterial cell—Application of novel techniques in fluorescence microscopy. Postepy Hig. Med. Dosw. 2011, 65, 114–123. [Google Scholar] [CrossRef]
- Bergmans, L.; Moisiadis, P.; Van Meerbeek, B.; Quirynen, M.; Lambrechts, P. Microscopic observation of bacteria: Review highlighting the use of environmental SEM. Int. Endod. J. 2005, 38, 775–788. [Google Scholar] [CrossRef] [Green Version]
- Grin, I.; Schwarz, H.; Linkeet, D. Electron microscopy techniques to study bacterial adhesion. Adv. Exp. Med. Biol. 2011, 715, 257–269. [Google Scholar] [CrossRef] [PubMed]
- Gonzâlez-Machado, C.; Capita, R.; Riesco-Pelâez, F.; Alonso-Calleja, C. Visualization and quantification of the cellular and extracellular components of Salmonella Agona biofilms at different stages of development. PLoS ONE 2018, 3, e0200011. [Google Scholar] [CrossRef]
- Mackey, B.M.; Forestière, K.; Issaacs, N.S.; Stenning, R.; Brooker, B. The effect of high hydrostatic pressure on Salmonella thompson and Listeria monocytogenes examined by electron microscopy. Lett. Appl. Microbiol. 1994, 19, 429–432. [Google Scholar] [CrossRef]
- Prieto-Calvo, M.; Prieto, M.; Lopez, M.; Alvarez-Ordóñez, A. Effects of high hydrostatic pressure on Escherichia coli ultrastructure, membrane integrity and molecular composition as assessed by FTIR spectroscopy and microscopic imaging techniques. Molecules 2014, 19, 21310–21323. [Google Scholar] [CrossRef] [PubMed]
- Pillet, F.; Formosa-Dague, C.; Baaziz, H.; Dague, E.; Rols, M.P. Cell wall as a target for bacteria inactivation by pulsed electric fields. Sci. Rep. 2016, 6, 19778. [Google Scholar] [CrossRef]
- Khadi, N.; Raj Upreti, A.; Li, Y. Simultaneous bacterial inactivation and degradation of an emerging pollutant under visible light by ZnFe2O4 co-modified with Ag and rGO. RSC Adv. 2017, 7, 27007–27016. [Google Scholar] [CrossRef] [Green Version]
- Berney, M.; Hammes, F.; Bosshard, F.; Weilenmann, H.U.; Egli, T. Assessment and interpretation of bacterial viability by using the LIVE/DEAD BacLight Kit in combination with flow cytometry. Appl. Environ. Microbiol. 2007, 73, 3283–3290. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stewart, E.J. Growing unculturable bacteria. J. Bacteriol. 2012, 194, 4151–4160. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dos Santos Leandro, E.; Kunrath Lima, G.; Fernandes de Carvalho, A.; Gomes Pereira, O.; de Moraes, C.A. Flow cytometric assessment of Lactococcus lactis isolates viability after lyophilization. Int. J. Nutr. Food Sci. 2014, 5, 391–396. [Google Scholar] [CrossRef] [Green Version]
- Nishi, K.; Isobe, S.; Zhu, Y.; Kiyama, R. Fluorescence-based bioassays for the detection and evaluation of food materials. Sensors 2015, 15, 25831–25867. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Léonard, L.; Bouarab Chibane, L.; Ouled Bouhedda, B.; Degraeve, P.; Oulahal, N. Recent Advances on Multi-Parameter Flow Cytometry to Characterize Antimicrobial Treatments. Front. Microbiol. 2016, 7, 1225. [Google Scholar] [CrossRef] [Green Version]
- Moreno, Y.; Collado, M.C.; Ferrus, M.A.; Cobo, J.M.; Hernandez, E.; Hernandez, M. Viability assessment of lactic acid bacteria on commercial dairy products stored at 4 °C using LIVE/DEAD® BacLightTM staining and conventional plate counts. Int. J. Food Sci. Technol. 2006, 41, 275–280. [Google Scholar] [CrossRef]
- Sunny-Roberts, E.O.; Knorr, D. Evaluation of the response of Lactobacillus rhamnosus VTT E-97800 to sucrose-induced osmotic stress. Food Microbiol. 2008, 25, 183–189. [Google Scholar] [CrossRef]
- Zotta, T.; Guidone, A.; Tremonte, P.; Parente, E.; Ricciardi, A. A comparison of fluorescent stains for the assessment of viability and metabolic activity of lactic acid bacteria. World J. Microbiol. Biotechnol. 2012, 28, 919–927. [Google Scholar] [CrossRef] [PubMed]
- Kocot, A.M.; Olszewska, M.A. Interaction of Pseudomonas aeruginosa and Staphylococcus aureus with Listeria innocua in dual species biofilms and inactivation following disinfectant treatments. LWT Food Sci. Technol. 2020, 118, 108736. [Google Scholar] [CrossRef]
- Nikparvar, B.; Subires, A.; Capellas, M.; Hernandez-Herrero, M.; Crauwels, P.; Riedel, C.U.; Bar, N. A diffusion model to quantify membrane repair process in Listeria monocytogenes exposed to high pressure processing based on fluorescence microscopy. Data Front. Microbiol. 2021, 12, 598739. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.-W.; Lung, H.-M.; Yang, B.B.; Wang, C.-Y. Responses of microorganisms to high hydrostatic pressure processing. Food Control 2014, 40, 250–259. [Google Scholar] [CrossRef]
- Balasubramaniam, V.M.; Barbosa-Cánovas, G.V.; Lelieved, H.L.M. High Pressure Processing of Foods. Food Sci. Technol. Int. 2016, 14, 413–418. [Google Scholar] [CrossRef]
- Huang, H.-W.; Wu, S.-J.; Lu, J.-K.; Shyu, Y.-T.; Wang, C.-Y. Current status and future trends of high-pressure processing in food industry. Food Control 2017, 72, 1–8. [Google Scholar] [CrossRef]
- Cacace, F.; Bottani, E.; Rizzi, A.; Vignali, G. Evaluation of the economic and environmental sustainability of high pressure processing of foods. Innovative. Food Sci. Emerg. Technol. 2020, 60, 102281. [Google Scholar] [CrossRef]
- Wu, D.; Forghani, F.; Banan-Mwine Daliri, E.; Li, J.; Liao, X.; Liu, D.; Ye, X.; Chen, S.; Ding, T. Microbial response to some nonthermal physical technologies. Trends Food Sci. Technol. 2020, 95, 107–117. [Google Scholar] [CrossRef]
- Torress Bello, E.F.; González Martínez, G.; Klotz Ceberio, B.F.; Rodrigo, D.; Martínez López, A. High Pressure Treatment in Foods. Foods 2014, 3, 476–490. [Google Scholar] [CrossRef] [Green Version]
- Marszałek, K.; Woźniak, Ł.; Kruszewski, B.; Skąpska, S. The effect of high pressure techniques on the stability of anthocyanins in fruit and vegetables. Int. J. Mol. Sci. 2017, 18, 277. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Szczepańska, J.; Barba, F.J.; Skąpska, S.; Marszałek, K. High pressure processing of carrot juice: Effect of static and multi-pulsed pressure on the polyphenolic profile, oxidoreductases activity and colour. Food Chem. 2020, 307, 125549. [Google Scholar] [CrossRef] [PubMed]
- Balakrishna, A.K.; Wazed, M.A.; Farid, M.A. Review on the Effect of High Pressure Processing (HPP) on Gelatinization and Infusion of Nutrients. Molecules 2020, 25, 2369. [Google Scholar] [CrossRef]
- Mukhopadhyay, S.; Sokorai, K.; Ukuku, D.; Fan, X.; Juneja, V.; Sites, J.; Cassidy, J. Inactivation of Salmonella enterica and Listeria monocytogenes in cantaloupe puree by high hydrostatic pressure with/without added ascorbic acid. Int. J. Food Microbiol. 2016, 235, 77–84. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nasiłowska, J.; Sokołowska, B.; Fonberg-Broczek, M. Behavior of Listeria innocua strains under pressure treatment—Inactivation and sublethal injury. Pol. J. Food Nutr. Sci. 2019, 69, 45–52. [Google Scholar] [CrossRef]
- Sokołowska, B.; Nasiłowska, J.; Rutkowska, M.; Fonberg-Broczek, M.; Rzoska, S.J. The usage of high hydrostatic pressure (HHP) to control food-borne pathogens in hummus. High Press. Res. 2019, 39, 525–532. [Google Scholar] [CrossRef]
- Yamamoto, K.; Zhang, X.; Inaoka, T.; Morimatsu, K.; Kimura, K.; Nakaura, Y. Bacterial Injury Induced by High Hydrostatic Pressure. Food Eng. Rev. 2021, 13, 442–453. [Google Scholar] [CrossRef]
- Wesche, A.M.; Gurtler, J.B.; Marks, B.P.; Ryser, E.T. Stress, sublethal injury, resuscitation and virulence of bacterial foodborn pathogens. J. Food Prot. 2009, 72, 1121–1138. [Google Scholar] [CrossRef]
- Podolak, R.; Whitman, D.; Black, A.G. Factors Affecting Microbial Inactivation during High Pressure Processing in Juices and Beverages: A Review. J. Food Prot. 2020, 83, 1561–1575. [Google Scholar] [CrossRef]
- Hoover, D.G.; Metrick, C.; Papineau, A.M.; Farkas, D.F.; Knorr, D. Biological effects of high hydrostatic pressure on food microorganisms. Food Tech. 1989, 43, 99–107. [Google Scholar]
- Yuste, J.; Capellas, M.; Fung, D.Y.C.; Mor–Mur, M. Inactivation and sublethal injury of foodborne pathogens by high pressure processing: Evaluation with conventional media and thin agar layer method. Food Res. Int. 2004, 37, 861–866. [Google Scholar] [CrossRef]
- Maldonado, J.A.; Schaffner, D.W.; Cuitiño, A.M.; Karwe, M.V. In situ studies of microbial inactivation during high pressure processing. High Pressure Res. 2016, 36, 79–89. [Google Scholar] [CrossRef]
- Pavlov, M.Y.; Ehrenberg, M. Optimal control of gene expression for fast proteome adaptation to environmental change. Proc. Natl. Acad. Sci. USA 2013, 110, 20527–20532. [Google Scholar] [CrossRef] [Green Version]
- Evrendilek, G.A. Microbial stress response to high pressure processing. In Stress Response of Foodborne Microorganisms; Wong, H.C.H., Ed.; Nova Science Publishers, Inc.: Hauppauge, NY, USA, 2012. [Google Scholar]
- Reynolds, E.S. The use of lead citrate at high pH as electron-opaque stain for electron microscopy. J. Cell Biol. 1963, 17, 208–213. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- LIVE/DEAD® BacLight™ Bacterial Viability Kit Protocol. Available online: https://www.thermofisher.com/pl/en/home/references/protocols/cell-and-tissue-analysis/protocols/live-dead-baclight-bacterial-viability-protocol.html (accessed on 17 October 2021).
- Moussa, M.; Perrier-Cornet, J.-N.; Gervais, P. Damage in Escherichia coli Cells Treated with a Combination of High Hydrostatic Pressure and Subzero Temperature. Appl. Environ. Microbiol. 2007, 73, 6508–6518. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, B.; Shi, Y.; Xia, X.; Xi, M.; Wang, X.; Ji, B.; Meng, J. Inactivation of foodborne pathogens in raw milk using high hydrostatic pressure. Food Control 2012, 28, 273–278. [Google Scholar] [CrossRef]
- Hsu, H.Y.; Sheen, S.; Sites, J.; Huang, L.; Wu, J.S. Effect of high pressure treatment on the survival of shiga toxin producing Escherichia coli in strawberry puree. Food Microbiol. 2014, 40, 25–30. [Google Scholar] [CrossRef]
- Huang, H.W.; Lung, H.M.; Chang, Y.H.; Yang, B.B.; Wang, C.Y. Inactivation of pathogenic Listeria monocytogenes in raw milk by high hydrostatic pressure. Foodborne Pathog. Dis. 2015, 12, 139–144. [Google Scholar] [CrossRef] [PubMed]
- Basaran-Akgul, N.; Rasco, B. High pressure processing inactivation of Listeria innocua in minced trout (Oncorhynchus mykiss). J. Food Process. Preserv. 2010, 34, 191–206. [Google Scholar] [CrossRef]
- Ritz, M.; Tholozan, J.L.; Federighi, M.; Pilet, M.F. Physiological damages of Listeria monocytogenes treated by high hydrostatic pressure. Int. J. Food Microbiol. 2002, 15, 47–53. [Google Scholar] [CrossRef]
- Kimura, K.; Morimatsu, K.; Inaoka, T.; Yamamoto, K. Injury and recovery of Escherichia coli ATCC25922 cells treated by high hydrostatic pressure at 400–600 MPa. J. Biosci. Bioeng. 2017, 123, 698–706. [Google Scholar] [CrossRef]
- Nasiłowska, J.; Sokołowska, B.; Fonberg-Broczek, M. Long-Term Storage of Vegetable Juices Treated by High Hydrostatic Pressure: Assurance of the Microbial Safety. BioMed Res. Int. 2018, 2, 7389381. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nasiłowska, J.; Kocot, A.; Osuchowska, P.N.; Sokołowska, B. High-Pressure-Induced Sublethal Injuries of Food Pathogens—Microscopic Assessment. Foods 2021, 10, 2940. https://doi.org/10.3390/foods10122940
Nasiłowska J, Kocot A, Osuchowska PN, Sokołowska B. High-Pressure-Induced Sublethal Injuries of Food Pathogens—Microscopic Assessment. Foods. 2021; 10(12):2940. https://doi.org/10.3390/foods10122940
Chicago/Turabian StyleNasiłowska, Justyna, Aleksandra Kocot, Paulina Natalia Osuchowska, and Barbara Sokołowska. 2021. "High-Pressure-Induced Sublethal Injuries of Food Pathogens—Microscopic Assessment" Foods 10, no. 12: 2940. https://doi.org/10.3390/foods10122940
APA StyleNasiłowska, J., Kocot, A., Osuchowska, P. N., & Sokołowska, B. (2021). High-Pressure-Induced Sublethal Injuries of Food Pathogens—Microscopic Assessment. Foods, 10(12), 2940. https://doi.org/10.3390/foods10122940







