3.1. Resolution and Printing Quality of LIFT-Printed Arrays
LIFT bioprinting was conducted at 11.3 μJ (RMS stability 0.34%), corresponding to the threshold energy fluence of 400 mJ/cm
2 for 60 μm focal spot, which ensures minimal energy input while still enabling stable jet formation and material transfer as well as immobilization. Laser power was measured with a laser energy detector (QE8SP-B-MT-D0, Gentec-EO) in combination with a laser power monitor (11MAESTRO, Standa). Although droplet deposition was achievable at donor–receiver distances of up to 3 mm, the spatial resolution and integrity of printed patterns were found to be highly dependent on the separation distance. In this study, 4 × 4 spot arrays were printed at distances ranging from 1 mm to 3 mm in order to evaluate the influence of propagation length on printing accuracy (
Figure 3).
To assess the quality and accuracy of the printed arrays, image analysis was performed using Fiji (ImageJ 1.54p). Each droplet was approximated as a perfect circle, and its geometric center was calculated based on pixel coordinates, annotated as a red dot (
Figure 3). The overall centroid of each 4 × 4 array was then determined using the mean x and y positions of the 16 droplets according to the following equation:
Based on this reference centroid, the average center-to-center distances between adjacent droplets in both rows and columns were computed, with the expected nominal spacing being 1 mm. Subsequently a reference grid with 1 mm
2 square cells was con-structed and centered on the array centroid. The center of each unit cell of the grid is demonstrated as a black dot and represents the intended position of each droplet (
Figure 3). The grid was generated for the 1 mm donor–receiver distance and applied to the rest of the arrays. Since the intended spacing was 1000 µm, the relative percentage deviation for each droplet was computed as follows:
where
is the distance between each droplet’s actual position and its intended position in μm, measured in ImageJ.
Image acquisition was carried out using a Basler acA1920-40uc USB 3.0 camera mounted on a conventional upright microscope equipped with a 4× objective lens. Overlapping images were captured and later combined into a single high-resolution composite using the GNU Image Manipulation Program (GIMP).
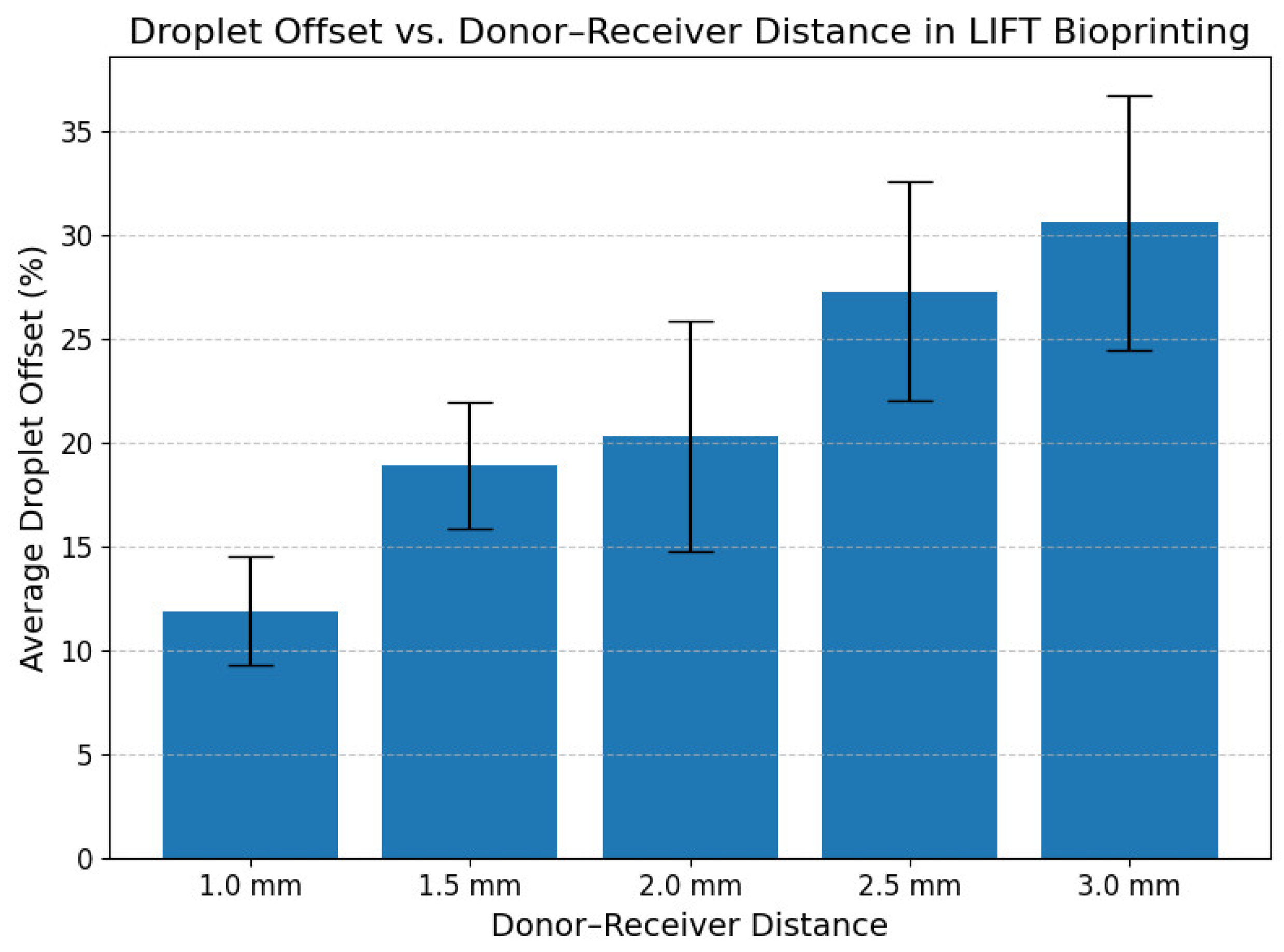
Quantitative analysis demonstrated a progressive loss of resolution as the donor–receiver distance increased. The average deviation of the droplets from the intended array geometry was measured as follows: 11.9 ± 5.2% at 1.0 mm, 18.9 ± 6.1% at 1.5 mm, 20.3 ± 10.9% at 2.0 mm, 27.3 ± 10.6% at 2.5 mm, and 30.6 ± 12.3% at 3.0 mm (
Figure 4). These deviations are primarily attributed to the complex fluid dynamic phenomena during the early phases of jet formation and droplet flight, which become more pronounced at longer travel distances.
In addition to geometrical inaccuracies, droplet–droplet interactions emerged as a secondary challenge. At distances beyond 2.0 mm, droplet trajectories exhibited increasing spatial instability, frequently resulting in droplet-to-droplet collisions during deposition. These interactions severely disrupted the spatial organization of the printed arrays, further degrading the overall print resolution. As such, a critical donor–receiver distance of 2.0 mm was identified as the upper limit for maintaining acceptable pattern fidelity under the applied energy fluence.
3.2. Jet Propagation Dynamics in LIFT Bioprinting
To quantitatively analyze the droplet propagation dynamics captured via high-speed imaging, two custom Python scripts were developed. These scripts enabled frame-by-frame inspection of the LIFT videos and enabled manual annotation of key jet features, such as the position of the primary and secondary jets at each time point. Pixel-to-micrometer calibration was performed using predefined reference scales. Additionally, data extraction includes droplet displacement for each frame, in structured formats (CSV files) for analysis.
Specifically, the first script allowed for the definition of reference distances and start/end points using user-interactive mouse input within the video frame, while automatically converting pixel displacement into micrometer-scale distances based on predefined calibration values. This enabled the extraction of displacement data for each jet with high temporal precision. The second script expanded on this functionality by introducing a user-guided cursor system and segment-specific data export features for finer measurements. In both cases, video frames were indexed based on the effective frame rate (ranging from 102,000 to 170,000 fps depending on the experiment), and time was automatically computed for each data point. Output data were exported directly to structured formats (e.g., clipboard or CSV) for further numerical analysis. This semi-automated approach provided consistent, high-resolution tracking of jet behavior across all donor–receiver distances, ensuring accurate characterization of the dynamics throughout the LIFT process.
Multiple studies have consistently reported the formation of two successive jets during the printing process [
3,
11,
12,
13]. Observations from the high-speed video recordings clearly indicate that the primary jet exhibits a narrow diameter and high velocity, while the secondary jet appears broader and propagates at a reduced speed (
Figure 5).
Initially, the bioink undergoes a rapid acceleration phase, followed by stabilization to a constant propagation velocity. At the applied laser fluence, the secondary jet remained continuous and unfragmented at 1 mm, but beyond this distance, it consistently broke into discrete droplets, the front of which was measured with a diameter of 145 ± 12 μm. Further analysis of the stabilized propagation phase revealed that the primary jet exhibited an average propagation velocity of 9.14 ± 0.44 m/s, while the secondary jet followed at a lower average velocity of 1.76 ± 0.38 m/s (
Figure 6A). All data were derived from four independent replicates in each distance to ensure experimental reproducibility and enhance the reliability of the results. In certain cases, particularly at distances beyond 1.5 mm, the fragmented droplets exhibited visible oscillatory motion. This oscillation induced a minor deceleration; however, this value was deemed negligible with respect to the overall droplet dynamics and was not found to influence the stability or trajectory of the jet under the tested conditions.
High-speed imaging of jet propagation dynamics confirmed that donor–receiver distances exceeding 2.0 mm introduced significant spatial instability in droplet trajectories. Angular deviations of fragmented droplets from the initial jet trajectory path were found to amplify spatial deviations in the printed pattern, with increased propagation distance. Specifically, an angular deviation greater than 7° along a single axis (
Figure 6B) can produce a spatial offset of less than 150 μm at a 1.0 mm distance, while this offset increased to approximately 250 μm at 2.0 mm and reached nearly 400 μm at 3.0 mm. These findings highlight the critical impact of jet directionality on pattern fidelity and support the conclusion that distances above 2.0 mm are unsuitable for high-precision patterning using LIFT under the tested conditions.
3.4. CFD Simulation in Designed Microfluidic Chip
Computational fluid dynamics (CFD) simulations are powerful tools widely employed to optimize microfluidic designs by accurately modeling fluid behavior within microchannels [
14,
15,
16]. These simulations enable researchers to predict and refine flow dynamics and other critical parameters. This approach supports the development of improved OoC devices with well-characterized and reproducible performance.
To evaluate the fluid dynamics within microfluidic chip designed in this study, a CFD simulation was performed using physiologically relevant flow parameters. Capillary blood flow in human tissues typically ranges between 10
−3 and 10
−4 m/s [
17]. Based on this, an inlet velocity of 2 × 10
−4 m/s was selected, corresponding to a flow rate of approximately 2.4 μL/min within our system. This flow rate aligns well with estimated tissue blood flow (TBF) values for volumes comparable to our platform’s configuration—a monolayer of cells with a 6 mm radius and 20 μm height—where TBF is reported to be between 1.5 and 2.6 μL/min [
18,
19].
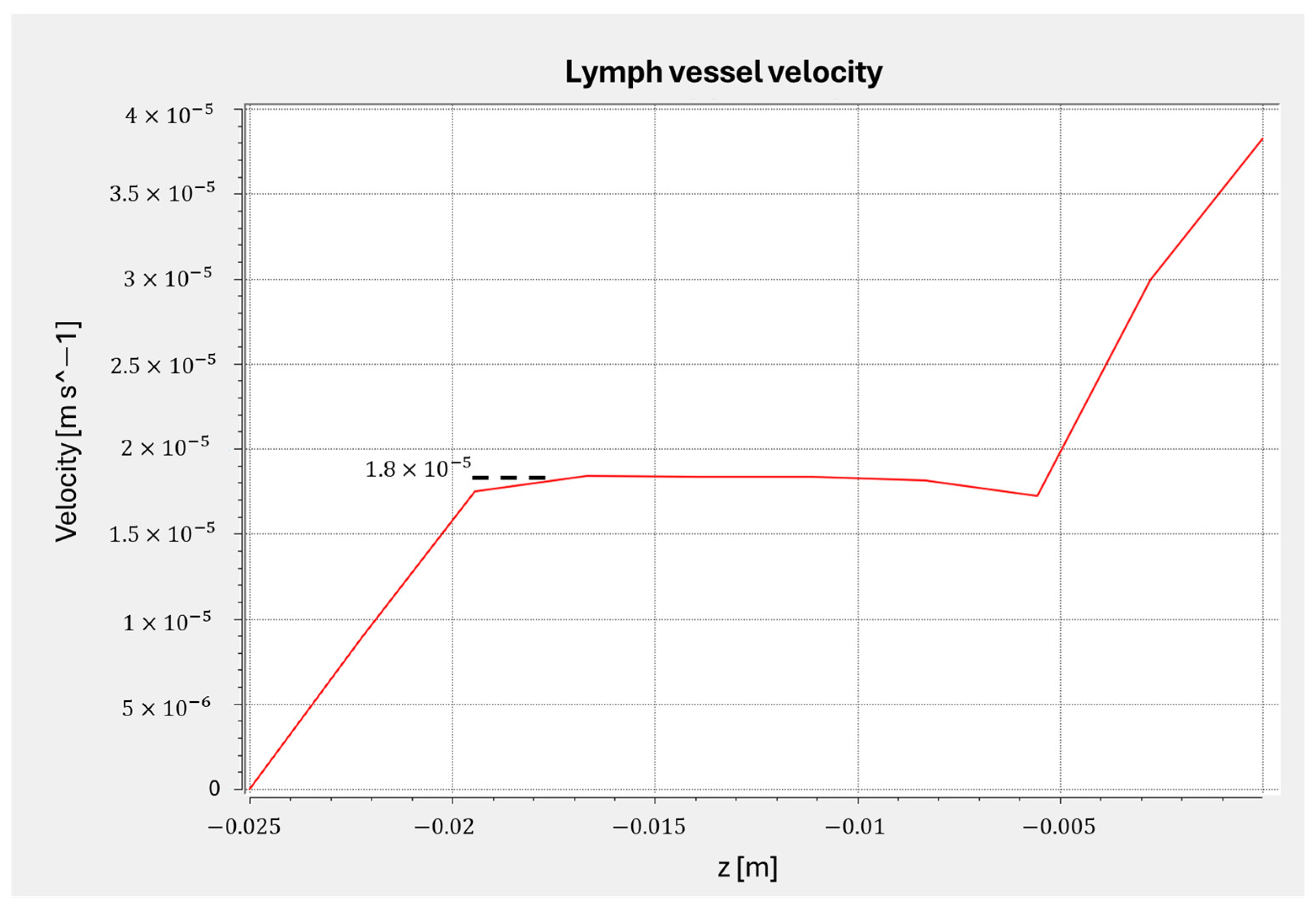
Simulation results confirmed uniform flow distribution within the tumor and lymph compartments (
Figure 8) and revealed that the flow velocity in the interconnecting microchannel designed to mimic a lymphatic vessel was approximately 1.8 × 10
−5 m/s (
Figure A2). This value is consistent with physiological lymph flow velocities reported in the literature [
20]. To assess molecular transport within this region, the Péclet number (Pe = VL/D) was calculated for typical cytokines (diffusion coefficient D ≈ 10
−6 m
2/s [
21]), yielding Pe ≪ 1. This indicates that molecular transport in the channel is diffusion-dominated, supporting effective biochemical communication between the two compartments through passive protein exchange.
3.6. Flow Visualization—Particle Shadow Velocimetry
To experimentally validate the theoretical flow velocity values obtained from CFD simulations, flow visualization was performed using particle shadow velocimetry. Polystyrene microparticles are well-suited for water flow tracking in small channels [
22], with a density appropriate for suspension in aqueous media such as water and cell culture media. The particles were employed at a low concentration of 10
2 particles/mL to avoid aggregation and enable individual particle tracking. The experiment was conducted using the fluidic setup described in the Materials and Methods section. The microfluidic chip was mounted onto an inverted microscope, with focus placed on the interconnecting channel designed to mimic a lymphatic vessel.
Time-lapse imaging captured the motion of particles within the central region of the channel. Post-processing and tracking were conducted using Fiji (ImageJ) (
Figure A3). By tracking five representative particles, the experimentally measured average flow velocity was determined to be V
exp = 1.36 ± 0.02 × 10
−5 m/s. This value was compared to the theoretical velocity predicted by the CFD simulation (V
theo = 1.8 × 10
−5 m/s), revealing a 24% deviation. This deviation is attributable to the 3D printing process, which, as a layer-by-layer additive manufacturing technique, inherently introduces dimensional imperfections in cross sections of the microchannels, as well as geometric irregularities at channel junctions and corners. Such geometrical deviations, from the initial microfluidic design imported in the CFD calculations, have a notable impact on the hydraulic resistances of microchannels, which, in turn, affect the pressure and flow rate distributions within the multichannel microfluidic chip. Despite this, the experimentally observed flow remained within a physiologically relevant range, supporting the chip’s functional validity.
3.8. Cell Culture Under Flow Inside the Microfluidic Chip Chamber
In order to evaluate the ability of the microfluidic platform to support viable cell culture under continuous flow, two separate experiments were conducted using LLC cells by utilizing the fluidic setup described above (
Figure 2). In the first experiment, cells were seeded into the chamber of the microfluidic chip at a concentration of 3 × 10
4 cells/mL and subjected to continuous perfusion after a 4 h static incubation time. After 24 h under flow, cells were observed to have successfully adhered to the glass-bottom substrate and exhibited normal phenotypic characteristics, including elongated morphology typical of LLC cells under adherent conditions (
Figure A4). Cells were subsequently harvested by PBS washing and trypsinization and assessed for viability using Trypan blue exclusion. The viability exceeded 95%, indicating that the culture conditions within the device were non-toxic and supportive of sustained cancer cell growth.
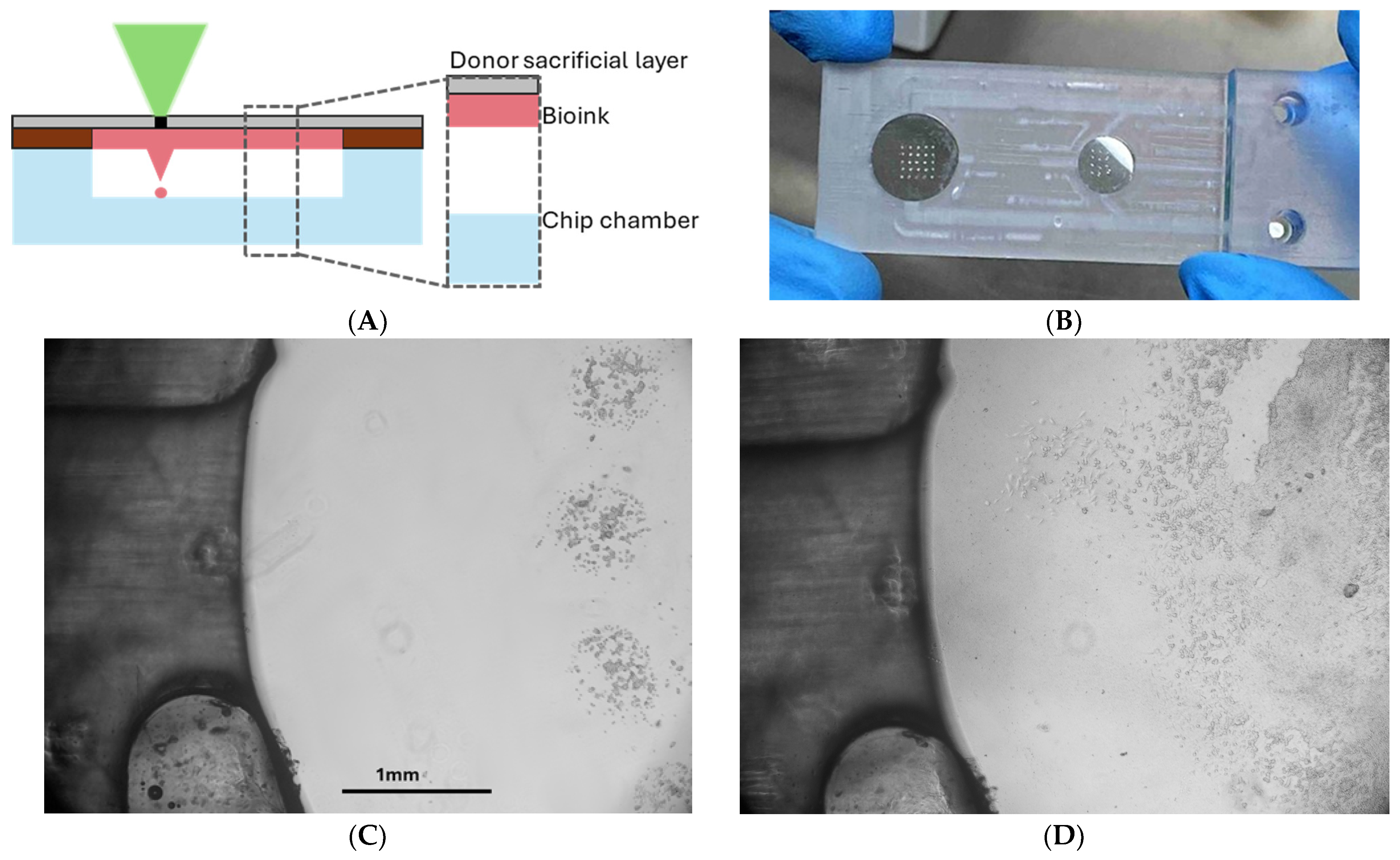
Building upon these results, a second experiment was performed using a microfluidic chip containing a bioprinted array of LLC cells. To accomplish this, the microfluidic chip chamber was brought into direct contact with the donor substrate containing the bioink, in order to ensure the 2 mm gap from the substrate (chamber bottom) (
Figure 9A). In this approach, perfusion was initiated immediately after cell introduction (
Figure 9C), as the LIFT process enables precise immobilization of cells at the printed positions. After 48 h of perfusion, a confluent monolayer formed around the printed spots, and signs of cell detachment were observed due to over-confluency (
Figure 9D). High-resolution images captured using a 10× objective lens further revealed cell migration toward the central microchannel, where fresh medium entered the chamber. These findings confirm that the platform can support viable, dynamic cultures under physiological flow conditions and suggest its potential for studying cancer cell behavior, migration, and cell–cell signaling in microengineered environments.
To summarize, this study demonstrates the potential of LIFT bioprinting as a high-precision, non-contact technique for cell patterning within microfluidic devices, offering significant advantages for OoC research. Through a detailed parametric analysis, we established that a donor–receiver distance of up to 2.0 mm allows consistent jet propagation and acceptable print resolution. At distances beyond this threshold, spatial instability increases, leading to compromised printed arrays.
Previous studies have explored the basic principles of LIFT printing for cell-laden materials (Guillotin et al., Catros et al.), yet most focused on either open substrate systems or 2D patterning on static surfaces. Our findings expand upon these works by systematically quantifying the droplet propagation behavior and spatial accuracy across a range of donor–receiver distances (1.0–3.0 mm), using high-speed videography and custom image analysis tools. While Duocastella et al. and Patrascioiu et al. discussed jet formation and dynamics, the current study bridges this physical understanding with practical device integration, particularly within a microfluidic chip environment.
To our knowledge, this is one of the first studies to directly integrate LIFT bioprinting into a closed OoC device and validate compatibility under perfusion. The successful deposition and proliferation of LLC cells within the chamber demonstrates that LIFT can overcome one of the key challenges of bioprinting in microenvironments, which is precise delivery in confined spaces without damaging sensitive bioinks or disrupting printing geometries. Compared to inkjet or extrusion-based methods, which suffer from lower resolution and shear-induced cell stress (Koch et al.), LIFT offers a highly localized, low-shear alternative suited for advanced tissue models.
Furthermore, CFD simulation and particle velocimetry confirm that the fabricated chip provides physiologically relevant flow rates and diffusion-dominated transport in the inter-compartmental channel. This is critical for mimicking tumor–lymphatic interactions, which remain challenging to model in vitro. Similar approaches have been attempted (Carvalho et al.), but often without integration of bioprinting techniques or quantitative print resolution analysis.
Despite these advances, limitations remain. The printing resolution deteriorates rapidly beyond 2.0 mm, limiting the flexibility of chip architectures. Additionally, while short-term cell viability was confirmed, long-term behavior, differentiation potential, and inter-compartmental signaling were not evaluated in this study. Future work should focus on multi-cellular and multi-material printing and explore chronic perfusion conditions in tumor-on-chip platforms.
In summary, this work presents a robust framework for LIFT bioprinting integration with microfluidic systems, providing both a physical basis for operational limits and a functional demonstration in OoC applications. By aligning jet dynamics with chip geometry, we contribute a reproducible design rule for future OoC platforms seeking to incorporate non-contact, high-resolution bioprinting. Our study is among the first to experimentally demonstrate and validate high-resolution, non-contact bioprinting inside a closed microfluidic environment under flow. This not only corroborates the feasibility of LIFT for biofabrication but also establishes a critical design threshold (≤2.0 mm) for effective jet propagation, which has not been explicitly defined in prior literature. By introducing quantifiable guidelines and demonstrating functional cell integration, our work lays the foundation for developing advanced in vitro cancer models with improved physiological relevance and greater translational potential for drug screening and personalized medicine.