Abstract
This study investigated the potential of using the changes in polyphenol composition of red wine to enable a more comprehensive chemometric differentiation and suitable identification of authentication markers. Based on high performance liquid chromatography-mass spectrometry (HPLC-MS) data collected from Feteasca Neagra, Merlot, and Cabernet Sauvignon finished wines, phenolic profiles of relevant classes were investigated immediately after vinification (Stage 1), after three months (Stage 2) and six months (Stage 3) of storage, respectively. The data were subjected to multivariate analysis, and resulted in an initial vintage differentiation by principal component analysis (PCA), and variety grouping by canonical discriminant analysis (CDA). Based on polyphenol common biosynthesis route and on the PCA correlation matrix, additional descriptors were investigated. We observed that the inclusion of specific compositional ratios into the data matrix allowed for improved sample differentiation. We obtained simultaneous discrimination according to the considered oenological factors (variety, vintage, and geographical origin) as well as the respective clustering applied during the storage period. Subsequently, further discriminatory investigations to assign wine samples to their corresponding classes relied on partial least squares-discriminant analysis (PLS-DA); the classification models confirmed the clustering initially obtained by PCA. The benefits of the presented fingerprinting approach might justify its selection and warrant its potential as an applicable tool with improved authentication capabilities in red wines.
1. Introduction
The fierce competition on the world market of wine, in general, and red wine, in particular, highlights the need for authenticity verification and confirmation. Strongly relying on these features, quality assurance has gained significant importance both for producers and consumers [1,2]. While the terms authenticity and traceability have been extensively discussed in the literature, these important features are yet to be entirely handled and well differentiated [3,4,5]. From the sensorial perspective, considering that an authentic red wine is not always a typic red wine, authenticity would be better correlated with typicity, and not with traceability [6,7]; the latter approach complicates the scientific endeavors more, taking into account the complexity of the involved influence factors [8,9,10,11,12]. In this setting, authenticity and typicity features are influenced by grape variety, growing conditions as well as the employed winemaking techniques [8,13,14,15,16,17]. For instance, aside from the year of harvest, the winemaking process may render two authentic wines to exhibit different typicity attributes [8,9].
Polyphenols represent a large class of secondary metabolites, with an important role in plant protection against environmental factors [18]. Besides their nutritional benefits, polyphenolic compounds from wine act as endogenous antioxidants, with anti-inflammatory potential as well as carcinogenic preventive capacity [19]. Polyphenol biosynthesis plays a key role in the diversity and accumulation of polyphenols in grapes and by-products, thus affecting the compositional and sensory profile of red wines [18,20,21,22,23,24]. To this extent, polyphenols are compounds with a decisive role in evaluating the notions of authenticity, typicity, and traceability of these wines [21,25,26,27,28].
The distribution of individual polyphenolic compounds varies along with their concentrations in wines [3]. Their extraction is generally affected by the fermentation temperature [29] and the duration of maceration [13,30,31]. Similar to the sensory constituents, individual polyphenolic species in each group or class are released differently in wines; this ultimately results in distinct attributes, and are perceived as quality descriptors and as potential traceability markers [32,33].
Considering the structural transformations during wine elaboration and maturation, determining the polyphenols’ composition is necessary to distinguish grape and wine quality [34]. Thus, polyphenol analytical fingerprints are commonly used for general characterization (variety differentiation and adulteration) [9,16,35], being regarded as indirect indicators for traceability [35]. When we discuss the applicability of utilizing polyphenolic compounds as appropriate chemical markers for wine traceability [9], a series of factors have to be taken into account (reliability, selectivity, accuracy, and reproducibility) with respect to the involved instrumentation [16].
In this regard, given that red wine style and quality is significantly influenced by polyphenols [31], their analysis is conducted using various analytical [36] and statistical methods [37]. Moreover, the mainly studied application toward differentiating grape cultivars is the use of phenolic compounds as chemical markers through combined analytical techniques [38]. Among these, polyphenol datasets obtained by high performance liquid chromatography-mass spectrometry (HPLC-MS) have been widely applied in conjunction with chemometrics [35]. This is a suitable approach for the analysis of various known and unknown samples in fingerprinting and profiling assessments in order to improve the interpretation of the results [35,39].
Anthocyanins, flavonols, and flavan-3-ols are among the main flavonoid classes exploited as chemical markers in wine quality [38]. Previous studies have reported that polyphenol compositional profiles in wine show notable botanical and environmental variations [14,39,40,41,42]. For example, anthocyanin profile analysis has provided important information used for the varietal differentiation [9,14,43] as well as information on their implication in vintage discrimination of red wines [43,44]. In addition, other compounds such as flavanols, phenolic acids, and resveratrol could be of use for vintage classification [45]. Additionally, in the attempt to classify red wines according to geographical origin, flavanols and flavonols have been proposed as production area indicators [46,47]. Moreover, red wine regional identity was correctly assigned by involving phenolic acid profiles (gallic acid, p-coumaric acid, ferulic acid, and caffeic acid) [14,46,48,49,50]. Very recently, Wu et al. (2021) effectively differentiated Chinese red wines according to geographical origin on the basis of phenolic compounds by employing HPLC-DAD (diode array detector) method combined with chemometrics [51]. Furthermore, Pisano noted that three malvidin-derived anthocyanins contributed mainly to the geographical origin discrimination of the studied samples [52].
Given the benefits of combined analytical and statistical methods for reporting polyphenol levels in grapes and wine [53,54], in this paper, we used the phenolic profiles obtained by HPLC-MS to assess the variety and terroir characteristics of three red wines from Feteasca Neagra (FN), Cabernet Sauvignon (CS), and Merlot (M) grape varieties grown in two viticultural areas in Romania. The compounds assessed herein belong to relevant phenolic classes (phenolic acids, flavonols, flavan-3-ols, anthocyanins, anthocyanidins) and are important markers generally focused on during authenticity assessment [38].
The analyses targeted the improvement of distribution and grouping of the wine samples according to varietal and zonal factors. In addition, the use of multivariate statistical analysis sought to identify additional descriptors as certain ratios of individual polyphenols; these would entail potential fingerprinting markers for red wine authentication, applicable on compositional data for traceability-wise characterization studies.
2. Materials and Methods
2.1. Reagents and Standards
Caffeic acid, quercetin, rutin, (−)-catechin, epicatechin, cyanidin, delphinidin, malvidin, peonidin-3-glucoside, and pelargonidin, all of analytical grade, were supplied by Sigma-Aldrich, Germany. Ellagic acid, gallic acid, myricetin, and luteolin were obtained from Fluka, Germany. All solvents used for chromatography were of HPLC analysis grade. All other reagents were of analytical purity or chromatographic grade and were used after filtration. The ultra-pure water was obtained using a water purification system, Elix 3 (Millipore). If not indicated otherwise, all solutions were stored at 4 °C and protected from light, under inert atmosphere, and were filtered before analysis using a Syringe Driven Filter Unit 0.2 μm (Chromafil, PTFE-polytetrafluoroethylene, Macherey-Nagel, Düren, Germany).
2.2. Wine Samples
The studied samples were red wines obtained from international (Merlot—M, and Cabernet Sauvignon—CS) and Romanian (Feteasca Neagra—FN) varieties of V. vinifera species. They were cultivated in two well-known viticultural areas in Romania. Grapes were harvested at their technological maturity, from the Murfatlar vineyard (years 2014 and 2015) and Valea Calugareasca vineyard (year 2015).
Wine samples (n = 9), obtained by traditional vinification from the aforementioned grape varieties, were collected at three different stages of maturation (aging): immediately after the end of the vinification process (Stage 1), at three months (Stage 2) and six months (Stage 3) of storage in stainless steel containers. Sample code designation was conducted according to area and vintage, as follows: Murfatlar 2014—FN1, CS1, and M1; Murfatlar 2015—FN2, CS2, and M2; and Valea Calugareasca 2015—FN3, CS3, and M3.
In order to avoid polyphenol degradation, all wine samples were stored under controlled conditions and were supplied to the laboratory prior to analysis. The bottles were opened, filtered (0.2 μm PTFE membrane filter), and injected into the HPLC-MS without any selective extraction.
2.3. HPLC-MS Analysis
All determinations were performed on a Shimadzu HLPC system (Kyoto, Japan) coupled to a LCMS-2010 mass spectrometer detector with an electrospray ionization interface. The equipment comprised two LC-20ADsp pumps, a SIL-20AC autosampler, a CTO-20AC column oven, a DGU-20A5 degasser, and LC Solution software.
The identification of phenolic acids, flavonols, and flavan-3-ols was carried out according to a method previously reported by Alecu et al. [15]. Briefly, 20 μL of the sample was injected into a Nucleosil 100-3.5C18 Kromasil (100 × 2.1 mm) column and eluted for 70 min at 20 °C (column temperature). The used mobile phase consisted of water (solvent A) and acetonitrile (solvent B), adjusted to pH 3.0 with formic acid. The negative ionization mode was used for polyphenolic compound assessment. The phenolic compound separation was performed using binary gradient elution as follows: the initial 5% solvent B was increased to 30% for 20 min (0.01–20 min) maintained at 30% for 20 min (20.01–40 min), increased to 50% for 10 min (40.01–50 min), decreased to 5% (50.01–52 min), and maintained at 5% for the remainder of the elution (52.01–70 min). The initial flow rate (0–5 min) was 0.1 mL/min; it then ramped to 0.2 mL/min (5.01–15 min), decreased to 0.1 mL/min (15.01–35 min), increased to 0.2 mL/min (35.01–60 min), and dropped to 0.1 mL/min on the final stage (60.01–70 min). The column was equilibrated for 1 h before injection.
Anthocyanin and anthocyanidins were quantified using reversed-phase HPLC-MS (positive ionization mode) using a method developed by Albu et al. [55]. The analytes were eluted with water–formic acid (95:5, v/v) (solvent A) and methanol–formic acid (95:5, v/v) (solvent B) by a Nucleosil 100-3.5C18, KROMASIL, 100 × 2.1 mm column, at 40 °C (column temperature) for 47 min. The compound separation was performed using a flow rate of 0.15 mL/min and a binary gradient. The elution was initiated with 6% solvent B (0 min), increased to 30% solvent B (0.01–10 min), ramped to 50% (10.01–25 min), and then to 60% (25.01–30 min), decreased to 6% (30.01–36 min), and further kept at 6% solvent B until the end (36.01–47 min).
For the qualitative analysis, the full scan acquisition mode (SCAN) was used; for the quantitative analysis, the monitoring of the selected ions acquisition mode (SIM) was used, screening [M-H]−1 ions for phenolic acids, flavonols, and flavan-3-ols, and [M-H]+1 ions for anthocyanin and anthocyanidins, respectively.
2.4. Statistical Analysis
Each wine sample (n = 9) was tested in triplicate and the obtained dataset comprised concentration values of quantified polyphenols. The obtained data were processed according to variety, vintage, and wine-growing region and expressed by mean values and standard deviations. Results were analyzed by one-way analysis of variance (ANOVA), and statistical significance was assessed by the Tukey HSD post hoc test (p < 0.05).
Bootstrapping (resampling method) was performed in order to assess the appropriateness of the obtained dataset (polyphenol profiles) for the present application. Briefly, bootstrapping approximates the process of taking repeated samples from the target population with replacement, resulting in new datasets; the output is a large number of bootstrapped samples used to estimate the standard error and confidence intervals for the statistic of interest.
Principal component analysis (PCA), an unsupervised pattern recognition technique, was employed to separate the wine samples into groups according to the previously mentioned criteria. In JMP® (SAS Institute Inc., Cary, NC, USA, 1989–2021.), when calculating principal components “on correlations”, the input variables are centered and scaled. Loadings are calculated by multiplying the eigenvectors by the square root of their eigenvalues.
Sample patterns were deduced form the simultaneous interpretation of score and loading plots of the first two principal components. Similar to PCA, canonical discriminant analysis (CDA) is an unsupervised method used to determine the extent to which variables are related and find patterns in the data. It was used for the validation of the obtained varietal separation. Mutual polyphenol dependencies based on the PCA correlation matrix (pairwise) and the cluster analysis (Ward method) were further exploited in the classification. After designating the suitable parameters for the classification of the samples, a combination of HPLC-MS variables was included in the data matrix in order to improve the distribution by PCA.
Further discriminatory studies to assign wine samples to their corresponding classes relied on partial least squares-discriminant analysis (PLS-DA), a supervised pattern recognition technique employing the NIPALS (nonlinear iterative partial least squares) algorithm with two latent variables to build the classification models, and cross validation under the KFold approach (K = 7). All statistical analyses were performed using SAS JMP® software.
3. Results and Discussion
3.1. Analytical Performance
Identification of compounds was based on the retention times and the mass spectra of individual compounds, and polyphenol quantification was made by calibration curves obtained with standard solutions of individual compounds. The performance characteristics of the HPLC-MS method are presented Supplementary Materials Table S1.
3.2. Evaluation of Wine Polyphenol Profiles
In this study, the evolution of the polyphenolic compounds from the Murfatlar (2014), Murfatlar (2015), and Valea Calugareasca (2015) wines is shown in Table 1. The differences in the polyphenol profiles were evaluated by means of one-way ANOVA, employing Tukey’s HSD test (p < 0.05). Within their respective technological stage, the concentrations of polyphenols varied significantly in red wines, depending on the grape variety vintage and area of origin.

Table 1.
Polyphenolic profile of wines from Murfatlar (2014), Murfatlar (2015), and Valea Calugareasca (2015).
During post-harvest processing, the maceration-fermentation step has strong implications on the optimum extraction of polyphenols [3,8,56,57,58].
The assayed technological stages revealed variations mainly in the case of the phenolic acids; their concentrations registered a drop during the first three months of storage, followed by a slight increase until the end of the study period (6 months of storage). Among the most frequently present phenolic acids in red wines, gallic acid is the most abundant [1,59]. In our study, its highest concentration was registered in Murfatlar (2014) in Stage 1 and averaged between 208.48–246.74 µg/mL.
The predominant detected flavonol was quercetin and had the highest concentrations in Murfatlar (2014) wines (4.89–12.52 µg/mL); its content follows a decreasing trend during storage in Murfatlar, compared with the steady levels in Valea Calugareasca.
Among the flavan-3-ols, epicatechin registered a higher concentration in 2014 (Murfatlar), compared with 2015 (irrespective of viticultural area), and highlighted a decreasing trend after the fermentative processes in all wine samples. On the contrary, (−)-catechin was the most abundant in 2015 (8.82–14.98 µg/mL in Murfatlar, and 9.65–12.03 µg/mL in Valea Calugareasca), and had an increasing evolution overall.
The distribution of chromatic compounds in the analyzed red wines revealed the anthocyanin peonidin 3-O-glucoside (128.34–155.86 µg/mL) as the most abundant in Murfatlar (2014), whereas among anthocyanidins, malvidin registered on average higher concentrations in the three analyzed varieties harvested in 2015 from both geographical areas. Similarly, it has been observed that peonidin-3-O-glucoside was the predominant pigment in the skins of the Garnacha Tintorera grape variety [60] as well as in several Italian varieties such as Moscato Rosa, Galliopo, Nebbiolo, etc. [61]. Additionally, in the Portuguese variety Alvarilhão, it was determined in amounts similar to those registered for malvidin-3-O-glucoside [62]. More recently, Kyraleou et al. (2020) observed that, while discriminating five Greek red grape varieties based on the anthocyanin profiles, peonidin-3-O-glucoside was the most abundant anthocyanin in Kotsifali variety, in both the 2017 and 2018 studied seasons [63]. Although the coloring matter exhibited an overall decreasing trend throughout the study period, delphinidin and cyanidin had a more static progress during the analyses.
During aging, wines are subjected to slow oxidation reactions, resulting in color stabilization by affecting the composition of volatile compounds and polyphenols [64]. For example, anthocyanins are prone to structural transformations and are unstable in aqueous solutions, resulting in changes in their original concentrations (degradation reactions) [43,65,66].
These results are in agreement with the findings of other reports [67,68,69], which inform on the occurrence of various transformations (oxidative and polymerization processes) at the level of major compounds.
Accordingly, on the basis of the evolution shown thus far, the observed differences are further exploited for authenticity evaluation.
3.3. Chemometric Differentiation of Wine Samples after Vinification
To support the relevance of the sample set, prior to the PCA analysis, bootstrapping was performed on the target variables by repeatedly sampling from the original dataset. The estimated sampling distribution is presented in Figure S1. The outcome shows that the occurring resample averages are between the confidence limits, and near the original estimate, which provides practical significance for the assessed parameters.
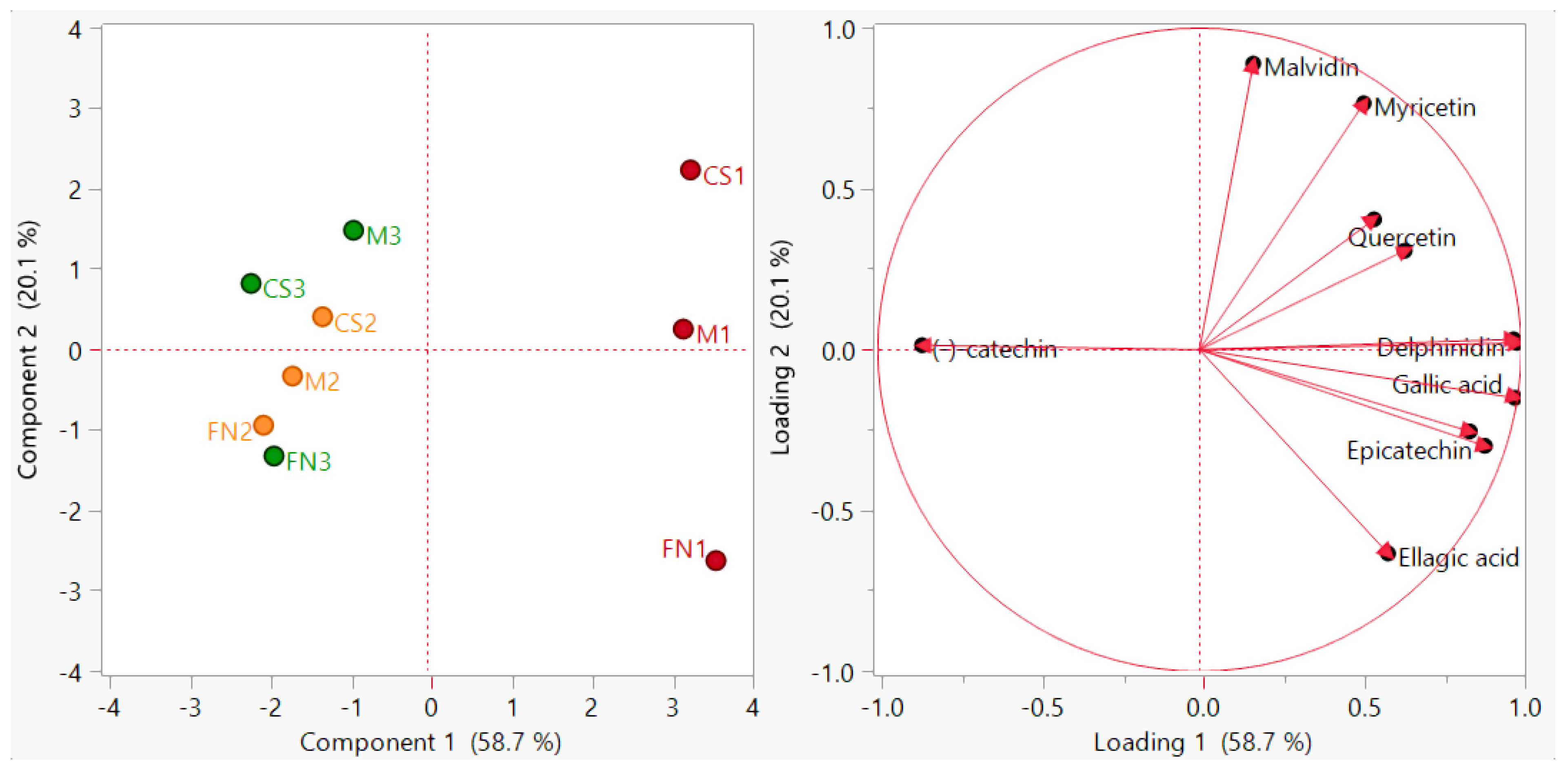
The PCA distribution diagram presented in Figure 1 was obtained by excluding several redundant variables (rutin, quercetin-3-β-d-glucoside, luteolin, and pelargonidin) from the initial data matrix due to their low concentrations (<LoD). The application of the PCA method on the analyzed red wines managed to differentiate them according to vintage; the first two principal components were retained and accounted for a total of 78.8% of the variance (PC1 = 58.7%, PC2 = 20.1%).
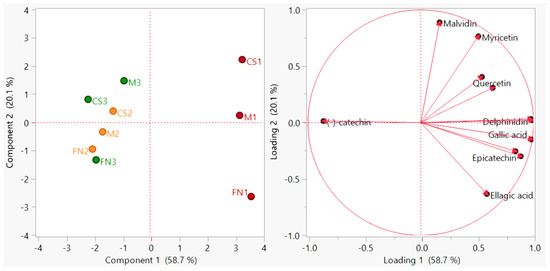
Figure 1.
PCA diagram—sample grouping (scores plot) and polyphenol distribution (loadings plot) for Stage 1 wines. Color coding: red—Murfatlar 2014; orange—Murfatlar 2015; green—Valea Călugărească 2015.
In this case, the associated loading vectors indicate the variables which significantly influenced data rearrangement in the new system of axes; among the analyzed polyphenolic compounds, gallic acid, delphinidin, epicatechin, peonidin-3-O-glucoside and cyanidin had a major impact on the first principal component, while malvidin, myricetin and caffeic acid highly influenced the second principal component. These observations revealed that the discrimination capability of the polyphenols to classify the studied samples has certain limitations.
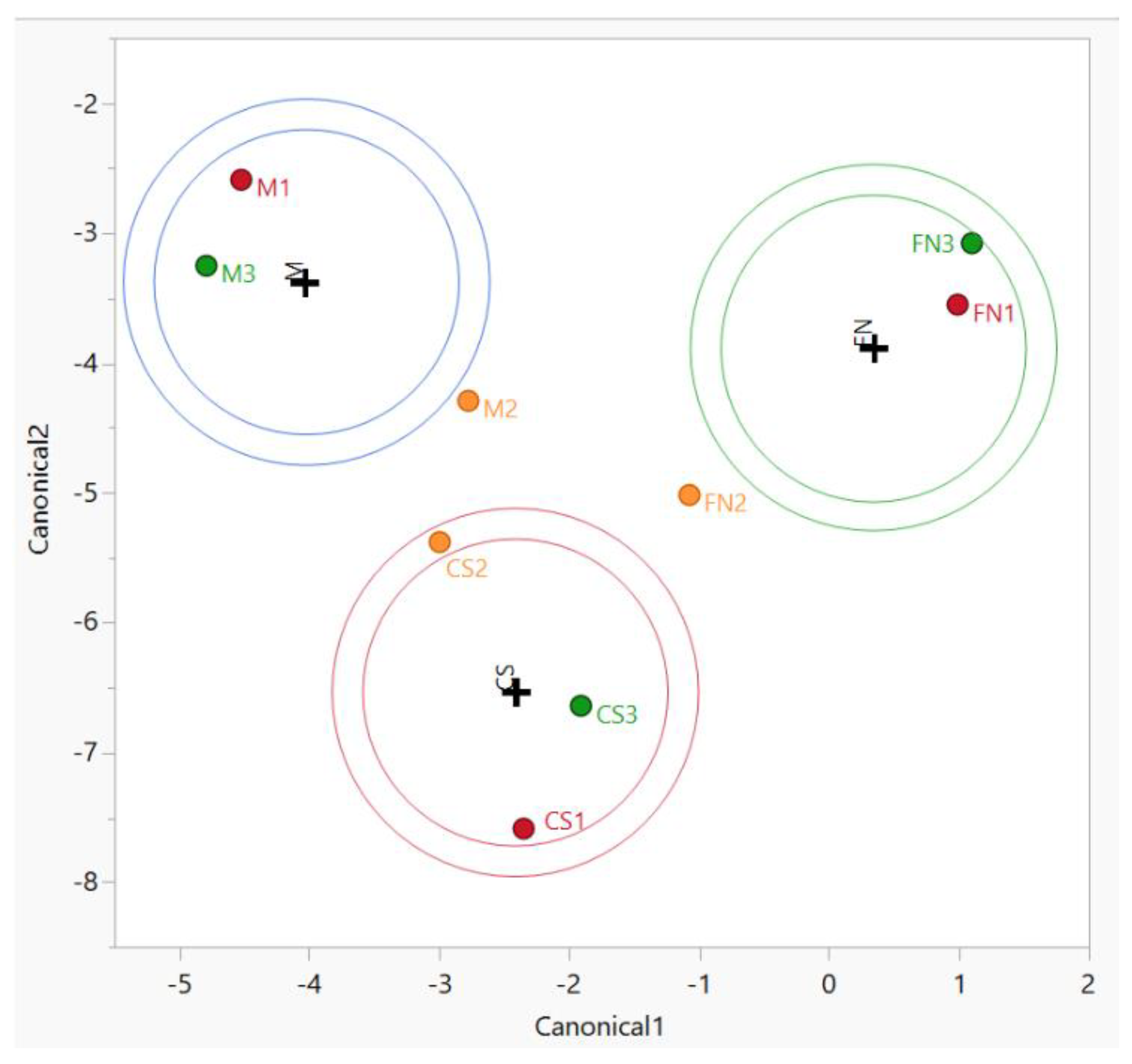
Consequently, we examined further approaches in order to supply additional biomarker candidates toward distinguishing wine classes (groups) through PCA. We used canonical discriminant analysis (CDA), which involves the derivation of some canonical variables that can explain inter-class variation in a similar manner to that of PCA [1,49]. Due to the lack of a complete dataset for the other factors (vintage and geographical origin), only the variety-based authenticity was assayed by CDA (Figure 2).
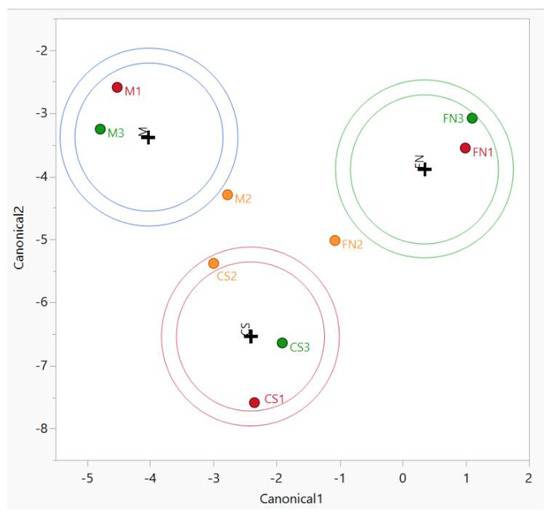
Figure 2.
CDA diagram—sample grouping based on variety for Stage 1 wines. Color coding: red—Murfatlar 2014; orange—Murfatlar 2015; green—Valea Călugărească 2015.
Using the same variables as for PCA, the variance for the separation resulted in 63% for Canonical 1 and 37% for Canonical 2, with significant differences between grape varieties (p < 0.0001).
Based on the obtained standardized coefficients (Table 2), ellagic and caffeic acids show a strong positive link to Can. 1, while epicatechin and myricetin are negatively correlated. For Can. 2, epicatechin, quercetin, and caffeic acid are positively correlated, whereas myricetin and gallic acid revealed a negative influence.

Table 2.
Standardized canonical discriminant functions for the differentiation based on variety using CDA.
These findings provide further insight regarding red wine authentication according to variety based on polyphenol fingerprints. Additionally, they outline future research directions in analyzing the transformations of “target” polyphenols in red wines during storage [1,70,71].
3.4. Preliminary Assessment of Tentative Markers
Metabolic changes during grape berry development have been outlined as important chemical parameters used in variety-based classification [21,72], with a strong relationship between the biosynthesis pathway and polyphenol distribution in wine [73]. The study of Muccillo et al. (2014) revealed the successful varietal classification of the analyzed wines, involving (+)-catechin/(−)epicatechin and malvidin-3-acetylglucoside/malvidin-3-coumaroylglucoside ratios, biochemical attributes related to the phenylpropanoid pathway [34], based on the common features among polyphenolic compounds [20,74].
In this context, the next experimental step was the inclusion of several new variables (ratios between the individual polyphenol concentrations) into the data matrix.
3.4.1. Ratios between Individual Polyphenol Concentrations
Their selection was achieved taking into consideration their common biosynthesis route and branching point [75,76], generating the potential for using combinations of downstream derivatives as traceability markers [66,72,77].
In addition, we also investigated the correlation matrix of the initial PCA distribution. This allowed the identification of several significant (* symbol) relationships among the tested polyphenols: ellagic acid and epicatechin (0.6971, p = 0.0369 *), quercetin and myricetin (0.7902, p = 0.0113 *), gallic acid and epicatechin (0.8919, p = 0.0012 *), (−)-catechin and epicatechin (−0.6838, p = 0.0422 *), (−)-catechin and gallic acid (−0.8608, p = 0.0029 *), delphinidin and epicatechin (0.8635, p = 0.0027 *), delphinidin and gallic acid (0.9817, p < 0.0001 *), delphinidin and (−)-catechin (−0.8767, p = 0.0019 *), peonidin-3-O-glucoside and epicatechin (0.8502, p = 0.0037 *), peonidin-3-O-glucoside and gallic acid (0.9665, p < 0.0001 *), peonidin-3-O-glucoside and (−)-catechin (−0.8914, p = 0.0012 *), peonidin-3-O-glucoside and delphinidin (0.9894, p < 0.0001 *), malvidin and myricetin (0.6918, p = 0.0390 *), cyanidin and epicatechin (0.8074, p = 0.0085 *), cyanidin and gallic acid (0.8418, p = 0.0044 *), cyanidin and delphinidin (0.7804, p = 0.0131 *), and cyanidin and peonidin-3-O-glucoside (0.7791, p = 0.0133 *). These interactions are considered to be influenced by the chemical reactions occurring during the vinification process and are involved in polyphenol fingerprinting [1,19,70].
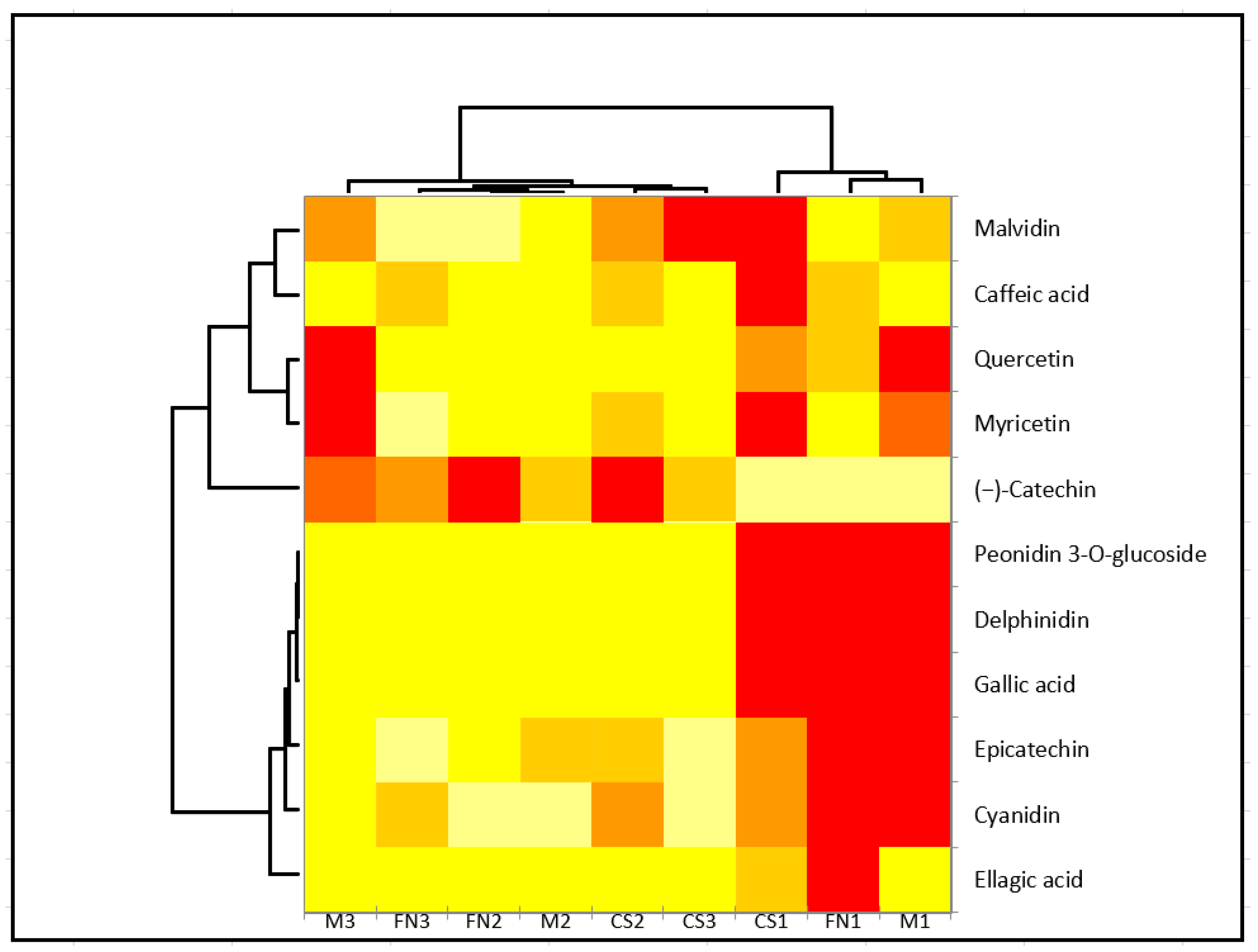
Subsequent application of cluster analysis (Ward method) supported the observed mutual dependencies and verified the initial vintage differentiation by PCA (Figure 3).
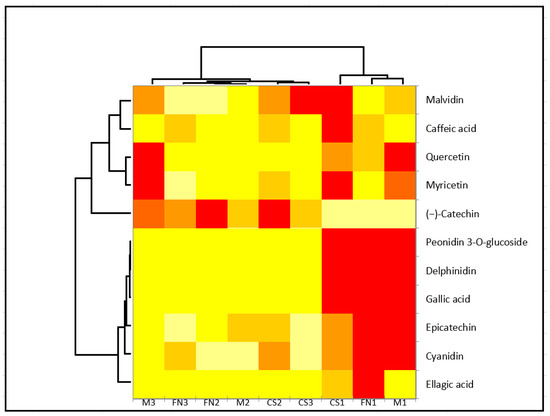
Figure 3.
Heatmap of quantified polyphenolic compounds based on variety. Clustering between variety and also polyphenols was selected. The correlation is colored in shades of yellow (low correlation) to red (high correlation). Sample code designation was conducted according to area and vintage: Murfatlar 2014—FN1, CS1 and M1; Murfatlar 2015—FN2, CS2 and M2; and Valea Calugareasca 2015—FN3, CS3 and M3.
The new variables were represented by the following ratios: peonidin-3-O-glucoside/cyanidin; peonidin-3-O-glucoside/(−)-catechin; quercetin/cyanidin; quercetin/peonidin-3-O-glucoside; (−)-catechin/epicatechin; malvidin/delphinidin. In a similar manner, besides the individual color components, Geana et al. (2016) showed that the use of anthocyanin ratios allowed a fine variety and vintage separation of five Romanian red wines, using a linear discriminant analysis [78].
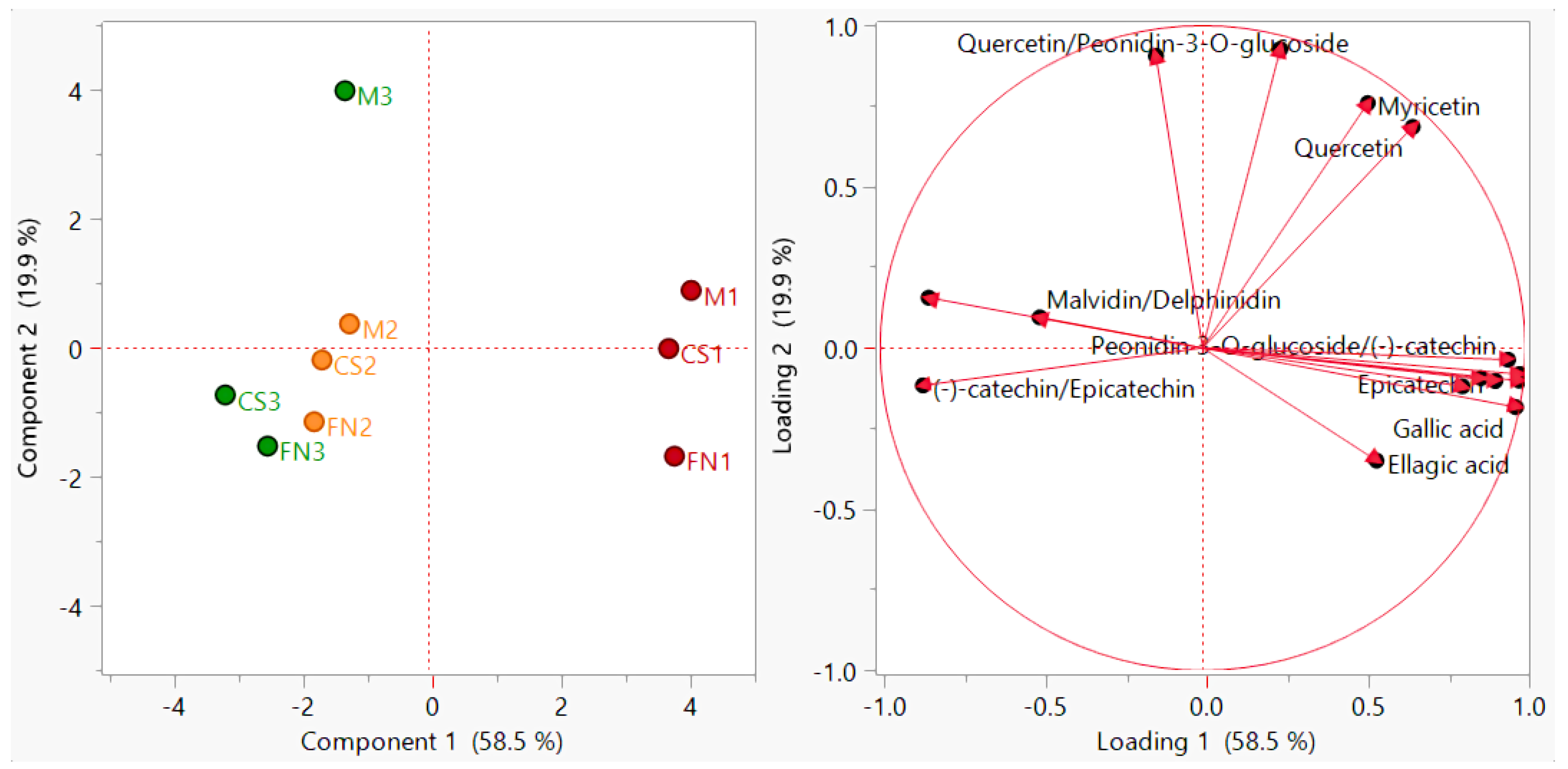
The addition of these supplementary variables (Figure 4) delineates the simultaneous sample clustering according to geographical origin, harvest year, and variety. The observed differentiation did not result in an enhanced discrimination value (PC1—58.5% and PC2—19.9%), but refined the separation. In this regard, several literature reports confirm the interest in raising the accuracy of polyphenol fingerprint-based authentication of red wines [40,79,80].
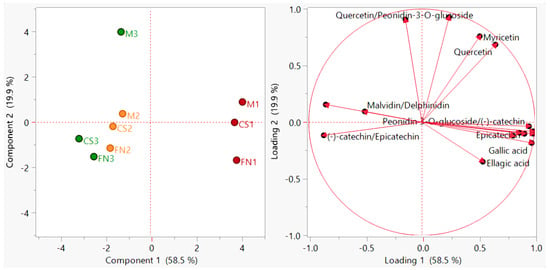
Figure 4.
PCA diagram—improved sample grouping (scores plot) and polyphenol distribution (loadings plot) for Stage 1 wines. Color coding: red—Murfatlar 2014; orange—Murfatlar 2015; green—Valea Călugărească 2015.
3.4.2. Fingerprinting Application of Proposed Polyphenol Markers during Storage
The chemical mechanisms and pathways involved in the formation and/or extraction of phenolic compounds have been thoroughly investigated [30,81]. In this regard, mass transport and reaction kinetics for these species are closely related to the transformations occurring during wine elaboration and storage [30,82].
As such, different evolution trends have been observed depending on the initial composition of wine samples such as color or polyphenolic composition [81]. For example, a decrease in anthocyanin content was observed in the first months of aging [83]. The initial drop might be attributed to precipitation and oxidation reactions as well as the formation of anthocyanin–anthocyanin complexes, or the formation of more stable pigments [33]. In contrast, a substantial decrease in non-anthocyanin polyphenols occurs after three months of storage, especially in the case of flavan-3-ols [83]. Other authors indicated the use of additional markers besides monomeric anthocyanins such as dimers of flavan-3-ols and polymerized pigments [84].
In our case, the importance of the modifications during storage is illustrated by the PCA. In order to verify the potential use of the introduced markers, we assessed the separation of the red wines subjected to storage conditions. Figure 5 shows the loading plots for wine sample grouping at Stages 2 and 3.
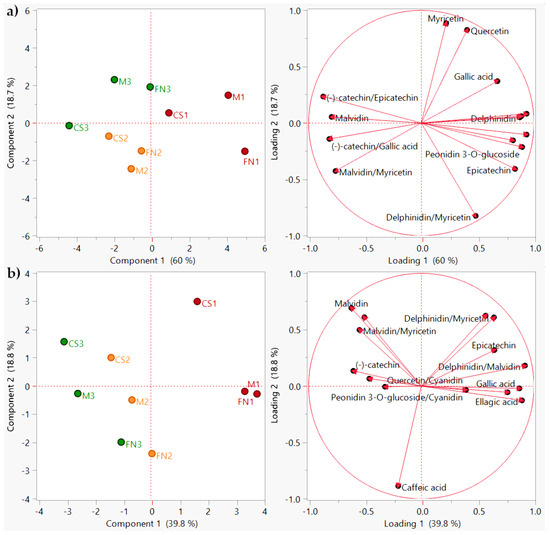
Figure 5.
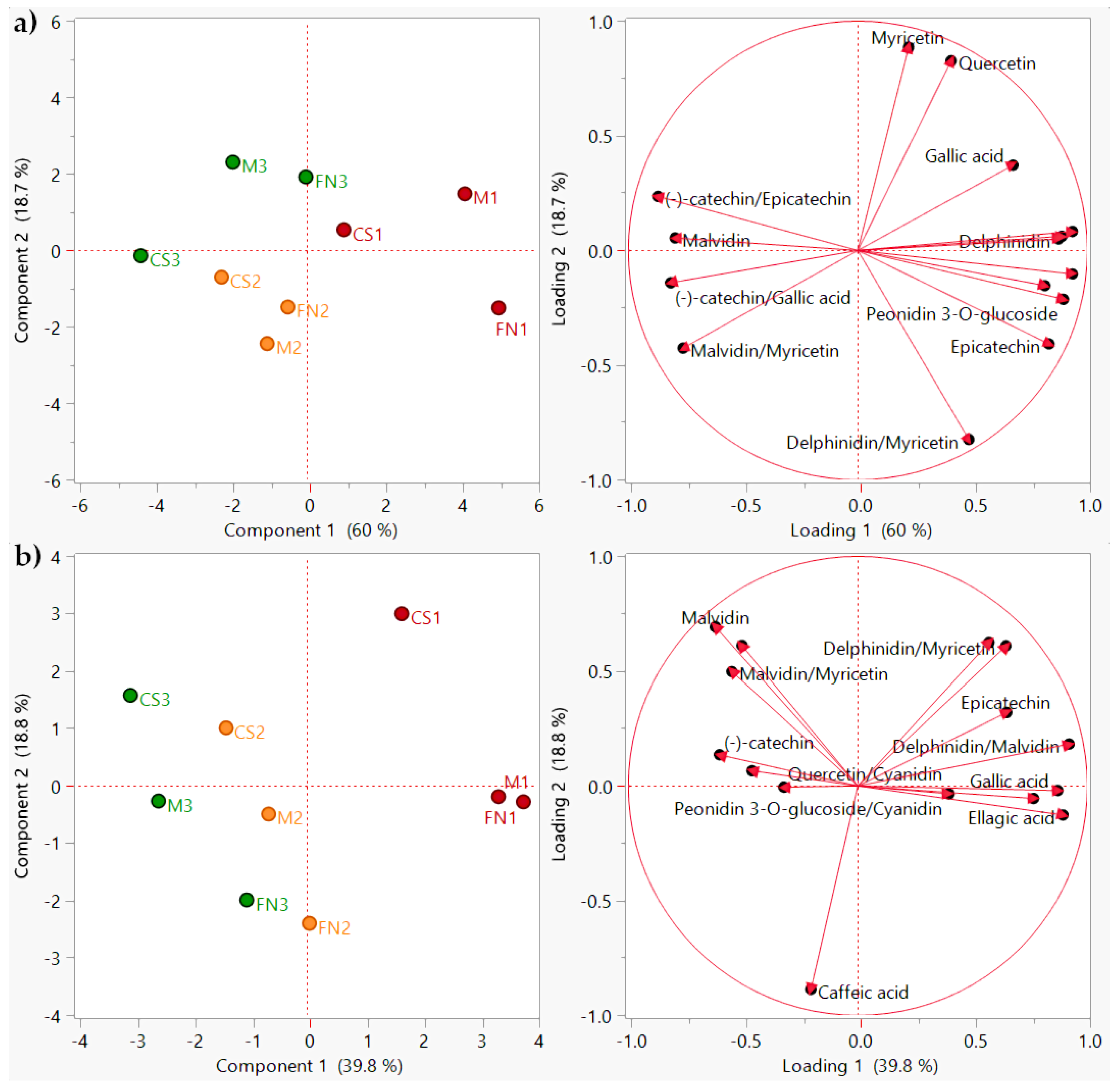
PCA diagram showing sample clustering (scores plot) and polyphenol distribution (loadings plot) during storage. (a) Stage 2–3 months; (b) Stage 3–6 months. Color coding: red—Murfatlar 2014; orange—Murfatlar 2015; green—Valea Călugărească 2015.
For the 3-month storage period, the clustering was achieved only for vintage and area of origin (Figure 5a), showing mild varietal misclassification. Results indicate that the first two components account for 78.7% of the total variance, first component 60.0%, and second component 18.7%. The PCA was constructed based on the addition of several phenolic ratios to the initial data matrix: delphinidin/malvidin, peonidin-3-O-glucoside/cyanidin, delphinidin/myricetin, peonidin-3-O-glucoside/(−)-catechin, malvidin/myricetin, (−)-catechin/epicatechin, and (−)-catechin/gallic acid.
Similarly, for the 6-month storage period, the PCA model using the refined dataset revealed that the retained variance was 58.6%, with 39.8% for PC1, and 18.8% for PC2 (Figure 5b). Polyphenol profiles have been exploited to include additional descriptors: delphinidin/malvidin, peonidin-3-O-glucoside/cyanidin, delphinidin/myricetin, peonidin-3-O-glucoside/(−)-catechin, malvidin/(−)-catechin, malvidin/myricetin, quercetin/cyanidin, and quercetin/peonidin-3-O-glucoside. Even though the score plots showed scattered sample distribution, clear groups are displaying simultaneous variety, vintage, and area differentiation.
3.4.3. Classification Studies by PLS-DA
This section is focused on the assessment of classification models by PLS-DA as an initial survey toward red wine authentication. Encouraged by the classification results obtained by PCA, the supervised PLS-DA method was employed to predict the category attribution according to the polyphenolic profiles in red wines.
The PLS-DA score plot for the final model for Stage 1 is shown in Figure 6. It delineates a clear separation according to vintage as well as a tendency for variety clustering with mild miss-classification. The variable importance table is useful to assess the real contribution of each VIP (variable importance for projection). The addition of polyphenol ratios as supplementary variables, similar to the PCA results, indicates their potential to be integrated as authenticity in future studies. A further application of the PLS-DA method was carried out for the wines from Stage 2 and 3, and is presented in Figure 7 and Figure 8.
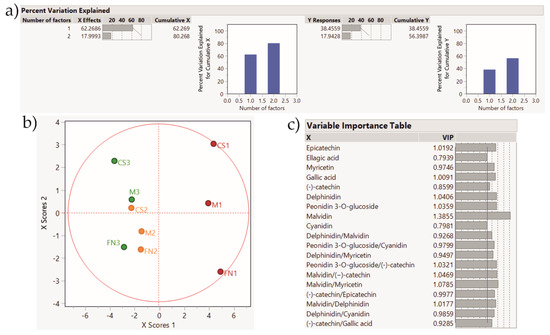
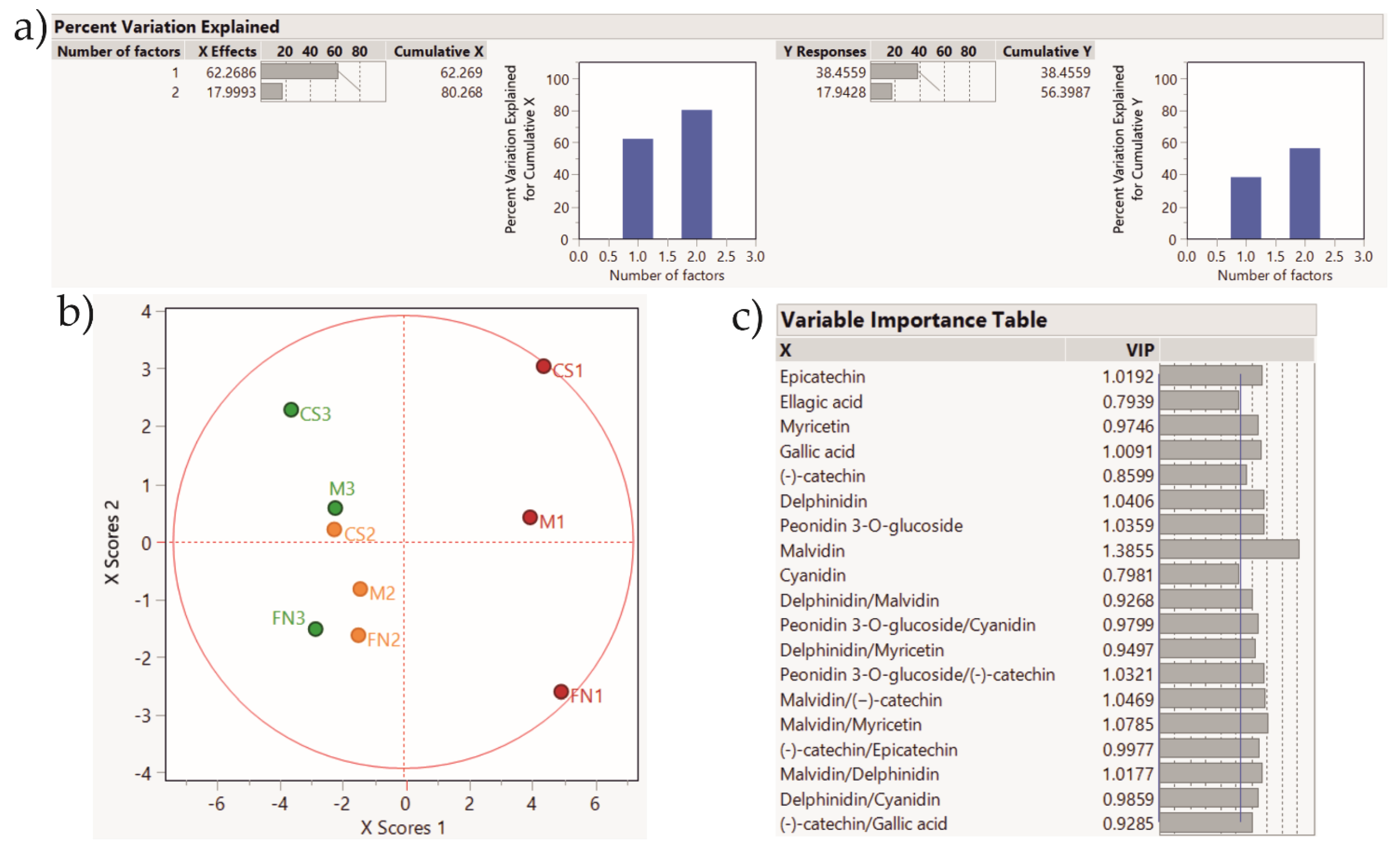
Figure 6.
PLS-DA score plot showing sample clustering for Stage 1 wines. Color coding: red—Murfatlar 2014; orange—Murfatlar 2015; green—Valea Călugărească 2015. (a) Percent of variation explained by the two factors; (b) PLS-DA score plot; (c) VIPs (Variable Importance for the Projection) showing the predictors and their importance in determining the PLS projection model.
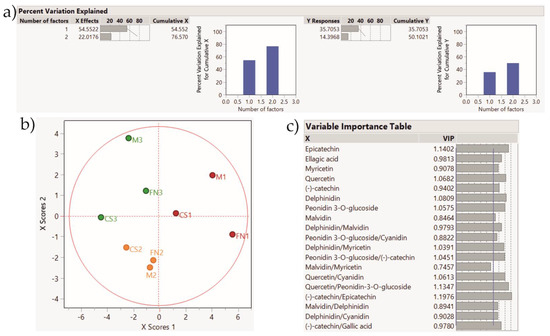
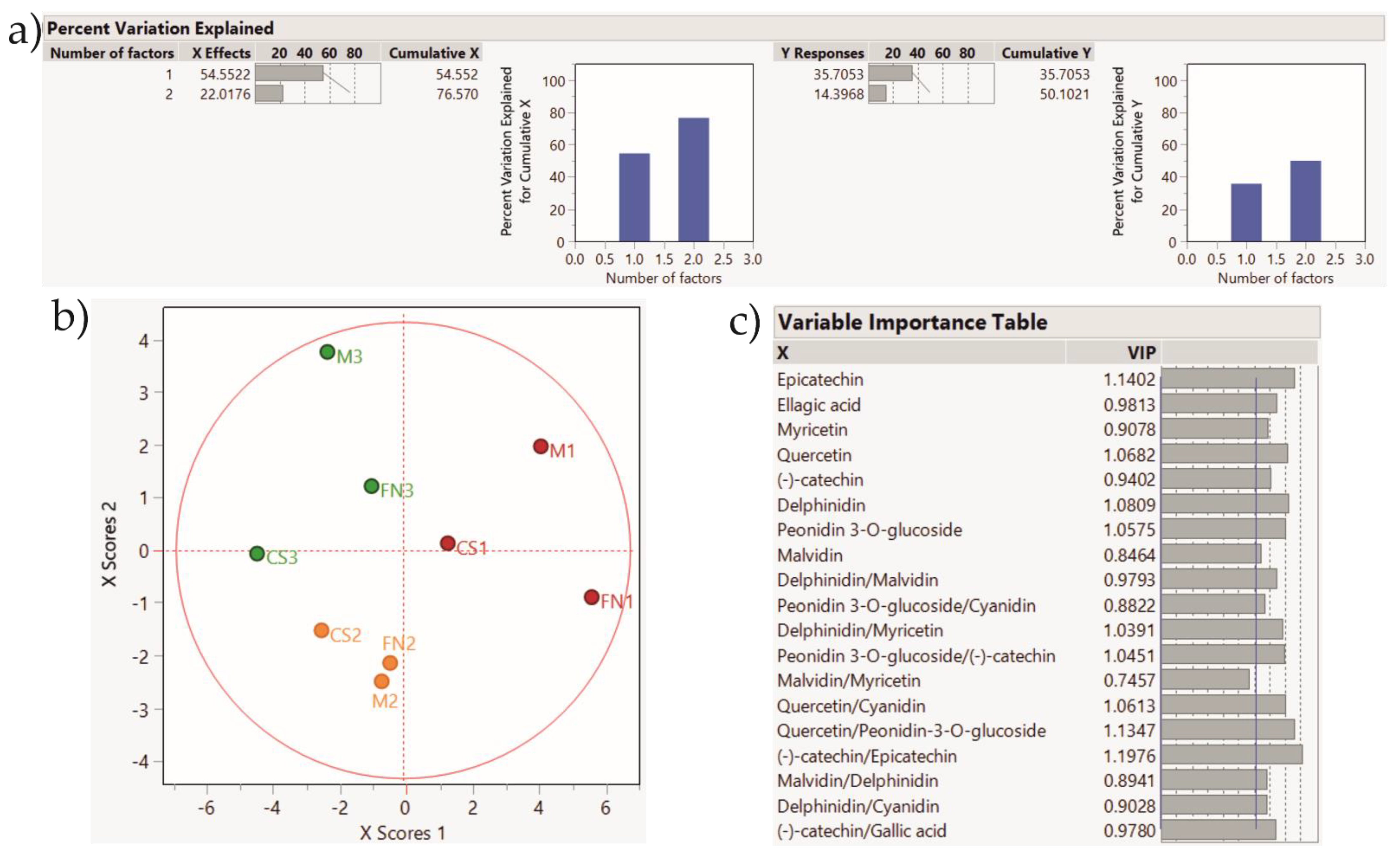
Figure 7.
PLS-DA score plot showing sample clustering for Stage 2 wines. Color coding: red—Murfatlar 2014; orange—Murfatlar 2015; green—Valea Călugărească 2015. (a) Percent of variation explained by the two factors; (b) PLS-DA score plot; (c) VIPs (Variable Importance for the Projection) showing the predictors and their importance in determining the PLS projection model.
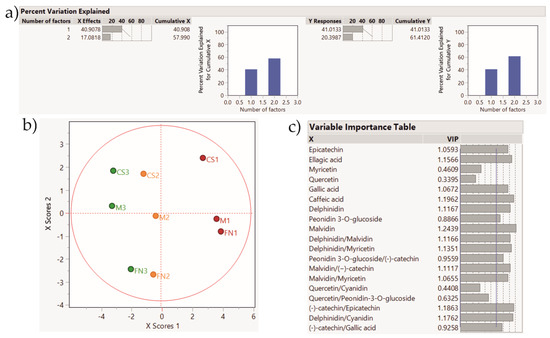
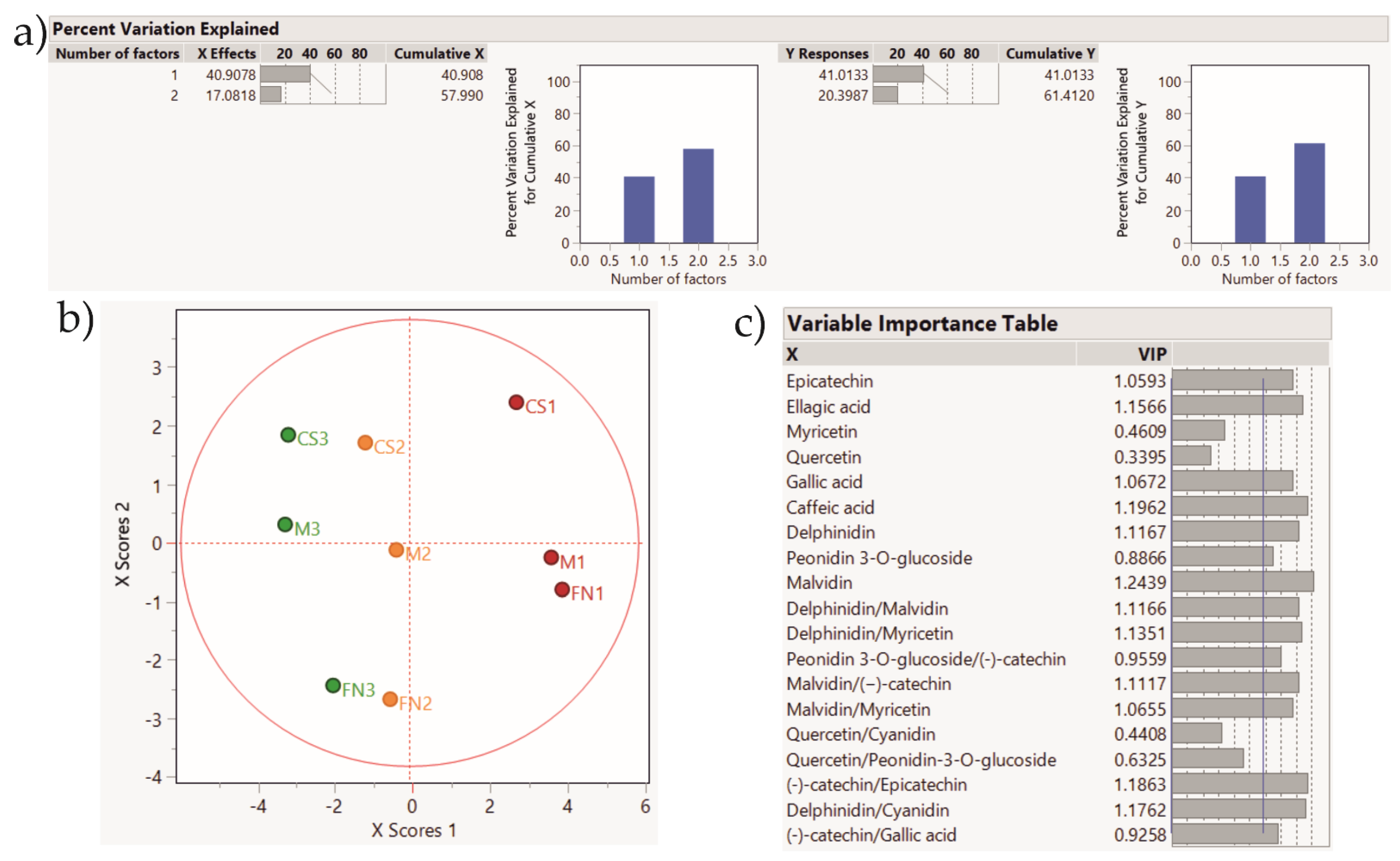
Figure 8.
PLS-DA score plot showing sample clustering for Stage 3 wines. Color coding: red—Murfatlar 2014; orange—Murfatlar 2015; green—Valea Călugărească 2015. (a) Percent of variation explained by the two factors; (b) PLS-DA score plot; (c) VIPs (Variable Importance for the Projection) showing the predictors and their importance in determining the PLS projection model.
The application of the PLS-DA method based on the tentative markers for Stage 2 red wines rendered a clear clustering of the samples according to vintage and geographical area. Accordingly, the PLS-DA method applied on Stage 3 wines depict sample grouping based on vintage and geographical area, and the tendency for variety clustering.
Using the polyphenol generated fingerprints along with the tentative markers (polyphenol ratios), we observed sample separation according to the tested factors, and validated the results obtained by PCA. We believe that the method is not limited to the cases presented herein and merit further investigation, and could be adapted to more complex studies.
3.5. Study Limitations
The present study was intended to stand as a potential tool in support to wine traceability and authenticity, offering added-value functionality. However, given the nature of the employed methods, some drawbacks of our study include (a) the use of small sample sets, which may render the application of the tentative markers not conclusive, and (b) the use of a small number of relevant analytes to generate red wine fingerprints, which is far too few for the polyphenolic compounds in red wines.
As a means of providing practical relevance for the small sample number, bootstrap was carried out (100 resamples), and afforded good resampling distribution of the assessed parameters, within the confidence limits and near the original estimates. These observations warrant the applicability for the purpose at hand. With regard to the employed polyphenol fingerprints, the array of compounds used in the present study does not match the variety found in red wines; however, they belong to relevant phenolic classes generally targeted during authenticity assessment. In this context, there are other studies employing a small number of polyphenolic compounds [50,54,84,85,86].
Additionally, wine authenticity is a concern for red wines, especially for expensive red wines, which involve long-time storage. A fair assumption would be that the short-aged wines used herein may not be representative enough for the sought application. Nonetheless, having achieved refined classification pertaining the proposed polyphenol ratios as biomarkers, this tool merits further exploration; it may serve as a preliminary indicator entailing its potential application on expensive wines, subjected to longer aging periods.
Moreover, considering the novel observation of tandem discrimination according to variety, vintage, and area of origin by PCA and confirmed by PLS-DA, the benefits of the proposed approach would be desirable. Taking into account both the drawbacks and the strengths of the study, the tentative traceability markers (phenolic ratios) could be effectively employed as additional descriptors to other study models on larger datasets involving category attribution, and under more thorough analytical conditions.
4. Conclusions
The analysis of the samples under study showed a strong coordination between polyphenol pathways (metabolism) and the associated profiles in finished wines. This allowed additional variables to be integrated into the system as potential authenticity markers–ratios between compositional values of individual polyphenols. In the case of storage evaluation, overall improved clustering was achieved through the addition of polyphenol ratios to the original PCA matrix. Subsequently, the discrimination of wine samples by PLS-DA confirmed the PCA classification. Thus, the proposed chemical descriptors point to a promising approach toward the differentiation of red wines by evaluating variety and terroir among the analyzed sample sets. Taken together, the profiling findings provided herein corroborated with the ongoing endeavors in wine polyphenol fingerprinting, and recommend the considered markers for routine integration in future explorations.
Supplementary Materials
The following are available online at https://www.mdpi.com/article/10.3390/separations8120233/s1, Table S1: Performance characteristics of the HPLC-MS method applicable to polyphenolic compounds evaluation in red wine samples. Figure S1: Bootstrap report and estimated sampling distribution for 100 resamples on the assessed variables.
Author Contributions
Conceptualization, L.M.P., C.C. and M.E.P.; Methodology, L.M.P. and C.A.; Formal analysis, L.M.P. and C.A.; Investigation, L.M.P. and C.A.; Resources, G.L.R.; Data curation, L.M.P.; Writing—original draft preparation, L.M.P. and C.C.; Writing—review and editing, L.M.P., C.C., C.A., G.L.R. and M.E.P.; Visualization, L.M.P.; Supervision, C.C, G.L.R. and M.E.P. All authors have read and agreed to the published version of the manuscript.
Funding
This research was supported by the Romanian National Center for Program Management-PN 19270101-Contract No. 25N/2019. The APC was funded by the authors.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The authors confirm that the data supporting the findings of this study are available within the article and its supplementary material.
Acknowledgments
This work is part of the thesis of L.M.P., supported by the Romanian Ministry of Education.
Conflicts of Interest
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
References
- Pavloušek, P.; Kumšta, M. Authentication of Riesling wines from the Czech Republic on the basis of the non-flavonoid phenolic compounds. Czech J. Food Sci. 2013, 31, 474–482. [Google Scholar] [CrossRef] [Green Version]
- Bonada, M.; Jeffery, D.; Petrie, P.; Moran, M.A.; Sadras, V. Impact of elevated temperature and water deficit on the chemical and sensory profiles of Barossa Shiraz grapes and wines. Aust. J. Grape Wine Res. 2015, 21, 240–253. [Google Scholar] [CrossRef]
- Palade, L.M.; Popa, M.E. Polyphenol Fingerprinting Approaches in Wine Traceability and Authenticity: Assessment and Implications of Red Wines. Beverages 2018, 4, 75. [Google Scholar] [CrossRef] [Green Version]
- Kamiloglu, S. Authenticity and traceability in beverages. Food Chem. 2019, 277, 12–24. [Google Scholar] [CrossRef]
- Wadood, S.A.; Boli, G.; Xiaowen, Z.; Hussain, I.; Yimin, W. Recent development in the application of analytical techniques for the traceability and authenticity of food of plant origin. Microchem. J. 2020, 152, 104295. [Google Scholar] [CrossRef]
- Tonietto, J.; Ruiz, V.S.; Zanus, M.C.; Montes, C. The Effect of Viticultural Climate on Red and White Wine Typicity. Characterization in Ibero-American grape-growing regions. Spécial Laccave J. Int. Sci. Vigne Vin 2014, 19–23. [Google Scholar]
- de Oliveira, J.B.; Egipto, R.; Laureano, O.; de Castro, R.; Pereira, G.E.; Ricardo-Da-Silva, J.M. Chemical composition and sensory profile of Syrah wines from semiarid tropical Brazil—Rootstock and harvest season effects. LWT 2019, 114, 108415. [Google Scholar] [CrossRef]
- Croitoru, C. Oenologie. Inovari si Noutati; Editura AGIR: Bucuresti, Romania, 2012; ISBN 978-973-720-463-9. [Google Scholar]
- Versari, A.; Laurie, F.; Ricci, A.; Laghi, L.; Parpinello, G.P. Progress in authentication, typification and traceability of grapes and wines by chemometric approaches. Food Res. Int. 2014, 60, 2–18. [Google Scholar] [CrossRef]
- Cravero, C.M. Wine Traceability. Beverages 2019, 5, 59. [Google Scholar] [CrossRef] [Green Version]
- Perini, M.; Strojnik, L.; Paolini, M.; Camin, F. Gas Chromatography Combustion Isotope Ratio Mass Spectrometry for Improving the Detection of Authenticity of Grape Must. J. Agric. Food Chem. 2020, 68, 3322–3329. [Google Scholar] [CrossRef] [PubMed]
- de Lima, C.M.; Fernandes, D.D.S.; Pereira, G.E.; Gomes, A.D.A.; de Araújo, M.C.U.; Diniz, P.H.G.D. Digital image-based tracing of geographic origin, winemaker, and grape type for red wine authentication. Food Chem. 2020, 312, 126060. [Google Scholar] [CrossRef]
- Sacchi, K.L.; Bisson, L.F.; Adams, D.O. A review of the effect of winemaking techniques on phenolic extraction in red wines. Am. J. Enol. Vitic. 2005, 56, 197–206. [Google Scholar]
- Di Paola-Naranjo, R.D.; Baroni, M.V.; Podio, N.S.; Rubinstein, H.R.; Fabani, M.P.; Badini, R.G.; Inga, M.; Ostera, H.A.; Cagnoni, M.; Gallegos, E.; et al. Fingerprints for Main Varieties of Argentinean Wines: Terroir Differentiation by Inorganic, Organic, and Stable Isotopic Analyses Coupled to Chemometrics. J. Agric. Food Chem. 2011, 59, 7854–7865. [Google Scholar] [CrossRef] [PubMed]
- Alecu, A.; Albu, C.; Litescu, S.C.; Eremia, S.A.V.; Radu, G.L. Phenolic and Anthocyanin Profile of Valea Calugareasca Red Wines by HPLC-PDA-MS and MALDI-TOF Analysis. Food Anal. Methods 2015, 9, 300–310. [Google Scholar] [CrossRef]
- Siracusa, L.; Ruberto, G. Plant Polyphenol Profiles as a Tool for Traceability and Valuable Support to Biodiversity. In Polyphenols in Plants: Isolation, Purification and Extract Preparation; Watson, R.R., Ed.; Academic Press: Cambridge, MA, USA, 2014; pp. 15–33. ISBN 9780123979346. [Google Scholar]
- Donno, D.; Boggia, R.; Zunin, P.; Cerutti, A.; Guido, M.; Mellano, M.G.; Prgomet, Z.; Beccaro, G.L. Phytochemical fingerprint and chemometrics for natural food preparation pattern recognition: An innovative technique in food supplement quality control. J. Food Sci. Technol. 2016, 53, 1071–1083. [Google Scholar] [CrossRef] [Green Version]
- Jackson, R. Grapevine Structure and Function. In Wine Science, 4th ed.; Academic Press: San Diego, CA, USA, 2014; pp. 69–141. ISBN 9780123814685. [Google Scholar]
- Lingua, M.S.; Fabani, M.; Wunderlin, D.A.; Baroni, M.V. From grape to wine: Changes in phenolic composition and its influence on antioxidant activity. Food Chem. 2016, 208, 228–238. [Google Scholar] [CrossRef] [PubMed]
- Teixeira, A.; Eiras-Dias, J.; Castellarin, S.D.; Gerós, H. Berry Phenolics of Grapevine under Challenging Environments. Int. J. Mol. Sci. 2013, 14, 18711–18739. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Degu, A.; Hochberg, U.; Sikron, N.; Venturini, L.; Buson, G.; Ghan, R.; Plaschkes, I.; Batushansky, A.; Chalifa-Caspi, V.; Mattivi, F.; et al. Metabolite and transcript profiling of berry skin during fruit development elucidates differential regulation between Cabernet Sauvignon and Shiraz cultivars at branching points in the polyphenol pathway. BMC Plant Biol. 2014, 14, 188. [Google Scholar] [CrossRef] [Green Version]
- Harrison, R. Practical interventions that influence the sensory attributes of red wines related to the phenolic composition of grapes: A review. Int. J. Food Sci. Technol. 2017, 53, 3–18. [Google Scholar] [CrossRef]
- Gambacorta, G.; Trani, A.; Fasciano, C.; Paradiso, V.M.; Faccia, M. Effects of prefermentative cold soak on polyphenols and volatiles of Aglianico, Primitivo and Nero di Troia red wines. Food Sci. Nutr. 2019, 7, 483–491. [Google Scholar] [CrossRef] [Green Version]
- Rousserie, P.; Rabot, A.; Geny-Denis, L. From Flavanols Biosynthesis to Wine Tannins: What Place for Grape Seeds? J. Agric. Food Chem. 2019, 67, 1325–1343. [Google Scholar] [CrossRef]
- Jackson, R. Vineyard Practice. In Wine Science, 4th ed.; Academic Press: San Diego, CA, USA, 2014; pp. 143–306. ISBN 9780123814685. [Google Scholar]
- de Freitas, V. Revisiting Wine Polyphenols Chemistry in Relation to Their Sensory Characteristics. In Recent Advances in Polyphenol Research; Halbwirth, H., Stich, K., Cheynier, V., Quideau, S., Eds.; Wiley Online Books: Hoboken, NJ, USA, 2019; pp. 263–284. ISBN 9781119427896. [Google Scholar]
- Li, S.-Y.; Duan, C.-Q. Astringency, bitterness and color changes in dry red wines before and during oak barrel aging: An updated phenolic perspective review. Crit. Rev. Food Sci. Nutr. 2019, 59, 1840–1867. [Google Scholar] [CrossRef]
- Santos, M.C.; Nunes, C.; Ferreira, A.S.; Jourdes, M.; Teissedre, P.-L.; Rodrigues, A.; Amado, O.; Saraiva, J.A.; Coimbra, M.A. Comparison of high pressure treatment with conventional red wine aging processes: Impact on phenolic composition. Food Res. Int. 2019, 116, 223–231. [Google Scholar] [CrossRef] [PubMed]
- Lerno, L.; Reichwage, M.; Ponangi, R.; Hearne, L.; Block, D.E.; Oberholster, A. Effects of Cap and Overall Fermentation Temperature on Phenolic Extraction in Cabernet Sauvignon Fermentations. Am. J. Enol. Vitic. 2015, 66, 444–453. [Google Scholar] [CrossRef]
- Setford, P.C.; Jeffery, D.W.; Grbin, P.R.; Muhlack, R.A. Factors affecting extraction and evolution of phenolic compounds during red wine maceration and the role of process modelling. Trends Food Sci. Technol. 2017, 69, 106–117. [Google Scholar] [CrossRef]
- Pérez-Navarro, J.; Romero, E.G.; Gómez-Alonso, S.; Cañas, P.M.I. Comparison between the phenolic composition of Petit Verdot wines elaborated at different maceration/fermentation temperatures. Int. J. Food Prop. 2018, 21, 996–1007. [Google Scholar] [CrossRef] [Green Version]
- Ortega-Regules, A.; Romero-Cascales, I.; López-Roca, J.M.; Ros-García, J.M.; Gómez-Plaza, E. Anthocyanin fingerprint of grapes: Environmental and genetic variations. J. Sci. Food Agric. 2006, 86, 1460–1467. [Google Scholar] [CrossRef]
- Mayr, C.M.; De Rosso, M.; Vedova, A.D.; Flamini, R. High-Resolution Mass Spectrometry Identification of Secondary Metabolites in Four Red Grape Varieties Potentially Useful as Traceability Markers of Wines. Beverages 2018, 4, 74. [Google Scholar] [CrossRef] [Green Version]
- Muccillo, L.; Gambuti, A.; Frusciante, L.; Iorizzo, M.; Moio, L.; Raieta, K.; Rinaldi, A.; Colantuoni, V.; Aversano, R. Biochemical features of native red wines and genetic diversity of the corresponding grape varieties from Campania region. Food Chem. 2014, 143, 506–513. [Google Scholar] [CrossRef] [Green Version]
- Bertacchini, L.; Cocchi, M.; Vigni, M.L.; Marchetti, A.; Salvatore, E.; Sighinolfi, S.; Silvestri, M.; Durante, C. The Impact of Chemometrics on Food Traceability. In Data Handling in Science and Technology; Elsevier: Amsterdam, The Netherlands, 2013; Volume 28, pp. 371–410. ISBN 0922-3487. [Google Scholar]
- Schlesier, K.; Fauhl-Hassek, C.; Forina, M.; Cotea, V.; Kocsi, E.; Schoula, R.; van Jaarsveld, F.; Wittkowski, R. Characterisation and determination of the geographical origin of wines. Part I: Overview. Eur. Food Res. Technol. 2009, 230, 1–13. [Google Scholar] [CrossRef]
- Römisch, U.; Jäger, H.; Capron, X.; Lanteri, S.; Forina, M.; Smeyers-Verbeke, J. Characterization and determination of the geographical origin of wines. Part III: Multivariate discrimination and classification methods. Eur. Food Res. Technol. 2009, 230, 31–45. [Google Scholar] [CrossRef]
- Merkytė, V.; Longo, E.; Windisch, G.; Boselli, E. Phenolic Compounds as Markers of Wine Quality and Authenticity. Foods 2020, 9, 1785. [Google Scholar] [CrossRef]
- Saurina, J. Characterization of wines using compositional profiles and chemometrics. TrAC Trends Anal. Chem. 2010, 29, 234–245. [Google Scholar] [CrossRef]
- Rubert, J.; Lacina, O.; Fauhl-Hassek, C.; Hajslova, J. Metabolic fingerprinting based on high-resolution tandem mass spectrometry: A reliable tool for wine authentication? Anal. Bioanal. Chem. 2014, 406, 6791–6803. [Google Scholar] [CrossRef]
- Jackson, R.S. Site Selection and Climate. In Wine Science, 4th ed.; Academic Press: San Diego, CA, USA, 2014; pp. 307–346. ISBN 9780123814685. [Google Scholar]
- Anesi, A.; Stocchero, M.; Santo, S.D.; Commisso, M.; Zenoni, S.; Ceoldo, S.; Tornielli, G.B.; Siebert, T.E.; Herderich, M.; Pezzotti, M.; et al. Towards a scientific interpretation of the terroir concept: Plasticity of the grape berry metabolome. BMC Plant Biol. 2015, 15, 191. [Google Scholar] [CrossRef] [Green Version]
- Villano, C.; Lisanti, M.T.; Gambuti, A.; Vecchio, R.; Moio, L.; Frusciante, L.; Aversano, R.; Carputo, D. Wine varietal authentication based on phenolics, volatiles and DNA markers: State of the art, perspectives and drawbacks. Food Control. 2017, 80, 1–10. [Google Scholar] [CrossRef]
- González-Neves, G.; Favre, G.; Gil, G.; Ferrer, M.; Charamelo, D. Effect of cold pre-fermentative maceration on the color and composition of young red wines cv. Tannat. J. Food Sci. Technol. 2014, 52, 3449–3457. [Google Scholar] [CrossRef] [Green Version]
- Vilanova, M.; Rodríguez, I.; Canosa, P.; Otero, I.; Gamero, E.; Moreno, D.; Talaverano, I.; Valdés, E. Variability in chemical composition of Vitis vinifera cv Mencía from different geographic areas and vintages in Ribeira Sacra (NW Spain). Food Chem. 2015, 169, 187–196. [Google Scholar] [CrossRef]
- Makris, D.P.; Kallithraka, S.; Mamalos, A. Differentiation of young red wines based on cultivar and geographical origin with application of chemometrics of principal polyphenolic constituents. Talanta 2006, 70, 1143–1152. [Google Scholar] [CrossRef] [PubMed]
- Radovanović, B.C.; Radovanović, A.N.; Souquet, J.-M. Phenolic profile and free radical-scavenging activity of Cabernet Sauvignon wines of different geographical origins from the Balkan region. J. Sci. Food Agric. 2010, 90, 2455–2461. [Google Scholar] [CrossRef] [PubMed]
- Kallithraka, S.; Tsoutsouras, E.; Tzourou, E.; Lanaridis, P. Principal phenolic compounds in Greek red wines. Food Chem. 2006, 99, 784–793. [Google Scholar] [CrossRef]
- Jaitz, L.; Siegl, K.; Eder, R.; Rak, G.; Abranko, L.; Koellensperger, G.; Hann, S. LC–MS/MS analysis of phenols for classification of red wine according to geographic origin, grape variety and vintage. Food Chem. 2010, 122, 366–372. [Google Scholar] [CrossRef]
- Salvatore, E.; Cocchi, M.; Marchetti, A.; Marini, F.; de Juan, A. Determination of phenolic compounds and authentication of PDO Lambrusco wines by HPLC-DAD and chemometric techniques. Anal. Chim. Acta 2013, 761, 34–45. [Google Scholar] [CrossRef]
- Wu, Q.; Gu, H.-W.; Yin, X.-L.; Zhou, H.-H.; Chang, H.-Y.; Shi, J.; Chen, Y.; Yan, X.-F.; Liu, Z. Development of an HPLC-DAD Method Combined with Chemometrics for Differentiating Geographical Origins of Chinese Red Wines on the Basis of Phenolic Compounds. Food Anal. Methods 2021, 14, 1895–1907. [Google Scholar] [CrossRef]
- Pisano, P.L.; Silva, M.F.; Olivieri, A.C. Anthocyanins as markers for the classification of Argentinean wines according to botanical and geographical origin. Chemometric modeling of liquid chromatography–mass spectrometry data. Food Chem. 2015, 175, 174–180. [Google Scholar] [CrossRef] [PubMed]
- Cozzolino, D. Metabolomics in Grape and Wine: Definition, Current Status and Future Prospects. Food Anal. Methods 2016, 9, 2986–2997. [Google Scholar] [CrossRef]
- Aleixandre-Tudo, J.L.; Nieuwoudt, H.; Olivieri, A.; Aleixandre, J.L.; du Toit, W. Phenolic profiling of grapes, fermenting samples and wines using UV-Visible spectroscopy with chemometrics. Food Control 2018, 85, 11–22. [Google Scholar] [CrossRef]
- Albu, C.; Eremia, S.A.V.; Penu, R.; Vasilescu, I.; Litescu, S.C.; Radu, G.L. Characterization of the Phenolics and Free Radical Scavenging of Romanian Red Wine. Anal. Lett. 2017, 50, 591–606. [Google Scholar] [CrossRef]
- Favre, G.; Hermosín-Gutiérrez, I.; Piccardo, D.; Gómez-Alonso, S.; González-Neves, G. Selectivity of pigments extraction from grapes and their partial retention in the pomace during red-winemaking. Food Chem. 2019, 277, 391–397. [Google Scholar] [CrossRef]
- Piccardo, D.; González-Neves, G.; Favre, G.; Pascual, O.; Canals, J.M.; Zamora, F. Impact of Must Replacement and Hot Pre-Fermentative Maceration on the Color of Uruguayan Tannat Red Wines. Fermentation 2019, 5, 80. [Google Scholar] [CrossRef] [Green Version]
- Arcena, M.R.; Kebede, B.; Leong, S.Y.; Silcock, P.; Oey, I. Feasibility of using integrated fingerprinting, profiling and chemometrics approach to understand (bio) chemical changes throughout commercial red winemaking: A case study on Merlot. Food Res. Int. 2020, 127, 108767. [Google Scholar] [CrossRef]
- Rentzsch, M.; Wilkens, A.; Winterhalter, P. Non-flavonoid Phenolic Compounds. In Wine Chemistry and Biochemistry; Moreno-Arribas, M.V., Polo, M.C., Eds.; Springer: New York, NY, USA, 2008; pp. 509–527. ISBN 978-0-387-74118-5. [Google Scholar]
- García-Beneytez, E.; Revilla, E.; Cabello, F. Anthocyanin pattern of several red grape cultivars and wines made from them. Eur. Food Res. Technol. 2002, 215, 32–37. [Google Scholar] [CrossRef]
- Mattivi, F.; Guzzon, R.; Vrhovsek, U.; Stefanini, A.M.; Velasco, R. Metabolite Profiling of Grape: Flavonols and Anthocyanins. J. Agric. Food Chem. 2006, 54, 7692–7702. [Google Scholar] [CrossRef] [PubMed]
- Costa, E.; Cosme, F.; Jordão, A.M.; Mendes-Faia, A. Anthocyanin profile and antioxidant activity from 24 grape varieties cultivated in two Portuguese wine regions. OENO One 2014, 48, 51–62. [Google Scholar] [CrossRef]
- Kyraleou, M.; Kallithraka, S.; Gkanidi, E.; Koundouras, S.; Mannion, D.T.; Kilcawley, K.N. Discrimination of five Greek red grape varieties according to the anthocyanin and proanthocyanidin profiles of their skins and seeds. J. Food Compos. Anal. 2020, 92, 103547. [Google Scholar] [CrossRef]
- Jackson, R. Post-Fermentation Treatments and Related Topics. In Wine Science, 4th ed.; Academic Press: San Diego, CA, USA, 2014; pp. 535–676. ISBN 9780123814685. [Google Scholar]
- Zhang, B.; He, F.; Zhou, P.-P.; Liu, Y.; Duan, C.-Q. Copigmentation between malvidin-3-O-glucoside and hydroxycinnamic acids in red wine model solutions: Investigations with experimental and theoretical methods. Food Res. Int. 2015, 78, 313–320. [Google Scholar] [CrossRef]
- He, F.; Liang, N.-N.; Mu, L.; Pan, Q.-H.; Wang, J.; Reeves, M.J.; Duan, C.-Q. Anthocyanins and Their Variation in Red Wines I. Monomeric Anthocyanins and Their Color Expression. Molecules 2012, 17, 1571–1601. [Google Scholar] [CrossRef] [Green Version]
- Garrido, J.; Borges, F. Wine and grape polyphenols—A chemical perspective. Food Res. Int. 2013, 54, 1844–1858. [Google Scholar] [CrossRef] [Green Version]
- Blanco-Vega, D.; Gómez-Alonso, S.; Hermosín-Gutiérrez, I. Identification, content and distribution of anthocyanins and low molecular weight anthocyanin-derived pigments in Spanish commercial red wines. Food Chem. 2014, 158, 449–458. [Google Scholar] [CrossRef]
- Marquez, A.; Serratosa, M.P.; Merida, J. Influence of bottle storage time on colour, phenolic composition and sensory properties of sweet red wines. Food Chem. 2014, 146, 507–514. [Google Scholar] [CrossRef] [PubMed]
- Kumšta, M.; Pavloušek, P.; Kárník, P. Use of Anthocyanin Profiles When Differentiating Individual Varietal Wines and Terroirs. Food Technol. Biotechnol. 2014, 52, 383–390. [Google Scholar] [CrossRef]
- Malacarne, M.; Nardin, T.; Bertoldi, D.; Nicolini, G.; Larcher, R. Verifying the botanical authenticity of commercial tannins through sugars and simple phenols profiles. Food Chem. 2016, 206, 274–283. [Google Scholar] [CrossRef]
- Figueiredo-González, M.; Martínez-Carballo, E.; Cancho-Grande, B.; Santiago, J.; Martínez, M.; Simal-Gándara, J. Pattern recognition of three Vitis vinifera L. red grapes varieties based on anthocyanin and flavonol profiles, with correlations between their biosynthesis pathways. Food Chem. 2012, 130, 9–19. [Google Scholar] [CrossRef]
- Díaz, R.; Gallart-Ayala, H.; Sancho, J.V.; Nuñez, O.; Zamora, T.; Martins, C.P.; Hernandez, F.; Hernández-Cassou, S.; Saurina, J.; Checa, A. Told through the wine: A liquid chromatography–mass spectrometry interplatform comparison reveals the influence of the global approach on the final annotated metabolites in non-targeted metabolomics. J. Chromatogr. A 2016, 1433, 90–97. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Flamini, R.; Mattivi, F.; De Rosso, M.; Arapitsas, P.; Bavaresco, L. Advanced Knowledge of Three Important Classes of Grape Phenolics: Anthocyanins, Stilbenes and Flavonols. Int. J. Mol. Sci. 2013, 14, 19651–19669. [Google Scholar] [CrossRef] [PubMed]
- Terrier, N.; Poncet-Legrand, C.; Cheynier, V. Flavanols, Flavonols and Dihydroflavonols. In Wine Chemistry and Biochemistry; Moreno-Arribas, M.V., Polo, M.C., Eds.; Springer: New York, NY, USA, 2009; pp. 463–507. ISBN 9780387741161. [Google Scholar]
- Moreno, J.; Peinado, R. Polyphenols. In Enological Chemistry; Elsevier: Amsterdam, The Netherlands, 2012; pp. 53–76. ISBN 9780123884381. [Google Scholar]
- Castellarin, S.; Bavaresco, L.; Falginella, L.; Goncalves, M.; Di Gaspero, G. Phenolics in Grape Berry and Key Antioxidants. In The Biochemistry of the Grape Berry; Geros, H., Chaves, M.M., Delrot, S., Eds.; Bentham Science Publishers: Sharja, United Arab Emirates, 2012; pp. 89–110. [Google Scholar]
- Geana, E.-I.; Popescu, R.; Costinel, D.; (Dinca), O.R.B.; Stefanescu, I.; Ionete, R.E.; Bala, C. Verifying the red wines adulteration through isotopic and chromatographic investigations coupled with multivariate statistic interpretation of the data. Food Control 2016, 62, 1–9. [Google Scholar] [CrossRef]
- Fabani, M.; Ravera, M.J.; Wunderlin, D.A. Markers of typical red wine varieties from the Valley of Tulum (San Juan-Argentina) based on VOCs profile and chemometrics. Food Chem. 2013, 141, 1055–1062. [Google Scholar] [CrossRef]
- Ferreira, V.; Fernandes, F.; Pinto-Carnide, O.; Valentão, P.; Falco, V.; Martín, J.P.; Ortiz, J.M.; Arroyo-Garcia, R.; Andrade, P.B.; Castro, I. Identification of Vitis vinifera L. grape berry skin color mutants and polyphenolic profile. Food Chem. 2016, 194, 117–127. [Google Scholar] [CrossRef]
- Avizcuri, J.-M.; Saenz-Navajas, M.-P.; Echavarri, J.F.; Ferreira, V.; Fernández-Zurbano, P. Evaluation of the impact of initial red wine composition on changes in color and anthocyanin content during bottle storage. Food Chem. 2016, 213, 123–134. [Google Scholar] [CrossRef] [Green Version]
- Carrascón, V.; Vallverdu-Queralt, A.; Meudec, E.; Sommerer, N.; Fernandez-Zurbano, P.; Ferreira, V. The kinetics of oxygen and SO2 consumption by red wines. What do they tell about oxidation mechanisms and about changes in wine composition? Food Chem. 2018, 241, 206–214. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Puértolas, E.; Saldaña, G.; Condón, S.; Alvarez, I.; Raso, J. Evolution of polyphenolic compounds in red wine from Cabernet Sauvignon grapes processed by pulsed electric fields during aging in bottle. Food Chem. 2010, 119, 1063–1070. [Google Scholar] [CrossRef]
- Quaglieri, C.; Prieto-Perea, N.; Berrueta, L.A.; Gallo, B.; Rasines-Perea, Z.; Jourdes, M.; Teissedre, P.-L. Comparison of Aquitaine and Rioja Red Wines: Characterization of Their Phenolic Composition and Evolution from 2000 to 2013. Molecules 2017, 22, 192. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bellomarino, S.; Conlan, X.; Parker, R.; Barnett, N.; Adams, M. Geographical classification of some Australian wines by discriminant analysis using HPLC with UV and chemiluminescence detection. Talanta 2009, 80, 833–838. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Li, L.; Ma, T.; Liu, X.; Huang, W.; Zhan, J. Profiles of Phenolic Acids and Flavan-3-ols for Select Chinese Red Wines: A Comparison and Differentiation According to Geographic Origin and Grape Variety. J. Food Sci. 2015, 80, C2170–C2179. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).