Analysis and Anticancer Effects of Active Compounds from Spatholobi Caulis in Human Breast Cancer Cells
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material
2.2. Chemicals and Reagents
2.3. Making 70% Ethyl Alcohol Extract of Spatholobi Caulis Spatholobi Caulis
2.4. High-Performance Liquid Chromatography Analysis of the Nine Marker Compounds in Spatholobi Caulis
2.5. Cell Culture and Drug Preparation
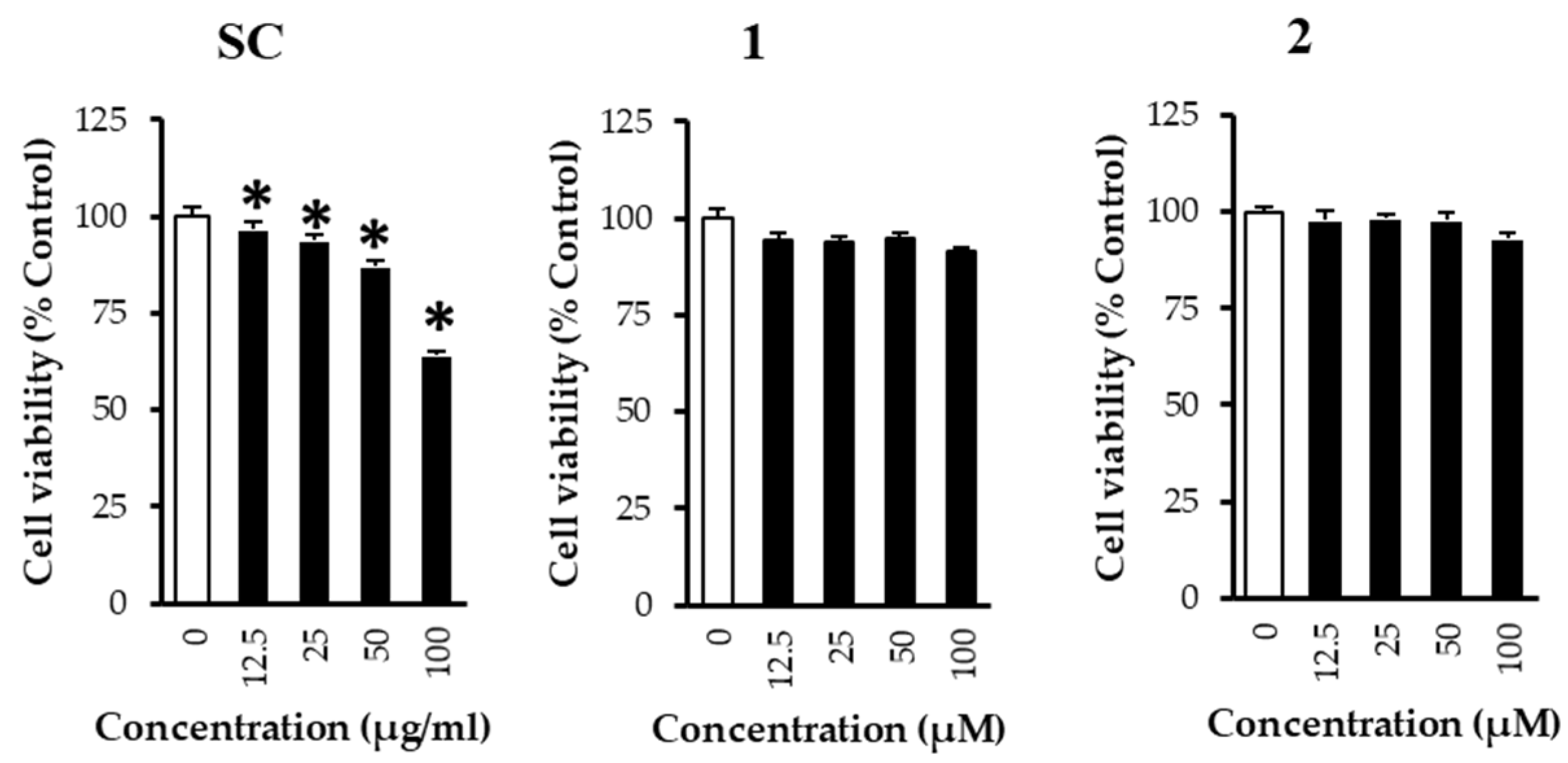
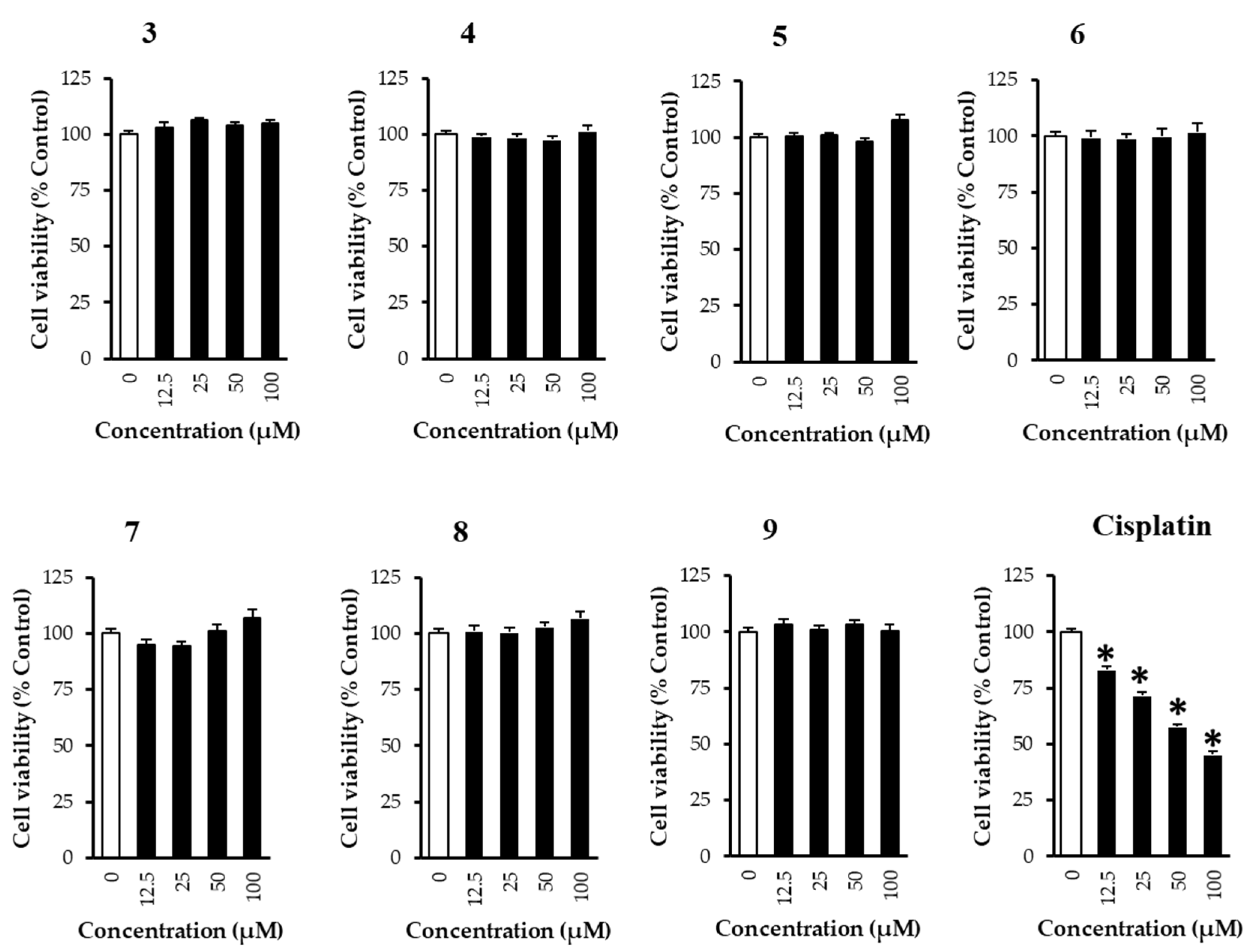
2.6. Cell Viability Assay
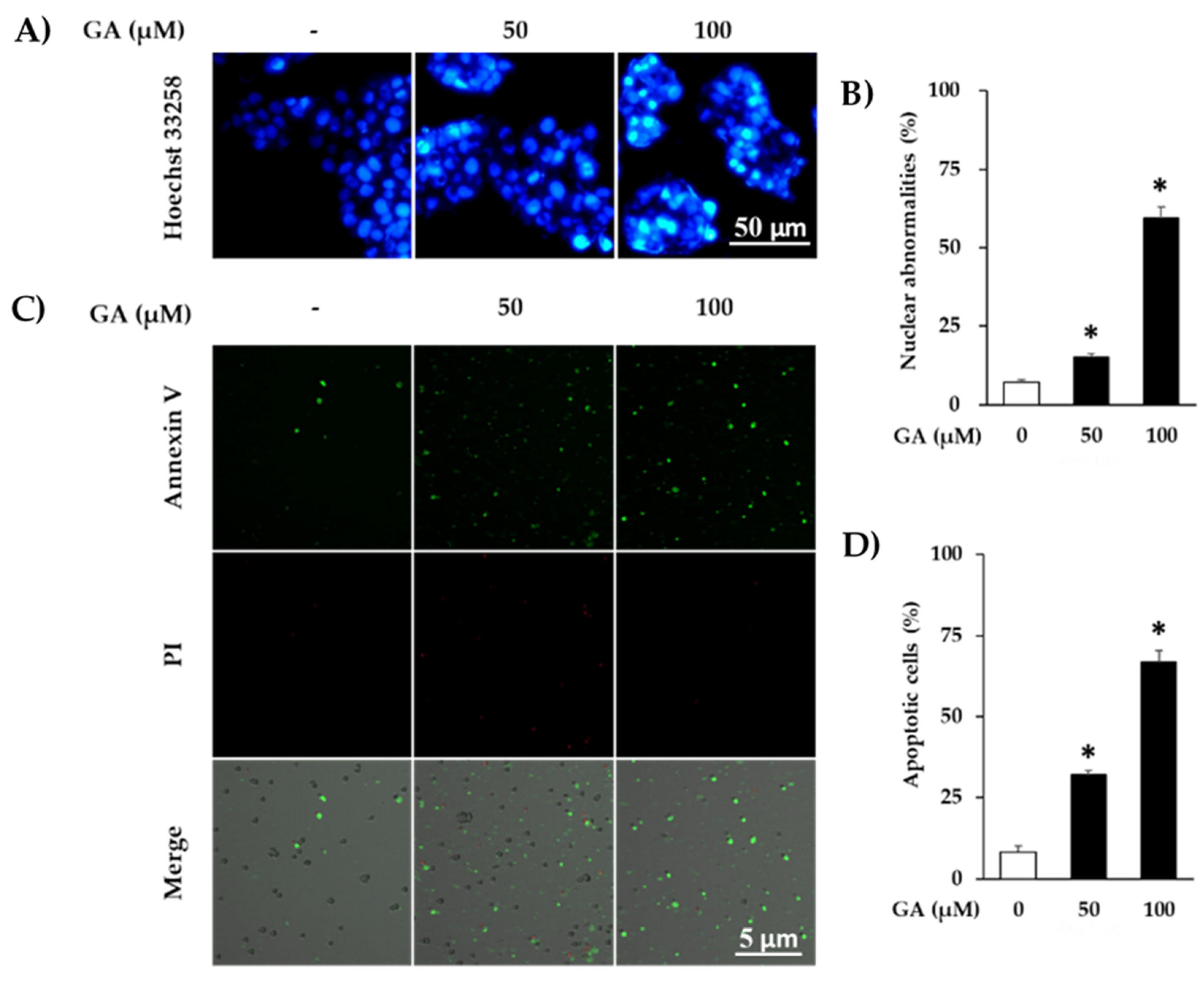
2.7. Nuclear Staining
2.8. Tali Assay
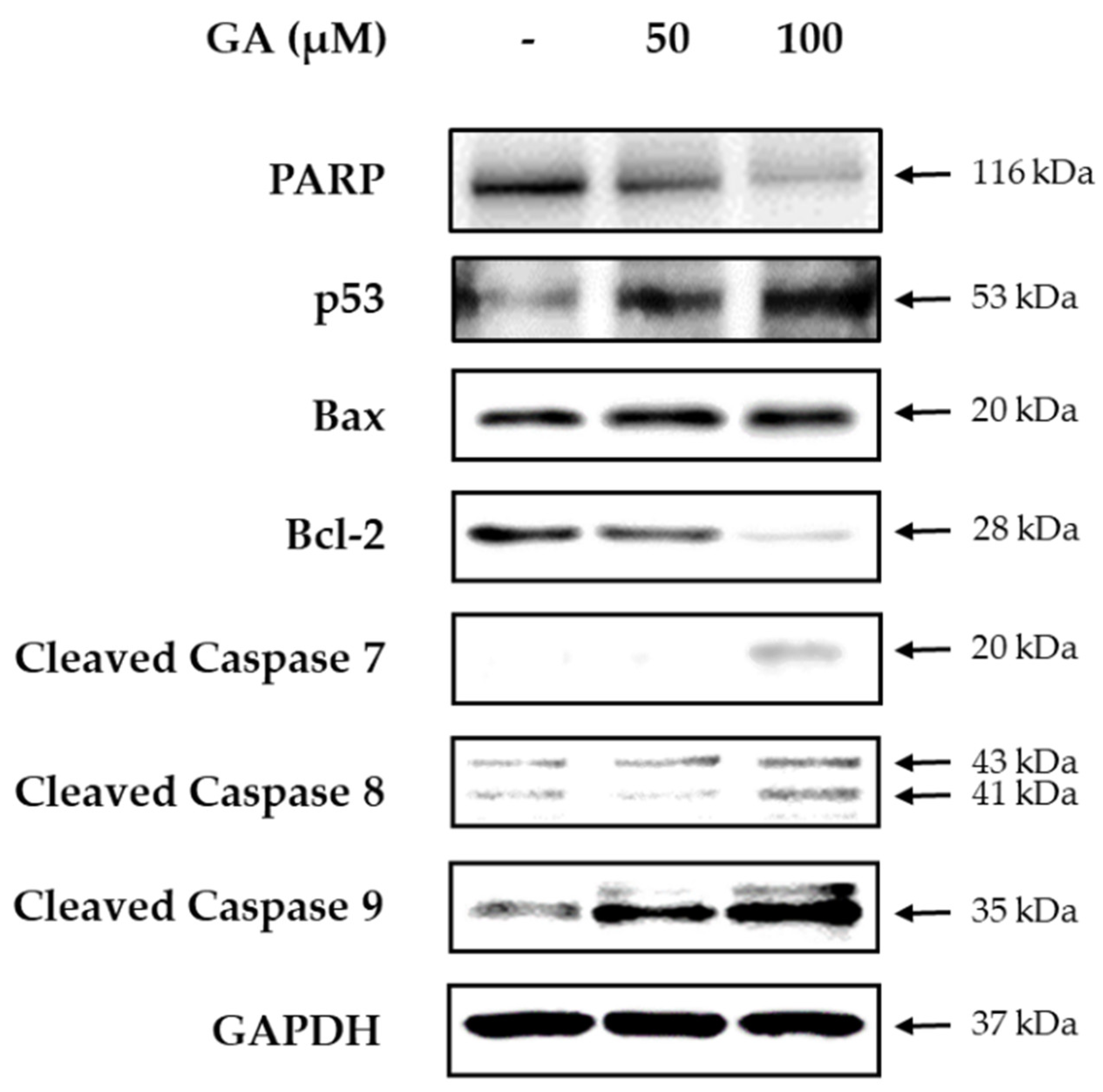
2.9. Western Blotting
2.10. Network Pharmacological Analyses
2.11. Statistical Method
3. Results and Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2019. CA Cancer J.Clin. 2019, 69, 7–34. [Google Scholar] [CrossRef] [PubMed]
- Wörmann, B. Breast cancer: Basics, screening, diagnostics and treatment. Med. Mon. Pharm. 2017, 40, 55–64. [Google Scholar]
- Chang-Claude, J.; Popanda, O.; Tan, X.-L.; Kropp, S.; Helmbold, I.; von Fournier, D.; Haase, W.; Sautter-Bihl, M.L.; Wenz, F.; Schmezer, P. Association between polymorphisms in the DNA repair genes, XRCC1, APE1, and XPD and acute side effects of radiotherapy in breast cancer patients. Clin. Cancer Res. 2005, 11, 4802–4809. [Google Scholar] [CrossRef]
- Partridge, A.H.; Burstein, H.J.; Winer, E.P. Side effects of chemotherapy and combined chemohormonal therapy in women with early-stage breast cancer. JNCI Monogr. 2001, 2001, 135–142. [Google Scholar] [CrossRef] [PubMed]
- Khan, T.; Ali, M.; Khan, A.; Nisar, P.; Jan, S.A.; Afridi, S.; Shinwari, Z.K. Anticancer plants: A Review of the active phytochemicals, applications in animal models, and regulatory aspects. Biomolecules 2020, 10, 47. [Google Scholar] [CrossRef]
- Majolo, F.; Delwing, L.K.D.O.B.; Marmitt, D.J.; Bustamante-Filho, I.C.; Goettert, M.I. Medicinal plants and bioactive natural compounds for cancer treatment: Important advances for drug discovery. Phytochem. Lett. 2019, 31, 196–207. [Google Scholar] [CrossRef]
- Olaku, O.; White, J.D. Herbal therapy use by cancer patients: A literature review on case reports. Eur. J. Cancer 2011, 47, 508–514. [Google Scholar] [CrossRef]
- Kamei, T.; Kumano, H.; Iwata, K.; Nariai, Y.; Matsumoto, T. The effect of a traditional Chinese prescription for a case of lung carcinoma. J. Altern. Complement. Med. 2000, 6, 557–559. [Google Scholar] [CrossRef]
- Sun, L.-R.; Zhou, W.; Zhang, H.-M.; Guo, Q.-S.; Yang, W.; Li, B.-J.; Sun, Z.-H.; Gao, S.-H.; Cui, R.-J. Modulation of multiple signaling pathways of the plant-derived natural products in cancer. Front. Oncol. 2019, 9, 9. [Google Scholar] [CrossRef]
- Millimouno, F.M.; Dong, J.; Yang, L.; Li, J.; Li, X. Targeting apoptosis pathways in cancer and perspectives with natural compounds from mother nature. Cancer Prev. Res. 2014, 7, 1081–1107. [Google Scholar] [CrossRef]
- Im, N.-K.; Lee, S.-G.; Lee, D.-S.; Park, P.-H.; Lee, I.-S.; Jeong, G.-S. Spatholobus suberectus inhibits osteoclastogenesis and stimulates chondrogenesis. Am. J. Chin. Med. 2014, 42, 1123–1138. [Google Scholar] [CrossRef] [PubMed]
- Li, R.W.; Lin, G.D.; Myers, S.P.; Leach, D.N. Anti-inflammatory activity of Chinese medicinal vine plants. J. Ethnopharmacol. 2003, 85, 61–67. [Google Scholar] [CrossRef]
- Pang, J.; Guo, J.-P.; Jin, M.; Chen, Z.-Q.; Wang, X.-W.; Li, J.-W. Antiviral effects of aqueous extract from Spatholobus suberectus Dunn. against coxsackievirus B3 in mice. Chin. J. Integr. Med. 2011, 17, 764–769. [Google Scholar] [CrossRef] [PubMed]
- Park, H.R.; Lee, H.; Lee, J.-J.; Yim, N.-H.; Gu, M.-J.; Ma, J.Y. Protective effects of spatholobi caulis extract on neuronal damage and focal ischemic stroke/reperfusion injury. Mol. Neurobiol. 2018, 55, 4650–4666. [Google Scholar] [CrossRef] [PubMed]
- Su, E.; Fang, Y.; Chen, H. Clinical observation of treating 62 patients with severe aplastic anemia failing in immunosuppressive therapy by integrative medicine. Chin. J. Integr. Tradit. West. Med. 2012, 32, 1616–1620. [Google Scholar]
- Ha, E.-S.; Lee, E.-O.; Yoon, T.-J.; Kim, J.-H.; Park, J.-O.; Lim, N.-C.; Jung, S.-K.; Yoon, B.-S.; Kim, S.-H. Methylene chloride fraction of Spatholobi Caulis induces apoptosis via caspase dependent pathway in U937 cells. Biol. Pharm. Bull. 2004, 27, 1348–1352. [Google Scholar] [CrossRef]
- Liu, B.; Liu, J.; Chen, J.; Zhu, D.; Zhou, H.; Wang, X. Study on anticancer activity of caulis spatholobi extract on human osteosarcoma saos-2 cells. Afr. J. Tradit. Complement. Altern. Med. 2013, 10, 256–260. [Google Scholar] [CrossRef]
- Sun, L.; Li, Q.; Guo, Y.; Yang, Q.; Yin, J.; Ran, Q.; Liu, L.; Zhao, Z.; Wang, Y.; Li, Y. Extract of Caulis Spatholobi, a novel platelet inhibitor, efficiently suppresses metastasis of colorectal cancer by targeting tumor cell-induced platelet aggregation. Biomed. Pharmacother. 2020, 123, 109718. [Google Scholar] [CrossRef]
- Wang, N.; Wang, J.; Meng, X.; Bao, Y.; Wang, S.; Li, T. 3D microfluidic in vitro model and bioinformatics integration to study the effects of Spatholobi Caulis tannin in cervical cancer. Sci. Rep. 2018, 8, 1–11. [Google Scholar] [CrossRef]
- Wang, N.; Wang, J.; Meng, X.; Li, T.; Wang, S.; Bao, Y. The pharmacological effects of spatholobi caulis tannin in cervical cancer and its precise therapeutic effect on related circRNA. Mol. Ther. Oncolytics 2019, 14, 121–129. [Google Scholar] [CrossRef]
- Huang, Y.; Chen, L.; Feng, L.; Guo, F.; Li, Y. Characterization of total phenolic constituents from the stems of Spatholobus suberectus using LC-DAD-MSn and their inhibitory effect on human neutrophil elastase activity. Molecules 2013, 18, 7549–7556. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Guo, L.; Duan, L.; Dong, X.; Zhou, P.; Liu, E.-H.; Li, P. Simultaneous determination of 16 phenolic constituents in Spatholobi Caulis by high performance liquid chromatography/electrospray ionization triple quadrupole mass spectrometry. J. Pharm. Biomed. Anal. 2015, 102, 110–118. [Google Scholar] [CrossRef] [PubMed]
- Lu, D.; He, H.; Wu, B.; Yao, S. Cytotoxic Effect on Cancer Cells and Structural Identification of Phenols from Spatholobi Caulis by HPLC-ESI-MSn. Nat. Prod. Commun. 2009, 4. [Google Scholar] [CrossRef]
- Baek, S.J.; Kim, J.-S.; Jackson, F.R.; Eling, T.E.; McEntee, M.F.; Lee, S.-H. Epicatechin gallate-induced expression of NAG-1 is associated with growth inhibition and apoptosis in colon cancer cells. Carcinogenesis 2004, 25, 2425–2432. [Google Scholar] [CrossRef]
- Lim, Y.C.; Lee, S.-H.; Song, M.H.; Yamaguchi, K.; Yoon, J.-H.; Choi, E.C.; Baek, S.J. Growth inhibition and apoptosis by (−)-epicatechin gallate are mediated by cyclin D1 suppression in head and neck squamous carcinoma cells. Eur. J. Cancer 2006, 42, 3260–3266. [Google Scholar] [CrossRef]
- Chen, H.-M.; Wu, Y.-C.; Chia, Y.-C.; Chang, F.-R.; Hsu, H.-K.; Hsieh, Y.-C.; Chen, C.-C.; Yuan, S.-S. Gallic acid, a major component of Toona sinensis leaf extracts, contains a ROS-mediated anti-cancer activity in human prostate cancer cells. Cancer Lett. 2009, 286, 161–171. [Google Scholar] [CrossRef] [PubMed]
- Ohno, Y.; Fukuda, K.; Takemura, G.; Toyota, M.; Watanabe, M.; Yasuda, N.; Xinbin, Q.; Maruyama, R.; Akao, S.; Gotou, K. Induction of apoptosis by gallic acid in lung cancer cells. Anticancer Drugs 1999, 10, 845–851. [Google Scholar] [CrossRef]
- Xia, T.; Zhang, J.; Zhou, C.; Li, Y.; Duan, W.; Zhang, B.; Wang, M.; Fang, J. 20 (S)-Ginsenoside Rh2 displays efficacy against T-cell acute lymphoblastic leukemia through the PI3K/Akt/mTOR signal pathway. J. Ginseng Res. 2019. [Google Scholar] [CrossRef]
- Jang, M.G.; Ko, H.C.; Kim, S.J. Effect of sasa quelpaertensis Nakai extracts and its constituent p-coumaric acid on the apoptosis of human cancer cell lines. Nat. Prod. Sci. 2018, 24, 293–297. [Google Scholar] [CrossRef]
- Kuhn, M.; Szklarczyk, D.; Pletscher-Frankild, S.; Blicher, T.H.; Von Mering, C.; Jensen, L.J.; Bork, P. STITCH 4: Integration of protein–chemical interactions with user data. Nucleic Acids Res. 2014, 42, D401–D407. [Google Scholar] [CrossRef]
- Kuleshov, M.V.; Jones, M.R.; Rouillard, A.D.; Fernandez, N.F.; Duan, Q.; Wang, Z.; Koplev, S.; Jenkins, S.L.; Jagodnik, K.M.; Lachmann, A. Enrichr: A comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 2016, 44, W90–W97. [Google Scholar] [CrossRef] [PubMed]
- Sledge, G.W., Jr.; Loehrer, P.J., Sr.; Roth, B.J.; Einhorn, L.H. Cisplatin as first-line therapy for metastatic breast cancer. J. Clin. Oncol. 1988, 6, 1811–1814. [Google Scholar] [CrossRef] [PubMed]
- Fujiki, H.; Sueoka, E.; Watanabe, T.; Suganuma, M. Synergistic enhancement of anticancer effects on numerous human cancer cell lines treated with the combination of EGCG, other green tea catechins, and anticancer compounds. J. Cancer Res. Clin. Oncol. 2015, 141, 1511–1522. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Zheng, C.; Han, L.; Xie, B.; Jia, J.; Cao, Z.; Li, Y.; Chen, Y. Synergistic therapeutic actions of herbal ingredients and their mechanisms from molecular interaction and network perspectives. Drug Discov. Today 2009, 14, 579–588. [Google Scholar] [CrossRef] [PubMed]
- Yu, R.-J.; Liu, H.-B.; Yu, Y.; Liang, L.; Xu, R.; Liang, C.; Tang, J.-S.; Yao, X.-S. Anticancer activities of proanthocyanidins from the plant Urceola huaitingii and their synergistic effects in combination with chemotherapeutics. Fitoterapia 2016, 112, 175–182. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D.; Weinberg, R.A. The hallmarks of cancer. Cell 2000, 100, 57–70. [Google Scholar] [CrossRef]
- Pfeffer, C.M.; Singh, A.T. Apoptosis: A target for anticancer therapy. Int. J. Mol. Sci. 2018, 19, 448. [Google Scholar]
- Elmore, S. Apoptosis: A review of programmed cell death. Toxicol. Pathol. 2007, 35, 495–516. [Google Scholar] [CrossRef]
- Murphy, K.; Ranganathan, V.; Farnsworth, M.; Kavallaris, M.; Lock, R.B. Bcl-2 inhibits Bax translocation from cytosol to mitochondria during drug-induced apoptosis of human tumor cells. Cell Death Differ. 2000, 7, 102–111. [Google Scholar] [CrossRef]
- Faried, A.; Kurnia, D.; Faried, L.; Usman, N.; Miyazaki, T.; Kato, H.; Kuwano, H. Anticancer effects of gallic acid isolated from Indonesian herbal medicine, Phaleria macrocarpa (Scheff.) Boerl, on human cancer cell lines. Int. J. Oncol. 2007, 30, 605–613. [Google Scholar] [CrossRef]
- Lo, C.; Lai, T.-Y.; Yang, J.-H.; Yang, J.-S.; Ma, Y.-S.; Weng, S.-W.; Chen, Y.-Y.; Lin, J.-G.; Chung, J.-G. Gallic acid induces apoptosis in A375. S2 human melanoma cells through caspase-dependent and-independent pathways. Int. J. Oncol. 2010, 37, 377–385. [Google Scholar] [PubMed]
- Sun, G.; Zhang, S.; Xie, Y.; Zhang, Z.; Zhao, W. Gallic acid as a selective anticancer agent that induces apoptosis in SMMC-7721 human hepatocellular carcinoma cells. Oncol. Lett. 2016, 11, 150–158. [Google Scholar] [CrossRef] [PubMed]
- Tsai, C.-L.; Chiu, Y.-M.; Ho, T.-Y.; Hsieh, C.-T.; Shieh, D.-C.; Lee, Y.-J.; Tsay, G.J.; Wu, Y.-Y. Gallic acid induces apoptosis in human gastric adenocarcinoma cells. Anticancer Res. 2018, 38, 2057–2067. [Google Scholar] [PubMed]
- Jänicke, R.U. MCF-7 breast carcinoma cells do not express caspase-3. Breast Cancer Res. Treat. 2009, 117, 219–221. [Google Scholar] [CrossRef]
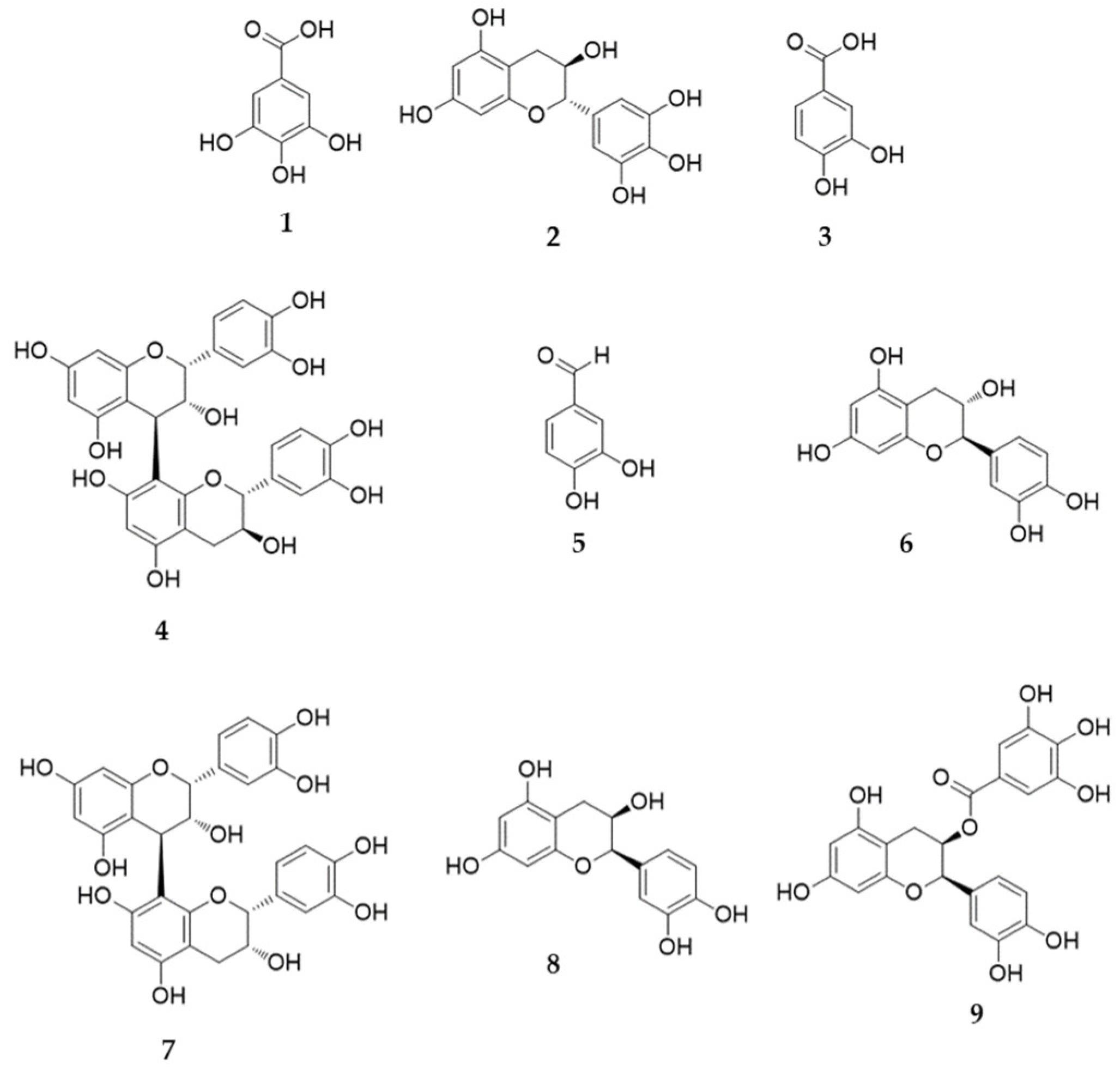
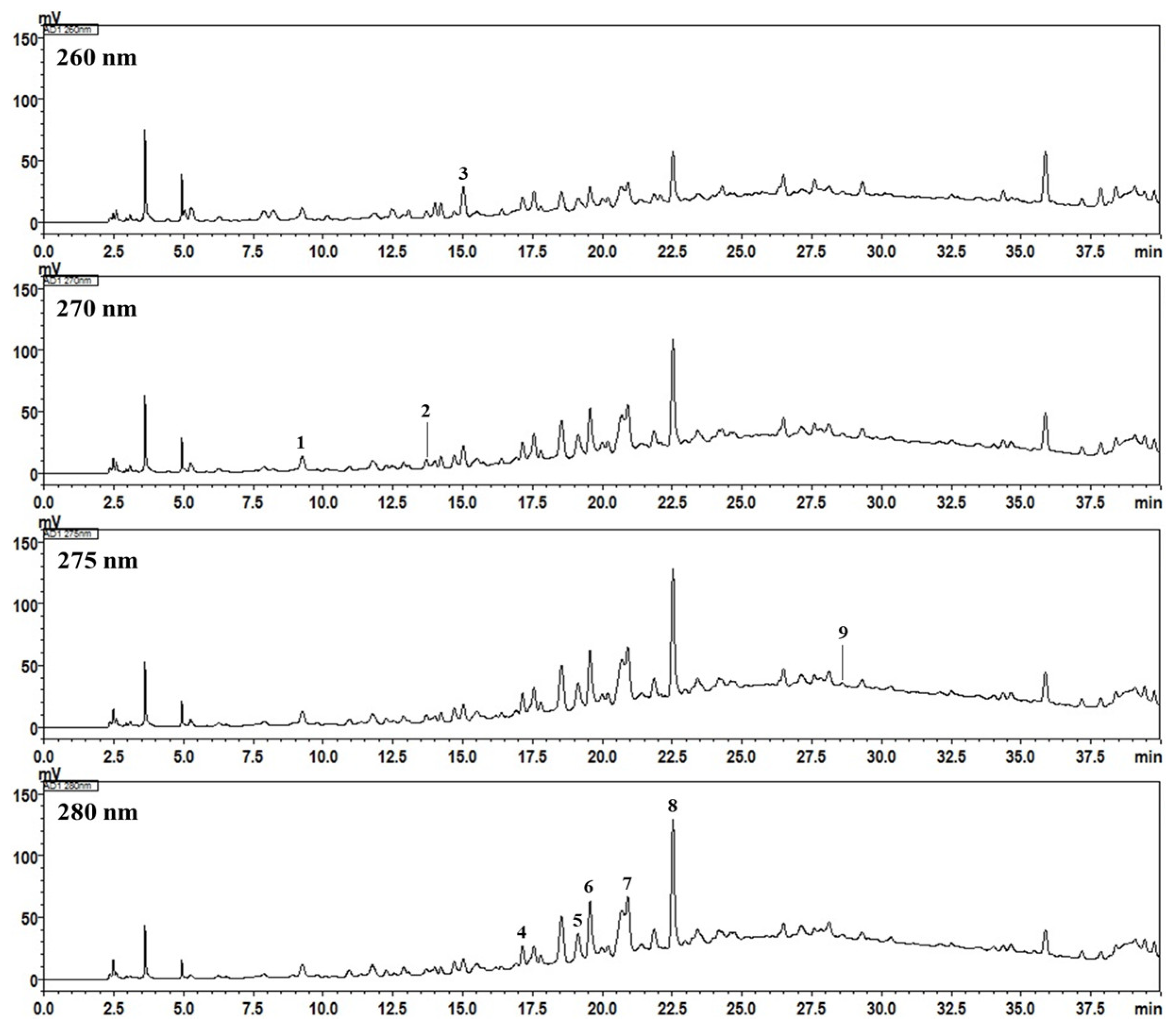
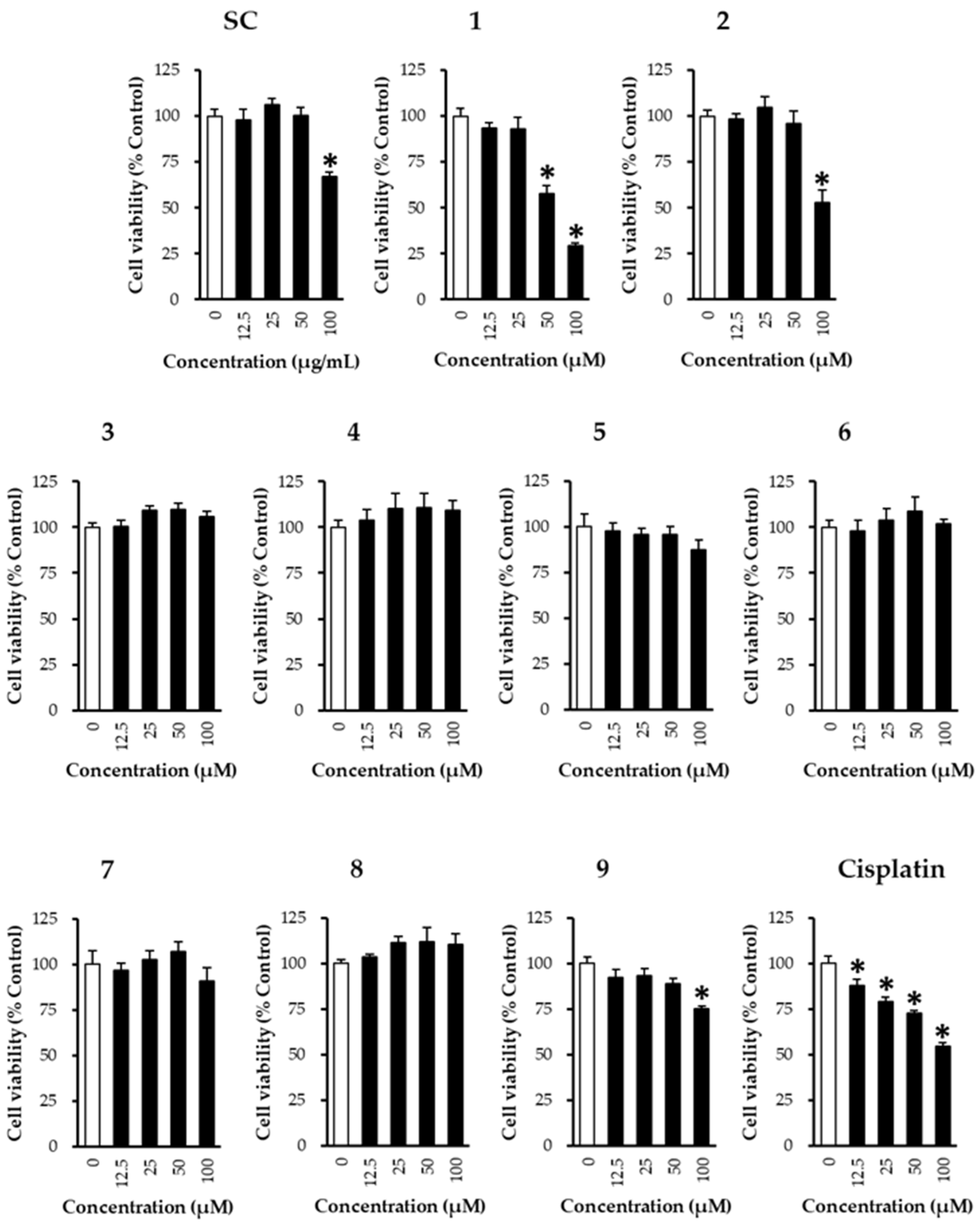
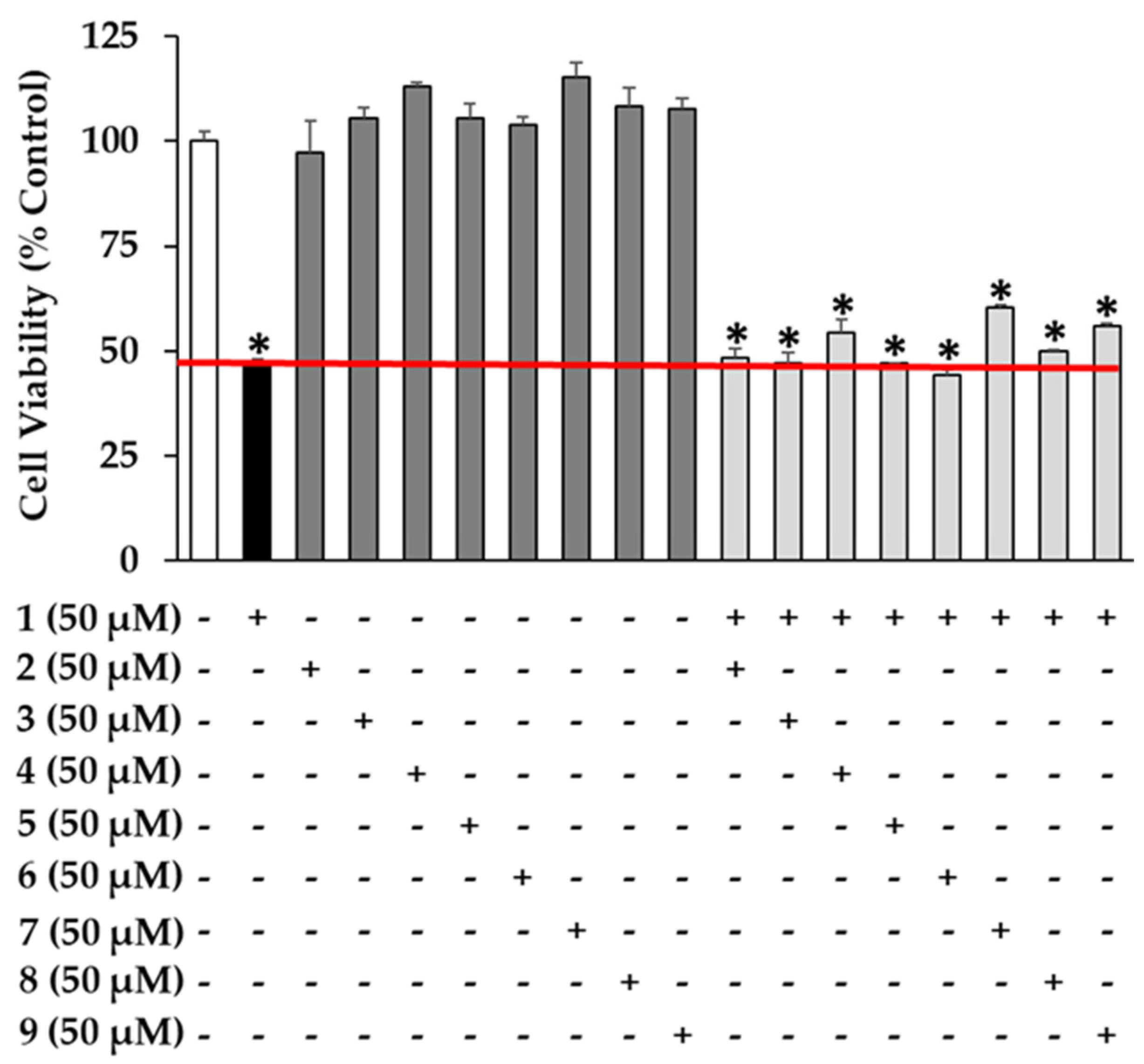
| No | Compound | Linear Range (μg/mL) | Regression Equation (y = ax + b) a | r2 | LOD (μg/mL) b | LOQ (μg/mL) c |
|---|---|---|---|---|---|---|
| 1 | Gallic acid | 0.16–10.00 | y = 51,544.39x − 3596.01 | 0.9996 | 0.03 | 0.08 |
| 2 | (−)-Gallocatechin | 0.78–50.00 | y = 3788.85x − 539.68 | 1.0000 | 0.23 | 0.71 |
| 3 | 3,4-Dihydroxybenzoic acid | 0.16–10.00 | y = 68,850.48x − 3189.43 | 0.9999 | 0.03 | 0.08 |
| 4 | Procyanidin B1 | 1.56–100.00 | y = 8959.72x − 3750.11 | 0.9999 | 0.26 | 0.78 |
| 5 | 3,4-Dihydroxybenzaldehyde | 0.31–20.00 | y = 116,002.12x – 10,175.98 | 0.9999 | 0.02 | 0.07 |
| 6 | (+)-Catechin | 1.56–100.00 | y = 18,339.66x − 6373.92 | 0.9999 | 0.19 | 0.58 |
| 7 | Procyanidin B2 | 4.69–300.00 | y = 8625.79x − 8476.36 | 0.9999 | 0.72 | 2.19 |
| 8 | Epicatechin | 4.69–300.00 | y = 10,663.73x − 8145.46 | 0.9999 | 0.57 | 1.72 |
| 9 | (−)-Epicatechin gallate | 0.31–20.00 | y = 25,065.19x − 3031.59 | 0.9997 | 0.08 | 0.24 |
| No | Compound | Mean (mg/g) | SD × 10−2 | RSD (%) |
|---|---|---|---|---|
| 1 | Gallic acid | 0.32 | 0.63 | 1.97 |
| 2 | (−)-Gallocatechin | 3.40 | 6.19 | 1.82 |
| 3 | 3,4-Dihydroxybenzoic acid | 0.43 | 0.56 | 1.31 |
| 4 | Procyanidin B1 | 1.63 | 0.86 | 0.53 |
| 5 | 3,4-Dihydroxybenzaldehyde | 0.23 | 0.06 | 0.27 |
| 6 | (+)-Catechin | 2.57 | 0.75 | 0.29 |
| 7 | Procyanidin B2 | 5.69 a | 4.31 | 0.76 |
| 8 | Epicatechin | 8.53 | 5.64 | 0.66 |
| 9 | (−)-Epicatechin gallate | 0.11 | 0.03 | 0.31 |
| Term | Overlap | Adjusted p-Value | Odds Ratio | Combined Score | Genes |
|---|---|---|---|---|---|
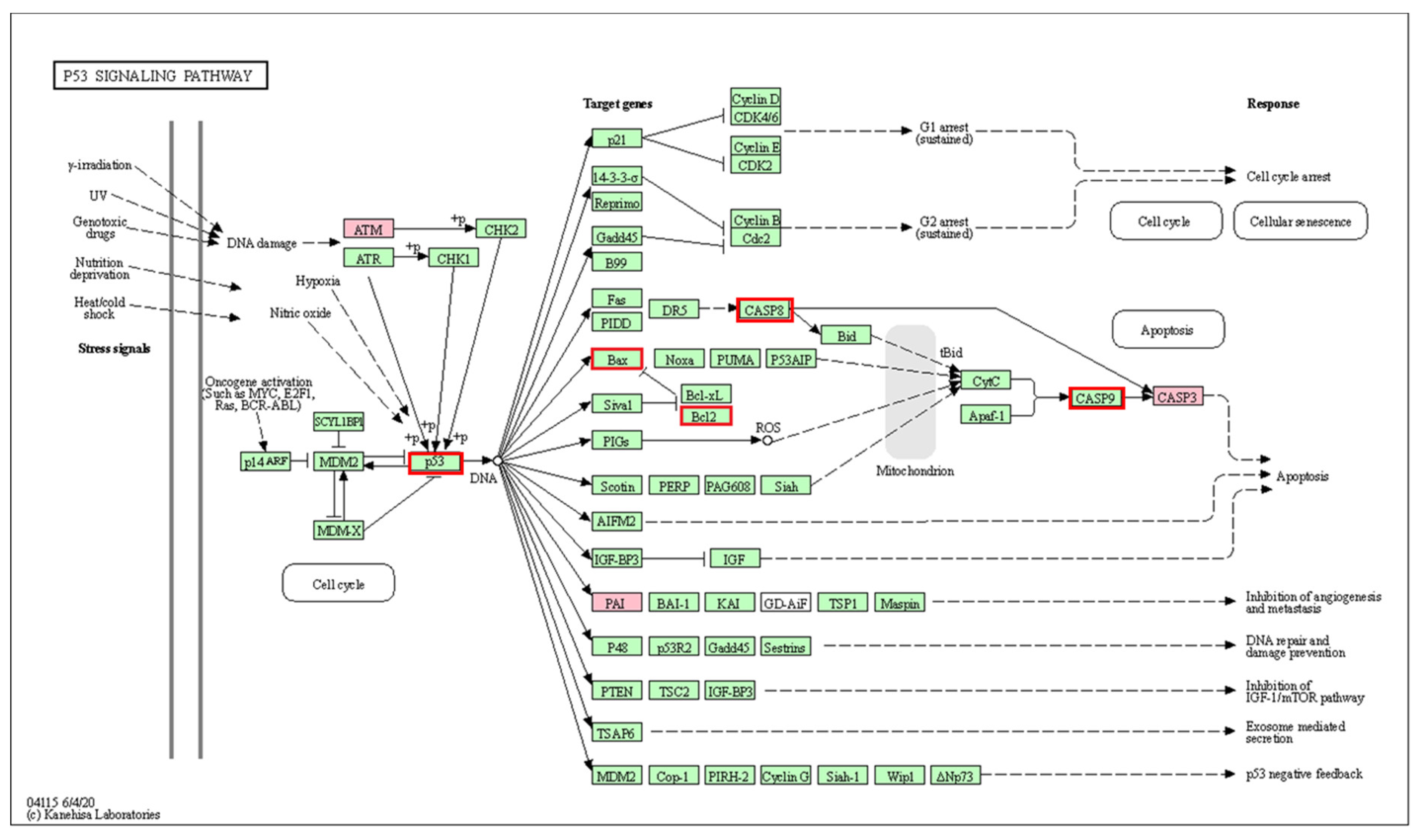
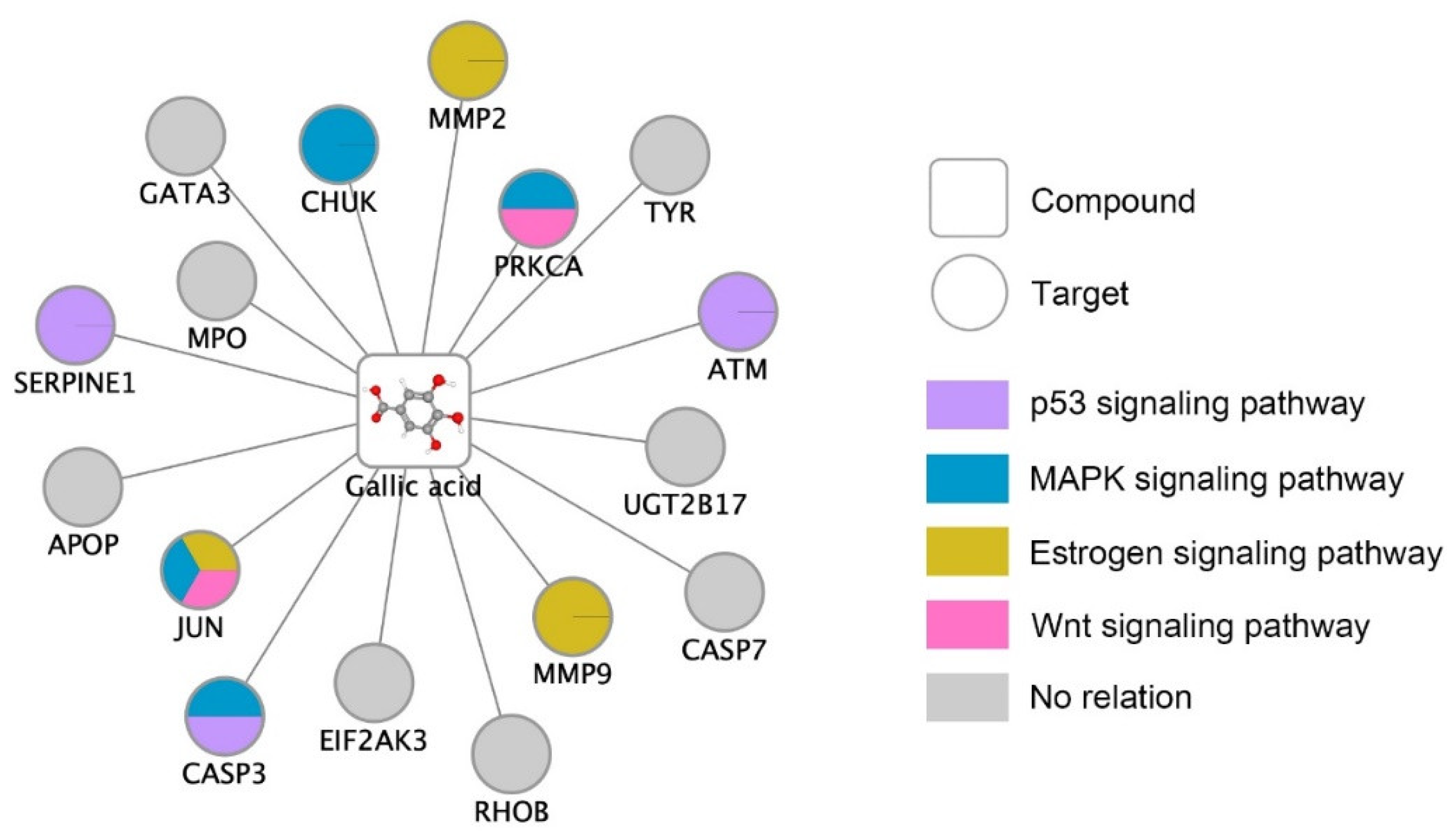
| p53 signaling pathway *** | 3/72 | 0.00053 | 52.08 | 553.55 | CASP3; SERPINE1; ATM |
| MAPK signaling pathway ** | 4/295 | 0.0019 | 16.95 | 161.35 | JUN; CHUK; CASP3; PRKCA |
| Estrogen signaling pathway ** | 3/137 | 0.0022 | 27.37 | 238.41 | JUN; MMP2; MMP9 |
| Wnt signaling pathway * | 2/158 | 0.034 | 15.82 | 78.69 | JUN; PRKCA |
| PI3K-Akt signaling pathway | 2/354 | 0.13 | 7.06 | 24.35 | CHUK; PRKCA |
| Homologous recombination | 1/41 | 0.13 | 30.49 | 104.64 | ATM |
| Cell cycle | 1/124 | 0.24 | 10.08 | 23.76 | ATM |
| Breast cancer | 1/147 | 0.26 | 8.50 | 18.66 | JUN |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Phung, H.M.; Lee, H.; Lee, S.; Jang, D.; Kim, C.-E.; Kang, K.S.; Seo, C.-S.; Choi, Y.-K. Analysis and Anticancer Effects of Active Compounds from Spatholobi Caulis in Human Breast Cancer Cells. Processes 2020, 8, 1193. https://doi.org/10.3390/pr8091193
Phung HM, Lee H, Lee S, Jang D, Kim C-E, Kang KS, Seo C-S, Choi Y-K. Analysis and Anticancer Effects of Active Compounds from Spatholobi Caulis in Human Breast Cancer Cells. Processes. 2020; 8(9):1193. https://doi.org/10.3390/pr8091193
Chicago/Turabian StylePhung, Hung Manh, Hesol Lee, Sullim Lee, Dongyeop Jang, Chang-Eop Kim, Ki Sung Kang, Chang-Seob Seo, and You-Kyung Choi. 2020. "Analysis and Anticancer Effects of Active Compounds from Spatholobi Caulis in Human Breast Cancer Cells" Processes 8, no. 9: 1193. https://doi.org/10.3390/pr8091193
APA StylePhung, H. M., Lee, H., Lee, S., Jang, D., Kim, C.-E., Kang, K. S., Seo, C.-S., & Choi, Y.-K. (2020). Analysis and Anticancer Effects of Active Compounds from Spatholobi Caulis in Human Breast Cancer Cells. Processes, 8(9), 1193. https://doi.org/10.3390/pr8091193









