Modelling and Development of Electrical Aptasensors: A Short Review
Abstract
1. Introduction
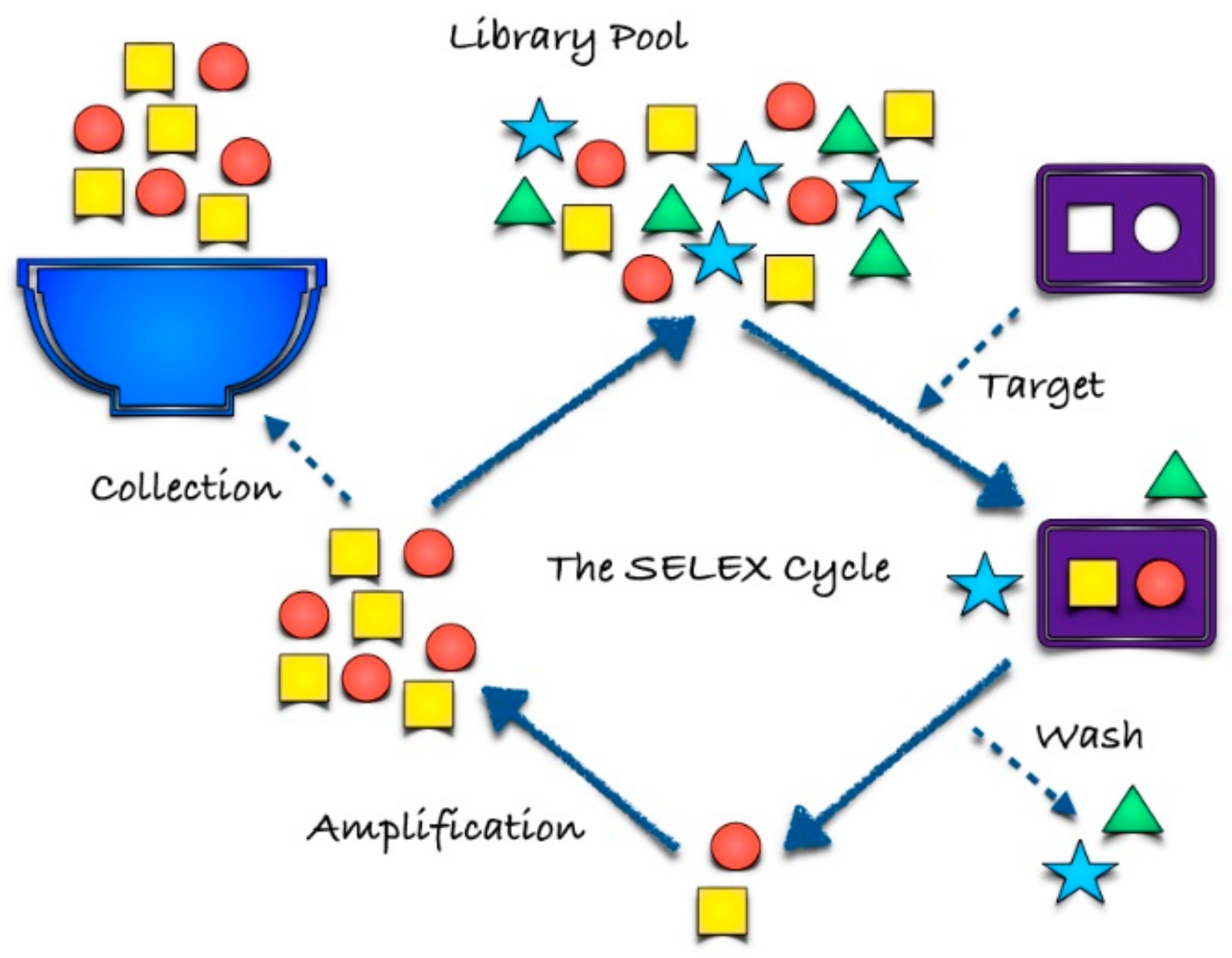
2. Aptamer Selection
2.1. SELEX and Cell-SELEX
2.2. Aptamer vs. Antibody in Cancer Therapy
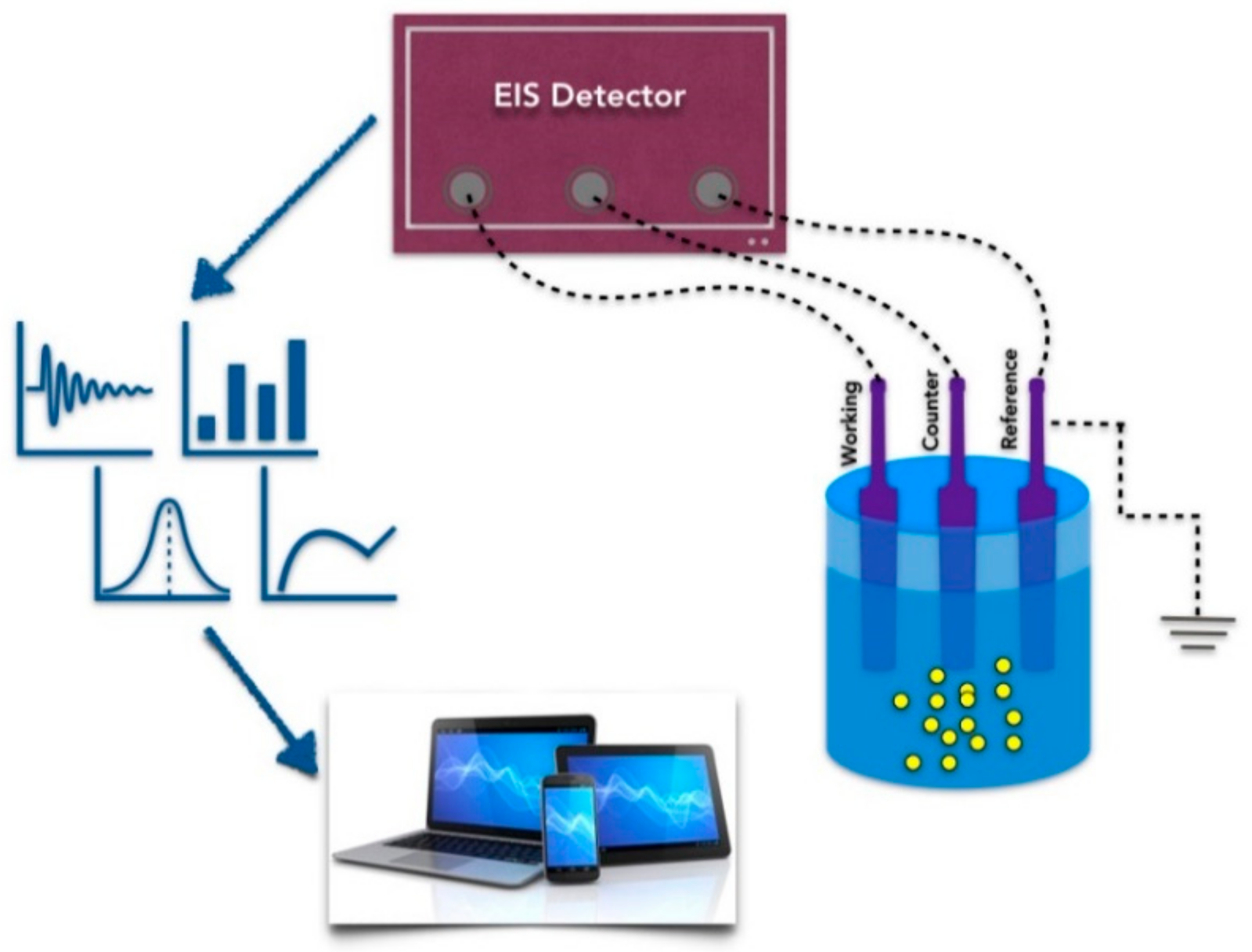
3. Aptamers in Electrical Nanobiosensors
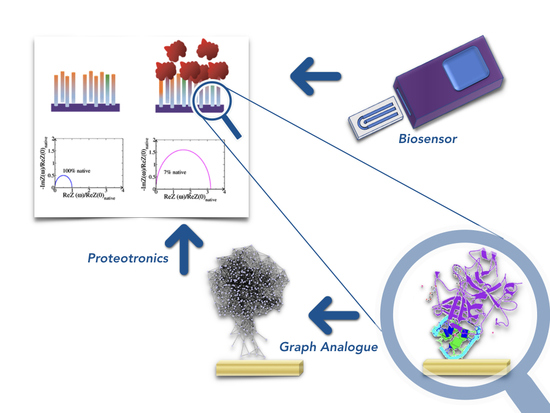
4. Modelling the Electrical Properties of Aptamers
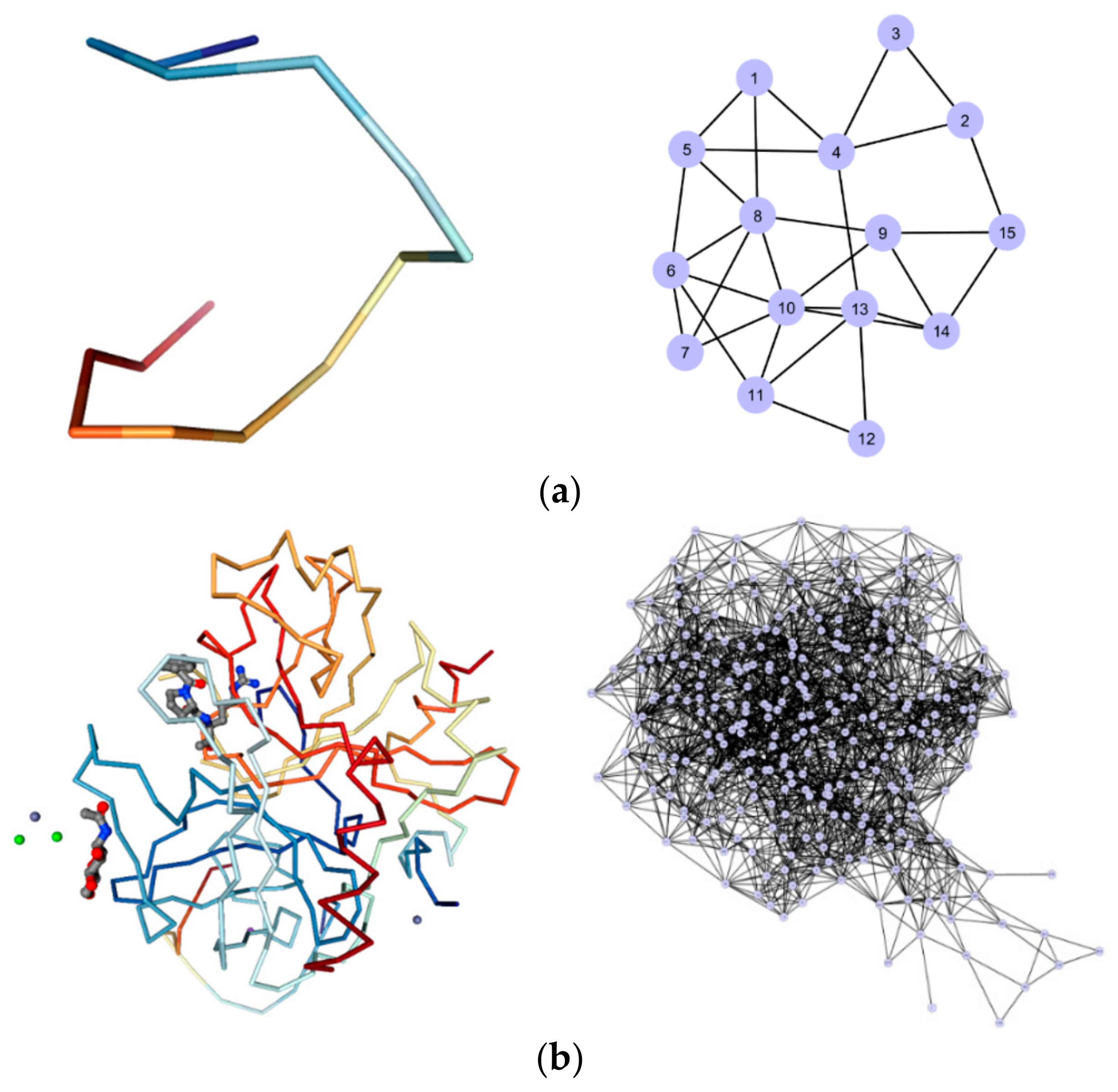
4.1. Complex Networks for Bioelectronics
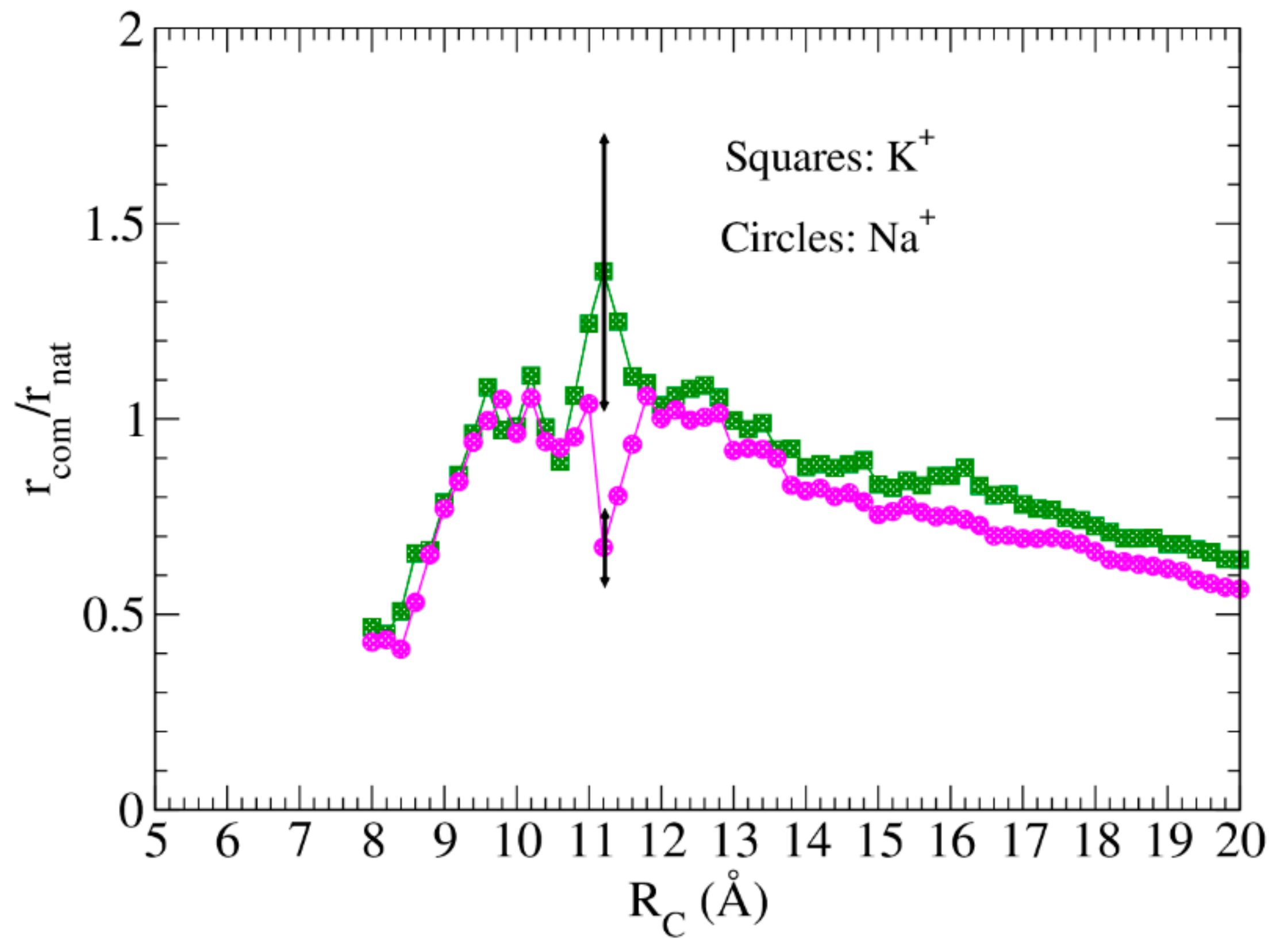
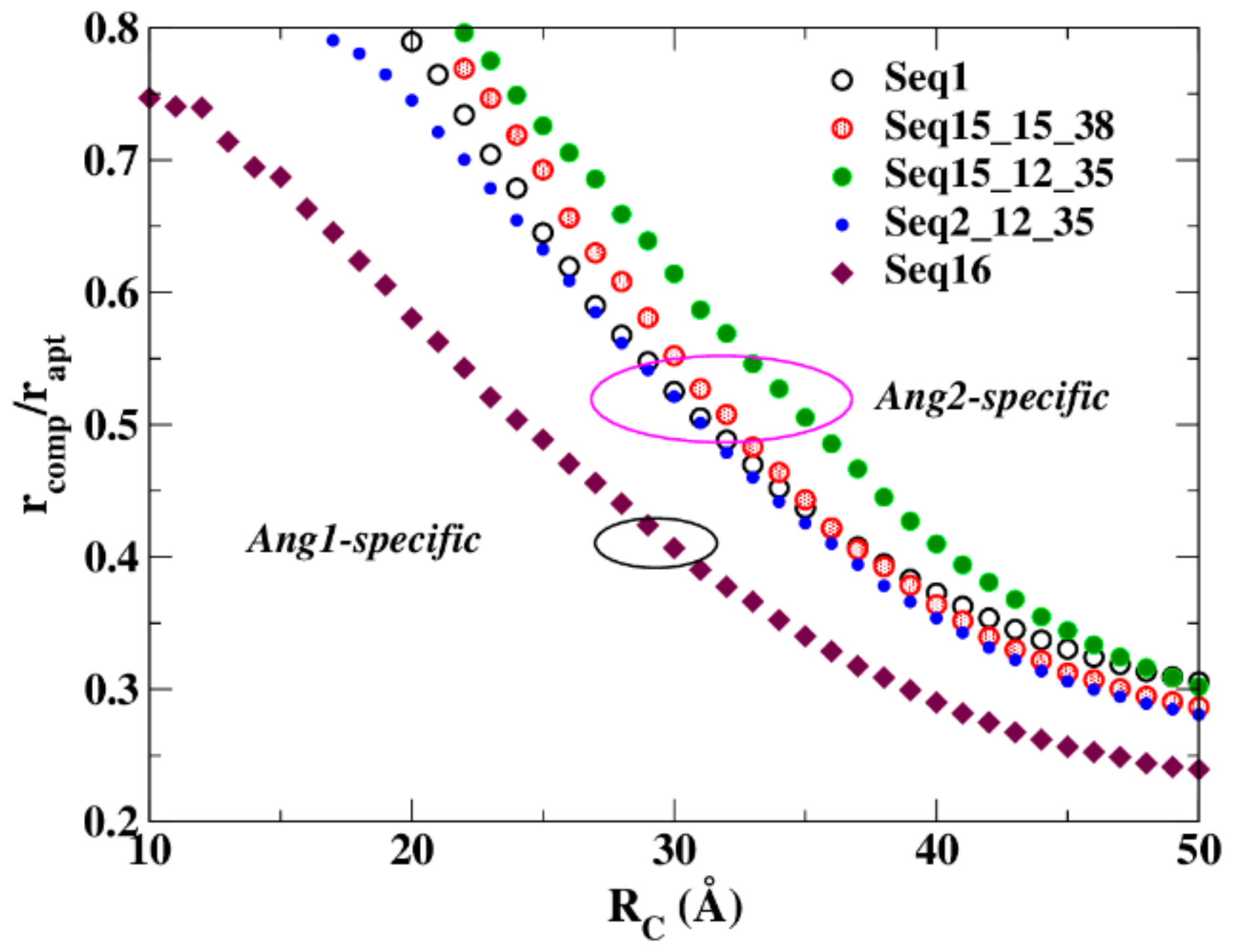
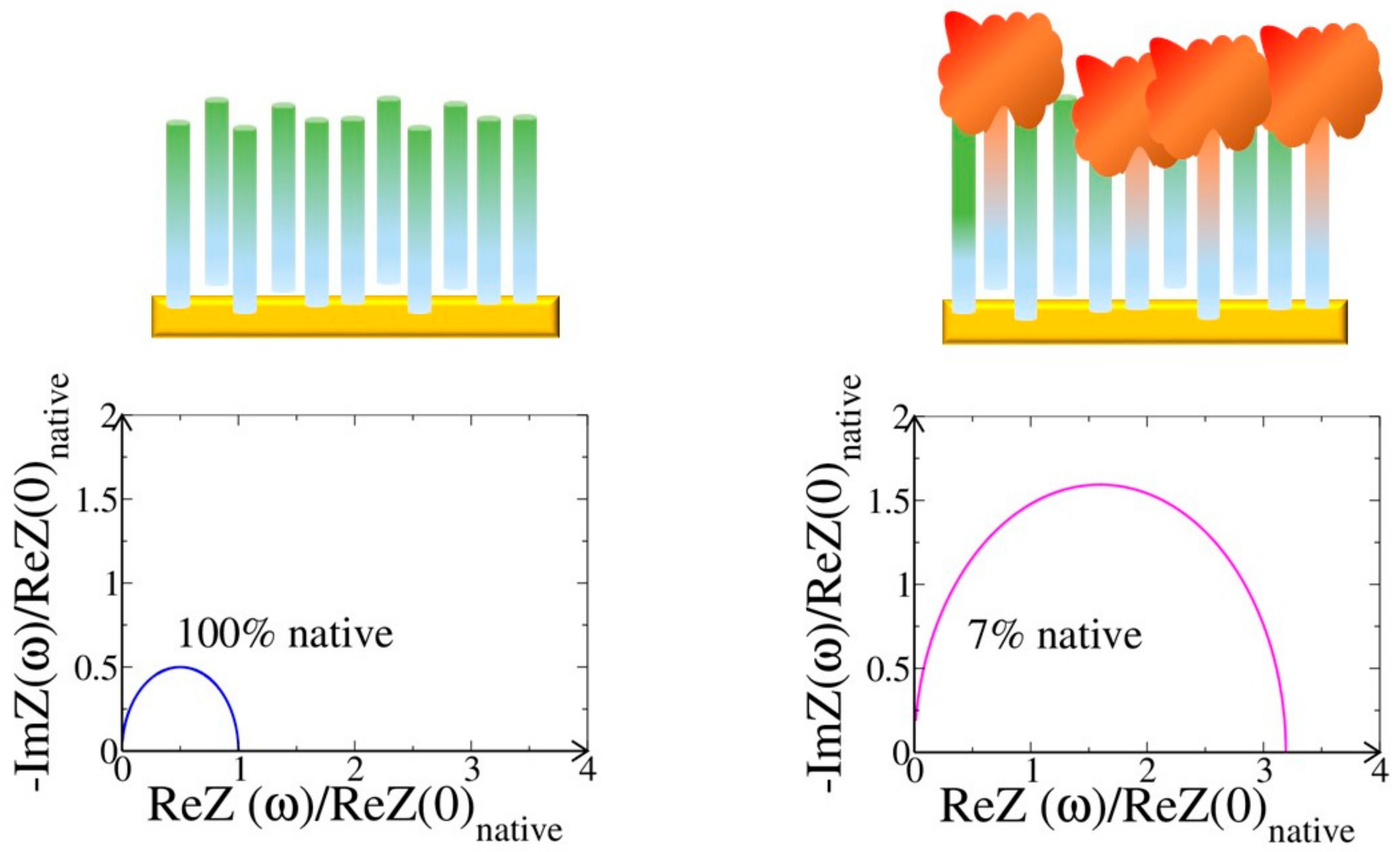
4.2. Resistance to Evaluate Binding Affinity
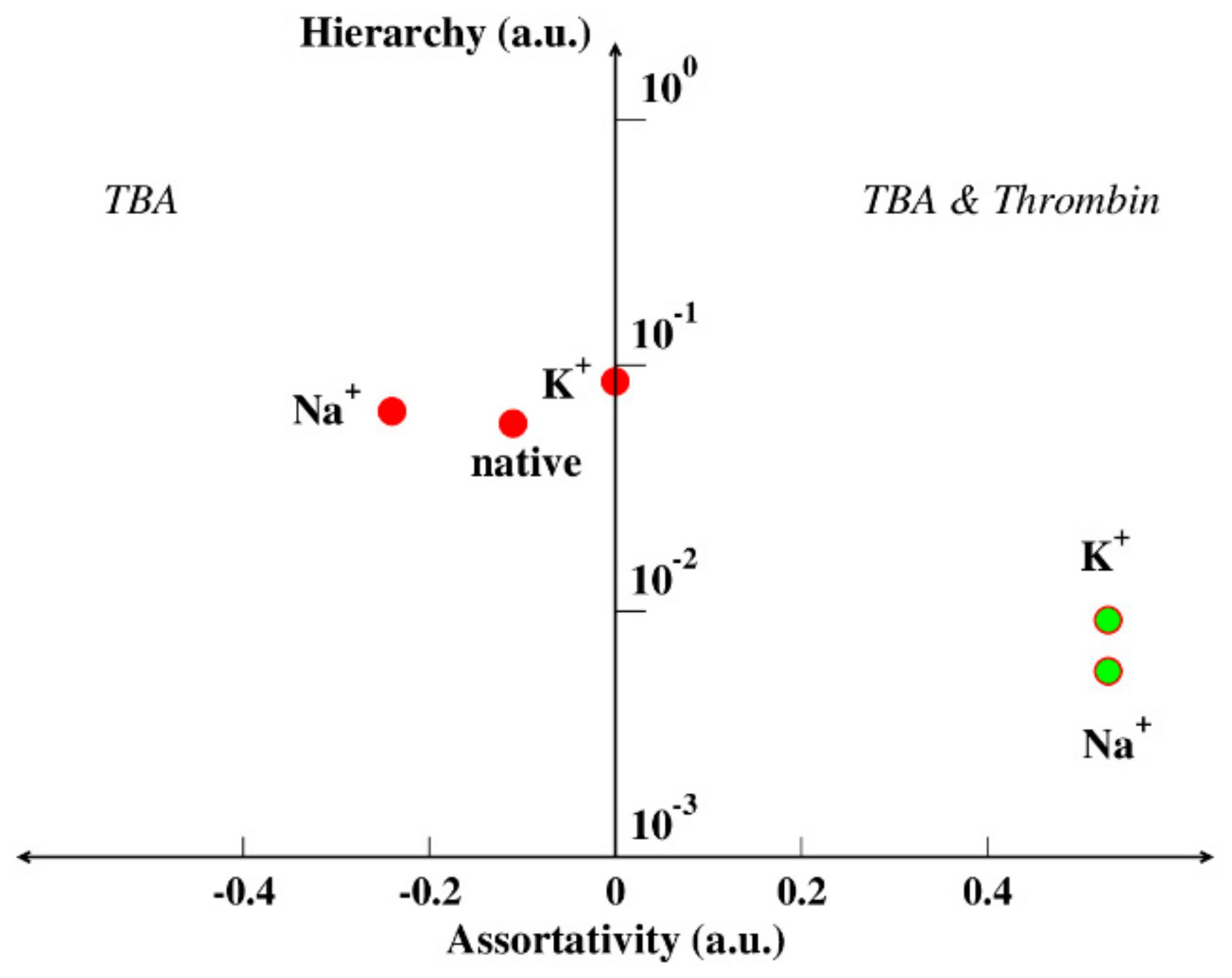
4.3. Hierarchy and Assortativity as a Measure of Binding Affinity
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Wu, X.; Chen, J.; Wu, M.; Zhao, J.X. Aptamers Active Targeting Ligands for Cancer Diagnosis and Therapy. Theranostics 2015, 5, 322–344. [Google Scholar] [CrossRef] [PubMed]
- Strehlitz, B. Selex and its recent optimizations. In Aptamers in Bioanalysis; Mascini, M., Ed.; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2003; p. 33. [Google Scholar]
- Yang, J.; Palla, M.; Bosco, F.G.; Rindzevicius, T.; Alstrøm, T.S.; Schmidt, M.S.; Boisen, A.; Ju, J.; Lin, Q. Surface-enhanced raman spectroscopy based quantitative bioassay on aptamer-functionalized nanopillars using large-area raman mapping. ACS Nano 2013, 7, 5350–5359. [Google Scholar]
- Noonan, P.S.; Roberts, R.H.; Schwartz, D.K. Liquid crystal reorientation induced by aptamer conformational changes. J. Am. Chem. Soc. 2013, 135, 5183–5189. [Google Scholar] [CrossRef] [PubMed]
- Famulok, M.; Mayer, G. Aptamers and SELEX in Chemistry & Biology. Chem. Biol. 2014, 21, 1054–1058. [Google Scholar]
- Chen, T.; Shukoor, M.I.; Chen, Y.; Yuan, Q.; Zhu, Z.; Zhao, Z.; Gulbakan, B.; Tan, W. Aptamer-conjugated nanomaterials for bioanalysis and biotechnology applications. Nanoscale 2011, 3, 546–556. [Google Scholar] [CrossRef] [PubMed]
- Tan, W.; Donovan, M.J.; Jiang, J. Aptamers from cell-based selection for bioanalytical applications. Chem. Rev. 2013, 113, 2842–2862. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.; Zu, Y. A Highlight of Recent Advances in Aptamer Technology and Its Application. Molecules 2015, 20, 11959–11980. [Google Scholar] [CrossRef] [PubMed]
- Tuerk, C.; Gold, L. Systematic evolution of ligands by exponential enrichment, RNA ligands to bacteriophage T4 DNA polymerase. Science 1990, 10, 505–510. [Google Scholar] [CrossRef]
- Gopinath, S.C.B. Methods developed for SELEX. Anal. Bioanal. Chem. 2007, 387, 171–182. [Google Scholar] [CrossRef] [PubMed]
- Shangguan, D.; Bing, T.; Zhang, N. Cell-SELEX, Aptamer Selection Against Whole Cells. In Aptamers Selected by Cell-SELEX for Theranostics; Tan, W., Fang, X., Eds.; Springer-Verlag: Berlin/Heidelberg, Germany, 2015. [Google Scholar] [CrossRef]
- Lyu, Y.; Chen, G.; Shangguan, D.; Zhang, L.; Wan, S.; Wu, Y.; Zhang, H.; Duan, L.; Liu, C.; You, M.; et al. Generating Cell Targeting Aptamers for Nanotheranostics Using Cell-SELEX. Theranostics 2016, 6, 1440–1452. [Google Scholar] [CrossRef] [PubMed]
- Ravalli, A.; Voccia, D.; Palchetti, I.; Marrazza, G. Electrochemical, electrochemiluminescence, and photoelectrochemical aptamer-based nanostructured sensors for biomarker analysis. Biosensors 2016, 6, 39. [Google Scholar] [CrossRef] [PubMed]
- Scott, A.M.; Wolchok, J.D.; Old, L.J. Antibody therapy of cancer. Nat. Rev. Cancer 2012, 12, 278–287. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.K. The history of monoclonal antibody development—Progress, remaining challenges and future innovations. Ann. Med. Surg. 2014, 3, 113–116. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.W.; Kim, H.J.; Heo, K. Therapeutic aptamers: Developmental potential as anticancer drugs. BMB Rep. 2015, 48, 234–237. [Google Scholar] [CrossRef] [PubMed]
- Lao, Y.H.; Phua, K.K.L.; Leong, K.W. Aptamer Nanomedicine for Cancer Therapeutics, Barriers and Potential for Translation. ACS Nano 2015, 9, 2235–2254. [Google Scholar] [CrossRef] [PubMed]
- Peer, D.; Karp, J.M.; Hong, S.; Farokhzad, O.C.; Margalit, R.; Langer, R. Nanocarriers as an emerging platform for cancer therapy. Nat. Nanotechnol. 2007, 2, 751–760. [Google Scholar] [CrossRef] [PubMed]
- Shangguan, D.; Li, Y. Aptamers evolved from live cells as effective molecular probes for cancer study. Proc. Natl. Acad. Sci. USA 2006, 103, 11838–11843. [Google Scholar] [CrossRef] [PubMed]
- Do Carmo, F.S.; Rocha Pinto, S.; Camões Orlando, M.M.; de Souza Albernaz, M.; de Souza Junqueira, M.; Soares Bernardes, E.; Cerecetto, H.; Moglioni, A.; Kozempel, J.; Szwed, M.; et al. Nano-Aptamer for Breast Cancer Imaging, Initial Considerations. J. Diagn. Imaging Ther. 2015, 2, 41–49. [Google Scholar] [CrossRef][Green Version]
- Do Carmo, F.S.; Ricci-Junior, E.; Cerqueira-Coutinho, C.; de Souza Albernaz, M.; Bernardes, E.S.; Missailidis, S.; Santos-Oliveira, R. Anti-MUC1 nano-aptamers for triple-negative breast cancer imaging by single-photon emission computed tomography in inducted animals: Initial considerations. Int. J. Nanomed. 2017, 12, 53–60. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Zhou, S.; Cai, Y.; Tang, F. Nucleic acid aptamer application in diagnosis and therapy of colorectal cancer based on cell-SELEX technology. Precis. Oncol. 2017, 1, 37. [Google Scholar] [CrossRef]
- Lee, J.-O.; So, H.-M.; Jeon, E.-K.; Chang, H.; Won, K.; Kim, Y.H. Aptamers as molecular recognition elements for electrical nanobiosensors. Anal. Bioanal. Chem. 2008, 390, 1023–1032. [Google Scholar] [CrossRef] [PubMed]
- Daniels, J.S.; Pourmand, N. Label-Free Impedance Biosensors, Opportunities and Challenges. Electroanalysis 2007, 19, 1239–1257. [Google Scholar] [CrossRef] [PubMed]
- Radi, A.-E. Electrochemical Aptamer-Based Biosensors, Recent Advances and Perspectives. Int. J. Electrochem. 2011, 2011, 863196. [Google Scholar] [CrossRef]
- Ikebukuro, K.; Kiyohara, C.; Sode, K. Electrochemical detection of protein using a double aptamer sandwich. Anal. Lett. 2004, 37, 2901–2909. [Google Scholar] [CrossRef]
- Putzbach, W.; Ronkainen, N.J. Immobilization techniques in the fabrication of nanomaterial-based electrochemical biosensors: A review. Sensors 2013, 13, 4811–4840. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Xu, J.; Liu, J.; Wang, X.; Chen, B. Disease-Related Detection with Electrochemical, Biosensors, A Review. Sensors 2017, 17, 2375. [Google Scholar] [CrossRef] [PubMed]
- Kaisti, M. Detection principles of biological and chemical FET sensors. Biosens. Bioelectron. 2017, 98, 437–448. [Google Scholar] [CrossRef] [PubMed]
- Meirinho, S.G.; Dias, L.G.; Peres, A.M.; Rodrigues, L.R. Voltammetric aptasensors for protein disease biomarkers detection: A. review. Biotechnol. Adv. 2016, 34, 941–953. [Google Scholar] [CrossRef] [PubMed]
- Abi, A.; Mohammadpour, Z.; Zuo, X.; Safavi, A. Nucleic Acid-Based Electrochemical Nanobiosensors. Biosens. Bioelectron. 2018, 102, 479–489. [Google Scholar] [CrossRef] [PubMed]
- Rapini, R.; Marrazza, G. Electrochemical aptasensors for contaminants detection in food and environment: Recent advances. Bioelectrochemistry 2017, 118, 47–61. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Song, C.; Yang, K.; Hong, W.; Lu, Y.; Yu, P.; Mao, L. Photo-Induced Regeneration of an Aptamer-based Electrochemical Sensor for Sensitively Detecting Adenosine Triphosphate. Anal. Chem. 2018, in press. [Google Scholar]
- Tang, Y.; Liu, P.; Xu, J.; Li, L.; Yang, L.; Liu, X.; Zhou, Y. Electrochemical aptasensor based on a novel flower-like TiO2 nanocomposite for the detection of tetracycline. Sens. Actuators B Chem. 2018, 258, 906–912. [Google Scholar] [CrossRef]
- Yu, Z.; Lai, R.Y. A reagentless and reusable electrochemical aptamer-based sensor for rapid detection of ampicillin in complex samples. Talanta 2018, 176, 619–624. [Google Scholar] [CrossRef] [PubMed]
- Chekin, F.; Bagga, K.; Subramanian, P.; Jijie, R.; Singh, S.K.; Kurungot, S.; Szunerits, S. Nucleic aptamer modified porous reduced graphene oxide/MoS2 based electrodes for viral detection: Application to human papillomavirus (HPV). Sens. Actuators B Chem. 2018, 262, 991–1000. [Google Scholar] [CrossRef]
- Figueroa-Miranda, G.; Feng, L.; Shiu, S.C.C.; Dirkzwager, R.M.; Cheung, Y.W.; Tanner, J.A.; Mayer, D. Aptamer-based electrochemical biosensor for highly sensitive and selective malaria detection with adjustable dynamic response range and reusability. Sens. Actuators B Chem. 2018, 255, 235–243. [Google Scholar] [CrossRef]
- Khosravi, F.; Loeian, S.M.; Panchapakesan, B. Ultrasensitive Label-Free Sensing of IL-6 Based on PASE Functionalized Carbon Nanotube Micro-Arrays with RNA-Aptamers as Molecular Recognition Elements. Biosensors 2017, 7, 17. [Google Scholar] [CrossRef] [PubMed]
- Pulikkathodi, A.K.; Sarangadharan, I.; Hsu, C.P.; Chen, Y.H.; Hung, L.Y.; Lee, G.Y.; Wang, Y.L. Enumeration of circulating tumor cells and investigation of cellular responses using aptamer-immobilized AlGaN/GaN high electron mobility transistor sensor array. Sens. Actuators B Chem. 2018, 257, 96–104. [Google Scholar] [CrossRef]
- Meirinho, S.G.; Dias, L.G.; Peres, A.M.; Rodrigues, L.R. Electrochemical aptasensor for human osteopontin detection using a DNA aptamer selected by SELEX. Anal. Chim. Acta 2017, 987, 25–37. [Google Scholar] [CrossRef] [PubMed]
- Majd, S.M.; Salimi, A. Ultrasensitive flexible FET-type aptasensor for CA 125 cancer marker detection based on carboxylated multiwalled carbon nanotubes immobilized onto reduced graphene oxide film. Anal. Chim. Acta 2018, 1000, 273–282. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Guo, Z.; Tang, Y.; Shen, Y.; Miao, P. A highly sensitive gold nanoparticle-based electrochemical aptasensor for theophylline detection. Anal. Chim. Acta 2018, 999, 54–59. [Google Scholar] [CrossRef] [PubMed]
- Gutierrez, R.; Porath, D.; Cuniberti, G.; Baranowski, S. Charge Transport in Disordered Solids with Applications in Electronics; Wiley: New York, NY, USA, 2006. [Google Scholar]
- Regan, J.J.; Risser, S.M.; Beratan, D.N.; Onuchic, J.N. Protein electron transport: Single versus multiple pathways. J. Phys. Chem. 1993, 97, 13083–13088. [Google Scholar] [CrossRef]
- Alfinito, E.; Pousset, J.; Reggiani, L. Proteotronics: Development of Protein-Based Electronics; CRC Press: Boca Raton, FL, USA, 2015. [Google Scholar]
- Berman, H.M.; Bhat, T.N.; Bourne, P.E.; Feng, Z.; Gilliland, G.; Weissig, H.; Westbrook, J. The Protein Data Bank and the challenge of structural genomics. Nat. Struct. Mol. Biol. 2000, 7, 957–959. [Google Scholar] [CrossRef] [PubMed]
- Gilson, M.K.; Zhou, H.X. Calculation of protein-ligand binding affinities. Annu. Rev. Biophys. Biomol. Struct. 2007, 36, 21–42. [Google Scholar] [CrossRef] [PubMed]
- Rother, K.; Rother, M.; Boniecki, M.; Puton, T.; Bujnicki, J.M. RNA and protein 3D structure modeling: Similarities and differences. J. Mol. Model. 2011, 17, 2325–2336. [Google Scholar] [CrossRef] [PubMed]
- Rabal, O.; Pastor, F.; Villanueva, H.; Soldevilla, M.S.; Hervas-Stubbs, S.; Oyarzabal, J. In Silico Aptamer Docking Studies, From a Retrospective Validation to a Prospective Case Study—TIM3 Aptamers Binding. Mol. Ther. Nucleic Acids 2016, 5, e376. [Google Scholar] [CrossRef] [PubMed]
- Poblete, S.; Bottaro, S.; Bussi, G. A nucleobase-centered coarse-grained representation for structure prediction of RNA motifs. Nucleic Acids Res. 2017, 46, 1674–1683. [Google Scholar] [CrossRef] [PubMed]
- Cataldo, R.; Alfinito, E.; Reggiani, L. Hierarchy and Assortativity as New Tools for Binding-Affinity Investigation, the Case of the TBA Aptamer-Ligand Complex. IEEE Trans. Nanobiosci. 2018, 16, 896–904. [Google Scholar] [CrossRef] [PubMed]
- Alfinito, E.; Reggiani, L. Role of topology in electrical properties of bacterio-rhodopsin and rat olfactory receptor I7. Phys. Rev. E 2010, 81, 032902. [Google Scholar] [CrossRef] [PubMed]
- Alfinito, E.; Millithaler, J.F.; Reggiani, L.; Zine, N.; Jaffrezic-Renault, N. Human olfactory receptor 17–40 as an active part of a nanobiosensor: A microscopic investigation of its electrical properties. RSC Adv. 2011, 1, 123–127. [Google Scholar] [CrossRef]
- Alfinito, E.; Millithaler, J.F.; Reggiani, L. Charge transport in purple membrane monolayers: A sequential tunneling approach. Phys. Rev. E 2011, 83, 042902. [Google Scholar] [CrossRef] [PubMed]
- Alfinito, E.; Reggiani, L. Modeling current-voltage characteristics of proteorhodopsin and bacteriorhodopsin: Towards an optoelectronics based on proteins. IEEE Trans. Nanobiosci. 2016, 15, 775–780. [Google Scholar] [CrossRef] [PubMed]
- Alfinito, E.; Reggiani, L.; Cataldo, R.; De Nunzio, G.; Giotta, L.; Guascito, M.R. Modeling the microscopic electrical properties of thrombin binding aptamer (TBA) for label-free biosensors. Nanotechnology 2017, 28, 065502. [Google Scholar] [CrossRef] [PubMed]
- Cataldo, R.; Ciriaco, F.; Alfinito, E. A new strategy to evaluate aptamer binding affinity. arXiv, 2017; arXiv:1704.00341. [Google Scholar]
- Alfinito, E.; Pousset, J.; Reggiani, L.; Lee, K. Photoreceptors for a light biotransducer: A comparative study of the electrical responses of two (type-1) opsins. Nanotechnology 2013, 24, 395501. [Google Scholar] [CrossRef] [PubMed]
- Alfinito, E.; Pennetta, C.; Reggiani, L. A network model to correlate conformational change and the impedance spectrum of single proteins. Nanotechnology 2008, 19, 065202. [Google Scholar] [CrossRef] [PubMed]
- Russo Krauss, I.; Merlino, A.; Randazzo, A.; Novellino, E.; Mazzarella, L.; Sica, F. High-resolution structures of two complexes between thrombin and thrombin-binding aptamer shed light on the role of cations in the aptamer inhibitory activity. Nucleic Acids Res. 2012, 40, 8119–8128. [Google Scholar] [CrossRef] [PubMed]
- Schultze, P.; Macaya, R.F.; Feigon, J. Three-dimensional solution structure of the thrombin-binding DNA aptamer d (GGTTGGTGTGGTTGG). J. Mol. Boil. 1994, 235, 1532–1547. [Google Scholar] [CrossRef] [PubMed]
- Cai, H.; Lee, T.M.H.; Hsing, I.M. Label-free protein recognition using an aptamer-based impedance measurement assay. Sens. Actuators B Chem. 2006, 114, 433–437. [Google Scholar] [CrossRef]
- Hu, W.P.; Kumar, J.V.; Huang, C.J.; Chen, W.Y. Computational selection of RNA aptamer against angiopoietin-2 and experimental evaluation. BioMed Res. Int. 2015. [Google Scholar] [CrossRef] [PubMed]
- Perry, C.; Pyatt, R. Network theory’s contribution to the development of marketing research. In Proceedings of the 1995 World Marketing Congress; Springer: Cham, Switzerland, 2015; pp. 188–196. [Google Scholar]
- Azevedo, H.; Moreira-Filho, C.A. Topological robustness analysis of protein interaction networks reveals key targets for overcoming chemotherapy resistance in glioma. Sci. Rep. 2015, 19, 16830. [Google Scholar] [CrossRef] [PubMed]
| Target | Experimental Technique | Innovation | LOD | Reference |
|---|---|---|---|---|
| ATP | EIS/switch | UV regeneration | 1 nM | [33] |
| Tetracycline | EIS | MoS2-TiO2@Au arrays | 10 pM | [34] |
| Ampicillin | ACV SWV | Fast response; reusability for at least 3 times | 1 μM (ACV) 30 μM (SWV) | [35] |
| Papilloma virus | CV | Glassy carbon electrodes | 1.75 pM | [36] |
| Malaria biomarker | EIS | Adjustable detection range, reusability | 0.84 pM | [37] |
| Interleukin-6 cancer marker | FET | Nanotube arrays | 10 pg/mL | [38] |
| Cancer cells | FET | Whole cell detection | 1 cell/mL | [39] |
| Human osteopontin | SWV CV | SWV vs. CV | 1.4 nM (SWV) 4.2 nM (ACV) | [40] |
| CA125-cancer marker | FET | Multiwalled carbon nanotubes | 5 × 10−10 U/mL | [41] |
| Theophylline | EIS | Gold nanoparticle | 0.7 μM | [42] |
| Biomolecule | Target/Inhibitor | Experimental Technique | References |
|---|---|---|---|
| Anti-thrombin aptamer (TBA) | Thrombin | SPR | [51,56] |
| Bacteriorhodopsin | Light | Conductive atomic force spectroscopy | [52,54,55] |
| Bovine rhodopsin | Light | EIS | [55] |
| OR I7 | Octanal, heptanal | EIS | [52] |
| OR 17-40 | Helional | EIS, SPR | [53] |
| Anti-Ang2 aptamer | Ang2 | EIS | [57] |
| Proteorhodopsin | Light | UV-spectroscopy | [58] |
| AChE | Several toxins | Potentiometric | [59] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cataldo, R.; Leuzzi, M.; Alfinito, E. Modelling and Development of Electrical Aptasensors: A Short Review. Chemosensors 2018, 6, 20. https://doi.org/10.3390/chemosensors6020020
Cataldo R, Leuzzi M, Alfinito E. Modelling and Development of Electrical Aptasensors: A Short Review. Chemosensors. 2018; 6(2):20. https://doi.org/10.3390/chemosensors6020020
Chicago/Turabian StyleCataldo, Rosella, Maria Leuzzi, and Eleonora Alfinito. 2018. "Modelling and Development of Electrical Aptasensors: A Short Review" Chemosensors 6, no. 2: 20. https://doi.org/10.3390/chemosensors6020020
APA StyleCataldo, R., Leuzzi, M., & Alfinito, E. (2018). Modelling and Development of Electrical Aptasensors: A Short Review. Chemosensors, 6(2), 20. https://doi.org/10.3390/chemosensors6020020