Identification of Protein Biomarker Signatures for Acute Myeloid Leukemia (AML) Using Both Nontargeted and Targeted Approaches
Abstract
1. Introduction
2. Materials and Methods
2.1. Clinical Samples
2.2. Sample Preparation for Mass Spectrometry
2.3. Label-Free Liquid Chromatography Mass Spectrometry
2.4. Protein Identification and Quantification
2.5. Luminex Assay
3. Results
3.1. Proteomics Profiling of Human Bone Marrow Cells
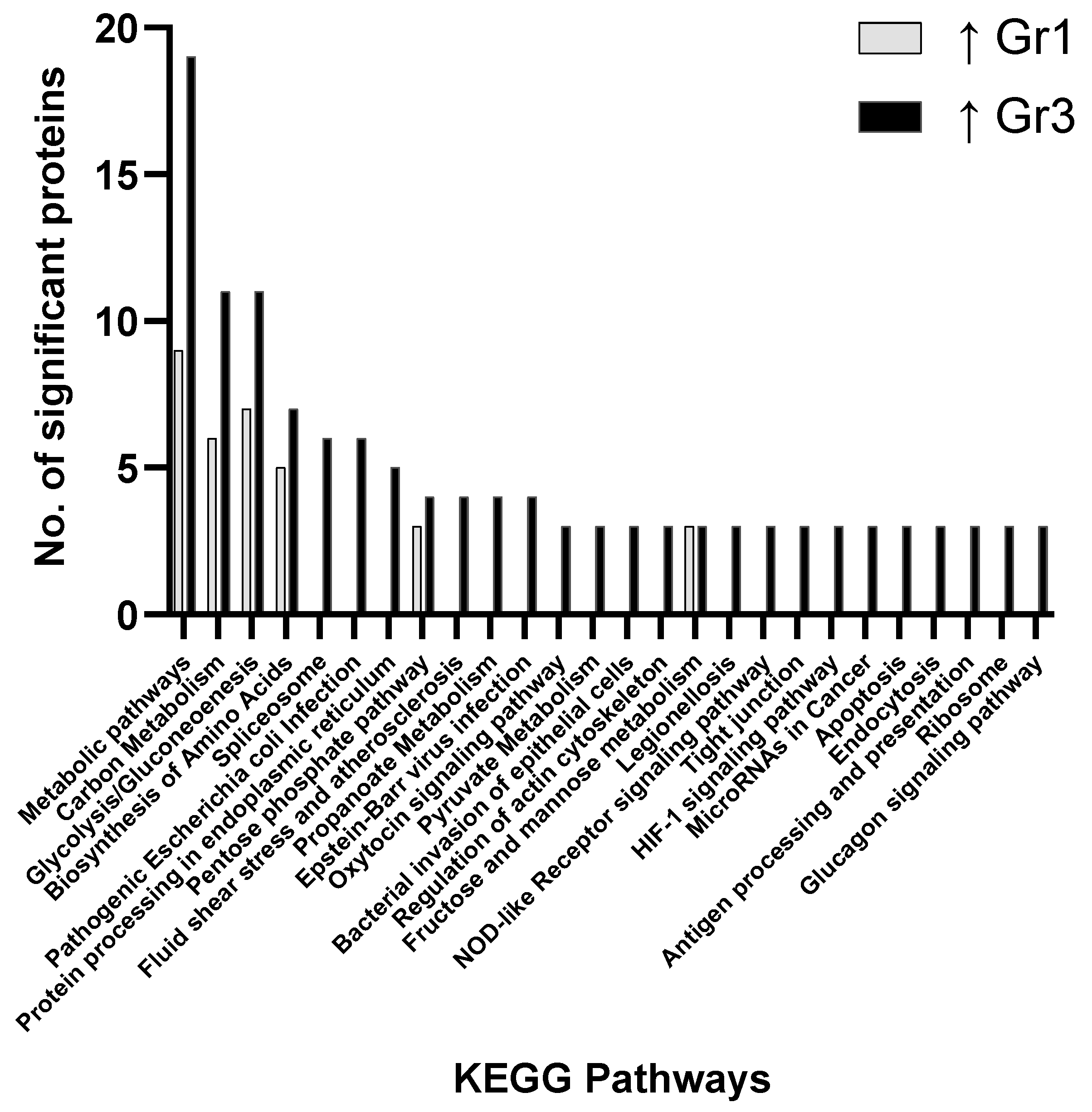
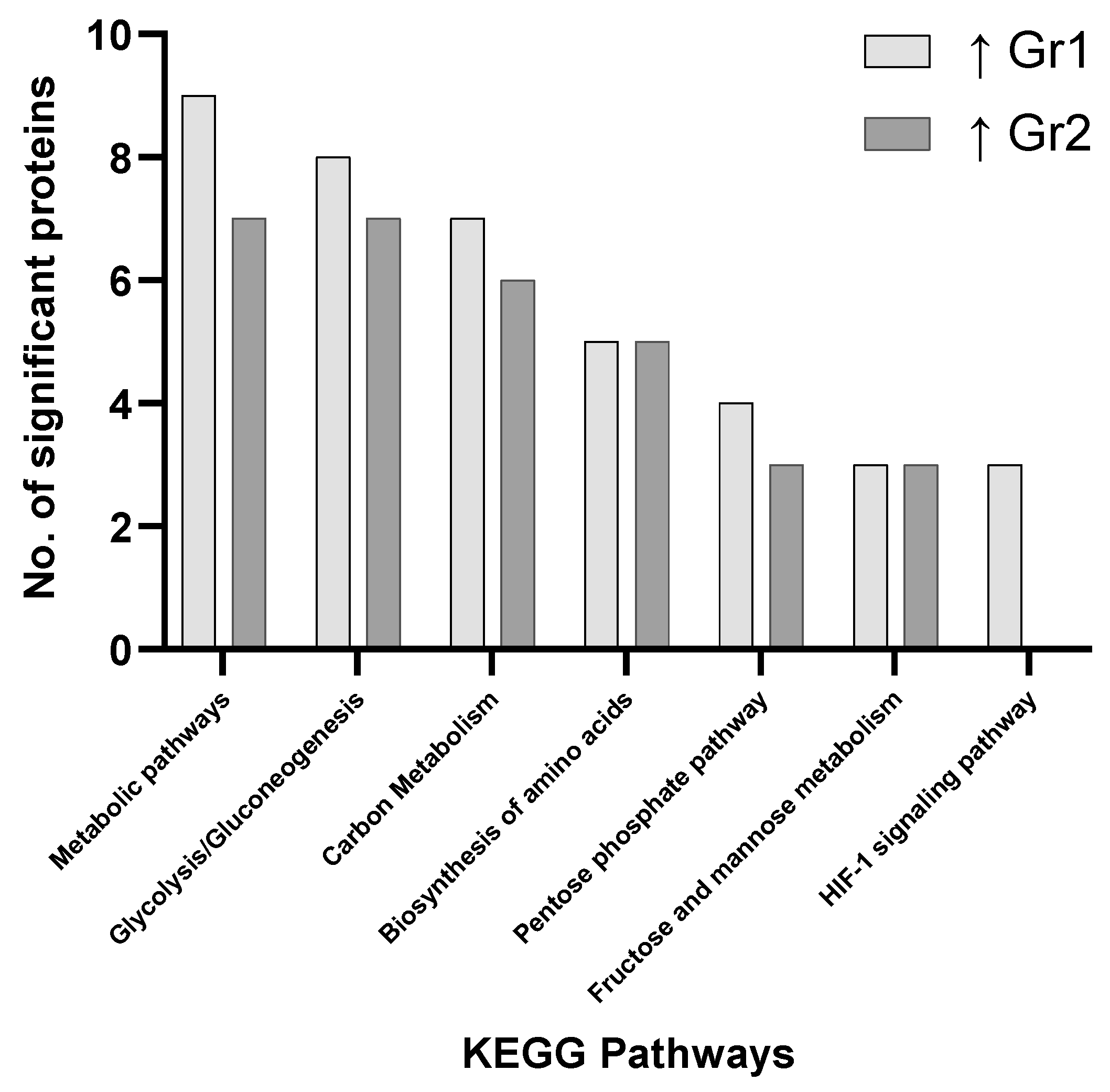
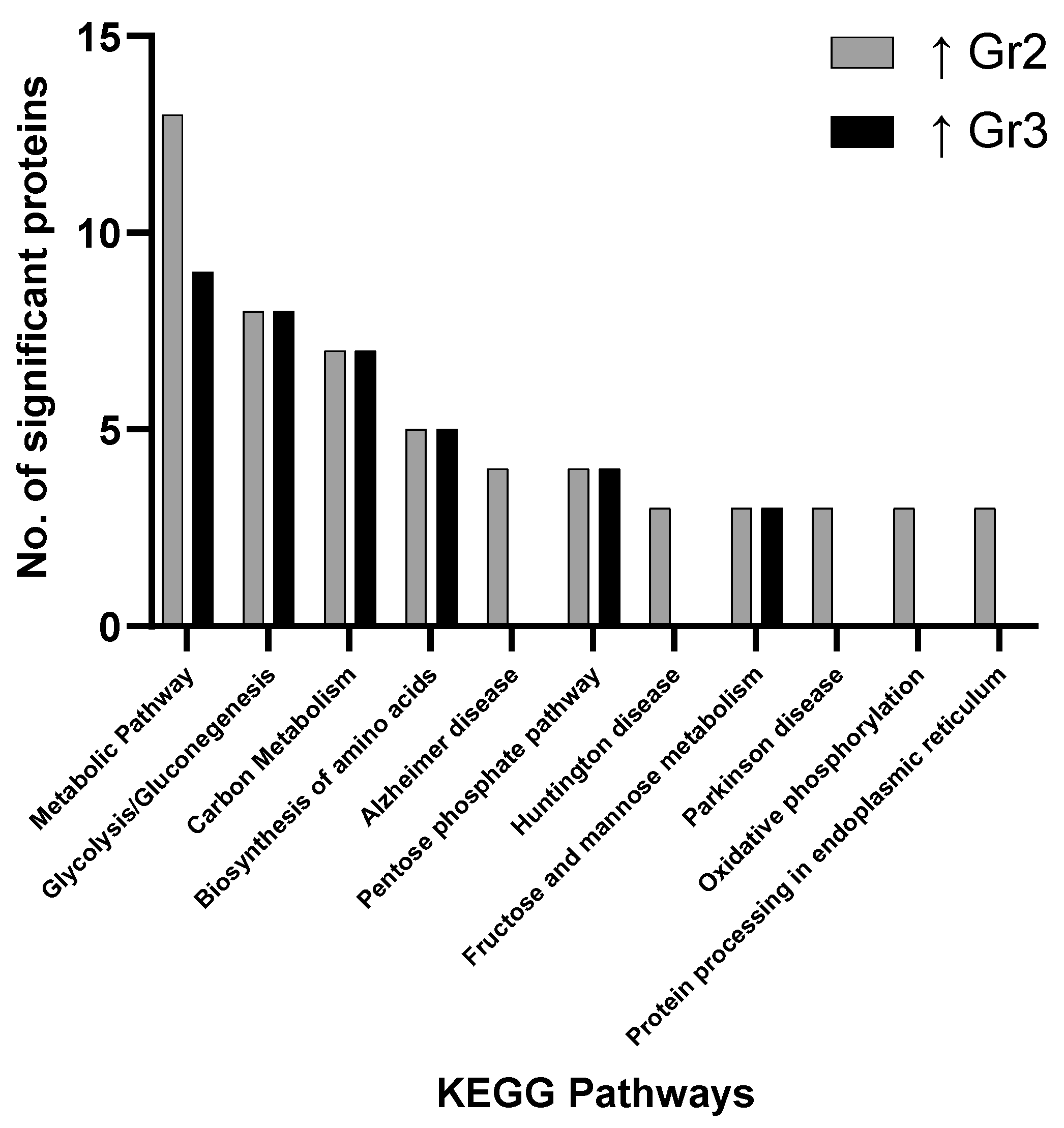
3.2. Pathway Analysis
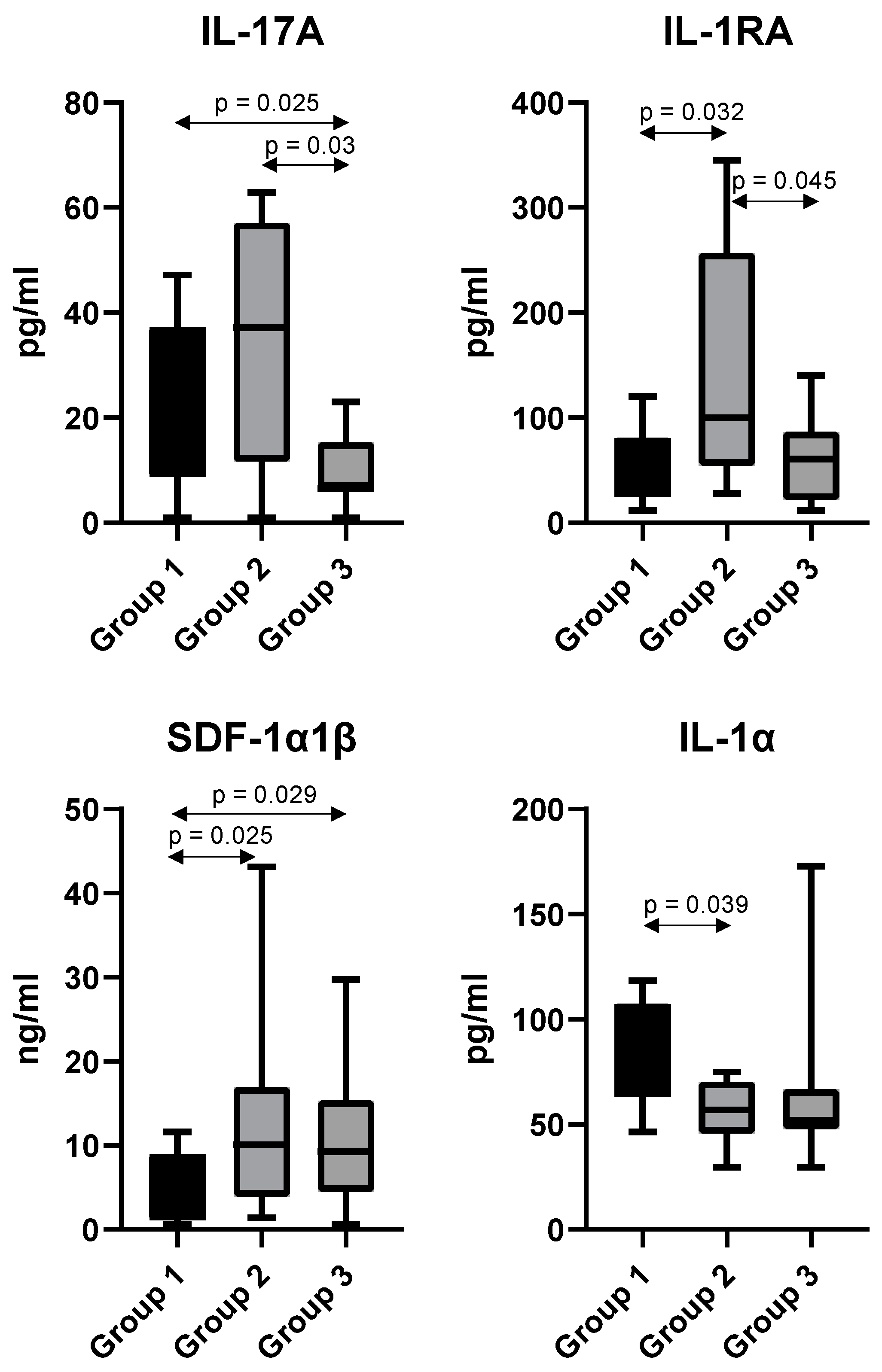
3.3. Targeted Proteomics Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- De Kouchkovsky, I.; Abdul-Hay, M. Acute myeloid leukemia: A comprehensive review and 2016 update. Blood Cancer J. 2016, 6, e441. [Google Scholar] [CrossRef]
- Hao, T.; Li-Talley, M.; Buck, A.; Chen, W. An emerging trend of rapid increase of leukemia but not all cancers in the aging population in the United States. Sci. Rep. 2019, 9, 12070. [Google Scholar] [CrossRef] [PubMed]
- Shallis, R.M.; Wang, R.; Davidoff, A.; Ma, X.; Zeidan, A.M. Epidemiology of acute myeloid leukemia: Recent progress and enduring challenges. Blood Rev. 2019, 36, 70–87. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Gu, Y.; Chen, B. Mechanisms of drug resistance in acute myeloid leukemia. OncoTargets Ther. 2019, 12, 1937–1945. [Google Scholar] [CrossRef] [PubMed]
- Bennett, J.M.; Catovsky, D.; Daniel, M.T.; Flandrin, G.; Galton, D.A.; Gralnick, H.R.; Sultan, C. Proposed revised criteria for the classification of acute myeloid leukemia. A report of the French-American-British Cooperative Group. Ann. Intern. Med. 1985, 103, 620–625. [Google Scholar] [CrossRef]
- Vardiman, J.W.; Thiele, J.; Arber, D.A.; Brunning, R.D.; Borowitz, M.J.; Porwit, A.; Harris, N.L.; Le Beau, M.M.; Hellström-Lindberg, E.; Tefferi, A.; et al. The 2008 revision of the World Health Organization (WHO) classification of myeloid neoplasms and acute leukemia: Rationale and important changes. Blood 2009, 114, 937–951. [Google Scholar] [CrossRef] [PubMed]
- Patel, J.P.; Gönen, M.; Figueroa, M.E.; Fernandez, H.; Sun, Z.; Racevskis, J.; Van Vlierberghe, P.; Dolgalev, I.; Thomas, S.; Aminova, O.; et al. Prognostic relevance of integrated genetic profiling in acute myeloid leukemia. N. Engl. J. Med. 2012, 366, 1079–1089. [Google Scholar] [CrossRef]
- Döhner, H.; Estey, E.; Grimwade, D.; Amadori, S.; Appelbaum, F.R.; Büchner, T.; Dombret, H.; Ebert, B.L.; Fenaux, P.; Larson, R.A.; et al. Diagnosis and management of AML in adults: 2017 ELN recommendations from an international expert panel. Blood 2017, 129, 424–447. [Google Scholar] [CrossRef]
- Hernandez-Valladares, M.; Bruserud, Ø.; Selheim, F. The Implementation of Mass Spectrometry-Based Proteomics Workflows in Clinical Routines of Acute Myeloid Leukemia: Applicability and Perspectives. Int. J. Mol. Sci. 2020, 21, 6830. [Google Scholar] [CrossRef]
- Eisfeld, A.K.; Kohlschmidt, J.; Mrózek, K.; Blachly, J.S.; Walker, C.J.; Nicolet, D.; Orwick, S.; Maharry, S.E.; Carroll, A.J.; Stone, R.M.; et al. Mutation patterns identify adult patients with de novo acute myeloid leukemia aged 60 years or older who respond favorably to standard chemotherapy: An analysis of Alliance studies. Leukemia 2018, 32, 1338–1348. [Google Scholar] [CrossRef]
- Patel, S.S.; Kuo, F.C.; Gibson, C.J.; Steensma, D.P.; Soiffer, R.J.; Alyea, E.P., 3rd; Chen, Y.A.; Fathi, A.T.; Graubert, T.A.; Brunner, A.M.; et al. High NPM1-mutant allele burden at diagnosis predicts unfavorable outcomes in de novo AML. Blood 2018, 131, 2816–2825. [Google Scholar] [CrossRef] [PubMed]
- Papaemmanuil, E.; Gerstung, M.; Bullinger, L.; Gaidzik, V.I.; Paschka, P.; Roberts, N.D.; Potter, N.E.; Heuser, M.; Thol, F.; Bolli, N.; et al. Genomic Classification and Prognosis in Acute Myeloid Leukemia. N. Engl. J. Med. 2016, 374, 2209–2221. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, A.; Creek, D.J. Discovery and Validation of Clinical Biomarkers of Cancer: A Review Combining Metabolomics and Proteomics. Proteomics 2019, 19, e1700448. [Google Scholar] [CrossRef]
- Aasebø, E.; Berven, F.S.; Bartaula-Brevik, S.; Stokowy, T.; Hovland, R.; Vaudel, M.; Døskeland, S.O.; McCormack, E.; Batth, T.S.; Olsen, J.V.; et al. Proteome and Phosphoproteome Changes Associated with Prognosis in Acute Myeloid Leukemia. Cancers 2020, 12, 709. [Google Scholar] [CrossRef] [PubMed]
- Grønningsæter, I.S.; Reikvam, H.; Aasebø, E.; Bartaula-Brevik, S.; Tvedt, T.H.; Bruserud, Ø.; Hatfield, K.J. Targeting Cellular Metabolism in Acute Myeloid Leukemia and The Role of Patient Heterogeneity. Cells 2020, 9, 1155. [Google Scholar] [CrossRef]
- Stockard, B.; Garrett, T.; Guingab-Cagmat, J.; Meshinchi, S.; Lamba, J. Distinct Metabolic features differentiating FLT3-ITD AML from FLT3-WT childhood Acute Myeloid Leukemia. Sci. Rep. 2018, 8, 5534. [Google Scholar] [CrossRef]
- Riether, C.; Schürch, C.M.; Bührer, E.D.; Hinterbrandner, M.; Huguenin, A.L.; Hoepner, S.; Zlobec, I.; Pabst, T.; Radpour, R.; Ochsenbein, A.F. CD70/CD27 signaling promotes blast stemness and is a viable therapeutic target in acute myeloid leukemia. J. Exp. Med. 2017, 214, 359–380. [Google Scholar] [CrossRef] [PubMed]
- Mi, H.; Muruganujan, A.; Ebert, D.; Huang, X.; Thomas, P.D. PANTHER version 14: More genomes, a new PANTHER GO-slim and improvements in enrichment analysis tools. Nucleic Acids Res. 2019, 47, D419–D426. [Google Scholar] [CrossRef]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Soc. Ser. B Methodol. 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Stanta, G.; Bonin, S. Overview on Clinical Relevance of Intra-Tumor Heterogeneity. Front. Med. 2018, 5, 85. [Google Scholar] [CrossRef]
- Fisher, R.; Pusztai, L.; Swanton, C. Cancer heterogeneity: Implications for targeted therapeutics. Br. J. Cancer 2013, 108, 479–485. [Google Scholar] [CrossRef]
- Pogosova-Agadjanyan, E.L.; Moseley, A.; Othus, M.; Appelbaum, F.R.; Chauncey, T.R.; Chen, I.L.; Erba, H.P.; Godwin, J.E.; Fang, M.; Kopecky, K.J.; et al. Impact of Specimen Heterogeneity on Biomarkers in Repository Samples from Patients with Acute Myeloid Leukemia: A SWOG Report. Biopreserv. Biobank. 2018, 16, 42–52. [Google Scholar] [CrossRef]
- Klco, J.M.; Spencer, D.H.; Miller, C.A.; Griffith, M.; Lamprecht, T.L.; O’Laughlin, M.; Fronick, C.; Magrini, V.; Demeter, R.T.; Fulton, R.S.; et al. Functional heterogeneity of genetically defined subclones in acute myeloid leukemia. Cancer Cell 2014, 25, 379–392. [Google Scholar] [CrossRef] [PubMed]
- Kaushik, A.K.; DeBerardinis, R.J. Applications of metabolomics to study cancer metabolism. Biochim. Biophys. Acta Rev. Cancer 2018, 1870, 2–14. [Google Scholar] [CrossRef] [PubMed]
- Henkenius, K.; Greene, B.H.; Barckhausen, C.; Hartmann, R.; Märken, M.; Kaiser, T.; Rehberger, M.; Metzelder, S.K.; Parak, W.J.; Neubauer, A.; et al. Maintenance of cellular respiration indicates drug resistance in acute myeloid leukemia. Leuk Res. 2017, 62, 56–63. [Google Scholar] [CrossRef] [PubMed]
- Ryu, M.J.; Han, J.; Kim, S.J.; Lee, M.J.; Ju, X.; Lee, Y.L.; Son, J.H.; Cui, J.; Jang, Y.; Chung, W.; et al. PTEN/AKT signaling mediates chemoresistance in refractory acute myeloid leukemia through enhanced glycolysis. Oncol. Rep. 2019, 42, 2149–2158. [Google Scholar] [CrossRef]
- Stuani, L.; Riols, F.; Millard, P.; Sabatier, M.; Batut, A.; Saland, E.; Viars, F.; Tonini, L.; Zaghdoudi, S.; Linares, L.K.; et al. Stable Isotope Labeling Highlights Enhanced Fatty Acid and Lipid Metabolism in Human Acute Myeloid Leukemia. Int. J. Mol. Sci. 2018, 19, 3325. [Google Scholar] [CrossRef]
- Chen, Y.; Xu, Q.; Ji, D.; Wei, Y.; Chen, H.; Li, T.; Wan, B.; Yuan, L.; Huang, R.; Chen, G. Inhibition of pentose phosphate pathway suppresses acute myelogenous leukemia. Tumour Biol. 2016, 37, 6027–6034. [Google Scholar] [CrossRef]
- Forte, D.; García-Fernández, M.; Sánchez-Aguilera, A.; Stavropoulou, V.; Fielding, C.; Martín-Pérez, D.; López, J.A.; Costa, A.S.H.; Tronci, L.; Nikitopoulou, E.; et al. Bone Marrow Mesenchymal Stem Cells Support Acute Myeloid Leukemia Bioenergetics and Enhance Antioxidant Defense and Escape from Chemotherapy. Cell Metab. 2020, 32, 829–843. [Google Scholar] [CrossRef]
- Pikman, Y.; Puissant, A.; Alexe, G.; Furman, A.; Chen, L.M.; Frumm, S.M.; Ross, L.; Fenouille, N.; Bassil, C.F.; Lewis, C.A.; et al. Targeting MTHFD2 in acute myeloid leukemia. J. Exp. Med. 2016, 213, 1285–1306. [Google Scholar] [CrossRef]
- Schnittger, S.; Haferlach, C.; Ulke, M.; Alpermann, T.; Kern, W.; Haferlach, T. IDH1 mutations are detected in 6.6% of 1414 AML patients and are associated with intermediate risk karyotype and unfavorable prognosis in adults younger than 60 years and unmutated NPM1 status. Blood 2010, 116, 5486–5496. [Google Scholar] [CrossRef] [PubMed]
- DiNardo, C.D.; Propert, K.J.; Loren, A.W.; Paietta, E.; Sun, Z.; Levine, R.L.; Straley, K.S.; Yen, K.; Patel, J.P.; Agresta, S.; et al. Serum 2-hydroxyglutarate levels predict isocitrate dehydrogenase mutations and clinical outcome in acute myeloid leukemia. Blood 2013, 121, 4917–4924. [Google Scholar] [CrossRef] [PubMed]
- Miller, A.; Tong, A.W.; Sweetman, L.; Theiss, A.; Murtaza, M.; Daoud, Y.; Wong, L. Characterization of Acute Myeloid Leukaemia (AML) Patients with Elevated Peripheral Blood Plasma D-2-Hydroxyglutarate (D-2HG) and/or Isocitrate Dehydrogenase (IDH) Mutational Status. Blood 2017, 130, 3923. [Google Scholar] [CrossRef]
- Chen, W.L.; Wang, J.H.; Zhao, A.H.; Xu, X.; Wang, Y.H.; Chen, T.L.; Li, J.M.; Mi, J.Q.; Zhu, Y.M.; Liu, Y.F.; et al. A distinct glucose metabolism signature of acute myeloid leukemia with prognostic value. Blood 2014, 124, 1645–1654. [Google Scholar] [CrossRef] [PubMed]
- Vitkevičienė, A.; Janulis, V.; Žučenka, A.; Borutinskaitė, V.; Kaupinis, A.; Valius, M.; Griškevičius, L.; Navakauskienė, R. Oxidative phosphorylation inhibition induces anticancerous changes in therapy-resistant-acute myeloid leukemia patient cells. Mol. Carcinog. 2019, 58, 2008–2016. [Google Scholar] [CrossRef] [PubMed]
- Advani, A.S.; Li, H.; Michaelis, L.C.; Medeiros, B.C.; Liedtke, M.; List, A.F.; O’Dwyer, K.; Othus, M.; Erba, H.P.; Appelbaum, F.R. Report of the relapsed/refractory cohort of SWOG S0919: A phase 2 study of idarubicin and cytarabine in combination with pravastatin for acute myelogenous leukemia (AML). Leuk Res. 2018, 67, 17–20. [Google Scholar] [CrossRef]
- Rector, R.S.; Payne, R.M.; Ibdah, J.A. Mitochondrial trifunctional protein defects: Clinical implications and therapeutic approaches. Adv. Drug Deliv. Rev. 2008, 60, 1488–1496. [Google Scholar] [CrossRef]
- Farge, T.; Saland, E.; de Toni, F.; Aroua, N.; Hosseini, M.; Perry, R.; Bosc, C.; Sugita, M.; Stuani, L.; Fraisse, M.; et al. Chemotherapy-Resistant Human Acute Myeloid Leukemia Cells Are Not Enriched for Leukemic Stem Cells but Require Oxidative Metabolism. Cancer Discov. 2017, 7, 716–735. [Google Scholar] [CrossRef]
- Shi, J.; Fu, H.; Jia, Z.; He, K.; Fu, L.; Wang, W. High Expression of CPT1A Predicts Adverse Outcomes: A Potential Therapeutic Target for Acute Myeloid Leukemia. EBioMedicine 2016, 14, 55–64. [Google Scholar] [CrossRef] [PubMed]
- Tabe, Y.; Yamamoto, S.; Saitoh, K.; Sekihara, K.; Monma, N.; Ikeo, K.; Mogushi, K.; Shikami, M.; Ruvolo, V.; Ishizawa, J.; et al. Bone Marrow Adipocytes Facilitate Fatty Acid Oxidation Activating AMPK and a Transcriptional Network Supporting Survival of Acute Monocytic Leukemia Cells. Cancer Res. 2017, 77, 1453–1464. [Google Scholar] [CrossRef]
- Lee, E.A.; Angka, L.; Rota, S.G.; Hanlon, T.; Mitchell, A.; Hurren, R.; Wang, X.M.; Gronda, M.; Boyaci, E.; Bojko, B.; et al. Targeting Mitochondria with Avocatin B Induces Selective Leukemia Cell Death. Cancer Res. 2015, 75, 2478–2488. [Google Scholar] [CrossRef]
- Kornberg, A.; Polliack, A. Serum lactic dehydrogenase (LDH) levels in acute leukemia: Marked elevations in lymphoblastic leukemia. Blood 1980, 56, 351–355. [Google Scholar] [CrossRef]
- Liu, D.; Wang, D.; Wu, C.; Zhang, L.; Mei, Q.; Hu, G.; Long, G.; Sun, W. Prognostic significance of serum lactate dehydrogenase in patients with breast cancer: A meta-analysis. Cancer Manag. Res. 2019, 11, 3611–3619. [Google Scholar] [CrossRef] [PubMed]
- Lippert, M.C.; Javadpour, N. Lactic dehydrogenase in the monitoring and prognosis of testicular cancer. Cancer 1981, 48, 2274–2278. [Google Scholar] [CrossRef]
- Krykowski, E.; Polkowska-Kulesza, E.; Robak, T.; Matusewicz, W.; Urbańska-Rys, H.; Hołub, A. Analysis of Prognostic Factors in Acute Leukemias in Adults. In Acute Leukemias; Springer: Berlin/Heidelberg, Germany, 1987; pp. 369–372. [Google Scholar]
- Geva, M.; Shouval, R.; Fein, J.A.; Danylesko, I.; Shem-Tov, N.; Yerushalmi, R.; Shimoni, A.; Nagler, A. Lactate Dehydrogenase Is a Key Prognostic Factor in Acute Myeloid Leukemia and Lymphoma Patients Undergoing Allogeneic Hematopoietic Stem Cell Transplantation. Blood 2019, 134, 3304. [Google Scholar] [CrossRef]
- Kalaycio, M.; Rybicki, L.; Pohlman, B.; Dean, R.; Sweetenham, J.W.; Andresen, S.; Sobecks, R.; Sekeres, M.A.; Advani, A.; Davis, R.; et al. Lactate Dehydrogenase (LDH) Level Predicts the Outcome of Patients with Acute Myelogenous Leukemia (AML) Following HLA-Matched Sibling Bone Marrow Transplant (BMT). Blood 2006, 108, 3013. [Google Scholar] [CrossRef]
- Stuart, S.D.; Schauble, A.; Gupta, S.; Kennedy, A.D.; Keppler, B.R.; Bingham, P.M.; Zachar, Z. A strategically designed small molecule attacks alpha-ketoglutarate dehydrogenase in tumor cells through a redox process. Cancer Metab. 2014, 2, 4. [Google Scholar] [CrossRef]
- Wakimoto, N.; Yokoyama, A.; Okabe-Kado, J.; Nagata, N.; Motoyoshi, K.; Honma, Y. Combined analysis of differentiation inhibitory factor nm23-H1 and nm23-H2 as prognostic factors in acute myeloid leukaemia. Br. J. Cancer 1998, 77, 2298–2303. [Google Scholar] [CrossRef][Green Version]
- Oliveira, P.J.; Urbano, A.M. Oncometabolism: The switchboard of cancer—An editorial. Biochim. Biophys. Acta BBA-Mol. Basis Dis. 2021, 1867, 166031. [Google Scholar] [CrossRef]
- Sallmyr, A.; Fan, J.; Datta, K.; Kim, K.-T.; Grosu, D.; Shapiro, P.; Small, D.; Rassool, F. Internal tandem duplication of FLT3 (FLT3/ITD) induces increased ROS production, DNA damage, and misrepair: Implications for poor prognosis in AML. Blood 2008, 111, 3173–3182. [Google Scholar] [CrossRef]
- Lo Presti, C.; Fauvelle, F.; Jacob, M.-C.; Mondet, J.; Mossuz, P. The metabolic reprogramming in acute myeloid leukemia patients depends on their genotype and is a prognostic marker. Blood Adv. 2021, 5, 156–166. [Google Scholar] [CrossRef] [PubMed]
- Döhner, K.; Paschka, P. Intermediate-risk acute myeloid leukemia therapy: Current and future. Hematology 2014, 2014, 34–43. [Google Scholar] [CrossRef] [PubMed]
- Lin, P.-H.; Li, H.-Y.; Fan, S.-C.; Yuan, T.-H.; Chen, M.; Hsu, Y.-H.; Yang, Y.-H.; Li, L.-Y.; Yeh, S.-P.; Bai, L.-Y.; et al. A targeted next-generation sequencing in the molecular risk stratification of adult acute myeloid leukemia: Implications for clinical practice. Cancer Med. 2017, 6, 349–360. [Google Scholar] [CrossRef] [PubMed]
- Pallarès, V.; Hoyos, M.; Chillón, M.C.; Barragán, E.; Prieto Conde, M.I.; Llop, M.; Falgàs, A.; Céspedes, M.V.; Montesinos, P.; Nomdedeu, J.F.; et al. Focal Adhesion Genes Refine the Intermediate-Risk Cytogenetic Classification of Acute Myeloid Leukemia. Cancers 2018, 10, 436. [Google Scholar] [CrossRef] [PubMed]
- Pardini, B.; Sabo, A.A.; Birolo, G.; Calin, G.A. Noncoding RNAs in Extracellular Fluids as Cancer Biomarkers: The New Frontier of Liquid Biopsies. Cancers 2019, 11, 1170. [Google Scholar] [CrossRef]
- Percival, M.E.; Lai, C.; Estey, E.; Hourigan, C.S. Bone marrow evaluation for diagnosis and monitoring of acute myeloid leukemia. Blood Rev. 2017, 31, 185–192. [Google Scholar] [CrossRef]
- Dinarello, C.A. Immunological and inflammatory functions of the interleukin-1 family. Annu. Rev. Immunol. 2009, 27, 519–550. [Google Scholar] [CrossRef]
- Tsimberidou, A.M.; Estey, E.; Wen, S.; Pierce, S.; Kantarjian, H.; Albitar, M.; Kurzrock, R. The prognostic significance of cytokine levels in newly diagnosed acute myeloid leukemia and high-risk myelodysplastic syndromes. Cancer 2008, 113, 1605–1613. [Google Scholar] [CrossRef] [PubMed]
- Fredly, H.; Reikvam, H.; Gjertsen, B.T.; Bruserud, Ø. Disease-stabilizing treatment with all-trans retinoic acid and valproic acid in acute myeloid leukemia: Serum hsp70 and hsp90 levels and serum cytokine profiles are determined by the disease, patient age, and anti-leukemic treatment. Am. J. Hematol. 2012, 87, 368–376. [Google Scholar] [CrossRef] [PubMed]
- Tao, M.; Li, B.; Nayini, J.; Andrews, C.B.; Huang, R.W.; Devemy, E.; Song, S.; Venugopal, P.; Preisler, H.D. SCF, IL-1beta, IL-1ra and GM-CSF in the bone marrow and serum of normal individuals and of AML and CML patients. Cytokine 2000, 12, 699–707. [Google Scholar] [CrossRef]
- Stosić-Grujicić, S.; Basara, N.; Dinarello, C.A. Modulatory in vitro effects of interleukin-1 receptor antagonist (IL-1Ra) or antisense oligonucleotide to interleukin-1 beta converting enzyme (ICE) on acute myeloid leukaemia (AML) cell growth. Clin. Lab. Haematol. 1999, 21, 173–185. [Google Scholar] [CrossRef]
- Baker, K.J.; Houston, A.; Brint, E. IL-1 Family Members in Cancer; Two Sides to Every Story. Front. Immunol. 2019, 10, 1197. [Google Scholar] [CrossRef]
- Carey, A.; Edwards, D.K.t.; Eide, C.A.; Newell, L.; Traer, E.; Medeiros, B.C.; Pollyea, D.A.; Deininger, M.W.; Collins, R.H.; Tyner, J.W.; et al. Identification of Interleukin-1 by Functional Screening as a Key Mediator of Cellular Expansion and Disease Progression in Acute Myeloid Leukemia. Cell Rep. 2017, 18, 3204–3218. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Ye, A.; Bi, L.; Wu, J.; Yu, K.; Zhang, S. Th17 cells and interleukin-17 increase with poor prognosis in patients with acute myeloid leukemia. Cancer Sci. 2014, 105, 933–942. [Google Scholar] [CrossRef]
- Abousamra, N.K.; Salah El-Din, M.; Helal, R. Prognostic value of Th17 cells in acute leukemia. Med. Oncol. 2013, 30, 732. [Google Scholar] [CrossRef]
- Kryczek, I.; Wei, S.; Szeliga, W.; Vatan, L.; Zou, W. Endogenous IL-17 contributes to reduced tumor growth and metastasis. Blood 2009, 114, 357–359. [Google Scholar] [CrossRef] [PubMed]
- Bhat, S.; Gardi, N.; Hake, S.; Kotian, N.; Sawant, S.; Kannan, S.; Parmar, V.; Desai, S.; Dutt, A.; Joshi, N.N. Impact of intra-tumoral IL17A and IL32 gene expression on T-cell responses and lymph node status in breast cancer patients. J. Cancer Res. Clin. Oncol. 2017, 143, 1745–1756. [Google Scholar] [CrossRef] [PubMed]
- Armstrong, D.; Chang, C.-Y.; Lazarus, D.R.; Corry, D.; Kheradmand, F. Lung Cancer Heterogeneity in Modulation of Th17/IL17A Responses. Front. Oncol. 2019, 9, 1384. [Google Scholar] [CrossRef] [PubMed]
- Moore, N.; Moreno Gonzales, M.; Bonner, K.; Smith, B.; Park, W.; Stegall, M. Impact of CXCR4/CXCL12 Blockade on Normal Plasma Cells In Vivo. Am. J. Transplant. 2017, 17, 1663–1669. [Google Scholar] [CrossRef] [PubMed]
- Peled, A.; Klein, S.; Beider, K.; Burger, J.A.; Abraham, M. Role of CXCL12 and CXCR4 in the pathogenesis of hematological malignancies. Cytokine 2018, 109, 11–16. [Google Scholar] [CrossRef]
- Pitt, L.A.; Tikhonova, A.N.; Hu, H.; Trimarchi, T.; King, B.; Gong, Y.; Sanchez-Martin, M.; Tsirigos, A.; Littman, D.R.; Ferrando, A.A.; et al. CXCL12-Producing Vascular Endothelial Niches Control Acute T Cell Leukemia Maintenance. Cancer Cell 2015, 27, 755–768. [Google Scholar] [CrossRef] [PubMed]
- Kaverina, N.; Borovjagin, A.V.; Kadagidze, Z.; Baryshnikov, A.; Baryshnikova, M.; Malin, D.; Ghosh, D.; Shah, N.; Welch, D.R.; Gabikian, P.; et al. Astrocytes promote progression of breast cancer metastases to the brain via a KISS1-mediated autophagy. Autophagy 2017, 13, 1905–1923. [Google Scholar] [CrossRef] [PubMed]
- Kryczek, I.; Lange, A.; Mottram, P.; Alvarez, X.; Cheng, P.; Hogan, M.; Moons, L.; Wei, S.; Zou, L.; Machelon, V.; et al. CXCL12 and vascular endothelial growth factor synergistically induce neoangiogenesis in human ovarian cancers. Cancer Res. 2005, 65, 465–472. [Google Scholar] [PubMed]
- Samarendra, H.; Jones, K.; Petrinic, T.; Silva, M.A.; Reddy, S.; Soonawalla, Z.; Gordon-Weeks, A. A meta-analysis of CXCL12 expression for cancer prognosis. Br. J. Cancer 2017, 117, 124–135. [Google Scholar] [CrossRef]
- Möhle, R.; Bautz, F.; Rafii, S.; Moore, M.A.; Brugger, W.; Kanz, L. The chemokine receptor CXCR-4 is expressed on CD34+ hematopoietic progenitors and leukemic cells and mediates transendothelial migration induced by stromal cell-derived factor-1. Blood 1998, 91, 4523–4530. [Google Scholar] [CrossRef]
- Yazdani, Z.; Mousavi, Z.; Moradabadi, A.; Hassanshahi, G. Significance of CXCL12/CXCR4 Ligand/Receptor Axis in Various Aspects of Acute Myeloid Leukemia. Cancer Manag. Res. 2020, 12, 2155–2165. [Google Scholar] [CrossRef]
- Abe-Suzuki, S.; Kurata, M.; Abe, S.; Onishi, I.; Kirimura, S.; Nashimoto, M.; Murayama, T.; Hidaka, M.; Kitagawa, M. CXCL12+ stromal cells as bone marrow niche for CD34+ hematopoietic cells and their association with disease progression in myelodysplastic syndromes. Lab. Investig. 2014, 94, 1212–1223. [Google Scholar] [CrossRef] [PubMed]
- Fiegl, M.; Samudio, I.; Clise-Dwyer, K.; Burks, J.K.; Mnjoyan, Z.; Andreeff, M. CXCR4 expression and biologic activity in acute myeloid leukemia are dependent on oxygen partial pressure. Blood 2009, 113, 1504–1512. [Google Scholar] [CrossRef] [PubMed]
- Burger, J.A.; Peled, A. CXCR4 antagonists: Targeting the microenvironment in leukemia and other cancers. Leukemia 2009, 23, 43–52. [Google Scholar] [CrossRef] [PubMed]
- Kittang, A.O.; Hatfield, K.; Sand, K.; Reikvam, H.; Bruserud, Ø. The chemokine network in acute myelogenous leukemia: Molecular mechanisms involved in leukemogenesis and therapeutic implications. Curr. Top. Microbiol. Immunol. 2010, 341, 149–172. [Google Scholar] [CrossRef] [PubMed]
- Du, W.; Lu, C.; Zhu, X.; Hu, D.; Chen, X.; Li, J.; Liu, W.; Zhu, J.; He, Y.; Yao, J. Prognostic significance of CXCR4 expression in acute myeloid leukemia. Cancer Med. 2019, 8, 6595–6603. [Google Scholar] [CrossRef]
- Zhang, Y.; Saavedra, E.; Tang, R.; Gu, Y.; Lappin, P.; Trajkovic, D.; Liu, S.-H.; Smeal, T.; Fantin, V.; De Botton, S.; et al. Targeting primary acute myeloid leukemia with a new CXCR4 antagonist IgG1 antibody (PF-06747143). Sci. Rep. 2017, 7, 7305. [Google Scholar] [CrossRef]
- Shumilov, E.; Novak, U.; Jeker, B.; Mansouri Taleghani, B.; Bacher, U.; Pabst, T. Hematopoietic Stem Cell Mobilization With Plerixafor Is Safe and Effective in Poorly Mobilizing Acute Myeloid Leukemia Patients. Hemasphere 2019, 3, e176. [Google Scholar] [CrossRef] [PubMed]
- Walters, H.M.; Pan, N.; Lehman, T.J.; Adams, A.; Kalliolias, G.D.; Zhu, Y.S.; Santiago, F.; Nguyen, J.; Sitaras, L.; Cunningham-Rundles, S.; et al. The impact of disease activity and tumour necrosis factor-α inhibitor therapy on cytokine levels in juvenile idiopathic arthritis. Clin. Exp. Immunol. 2016, 184, 308–317. [Google Scholar] [CrossRef]
- Sack, U.; Burkhardt, U.; Borte, M.; Schädlich, H.; Berg, K.; Emmrich, F. Age-dependent levels of select immunological mediators in sera of healthy children. Clin. Diagn Lab. Immunol. 1998, 5, 28–32. [Google Scholar] [CrossRef] [PubMed]
- Eide, H.A.; Knudtsen, I.S.; Sandhu, V.; Løndalen, A.M.; Halvorsen, A.R.; Abravan, A.; Kure, E.H.; Bogsrud, T.V.; Brustugun, O.T.; Kyte, J.A.; et al. Serum cytokine profiles and metabolic tumor burden in patients with non-small cell lung cancer undergoing palliative thoracic radiation therapy. Adv. Radiat. Oncol. 2018, 3, 130–138. [Google Scholar] [CrossRef] [PubMed]
- Daver, N.; Schlenk, R.F.; Russell, N.H.; Levis, M.J. Targeting FLT3 mutations in AML: Review of current knowledge and evidence. Leukemia 2019, 33, 299–312. [Google Scholar] [CrossRef]
- Grunwald, M.R.; Levis, M.J. FLT3 inhibitors for acute myeloid leukemia: A review of their efficacy and mechanisms of resistance. Int. J. Hematol. 2013, 97, 683–694. [Google Scholar] [CrossRef] [PubMed]
- Dang, L.; Yen, K.; Attar, E.C. IDH mutations in cancer and progress toward development of targeted therapeutics. Ann. Oncol. 2016, 27, 599–608. [Google Scholar] [CrossRef] [PubMed]
- Chan, S.M.; Thomas, D.; Corces-Zimmerman, M.R.; Xavy, S.; Rastogi, S.; Hong, W.-J.; Zhao, F.; Medeiros, B.C.; Tyvoll, D.A.; Majeti, R. Isocitrate dehydrogenase 1 and 2 mutations induce BCL-2 dependence in acute myeloid leukemia. Nat. Med. 2015, 21, 178–184. [Google Scholar] [CrossRef]
- Othman, T.A.; Azenkot, T.; Moskoff, B.N.; Tenold, M.E.; Jonas, B.A. Venetoclax-based combinations for the treatment of newly diagnosed acute myeloid leukemia. Future Oncol. 2021, 17, 2989–3005. [Google Scholar] [CrossRef] [PubMed]
- Thompson, D.L.; Moore, D.C. Glasdegib: A Novel Hedgehog Pathway Inhibitor for Acute Myeloid Leukemia. J. Adv. Pract. Oncol. 2020, 11, 196–200. [Google Scholar] [CrossRef] [PubMed]
- Isidori, A.; Cerchione, C.; Daver, N.; DiNardo, C.; Garcia-Manero, G.; Konopleva, M.; Jabbour, E.; Ravandi, F.; Kadia, T.; Burguera, A.F.; et al. Immunotherapy in Acute Myeloid Leukemia: Where We Stand. Front. Oncol. 2021, 11, 656218. [Google Scholar] [CrossRef]
- Kumar, S.; Nagpal, R.; Kumar, A.; Ashraf, M.U.; Bae, Y.-S. Immunotherapeutic Potential of m6A-Modifiers and MicroRNAs in Controlling Acute Myeloid Leukaemia. Biomedicines 2021, 9, 690. [Google Scholar] [CrossRef] [PubMed]
- Carbonara, K.; Andonovski, M.; Coorssen, J.R. Proteomes Are of Proteoforms: Embracing the Complexity. Proteomes 2021, 9, 38. [Google Scholar] [CrossRef]
| Sample ID | Gender | Diagnosis Age | Risk Class | Diagnosis Type | % Blasts in BM |
|---|---|---|---|---|---|
| 1 | Female | 46.4 | 1 | 9871 Ac. myelomonocytic leuk. w abn. mar. eosinophils | 50 |
| 2 | Female | 35.3 | 1 | 9896 Acute myeloid leukemia, t(8;21)(q22;q22) | 60 |
| 3 | Male | 21.6 | 1 | 9896 Acute myeloid leukemia, t(8;21)(q22;q22) | 40 |
| 4 | Female | 67.3 | 1 | 9861 Acute myeloid leukemia | 75 |
| 5 | Female | 68.8 | 1 | 9861 Acute myeloid leukemia | n/a |
| 6 | Male | 16.8 | 1 | 9896 Acute myeloid leukemia, t(8;21)(q22;q22) | 26 |
| 7 | Female | 55.5 | 1 | 9873 Acute myeloid leukemia without maturation | 90 |
| 8 | Female | 44.8 | 1 | 9861 Acute myeloid leukemia | 65 |
| 9 | Female | 53.5 | 1 | 9874 Acute myeloid leukemia with maturation | 20 |
| 10 | Male | 72.8 | 1 | 9861 Acute myeloid leukemia | n/a |
| 11 | Female | 48.7 | 1 | 9891 Acute monocytic leukemia | 8 |
| 12 | Male | 76.9 | 2 | 9861 Acute myeloid leukemia | n/a |
| 13 | Female | 62.9 | 2 | 9874 Acute myeloid leukemia with maturation | 30 |
| 14 | Male | 56.5 | 2 | 9861 Acute myeloid leukemia | 23 |
| 15 | Female | 63.8 | 2 | 9861 Acute myeloid leukemia | 70 |
| 16 | Female | 78.1 | 2 | 9891 Acute monocytic leukemia | 60 |
| 17 | Female | 24.3 | 2 | 9861 Acute myeloid leukemia | 63 |
| 18 | Male | 67.3 | 2 | 9895 Acute myeloid leuk. with multilineage dysplasia | 37 |
| 19 | Female | 48.6 | 2 | 9873 Acute myeloid leukemia without maturation | 60 |
| 20 | Male | 72.6 | 2 | 9874 Acute myeloid leukemia with maturation | 33 |
| 21 | Male | 16.5 | 2 | 9891 Acute monocytic leukemia | 80 |
| 22 | Female | 62.9 | 2 | 9861 Acute myeloid leukemia | 22 |
| 23 | Female | 61.5 | 2 | 9891 Acute monocytic leukemia | 40 |
| 24 | Female | 66.7 | 2 | 9897 Acute myeloid leukemia, 11q23 abnormalities | 15 |
| 25 | Male | 57 | 2 | 9874 Acute myeloid leukemia with maturation | 42 |
| 26 | Female | 35.4 | 2 | 9920 Therapy-related acute myeloid leukemia, NOS | 95 |
| 27 | Female | 68.2 | 2 | del(9q)w23 | 60 |
| 28 | Female | 76.6 | 3 | 9873 Acute myeloid leukemia without maturation | 91 |
| 29 | Female | 54.3 | 3 | 9867 Acute myelomonocytic leukemia | 12 |
| 30 | Male | 28.6 | 3 | 9891 Acute monocytic leukemia | 45 |
| 31 | Male | 66.7 | 3 | 9873 Acute myeloid leukemia without maturation | 85 |
| 32 | Female | 52 | 3 | 9896 Acute myeloid leukemia, t(8;21)(q22;q22) | 91 |
| 33 | Female | 21.8 | 3 | 9873 Acute myeloid leukemia without maturation | 79 |
| 34 | Male | 44.6 | 3 | 9873 Acute myeloid leukemia without maturation | 73 |
| 35 | Female | 71.1 | 3 | 9873 Acute myeloid leukemia without maturation | 70 |
| 36 | Female | 39.7 | 3 | 9891 Acute monocytic leukemia | 40 |
| 37 | Male | 40.6 | 3 | 9861 Acute myeloid leukemia | 85 |
| 38 | Female | 59.4 | 3 | 9865 Acute myeloid leukemia with t(6;9)(p23;q34) DEK-NUP214 | 85 |
| 39 | Male | 77.7 | 3 | 9895 Acute myeloid leuk. with multilineage dysplasia | 16 |
| 40 | Male | 62.5 | 3 | 9727 Precursor cell lymphoblastic lymphoma, NOS | 91 |
| 41 | Female | 64.7 | 3 | 9920 Therapy-related acute myeloid leukemia, NOS | 65 |
| Group 1 vs. Group 2 | |||
| Gene Name | ANOVA p-Value | ↑ in Gr1 (Fold-Change) | ↑ in Gr2 (Fold-Change) |
| UBP7 | 0.001 | 1.4 | |
| HS105 | 0.004 | 2.0 | |
| DPYL2 | 0.006 | 1.2 | |
| SRSF2 | 0.007 | 1.1 | |
| FUS | 0.010 | 1.6 | |
| RTCB | 0.012 | 1.4 | |
| ANM1 | 0.017 | 1.3 | |
| PSA1 | 0.020 | 1.2 | |
| HNRL1 | 0.020 | 1.1 | |
| RAB5C | 0.022 | 1.3 | |
| SYVC | 0.030 | 1.3 | |
| 1433Z | 0.032 | 1.2 | |
| CAH1 | 0.035 | 5.2 | |
| SPTN1 | 0.035 | 2.3 | |
| LDHA | 0.043 | 1.3 | |
| FLNA | 0.045 | 1.5 | |
| ANXA6 | 0.046 | 1.3 | |
| G6PD | 0.048 | 1.8 | |
| Group 2 vs. Group 3 | |||
| Gene Name | ANOVA p-Value | ↑ in Gr2 (Fold-Change) | ↑ in Gr3 (Fold-Change) |
| DHX9 | 0.000 | 3.4 | |
| ATPB | 0.001 | 6.1 | |
| GSTK1 | 0.001 | 6.7 | |
| AHNK | 0.004 | 6.5 | |
| SYNC | 0.004 | 1.4 | |
| TCPA | 0.005 | 2.2 | |
| 1433G | 0.007 | 1.3 | |
| CH60 | 0.010 | 2.9 | |
| VATA | 0.010 | 2.3 | |
| PRKDC | 0.010 | 12.0 | |
| TAGL2 | 0.011 | 1.7 | |
| RPN1 | 0.012 | 1.9 | |
| TCPH | 0.013 | 1.7 | |
| UB2V1 | 0.013 | 1.4 | |
| PA2G4 | 0.016 | 1.1 | |
| ROA2 | 0.016 | 1.5 | |
| ATPA | 0.018 | 5.9 | |
| UBA1 | 0.020 | 1.6 | |
| FUBP1 | 0.020 | 1.9 | |
| TCPG | 0.020 | 1.6 | |
| TBB4B | 0.021 | 4.4 | |
| FUBP2 | 0.022 | 2.8 | |
| PNPH | 0.023 | 2.2 | |
| GSTO1 | 0.025 | 1.9 | |
| CAN1 | 0.026 | 1.5 | |
| HBB | 0.029 | 4.7 | |
| BAX | 0.029 | 1.9 | |
| EF2 | 0.030 | 1.4 | |
| DDX1 | 0.031 | 3.3 | |
| URP2 | 0.031 | 1.8 | |
| HBA | 0.032 | 5.4 | |
| ESTD | 0.032 | 1.4 | |
| HBD | 0.034 | 8.2 | |
| ACTZ | 0.038 | 1.9 | |
| TCPB | 0.039 | 1.6 | |
| CBX3 | 0.040 | 1.2 | |
| TIF1B | 0.043 | 2.8 | |
| PGM1 | 0.045 | 1.1 | |
| IF4A1 | 0.045 | 2.9 | |
| CPNS1 | 0.047 | 3.5 | |
| TCPE | 0.048 | 1.6 | |
| Group 1 vs. Group 3 | |||
| Gene Name | ANOVA p-Value | ↑ in Gr1 (Fold-Change) | ↑ in Gr3 (Fold-Change) |
| LA | 0.001 | 4.1 | |
| OTUB1 | 0.001 | 2.2 | |
| CNDP2 | 0.001 | 5.3 | |
| RAN | 0.001 | 2.5 | |
| HNRPC | 0.002 | 4.1 | |
| HNRPQ | 0.003 | 4.2 | |
| CH60 | 0.003 | 6.6 | |
| PRDX6 | 0.004 | 2.9 | |
| TBA1B | 0.005 | 3.7 | |
| TERA | 0.006 | 2.2 | |
| SET | 0.006 | 2.2 | |
| ROA2 | 0.006 | 2.8 | |
| CAPZB | 0.007 | 1.4 | |
| RCC2 | 0.007 | 2.0 | |
| ECHA | 0.007 | 4.2 | |
| ARPC4 | 0.007 | 1.3 | |
| PTPRC | 0.007 | 2.0 | |
| NONO | 0.008 | 2.5 | |
| THIO | 0.009 | 2.9 | |
| ILF3 | 0.011 | 2.0 | |
| VIME | 0.011 | 3.5 | |
| TALDO | 0.012 | 2.1 | |
| LDHA | 0.013 | 2.0 | |
| TCPH | 0.013 | 2.3 | |
| NUCL | 0.014 | 2.8 | |
| NAGK | 0.016 | 1.7 | |
| DHX9 | 0.016 | 4.1 | |
| PRDX4 | 0.016 | 1.0 | |
| TCP4 | 0.017 | 2.5 | |
| HS90A | 0.018 | 1.9 | |
| ROA1 | 0.018 | 2.5 | |
| LDHB | 0.019 | 2.6 | |
| EF1A3 | 0.020 | 2.4 | |
| FEN1 | 0.020 | 1.8 | |
| EF2 | 0.021 | 1.9 | |
| NPM | 0.024 | 2.6 | |
| F10A1 | 0.025 | 2.4 | |
| 1433Z | 0.026 | 1.6 | |
| TIF1B | 0.027 | 7.0 | |
| ESTD | 0.028 | 2.1 | |
| HNRH1 | 0.029 | 2.4 | |
| LC7L2 | 0.030 | 2.1 | |
| TCPZ | 0.030 | 1.7 | |
| GANAB | 0.030 | 2.3 | |
| PGAM1 | 0.031 | 1.3 | |
| ACTB | 0.031 | 1.7 | |
| PARP1 | 0.032 | 2.9 | |
| RUVB2 | 0.032 | 2.1 | |
| NPS3A | 0.034 | 1.2 | |
| NDKB | 0.034 | 2.2 | |
| RHOA | 0.035 | 1.6 | |
| SFPQ | 0.035 | 1.9 | |
| IF4A3 | 0.035 | 2.3 | |
| HNRPU | 0.037 | 2.4 | |
| DLDH | 0.039 | 2.6 | |
| RSSA | 0.041 | 3.6 | |
| ROA3 | 0.042 | 2.4 | |
| G3P | 0.042 | 2.8 | |
| RS3 | 0.042 | 4.5 | |
| FSCN1 | 0.044 | 1.0 | |
| RL40 | 0.046 | 1.2 | |
| PDIA3 | 0.049 | 1.7 | |
| HSP7C | 0.049 | 1.7 | |
| TSN | 0.050 | 1.2 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dowling, P.; Tierney, C.; Dunphy, K.; Miettinen, J.J.; Heckman, C.A.; Bazou, D.; O’Gorman, P. Identification of Protein Biomarker Signatures for Acute Myeloid Leukemia (AML) Using Both Nontargeted and Targeted Approaches. Proteomes 2021, 9, 42. https://doi.org/10.3390/proteomes9040042
Dowling P, Tierney C, Dunphy K, Miettinen JJ, Heckman CA, Bazou D, O’Gorman P. Identification of Protein Biomarker Signatures for Acute Myeloid Leukemia (AML) Using Both Nontargeted and Targeted Approaches. Proteomes. 2021; 9(4):42. https://doi.org/10.3390/proteomes9040042
Chicago/Turabian StyleDowling, Paul, Ciara Tierney, Katie Dunphy, Juho J. Miettinen, Caroline A. Heckman, Despina Bazou, and Peter O’Gorman. 2021. "Identification of Protein Biomarker Signatures for Acute Myeloid Leukemia (AML) Using Both Nontargeted and Targeted Approaches" Proteomes 9, no. 4: 42. https://doi.org/10.3390/proteomes9040042
APA StyleDowling, P., Tierney, C., Dunphy, K., Miettinen, J. J., Heckman, C. A., Bazou, D., & O’Gorman, P. (2021). Identification of Protein Biomarker Signatures for Acute Myeloid Leukemia (AML) Using Both Nontargeted and Targeted Approaches. Proteomes, 9(4), 42. https://doi.org/10.3390/proteomes9040042








