Role of the Cytokinin-Activated Type-B Response Regulators in Hormone Crosstalk
Abstract
1. Introduction
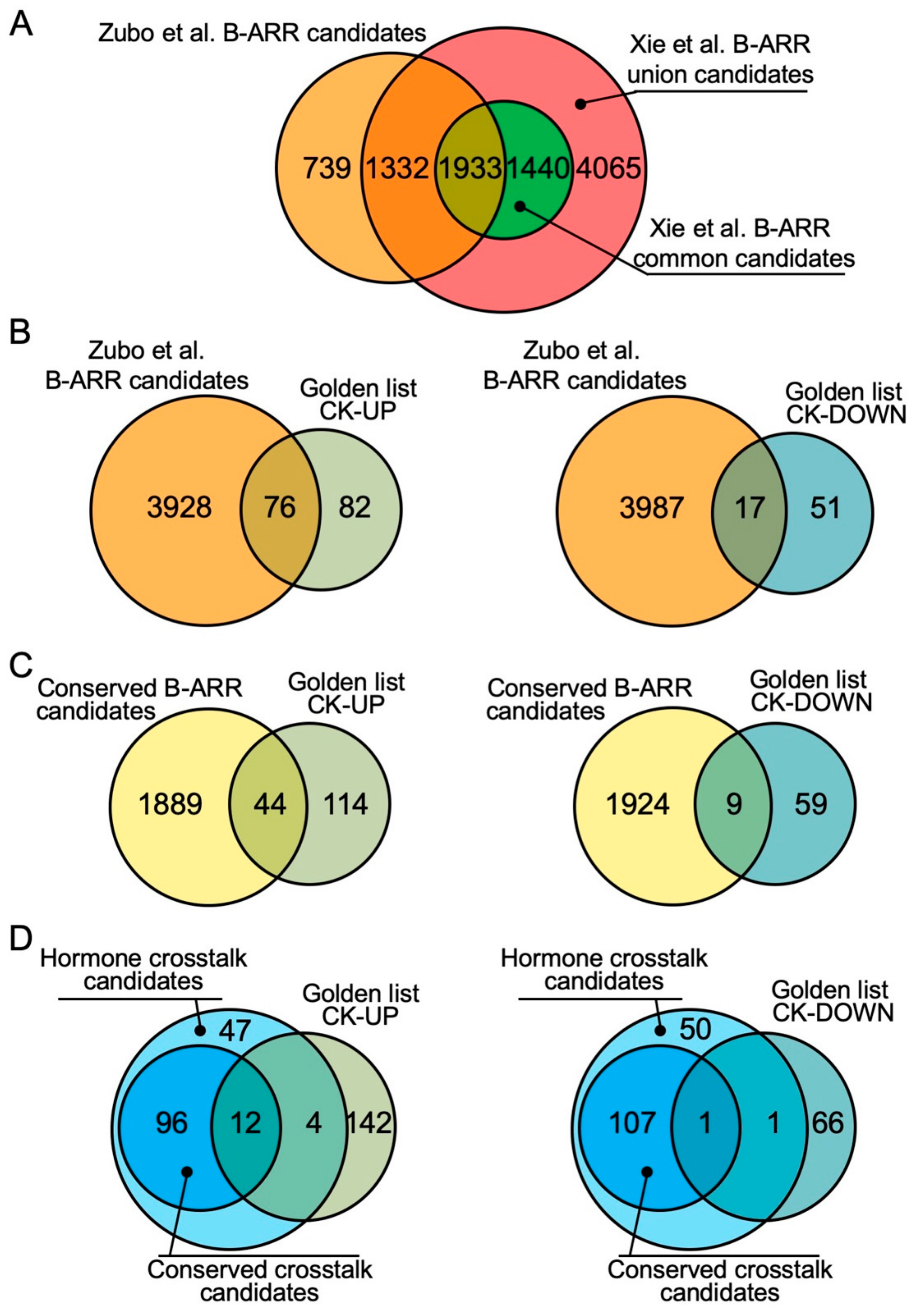
2. Characteristics of the type-B ARR ChIP-seq Studies
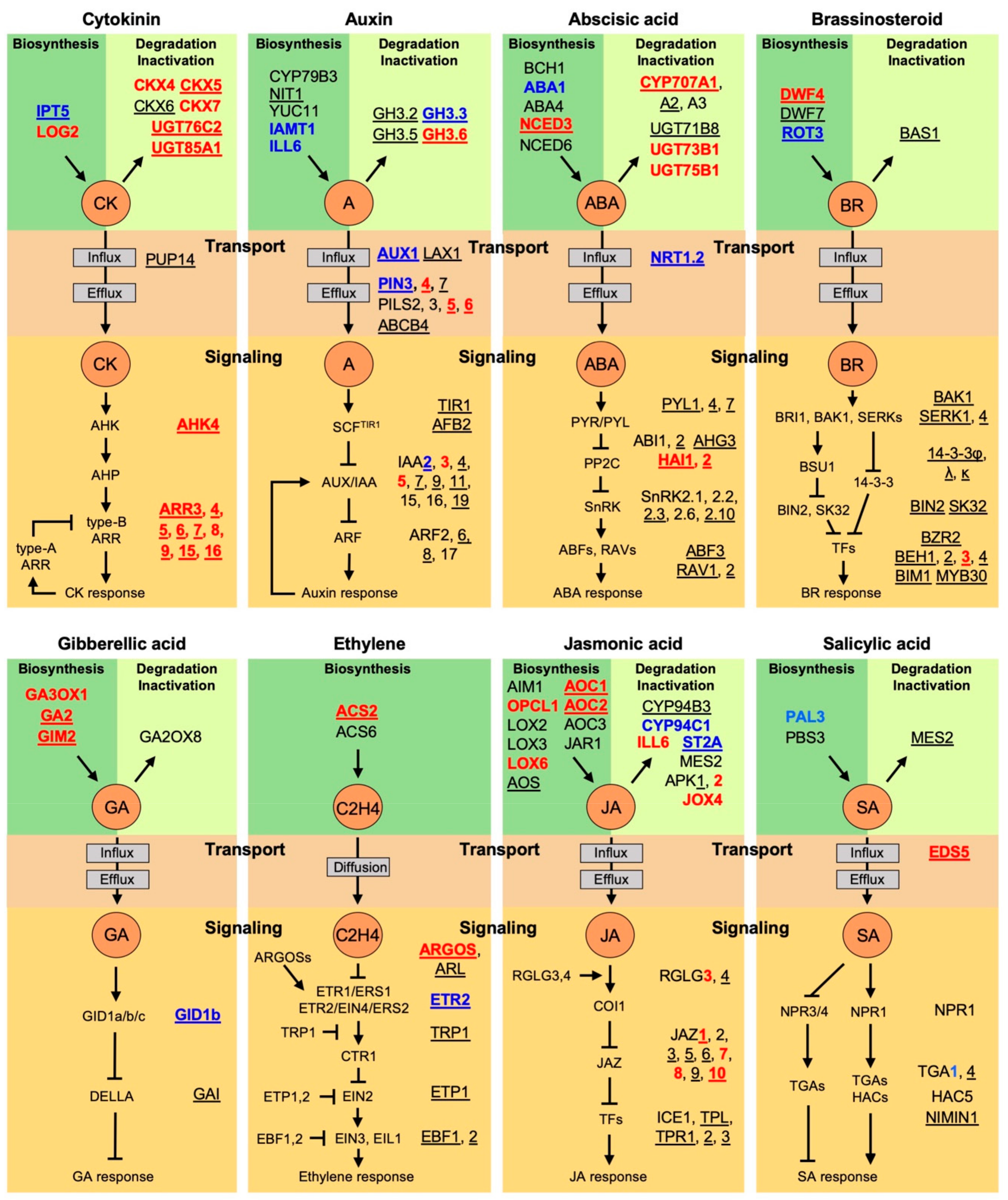
3. Hormone Crosstalk Revealed by ChIP-seq Studies
3.1. Cytokinin Negative Feedback Regulation
3.2. Cytokinin-Auxin Crosstalk
3.3. Cytokinin-Abscisic Acid Crosstalk
3.4. Cytokinin-Brassinosteroid Crosstalk
3.5. Cytokinin-Gibberellic Acid Crosstalk
3.6. Cytokinin-Ethylene Crosstalk
3.7. Cytokinin-Jasmonic Acid Crosstalk
3.8. Cytokinin-Salicylic Acid Crosstalk
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Davies, P.J. Plant Hormones: Biosynthesis, Signal Transduction, Action; Kluwer Academic Publishers: Dordrecht, The Netherlands, 2004. [Google Scholar]
- Hwang, I.; Sheen, J.; Müller, B. Cytokinin signaling networks. Annu. Rev. Plant Biol. 2012, 63, 353–380. [Google Scholar] [CrossRef]
- Kieber, J.J.; Schaller, G.E. Cytokinins. Arabidopsis Book 2014, 12, e0168. [Google Scholar] [CrossRef]
- Mok, D.W.; Mok, M.C. Cytokinin metabolism and action. Annu. Rev. Plant Physiol. Plant Mol. Biol. 2001, 52, 89–118. [Google Scholar] [CrossRef] [PubMed]
- Sakakibara, H. Cytokinins: Activity, biosynthesis, and translocation. Annu Rev Plant Biol 2006, 57, 431–449. [Google Scholar] [CrossRef] [PubMed]
- Choi, J.; Choi, D.; Lee, S.; Ryu, C.M.; Hwang, I. Cytokinins and plant immunity: Old foes or new friends? Trends Plant Sci. 2011, 16, 388–394. [Google Scholar] [CrossRef] [PubMed]
- Argueso, C.T.; Ferreira, F.J.; Epple, P.; To, J.P.; Hutchison, C.E.; Schaller, G.E.; Dangl, J.L.; Kieber, J.J. Two-component elements mediate interactions between cytokinin and salicylic acid in plant immunity. PLoS Genet. 2012, 8, e1002448. [Google Scholar] [CrossRef]
- O’Brien, J.A.; Benková, E. Cytokinin cross-talking during biotic and abiotic stress responses. Front. Plant Sci. 2013, 4, 451. [Google Scholar] [CrossRef]
- Schaller, G.E.; Kieber, J.J.; Shiu, S.-H. Two-component signaling elements and histidyl-aspartyl phosphorelays. Arabidopsis Book 2008, 6, e0112. [Google Scholar] [CrossRef]
- Brandstatter, I.; Kieber, J.J. Two genes with similarity to bacterial response regulators are rapidly and specifically induced by cytokinin in Arabidopsis. Plant Cell 1998, 10, 1009–1019. [Google Scholar] [CrossRef]
- To, J.P.; Haberer, G.; Ferreira, F.J.; Deruere, J.; Mason, M.G.; Schaller, G.E.; Alonso, J.M.; Ecker, J.R.; Kieber, J.J. Type-A Arabidopsis response regulators are partially redundant negative regulators of cytokinin signaling. Plant Cell 2004, 16, 658–671. [Google Scholar] [CrossRef]
- To, J.P.C.; Deruere, J.; Maxwell, B.B.; Morris, V.F.; Hutchison, C.E.; Ferreira, F.J.; Schaller, G.E.; Kieber, J.J. Cytokinin regulates type-A Arabidopsis response regulator activity and protein stability via two-component phosphorelay. Plant Cell 2007, 19, 3901–3914. [Google Scholar] [CrossRef] [PubMed]
- Heyl, A.; Riefler, M.; Romanov, G.A.; Schmülling, T. Properties, functions and evolution of cytokinin receptors. Eur. J. Cell Biol. 2012, 91, 246–256. [Google Scholar] [CrossRef] [PubMed]
- Pils, B.; Heyl, A. Unraveling the evolution of cytokinin signaling. Plant Physiol. 2009, 151, 782–791. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.J.; Ryu, H.; Hong, S.H.; Woo, H.R.; Lim, P.O.; Lee, I.C.; Sheen, J.; Nam, H.G.; Hwang, I. Cytokinin-mediated control of leaf longevity by AHK3 through phosphorylation of ARR2 in Arabidopsis. Proc. Natl. Acad. Sci. USA 2006, 103, 814–819. [Google Scholar] [CrossRef] [PubMed]
- Kurepa, J.; Li, Y.; Smalle, J.A. Cytokinin signaling stabilizes the response activator ARR1. Plant J. 2014, 78, 157–168. [Google Scholar] [CrossRef]
- Hass, C.; Lohrmann, J.; Albrecht, V.; Sweere, U.; Hummel, F.; Yoo, S.D.; Hwang, I.; Zhu, T.; Schafer, E.; Kudla, J.; et al. The response regulator 2 mediates ethylene signalling and hormone signal integration in Arabidopsis. EMBO J. 2004, 23, 3290–3302. [Google Scholar] [CrossRef]
- Imamura, A.; Hanaki, N.; Nakamura, A.; Suzuki, T.; Taniguchi, M.; Kiba, T.; Ueguchi, C.; Sugiyama, T.; Mizuno, T. Compilation and characterization of Arabidopsis thaliana response regulators implicated in His-Asp phosphorelay signal transduction. Plant Cell Physiol. 1999, 40, 733–742. [Google Scholar] [CrossRef]
- Imamura, A.; Hanaki, N.; Umeda, H.; Nakamura, A.; Suzuki, T.; Ueguchi, C.; Mizuno, T. Response regulators implicated in his-to-asp phosphotransfer signaling in Arabidopsis. Proc. Natl. Acad. Sci. USA 1998, 95, 2691–2696. [Google Scholar] [CrossRef]
- Suzuki, T.; Sakurai, K.; Ueguchi, C.; Mizuno, T. Two types of putative nuclear factors that physically interact with histidine-containing phosphotransfer (Hpt) domains, signaling mediators in His–to-Asp phosphorelay, in Arabidopsis thaliana. Plant Cell Physiol. 2001, 42, 37–45. [Google Scholar] [CrossRef]
- Hosoda, K.; Imamura, A.; Katoh, E.; Hatta, T.; Tachiki, M.; Yamada, H.; Mizuno, T.; Yamazaki, T. Molecular structure of the GARP family of plant Myb-related DNA binding motifs of the Arabidopsis response regulators. Plant Cell 2002, 14, 2015–2029. [Google Scholar] [CrossRef]
- Franco-Zorrilla, J.M.; Lopez-Vidriero, I.; Carrasco, J.L.; Godoy, M.; Vera, P.; Solano, R. DNA-binding specificities of plant transcription factors and their potential to define target genes. Proc. Natl. Acad. Sci. USA 2014, 111, 2367–2372. [Google Scholar] [CrossRef] [PubMed]
- Weirauch, M.T.; Yang, A.; Albu, M.; Cote, A.G.; Montenegro-Montero, A.; Drewe, P.; Najafabadi, H.S.; Lambert, S.A.; Mann, I.; Cook, K.; et al. Determination and inference of eukaryotic transcription factor sequence specificity. Cell 2014, 158, 1431–1443. [Google Scholar] [CrossRef] [PubMed]
- Zubo, Y.O.; Blakley, I.C.; Yamburenko, M.V.; Worthen, J.M.; Street, I.H.; Franco-Zorrilla, J.M.; Zhang, W.; Hill, K.; Raines, T.; Solano, R.; et al. Cytokinin induces genome-wide binding of the type-B response regulator ARR10 to regulate growth and development in Arabidopsis. Proc. Natl. Acad. Sci. USA 2017, 114, E5995–E6004. [Google Scholar] [CrossRef] [PubMed]
- Sakai, H.; Honma, T.; Aoyama, T.; Sato, S.; Kato, T.; Tabata, S.; Oka, A. ARR1, a transcription factor for genes immediately responsive to cytokinins. Science 2001, 294, 1519–1521. [Google Scholar] [CrossRef]
- Kim, K.; Ryu, H.; Cho, Y.H.; Scacchi, E.; Sabatini, S.; Hwang, I. Cytokinin-facilitated proteolysis of ARABIDOPSIS RESPONSE REGULATOR 2 attenuates signaling output in two-component circuitry. Plant J. 2012, 69, 934–945. [Google Scholar] [CrossRef]
- Kurepa, J.; Li, Y.; Perry, S.E.; Smalle, J.A. Ectopic expression of the phosphomimic mutant version of Arabidopsis response regulator 1 promotes a constitutive cytokinin response phenotype. BMC Plant Biol. 2014, 14, 28. [Google Scholar] [CrossRef]
- Mason, M.G.; Mathews, D.E.; Argyros, D.A.; Maxwell, B.B.; Kieber, J.J.; Alonso, J.M.; Ecker, J.R.; Schaller, G.E. Multiple type-B response regulators mediate cytokinin signal transduction in Arabidopsis. Plant Cell 2005, 17, 3007–3018. [Google Scholar] [CrossRef]
- Ishida, K.; Yamashino, T.; Yokoyama, A.; Mizuno, T. Three type-B response regulators, ARR1, ARR10, and ARR12, play essential but redundant roles in cytokinin signal transduction throughout the life cycle of Arabidopsis thaliana. Plant Cell Physiol. 2008, 49, 47–57. [Google Scholar] [CrossRef]
- Argyros, R.D.; Mathews, D.E.; Chiang, Y.-H.; Palmer, C.M.; Thibault, D.M.; Etheridge, N.; Argyros, D.A.; Mason, M.G.; Kieber, J.J.; Schaller, G.E. Type B response regulators of Arabidopsis play key roles in cytokinin signaling and plant development. Plant Cell 2008, 20, 2102–2116. [Google Scholar] [CrossRef]
- Yokoyama, A.; Yamashino, T.; Amano, Y.; Tajima, Y.; Imamura, A.; Sakakibara, H.; Mizuno, T. Type-B ARR transcription factors, ARR10 and ARR12, are implicated in cytokinin-mediated regulation of protoxylem differentiation in roots of Arabidopsis thaliana. Plant Cell Physiol. 2007, 48, 84–96. [Google Scholar] [CrossRef]
- Brenner, W.G.; Ramireddy, E.; Heyl, A.; Schmülling, T. Gene regulation by cytokinin in Arabidopsis. Front. Plant Sci. 2012, 3, 8. [Google Scholar] [CrossRef] [PubMed]
- Bhargava, A.; Clabaugh, I.; To, J.P.; Maxwell, B.B.; Chiang, Y.H.; Schaller, G.E.; Loraine, A.; Kieber, J.J. Identification of cytokinin-responsive genes using microarray meta-analysis and RNA-Seq in Arabidopsis. Plant Physiol. 2013, 162, 272–294. [Google Scholar] [CrossRef] [PubMed]
- Xie, M.; Chen, H.; Huang, L.; O’Neil, R.C.; Shokhirev, M.N.; Ecker, J.R. A B-ARR-mediated cytokinin transcriptional network directs hormone cross-regulation and shoot development. Nat. Commun. 2018, 9, 1604. [Google Scholar] [CrossRef] [PubMed]
- Grefen, C.; Harter, K. Plant two-component systems: Principles, functions, complexity and cross talk. Planta 2004, 219, 733–742. [Google Scholar] [CrossRef]
- El-Showk, S.; Ruonala, R.; Helariutta, Y. Crossing paths: Cytokinin signalling and crosstalk. Development 2013, 140, 1373–1383. [Google Scholar] [CrossRef]
- Rashotte, A.M.; Chae, H.S.; Maxwell, B.B.; Kieber, J.J. The interaction of cytokinin with other signals. Physiol. Plant. 2005, 123, 184–194. [Google Scholar] [CrossRef]
- Pacifici, E.; Polverari, L.; Sabatini, S. Plant hormone cross-talk: The pivot of root growth. J. Exp. Bot. 2015, 66, 1113–1121. [Google Scholar] [CrossRef]
- Jaillais, Y.; Chory, J. Unraveling the paradoxes of plant hormone signaling integration. Nat. Struct. Mol. Biol. 2010, 17, 642–645. [Google Scholar] [CrossRef]
- D’Agostino, I.B.; Deruere, J.; Kieber, J.J. Characterization of the response of the Arabidopsis response regulator gene family to cytokinin. Plant Physiol. 2000, 124, 1706–1717. [Google Scholar] [CrossRef]
- Rashotte, A.M.; Carson, S.D.; To, J.P.; Kieber, J.J. Expression profiling of cytokinin action in Arabidopsis. Plant Physiol. 2003, 132, 1998–2011. [Google Scholar] [CrossRef]
- Brenner, W.G.; Schmülling, T. Transcript profiling of cytokinin action in Arabidopsis roots and shoots discovers largely similar but also organ-specific responses. BMC Plant Biol. 2012, 12, 112. [Google Scholar] [CrossRef] [PubMed]
- Meng, W.J.; Cheng, Z.J.; Sang, Y.L.; Zhang, M.M.; Rong, X.F.; Wang, Z.W.; Tang, Y.Y.; Zhang, X.S. Type-B ARABIDOPSIS RESPONSE REGULATORs Specify the Shoot Stem Cell Niche by Dual Regulation of WUSCHEL. Plant Cell 2017, 29, 1357–1372. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Tian, C.; Zhang, C.; Shi, B.; Cao, X.; Zhang, T.Q.; Zhao, Z.; Wang, J.W.; Jiao, Y. Cytokinin Signaling Activates WUSCHEL Expression during Axillary Meristem Initiation. Plant Cell 2017, 29, 1373–1387. [Google Scholar] [CrossRef] [PubMed]
- Kiba, T.; Yamada, H.; Sato, S.; Kato, T.; Tabata, S.; Yamashino, T.; Mizuno, T. The type-A response regulator, ARR15, acts as a negative regulator in the cytokinin-mediated signal transduction in Arabidopsis thaliana. Plant Cell Physiol. 2003, 44, 868–874. [Google Scholar] [CrossRef] [PubMed]
- Taniguchi, M.; Kiba, T.; Sakakibara, H.; Ueguchi, C.; Mizuno, T.; Sugiyama, T. Expression of Arabidopsis response regulator homologs is induced by cytokinins and nitrate. FEBS Lett. 1998, 429, 259–262. [Google Scholar] [CrossRef]
- Taniguchi, M.; Sasaki, N.; Tsuge, T.; Aoyama, T.; Oka, A. ARR1 directly activates cytokinin response genes that encode proteins with diverse regulatory functions. Plant Cell Physiol 2007, 48, 263–277. [Google Scholar] [CrossRef]
- Niemann, M.C.E.; Weber, H.; Hluska, T.; Leonte, G.; Anderson, S.M.; Novak, O.; Senes, A.; Werner, T. The cytokinin oxidase/dehydrogenase CKX1 is a membrane-bound protein requiring homooligomerization in the endoplasmic reticulum for its cellular activity. Plant Physiol. 2018, 176, 2024–2039. [Google Scholar] [CrossRef]
- Schmulling, T.; Werner, T.; Riefler, M.; Krupkova, E.; Bartrina y Manns, I. Structure and function of cytokinin oxidase/dehydrogenase genes of maize, rice, Arabidopsis and other species. J. Plant Res. 2003, 116, 241–252. [Google Scholar] [CrossRef]
- Dortay, H.; Mehnert, N.; Bürkle, L.; Schmülling, T.; Heyl, A. Analysis of protein interactions within the cytokinin-signaling pathway of Arabidopsis thaliana. FEBS J. 2006, 273, 4631–4644. [Google Scholar] [CrossRef]
- Hwang, I.; Sheen, J. Two-component circuitry in Arabidopsis cytokinin signal transduction. Nature 2001, 413, 383–389. [Google Scholar] [CrossRef]
- Huang, X.; Zhang, X.; Gong, Z.; Yang, S.; Shi, Y. ABI4 represses the expression of type-A ARRs to inhibit seed germination in Arabidopsis. Plant J. 2017, 89, 354–365. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Li, L.; Ye, T.; Zhao, S.; Liu, Z.; Feng, Y.Q.; Wu, Y. Cytokinin antagonizes ABA suppression to seed germination of Arabidopsis by downregulating ABI5 expression. Plant J 2011, 68, 249–261. [Google Scholar] [CrossRef] [PubMed]
- Koshland, D.E., Jr.; Goldbeter, A.; Stock, J.B. Amplification and adaptation in regulatory and sensory systems. Science 1982, 217, 220–225. [Google Scholar] [CrossRef] [PubMed]
- Lan, G.; Sartori, P.; Neumann, S.; Sourjik, V.; Tu, Y. The energy-speed-accuracy tradeoff in sensory adaptation. Nat. Phys. 2012, 8, 422–428. [Google Scholar] [CrossRef]
- Miyawaki, K.; Matsumoto-Kitano, M.; Kakimoto, T. Expression of cytokinin biosynthetic isopentenyltransferase genes in Arabidopsis: Tissue specificity and regulation by auxin, cytokinin, and nitrate. Plant J. 2004, 37, 128–138. [Google Scholar] [CrossRef]
- Tokunaga, H.; Kojima, M.; Kuroha, T.; Ishida, T.; Sugimoto, K.; Kiba, T.; Sakakibara, H. Arabidopsis lonely guy (LOG) multiple mutants reveal a central role of the LOG-dependent pathway in cytokinin activation. Plant J. 2012, 69, 355–365. [Google Scholar] [CrossRef]
- Mähönen, A.P.; Higuchi, M.; Törmäkangas, K.; Miyawaki, K.; Pischke, M.S.; Sussman, M.R.; Helariutta, Y.; Kakimoto, T. Cytokinins regulate a bidirectional phosphorelay network in Arabidopsis. Curr. Biol. 2006, 16, 1116–1122. [Google Scholar] [CrossRef]
- Lavy, M.; Estelle, M. Mechanisms of auxin signaling. Development 2016, 143, 3226–3229. [Google Scholar] [CrossRef]
- Schaller, G.E.; Bishopp, A.; Kieber, J.J. The yin-yang of hormones: Cytokinin and auxin interactions in plant development. Plant Cell 2015, 27, 44–63. [Google Scholar] [CrossRef]
- Weijers, D.; Nemhauser, J.; Yang, Z. Auxin: Small molecule, big impact. J. Exp. Bot. 2018, 69, 133–136. [Google Scholar] [CrossRef]
- Woodward, A.W.; Bartel, B. Auxin: Regulation, action, and interaction. Ann. Bot. 2005, 95, 707–735. [Google Scholar] [CrossRef] [PubMed]
- Chandler, J.W.; Werr, W. Cytokinin-auxin crosstalk in cell type specification. Trends Plant Sci. 2015, 20, 291–300. [Google Scholar] [CrossRef] [PubMed]
- Moubayidin, L.; Di Mambro, R.; Sabatini, S. Cytokinin-auxin crosstalk. Trends Plant Sci. 2009, 14, 557–562. [Google Scholar] [CrossRef] [PubMed]
- Müller, B.; Sheen, J. Cytokinin and auxin interaction in root stem-cell specification during early embryogenesis. Nature 2008, 453, 1094–1097. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Andersen, S.U.; Ljung, K.; Dolezal, K.; Miotk, A.; Schultheiss, S.J.; Lohmann, J.U. Hormonal control of the shoot stem-cell niche. Nature 2010, 465, 1089–1092. [Google Scholar] [CrossRef]
- Dello Ioio, R.; Nakamura, K.; Moubayidin, L.; Perilli, S.; Taniguchi, M.; Morita, M.T.; Aoyama, T.; Costantino, P.; Sabatini, S. A genetic framework for the control of cell division and differentiation in the root meristem. Science 2008, 322, 1380–1384. [Google Scholar] [CrossRef] [PubMed]
- Moubayidin, L.; Perilli, S.; Dello Ioio, R.; Di Mambro, R.; Costantino, P.; Sabatini, S. The rate of cell differentiation controls the Arabidopsis root meristem growth phase. Curr. Biol. 2010, 20, 1138–1143. [Google Scholar] [CrossRef]
- Street, I.H.; Mathews, D.E.; Yamburenko, M.V.; Sorooshzadeh, A.; John, R.T.; Swarup, R.; Bennett, M.J.; Kieber, J.J.; Schaller, G.E. Cytokinin acts through the auxin influx carrier AUX1 to regulate cell elongation in the root. Development 2016, 143, 3982–3993. [Google Scholar] [CrossRef]
- Hill, K.; Mathews, D.E.; Kim, H.J.; Street, I.H.; Wildes, S.L.; Chiang, Y.H.; Mason, M.G.; Alonso, J.M.; Ecker, J.R.; Kieber, J.J.; et al. Functional characterization of type-B response regulators in the Arabidopsis cytokinin response. Plant Physiol. 2013, 162, 212–224. [Google Scholar] [CrossRef]
- Mason, M.G.; Li, J.; Mathews, D.E.; Kieber, J.J.; Schaller, G.E. Type-B response regulators display overlapping expression patterns in Arabidopsis. Plant Physiol. 2004, 135, 927–937. [Google Scholar] [CrossRef]
- Tajima, Y.; Imamura, A.; Kiba, T.; Amano, Y.; Yamashino, T.; Mizuno, T. Comparative studies on the type-B response regulators revealing their distinctive properties in the His-to-Asp phosphorelay signal transduction of Arabidopsis thaliana. Plant Cell Physiol. 2004, 45, 28–39. [Google Scholar] [CrossRef] [PubMed]
- Sauer, M.; Kleine-Vehn, J. PIN-FORMED and PIN-LIKES auxin transport facilitators. Development 2019, 146, dev168088. [Google Scholar] [CrossRef] [PubMed]
- Zwiewka, M.; Bilanovicova, V.; Seifu, Y.W.; Nodzynski, T. The nuts and bolts of PIN auxin efflux carriers. Front. Plant Sci. 2019, 10, 985. [Google Scholar] [CrossRef] [PubMed]
- Adamowski, M.; Friml, J. PIN-dependent auxin transport: Action, regulation, and evolution. Plant Cell 2015, 27, 20–32. [Google Scholar] [CrossRef]
- Nakashima, K.; Yamaguchi-Shinozaki, K. ABA signaling in stress-response and seed development. Plant Cell Rep. 2013, 32, 959–970. [Google Scholar] [CrossRef]
- Parent, B.; Hachez, C.; Redondo, E.; Simonneau, T.; Chaumont, F.; Tardieu, F. Drought and abscisic acid effects on aquaporin content translate into changes in hydraulic conductivity and leaf growth rate: A trans-scale approach. Plant Physiol. 2009, 149, 2000–2012. [Google Scholar] [CrossRef]
- Raghavendra, A.S.; Gonugunta, V.K.; Christmann, A.; Grill, E. ABA perception and signalling. Trends Plant Sci. 2010, 15, 395–401. [Google Scholar] [CrossRef]
- Jia, H.; Jiu, S.; Zhang, C.; Wang, C.; Tariq, P.; Liu, Z.; Wang, B.; Cui, L.; Fang, J. Abscisic acid and sucrose regulate tomato and strawberry fruit ripening through the abscisic acid-stress-ripening transcription factor. Plant Biotechnol. J. 2016, 14, 2045–2065. [Google Scholar] [CrossRef]
- Munemasa, S.; Hauser, F.; Park, J.; Waadt, R.; Brandt, B.; Schroeder, J.I. Mechanisms of abscisic acid-mediated control of stomatal aperture. Curr. Opin. Plant Biol. 2015, 28, 154–162. [Google Scholar] [CrossRef]
- Tuan, P.A.; Kumar, R.; Rehal, P.K.; Toora, P.K.; Ayele, B.T. Molecular mechanisms underlying abscisic acid/gibberellin balance in the control of seed dormancy and germination in cereals. Front. Plant Sci. 2018, 9, 668. [Google Scholar] [CrossRef]
- Tran, L.-S.P.; Urao, T.; Qin, F.; Maruyama, K.; Kakimoto, T.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Functional analysis of AHK1/ATHK1 and cytokinin receptor histidine kinases in response to abscisic acid, drought, and salt stress in Arabidopsis. Proc. Natl. Acad. Sci. USA 2007, 104, 20623–20628. [Google Scholar] [CrossRef]
- Huang, X.; Hou, L.; Meng, J.; You, H.; Li, Z.; Gong, Z.; Yang, S.; Shi, Y. The antagonistic action of abscisic acid and cytokinin signaling mediates drought stress response in Arabidopsis. Mol. Plant 2018, 11, 970–982. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, K.H.; Van Ha, C.; Nishiyama, R.; Watanabe, Y.; Leyva-Gonzalez, M.A.; Fujita, Y.; Tran, U.T.; Li, W.Q.; Tanaka, M.; Seki, M.; et al. Arabidopsis type B cytokinin response regulators ARR1, ARR10, and ARR12 negatively regulate plant responses to drought. Proc. Natl. Acad. Sci. USA 2016, 113, 3090–3095. [Google Scholar] [CrossRef] [PubMed]
- Clouse, S.D. Brassinosteroids. Arabidopsis Book 2011, 9, e0151. [Google Scholar] [CrossRef] [PubMed]
- Gruszka, D. The brassinosteroid signaling pathway-new key players and interconnections with other signaling networks crucial for plant development and stress tolerance. Int. J. Mol. Sci. 2013, 14, 8740–8774. [Google Scholar] [CrossRef] [PubMed]
- Sakurai, A. Brassinosteroid biosynthesis. Plant Physiol. Biochem. 1999, 37, 351–361. [Google Scholar] [CrossRef]
- Jibran, R.; Hunter, D.A.; Dijkwel, P.P. Hormonal regulation of leaf senescence through integration of developmental and stress signals. Plant Mol. Biol. 2013, 82, 547–561. [Google Scholar] [CrossRef]
- Hansen, M.; Chae, H.S.; Kieber, J.J. Regulation of ACS protein stability by cytokinin and brassinosteroid. Plant J. 2009, 57, 606–614. [Google Scholar] [CrossRef]
- Saini, S.; Sharma, I.; Pati, P.K. Versatile roles of brassinosteroid in plants in the context of its homoeostasis, signaling and crosstalks. Front. Plant Sci. 2015, 6, 950. [Google Scholar] [CrossRef]
- Yuan, L.B.; Peng, Z.H.; Zhi, T.T.; Zho, Z.; Liu, Y.; Zhu, Q.; Xiong, X.Y.; Ren, C.M. Brassinosteroid enhances cytokinin-induced anthocyanin biosynthesis in Arabidopsis seedlings. Biol. Plant 2015, 59, 99–105. [Google Scholar] [CrossRef]
- Hu, Y.; Bao, F.; Li, J. Promotive effect of brassinosteroids on cell division involves a distinct CycD3-induction pathway in Arabidopsis. Plant J. 2000, 24, 693–701. [Google Scholar] [CrossRef] [PubMed]
- Yuldashev, R.; Avalbaev, A.; Bezrukova, M.; Vysotskaya, L.; Khripach, V.; Shakirova, F. Cytokinin oxidase is involved in the regulation of cytokinin content by 24-epibrassinolide in wheat seedlings. Plant Physiol. Biochem. 2012, 55, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Daviere, J.M.; Achard, P. A Pivotal Role of DELLAs in Regulating Multiple Hormone Signals. Mol. Plant 2016, 9, 10–20. [Google Scholar] [CrossRef] [PubMed]
- Sun, T.P. Gibberellin metabolism, perception and signaling pathways in Arabidopsis. Arabidopsis Book 2008, 6, e0103. [Google Scholar] [CrossRef]
- Xu, H.; Liu, Q.; Yao, T.; Fu, X. Shedding light on integrative GA signaling. Curr. Opin. Plant Biol. 2014, 21, 89–95. [Google Scholar] [CrossRef]
- Ezura, H.; Harberd, N.P. Endogenous gibberellin levels influence in-vitro shoot regeneration in Arabidopsis thaliana (L.) Heynh. Planta 1995, 197, 301–305. [Google Scholar] [CrossRef]
- Greenboim-Wainberg, Y.; Maymon, I.; Borochov, R.; Alvarez, J.; Olszewski, N.; Ori, N.; Eshed, Y.; Weiss, D. Cross talk between gibberellin and cytokinin: The Arabidopsis GA response inhibitor SPINDLY plays a positive role in cytokinin signaling. Plant Cell 2005, 17, 92–102. [Google Scholar] [CrossRef]
- Marin-de la Rosa, N.; Pfeiffer, A.; Hill, K.; Locascio, A.; Bhalerao, R.P.; Miskolczi, P.; Gronlund, A.L.; Wanchoo-Kohli, A.; Thomas, S.G.; Bennett, M.J.; et al. Genome wide binding site analysis reveals transcriptional coactivation of cytokinin-responsive genes by DELLA proteins. PLoS Genet 2015, 11, e1005337. [Google Scholar] [CrossRef]
- Huang, S.; Cerny, R.E.; Qi, Y.; Bhat, D.; Aydt, C.M.; Hanson, D.D.; Malloy, K.P.; Ness, L.A. Transgenic studies on the involvement of cytokinin and gibberellin in male development. Plant Physiol. 2003, 131, 1270–1282. [Google Scholar] [CrossRef]
- Abeles, F.B.; Morgan, P.W.; Saltveit, M.E., Jr. Ethylene in Plant Biology, 2nd ed.; Academic Press: San Diego, CA, USA, 1992. [Google Scholar]
- McManus, M.T. The Plant Hormone Ethylene; Wiley-Blackwell: Chichester, UK, 2012. [Google Scholar]
- Schaller, G.E. Ethylene and the regulation of plant development. BMC Biol. 2012, 10, 9. [Google Scholar] [CrossRef]
- Rai, M.I.; Wang, X.; Thibault, D.M.; Kim, H.J.; Bombyk, M.M.; Binder, B.M.; Shakeel, S.N.; Schaller, G.E. The ARGOS gene family functions in a negative feedback loop to desensitize plants to ethylene. BMC Plant Biol. 2015, 15, 157. [Google Scholar] [CrossRef] [PubMed]
- Qin, H.; He, L.; Huang, R. The coordination of ethylene and other hormones in primary root development. Front. Plant Sci. 2019, 10, 874. [Google Scholar] [CrossRef] [PubMed]
- Zdarska, M.; Dobisova, T.; Gelova, Z.; Pernisova, M.; Dabravolski, S.; Hejatko, J. Illuminating light, cytokinin, and ethylene signalling crosstalk in plant development. J. Exp. Bot. 2015, 66, 4913–4931. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Street, I.H.; Aman, S.; Zubo, Y.; Ramzan, A.; Wang, X.; Shakeel, S.N.; Kieber, J.J.; Schaller, G.E. Ethylene inhibits cell proliferation of the Arabidopsis root meristem. Plant Physiol. 2015, 169, 338–350. [Google Scholar] [CrossRef]
- Růžička, K.; Ljung, K.; Vanneste, S.; Podhorská, R.; Beeckman, T.; Friml, J.; Benková, E. Ethylene regulates root growth through effects on auxin biosynthesis and transport-dependent auxin distribution. Plant Cell 2007, 19, 2197–2212. [Google Scholar] [CrossRef]
- Růžička, K.; Simásková, M.; Duclercq, J.; Petrásek, J.; Zazímalová, E.; Simon, S.; Friml, J.; Van Montagu, M.C.; Benková, E. Cytokinin regulates root meristem activity via modulation of the polar auxin transport. Proc. Natl. Acad. Sci. USA 2009, 106, 4284–4289. [Google Scholar] [CrossRef]
- Zdarska, M.; Cuyacot, A.R.; Tarr, P.T.; Yamoune, A.; Szmitkowska, A.; Hrdinova, V.; Gelova, Z.; Meyerowitz, E.M.; Hejatko, J. ETR1 integrates response to ethylene and cytokinins into a single multistep phosphorelay pathway to control root growth. Mol. Plant 2019, 12, 1338–1352. [Google Scholar] [CrossRef]
- Shi, J.; Drummond, B.J.; Wang, H.; Archibald, R.L.; Habben, J.E. Maize and Arabidopsis ARGOS proteins interact with ethylene receptor signaling complex, supporting a regulatory role for ARGOS in ethylene signal transduction. Plant Physiol. 2016, 171, 2783–2797. [Google Scholar] [CrossRef]
- Azhar, B.J.; Zulfiqar, A.; Shakeel, S.N.; Schaller, G.E. Amplification and adaptation in the ethylene signaling pathway. Small Methods 2019, 2019, 1900452. [Google Scholar] [CrossRef]
- Lin, Z.; Ho, C.W.; Grierson, D. AtTRP1 encodes a novel TPR protein that interacts with the ethylene receptor ERS1 and modulates development in Arabidopsis. J. Exp. Bot. 2009, 60, 3697–3714. [Google Scholar] [CrossRef]
- Qiao, H.; Chang, K.N.; Yazaki, J.; Ecker, J.R. Interplay between ethylene, ETP1/ETP2 F-box proteins, and degradation of EIN2 triggers ethylene responses in Arabidopsis. Genes Dev. 2009, 23, 512–521. [Google Scholar] [CrossRef] [PubMed]
- Guo, H.; Ecker, J.R. Plant responses to ethylene gas are mediated by SCF(EBF1/EBF2)-dependent proteolysis of EIN3 transcription factor. Cell 2003, 115, 667–677. [Google Scholar] [CrossRef]
- Potuschak, T.; Lechner, E.; Parmentier, Y.; Yanagisawa, S.; Grava, S.; Koncz, C.; Genschik, P. EIN3-dependent regulation of plant ethylene hormone signaling by two arabidopsis F box proteins: EBF1 and EBF2. Cell 2003, 115, 679–689. [Google Scholar] [CrossRef]
- Huang, H.; Liu, B.; Liu, L.; Song, S. Jasmonate action in plant growth and development. J. Exp. Bot. 2017, 68, 1349–1359. [Google Scholar] [CrossRef]
- Wasternack, C.; Hause, B. Jasmonates: Biosynthesis, perception, signal transduction and action in plant stress response, growth and development. An update to the 2007 review in Annals of Botany. Ann. Bot. 2013, 111, 1021–1058. [Google Scholar] [CrossRef]
- Yang, J.; Duan, G.; Li, C.; Liu, L.; Han, G.; Zhang, Y.; Wang, C. The crosstalks between jasmonic acid and other plant hormone signaling highlight the involvement of jasmonic acid as a core component in plant response to biotic and abiotic stresses. Front. Plant Sci. 2019, 10, 1349. [Google Scholar] [CrossRef]
- Jang, G.; Chang, S.H.; Um, T.Y.; Lee, S.; Kim, J.K.; Choi, Y.D. Antagonistic interaction between jasmonic acid and cytokinin in xylem development. Sci. Rep. 2017, 7, 10212. [Google Scholar] [CrossRef]
- Ueda, J.; Kato, j. Inhibition of cytokinin-induced plant growth by jasmonic acid and its methyl ester. Physiol. Plant 1982, 54, 249–252. [Google Scholar] [CrossRef]
- Zhu, X.; Chen, J.; Xie, Z.; Gao, J.; Ren, G.; Gao, S.; Zhou, X.; Kuai, B. Jasmonic acid promotes degreening via MYC2/3/4- and ANAC019/055/072-mediated regulation of major chlorophyll catabolic genes. Plant J. 2015, 84, 597–610. [Google Scholar] [CrossRef]
- Seo, S.; Sano, H.; Ohashi, Y. Jasmonic acid in wound signal transduction pathways. Physiol. Plant 1997, 101, 740–745. [Google Scholar] [CrossRef]
- Dermastia, M.; Ravnikar, M.; Vilhar, B.; Kovac, M. Increased level of cytokinin ribosides in jasmonic acid-treated potato (Solanum tuberosum) stem node cultures. Physiol. Plant 1994, 92, 241–246. [Google Scholar] [CrossRef]
- Sano, H.; Seo, S.; Koizumi, N.; Niki, T.; Iwamura, H.; Ohashi, Y. Regulation by cytokinins of endogenous levels of jasmonic and salicylic acids in mechanically wounded tobacco plants. Plant Cell Physiol. 1996, 37, 762–769. [Google Scholar] [CrossRef]
- Klessig, D.F.; Choi, H.W.; Dempsey, D.A. Systemic acquired resistance and salicylic acid: Past, present, and future. Mol. Plant Microbe. Interact. 2018, 31, 871–888. [Google Scholar] [CrossRef] [PubMed]
- Maruri-Lopez, I.; Aviles-Baltazar, N.Y.; Buchala, A.; Serrano, M. Intra and extracellular journey of the phytohormone salicylic acid. Front. Plant Sci. 2019, 10, 423. [Google Scholar] [CrossRef]
- Seyfferth, C.; Tsuda, K. Salicylic acid signal transduction: The initiation of biosynthesis, perception and transcriptional reprogramming. Front. Plant Sci. 2014, 5, 697. [Google Scholar] [CrossRef]
- Choi, J.; Huh, S.U.; Kojima, M.; Sakakibara, H.; Paek, K.H.; Hwang, I. The cytokinin-activated transcription factor ARR2 promotes plant immunity via TGA3/NPR1-dependent salicylic acid signaling in Arabidopsis. Dev. Cell 2010, 19, 284–295. [Google Scholar] [CrossRef]
- Naseem, M.; Philippi, N.; Hussain, A.; Wangorsch, G.; Ahmed, N.; Dandekar, T. Integrated systems view on networking by hormones in Arabidopsis immunity reveals multiple crosstalk for cytokinin. Plant Cell 2012, 24, 1793–1814. [Google Scholar] [CrossRef][Green Version]
- Serrano, M.; Wang, B.; Aryal, B.; Garcion, C.; Abou-Mansour, E.; Heck, S.; Geisler, M.; Mauch, F.; Nawrath, C.; Metraux, J.P. Export of salicylic acid from the chloroplast requires the multidrug and toxin extrusion-like transporter EDS5. Plant Physiol. 2013, 162, 1815–1821. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zubo, Y.O.; Schaller, G.E. Role of the Cytokinin-Activated Type-B Response Regulators in Hormone Crosstalk. Plants 2020, 9, 166. https://doi.org/10.3390/plants9020166
Zubo YO, Schaller GE. Role of the Cytokinin-Activated Type-B Response Regulators in Hormone Crosstalk. Plants. 2020; 9(2):166. https://doi.org/10.3390/plants9020166
Chicago/Turabian StyleZubo, Yan O., and G. Eric Schaller. 2020. "Role of the Cytokinin-Activated Type-B Response Regulators in Hormone Crosstalk" Plants 9, no. 2: 166. https://doi.org/10.3390/plants9020166
APA StyleZubo, Y. O., & Schaller, G. E. (2020). Role of the Cytokinin-Activated Type-B Response Regulators in Hormone Crosstalk. Plants, 9(2), 166. https://doi.org/10.3390/plants9020166




