Plant Roots Release Small Extracellular Vesicles with Antifungal Activity
Abstract
1. Introduction
2. Results and Discussion
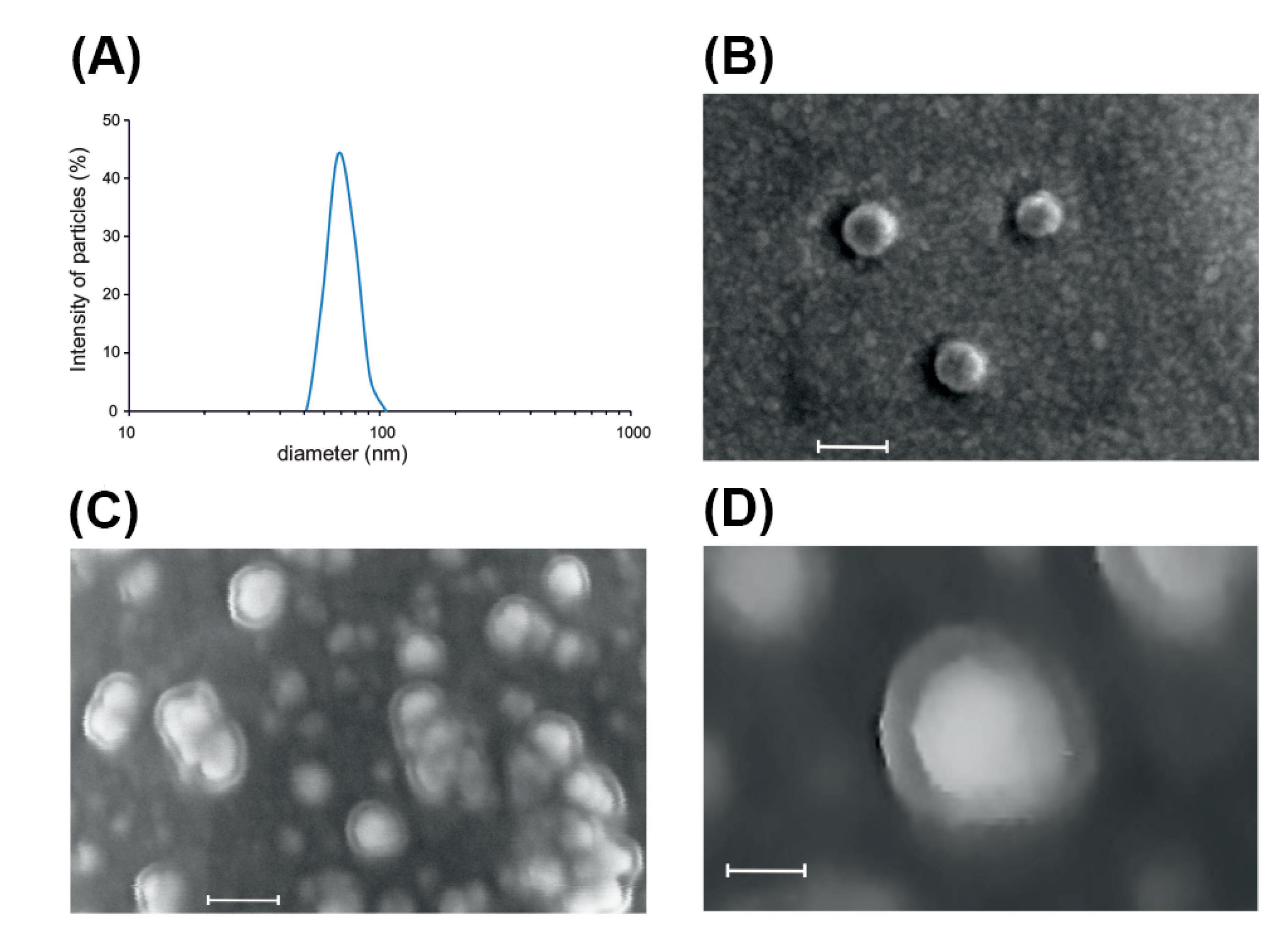
2.1. Isolation and Characterization of Root-Derived EVs
2.2. Shot-Gun Proteomics of Root-Derived EVs
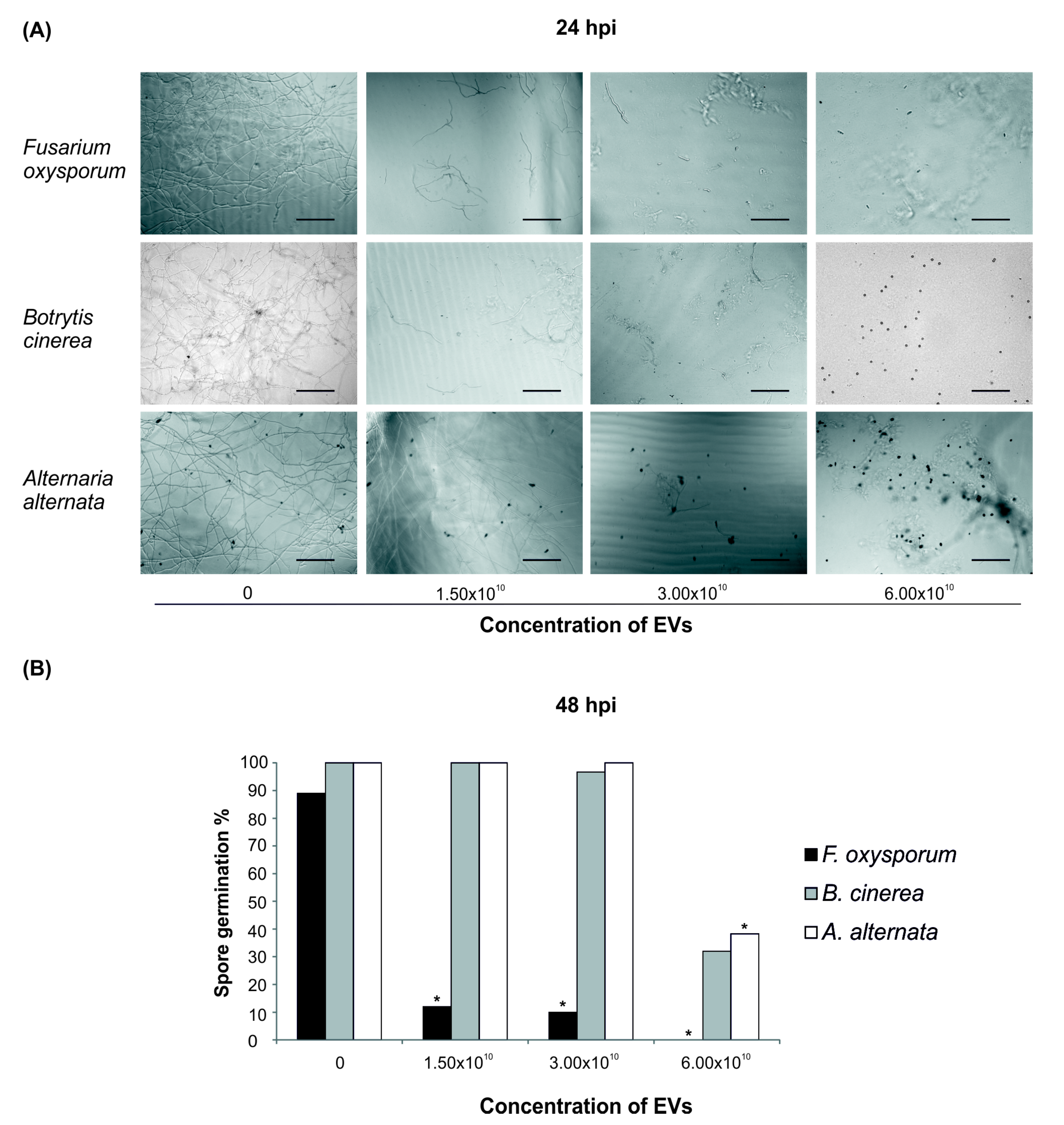
2.3. Bioactivity of Root-Derived EVs on Plant Pathogens
3. Conclusions
4. Materials and Methods
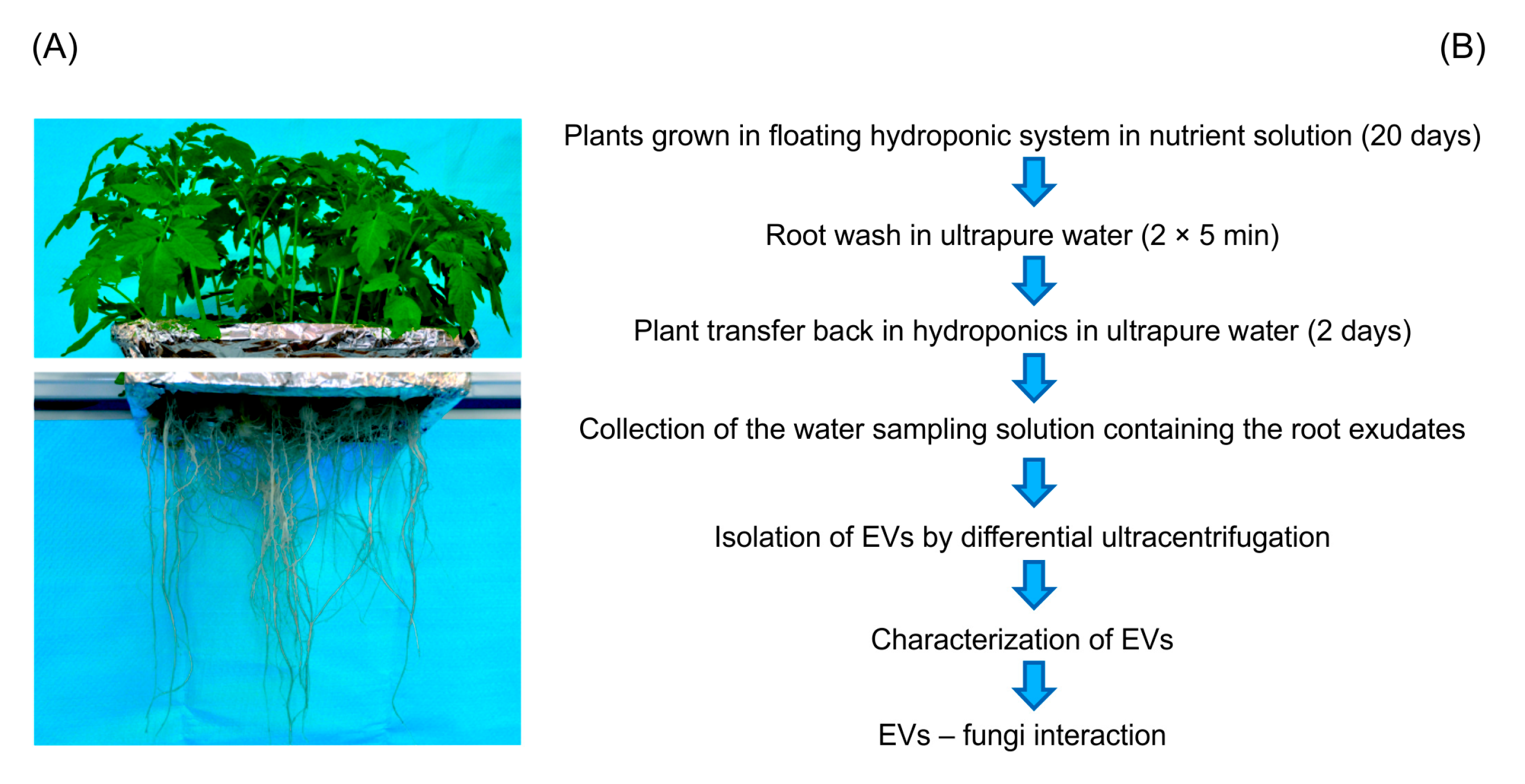
4.1. Plant Growth and Root Exudate Sampling Strategy
4.2. Isolation of Extracellular Vesicles
4.3. Protein Profiling by SDS-PAGE
4.4. Liquid Chromatography-Electrospray Ionization Mass Septrometry
4.5. Bioinformatics
4.6. Dynamic Light Scattering (DLS)
4.7. Microfluidic Resistive Pulse Sensing (MRPS)
4.8. Scanning Electron Microscopy (SEM)
4.9. Preparation of Fungal Spore Suspension
4.10. Vesicle/Spore Interaction Tests
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Pathan, M.; Fonseka, P.; Chitti, S.V.; Kang, T.; Sanwlani, R.; Van Deun, J.; Hendrix, A.; Mathivanan, S. Vesiclepedia 2019: A compendium of RNA, proteins, lipids and metabolites in extracellular vesicles. Nucleic Acids Res. 2019, 47, D516–D519. [Google Scholar] [CrossRef] [PubMed]
- Sedgwick, A.E.; D’Souza-Schorey, C. The biology of extracellular microvesicles. Traffic 2018, 19, 319–327. [Google Scholar] [CrossRef] [PubMed]
- Marchant, R.; Robards, A.W. Membrane Systems Associated with the Plasmalemma of Plant Cells. Ann. Bot. 1968, 32, 457–471. [Google Scholar] [CrossRef]
- Regente, M.; Pinedo, M.; San Clemente, H.; Balliau, T.; Jamet, E.; de la Canal, L. Plant extracellular vesicles are incorporated by a fungal pathogen and inhibit its growth. J. Exp. Bot. 2017, 68, 5485–5495. [Google Scholar] [CrossRef]
- Rutter, B.D.; Innes, R.W. Extracellular Vesicles Isolated from the Leaf Apoplast Carry Stress-Response Proteins. Plant Physiol. 2017, 173, 728–741. [Google Scholar] [CrossRef] [PubMed]
- Samuel, M.; Bleackley, M.; Anderson, M.; Mathivanan, S. Extracellular vesicles including exosomes in cross kingdom regulation: A viewpoint from plant-fungal interactions. Front. Plant Sci. 2015, 6. [Google Scholar] [CrossRef]
- Cai, Q.; Qiao, L.; Wang, M.; He, B.; Lin, F.M.; Palmquist, J.; Huang, S.D.; Jin, H. Plants send small RNAs in extracellular vesicles to fungal pathogen to silence virulence genes. Science 2018, 360, 1126–1129. [Google Scholar] [CrossRef]
- Rutter, B.D.; Innes, R.W. Extracellular vesicles as key mediators of plant-microbe interactions. Curr. Opin. Plant Biol. 2018, 44, 16–22. [Google Scholar] [CrossRef]
- Rybak, K.; Robatzek, S. Functions of Extracellular Vesicles in Immunity and Virulence. Plant Physiol. 2019, 179, 1236–1247. [Google Scholar] [CrossRef]
- Baldrich, P.; Rutter, B.D.; Karimi, H.Z.; Podicheti, R.; Meyers, B.C.; Innes, R.W. Plant Extracellular Vesicles Contain Diverse Small RNA Species and Are Enriched in 10- to 17-Nucleotide “Tiny” RNAs. Plant Cell 2019, 31, 315–324. [Google Scholar] [CrossRef]
- Mu, J.; Zhuang, X.; Wang, Q.; Jiang, H.; Deng, Z.B.; Wang, B.; Zhang, L.; Kakar, S.; Jun, Y.; Miller, D.; et al. Interspecies communication between plant and mouse gut host cells through edible plant derived exosome-like nanoparticles. Mol. Nutr. Food Res. 2014, 58, 1561–1573. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, X.; Deng, Z.B.; Mu, J.; Zhang, L.; Yan, J.; Miller, D.; Feng, W.; McClain, C.J.; Zhang, H.G. Ginger-derived nanoparticles protect against alcohol-induced liver damage. J. Extracell. Vesicles 2015, 4, 28713. [Google Scholar] [CrossRef] [PubMed]
- Pocsfalvi, G.; Turiák, L.; Ambrosone, A.; Del Gaudio, P.; Puska, G.; Fiume, I.; Silvestre, T.; Vékey, K. Protein biocargo of citrus fruit-derived vesicles reveals heterogeneous transport and extracellular vesicle populations. J. Plant Physiol. 2018, 229, 111–121. [Google Scholar] [CrossRef] [PubMed]
- Ju, S.; Mu, J.; Dokland, T.; Zhuang, X.; Wang, Q.; Jiang, H.; Xiang, X.; Deng, Z.B.; Wang, B.; Zhang, L.; et al. Grape exosome-like nanoparticles induce intestinal stem cells and protect mice from DSS-induced colitis. Mol. Ther. 2013, 21, 1345–1357. [Google Scholar] [CrossRef]
- Raimondi, M.T.; Albani, D.; Giordano, C. An Organ-On-A-Chip Engineered Platform to Study the Microbiota-Gut-Brain Axis in Neurodegeneration. Trends Mol. Med. 2019, 25, 737–740. [Google Scholar] [CrossRef]
- Akuma, P.; Okagu, O.D.; Udenigwe, C.C. Naturally Occurring Exosome Vesicles as Potential Delivery Vehicle for Bioactive Compounds. Front. Sustain. Food Syst. 2019, 3, 1–8. [Google Scholar] [CrossRef]
- Webber, J.; Clayton, A. How pure are your vesicles? J. Extracell. Vesicles 2013, 2, 19861. [Google Scholar] [CrossRef]
- Grabarek, A.D.; Weinbuch, D.; Jiskoot, W.; Hawe, A. Critical Evaluation of Microfluidic Resistive Pulse Sensing for Quantification and Sizing of Nanometer- and Micrometer-Sized Particles in Biopharmaceutical Products. J. Pharm. Sci. 2019, 108, 563–573. [Google Scholar] [CrossRef] [PubMed]
- 1Woith, E.; Melzig, M.F. Extracellular vesicles from fresh and dried plants—Simultaneous purification and visualization using gel electrophoresis. Int. J. Mol. Sci. 2019, 20, 357. [Google Scholar] [CrossRef]
- Kim, D.K.; Lee, J.; Kim, S.R.; Choi, D.-S.; Yoon, Y.J.; Kim, J.H.; Go, G.; Nhung, D.; Hong, K.; Jang, S.C.; et al. EVpedia: A community web portal for extracellular vesicles research. Bioinformatics 2015, 31, 933–939. [Google Scholar] [CrossRef]
- Lo Cicero, A.; Stahl, P.D.; Raposo, G. Extracellular vesicles shuffling intercellular messages: For good or for bad. Curr. Opin. Cell Biol. 2015, 35, 69–77. [Google Scholar] [CrossRef] [PubMed]
- Hurwitz, S.N.; Rider, M.A.; Bundy, J.L.; Liu, X.; Singh, R.K.; Meckes, D.G. Proteomic profiling of NCI-60 extracellular vesicles uncovers common protein cargo and cancer type-specific biomarkers. Oncotarget 2016, 7, 86999–87015. [Google Scholar] [CrossRef] [PubMed]
- Oshikawa, S.; Sonoda, H.; Ikeda, M. Aquaporins in Urinary Extracellular Vesicles (Exosomes). Int. J. Mol. Sci. 2016, 17, 957. [Google Scholar] [CrossRef] [PubMed]
- Foolad, M.R.; Merk, H.L.; Ashrafi, H. Genetics, Genomics and Breeding of Late Blight and Early Blight Resistance in Tomato. Crit. Rev. Plant Sci. 2008, 27, 75–107. [Google Scholar] [CrossRef]
- Faino, L.; Carli, P.; Testa, A.; Cristinzio, G.; Frusciante, L.; Ercolano, M.R. Potato R1 resistance gene confers resistance against Phytophthora infestans in transgenic tomato plants. Eur. J. Plant Pathol. 2010, 128, 233–241. [Google Scholar] [CrossRef]
- Sedbrook, J.C.; Carroll, K.L.; Hung, K.F.; Masson, P.H.; Somerville, C.R. The Arabidopsis SKU5 Gene Encodes an Extracellular Glycosyl Phosphatidylinositol—Anchored Glycoprotein Involved in Directional Root Growth. Plant Cell 2002, 14, 1635–1648. [Google Scholar] [CrossRef]
- Wei, Y.; Zhang, Z.; Andersen, C.H.; Schmelzer, E.; Gregersen, P.L.; Collinge, D.B.; Smedegaard-Petersen, V.; Thordal-Christensen, H. An epidermis/papilla-specific oxalate oxidase-like protein in the defence response of barley attacked by the powdery mildew fungus. Plant Mol. Biol. 1998, 36, 101–112. [Google Scholar] [CrossRef]
- Liu, Q.; Yang, J.; Yan, S.; Zhang, S.; Zhao, J.; Wang, W.; Yang, T.; Wang, X.; Mao, X.; Dong, J.; et al. The germin-like protein OsGLP2-1 enhances resistance to fungal blast and bacterial blight in rice. Plant Mol. Biol. 2016, 92, 411–423. [Google Scholar] [CrossRef]
- Kumar, M.; Brar, A.; Yadav, M.; Chawade, A.; Vivekanand, V.; Pareek, N. Chitinases-Potential Candidates for Enhanced Plant Resistance towards Fungal Pathogens. Agriculture 2018, 8, 88. [Google Scholar] [CrossRef]
- Balasubramanian, V.; Vashisht, D.; Cletus, J.; Sakthivel, N. Plant β-1,3-glucanases: Their biological functions and transgenic expression against phytopathogenic fungi. Biotechnol. Lett. 2012, 34, 1983–1990. [Google Scholar] [CrossRef]
- Li, C.; Tian, Q.; u Rahman, M.K.; Wu, F. Effect of anti-fungal compound phytosphingosine in wheat root exudates on the rhizosphere soil microbial community of watermelon. Plant Soil 2020, 456, 223–240. [Google Scholar] [CrossRef]
- Lombardi, N.; Vitale, S.; Turrà, D.; Reverberi, M.; Fanelli, C.; Vinale, F.; Marra, R.; Ruocco, M.; Pascale, A.; d’Errico, G.; et al. Root Exudates of Stressed Plants Stimulate and Attract Trichoderma Soil Fungi. Mol. Plant Microbe Interact. 2018, 31, 982–994. [Google Scholar] [CrossRef] [PubMed]
- Cletus, J.; Balasubramanian, V.; Vashisht, D.; Sakthivel, N. Transgenic expression of plant chitinases to enhance disease resistance. Biotechnol. Lett. 2013, 35, 1719–1732. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Ouyang, B.; Zhang, J.; Wang, T.; Lu, C.; Han, Q.; Zhao, S.; Ye, Z.; Li, H. Genomic organization, phylogenetic comparison and expression profiles of annexin gene family in tomato (Solanum lycopersicum). Gene 2012, 499, 14–24. [Google Scholar] [CrossRef] [PubMed]
- Fernández, M.B.; Pagano, M.R.; Daleo, G.R.; Guevara, M.G. Hydrophobic proteins secreted into the apoplast may contribute to resistance against Phytophthora infestans in potato. Plant Physiol. Biochem. 2012, 60, 59–66. [Google Scholar] [CrossRef] [PubMed]
- Peng, H.; Yang, T.; Jurick, W.M., II. Calmodulin Gene Expression in Response to Mechanical Wounding and Botrytis cinerea Infection in Tomato Fruit. Plants 2014, 3, 427–441. [Google Scholar] [CrossRef]
- La Camera, S.; Geoffroy, P.; Samaha, H.; Ndiaye, A.; Rahim, G.; Legrand, M.; Heitz, T. A pathogen-inducible patatin-like lipid acyl hydrolase facilitates fungal and bacterial host colonization in Arabidopsis. Plant J. 2005, 44, 810–825. [Google Scholar] [CrossRef]
- De Palma, M.; Salzano, M.; Villano, C.; Aversano, R.; Lorito, M.; Ruocco, M.; Docimo, T.; Piccinelli, A.L.; D’Agostino, N.; Tucci, M. Transcriptome reprogramming, epigenetic modifications and alternative splicing orchestrate the tomato root response to the beneficial fungus Trichoderma harzianum. Hortic. Res. 2019, 6. [Google Scholar] [CrossRef]
- Livshits, M.A.; Khomyakova, E.; Evtushenko, E.G.; Lazarev, V.N.; Kulemin, N.A.; Semina, S.E.; Generozov, E.V.; Govorun, V.M. Isolation of exosomes by differential centrifugation: Theoretical analysis of a commonly used protocol. Sci. Rep. 2015, 5, 17319. [Google Scholar] [CrossRef]
- Cox, J.; Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 2008, 26, 1367–1372. [Google Scholar] [CrossRef]
- Van Deun, J.; Hendrix, A. Is your article EV-TRACKed? J. Extracell. Vesicles 2017, 6, 1379835. [Google Scholar] [CrossRef] [PubMed]
- Kalra, H.; Simpson, R.J.; Ji, H.; Aikawa, E.; Altevogt, P.; Askenase, P.; Bond, V.C.; Borràs, F.E.; Breakefield, X.; Budnik, V.; et al. Vesiclepedia: A compendium for extracellular vesicles with continuous community annotation. PLoS Biol. 2012, 10, e1001450. [Google Scholar] [CrossRef] [PubMed]
- Huerta-Cepas, J.; Szklarczyk, D.; Forslund, K.; Cook, H.; Heller, D.; Walter, M.C.; Rattei, T.; Mende, D.R.; Sunagawa, S.; Kuhn, M.; et al. eggNOG 4.5: A hierarchical orthology framework with improved functional annotations for eukaryotic, prokaryotic and viral sequences. Nucleic Acids Res. 2016, 44, D286–D293. [Google Scholar] [CrossRef] [PubMed]


| No. | Protein GI | Name |
|---|---|---|
| 1 | 1022943236 | H(+)-ATPase 4 |
| 2 | 1104626054 | hypersensitive-induced response protein 1 |
| 3 | 726965338 | probable aquaporin PIP2-1 |
| 4 | 350534408 | Plasma membrane intrinsic protein 1 |
| 5 | 1104594347 | 12S seed storage protein Cruciferin D (CRD) |
| 6 | 723685072 | senescence-induced receptor-like serine/threonine-protein kinase |
| 7 | 723688012 | high-affinity nitrate transporter 3.2-like |
| 8 | 1104611050 | patatin-like protein 2 |
| 9 | 921274020 | vacuolar H+-ATPase A1 subunit isoform |
| 10 | 460370435 | V-type proton ATPase subunit B 2 |
| 11 | 723754262 | ferric reduction oxidase 4 |
| 12 | 460376506 | phosphate transporter |
| 13 | 460404122 | uncharacterized protein LOC101257107 |
| 14 | 460401047 | polyphenol oxidase E, chloroplastic-like |
| 15 | 1104638510 | NDR1/HIN1-Like protein 3-like isoform X2 |
| 16 | 927928759 | aquaporin PIP2-7-like |
| 17 | 350537435 | glucan endo-1,3-beta-glucosidase B precursor |
| 18 | 1114439811 | probable aquaporin PIP-type pTOM75 |
| 19 | 10263446685 | calmodulin 1 |
| 20 | 902763248 | putative PIP-type aquaporin |
| Protein GI | Name |
|---|---|
| 460406368 | endochitinase 4 |
| 1104611050 | patatin-like protein 2 |
| 350537435 | glucan endo-1,3-beta-glucosidase B precursor |
| 1104626054 | hypersensitive-induced response protein 1 |
| 1026346657 | calmodulin 5 |
| 460408499 | trypsin inhibitor 1-like |
| 460403785 | probable linoleate 9S-lipoxygenase 5 |
| 350538805 | annexin p34 |
| 460370493 | lysM domain-containing GPI-anchored protein 2 |
| 124192 | ethylene-responsive proteinase inhibitor 1 |
| 723690477 | putative late blight resistance protein homolog R1A-10 |
| 460382464 | putative late blight resistance protein homolog R1A-3 |
| 1104638510 | NDR1/HIN1-like protein 3-like isoform X2 |
| 460409616 | putative LRR receptor-like serine/threonine protein kinase At4g00960 |
| 723690472 | putative late blight resistance protein homolog R1A-3 |
| 544011 | basic 30 kDa endochitinase |
| 460404505 | germin-like protein subfamily 1 member 19 |
| 460382045 | CASP-like protein PIMP1 |
| 460408543 | probable LRR receptor-like serine/threonine-protein kinase At1g06840 |
| 723687659 | hypersensitive-induced response protein 1 |
| 460400431 | monocopper oxidase-like protein SKU5 |
| 350537023 | wound/stress protein precursor |
| 1143632856 | MRLK1 serine/threonine protein kinase, partial |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
De Palma, M.; Ambrosone, A.; Leone, A.; Del Gaudio, P.; Ruocco, M.; Turiák, L.; Bokka, R.; Fiume, I.; Tucci, M.; Pocsfalvi, G. Plant Roots Release Small Extracellular Vesicles with Antifungal Activity. Plants 2020, 9, 1777. https://doi.org/10.3390/plants9121777
De Palma M, Ambrosone A, Leone A, Del Gaudio P, Ruocco M, Turiák L, Bokka R, Fiume I, Tucci M, Pocsfalvi G. Plant Roots Release Small Extracellular Vesicles with Antifungal Activity. Plants. 2020; 9(12):1777. https://doi.org/10.3390/plants9121777
Chicago/Turabian StyleDe Palma, Monica, Alfredo Ambrosone, Antonietta Leone, Pasquale Del Gaudio, Michelina Ruocco, Lilla Turiák, Ramesh Bokka, Immacolata Fiume, Marina Tucci, and Gabriella Pocsfalvi. 2020. "Plant Roots Release Small Extracellular Vesicles with Antifungal Activity" Plants 9, no. 12: 1777. https://doi.org/10.3390/plants9121777
APA StyleDe Palma, M., Ambrosone, A., Leone, A., Del Gaudio, P., Ruocco, M., Turiák, L., Bokka, R., Fiume, I., Tucci, M., & Pocsfalvi, G. (2020). Plant Roots Release Small Extracellular Vesicles with Antifungal Activity. Plants, 9(12), 1777. https://doi.org/10.3390/plants9121777







