Salt-Induced Damage is Alleviated by Short-Term Pre-Cold Treatment in Bermudagrass (Cynodon dactylon)
Abstract
:1. Introduction
2. Results
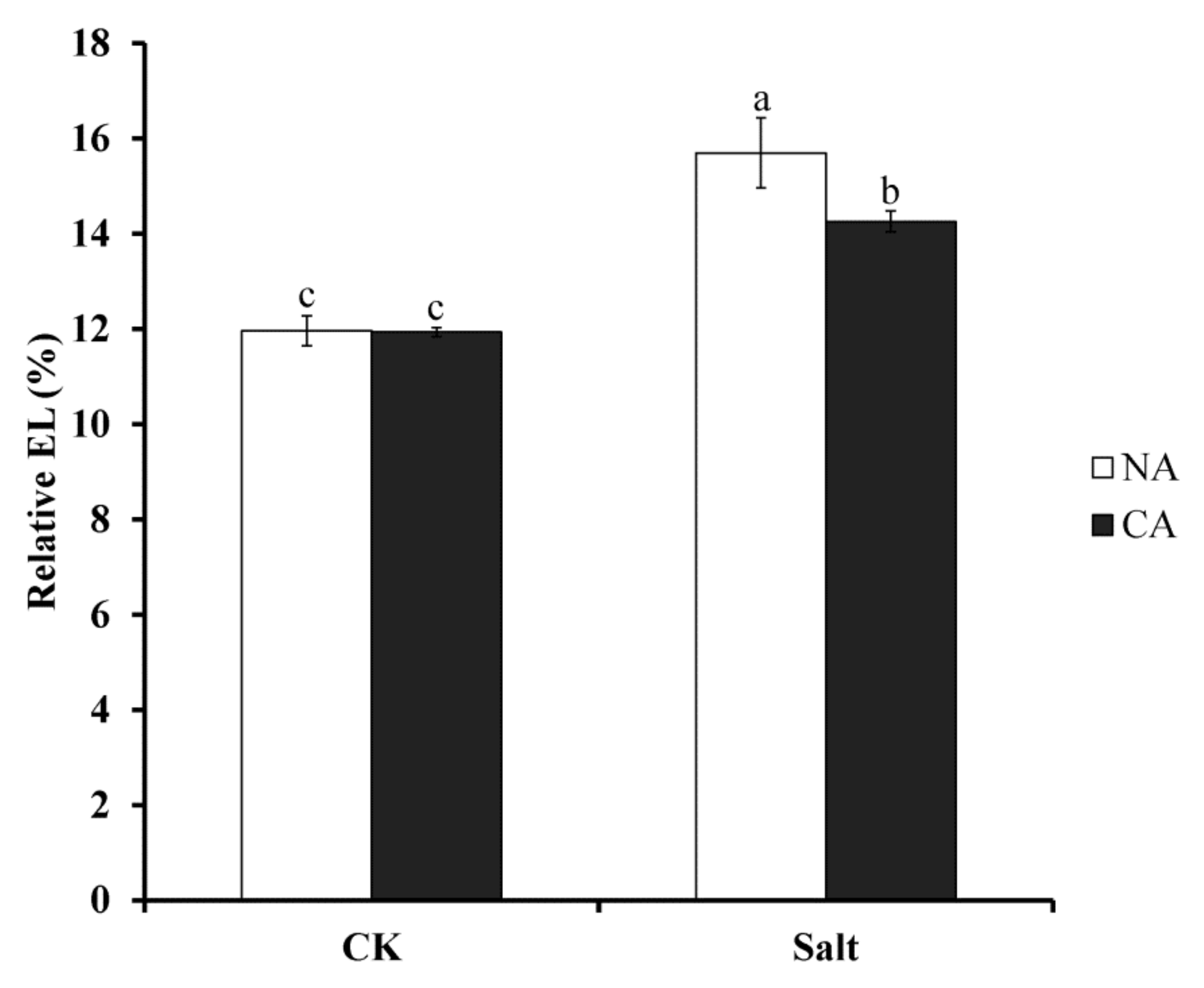
2.1. The Effect of Short-Term Pre-Cold Treatment on Cell Membrane Stability in Bermudagrass under Salt Stress
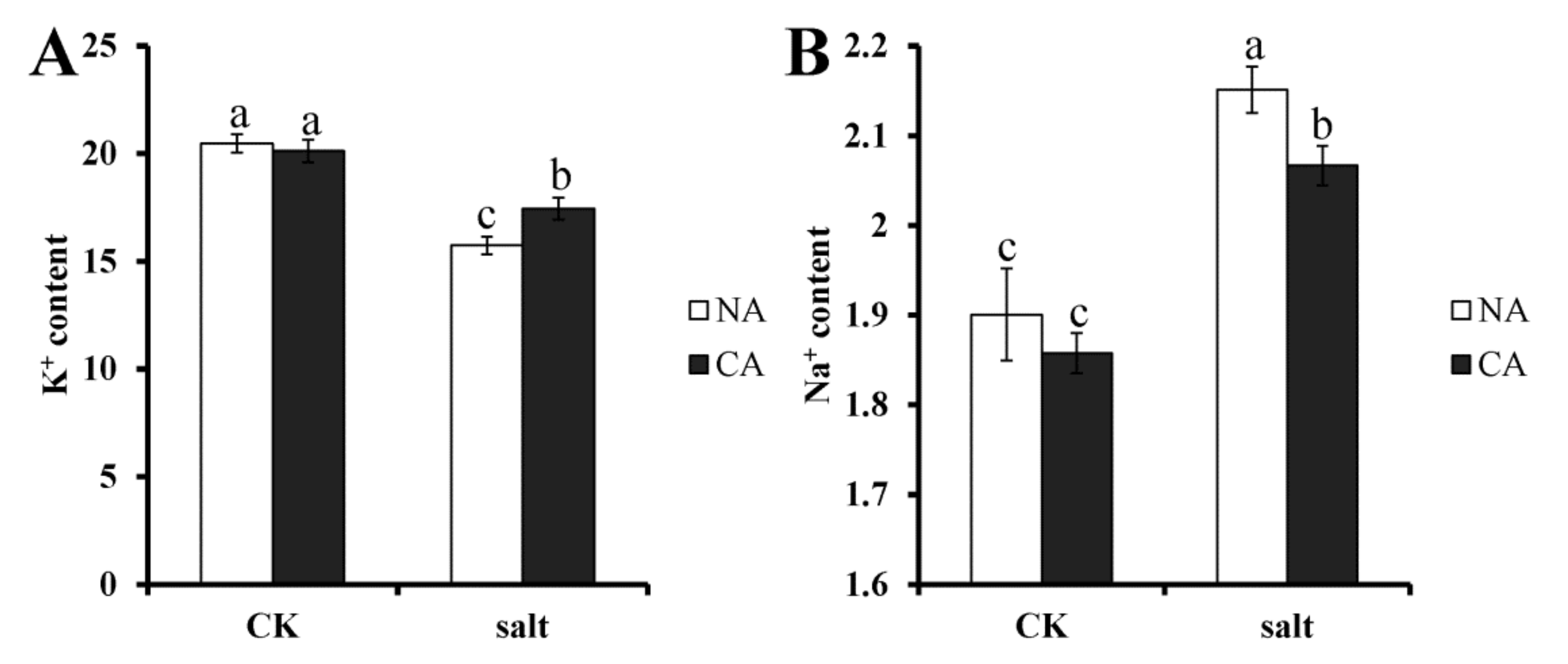
2.2. The Effect of Short-Term Pre-Cold Treatment on Ion Homeostasis in Bermudagrass under Salt Stress
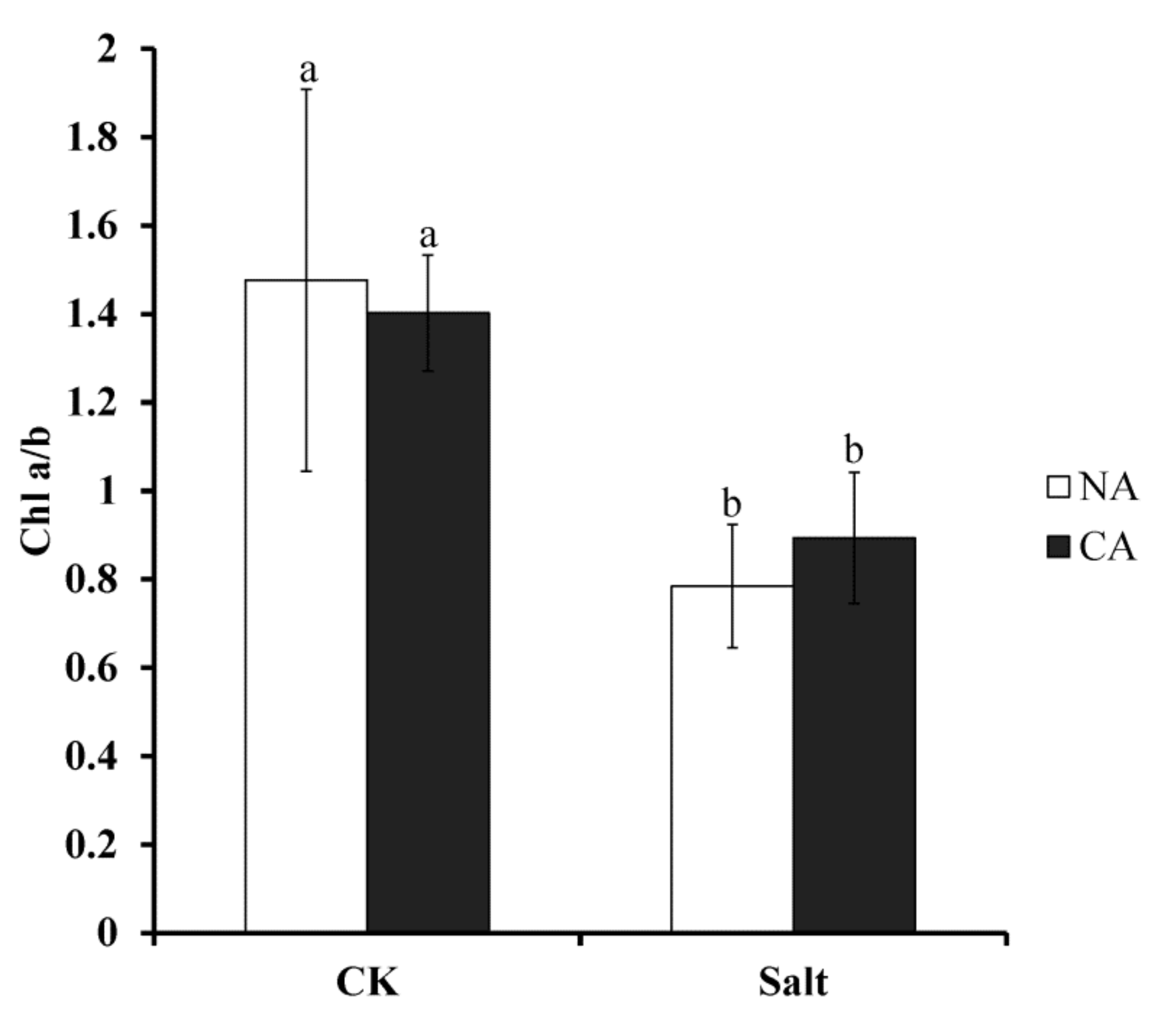
2.3. The Effect of Short-Term Pre-Cold Treatment on Chlorophyll a/b (Chl a/b) in Bermudagrass under Salt Stress
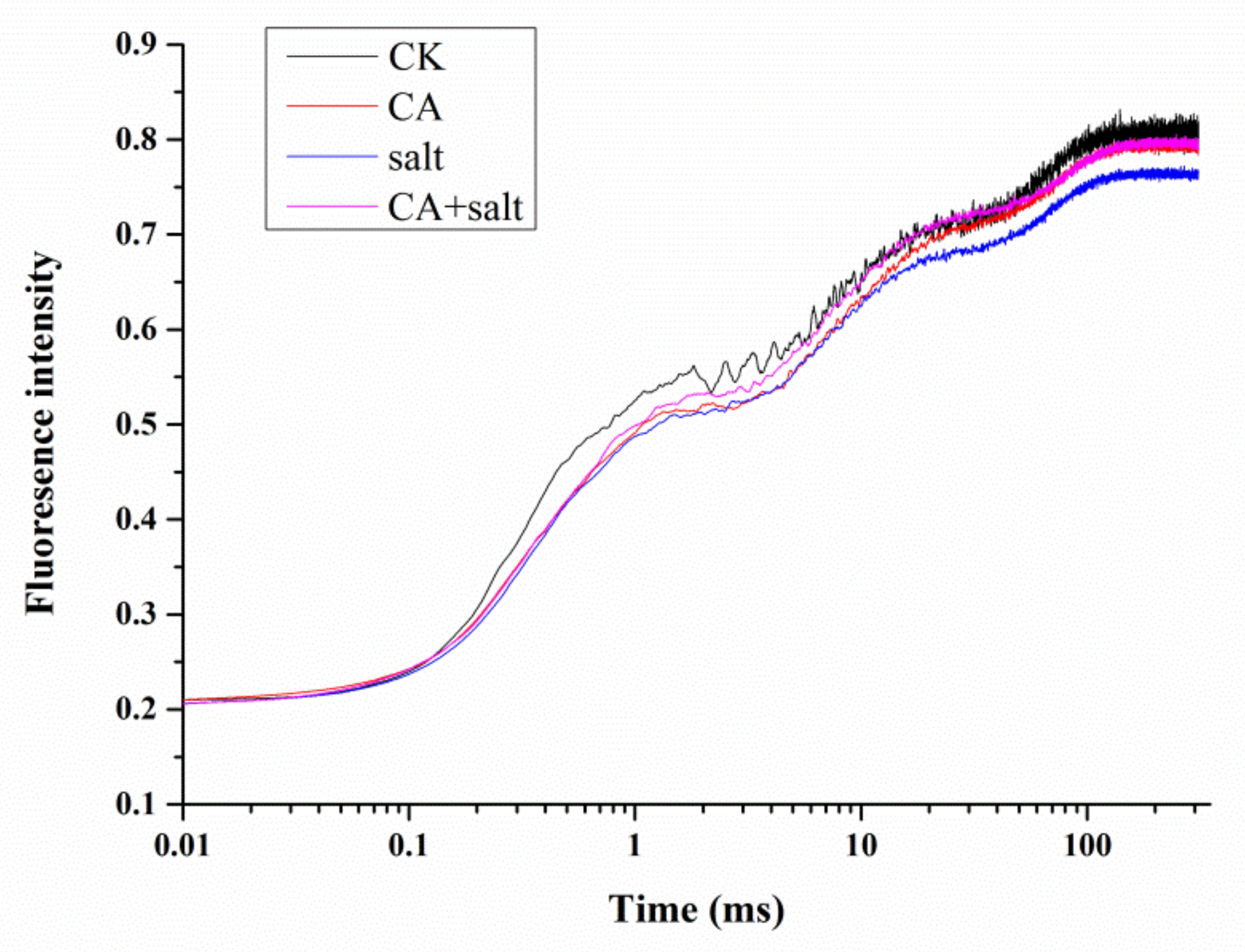
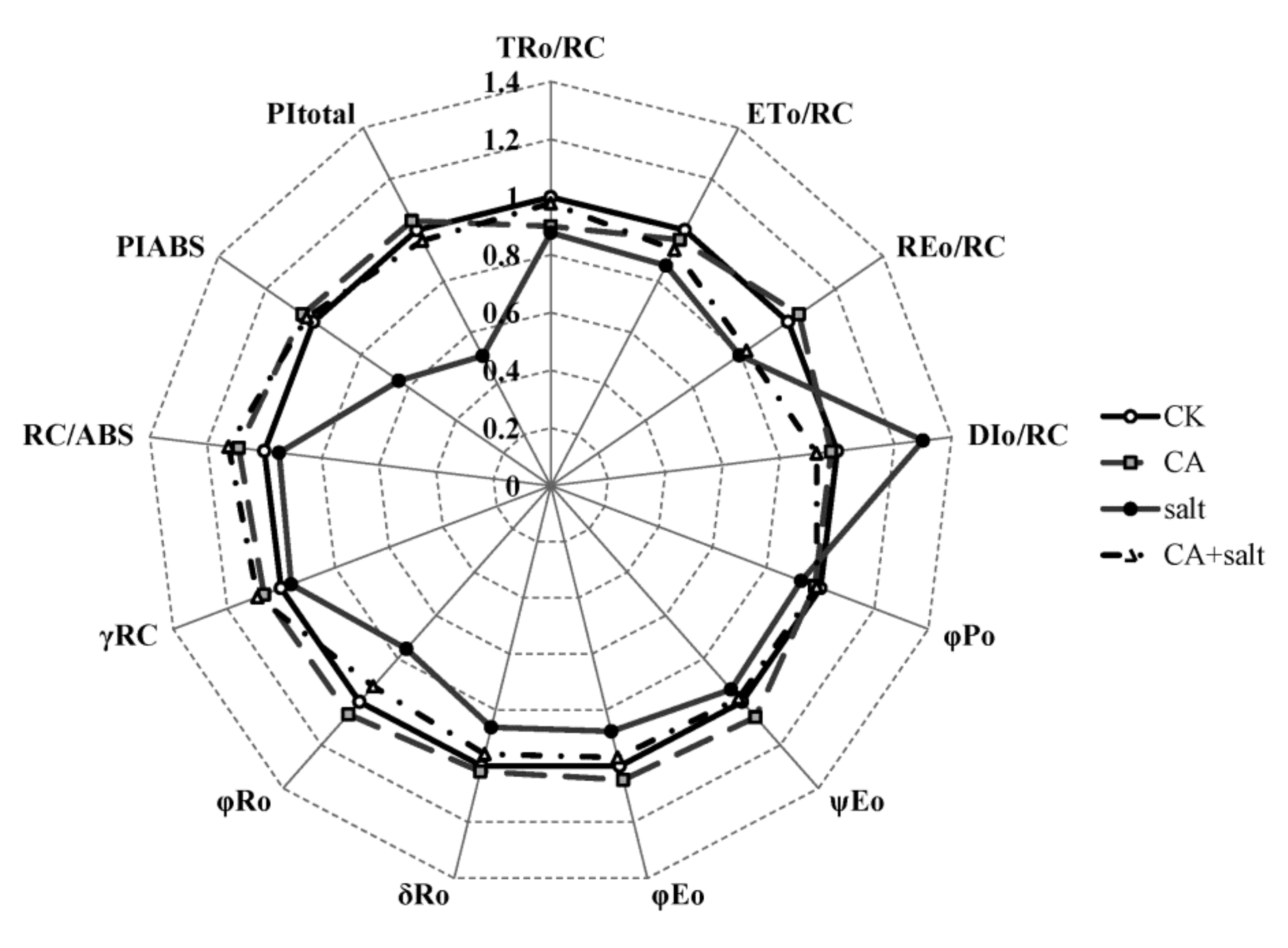
2.4. The Effect of Short-Term Pre-Cold Treatment on Chl a Fluorescence Transient (OJIP) in Bermudagrass under Salt Stress
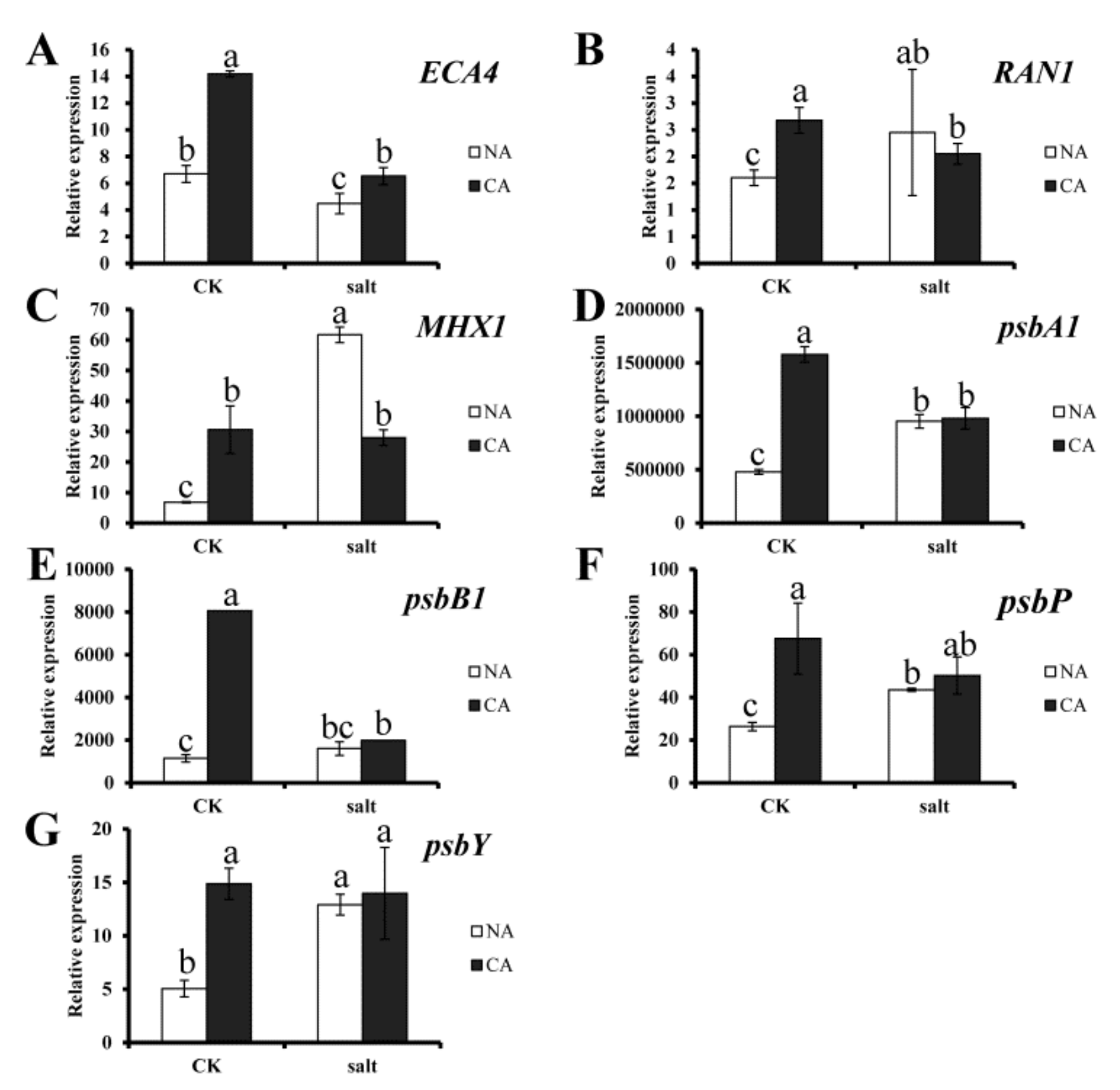
2.5. Gene Expression
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Plant Materials and Growth Conditions
5.2. Treatments
5.3. Measurements
5.3.1. Electrolyte Leakage
5.3.2. Ion Content
5.3.3. Chlorophyll Content
5.3.4. Chlorophyll (Chl) a Fluorescence Transient
5.3.5. The JIP-Test
5.3.6. RNA Extraction and Gene Expression Analysis
5.4. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Xiong, L.; Ishitani, M.; Zhu, J.K. Interaction of osmotic stress, temperature, and abscisic acid in the regulation of gene expression in Arabidopsis. Plant Physiol. 1999, 119, 205–212. [Google Scholar] [CrossRef] [PubMed]
- Choi, W.G.; Toyota, M.; Kim, S.H.; Hilleary, R.; Gilroy, S. Salt stress-induced Ca2+ waves are associated with rapid, long-distance root-to-shoot signaling in plants. Proc. Natl. Acad. Sci. USA 2014, 111, 6497–6502. [Google Scholar] [CrossRef] [PubMed]
- Pandey, G.K.; Kanwar, P.; Singh, A.; Steinhorst, L.; Pandey, A.; Yadav, A.K.; Tokas, I.; Sanyal, S.K.; Kim, B.G.; Lee, S.C.; et al. Calcineurin B-like protein-interacting protein kinase CIPK21 regulates osmotic and salt stress responses in Arabidopsis. Plant Physiol. 2015, 169, 780–792. [Google Scholar] [CrossRef] [PubMed]
- Demiral, T.; Türkan, İ. Comparative lipid peroxidation, antioxidant defense systems and proline content in roots of two rice cultivars differing in salt tolerance. Environ. Exp. Bot. 2005, 53, 247–257. [Google Scholar] [CrossRef]
- Tuna, A.L.; Kaya, C.; Ashraf, M.; Altunlu, H.; Yokas, I.; Yagmur, B. The effects of calcium sulphate on growth, membrane stability and nutrient uptake of tomato plants grown under salt stress. Environ. Exp. Bot. 2007, 59, 173–178. [Google Scholar] [CrossRef]
- Hu, L.; Hu, T.; Zhang, X.; Pang, H.; Fu, J. Exogenous glycine betaine ameliorates the adverse effect of salt stress on perennial ryegrass. J. Am. Soc. Hortic. Sci. 2012, 137, 38–46. [Google Scholar] [CrossRef]
- Kim, B.G.; Waadt, R.; Cheong, Y.H.; Pandey, G.K.; Dominguez-Solis, J.R.; Schültke, S.; Lee, S.C.; Kudla, J.; Luan, S. The calcium sensor CBL10 mediates salt tolerance by regulating ion homeostasis in Arabidopsis. Plant J. 2007, 52, 473–484. [Google Scholar] [CrossRef]
- Richter, J.A.; Behr, J.H.; Erban, A.; Kopka, J.; Zörb, C. Ion-dependent metabolic responses of Vicia faba L. to salt stress. Plant Cell Environ. 2019, 42, 295–309. [Google Scholar] [CrossRef]
- Meloni, D.A.; Oliva, M.A.; Martinez, C.A.; Cambraia, J. Photosynthesis and activity of superoxide dismutase, peroxidase and glutathione reductase in cotton under salt stress. Environ. Exp. Bot. 2003, 49, 69–76. [Google Scholar] [CrossRef]
- Sairam, R.K.; Srivastava, G.C. Changes in antioxidant activity in sub-cellular fractions of tolerant and susceptible wheat genotypes in response to long term salt stress. Plant Sci. 2002, 162, 897–904. [Google Scholar] [CrossRef]
- Regni, L.; Del Pino, A.M.; Mousavi, S.; Palmerini, C.A.; Baldoni, L.; Mariotti, R.; Mairech, H.; Gardi, T.; D’Amato, R.; Proietti, P. Behavior of four olive cultivars during salt stress. Front. Plant Sci. 2019, 10, 867. [Google Scholar] [CrossRef] [PubMed]
- Porcel, R.; Redondo-Gómez, S.; Mateos-Naranjo, E.; Aroca, R.; Garcia, R.; Ruiz-Lozano, J.M. Arbuscular mycorrhizal symbiosis ameliorates the optimum quantum yield of photosystem II and reduces non-photochemical quenching in rice plants subjected to salt stress. J. Plant Physiol. 2015, 185, 75–83. [Google Scholar] [CrossRef] [PubMed]
- Stirbet, A. On the relation between the Kautsky effect (chlorophyll a fluorescence induction) and Photosystem II: Basics and applications of the OJIP fluorescence transient. J. Photochem. Photobiol. B Biol. 2011, 104, 236–257. [Google Scholar] [CrossRef] [PubMed]
- Strasser, R.J.; Tsimilli-Michael, M.; Qiang, S.; Goltsev, V. Simultaneous in vivo recording of prompt and delayed fluorescence and 820-nm reflection changes during drying and after rehydration of the resurrection plant Haberlea rhodopensis. BBA-Bioenerg. 2010, 1797, 1313–1326. [Google Scholar] [CrossRef] [PubMed]
- Maxwell, K.; Johnson, G.N. Chlorophyll fluorescence—A practical guide. J. Exp. Bot. 2000, 51, 659–668. [Google Scholar] [CrossRef] [PubMed]
- Yusuf, M.A.; Kumar, D.; Rajwanshi, R.; Strasser, R.J.; Tsimilli-Michael, M.; Sarin, N.B. Overexpression of γ-tocopherol methyl transferase gene in transgenic Brassica juncea plants alleviates abiotic stress: Physiological and chlorophyll a fluorescence measurements. BBA-Bioenerg. 2010, 1797, 1428–1438. [Google Scholar] [CrossRef] [PubMed]
- Theocharis, A.; Clement, C.; Barka, E.A. Physiological and molecular changes in plants grown at low temperatures. Planta 2012, 235, 1091–1105. [Google Scholar] [CrossRef] [PubMed]
- Costa-Broseta, Á.; Perea-Resa, C.; Castillo, M.C.; Ruíz, M.F.; Salinas, J.; León, J. Nitric oxide deficiency decreases C-repeat bingding factor-dependent and -independent induction of cold acclimation. J. Exp. Bot. 2019, 70, 3283–3296. [Google Scholar] [CrossRef] [PubMed]
- Chai, F.; Liu, W.; Xiang, Y.; Meng, X.; Sun, X.; Cheng, C.; Liu, G.; Duan, L.; Xin, H.; Li, S. Comparative metabolic profiling of Vitis amurensis and Vitis vinifera during cold acclimation. Hortic. Res. 2019, 6, 8. [Google Scholar] [CrossRef]
- Takahashi, D.; Gorka, M.; Erban, A.; Graf, A.; Kopka, J.; Zuther, E.; Hincha, D.K. Both cold and sub-zero acclimation induce cell wall modification and changes in the extracellular proteome in Arabidopsis thaliana. Sci. Rep. 2019, 9, 2289. [Google Scholar] [CrossRef]
- Bertrand, A.; Bipfubusa, M.; Claessens, A.; Rocher, S.; Castonguay, Y. Effect of photoperiod prior to cold acclimation on freezing tolerance and carbohydrate metabolism in alfalfa (Medicago sativa L.). Plant Sci. 2017, 264, 122–128. [Google Scholar] [CrossRef]
- Gaete-Loyola, J.; Lagos, C.; Beltrán, M.F.; Valenzuela, S.; Emhart, V.; Fernández, M. Transcriptome profiling of Eucalyptus nitens reveals deeper insight into the molecular mechanism of cold acclimation and deacclimation process. Tree Genet. Genomes 2017, 13, 37. [Google Scholar] [CrossRef]
- Lv, X.; Li, H.; Chen, X.; Xiang, X.; Guo, Z.; Yu, J.; Zhou, Y. The role of calcium-dependent protein kinase in hydrogen peroxide, nitric oxide and ABA-dependent cold acclimation. J. Exp. Bot. 2018, 69, 4127–4139. [Google Scholar] [CrossRef] [PubMed]
- Zhao, C.; Zhang, Z.; Xie, S.; Si, T.; Li, Y.; Zhu, J.K. Mutational evidence for the critical role of CBF transcription factors in cold acclimation in Arabidopsis. Plant Physiol. 2016, 171, 2744–2759. [Google Scholar] [PubMed]
- Baier, M.; Bittner, A.; Prescher, A.; van Buer, J. Preparing plants for improved cold tolerance by priming. Plant Cell Environ. 2019, 42, 782–800. [Google Scholar] [CrossRef] [PubMed]
- Morton, M.J.L.; Awlia, M.; Al-Tamimi, N.; Saade, S.; Pailles, Y.; Negrão, S.; Tester, M. Salt stress under the scalpel—Dissecting the genetics of salt tolerance. Plant J. 2019, 97, 148–163. [Google Scholar] [CrossRef] [PubMed]
- Ding, Y.; Liu, N.; Virlouvet, L.; Riethoven, J.J.; Fromm, M.; Avramova, Z. Four distinct types of dehydration stress memory genes in Arabidopsis thaliana. BMC Plant Biol. 2013, 13, 229. [Google Scholar] [CrossRef] [PubMed]
- Tanou, G.; Minas, I.S.; Scossa, F.; Belghazi, M.; Xanthopoulou, A.; Ganopoulos, I.; Madesis, P.; Fernie, A.; Molassiotis, A. Exploring priming responses involved in peach fruit acclimation to cold stress. Sci. Rep. 2017, 7, 11358. [Google Scholar] [CrossRef]
- Huda, K.M.K.; Yadav, S.; Banu, M.S.A.; Trivedi, D.K.; Tuteja, N. Genome-wide analysis of plant-type II Ca2+ATPases gene family from rice and Arabidopsis: Potential role in abiotic stresses. Plant Physiol. Biochem. 2013, 65, 32–47. [Google Scholar] [CrossRef]
- Xu, P.; Cai, W. RAN1 is involved in plant cold resistance and development in rice (Oryza sativa). J. Exp. Bot. 2014, 65, 3277–3287. [Google Scholar] [CrossRef]
- Pittman, J.K.; Hirschi, K.D. Phylogenetic analysis and protein structure modelling identifies distinct Ca 2+/Cation antiporters and conservation of gene family structure within Arabidopsis and rice species. Rice 2016, 9, 3. [Google Scholar] [CrossRef] [PubMed]
- Hu, Z.; Liu, A.; Gitau, M.M.; Huang, X.; Chen, L.; Fu, J. Insights into the MicroRNA-regulated response of bermudagrass to cold and salt stress. Environ. Exp. Bot. 2018, 145, 64–74. [Google Scholar] [CrossRef]
- Chattopadhayay, M.K.; Tiwari, B.S.; Chattopadhyay, G.; Bose, A.; Sengupta, D.N.; Ghosh, B. Protective role of exogenous polyamines on salinity-stressed rice (Oryza sativa) plants. Physiol. Plant. 2002, 116, 192–199. [Google Scholar] [CrossRef]
- Shi, H.; Jiang, C.; Ye, T.; Yan, D.X.; Reiter, R.J.; Zhang, H.; Liu, R.; Chan, Z. Comparative physiological, metabolomic, and transcriptomic analyses reveal mechanisms of improved abiotic stress resistance in bermudagrass [Cynodon dactylon (L). Pers.] by exogenous melatonin. J. Exp. Bot. 2015, 66, 681–694. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Fan, J.; Hu, L.; Hu, Z.; Xie, Y.; Zhang, Y.; Lou, Y.; Nevo, E.; Fu, J. A transcriptomic analysis of bermudagrass (Cynodon dactylon) provides novel insights into the basis of low temperature tolerance. BMC Plant Biol. 2015, 15, 216. [Google Scholar] [CrossRef]
- Liu, A.; Hu, Z.; Bi, A.; Fan, J.; Gitau, M.M.; Amombo, E.; Chen, L.; Fu, J. Photosynthesis, antioxidant system and gene expression of bermudagrass in response to low temperature and salt stress. Ecotoxicology 2016, 25, 1445–1457. [Google Scholar] [CrossRef]
- Savvides, A.; Ali, S.; Tester, M.; Fotopoulos, V. Chemical priming of plants against multiple abiotic stresses: Mission possible? Trends Plant Sci. 2016, 21, 329–340. [Google Scholar] [CrossRef]
- Pandolfi, C.; Azzarello, E.; Mancuso, S.; Shabala, S. Acclimation improves salt stress tolerance in Zea mays plants. J. Plant Physiol. 2016, 201, 1–8. [Google Scholar] [CrossRef]
- Hoffmann, A.M.; Noga, G.; Hunsche, M. High blue light improves acclimation and photosynthetic recovery of pepper plants exposed to UV stress. Environ. Exp. Bot. 2015, 109, 254–263. [Google Scholar] [CrossRef]
- Tang, X.; Mu, X.; Shao, H.; Wang, H.; Brestic, M. Global plant-responding mechanisms to salt stress: Physiological and molecular levels and implications in biotechnology. Crit. Rev. Biotechnol. 2015, 35, 425–437. [Google Scholar] [CrossRef]
- You, J.; Chan, Z. ROS regulation during abiotic stress responses in crop plants. Front. Plant Sci. 2015, 6, 1092. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Smith, J.A.C.; Harberd, N.P.; Jiang, C. The regulatory roles of ethylene and reactive oxygen species (ROS) in plant salt stress responses. Plant Mol. Biol. 2016, 91, 651–659. [Google Scholar] [CrossRef] [PubMed]
- Petrov, V.; Hille, J.; Mueller-Roeber, B.; Gechev, T.S. ROS-mediated abiotic stress-induced programmed cell death in plants. Front. Plant Sci. 2015, 6, 69. [Google Scholar] [CrossRef] [PubMed]
- Koca, H.; Bor, M.; Özdemir, F.; Türkan, İ. The effect of salt stress on lipid peroxidation, antioxidative enzymes and proline content of sesame cultivars. Environ. Exp. Bot. 2007, 60, 344–351. [Google Scholar] [CrossRef]
- AbdElgawad, H.; Zinta, G.; Hegab, M.M.; Pandey, R.; Asard, H.; Abuelsoud, W. High salinity induces different oxidative stress and antioxidant responses in maize seedlings organs. Front. Plant Sci. 2016, 7, 276. [Google Scholar] [CrossRef] [PubMed]
- Farhangi-Abriz, S.; Torabian, S. Antioxidant enzyme and osmotic adjustment changes in bean seedlings as affected by biochar under salt stress. Ecotoxicol. Environ. Saf. 2017, 137, 64–70. [Google Scholar] [CrossRef] [PubMed]
- Forni, C.; Duca, D.; Glick, B.R. Mechanisms of plant response to salt and drought stress and their alteration by rhizobacteria. Plant Soil 2017, 410, 335–356. [Google Scholar] [CrossRef]
- Sheldon, A.R.; Dalal, R.C.; Kirchhof, G.; Kopittke, P.M.; Menzies, N.W. The effect of salinity on plant-available water. Plant Soil 2017, 418, 477–491. [Google Scholar] [CrossRef]
- Negrão, S.; Schmöckel, S.M.; Tester, M. Evaluating physiological responses of plants to salinity stress. Ann. Bot. 2017, 119, 1–11. [Google Scholar] [CrossRef]
- Egea, I.; Pineda, B.; Ortíz-Atienza, A.; Plasencia, F.A.; Drevensek, S.; García-Sogo, B.; Yuste-Lisbona, F.J.; Barrero-Gil, J.; Atarés, A.; Flores, F.B.; et al. The SlCBL10 calcineurin B-like protein ensures plant growth under salt stress by regulating Na+ and Ca2+ homeostasis. Plant Physiol. 2018, 176, 1676–1693. [Google Scholar] [CrossRef]
- El-Esawi, M.; Alaraidh, I.; Alsahli, A.; Alzahrani, S.; Ali, H.; Alayafi, A.; Ahmad, M. Serratia liquefaciens KM4 improves salt stress tolerance in maize by regulating redox potential, ion homeostasis, leaf gas exchange and stress-related gene expression. Int. J. Mol. Sci. 2018, 19, 3310. [Google Scholar] [CrossRef] [PubMed]
- Hamamoto, S.; Horie, T.; Hauser, F.; Deinlein, U.; Schroeder, J.I.; Uozumi, N. HKT transporters mediate salt stress resistance in plants: From structure and function to the field. Curr. Opin. Biotechnol. 2015, 32, 113–120. [Google Scholar] [CrossRef] [PubMed]
- Baker, N.R. A possible role for photosystem II in environmental perturbations of photosynthesis. Physiol. Plant. 1991, 81, 563–570. [Google Scholar] [CrossRef]
- Santos, C.V. Regulation of chlorophyll biosynthesis and degradation by salt stress in sunflower leaves. Sci. Hortic. 2004, 103, 93–99. [Google Scholar] [CrossRef]
- Rout, N.P.; Shaw, B.P. Salt tolerance in aquatic macrophytes: Possible involvement of the antioxidative enzymes. Plant Sci. 2001, 160, 415–423. [Google Scholar] [CrossRef]
- Kalaji, H.M.; Jajoo, A.; Oukarroum, A.; Brestic, M.; Zivcak, M.; Samborska, I.A.; Cetner, M.D.; Łukasik, I.; Goltsev, V.; Ladle, R.J. Chlorophyll a fluorescence as a tool to monitor physiological status of plants under abiotic stress conditions. Acta Physiol. Plant. 2016, 38, 102. [Google Scholar] [CrossRef]
- Oukarroum, A.; Bussotti, F.; Goltsev, V.; Kalaji, H.M. Correlation between reactive oxygen species production and photochemistry of photosystems I and II in Lemna gibba L. plants under salt stress. Environ. Exp. Bot. 2015, 109, 80–88. [Google Scholar] [CrossRef]
- Zhang, C.J.; Lim, S.H.; Kim, J.W.; Nah, G.; Fischer, A.; Kim, D.S. Leaf chlorophyll fluorescence discriminates herbicide resistance in Echinochloa species. Weed Res. 2016, 56, 424–433. [Google Scholar] [CrossRef]
- Chen, S.; Yang, J.; Zhang, M.; Strasser, R.J.; Qiang, S. Classification and characteristics of heat tolerance in Ageratina adenophora populations using fast chlorophyll a fluorescence rise O-J-I-P. Environ. Exp. Bot. 2016, 122, 126–140. [Google Scholar] [CrossRef]
- Živčák, M.; Olšovská, K.; Slamka, P.; Galambošová, J.; Rataj, V.; Shao, H.B.; Brestič, M. Application of chlorophyll fluorescence performance indices to assess the wheat photosynthetic functions influenced by nitrogen deficiency. Plant Soil Environ. 2014, 60, 210–215. [Google Scholar] [CrossRef]
- Akilan, S.; Halima, T.H.; Sasi, S.; Kappachery, S.; Baniekal-Hiremath, G.; Venkatesh, J.; Gururani, A. Evaluation of osmotic stress tolerance in transgenic Arabidopsis plants expressing Solanum tuberosum D200 gene. J. Plant Interact. 2018, 14, 79–86. [Google Scholar] [CrossRef]
- Bowler, C.; Fluhr, R. The role of calcium and activated oxygen as signals for controlling cross-tolerance. Trends Plant Sci. 2000, 5, 241–246. [Google Scholar] [CrossRef]
- Faralli, M.; Lektemur, C.; Rosellini, D.; Gürel, F. Effects of heat shock and salinity on barley growth and stress-related gene transcription. Biol. Plant. 2015, 59, 537–546. [Google Scholar] [CrossRef]
- Hilker, M.; Schwachtje, J.; Baier, M.; Balazadeh, S.; Bäurle, I.; Geiselhardt, S.; Hincha, D.K.; Kunze, R.; Mueller-Roeber, B.; Rillig, M.C.; et al. Priming and memory of stress responses in organisms lacking a nervous system. Biol. Rev. 2016, 91, 1118–1133. [Google Scholar] [CrossRef] [PubMed]
- Hossain, M.A.; Li, Z.G.; Hoque, T.S.; Burritt, D.J.; Fujita, M.; Munné-Bosch, S. Heat or cold priming-induced cross-tolerance to abiotic stresses in plants: Key regulators and possible mechanisms. Protoplasma 2018, 255, 399–412. [Google Scholar] [CrossRef] [PubMed]
- van Buer, J.; Prescher, A.; Baier, M. Cold-priming of chloroplast ROS signalling is developmentally regulated and is locally controlled at the thylakoid membrane. Sci. Rep. 2019, 9, 3022. [Google Scholar] [CrossRef] [PubMed]
- Fan, J.; Ren, J.; Zhu, W.; Amombo, E.; Fu, J.; Chen, L. Antioxidant responses and gene expression in bermudagrass under cold stress. J. Am. Soc. Hortic. Sci. 2014, 139, 699–705. [Google Scholar] [CrossRef]
- Chan, Z.; Shi, H. Improved abiotic stress tolerance of bermudagrass by exogenous small molecules. Plant Signal. Behav. 2015, 10, e991577. [Google Scholar] [CrossRef]
- Hu, L.; Li, H.; Pang, H.; Fu, J. Responses of antioxidant gene, protein and enzymes to salinity stress in two genotypes of perennial ryegrass (Lolium perenne) differing in salt tolerance. J. Plant Physiol. 2012, 169, 146–156. [Google Scholar] [CrossRef]
- Chen, K.; Chen, L.; Fan, J.; Fu, J. Alleviation of heat damage to photosystem II by nitric oxide in tall fescue. Photosynth. Res. 2013, 116, 21–31. [Google Scholar] [CrossRef]
- Chen, L.; Zhong, H.; Ren, F.; Guo, Q.; Hu, X.P. A novel coldregulated gene, COR25, of Brassica napus is involved in plant response and tolerance to cold stress. Plant Cell Rep. 2011, 30, 463–471. [Google Scholar] [CrossRef] [PubMed]
| Gene Name | Primer Direction | Primer Sequences (5′-3′) |
|---|---|---|
| ECA4 | F | GCCTTCGTCGAGCCGCTCGT |
| R | GCTGATAAGCTGGAGCACGC | |
| RAN1 | F | AATGCAGAAGGAAAATATTT |
| R | TCCTACTTTGGTCGCTTGTA | |
| MHX1 | F | ATGGCGAGCACTGCTCTGTC |
| R | GTAGTTCCACACCTTCTCAT | |
| psbA | F | AACAAGCCTTCTATTATCTATT |
| R | GGGCAGCGATGAAGGCGATA | |
| psbB | F | TGGGTTTACCTTGGTATCGT |
| R | TTCCTCCTGAAATACTCCAA | |
| psbP | F | AGGTGTACAAAGATGTGATT |
| R | GCACCCAGTGCATGTCTGGT | |
| psbY | F | TCTCATCTTCCTTCTCCACT |
| R | CTTCCATGGAAGCAGGACTT | |
| Actin | F | TCTGAAGGGTAAGTAGAGTAG |
| R | ACTCAGCACATTCCAGCAGAT |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fan, J.; Xu, J.; Zhang, W.; Amee, M.; Liu, D.; Chen, L. Salt-Induced Damage is Alleviated by Short-Term Pre-Cold Treatment in Bermudagrass (Cynodon dactylon). Plants 2019, 8, 347. https://doi.org/10.3390/plants8090347
Fan J, Xu J, Zhang W, Amee M, Liu D, Chen L. Salt-Induced Damage is Alleviated by Short-Term Pre-Cold Treatment in Bermudagrass (Cynodon dactylon). Plants. 2019; 8(9):347. https://doi.org/10.3390/plants8090347
Chicago/Turabian StyleFan, Jibiao, Jilei Xu, Weihong Zhang, Maurice Amee, Dalin Liu, and Liang Chen. 2019. "Salt-Induced Damage is Alleviated by Short-Term Pre-Cold Treatment in Bermudagrass (Cynodon dactylon)" Plants 8, no. 9: 347. https://doi.org/10.3390/plants8090347
APA StyleFan, J., Xu, J., Zhang, W., Amee, M., Liu, D., & Chen, L. (2019). Salt-Induced Damage is Alleviated by Short-Term Pre-Cold Treatment in Bermudagrass (Cynodon dactylon). Plants, 8(9), 347. https://doi.org/10.3390/plants8090347






