Intra-Sample Heterogeneity of Potato Starch Reveals Fluctuation of Starch-Binding Proteins According to Granule Morphology
Abstract
1. Introduction
2. Results
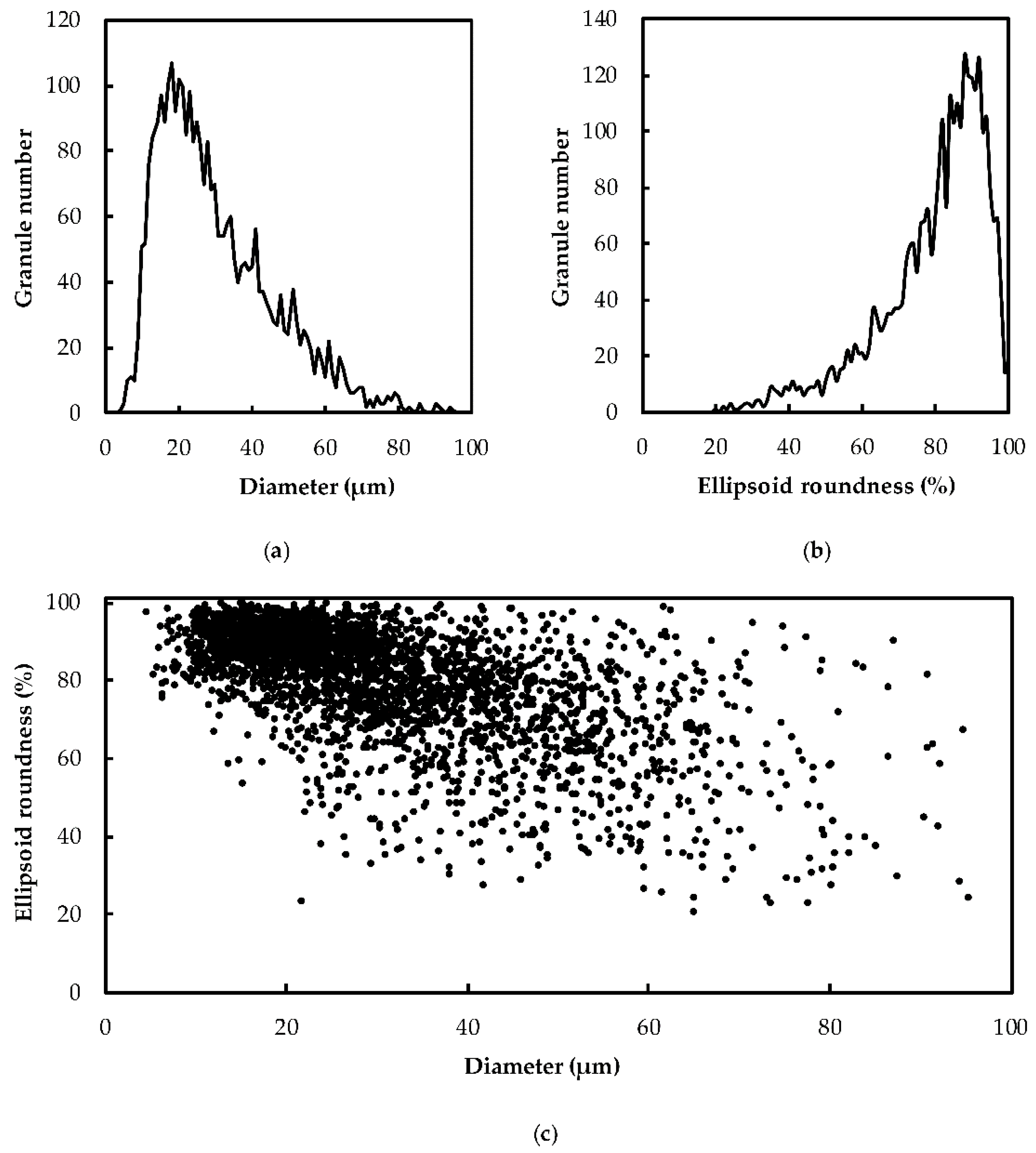
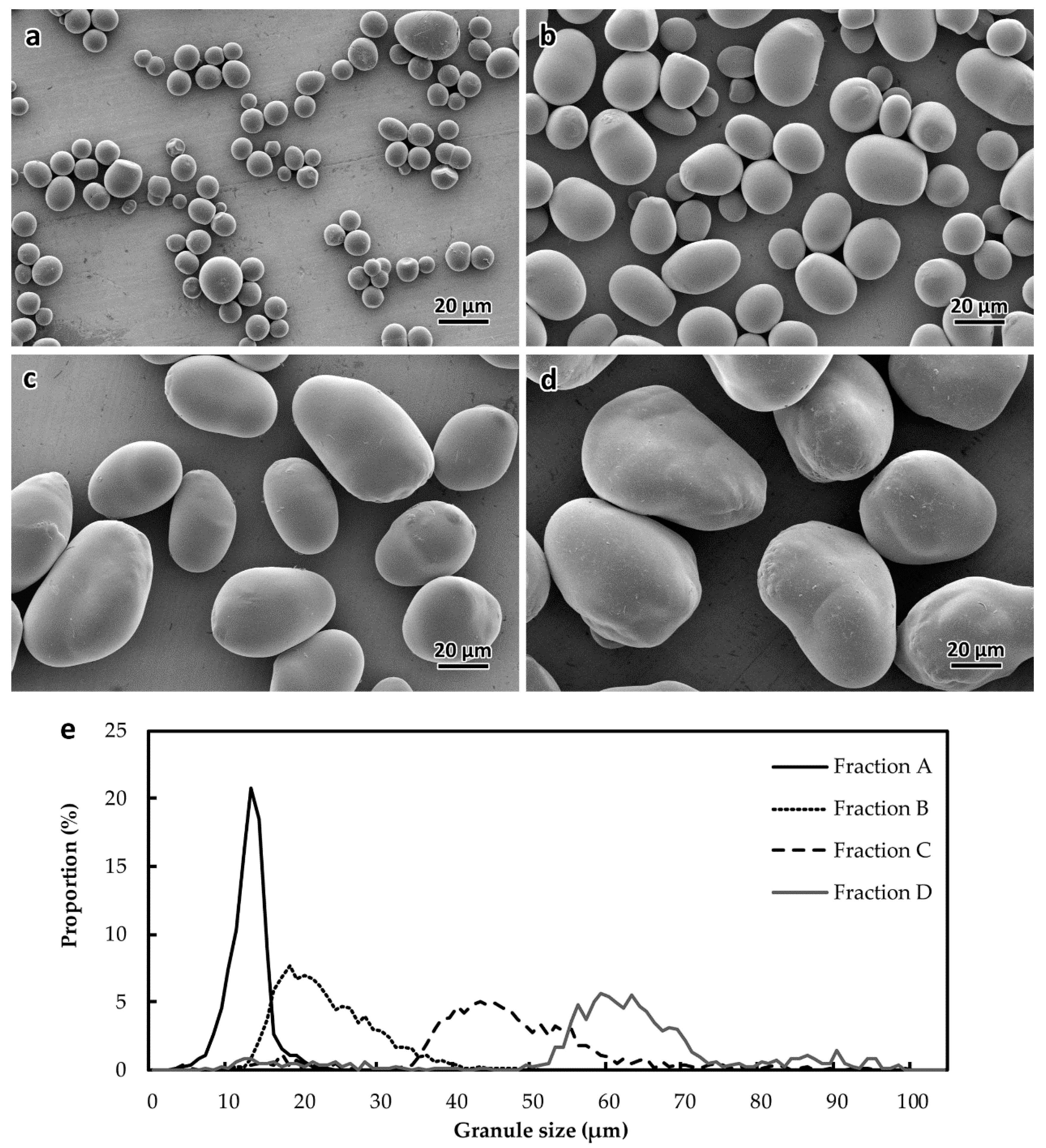
2.1. Heterogeneity of Starch Morphology and Size-Fractionation
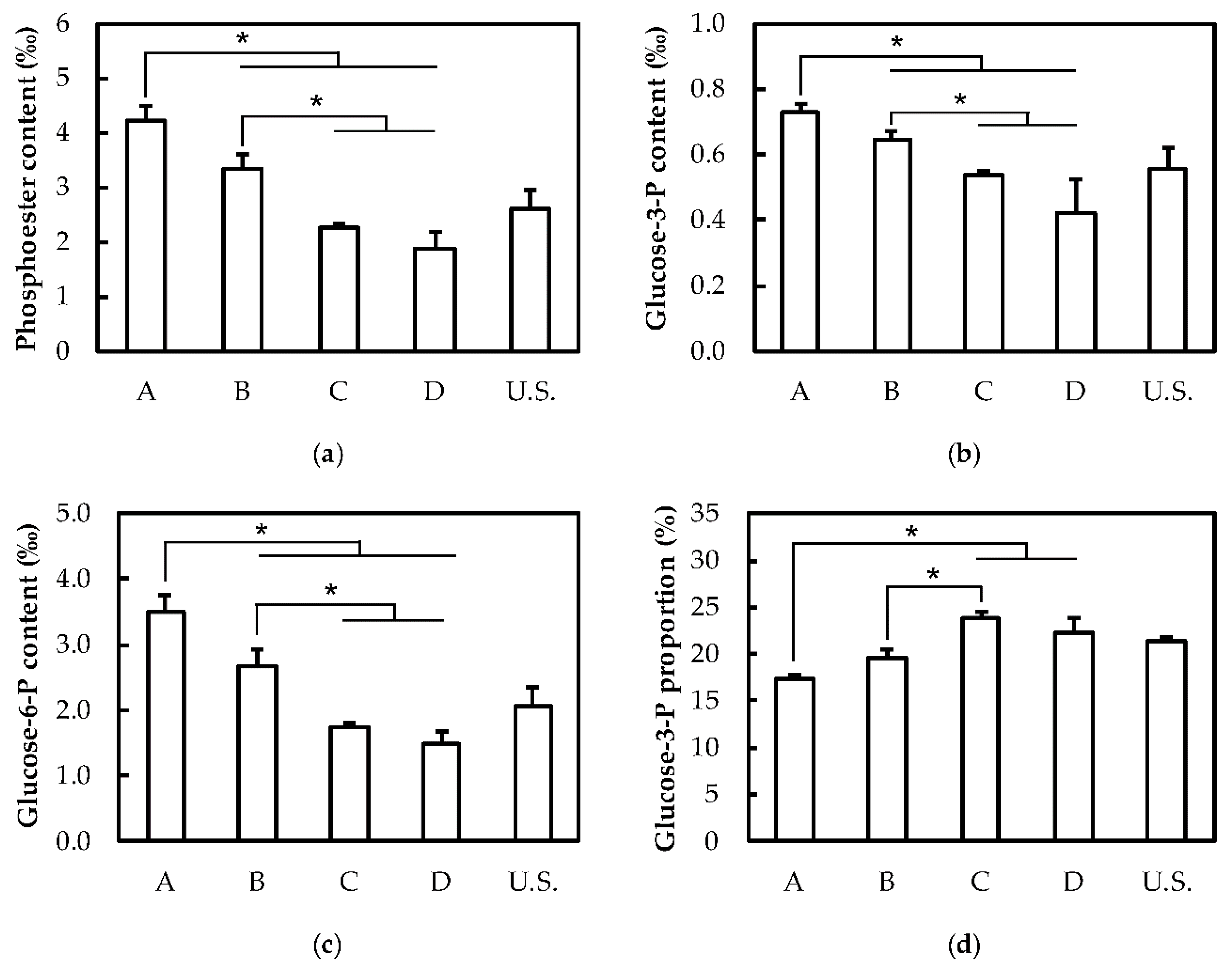
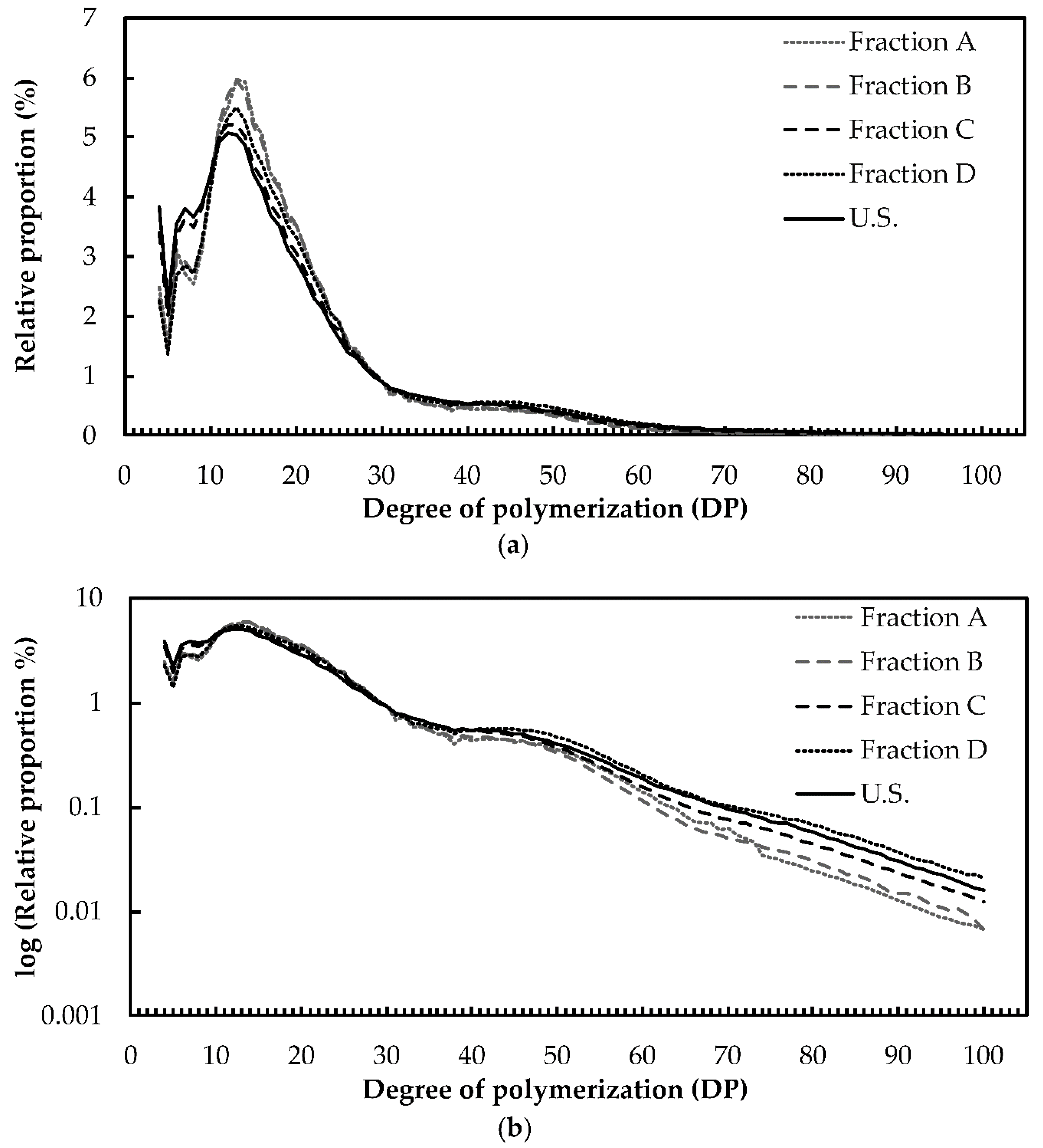
2.2. Structure and Composition of Starch
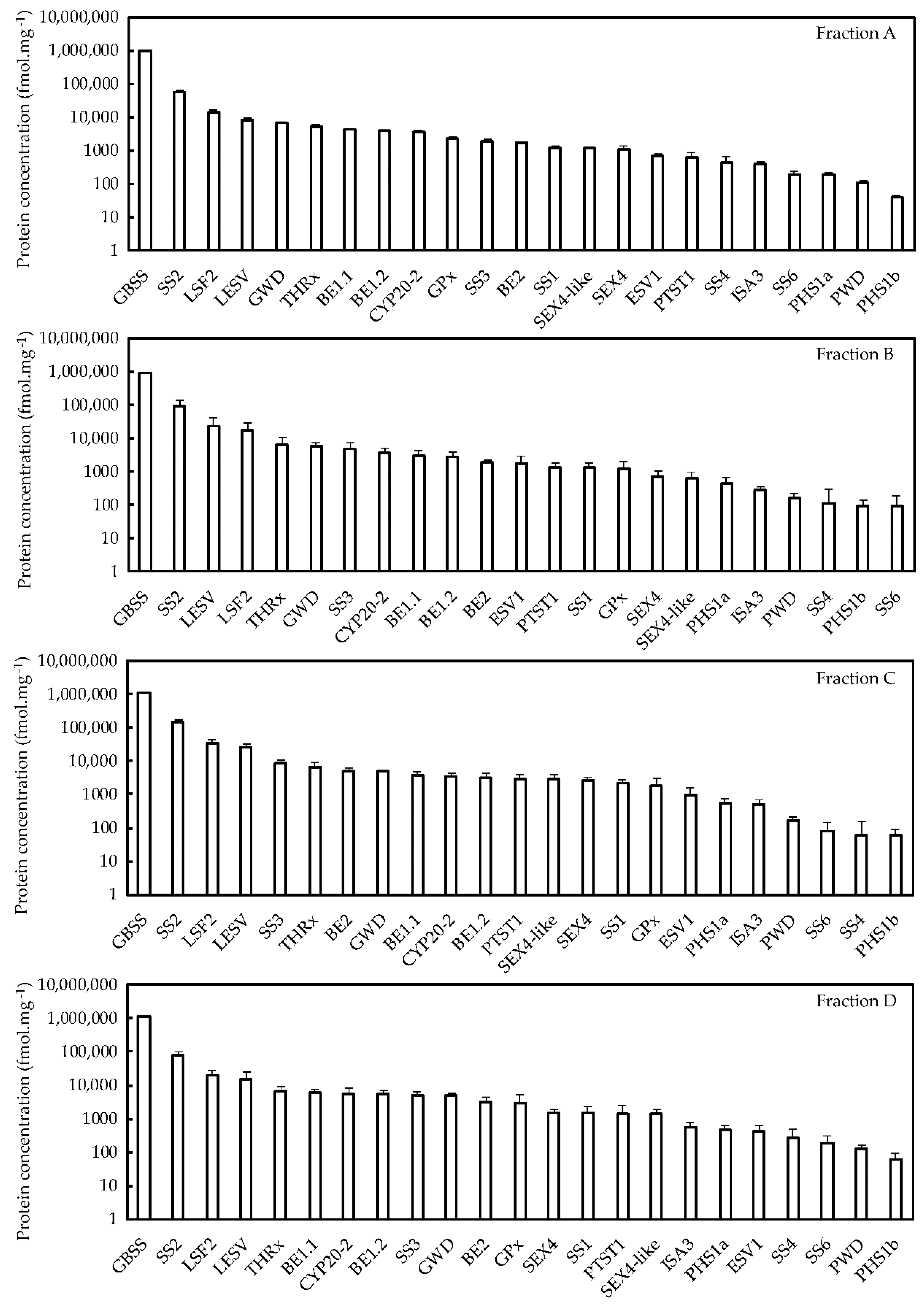
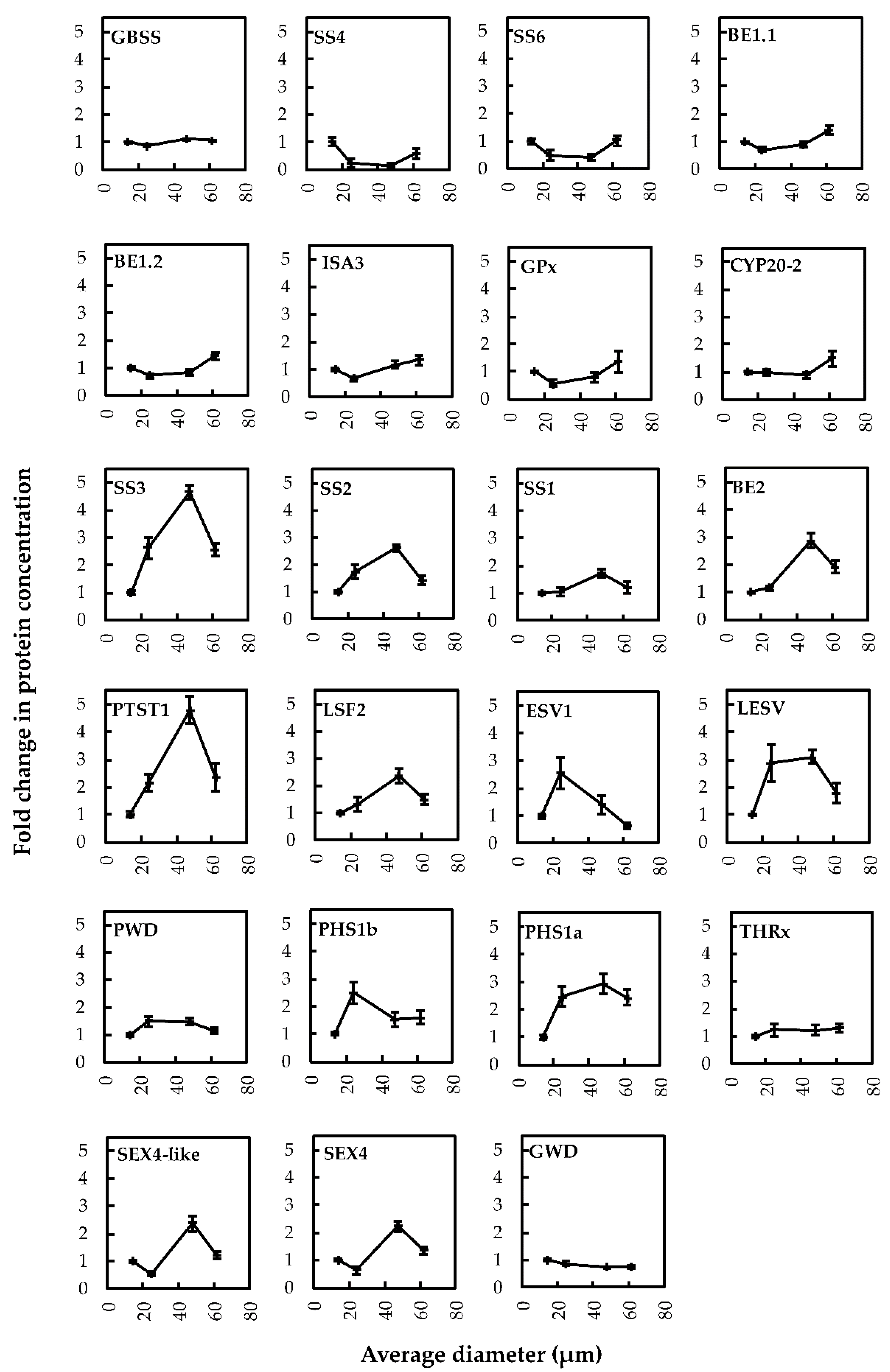
2.3. Quantitative Proteomics and Correlation Analysis
3. Discussion
4. Materials and Methods
4.1. Starch Extraction
4.2. Granulomorphometry
4.3. Filter-Fractionation of Starch Granules
4.4. Scanning Electron Microscopy (SEM)
4.5. X-Ray Diffraction (XRD)
4.6. Amylose Content
4.7. Amylopectin Chain Length Distribution
4.8. Phosphate Content
4.9. Starch Granule Associated Protein Extraction
4.10. Label-Free Quantitative Proteomics, Sample Preparation
4.11. Mass Spectrometry Analysis
4.12. Proteins Label Free Quantification with Maxquant and Perseus
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Manners, D.J. The structure and metabolism of starch. Essays Biochem. 1974, 10, 37–71. [Google Scholar] [PubMed]
- Curá, J.A.; Jansson, P.-E.; Krisman, C.R. Amylose is not strictly linear. Starch Stärke 1995, 47, 207–209. [Google Scholar] [CrossRef]
- Bertoft, E. Understanding starch structure: Recent Progress. Agronomy 2017, 7, 56. [Google Scholar] [CrossRef]
- Swinkels, J.J.M. Composition and properties of commercial native starches. Starch Stärke 1985, 37, 1–5. [Google Scholar] [CrossRef]
- Baldwin, P.M. Starch granule-associated proteins and polypeptides: A Review. Starch Stärke 2001, 53, 475–503. [Google Scholar] [CrossRef]
- Denyer, K.; Clarke, B.; Hylton, C.; Tatge, H.; Smith, A.M. The elongation of amylose and amylopectin chains in isolated starch granules. Plant J. 1996, 10, 1135–1143. [Google Scholar] [CrossRef]
- Commuri, P.D.; Keeling, P.L. Chain-length specificities of maize starch synthase I enzyme: Studies of glucan affinity and catalytic properties. Plant J. 2001, 25, 475–486. [Google Scholar] [CrossRef]
- Craig, J.; Lloyd, J.R.; Tomlinson, K.; Barber, L.; Edwards, A.; Wang, T.L.; Martin, C.; Hedley, C.L.; Smith, A.M. Mutations in the gene encoding starch synthase II profoundly alter amylopectin structure in pea embryos. Plant Cell 1998, 10, 413–426. [Google Scholar] [CrossRef]
- Lin, Q.; Huang, B.; Zhang, M.; Zhang, X.; Rivenbark, J.; Lappe, R.L.; James, M.G.; Myers, A.M.; Hennen-Bierwagen, T.A. Functional interactions between starch synthase III and isoamylase-type starch-debranching enzyme in maize endosperm. Plant Physiol. 2012, 158, 679–692. [Google Scholar] [CrossRef]
- Ragel, P.; Streb, S.; Feil, R.; Sahrawy, M.; Annunziata, M.G.; Lunn, J.E.; Zeeman, S.; Mérida, Á. Loss of starch granule initiation has a deleterious effect on the growth of arabidopsis plants due to an accumulation of ADP-glucose. Plant Physiol. 2013, 163, 75–85. [Google Scholar] [CrossRef]
- Raynaud, S.; Ragel, P.; Rojas, T.; Mérida, Á. The N-terminal part of Arabidopsis thaliana starch synthase 4 determines the localization and activity of the enzyme. J. Biol. Chem. 2016, 291, 10759–10771. [Google Scholar] [CrossRef] [PubMed]
- Roldán, I.; Wattebled, F.; Mercedes Lucas, M.; Delvallé, D.; Planchot, V.; Jiménez, S.; Pérez, R.; Ball, S.; D’Hulst, C.; Mérida, A. The phenotype of soluble starch synthase IV defective mutants of Arabidopsis thaliana suggests a novel function of elongation enzymes in the control of starch granule formation. Plant J. Cell Mol. Biol. 2007, 49, 492–504. [Google Scholar] [CrossRef] [PubMed]
- Seung, D.; Lu, K.-J.; Stettler, M.; Streb, S.; Zeeman, S.C. Degradation of glucan primers in the absence of starch synthase 4 disrupts starch granule initiation in Arabidopsis. J. Biol. Chem. 2016, 291, 20718–20728. [Google Scholar] [CrossRef] [PubMed]
- Szydlowski, N.; Ragel, P.; Raynaud, S.; Lucas, M.M.; Roldán, I.; Montero, M.; Muñoz, F.J.; Ovecka, M.; Bahaji, A.; Planchot, V.; et al. Starch granule initiation in Arabidopsis requires the presence of either class IV or class III starch synthases. Plant Cell 2009, 21, 2443–2457. [Google Scholar] [CrossRef] [PubMed]
- Van Harsselaar, J.K.; Lorenz, J.; Senning, M.; Sonnewald, U.; Sonnewald, S. Genome-wide analysis of starch metabolism genes in potato (Solanum tuberosum L.). BMC Genomics 2017, 18, 37. [Google Scholar] [CrossRef] [PubMed]
- Helle, S.; Bray, F.; Verbeke, J.; Devassine, S.; Courseaux, A.; Facon, M.; Tokarski, C.; Rolando, C.; Szydlowski, N. Proteome analysis of potato starch reveals the presence of new starch metabolic proteins as well as multiple protease inhibitors. Front. Plant Sci. 2018, 9, 746. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Wu, A.C.; Go, R.M.; Malouf, J.; Turner, M.S.; Malde, A.K.; Mark, A.E.; Gilbert, R.G. The characterization of modified starch branching enzymes: Toward the control of starch chain-length distributions. PLoS ONE 2015, 10, e0125507. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, S.J.; Senning, M.; Fischer-Stettler, M.; Streb, S.; Ast, M.; Neuhaus, H.E.; Zeeman, S.C.; Sonnewald, S.; Sonnewald, U. Simultaneous silencing of isoamylases ISA1, ISA2 and ISA3 by multi-target RNAi in potato tubers leads to decreased starch content and an early sprouting phenotype. PLoS ONE 2017, 12, e0181444. [Google Scholar] [CrossRef] [PubMed]
- Streb, S.; Zeeman, S.C. Replacement of the endogenous starch debranching enzymes ISA1 and ISA2 of Arabidopsis with the rice orthologs reveals a degree of functional conservation during starch synthesis. PLoS ONE 2014, 9, e92174. [Google Scholar] [CrossRef]
- Borén, M.; Larsson, H.; Falk, A.; Jansson, C. The barley starch granule proteome - Internalized granule polypeptides of the mature endosperm. Plant Sci. 2004, 166, 617–626. [Google Scholar] [CrossRef]
- Denyer, K.; Sidebottom, C.; Hylton, C.M.; Smith, A.M. Soluble isoforms of starch synthase and starch-branching enzyme also occur within starch granules in developing pea embryos. Plant J. 1993, 4, 191–198. [Google Scholar] [CrossRef] [PubMed]
- Grimaud, F.; Rogniaux, H.; James, M.G.; Myers, A.M.; Planchot, V. Proteome and phosphoproteome analysis of starch granule-associated proteins from normal maize and mutants affected in starch biosynthesis. J. Exp. Bot. 2008, 59, 3395–3406. [Google Scholar] [CrossRef] [PubMed]
- Peng, M.; Gao, M.; Båga, M.; Hucl, P.; Chibbar, R.N. Starch-branching enzymes preferentially associated with A-type starch granules in wheat endosperm. Plant Physiol. 2000, 124, 265–272. [Google Scholar] [CrossRef] [PubMed]
- Rahman, S.; Kosar-Hashemi, B.; Samuel, M.S.; Hill, A.; Abbott, D.C.; Skerritt, J.H.; Preiss, J.; Appels, R.; Morell, M.K. The major proteins of wheat endosperm starch granules. Funct. Plant Biol. 1995, 22, 793–803. [Google Scholar] [CrossRef]
- Regina, A.; Kosar-Hashemi, B.; Li, Z.; Pedler, A.; Mukai, Y.; Yamamoto, M.; Gale, K.; Sharp, P.J.; Morell, M.K.; Rahman, S. Starch branching enzyme IIb in wheat is expressed at low levels in the endosperm compared to other cereals and encoded at a non-syntenic locus. Planta 2005, 222, 899–909. [Google Scholar] [CrossRef] [PubMed]
- Stensballe, A.; Hald, S.; Bauw, G.; Blennow, A.; Welinder, K.G. The amyloplast proteome of potato tuber. FEBS J. 2008, 275, 1723–1741. [Google Scholar] [CrossRef] [PubMed]
- Xing, S.; Meng, X.; Zhou, L.; Mujahid, H.; Zhao, C.; Zhang, Y.; Wang, C.; Peng, Z. Proteome profile of starch granules purified from rice (Oryza sativa) endosperm. PLoS ONE 2016, 11, e0168467. [Google Scholar] [CrossRef] [PubMed]
- Ritte, G.; Lloyd, J.R.; Eckermann, N.; Rottmann, A.; Kossmann, J.; Steup, M. The starch-related R1 protein is an alpha -glucan, water dikinase. Proc. Natl. Acad. Sci. USA 2002, 99, 7166–7171. [Google Scholar] [CrossRef] [PubMed]
- Kötting, O.; Pusch, K.; Tiessen, A.; Geigenberger, P.; Steup, M.; Ritte, G. Identification of a novel enzyme required for starch metabolism in Arabidopsis leaves. The phosphoglucan, water dikinase. Plant Physiol. 2005, 137, 242–252. [Google Scholar] [CrossRef]
- Niittylä, T.; Comparot-Moss, S.; Lue, W.-L.; Messerli, G.; Trevisan, M.; Seymour, M.D.J.; Gatehouse, J.A.; Villadsen, D.; Smith, S.M.; Chen, J.; et al. Similar protein phosphatases control starch metabolism in plants and glycogen metabolism in mammals. J. Biol. Chem. 2006, 281, 11815–11818. [Google Scholar] [CrossRef] [PubMed]
- Santelia, D.; Kötting, O.; Seung, D.; Schubert, M.; Thalmann, M.; Bischof, S.; Meekins, D.A.; Lutz, A.; Patron, N.; Gentry, M.S.; et al. The phosphoglucan phosphatase like sex Four2 dephosphorylates starch at the C3-position in Arabidopsis. Plant Cell 2011, 23, 4096–4111. [Google Scholar] [CrossRef] [PubMed]
- Sokolov, L.N.; Dominguez-Solis, J.R.; Allary, A.-L.; Buchanan, B.B.; Luan, S. A redox-regulated chloroplast protein phosphatase binds to starch diurnally and functions in its accumulation. Proc. Natl. Acad. Sci. USA 2006, 103, 9732–9737. [Google Scholar] [CrossRef] [PubMed]
- Seung, D.; Soyk, S.; Coiro, M.; Maier, B.A.; Eicke, S.; Zeeman, S.C. PROTEIN TARGETING TO STARCH is required for localising GRANULE-BOUND STARCH SYNTHASE to starch granules and for normal amylose synthesis in Arabidopsis. PLoS Biol. 2015, 13, e1002080. [Google Scholar] [CrossRef] [PubMed]
- Feike, D.; Seung, D.; Graf, A.; Bischof, S.; Ellick, T.; Coiro, M.; Soyk, S.; Eicke, S.; Mettler-Altmann, T.; Lu, K.J.; et al. The starch granule-associated protein EARLY STARVATION1 is required for the control of starch degradation in Arabidopsis thaliana leaves. Plant Cell 2016, 28, 1472–1489. [Google Scholar] [CrossRef] [PubMed]
- Seung, D.; Boudet, J.; Monroe, J.; Schreier, T.B.; David, L.C.; Abt, M.; Lu, K.-J.; Zanella, M.; Zeeman, S.C. Homologs of PROTEIN TARGETING TO STARCH control starch granule initiation in Arabidopsis leaves. Plant Cell 2017, 29, 1657–1677. [Google Scholar] [CrossRef] [PubMed]
- Valerio, C.; Costa, A.; Marri, L.; Issakidis-Bourguet, E.; Pupillo, P.; Trost, P.; Sparla, F. Thioredoxin-regulated β-amylase (BAM1) triggers diurnal starch degradation in guard cells, and in mesophyll cells under osmotic stress. J. Exp. Bot. 2011, 62, 545–555. [Google Scholar] [CrossRef] [PubMed]
- Skryhan, K.; Cuesta-Seijo, J.A.; Nielsen, M.M.; Marri, L.; Mellor, S.B.; Glaring, M.A.; Jensen, P.E.; Palcic, M.M.; Blennow, A. The role of cysteine residues in redox regulation and protein stability of Arabidopsis thaliana starch synthase 1. PLoS ONE 2015, 10, e0136997. [Google Scholar] [CrossRef] [PubMed]
- Noctor, G.; Mhamdi, A.; Chaouch, S.; Han, Y.; Neukermans, J.; Marquez-Garcia, B.; Queval, G.; Foyer, C.H. Glutathione in plants: An integrated overview. Plant Cell Environ. 2012, 35, 454–484. [Google Scholar] [CrossRef]
- Pfister, B.; Zeeman, S.C. Formation of starch in plant cells. Cell. Mol. Life Sci. CMLS 2016, 73, 2781–2807. [Google Scholar] [CrossRef]
- Jane, J.-L.; Kasemsuwan, T.; Leas, S.; Zobel, H.; Robyt, J.F. Anthology of starch granule morphology by scanning electron microscopy. Starch Stärke 1994, 46, 121–129. [Google Scholar] [CrossRef]
- Tanaka, E.; Ral, J.-P.F.; Li, S.; Gaire, R.; Cavanagh, C.R.; Cullis, B.R.; Whan, A. Increased accuracy of starch granule type quantification using mixture distributions. Plant Methods 2017, 13, 107. [Google Scholar] [CrossRef] [PubMed]
- Tester, R.F.; Karkalas, J.; Qi, X. Starch structure and digestibility Enzyme-Substrate relationship. Worlds Poult. Sci. J. 2004, 60, 186–195. [Google Scholar] [CrossRef]
- Wang, C.; Tang, C.-H.; Fu, X.; Huang, Q.; Zhang, B. Granular size of potato starch affects structural properties, octenylsuccinic anhydride modification and flowability. Food Chem. 2016, 212, 453–459. [Google Scholar] [CrossRef] [PubMed]
- BeMiller, J.N.; Whistler, R.L. Starch: Chemistry and Technology; Elsevier Science: Amsterdam, The Netherlands, 2009; pp. 213–217. ISBN 978-0-12-746275-2. [Google Scholar]
- Santelia, D.; Lunn, J.E. Transitory starch metabolism in guard cells: Unique features for a unique function. Plant Physiol. 2017, 174, 539–549. [Google Scholar] [CrossRef] [PubMed]
- Crumpton-Taylor, M.; Grandison, S.; Png, K.M.Y.; Bushby, A.J.; Smith, A.M. Control of starch granule numbers in Arabidopsis chloroplasts. Plant Physiol. 2012, 158, 905–916. [Google Scholar] [CrossRef] [PubMed]
- Jane, J.; Shen, J.J. Internal structure of the potato starch granule revealed by chemical gelatinization. Carbohydr. Res. 1993, 247, 279–290. [Google Scholar] [CrossRef]
- Tetlow, I.J.; Beisel, K.G.; Cameron, S.; Makhmoudova, A.; Liu, F.; Bresolin, N.S.; Wait, R.; Morell, M.K.; Emes, M.J. Analysis of protein complexes in wheat amyloplasts reveals functional interactions among starch biosynthetic enzymes. Plant Physiol. 2008, 146, 1878–1891. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Ahmed, Z.; Lee, E.A.; Donner, E.; Liu, Q.; Ahmed, R.; Morell, M.K.; Emes, M.J.; Tetlow, I.J. Allelic variants of the amylose extender mutation of maize demonstrate phenotypic variation in starch structure resulting from modified protein–protein interactions. J. Exp. Bot. 2012, 63, 1167–1183. [Google Scholar] [CrossRef] [PubMed]
- Hayashi, M.; Crofts, N.; Oitome, N.F.; Fujita, N. Analyses of starch biosynthetic protein complexes and starch properties from developing mutant rice seeds with minimal starch synthase activities. BMC Plant Biol. 2018, 18, 59. [Google Scholar] [CrossRef] [PubMed]
- Yun, S.-H.; Matheson, N.K. Estimation of amylose content of starches after precipitation of amylopectin by concanavalin-A. Starch Stärke 1990, 42, 302–305. [Google Scholar] [CrossRef]
- Morrison, W.R.; Laignelet, B. An improved colorimetric procedure for determining apparent and total amylose in cereal and other starches. J. Cereal Sci. 1983, 1, 9–20. [Google Scholar] [CrossRef]
| Fraction | Average Diameter (µm) | Ellipsoid Roundness (%) | Protein Content (µg·mg−1) | Amylose Content (%) | λmax of the Amylopectin-I2 Complex (nm) |
|---|---|---|---|---|---|
| A | 14.2 ± 3.7 ** | 85.9 ± 12.5 ** | 2.6 ± 0.2 * | 22.6 ± 5.1 | 548 ± 2 |
| B | 24.5 ± 6.5 ** | 82.5 ± 13.6 ** | 2.8 ± 0.1 ** | 20.7 ± 2.1 | 546 ± 4 |
| C | 47.7 ± 12.8 ** | 72.0 ± 14.8 ** | 2.1 ± 0.4 * | 20.8 ± 2.9 | 552 ± 2 |
| D | 61.8 ± 17.4 ** | 73.2 ± 15.0 ** | 2.4 ± 0.1 * | 20.6 ± 1.9 | 549 ± 2 |
| U.S. | 30.7 ± 15.9 | 79.9 ± 14.6 | 2.5 ± 0.2 | 21.5 ± 1.0 | 554 ± 4 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Helle, S.; Bray, F.; Putaux, J.-L.; Verbeke, J.; Flament, S.; Rolando, C.; D’Hulst, C.; Szydlowski, N. Intra-Sample Heterogeneity of Potato Starch Reveals Fluctuation of Starch-Binding Proteins According to Granule Morphology. Plants 2019, 8, 324. https://doi.org/10.3390/plants8090324
Helle S, Bray F, Putaux J-L, Verbeke J, Flament S, Rolando C, D’Hulst C, Szydlowski N. Intra-Sample Heterogeneity of Potato Starch Reveals Fluctuation of Starch-Binding Proteins According to Granule Morphology. Plants. 2019; 8(9):324. https://doi.org/10.3390/plants8090324
Chicago/Turabian StyleHelle, Stanislas, Fabrice Bray, Jean-Luc Putaux, Jérémy Verbeke, Stéphanie Flament, Christian Rolando, Christophe D’Hulst, and Nicolas Szydlowski. 2019. "Intra-Sample Heterogeneity of Potato Starch Reveals Fluctuation of Starch-Binding Proteins According to Granule Morphology" Plants 8, no. 9: 324. https://doi.org/10.3390/plants8090324
APA StyleHelle, S., Bray, F., Putaux, J.-L., Verbeke, J., Flament, S., Rolando, C., D’Hulst, C., & Szydlowski, N. (2019). Intra-Sample Heterogeneity of Potato Starch Reveals Fluctuation of Starch-Binding Proteins According to Granule Morphology. Plants, 8(9), 324. https://doi.org/10.3390/plants8090324





