Comparative and Phylogenetic Analyses of Ginger (Zingiber officinale) in the Family Zingiberaceae Based on the Complete Chloroplast Genome
Abstract
:1. Introduction
2. Results and Discussion
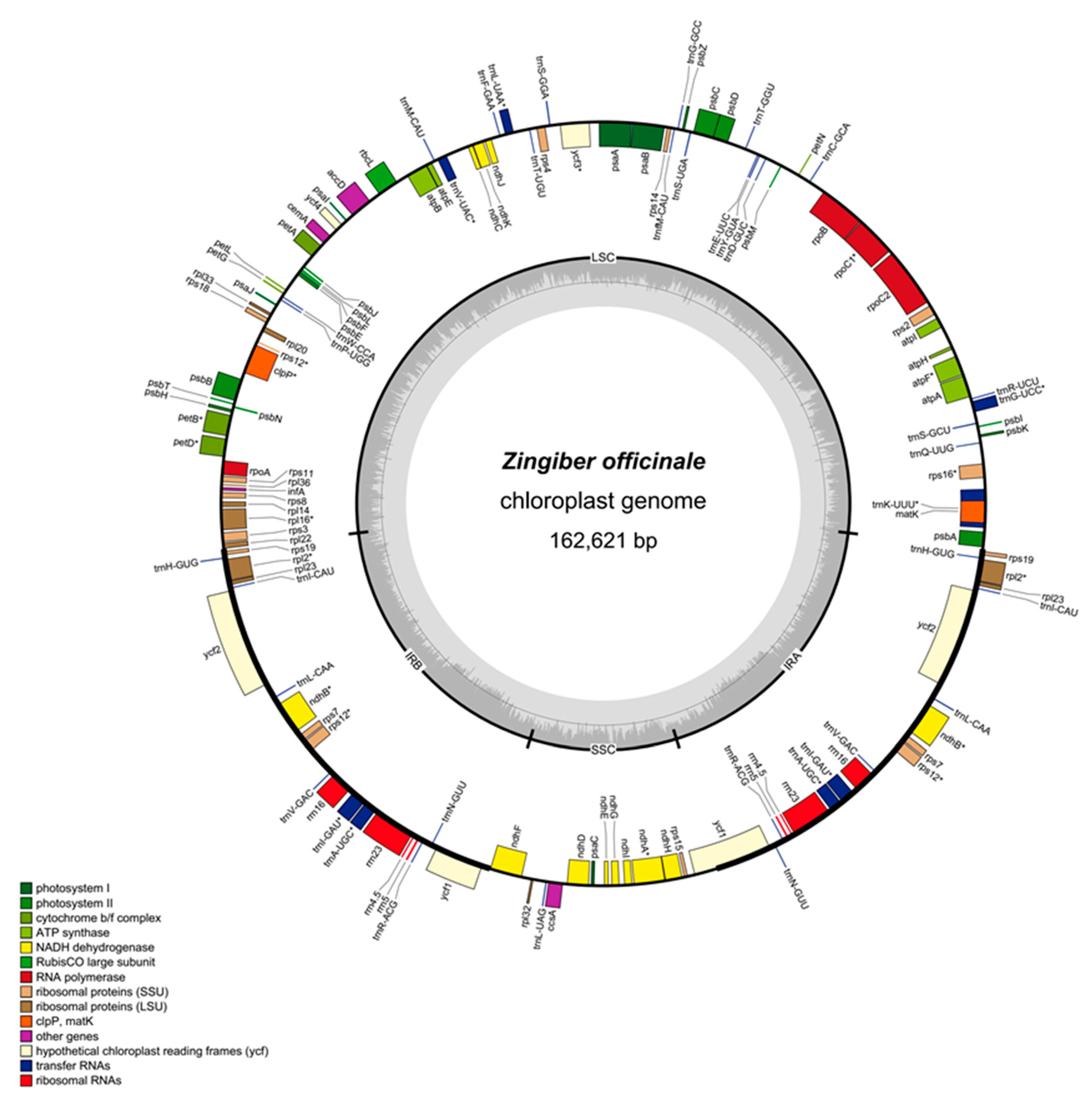
2.1. Zingiber Officinale Chloroplast Genome Features
2.2. Codon Usage Analysis
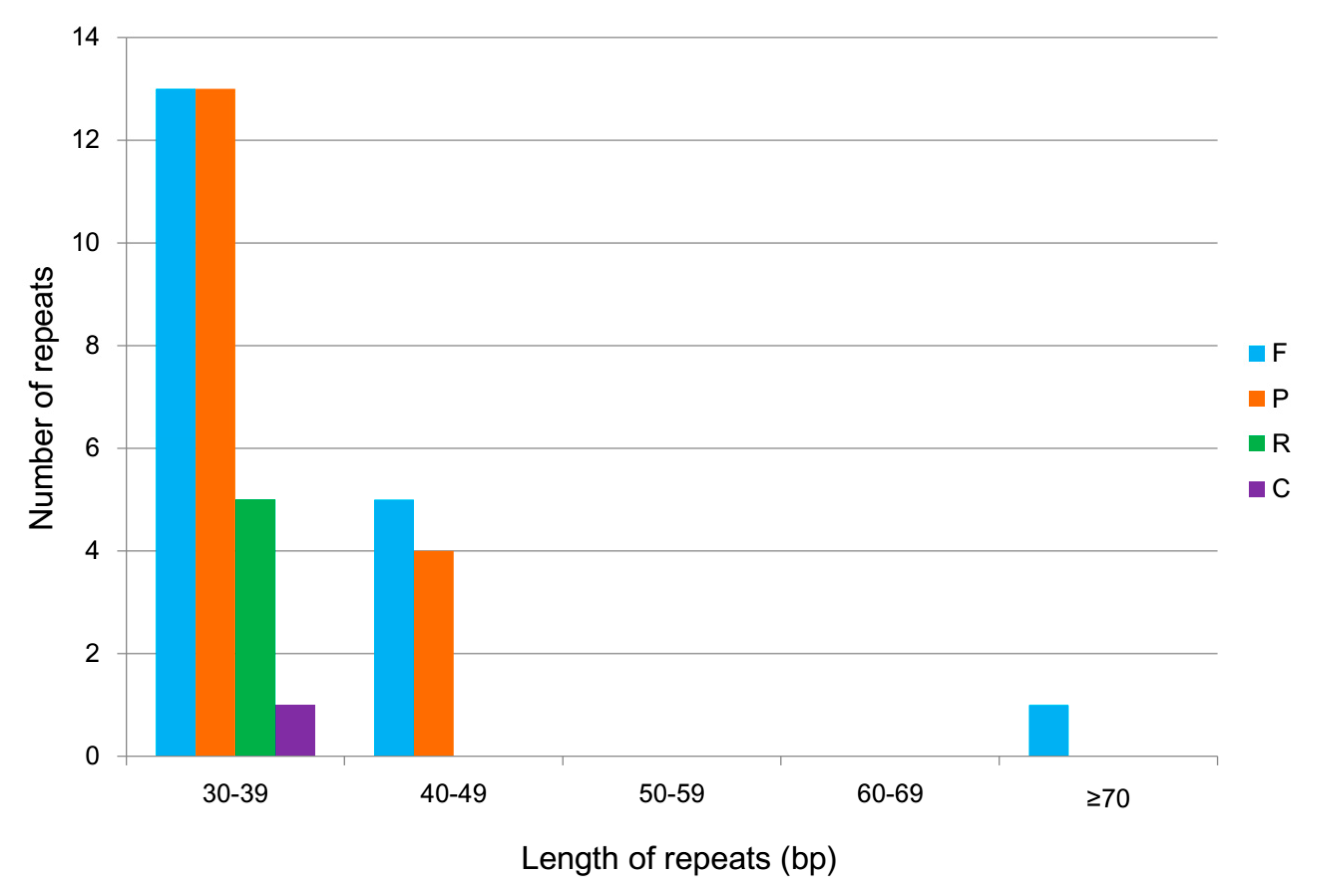
2.3. Repeat Structure Analysis
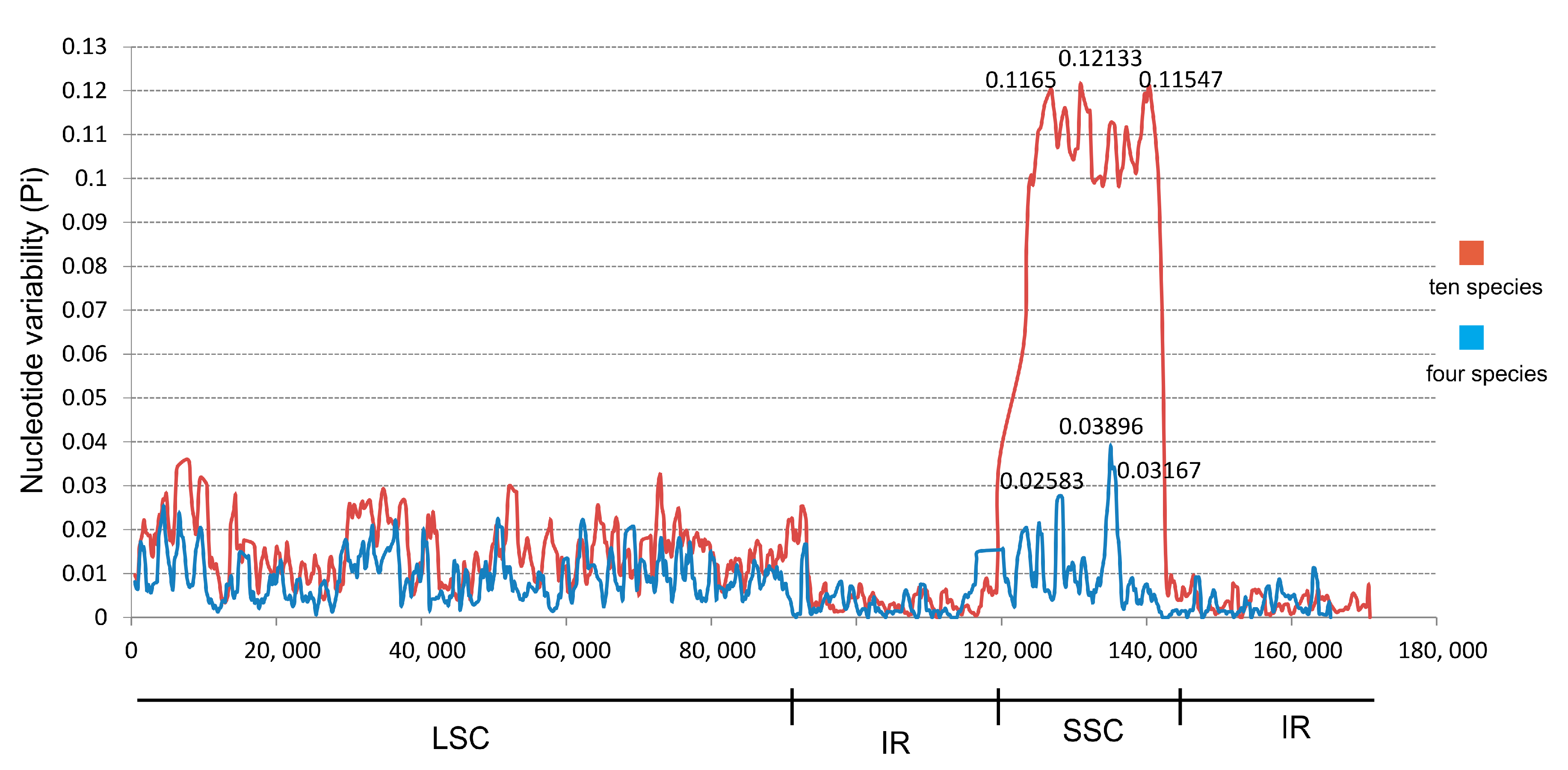
2.4. Comparative Analysis
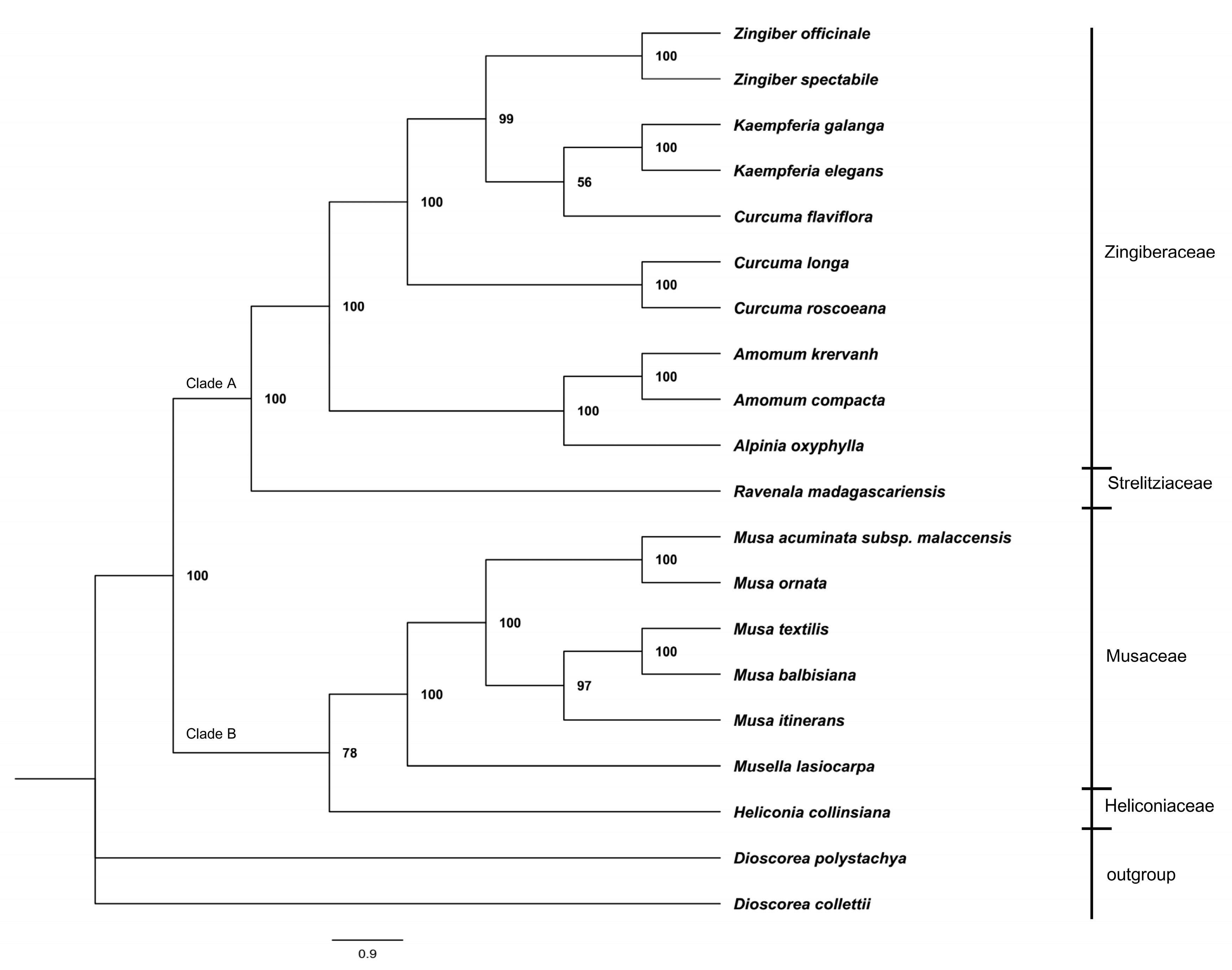
2.5. Phylogenetic Analysis
3. Materials and Methods
3.1. Plant and DNA Sources
3.2. Chloroplast Genome Assembly and Annotation
3.3. Chloroplast Genome Structure and Comparison Analysis
3.4. Phylogenetic Analyses
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| LSC | large single copy |
| SSC | small single copy |
| IR | inverted repeat |
| ML | maximum likelihood |
| SSR | Simple sequence repeats |
| ATP | Adenosine triphosphate |
| NADH | Nicotinamide adenine dinucleotide |
References
- Flora of China; Sci. Press: Beijing, China; Mo. Bot. Gard. Press: St. Louis, MO, USA, 2008; Volume 16, p. 141. Available online: http://www.efloras.org/flora_person_list.aspx?flora_id=2&volume_id=2016 (accessed on 16 August 2019).
- Commission, C.P. Pharmacopoeia of the People’s Republic of China; China Medical Science Press: Beijing, China, 2015. [Google Scholar]
- Kadnur, S.V.; Goyal, R.K. Beneficial effects of Zingiber officinale Roscoe on fructose induced hyperlipidemia and hyperinsulinemia in rats. Indian J. Exp. Biol. 2005, 43, 1161. [Google Scholar] [CrossRef] [PubMed]
- Thomson, M.; Al-Qattan, K.K.; Al-Sawan, S.M.; Alnaqeeb, M.A.; Khan, I.; Ali, M. The use of ginger (Zingiber officinale rosc.) As a potential anti-inflammatory and antithrombotic agent. Prostag. Leukotr. Ess. 2002, 67, 475–478. [Google Scholar] [CrossRef]
- Stoilova, I.; Krastanov, A.; Stoyanova, A.; Denev, P.; Gargova, S. Antioxidant activity of a ginger extract (Zingiber officinale). Food Chem. 2007, 102, 764–770. [Google Scholar] [CrossRef]
- Ahmed, R.S.; Seth, V.; Banerjee, B.D. Influence of dietary ginger (Zingiber officinales Rosc) on antioxidant defense system in rat: Comparison with ascorbic acid. Indian J. Exp. Biol. 2000, 38, 604–606. [Google Scholar] [PubMed]
- Habib, S.H.M.; Makpol, S.; Hamid, N.A.A.; Das, S.; Wan, Z.W.N.; Yusof, Y.A.M. Ginger extract (Zingiber officinale) has anti-cancer and anti-inflammatory effects on ethionine-induced hepatoma rats. Clinics 2008, 63, 807–813. [Google Scholar] [CrossRef] [PubMed]
- Pereira, M.M.; Haniadka, R.; Chacko, P.P.; Palatty, P.L.; Baliga, M.S. Zingiber officinale Roscoe (ginger) as an adjuvant in cancer treatment: A review. J. Buon 2011, 16, 414–424. [Google Scholar] [CrossRef]
- Phillips, S.; Ruggier, R.; Hutchinson, S.E. Zingiber officinale (ginger)-an antiemetic for day case surgery. Anaesthesia 2010, 48, 715–717. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.L.; Cheng, L.L.; Zhang, Z.X. Molecular cloning and characterization of violaxanthin de-epoxidase (VDE) in Zingiber officinale. Plant Sci. 2007, 172, 228–235. [Google Scholar] [CrossRef]
- Li, Y.; Zhou, J.; Chen, X.; Cui, Y.; Xu, Z.; Li, Y.; Song, J.; Duan, B.; Yao, H. Gene losses and partial deletion of small single-copy regions of the chloroplast genomes of two hemiparasitic Tax i llus species. Sci. Rep. 2017, 7, 12834. [Google Scholar] [CrossRef]
- Wolfe, K.H.; Li, W.H.; Sharp, P.M. Rates of nucleotide substitution vary greatly among plant mitochondrial, chloroplast, and nuclear DNAs. Proc. Natl. Acad. Sci. USA 1987, 84, 9054–9058. [Google Scholar] [CrossRef] [PubMed]
- Drouin, G.; Daoud, H.; Xia, J. Relative rates of synonymous substitutions in the mitochondrial, chloroplast and nuclear genomes of seed plants. Mol. Phylogenetics Evol. 2008, 49, 827–831. [Google Scholar] [CrossRef] [PubMed]
- Smith, D.R. Mutation Rates in Plastid Genomes: They Are Lower than You Might Think. Genome Biol. Evol. 2015, 7, 1227–1234. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chumley, T.W.; Palmer, J.D.; Mower, J.P.; Fourcade, H.M.; Calie, P.J.; Boore, J.L.; Jansen, R.K. The complete chloroplast genome sequence of Pelargonium x hortorum: Organization and evolution of the largest and most highly rearranged chloroplast genome of land plants. Mol. Biol. Evol. 2006, 23, 2175–2190. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Nie, J.; Xiao, L.; Hu, Z.; Wang, B. Comparative Chloroplast Genome Analysis of Rhubarb Botanical Origins and the Development of Specific Identification Markers. Molecules 2018, 23, 2811. [Google Scholar] [CrossRef] [PubMed]
- Park, I.; Yang, S.; Kim, W.J.; Noh, P.; Lee, H.O.; Moon, B.C. The Complete Chloroplast Genomes of Six Ipomoea Species and Indel Marker Development for the Discrimination of Authentic Pharbitidis Semen (Seeds of I. nil or I. purpurea). Front. Plant Sci. 2018, 9, 965. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Cui, Y.; Nie, L.; Hu, H.; Xu, Z.; Sun, W.; Gao, T.; Song, J.; Yao, H. Identification and Phylogenetic Analysis of the Complete Chloroplast Genomes of Three Ephedra Herbs Containing Ephedrine. Biomed. Res. Int. 2019, 2019, 5921725. [Google Scholar] [CrossRef] [PubMed]
- Lemieux, C.; Otis, C.; Turmel, M. Ancestral chloroplast genome in Mesostigma viride reveals an early branch of green plant evolution. Nature 2000, 403, 649–652. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhu, S.; Niu, Z.; Xue, Q.; Wang, H.; Xie, X.; Ding, X. Accurate authentication of Dendrobium officinale and its closely related species by comparative analysis of complete plastomes. Acta Pharm. Sin. B 2018, 8, 969–980. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.L.; Li, Q.; Xu, J.; Li, X.W. Complete chloroplast genome of the medicinal plant Amomum compactum: gene organization, comparative analysis, and phylogenetic relationships within Zingiberales. Chin. Med. 2018, 13, 1875. [Google Scholar] [CrossRef] [PubMed]
- Moore, M.J.; Soltis, P.S.; Bell, C.D.; J Gordon, B.; Soltis, D.E. Phylogenetic analysis of 83 plastid genes further resolves the early diversification of eudicots. PNAS 2010, 107, 4623–4628. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xiang, B.; Li, X.; Qian, J.; Wang, L.; Ma, L.; Tian, X.; Wang, Y. The Complete Chloroplast Genome Sequence of the Medicinal Plant Swertia mussotii Using the PacBio RS II Platform. Molecules 2016, 21, 1029. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.; Li, Q.; Hu, Z.; Li, X.; Chen, S. The Complete Amomum kravanh Chloroplast Genome Sequence and Phylogenetic Analysis of the Commelinids. Molecules 2017, 22, 1875. [Google Scholar] [CrossRef] [PubMed]
- Jiang, D.; Zhao, Z.; Zhang, T.; Zhong, W.; Liu, C.; Yuan, Q.; Huang, L. The Chloroplast Genome Sequence of Scutellaria baicalensis Provides Insight into Intraspecific and Interspecific Chloroplast Genome Diversity in Scutellaria. Genes (Basel) 2017, 8, 227. [Google Scholar] [CrossRef]
- Zuo, L.H.; Shang, A.Q.; Zhang, S.; Yu, X.Y.; Ren, Y.C.; Yang, M.S.; Wang, J.M. The first complete chloroplast genome sequences of Ulmus species by de novo sequencing: Genome comparative and taxonomic position analysis. PLoS ONE 2017, 12, e0171264. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Cui, Y.; Chen, X.; Li, Y.; Xu, Z.; Duan, B.; Li, Y.; Song, J.; Yao, H. Complete Chloroplast Genomes of Papaver rhoeas and Papaver orientale: Molecular Structures, Comparative Analysis and Phylogenetic Analysis. Molecules 2018, 23, 437. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Huang, Y.; An, W.; Zheng, X.; Huang, S.; Liang, L. Sequencing and Structural Analysis of the Complete Chloroplast Genome of the Medicinal Plant Lycium chinense Mill. Plants (Basel) 2019, 8, 87. [Google Scholar] [CrossRef] [PubMed]
- Powell, W.; Morgante, M.; Mcdevitt, R.; Vendramin, G.G.; Rafalski, J.A. Polymorphic simple sequence repeat regions in chloroplast genomes: Applications to the population genetics of pines. PNAS 1995, 92, 7759–7763. [Google Scholar] [CrossRef] [PubMed]
- Yang, A.H.; Zhang, J.J.; Yao, X.H.; Huang, H.W. Chloroplast microsatellite markers in Liriodendron tulipifera (Magnoliaceae) and cross-species amplification in L. chinense. Am. J. Bot. 2011, 98, 123–126. [Google Scholar] [CrossRef] [PubMed]
- Jiao, Y.J.H.; Li, X.W.; Chai, M.L.; Jia, H.J.; Chen, Z.; Wang, G.Y.; Chai, C.Y.; van de Weg, E.; Gao, Z.S. Development of simple sequence repeat (SSR) markers from a genome survey of Chinese bayberry (Myrica rubra). BMC Genom. 2012, 13, 201. [Google Scholar] [CrossRef]
- Jianhua, X.; Shuo, W.; Shi-Liang, Z. Polymorphic chloroplast microsatellite loci in Nelumbo (Nelumbonaceae). Am. J. Bot. 2012, 99, 240–244. [Google Scholar] [CrossRef]
- Park, I.; Yang, S.; Choi, G.; Kim, W.J.; Moon, B.C. The Complete Chloroplast Genome Sequences of Aconitum pseudolaeve and Aconitum longecassidatum, and Development of Molecular Markers for Distinguishing Species in the Aconitum Subgenus Lycoctonum. Molecules 2017, 22, 2012. [Google Scholar] [CrossRef] [PubMed]
- Tonti-Filippini, J.; Nevill, P.G.; Dixon, K.; Small, I. What can we do with 1000 plastid genomes? Plant J. 2017, 90, 808–818. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- NCBI. Genome. Available online: https://www.ncbi.nlm.nih.gov/genome/?term= (accessed on 26 June 2019).
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Jackman, S.D.; Vandervalk, B.P.; Mohamadi, H.; Chu, J.; Yeo, S.; Hammond, S.A.; Jahesh, G.; Khan, H.; Coombe, L.; Warren, R.L. ABySS 2.0: Resource-efficient assembly of large genomes using a Bloom filter. Genome Res. 2017, 27, 768–777. [Google Scholar] [CrossRef] [PubMed]
- Luo, R.; Liu, B.; Xie, Y.; Li, Z.; Huang, W.; Yuan, J.; He, G.; Chen, Y.; Pan, Q.; Liu, Y. SOAPdenovo2: An empirically improved memory-efficient short-read de novo assembler. Gigascience 2012, 1, 18. [Google Scholar] [CrossRef] [PubMed]
- Boetzer, M.; Henkel, C.V.; Jansen, H.J.; Butler, D.; Pirovano, W. Scaffolding pre-assembled contigs using SSPACE. Bioinformatics 2011, 27, 578–579. [Google Scholar] [CrossRef] [PubMed]
- Xu, G.C.; Xu, T.J.; Zhu, R.; Zhang, Y.; Li, S.Q.; Wang, H.W.; Li, J.T. LR_Gapcloser: A tiling path-based gap closer that uses long reads to complete genome assembly. Gigascience 2019, 8. [Google Scholar] [CrossRef]
- Liu, C.; Shi, L.; Zhu, Y.; Chen, H.; Zhang, J.; Lin, X.; Guan, X. CpGAVAS, an integrated web server for the annotation, visualization, analysis, and GenBank submission of completely sequenced chloroplast genome sequences. BMC Genomics 2012, 13, 715. [Google Scholar] [CrossRef]
- Wyman, S.K.; Jansen, R.K.; Boore, J.L. Automatic annotation of organellar genomes with DOGMA. Bioinformatics 2004, 20, 3252–3255. [Google Scholar] [CrossRef] [Green Version]
- Peter, S.; Brooks, A.N.; Lowe, T.M. The tRNAscan-SE, snoscan and snoGPS web servers for the detection of tRNAs and snoRNAs. Nucleic Acids Res. 2005, 33, 686–689. [Google Scholar] [CrossRef]
- Lohse, M.; Drechsel, O.; Bock, R. OrganellarGenomeDRAW (OGDRAW): A tool for the easy generation of high-quality custom graphical maps of plastid and mitochondrial genomes. Curr. Genet. 2007, 52, 267–274. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [PubMed]
- Beier, S.; Thiel, T.; Münch, T.; Scholz, U.; Mascher, M. MISA-web: A web server for microsatellite prediction. Bioinformatics 2017, 33, 2583–2585. [Google Scholar] [CrossRef] [PubMed]
- Kurtz, S.; Schleiermacher, C. REPuter: Fast computation of maximal repeats in complete genomes. Bioinformatics 1999, 15, 426–427. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Rozewicki, J.; Yamada, K.D. MAFFT online service: Multiple sequence alignment, interactive sequence choice and visualization. Brief. Bioinform. 2017. [Google Scholar] [CrossRef] [PubMed]
- Frazer, K.A.; Lior, P.; Alexander, P.; Rubin, E.M.; Inna, D. VISTA: Computational tools for comparative genomics. Nucleic Acids Res. 2004, 32. [Google Scholar] [CrossRef]
- Rozas, J.; Ferrermata, A.; Sánchezdelbarrio, J.C.; Guiraorico, S.; Librado, P.; Ramosonsins, S.E.; Sánchezgracia, A. Evolution. DnaSP 6: DNA Sequence Polymorphism Analysis of Large Datasets. Mol. Biol. 2017, 34, 3299–3302. [Google Scholar] [CrossRef]
- Nguyen, L.T.; Schmidt, H.A.; Von Haeseler, A.; Minh, B.Q. IQ-TREE: A Fast and Effective Stochastic Algorithm for Estimating Maximum-Likelihood Phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef]


| Region | Positions | T(U) (%) | C (%) | A (%) | G (%) | Length (bp) |
|---|---|---|---|---|---|---|
| Total | 32.2 | 18.4 | 31.7 | 17.7 | 162,621 | |
| IRa | 30.1 | 21.3 | 28.8 | 19.8 | 29,779 | |
| IRb | 28.8 | 19.8 | 30.1 | 21.3 | 29,779 | |
| LSC | 33.7 | 17.4 | 32.4 | 16.5 | 87,486 | |
| SSC | 34.3 | 15.8 | 36.0 | 13.9 | 15,577 | |
| CDS | 31.7 | 17.2 | 31.2 | 19.9 | 79,167 | |
| 1st position | 24 | 18.3 | 31.3 | 26.5 | 26,389 | |
| 2nd position | 33 | 20.1 | 29.7 | 17.7 | 26,389 | |
| 3rd position | 39 | 13.2 | 32.6 | 15.6 | 26,389 |
| Group of Genes | Gene Names | Amount |
|---|---|---|
| Photosystem I | psaA, psaB, psaC, psaI, psaJ | 5 |
| Photosystem II | psbA, psbB, psbC, psbD, psbE, psbF, psbH, psbI, psbJ, psbK, psbL, psbM, psbN, psbT, psbZ | 15 |
| Cytochrome b/f complex | petA, petB*, petD*, petG, petL, petN | 6 |
| ATP synthase | atpA, atpB, atpE, atpF*, atpH, atpI | 6 |
| NADH dehydrogenase | ndhA*, ndhB*(×2), ndhC, ndhD, ndhE, ndhF, ndhG, ndhH, ndhI, ndhJ, ndhK | 12 |
| RubisCO large subunit | rbcL | 1 |
| RNA polymerase | rpoA, rpoB, rpoC1*, rpoC2 | 4 |
| Ribosomal proteins (SSU) | rps2, rps3, rps4, rps7(×2), rps8, rps11, rps12** (×2), rps14, rps15, rps16*, rps18, rps19(×2) | 15 |
| Ribosomal proteins (LSU) | rpl2*(×2), rpl14, rpl16*, rpl20, rpl22, rpl23(×2), rpl32, rpl33, rpl36 | 11 |
| Proteins of unknown function | ycf1(×2), ycf2(×2), ycf3**, ycf4 | 6 |
| Transfer RNAs | 38 tRNAs (8 in the IRs(×2)) ** | 38 |
| Ribosomal RNAs | rrn4.5(×2), rrn5(×2), rrn16(×2), rrn23(×2) | 8 |
| Other genes | accD, clpP*, matK, ccsA, cemA, infA | 6 |
| SSR Type | Repeat Unit | Amount | Ratio (%) |
|---|---|---|---|
| Mono | A/T | 27 | 100.0 |
| Di | AG/CT | 2 | 8.7 |
| AT/TA | 21 | 91.3 | |
| Tri | AAG/CTT | 1 | 20.0 |
| AAT/ATT | 4 | 80.0 | |
| Tetra | AAAC/GTTT | 1 | 4.8 |
| AAAG/CTTT | 4 | 19.0 | |
| AAAT/ATTT | 13 | 61.9 | |
| AACT/AGTT | 1 | 4.8 | |
| AATG/ATTC | 2 | 9.5 | |
| Penta | AAAAT/ATTTT | 1 | 50.0 |
| AATAG/ATTCT | 1 | 50.0 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cui, Y.; Nie, L.; Sun, W.; Xu, Z.; Wang, Y.; Yu, J.; Song, J.; Yao, H. Comparative and Phylogenetic Analyses of Ginger (Zingiber officinale) in the Family Zingiberaceae Based on the Complete Chloroplast Genome. Plants 2019, 8, 283. https://doi.org/10.3390/plants8080283
Cui Y, Nie L, Sun W, Xu Z, Wang Y, Yu J, Song J, Yao H. Comparative and Phylogenetic Analyses of Ginger (Zingiber officinale) in the Family Zingiberaceae Based on the Complete Chloroplast Genome. Plants. 2019; 8(8):283. https://doi.org/10.3390/plants8080283
Chicago/Turabian StyleCui, Yingxian, Liping Nie, Wei Sun, Zhichao Xu, Yu Wang, Jing Yu, Jingyuan Song, and Hui Yao. 2019. "Comparative and Phylogenetic Analyses of Ginger (Zingiber officinale) in the Family Zingiberaceae Based on the Complete Chloroplast Genome" Plants 8, no. 8: 283. https://doi.org/10.3390/plants8080283
APA StyleCui, Y., Nie, L., Sun, W., Xu, Z., Wang, Y., Yu, J., Song, J., & Yao, H. (2019). Comparative and Phylogenetic Analyses of Ginger (Zingiber officinale) in the Family Zingiberaceae Based on the Complete Chloroplast Genome. Plants, 8(8), 283. https://doi.org/10.3390/plants8080283





