Factors Affecting the Regeneration, via Organogenesis, and the Selection of Transgenic Calli in the Peach Rootstock Hansen 536 (Prunus persica × Prunus amygdalus) to Express an RNAi Construct against PPV Virus
Abstract
1. Introduction
2. Results
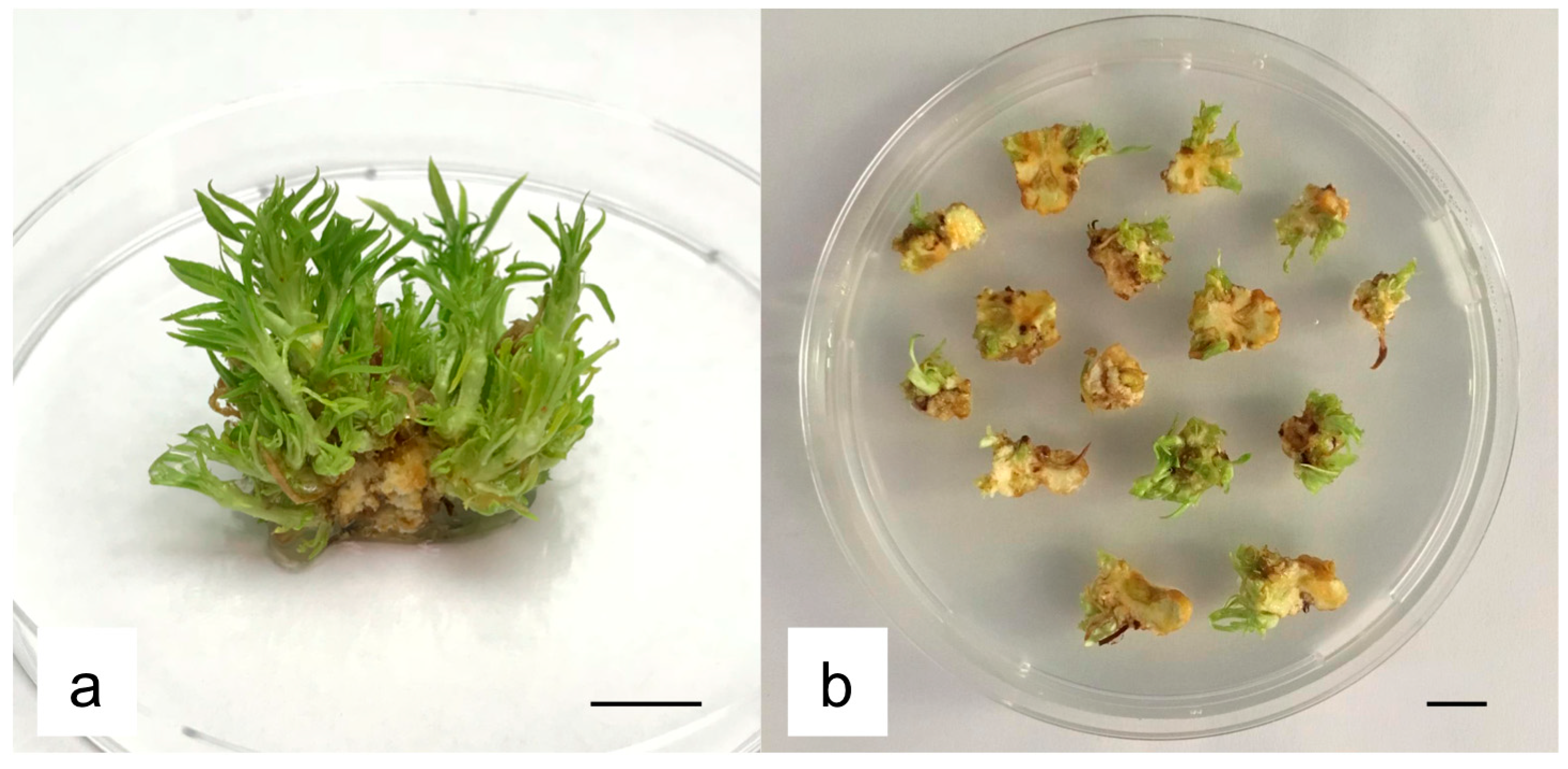
2.1. Influence of Basal Media, Growth Regulators and Gelling Agents on Regeneration Efficiency of MBs
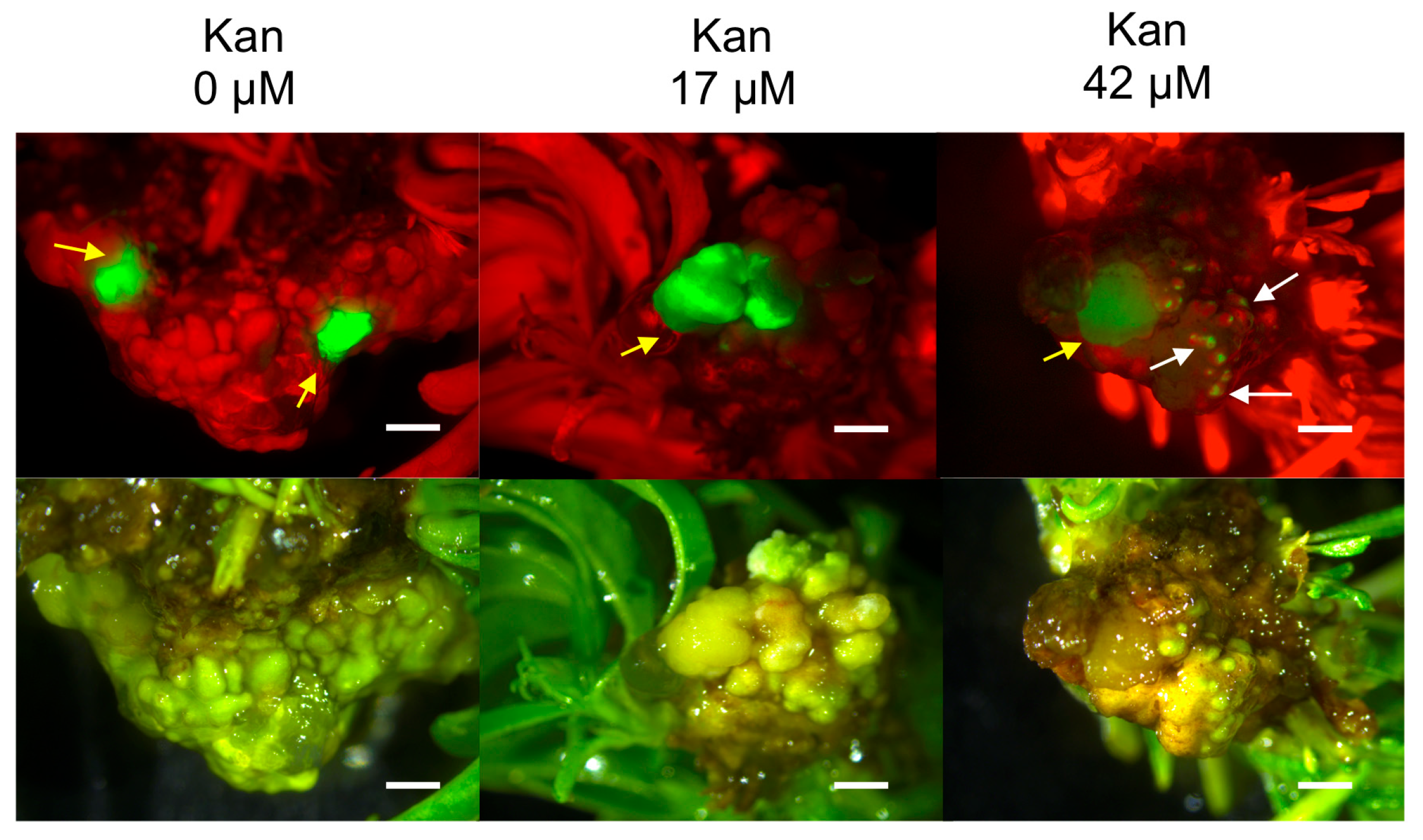
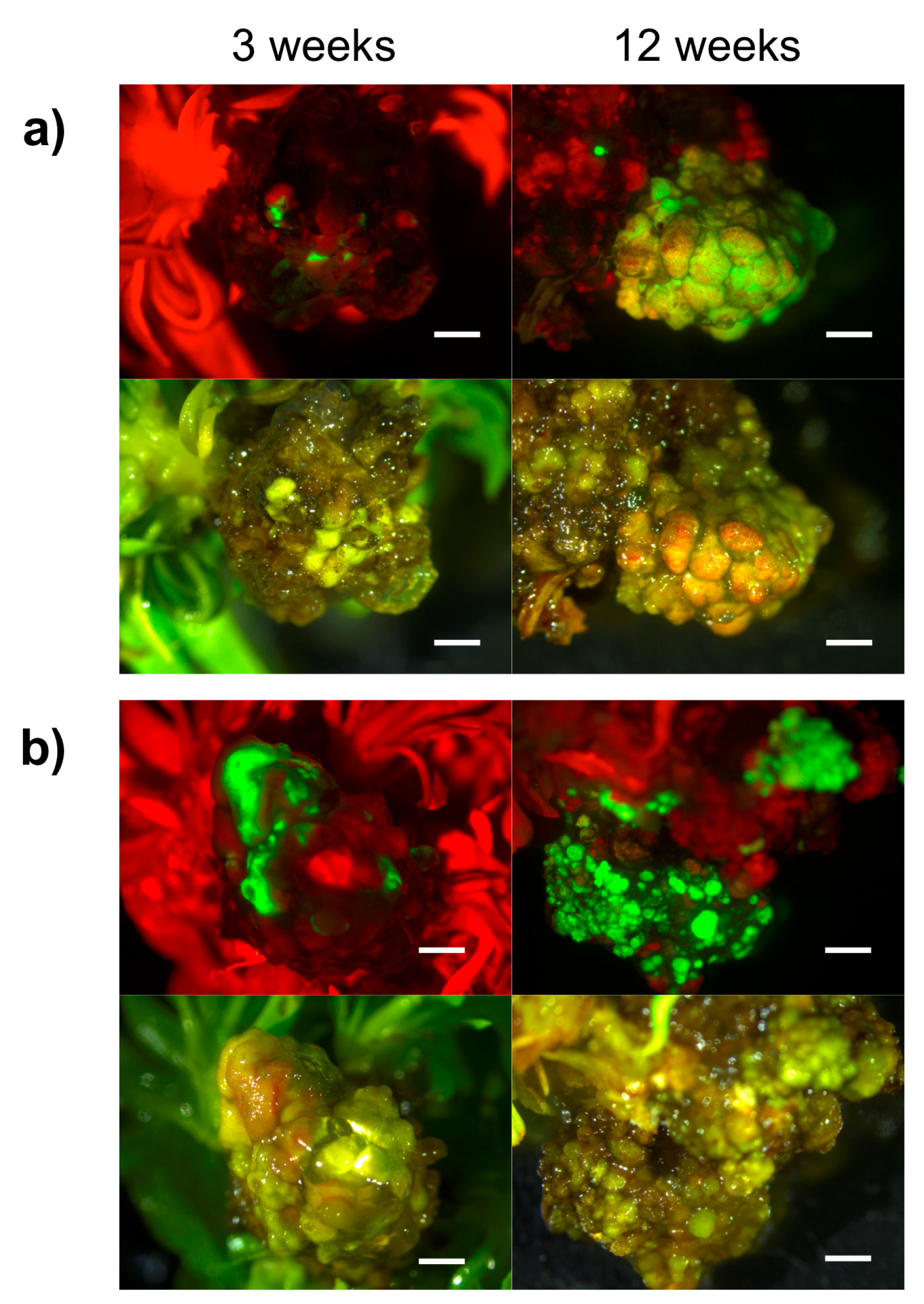
2.2. Hansen 536 MBs Transformation Efficiency and the Role of GFP and Kanamycin as Selectable Markers
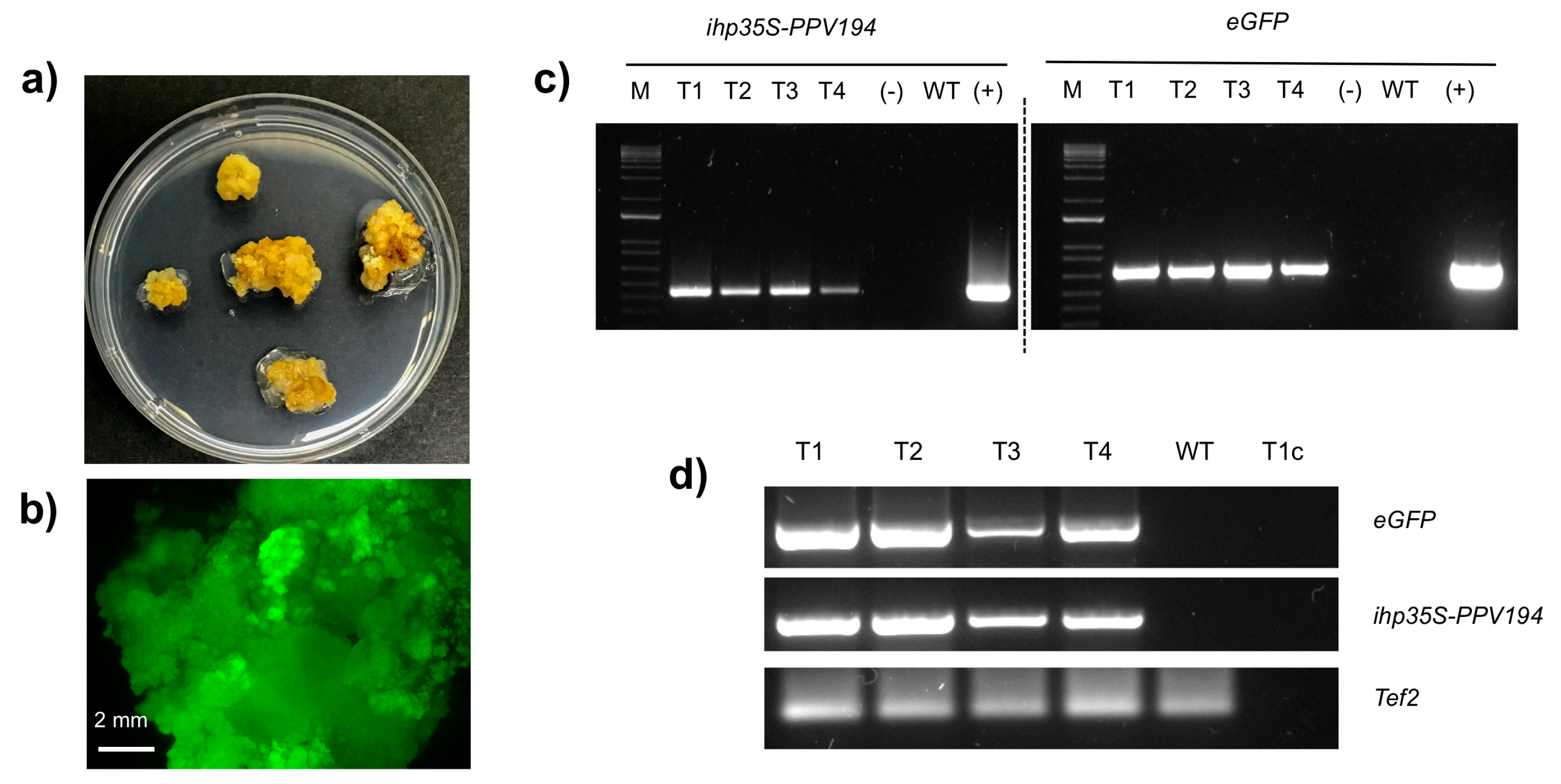
2.3. Molecular Analysis of Putatively Transgenic Callus Lines
3. Discussion
4. Material and Methods
4.1. Plant Material and Establishment of In Vitro Culture
4.2. Influence of Different Factors on the Regeneration Efficiency of Hansen 536 MBs
4.2.1. Effect of Different Basal Media and Plant Growth Regulator Combinations
4.2.2. Effect of Different Gelling Agents on Shoot Regeneration Efficiency and Shoot Hyperhydricity
4.3. Genetic Transformation
4.3.1. Gene Construct and Agrobacterium Tumefaciens Strain
4.3.2. Genetic Transformation Protocol and Selection of GFP Fluorescing Calli
4.4. Molecular Analysis of GFP Fluorescing Calli
4.5. Statistical Analysis
Author Contributions
Funding
Conflicts of Interest
References
- Carrasco, B.; Meisel, L.; Gebauer, M.; Garcia-Gonzales, R.; Silva, H. Breeding in peach, cherry and plum: from a tissue culture, genetic, transcriptomic and genomic perspective. Biol. Res. 2013, 46, 219–230. [Google Scholar] [CrossRef] [PubMed]
- Petri, C.; Hily, J.-M.; Vann, C.; Dardick, C.; Scorza, R. A high-throughput transformation system allows the regeneration of marker-free plum plants (Prunus domestica). Ann. Appl. Biol. 2011, 159, 302–315. [Google Scholar] [CrossRef]
- García-Almodóvar, R.C.; Clemente-Moreno, M.J.; Díaz-Vivancos, P.; Petri, C.; Rubio, M.; Padilla, I.M.G.; Ilardi, V.; Burgos, L. Greenhouse evaluation confirms in vitro sharka resistance of genetically engineered h-UTR/P1 plum plants. Plant Cell Tissue Organ Cult. 2015, 120, 791–796. [Google Scholar] [CrossRef]
- Hily, J.-M.; Ravelonandro, M.; Damsteegt, V.; Bassett, C.; Petri, C.; Liu, Z.; Scorza, R. Plum Pox Virus Coat Protein Gene Intron-hairpin-RNA (ihpRNA) Constructs Provide Resistance to Plum Pox Virus in Nicotiana benthamiana and Prunus domestica. J. Am. Soc. Hortic. Sci. 2007, 132, 850–858. [Google Scholar] [CrossRef]
- Lemgo, G.N.Y.; Sabbadini, S.; Pandolfini, T.; Mezzetti, B. Biosafety considerations of RNAi-mediated virus resistance in fruit-tree cultivars and in rootstock. Transgenic Res. 2013, 22, 1073–1088. [Google Scholar] [CrossRef] [PubMed]
- Limera, C.; Sabbadini, S.; Sweet, J.B.; Mezzetti, B. New Biotechnological Tools for the Genetic Improvement of Major Woody Fruit Species. Front. Plant Sci. 2017, 8, 1418. [Google Scholar] [CrossRef] [PubMed]
- Padilla, I.M.G.; Golis, A.; Gentile, A.; Damiano, C.; Scorza, R. Evaluation of transformation in peach Prunus persica explants using green fluorescent protein (GFP) and beta-glucuronidase (GUS) reporter genes. Plant Cell Tissue Organ Cult. 2006, 84, 309–314. [Google Scholar] [CrossRef]
- Scorza, R.; Callahan, A.; Dardick, C.; Ravelonandro, M.; Polak, J.; Malinowski, T.; Zagrai, I.; Cambra, M.; Kamenova, I. Genetic engineering of Plum pox virus resistance: ‘HoneySweet’ plum—from concept to product. Plant Cell Tissue Organ Cult. 2013, 115, 1–12. [Google Scholar] [CrossRef]
- Sidorova, T.; Pushin, A.; Miroshnichenko, D.; Dolgov, S. Generation of Transgenic Rootstock Plum ((Prunus pumila L. × P. salicina Lindl.) × (P. cerasifera Ehrh.)) Using Hairpin-RNA Construct for Resistance to the Plum pox virus. Agronomy 2018, 8, 2. [Google Scholar] [CrossRef]
- Song, G.; Sink, K.C.; Walworth, A.E.; Cook, M.A.; Allison, R.F.; Lang, G.A. Engineering cherry rootstocks with resistance to Prunus necrotic ring spot virus through RNAi-mediated silencing. Plant Biotechnol. J. 2013, 11, 702–708. [Google Scholar] [CrossRef]
- Febres, V.J.; Lee, R.F.; Moore, G.A. Transgenic resistance to Citrus tristeza virus in grapefruit. Plant Cell Rep. 2008, 27, 93–104. [Google Scholar] [CrossRef] [PubMed]
- Pessina, S.; Angeli, D.; Martens, S.; Visser, R.G.F.; Bai, Y.; Salamini, F.; Velasco, R.; Schouten, H.J.; Malnoy, M. The knock-down of the expression of MdMLO19 reduces susceptibility to powdery mildew (Podosphaera leucotricha) in apple (Malus domestica). Plant Biotechnol. J. 2016, 14, 2033–2044. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Thomas, N.; Jin, H. Cross-kingdom RNA trafficking and environmental RNAi for powerful innovative pre- and post-harvest plant protection. Curr. Opin. Plant Biol. 2017, 38, 133–141. [Google Scholar] [CrossRef] [PubMed]
- Zotti, M.; dos Santos, E.A.; Cagliari, D.; Christiaens, O.; Taning, C.N.T.; Smagghe, G. RNA interference technology in crop protection against arthropod pests, pathogens and nematodes. Pest Manag. Sci. 2017, 74, 1239–1250. [Google Scholar] [CrossRef] [PubMed]
- Frizzi, A.; Huang, S. Tapping RNA silencing pathways for plant biotechnology. Plant Biotechnol. J. 2010, 8, 655–677. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Li, Y. Secondary siRNAs rescue virus-infected plants. Nat. Plants 2018, 4, 136–137. [Google Scholar] [CrossRef]
- Li, S.; Castillo-González, C.; Yu, B.; Zhang, X. The functions of plant small RNAs in development and in stress responses. Plant J. 2016, 90, 654–670. [Google Scholar] [CrossRef]
- Molesini, B.; Pii, Y.; Pandolfini, T. Fruit improvement using intragenesis and artificial microRNA. Trends Biotechnol. 2012, 30, 80–88. [Google Scholar] [CrossRef]
- Melnyk, C.W.; Molnar, A.; Baulcombe, D.C. Intercellular and systemic movement of RNA silencing signals. EMBO J. 2011, 30, 3553–3563. [Google Scholar] [CrossRef]
- Molnar, A.; Melnyk, C.; Baulcombe, D.C. Silencing signals in plants: a long journey for small RNAs. Genome Biol. 2011, 12, 215. [Google Scholar] [CrossRef]
- Reagan, B.C.; Ganusova, E.E.; Fernandez, J.C.; McCray, T.N.; Burch-Smith, T.M. RNA on the move: The plasmodesmata perspective. Plant Sci. 2018, 275, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Haroldsen, V.; Szczerba, M.; Aktas, H.; Lopez, J.; Odias, M.; Chi-Ham, C.; Labavitch, J.; Bennett, A.; Powell, A. Mobility of Transgenic Nucleic Acids and Proteins within Grafted Rootstocks for Agricultural Improvement. Front. Plant Sci. 2012, 3, 39. [Google Scholar] [CrossRef] [PubMed]
- Zhao, D.; Song, G. Rootstock-to-scion transfer of transgene-derived small interfering RNAs and their effect on virus resistance in nontransgenic sweet cherry. Plant Biotechnol. J. 2014, 12, 1319–1328. [Google Scholar] [CrossRef]
- Litz, R.E.; Padilla, G. Genetic Transformation of Fruit Trees BT. In Genomics of Tree Crops; Schnell, R.J., Priyadarshan, P.M., Eds.; Springer: New York, NY, USA, 2012; pp. 117–153. [Google Scholar]
- Rai, M.K.; Shekhawat, N.S. Recent advances in genetic engineering for improvement of fruit crops. Plant Cell Tissue Organ Cult. 2014, 116, 1–15. [Google Scholar] [CrossRef]
- Sgamma, T.; Thomas, B.; Muleo, R. Ethylene inhibitor silver nitrate enhances regeneration and genetic transformation of Prunus avium (L.) cv Stella. Plant Cell Tissue Organ Cult. 2015, 120, 79–88. [Google Scholar] [CrossRef]
- Hammerschlag, F.A.; Bauchan, G.; Scorza, R. Regeneration of peach plants from callus derived from immature embryos. Theor. Appl. Genet. 1985, 70, 248–251. [Google Scholar] [CrossRef] [PubMed]
- Bhansali, R.R.; Driver, J.A.; Durzan, D.J. Rapid multiplication of adventitious somatic embryos in peach and nectarine by secondary embryogenesis. Plant Cell Rep. 1990, 9, 280–284. [Google Scholar] [CrossRef]
- Scorza, R.; Morgens, P.H.; Cordts, J.M.; Mante, S.; Callahan, A.M. Agrobacterium-mediated transformation of peach (Prunus persica L. batsch) leaf segments, immature embryos, and long-term embryogenic callus. In Vitro Cell. Dev. Biol. 1990, 26, 829–834. [Google Scholar] [CrossRef]
- Smigocki, A.C.; Hammerschlag, F.A. Regeneration of Plants from Peach Embryo Cells Infected with a Shooty Mutant Strain of Agrobacterium. J. Am. Soc. Hortic. Sci. 1991, 116, 1092–1097. [Google Scholar] [CrossRef]
- Pérez-Clemente, R.M.; Pérez-Sanjuán, A.; García-Férriz, L.; Beltrán, J.-P.; Cañas, L.A. Transgenic peach plants (Prunus persica L.) produced by genetic transformation of embryo sections using the green fluorescent protein (GFP) as an in vivo marker. Mol. Breed. 2005, 14, 419–427. [Google Scholar] [CrossRef]
- Declerck, V.; Korban, S.S. Influence of growth regulators and carbon sources on callus induction, growth and morphogenesis from leaf tissues of peach (Prunus persica L. Batsch). J. Hortic. Sci. 1996, 71, 49–55. [Google Scholar] [CrossRef]
- Gentile, A.; Monticelli, S.; Damiano, C. Adventitious shoot regeneration in peach [Prunus persica (L.) Batsch]. Plant Cell Rep. 2002, 20, 1011–1016. [Google Scholar] [CrossRef]
- San, B.; Li, Z.; Hu, Q.; Reighard, G.L.; Luo, H. Adventitious shoot regeneration from in vitro cultured leaf explants of peach rootstock Guardian® is significantly enhanced by silver thiosulfate. Plant Cell Tissue Organ Cult. 2015, 120, 757–765. [Google Scholar] [CrossRef]
- Zhou, H.; Li, M.; Zhao, X.; Fan, X.; Guo, A. Plant regeneration from in vitro leaves of the peach rootstock ‘Nemaguard’ (Prunus persica × P. davidiana). Plant Cell Tissue Organ Cult. 2010, 101, 79–87. [Google Scholar] [CrossRef]
- Pérez-Jiménez, M.; López-Soto, M.B.; Cos-Terrer, J. In vitro callus induction from adult tissues of peach (Prunus persica L. Batsch). In Vitro Cell. Dev. Biol. Plant 2013, 49, 79–84. [Google Scholar] [CrossRef]
- Mezzetti, B.; Pandolfini, T.; Navacchi, O.; Landi, L. Genetic transformation of Vitis vinifera via organogenesis. BMC Biotechnol. 2002, 2, 18. [Google Scholar] [CrossRef]
- Girolomini, L.; Sabbadini, S.; Mezzetti, B.; Palma, D.; Pandolfini, T.; Polverari, A. Regeneration and genetic transformation via organogenesis of different cultivars of Vitis Vinifera and Prunus Persica. In Proceedings of the XXVIII International Horticultural Congress on Science and Horticulture for People (IHC2010): International Symposium on Genomics and Genetic Transformation of Horticultural Crops, Leuven, Belgium, 22–27 August 2012; pp. 393–396. [Google Scholar] [CrossRef]
- Pérez-Jiménez, M.; Carrillo-Navarro, A.; Cos-Terrer, J. Regeneration of peach (Prunus persica L. Batsch) cultivars and Prunus persica × Prunus dulcis rootstocks via organogenesis. Plant Cell Tissue Organ Cult. 2012, 108, 55–62. [Google Scholar] [CrossRef]
- Sabbadini, S.; Pandolfini, T.; Girolomini, L.; Molesini, B.; Navacchi, O. Peach (Prunus persica L.). In Agrobacterium Protocols; Wang, K., Ed.; Springer: New York, NY, USA, 2015; Volume 2, pp. 205–215. [Google Scholar]
- Cappelletti, R.; Sabbadini, S.; Mezzetti, B. The use of TDZ for the efficient in vitro regeneration and organogenesis of strawberry and blueberry cultivars. Sci. Hortic. (Amst.) 2016, 207, 117–124. [Google Scholar] [CrossRef]
- Sabbadini, S.; Capriotti, L.; Molesini, B.; Pandolfini, T.; Navacchi, O.; Limera, C.; Ricci, A.; Mezzetti, B. Comparison of regeneration capacity and Agrobacterium-mediated cell transformation efficiency of different cultivars and rootstocks of Vitis spp. via organogenesis. Sci. Rep. 2019, 9. [Google Scholar] [CrossRef]
- Xie, X.; Agüero, C.B.; Wang, Y.; Walker, M.A. Genetic transformation of grape varieties and rootstocks via organogenesis. Plant Cell Tissue Organ Cult. 2016, 126, 541–552. [Google Scholar] [CrossRef]
- Jiménez, V.M. Involvement of Plant Hormones and Plant Growth Regulators on in vitro Somatic Embryogenesis. Plant Growth Regul. 2005, 47, 91–110. [Google Scholar] [CrossRef]
- Pérez-Jiménez, M.; Cantero-Navarro, E.; Pérez-Alfocea, F.; Cos-Terrer, J. Endogenous hormones response to cytokinins with regard to organogenesis in explants of peach (Prunus persica L. Batsch) cultivars and rootstocks (P. persica × Prunus dulcis). Plant Physiol. Biochem. 2014, 84, 197–202. [Google Scholar] [CrossRef] [PubMed]
- Abdollahi, H.; Muleo, R.; Rugini, E. Optimisation of regeneration and maintenance of morphogenic callus in pear (Pyrus communis L.) by simple and double regeneration techniques. Sci. Hortic. (Amst.) 2006, 108, 352–358. [Google Scholar] [CrossRef][Green Version]
- Escalettes, V.; Dosba, F. In vitro adventitious shoot regeneration from leaves of Prunus spp. Plant Sci. 1993, 90, 201–209. [Google Scholar] [CrossRef]
- Pérez-Tornero, O.; Egea, J.; Vanoostende, A.; Burgos, L. Assessment of factors affecting adventitious shoot regeneration from in vitro cultured leaves of apricot. Plant Sci. 2000, 158, 61–70. [Google Scholar] [CrossRef]
- Petri, C.; Scorza, R. Factors affecting adventitious regeneration from in vitro leaf explants of ‘Improved French’ plum, the most important dried plum cultivar in the USA. Ann. Appl. Biol. 2009, 156, 79–89. [Google Scholar] [CrossRef]
- Tang, H.; Ren, Z.; Reustle, G.; Krczal, G. Plant regeneration from leaves of sweet and sour cherry cultivars. Sci. Hortic. (Amst.) 2002, 93, 235–244. [Google Scholar] [CrossRef]
- Burgos, L.; Alburquerque, N. Ethylene inhibitors and low kanamycin concentrations improve adventitious regeneration from apricot leaves. Plant Cell Rep. 2003, 21, 1167–1174. [Google Scholar] [CrossRef]
- Chevreau, E.; Mourgues, F.; Neveu, M.; Chevalier, M. Effect of gelling agents and antibiotics on adventitious bud regeneration from in vitro leaves of pear. In Vitro Cell. Dev. Biol. 1997, 33, 173–179. [Google Scholar] [CrossRef]
- Debergh, P.C. Effects of agar brand and concentration on the tissue culture medium. Physiol. Plant. 1983, 59, 270–276. [Google Scholar] [CrossRef]
- Alanagh, E.N.; Garoosi, G.; Haddad, R.; Maleki, S.; Landín, M.; Gallego, P.P. Design of tissue culture media for efficient Prunus rootstock micropropagation using artificial intelligence models. Plant Cell Tissue Organ Cult. 2014, 117, 349–359. [Google Scholar] [CrossRef]
- Brand, M.H. Agar and ammonium nitrate influence hyperhydricity, tissue nitrate and total nitrogen content of serviceberry (Amelanchier arborea) shoots in vitro. Plant Cell. Tissue Organ Cult. 1993, 35, 203–209. [Google Scholar] [CrossRef]
- Ivanova, M.; van Staden, J. Nitrogen source, concentration, and NH4+:NO3− ratio influence shoot regeneration and hyperhydricity in tissue cultured Aloe polyphylla. Plant Cell Tissue Organ Cult. 2009, 99, 167–174. [Google Scholar] [CrossRef]
- Pandolfini, T.; Molesini, B.; Avesani, L.; Spena, A.; Polverari, A. Expression of self-complementary hairpin RNA under the control of the rolCpromoter confers systemic disease resistance to plum pox virus without preventing local infection. BMC Biotechnol. 2003, 3, 7. [Google Scholar] [CrossRef] [PubMed]
- Tee, C.; Marziah, M.; Tan, C.; Abdullah, M. Evaluation of different promoters driving the GFP reporter gene and selected target tissues for particle bombardment of Dendrobium Sonia 17. Plant Cell Rep. 2003, 21, 452–458. [Google Scholar] [CrossRef] [PubMed]
- Ghorbel, R.; Juarez, J.; Navarro, L.; Pena, L. Green fluorescent protein as a screenable marker to increase the efficiency of generating transgenic woody fruit plants. Theor. Appl. Genet. 1999, 99, 350–358. [Google Scholar] [CrossRef]
- Gonzalez Padilla, I.M.; Webb, K.; Scorza, R. Early antibiotic selection and efficient rooting and acclimatization improve the production of transgenic plum plants (Prunus domestica L.). Plant Cell Rep. 2003, 22, 38–45. [Google Scholar] [CrossRef]
- Yau, Y.-Y.; Davis, S.J.; Ipek, A.; Simon, P.W. Early identification of stable transformation events by combined use of antibiotic selection and vital detection of green fluorescent protein (GFP) in carrot (Daucus carota L.) callus. Agric. Sci. China 2008, 7, 664–671. [Google Scholar] [CrossRef]
- Ballester, A.; Cervera, M.; Pena, L. Evaluation of selection strategies alternative to nptII in genetic transformation of citrus. Plant Cell Rep. 2008, 27, 1005–1015. [Google Scholar] [CrossRef]
- Breyer, D.; Kopertekh, L.; Reheul, D. Alternatives to antibiotic resistance marker genes for in vitro selection of genetically modified plants–scientific developments, current use, operational access and biosafety considerations. CRC Crit. Rev. Plant Sci. 2014, 33, 286–330. [Google Scholar] [CrossRef]
- Rosellini, D. Selectable markers and reporter genes: a well furnished toolbox for plant science and genetic engineering. CRC Crit. Rev. Plant Sci. 2012, 31, 401–453. [Google Scholar] [CrossRef]
- Ainsley, P.; Collins, G.; Sedgley, M. Factors affecting Agrobacterium-mediated gene transfer and the selection of transgenic calli in paper shell almond (Prunus dulcis Mill.). J. Hortic. Sci. Biotechnol. 2001, 76, 522–528. [Google Scholar]
- Zhao, T.; Wang, Z.; Su, L.; Sun, X.; Cheng, J.; Zhang, L.; Karungo, S.K.; Han, Y.; Li, S.; Xin, H. An efficient method for transgenic callus induction from Vitis amurensis petiole. PLoS ONE 2017, 12, e0179730. [Google Scholar] [CrossRef] [PubMed]
- Martínez-Márquez, A.; Morante-Carriel, J.A.; Ramírez-Estrada, K.; Cusidó, R.M.; Palazon, J.; Bru-Martínez, R. Production of highly bioactive resveratrol analogues pterostilbene and piceatannol in metabolically engineered grapevine cell cultures. Plant Biotechnol. J. 2016, 14, 1813–1825. [Google Scholar] [CrossRef] [PubMed]
- Murashige, T.; Skoog, F. A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol. Plant. 1962, 15, 473–497. [Google Scholar] [CrossRef]
- Quoirin, M.; Lepoivre, P. Etude de milieux adaptes aux cultures in vitro de Prunus. Acta Hortic. 1977, 78, 437–442. [Google Scholar] [CrossRef]
- Driver, J.A.; Kuniyuki, A.H. In vitro propagation of Paradox walnut rootstock [Juglans hindsii × Juglans regia, tissue culture]. HortScience (USA) 1984, 18, 507–509. [Google Scholar]
- Lloyd, G.; McCown, B. Commercially-feasible micropropagation of mountain laurel, Kalmia latifolia, by use of shoot-tip culture. Comb. Proc. Int. Plant Propag. Soc. 1980, 30, 421–427. [Google Scholar]
- Bevan, M. Binary Agrobacterium vectors for plant transformation. Nucleic Acids Res. 1984, 12, 8711–8721. [Google Scholar] [CrossRef]
- Karimi, M.; Inzé, D.; Depicker, A. GATEWAYTM vectors for Agrobacterium-mediated plant transformation. Trends Plant Sci. 2002, 7, 193–195. [Google Scholar] [CrossRef]
- Sambrook, J.; Fritsch, E.F.; Maniatis, T. Molecular Cloning: A Laboratory Manual; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 1989. [Google Scholar]
- Tong, Z.; Gao, Z.; Wang, F.; Zhou, J.; Zhang, Z. Selection of reliable reference genes for gene expression studies in peach using real-time PCR. BMC Mol. Biol. 2009, 10, 71. [Google Scholar] [CrossRef] [PubMed]

| Regeneration Media | Frequency of Regeneration (%) ± SE * | Average Number of Regenerated Shoots/MB ± SE |
|---|---|---|
| QL1 | 61.1 ± 5.5 (abc) | 1.8 ± 0.5 (defg) |
| QL2 | 83.3 ± 9.6 (a) | 6.4 ± 1.2 (ab) |
| QL3 | 61.1 ± 11.1 (abc) | 3.8 ± 1 (bcde) |
| QL4 | 50 ± 0 (abc) | 2.8 ± 1.9 (cdefg) |
| QL5 | 66.7 ± 9.6 (abc) | 4.3 ± 1.2 (bcd) |
| QL6 | 83.3 ± 16.7 (a) | 7.2 ± 1.3 (a) |
| DKW1 | 38.9 ± 5.5 (abc) | 1.4 ± 0.6 (defg) |
| DKW2 | 38.9 ± 22.2 (abc) | 1.3 ± 0.5 (defg) |
| DKW3 | 27.8 ± 5.5 (bc) | 0.8 ± 0.3 (fg) |
| DKW4 | 22.2 ± 11.1 (c) | 0.4 ± 0.2 (g) |
| DKW5 | 27.8 ± 5.5 (bc) | 1.2 ± 0.5 (efg) |
| DKW6 | 72.2 ± 14.7 (ab) | 3.5 ± 0.8 (bcdef) |
| WPMm1 | 61.1 ± 22.2 (abc) | 2.6 ± 0.7 (cdefg) |
| WPMm2 | 72.2 ± 11.1 (ab) | 5.5 ± 1.4 (abc) |
| WPMm3 | 38.9 ± 5.5 (abc) | 1.4 ± 0.5 (defg) |
| WPMm4 | 72.2 ± 11.1 (ab) | 3.3 ± 0.8 (cdefg) |
| WPMm5 | 77.8 ± 5.5 (a) | 4.2 ± 0.9 (bcde) |
| WPMm6 | 50 ± 25.4 (abc) | 2.1 ± 0.7 (defg) |
| Gelling Agent Concentration (g L−1) | Average Number of Regenerated Shoots/MB ± SE | Frequency of Vitrified Shoots (%) ± SE * | |||||
|---|---|---|---|---|---|---|---|
| QL2 | QL6 | WPMm5 | QL2 | QL6 | WPMm5 | ||
| PA | 7 | 6.7 ± 0.5 (ab) | 7.4 ± 1.3 (ab) | 4.4 ± 0.7 (b) | 82.6 ± 5.7 (b) | 98.5 ± 8.9 (a) | 59.2 ± 5.9 (cd) |
| B&V | 6 | 5.6 ± 1 (ab) | 8.9 ± 0.7 (a) | 7.8 ± 1.4 (a) | 53.7 ± 4.3 (d) | 64.3 ± 7.4 (c) | 22.8 ± 4.9 (f) |
| Kan Concentration (μM) | Number of Weeks after Co-Culture | Mean Number of GFP Spot/Explant ± SE | GFP MB (%) ± SE * | Number of MB Slices with Lzs | GFP MB Lzs (%) ± SE ** |
|---|---|---|---|---|---|
| 0 | 3 | 1.17 ± 0.37 (b) | 26.67 ± 5.75 (c) | 12 | 20 ± 5.21 (ns) |
| 12 | 0.82 ± 0.38 (A) | 6.67 ± 3.25 (A) | 2 | 3.33 ± 2.34 (NS) | |
| 17 | 3 | 2.10 ± 0.33 (ab) | 63.33 ± 6.5 (a) | 15 | 25 ± 5.64 (ns) |
| 12 | 0.78 ± 0.29 (A) | 13.33 ± 4.42 (A) | 4 | 6.67 ± 3.25 (NS) | |
| 42 | 3 | 3.17 ± 0.24 (a) | 48.33 ± 6.27 (b) | 22 | 36.67 ± 6.27 (ns) |
| 12 | 0.68 ± 0.36 (A) | 8.33 ± 3.60 (A) | 1 | 1.67 ± 1.67 (NS) |
| Culture Media | Basal Salts | Growth Regulators (μM) | |||
|---|---|---|---|---|---|
| BAP | IBA | TDZ | NAA | ||
| QL1 | QL | 4.4 | 0.5 | ||
| QL2 | QL | 8.9 | 0.5 | ||
| QL3 | QL | 4.4 | 0.2 | ||
| QL4 | QL | 8.9 | 0.2 | ||
| QL5 | QL | 4.4 | 0.1 | ||
| QL6 | QL | 8.9 | 0.1 | ||
| DKW1 | DKW | 4.4 | 0.5 | ||
| DKW2 | DKW | 8.9 | 0.5 | ||
| DKW3 | DKW | 4.4 | 0.2 | ||
| DKW4 | DKW | 8.9 | 0.2 | ||
| DKW5 | DKW | 4.4 | 0.1 | ||
| DKW6 | DKW | 8.9 | 0.1 | ||
| WPMm1 | WPMm | 4.4 | 0.5 | ||
| WPMm2 | WPMm | 8.9 | 0.5 | ||
| WPMm3 | WPMm | 4.4 | 0.2 | ||
| WPMm4 | WPMm | 8.9 | 0.2 | ||
| WPMm5 | WPMm | 4.4 | 0.1 | ||
| WPMm6 | WPMm | 8.9 | 0.1 | ||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sabbadini, S.; Ricci, A.; Limera, C.; Baldoni, D.; Capriotti, L.; Mezzetti, B. Factors Affecting the Regeneration, via Organogenesis, and the Selection of Transgenic Calli in the Peach Rootstock Hansen 536 (Prunus persica × Prunus amygdalus) to Express an RNAi Construct against PPV Virus. Plants 2019, 8, 178. https://doi.org/10.3390/plants8060178
Sabbadini S, Ricci A, Limera C, Baldoni D, Capriotti L, Mezzetti B. Factors Affecting the Regeneration, via Organogenesis, and the Selection of Transgenic Calli in the Peach Rootstock Hansen 536 (Prunus persica × Prunus amygdalus) to Express an RNAi Construct against PPV Virus. Plants. 2019; 8(6):178. https://doi.org/10.3390/plants8060178
Chicago/Turabian StyleSabbadini, Silvia, Angela Ricci, Cecilia Limera, Dania Baldoni, Luca Capriotti, and Bruno Mezzetti. 2019. "Factors Affecting the Regeneration, via Organogenesis, and the Selection of Transgenic Calli in the Peach Rootstock Hansen 536 (Prunus persica × Prunus amygdalus) to Express an RNAi Construct against PPV Virus" Plants 8, no. 6: 178. https://doi.org/10.3390/plants8060178
APA StyleSabbadini, S., Ricci, A., Limera, C., Baldoni, D., Capriotti, L., & Mezzetti, B. (2019). Factors Affecting the Regeneration, via Organogenesis, and the Selection of Transgenic Calli in the Peach Rootstock Hansen 536 (Prunus persica × Prunus amygdalus) to Express an RNAi Construct against PPV Virus. Plants, 8(6), 178. https://doi.org/10.3390/plants8060178






