Polyamines as Quality Control Metabolites Operating at the Post-Transcriptional Level
Abstract
1. Introduction
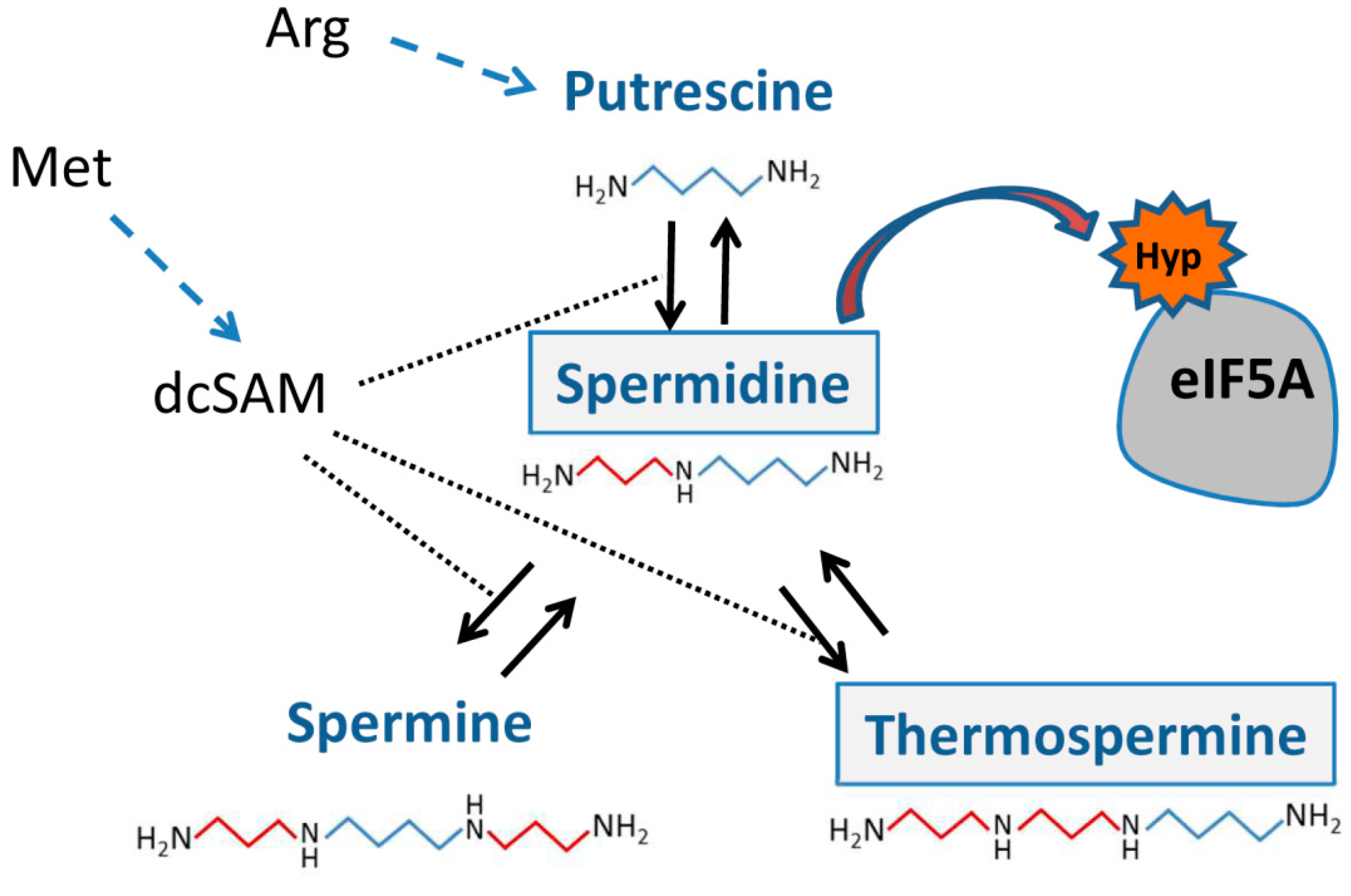
2. Polyamines and Interactions with the Translation Machinery
3. Polyamines and mRNA Quality Control Mechanisms
3.1. The NMD Pathway
3.2. The NGD Pathway
3.3. The NSD Pathway
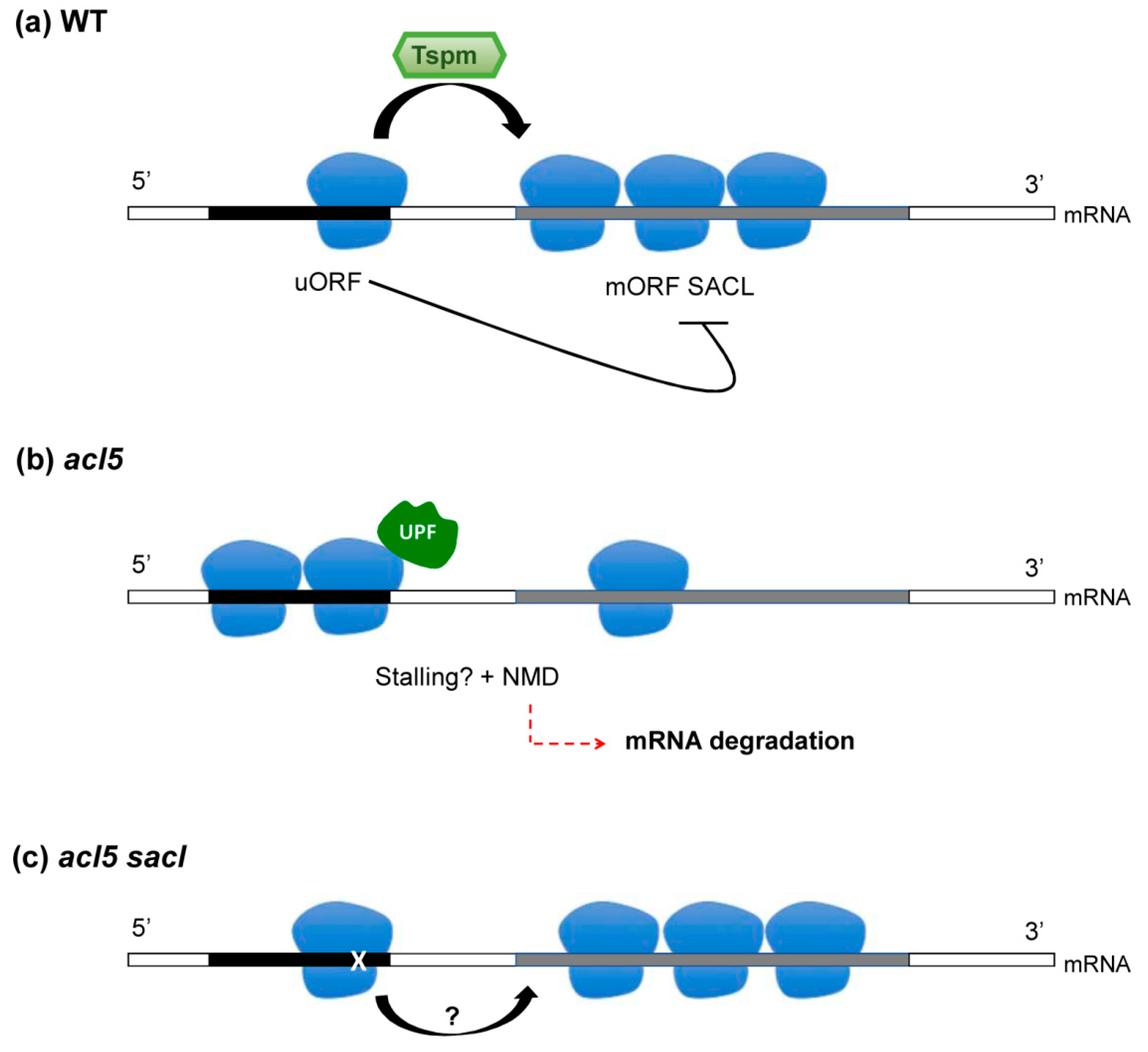
4. Thermospermine and uORF-Dependent Translational Inhibition: Is There A Connection to the NMD Pathway?
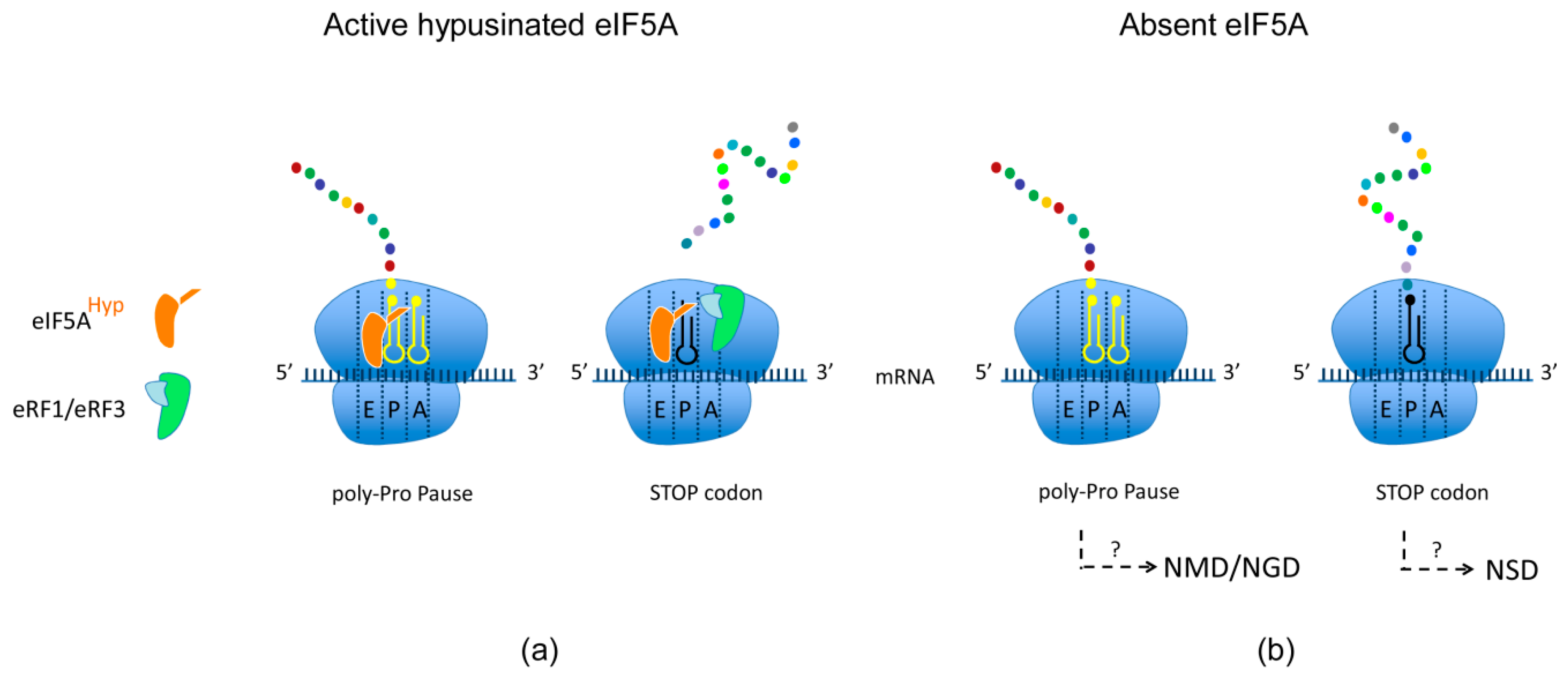
5. Spermidine and Hypusination of eIF5A: How to Avoid Reversible Ribosome Pausing and Links to the NMD Pathway
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Graille, M.; Séraphin, B. Surveillance pathways rescuing eukaryotic ribosomes lost in translation. Nat. Rev. Mol. Cell Biol. 2012, 13, 727. [Google Scholar] [CrossRef]
- Preissler, S.; Deuerling, E. Ribosome-associated chaperones as key players in proteostasis. Trends Biochem. Sci. 2012, 37, 274–283. [Google Scholar] [CrossRef] [PubMed]
- Fuell, C.; Elliott, K.A.; Hanfrey, C.C.; Franceschetti, M.; Michael, A.J. Polyamine biosynthetic diversity in plants and algae. Plant Physiol. Biochem. 2010, 48, 513–520. [Google Scholar] [CrossRef]
- Vera-Sirera, F.; Minguet, E.G.; Singh, S.K.; Ljung, K.; Tuominen, H.; Blázquez, M.A.; Carbonell, J. Role of polyamines in plant vascular development. Plant Physiol. Biochem. 2010, 48, 534–539. [Google Scholar] [CrossRef]
- Watanabe, S.; Kusama-Eguchi, K.; Kobayashi, H.; Igarashi, K. Estimation of polyamine binding to macromolecules and ATP in bovine lymphocytes and rat liver. J. Biol. Chem. 1991, 266, 20803–20809. [Google Scholar]
- Igarashi, K.; Sugawara, K.; Izumi, I.; Nagayama, C.; Hirose, S. Effect of Polyamines on Polyphenylalanine Synthesis by Escherichia coli and Rat-Liver Ribosomes. Eur. J. Biochem. 1974, 48, 495–502. [Google Scholar] [CrossRef] [PubMed]
- Igarashi, K.; Hashimoto, S.; Miyake, A.; KashiwagiI, K.; Hirose, S. Increase of Fidelity of Polypeptide Synthesis by Spermidine in Eukaryotic Cell-Free Systems. Eur. J. Biochem. 1982, 128, 597–604. [Google Scholar] [CrossRef] [PubMed]
- Echandi, G.; Algranati, I.D. Defective 30S ribosomal particles in a polyamine auxotroph of Escherichia coli. Biochem. Biophys. Res. Commun. 1975, 67, 1185–1191. [Google Scholar] [CrossRef]
- Igarashi, K.; Kishida, K.; Hirose, S. Stimulation by polyamines of enzymatic methylation of two adjacent adenines near the 3′ end of 16S ribosomal RNA of Escherichia coli. Biochem. Biophys. Res. Commun. 1980, 96, 678–684. [Google Scholar] [CrossRef]
- Hetrick, B.; Khade, P.K.; Lee, K.; Stephen, J.; Thomas, A.; Joseph, S. Polyamines Accelerate Codon Recognition by Transfer RNAs on the Ribosome. Biochemistry 2010, 49, 7179–7189. [Google Scholar] [CrossRef][Green Version]
- Amarantos, I.; Kalpaxis, D.L. Photoaffinity polyamines: interactions with AcPhe-tRNA free in solution or bound at the P-site of Escherichia coli ribosomes. Nucleic Acids Res. 2000, 28, 3733–3742. [Google Scholar] [CrossRef][Green Version]
- Kalpaxis, D.L.; Amarantos, I.; Zarkadis, I.K. The identification of spermine binding sites in 16S rRNA allows interpretation of the spermine effect on ribosomal 30S subunit functions. Nucleic Acids Res. 2002, 30, 2832–2843. [Google Scholar] [CrossRef]
- Xaplanteri, M.A.; Petropoulos, A.D.; Dinos, G.P.; Kalpaxis, D.L. Localization of spermine binding sites in 23S rRNA by photoaffinity labeling: parsing the spermine contribution to ribosomal 50S subunit functions. Nucleic Acids Res. 2005, 33, 2792–2805. [Google Scholar] [CrossRef]
- Dever, T.E.; Ivanov, I.P. Roles of polyamines in translation. J. Biol. Chem. 2018. [Google Scholar] [CrossRef]
- Ivanov, I.P.; Matsufuji, S.; Murakami, Y.; Gesteland, R.F.; Atkins, J.F. Conservation of polyamine regulation by translational frameshifting from yeast to mammals. Embo J. 2000, 19, 1907–1917. [Google Scholar] [CrossRef]
- Brandman, O.; Hegde, R.S. Ribosome-associated protein quality control. Nat. Struct. Mol. Biol. 2016, 23, 7. [Google Scholar] [CrossRef] [PubMed]
- Behm-Ansmant, I.; Kashima, I.; Rehwinkel, J.; Saulière, J.; Wittkopp, N.; Izaurralde, E. mRNA quality control: An ancient machinery recognizes and degrades mRNAs with nonsense codons. Feb. Lett. 2007, 581, 2845–2853. [Google Scholar] [CrossRef] [PubMed]
- Chang, Y.-F.; Imam, J.S.; Wilkinson, M.F. The Nonsense-Mediated Decay RNA Surveillance Pathway. Annu. Rev. Biochem. 2007, 76, 51–74. [Google Scholar] [CrossRef]
- Brogna, S.; Wen, J. Nonsense-mediated mRNA decay (NMD) mechanisms. Nat. Struct. Mol. Biol. 2009, 16, 107–113. [Google Scholar] [CrossRef]
- Amrani, N.; Sachs, M.S.; Jacobson, A. Early nonsense: mRNA decay solves a translational problem. Nat. Rev. Mol. Cell Biol. 2006, 7, 415. [Google Scholar] [CrossRef]
- Rebbapragada, I.; Lykke-Andersen, J. Execution of nonsense-mediated mRNA decay: what defines a substrate? Curr. Opin. Cell Biol. 2009, 21, 394–402. [Google Scholar] [CrossRef] [PubMed]
- Peccarelli, M.; Kebaara, B.W. Regulation of Natural mRNAs by the Nonsense-Mediated mRNA Decay Pathway. Eukaryot Cell 2014, 13, 1126–1135. [Google Scholar] [CrossRef] [PubMed]
- Kurihara, Y.; Matsui, A.; Hanada, K.; Kawashima, M.; Ishida, J.; Morosawa, T.; Tanaka, M.; Kaminuma, E.; Mochizuki, Y.; Matsushima, A.; et al. Genome-wide suppression of aberrant mRNA-like noncoding RNAs by NMD in Arabidopsis. Proc. Natl. Acad. Sci. USA 2009, 106, 2453–2458. [Google Scholar] [CrossRef] [PubMed]
- Drechsel, G.; Kahles, A.; Kesarwani, A.K.; Stauffer, E.; Behr, J.; Drewe, P.; Rätsch, G.; Wachter, A. Nonsense-Mediated Decay of Alternative Precursor mRNA Splicing Variants Is a Major Determinant of the Arabidopsis Steady State Transcriptome. Plant Cell 2013, 25, 3726–3742. [Google Scholar] [CrossRef]
- Kalyna, M.; Simpson, C.G.; Syed, N.H.; Lewandowska, D.; Marquez, Y.; Kusenda, B.; Marshall, J.; Fuller, J.; Cardle, L.; McNicol, J.; et al. Alternative splicing and nonsense-mediated decay modulate expression of important regulatory genes in Arabidopsis. Nucleic Acids Res. 2012, 40, 2454–2469. [Google Scholar] [CrossRef]
- Leeds, P.; Wood, J.M.; Lee, B.S.; Culbertson, M.R. Gene products that promote mRNA turnover in Saccharomyces cerevisiae. Mol. Cell Biol. 1992, 12, 2165–2177. [Google Scholar] [CrossRef]
- Kerényi, Z.; Mérai, Z.; Hiripi, L.; Benkovics, A.; Gyula, P.; Lacomme, C.; Barta, E.; Nagy, F.; Silhavy, D. Inter-kingdom conservation of mechanism of nonsense-mediated mRNA decay. Embo J. 2008, 27, 1585–1595. [Google Scholar] [CrossRef]
- Shaul, O. Unique Aspects of Plant Nonsense-Mediated mRNA Decay. Trends Plant Sci. 2015, 20, 767–779. [Google Scholar] [CrossRef]
- Rayson, S.; Arciga-Reyes, L.; Wootton, L.; De Torres Zabala, M.; Truman, W.; Graham, N.; Grant, M.; Davies, B. A role for nonsense-mediated mRNA decay in plants: Pathogen responses are induced in Arabidopsis thaliana NMD mutants. PLoS ONE 2012, 7, e31917. [Google Scholar] [CrossRef]
- Shi, C.; Baldwin, I.T.; Wu, J. Arabidopsis Plants Having Defects in Nonsense-mediated mRNA Decay Factors UPF1, UPF2, and UPF3 Show Photoperiod-dependent Phenotypes in Development and Stress Responses. J. Integr. Plant Biol. 2012, 54, 99–114. [Google Scholar] [CrossRef]
- Nasim, Z.; Fahim, M.; Ahn, J.H. Possible Role of MADS AFFECTING FLOWERING 3 and B-BOX DOMAIN PROTEIN 19 in Flowering Time Regulation of Arabidopsis Mutants with Defects in Nonsense-Mediated mRNA Decay. Front. Plant Sci. 2017, 8. [Google Scholar] [CrossRef]
- Degtiar, E.; Fridman, A.; Gottlieb, D.; Berezin, I.; Vexler, K.; Golani, L.; Farhi, R.; Shaul, O. The feedback control of UPF3 is crucial for RNA surveillance in plants. Nucleic Acids Res. 2015, 43, 4219–4235. [Google Scholar] [CrossRef] [PubMed]
- Popp, M.W.-L.; Maquat, L.E. Organizing Principles of Mammalian Nonsense-Mediated mRNA Decay. Annu. Rev. Genet. 2013, 47, 139–165. [Google Scholar] [CrossRef]
- Dai, Y.; Li, W.; An, L. NMD mechanism and the functions of Upf proteins in plant. Plant Cell Rep. 2016, 35, 5–15. [Google Scholar] [CrossRef] [PubMed]
- Karousis, E.D.; Mühlemann, O. Nonsense-Mediated mRNA Decay Begins Where Translation Ends. Cold Spring Harb. Perspect. Biol. 2019, 11. [Google Scholar] [CrossRef] [PubMed]
- Doma, M.K.; Parker, R. Endonucleolytic cleavage of eukaryotic mRNAs with stalls in translation elongation. Nature 2006, 440, 561. [Google Scholar] [CrossRef] [PubMed]
- Atkinson, G.C.; Baldauf, S.L.; Hauryliuk, V. Evolution of nonstop, no-go and nonsense-mediated mRNA decay and their termination factor-derived components. BMC Evol. Biol. 2008, 8, 290. [Google Scholar] [CrossRef] [PubMed]
- Szádeczky-Kardoss, I.; Gál, L.; Auber, A.; Taller, J.; Silhavy, D. The No-go decay system degrades plant mRNAs that contain a long A-stretch in the coding region. Plant Sci. 2018, 275, 19–27. [Google Scholar] [CrossRef] [PubMed]
- Shoemaker, C.J.; Eyler, D.E.; Green, R. Dom34:Hbs1 Promotes Subunit Dissociation and Peptidyl-tRNA Drop-Off to Initiate No-Go Decay. Science 2010, 330, 369–372. [Google Scholar] [CrossRef]
- Tsuboi, T.; Kuroha, K.; Kudo, K.; Makino, S.; Inoue, E.; Kashima, I.; Inada, T. Dom34:Hbs1 Plays a General Role in Quality-Control Systems by Dissociation of a Stalled Ribosome at the 3′ End of Aberrant mRNA. Mol. Cell 2012, 46, 518–529. [Google Scholar] [CrossRef]
- Beelman, C.A.; Parker, R. Differential effects of translational inhibition in cis and in trans on the decay of the unstable yeast MFA2 mRNA. J. Biol. Chem. 1994, 269, 9687–9692. [Google Scholar]
- Buchan, J.R.; Stansfield, I. Halting a cellular production line: responses to ribosomal pausing during translation. Biol. Cell 2007, 99, 475–487. [Google Scholar] [CrossRef]
- Simms, C.L.; Yan, L.L.; Zaher, H.S. Ribosome Collision Is Critical for Quality Control during No-Go Decay. Mol. Cell 2017, 68, 361–373. [Google Scholar] [CrossRef]
- Ozsolak, F.; Kapranov, P.; Foissac, S.; Kim, S.W.; Fishilevich, E.; Monaghan, A.P.; John, B.; Milos, P.M. Comprehensive Polyadenylation Site Maps in Yeast and Human Reveal Pervasive Alternative Polyadenylation. Cell 2010, 143, 1018–1029. [Google Scholar] [CrossRef] [PubMed]
- Dimitrova, L.N.; Kuroha, K.; Tatematsu, T.; Inada, T. Nascent Peptide-dependent Translation Arrest Leads to Not4p-mediated Protein Degradation by the Proteasome. J. Biol. Chem. 2009, 284, 10343–10352. [Google Scholar] [CrossRef]
- Koutmou, K.S.; Schuller, A.P.; Brunelle, J.L.; Radhakrishnan, A.; Djuranovic, S.; Green, R. Ribosomes slide on lysine-encoding homopolymeric A stretches. eLife 2015, 4, e05534. [Google Scholar] [CrossRef]
- van Hoof, A.; Frischmeyer, P.A.; Dietz, H.C.; Parker, R. Exosome-Mediated Recognition and Degradation of mRNAs Lacking a Termination Codon. Science 2002, 295, 2262–2264. [Google Scholar] [CrossRef] [PubMed]
- Frischmeyer, P.A.; van Hoof, A.; O’Donnell, K.; Guerrerio, A.L.; Parker, R.; Dietz, H.C. An mRNA Surveillance Mechanism That Eliminates Transcripts Lacking Termination Codons. Science 2002, 295, 2258–2261. [Google Scholar] [CrossRef] [PubMed]
- Auber, A.; Szádeczky-Kardoss, I.; Burgyán, J.; Csorba, T.; Nyikó, T.; Silhavy, D.; Schamberger, A.; Orbán, T.I.; Taller, J. The nonstop decay and the RNA silencing systems operate cooperatively in plants. Nucleic Acids Res. 2018, 46, 4632–4648. [Google Scholar] [CrossRef]
- Oshima, T. A new polyamine, thermospermine, 1,12-diamino-4,8-diazadodecane, from an extreme thermophile. J. Biol. Chem. 1979, 254, 8720–8722. [Google Scholar] [PubMed]
- Hanzawa, Y.; Takahashi, T.; Komeda, Y. ACL5: an Arabidopsis gene required for internodal elongation after flowering. Plant J. 1997, 12, 863–874. [Google Scholar] [CrossRef]
- Hanzawa, Y.; Takahashi, T.; Michael, A.J.; Burtin, D.; Long, D.; Pineiro, M.; Coupland, G.; Komeda, Y. ACAULIS5, an Arabidopsis gene required for stem elongation, encodes a spermine synthase. Embo J. 2000, 19, 4248–4256. [Google Scholar] [CrossRef]
- Knott, J.M.; Romer, P.; Sumper, M. Putative spermine synthases from Thalassiosira pseudonana and Arabidopsis thaliana synthesize thermospermine rather than spermine. Febs. Lett. 2007, 581, 3081–3086. [Google Scholar] [CrossRef] [PubMed]
- Minguet, E.G.; Vera-Sirera, F.; Marina, A.; Carbonell, J.; Blazquez, M.A. Evolutionary Diversification in Polyamine Biosynthesis. Mol. Biol. Evol. 2008, 25, 2119–2128. [Google Scholar] [CrossRef] [PubMed]
- Milhinhos, A.; Prestele, J.; Bollhöner, B.; Matos, A.; Vera-Sirera, F.; Rambla, J.L.; Ljung, K.; Carbonell, J.; Blázquez, M.A.; Tuominen, H.; et al. Thermospermine levels are controlled by an auxin-dependent feedback loop mechanism in Populus xylem. Plant J. 2013, 75, 685–698. [Google Scholar] [CrossRef]
- Baima, S.; Forte, V.; Possenti, M.; Peñalosa, A.; Leoni, G.; Salvi, S.; Felici, B.; Ruberti, I.; Morelli, G. Negative Feedback Regulation of Auxin Signaling by ATHB8/ACL5–BUD2 Transcription Module. Mol. Plant 2014, 7, 1006–1025. [Google Scholar] [CrossRef] [PubMed]
- Kakehi, J.i.; Kuwashiro, Y.; Niitsu, M.; Takahashi, T. Thermospermine is Required for Stem Elongation in Arabidopsis thaliana. Plant Cell Physiol. 2008, 49, 1342–1349. [Google Scholar] [CrossRef] [PubMed]
- Clay, N.K.; Nelson, T. Arabidopsis thickvein Mutation Affects Vein Thickness and Organ Vascularization, and Resides in a Provascular Cell-Specific Spermine Synthase Involved in Vein Definition and in Polar Auxin Transport. Plant Physiol. 2005, 138, 767–777. [Google Scholar] [CrossRef]
- Muñiz, L.; Minguet, E.G.; Singh, S.K.; Pesquet, E.; Vera-Sirera, F.; Moreau-Courtois, C.L.; Carbonell, J.; Blázquez, M.A.; Tuominen, H. ACAULIS5 controls Arabidopsis xylem specification through the prevention of premature cell death. Development 2008, 135, 2573–2582. [Google Scholar] [CrossRef]
- Imai, A.; Hanzawa, Y.; Komura, M.; Yamamoto, K.T.; Komeda, Y.; Takahashi, T. The dwarf phenotype of the Arabidopsis acl5 mutant is suppressed by a mutation in an upstream ORF of a bHLH gene. Development 2006, 133, 3575–3585. [Google Scholar] [CrossRef]
- Imai, A.; Komura, M.; Kawano, E.; Kuwashiro, Y.; Takahashi, T. A semi-dominant mutation in the ribosomal protein L10 gene suppresses the dwarf phenotype of the acl5 mutant in Arabidopsis thaliana. Plant J. 2008, 56, 881–890. [Google Scholar] [CrossRef] [PubMed]
- Kakehi, J.-I.; Kawano, E.; Yoshimoto, K.; Cai, Q.; Imai, A.; Takahashi, T. Mutations in Ribosomal Proteins, RPL4 and RACK1, Suppress the Phenotype of a Thermospermine-Deficient Mutant of Arabidopsis thaliana. PLoS ONE 2015, 10, e0117309. [Google Scholar] [CrossRef] [PubMed]
- Cai, Q.; Fukushima, H.; Yamamoto, M.; Ishii, N.; Sakamoto, T.; Kurata, T.; Motose, H.; Takahashi, T. The SAC51 Family Plays a Central Role in Thermospermine Responses in Arabidopsis. Plant Cell Physiol. 2016, 57, 1583–1592. [Google Scholar] [CrossRef] [PubMed]
- Vera-Sirera, F.; De Rybel, B.; Úrbez, C.; Kouklas, E.; Pesquera, M.; Álvarez-Mahecha, J.C.; Minguet, E.G.; Tuominen, H.; Carbonell, J.; Borst, J.W.; et al. A bHLH-Based Feedback Loop Restricts Vascular Cell Proliferation in Plants. Dev. Cell 2015, 35, 432–443. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, M.; Takahashi, T. Thermospermine enhances translation of SAC51 and SACL1 in Arabidopsis. Plant Signal. Behav. 2017, 12, e1276685. [Google Scholar] [CrossRef]
- von Arnim, A.G.; Jia, Q.; Vaughn, J.N. Regulation of plant translation by upstream open reading frames. Plant Sci. 2014, 214, 1–12. [Google Scholar] [CrossRef]
- Weiss, M.C.; Sousa, F.L.; Mrnjavac, N.; Neukirchen, S.; Roettger, M.; Nelson-Sathi, S.; Martin, W.F. The physiology and habitat of the last universal common ancestor. Nat. Microbiol. 2016, 1, 16116. [Google Scholar] [CrossRef]
- Imai, A.; Matsuyama, T.; Hanzawa, Y.; Akiyama, T.; Tamaoki, M.; Saji, H.; Shirano, Y.; Kato, T.; Hayashi, H.; Shibata, D.; et al. Spermidine Synthase Genes Are Essential for Survival of Arabidopsis. Plant Physiol. 2004, 135, 1565–1573. [Google Scholar] [CrossRef]
- Hamasaki-Katagiri, N.; Tabor, C.W.; Tabor, H. Spermidine biosynthesis in Saccharomyces cerevisiae: Polyamine requirement of a null mutant of the SPE3 gene (spermidine synthase). Gene 1997, 187, 35–43. [Google Scholar] [CrossRef]
- Mandal, S.; Mandal, A.; Johansson, H.E.; Orjalo, A.V.; Park, M.H. Depletion of cellular polyamines, spermidine and spermine, causes a total arrest in translation and growth in mammalian cells. Proc. Natl. Acad. Sci. USA 2013, 110, 2169–2174. [Google Scholar] [CrossRef]
- Park, M.H.; Wolff, E.C. Hypusine, a polyamine-derived amino acid critical for eukaryotic translation. J. Biol. Chem. 2018. [Google Scholar] [CrossRef]
- Park, M. The post-translational synthesis of a polyamine-derived amino acid, hypusine, in the eukaryotic translation initiation factor 5A (eIF5A). J. Biochem. 2006, 139, 161–169. [Google Scholar] [CrossRef] [PubMed]
- Park, M.H.; Wolff, E.C.; Folk, J.E. Hypusine - Its Posttranslational Formation in Eukaryotic Initiation Factor-5A and Its Potential Role in Cellular-Regulation. Biofactors 1993, 4, 95–104. [Google Scholar] [PubMed]
- Chattopadhyay, M.K.; Park, M.H.; Tabor, H. Hypusine modification for growth is the major function of spermidine in Saccharomyces cerevisiae polyamine auxotrophs grown in limiting spermidine. Proc. Natl. Acad. Sci. USA 2008, 105, 6554–6559. [Google Scholar] [CrossRef] [PubMed]
- Pällmann, N.; Braig, M.; Sievert, H.; Preukschas, M.; Hermans-Borgmeyer, I.; Schweizer, M.; Nagel, C.H.; Neumann, M.; Wild, P.; Haralambieva, E.; et al. Biological Relevance and Therapeutic Potential of the Hypusine Modification System. J. Biol. Chem. 2015, 290, 18343–18360. [Google Scholar] [CrossRef] [PubMed]
- Nishimura, K.; Lee, S.; Park, J.; Park, M. Essential role of eIF5A-1 and deoxyhypusine synthase in mouse embryonic development. Amino Acids 2012, 42, 703–710. [Google Scholar] [CrossRef] [PubMed]
- Pagnussat, G.C.; Yu, H.J.; Ngo, Q.A.; Rajani, S.; Mayalagu, S.; Johnson, C.S.; Capron, A.; Xie, L.F.; Ye, D.; Sundaresan, V. Genetic and molecular identification of genes required for female gametophyte development and function in Arabidopsis. Development 2005, 132, 603–614. [Google Scholar] [CrossRef]
- Thomas, A.; Goumans, H.; Amesz, H.; Benne, R.; Voorma, H.O. A Comparison of the Initiation Factors of Eukaryotic Protein Synthesis from Ribosomes and from the Postribosomal Supernatant. Eur. J. Biochem. 1979, 98, 329–337. [Google Scholar] [CrossRef]
- Cooper, H.L.; Park, M.H.; Folk, J.E.; Safer, B.; Braverman, R. Identification of the hypusine-containing protein hy+ as translation initiation factor eIF-4D. Proc. Natl. Acad. Sci. USA 1983, 80, 1854–1857. [Google Scholar] [CrossRef]
- Kemper, W.M.; Berry, K.W.; Merrick, W.C. Purification and properties of rabbit reticulocyte protein synthesis initiation factors M2Balpha and M2Bbeta. J. Biol. Chem. 1976, 251, 5551–5557. [Google Scholar]
- Shiba, T.; Mizote, H.; Kaneko, T.; Nakajima, T.; Yasuo, K.; Sano, I. Hypusine, a new amino acid occurring in bovine brain: Isolation and structural determination. Biochim. Biophys. Acta (BBA) Gen. Subj. 1971, 244, 523–531. [Google Scholar] [CrossRef]
- Park, M.H.; Cooper, H.L.; Folk, J.E. Identification of hypusine, an unusual amino acid, in a protein from human lymphocytes and of spermidine as its biosynthetic precursor. Proc. Natl. Acad. Sci. USA 1981, 78, 2869–2873. [Google Scholar] [CrossRef]
- Saini, P.; Eyler, D.E.; Green, R.; Dever, T.E. Hypusine-containing protein eIF5A promotes translation elongation. Nature 2009, 459, 118–121. [Google Scholar] [CrossRef]
- Schuller, A.P.; Wu, C.C.-C.; Dever, T.E.; Buskirk, A.R.; Green, R. eIF5A Functions Globally in Translation Elongation and Termination. Mol. Cell 2017, 66, 194–205.e195. [Google Scholar] [CrossRef] [PubMed]
- Gäbel, K.; Schmitt, J.; Schulz, S.; Näther, D.J.; Soppa, J. A Comprehensive Analysis of the Importance of Translation Initiation Factors for Haloferax volcanii Applying Deletion and Conditional Depletion Mutants. PLoS ONE 2013, 8, e77188. [Google Scholar] [CrossRef] [PubMed]
- Kyrpides, N.C.; Woese, C.R. Universally conserved translation initiation factors. Proc. Natl. Acad. Sci. USA 1998, 95, 224–228. [Google Scholar] [CrossRef]
- Navarre, W.W.; Zou, S.B.; Roy, H.; Xie, J.L.; Savchenko, A.; Singer, A.; Edvokimova, E.; Prost, L.R.; Kumar, R.; Ibba, M.; et al. PoxA, YjeK, and Elongation Factor P Coordinately Modulate Virulence and Drug Resistance in Salmonella enterica. Mol. Cell 2010, 39, 209–221. [Google Scholar] [CrossRef]
- Lassak, J.; Keilhauer, E.C.; Fürst, M.; Wuichet, K.; Gödeke, J.; Starosta, A.L.; Chen, J.-M.; Søgaard-Andersen, L.; Rohr, J.; Wilson, D.N.; et al. Arginine-rhamnosylation as new strategy to activate translation elongation factor P. Nat. Chem. Biol. 2015, 11, 266–270. [Google Scholar] [CrossRef]
- Bullwinkle, T.J.; Zou, S.B.; Rajkovic, A.; Hersch, S.J.; Elgamal, S.; Robinson, N.; Smil, D.; Bolshan, Y.; Navarre, W.W.; Ibba, M. (R)-β-Lysine-modified Elongation Factor P Functions in Translation Elongation. J. Biol. Chem. 2013, 288, 4416–4423. [Google Scholar] [CrossRef]
- Balibar, C.J.; Iwanowicz, D.; Dean, C.R. Elongation Factor P is Dispensable in Escherichia coli and Pseudomonas aeruginosa. Curr. Microbiol. 2013, 67, 293–299. [Google Scholar] [CrossRef]
- Blaha, G.; Stanley, R.E.; Steitz, T.A. Formation of the First Peptide Bond: The Structure of EF-P Bound to the 70S Ribosome. Science 2009, 325, 966–970. [Google Scholar] [CrossRef]
- Melnikov, S.; Mailliot, J.; Shin, B.-S.; Rigger, L.; Yusupova, G.; Micura, R.; Dever, T.E.; Yusupov, M. Crystal Structure of Hypusine-Containing Translation Factor eIF5A Bound to a Rotated Eukaryotic Ribosome. J. Mol. Biol. 2016. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, C.; Becker, T.; Heuer, A.; Braunger, K.; Shanmuganathan, V.; Pech, M.; Berninghausen, O.; Wilson, D.N.; Beckmann, R. Structure of the hypusinylated eukaryotic translation factor eIF-5A bound to the ribosome. Nucleic Acids Res. 2016, 44, 1944–1951. [Google Scholar] [CrossRef] [PubMed]
- Gutierrez, E.; Shin, B.-S.; Woolstenhulme, C.J.; Kim, J.-R.; Saini, P.; Buskirk, A.R.; Dever, T.E. eIF5A Promotes Translation of Polyproline Motifs. Mol. Cell 2013, 51, 35–45. [Google Scholar] [CrossRef]
- Doerfel, L.K.; Wohlgemuth, I.; Kothe, C.; Peske, F.; Urlaub, H.; Rodnina, M.V. EF-P Is Essential for Rapid Synthesis of Proteins Containing Consecutive Proline Residues. Science 2013, 339, 85–88. [Google Scholar] [CrossRef] [PubMed]
- Ude, S.; Lassak, J.; Starosta, A.L.; Kraxenberger, T.; Wilson, D.N.; Jung, K. Translation Elongation Factor EF-P Alleviates Ribosome Stalling at Polyproline Stretches. Science 2013, 339, 82–85. [Google Scholar] [CrossRef] [PubMed]
- Pavlov, M.Y.; Watts, R.E.; Tan, Z.; Cornish, V.W.; Ehrenberg, M.; Forster, A.C. Slow peptide bond formation by proline and other N-alkylamino acids in translation. Proc. Natl. Acad. Sci. USA 2009, 106, 50–54. [Google Scholar] [CrossRef] [PubMed]
- Belda Palazón, B.; Almendáriz, C.; Martí, E.; Carbonell, J.; Ferrando, A. Relevance of the axis spermidine/eIF5A for plant growth and development. Front. Plant Sci. 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Belda-Palazon, B.; Ferrando, A.; Alepuz, P. Fertility and Polarized Cell Growth Depends on eIF5A for Translation of Polyproline-Rich Formins in Saccharomyces cerevisiae. Genetics 2014, 197, 1191–1200. [Google Scholar] [CrossRef]
- Duguay, J.; Jamal, S.; Liu, Z.; Wang, T.W.; Thompson, J.E. Leaf-specific suppression of deoxyhypusine synthase in Arabidopsis thaliana enhances growth without negative pleiotropic effects. J. Plant Physiol. 2007, 164, 408–420. [Google Scholar] [CrossRef]
- Feng, H.; Chen, Q.; Feng, J.; Zhang, J.; Yang, X.; Zuo, J. Functional Characterization of the Arabidopsis Eukaryotic Translation Initiation Factor 5A-2 That Plays a Crucial Role in Plant Growth and Development by Regulating Cell Division, Cell Growth, and Cell Death. Plant Physiol. 2007, 144, 1531–1545. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Duguay, J.; Ma, F.; Wang, T.W.; Tshin, R.; Hopkins, M.T.; McNamara, L.; Thompson, J.E. Modulation of eIF5A1 expression alters xylem abundance in Arabidopsis thaliana. J. Exp. Bot. 2008, 59, 939–950. [Google Scholar] [CrossRef] [PubMed]
- Ma, F.; Liu, Z.; Wang, T.W.; Hopkins, M.T.; Peterson, C.A.; Thompson, J.E. Arabidopsis eIF5A3 influences growth and the response to osmotic and nutrient stress. Plant Cell Env. 2010, 33, 1682–1696. [Google Scholar] [CrossRef] [PubMed]
- Buskirk, A.R.; Green, R. Ribosome pausing, arrest and rescue in bacteria and eukaryotes. Philos. Trans. R. Soc. B: Biol. Sci. 2017, 372. [Google Scholar] [CrossRef]
- Dever, T.E.; Dinman, J.D.; Green, R. Translation Elongation and Recoding in Eukaryotes. Cold Spring Harb. Perspect. Biol. 2018, 10. [Google Scholar] [CrossRef] [PubMed]
- Zuk, D.; Jacobson, A. A single amino acid substitution in yeast eIF-5A results in mRNA stabilization. Embo J. 1998, 17, 2914–2925. [Google Scholar] [CrossRef]
- Schrader, R.; Young, C.; Kozian, D.; Hoffmann, R.; Lottspeich, F. Temperature-sensitive eIF5A Mutant Accumulates Transcripts Targeted to the Nonsense-mediated Decay Pathway. J. Biol. Chem. 2006, 281, 35336–35346. [Google Scholar] [CrossRef]
- Hoque, M.; Park, J.Y.; Chang, Y.-J.; Luchessi, A.D.; Cambiaghi, T.D.; Shamanna, R.; Hanauske-Abel, H.M.; Holland, B.; Pe’ery, T.; Tian, B.; et al. Regulation of gene expression by translation factor eIF5A: Hypusine-modified eIF5A enhances nonsense-mediated mRNA decay in human cells. Translation 2017, 5, e1366294. [Google Scholar] [CrossRef]
- Li, C.H.; Ohn, T.; Ivanov, P.; Tisdale, S.; Anderson, P. eIF5A Promotes Translation Elongation, Polysome Disassembly and Stress Granule Assembly. PLoS ONE 2010, 5, e9942. [Google Scholar] [CrossRef]
- Miller-Fleming, L.; Olin-Sandoval, V.; Campbell, K.; Ralser, M. Remaining Mysteries of Molecular Biology: The Role of Polyamines in the Cell. J. Mol. Biol. 2015, 427, 3389–3406. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Poidevin, L.; Unal, D.; Belda-Palazón, B.; Ferrando, A. Polyamines as Quality Control Metabolites Operating at the Post-Transcriptional Level. Plants 2019, 8, 109. https://doi.org/10.3390/plants8040109
Poidevin L, Unal D, Belda-Palazón B, Ferrando A. Polyamines as Quality Control Metabolites Operating at the Post-Transcriptional Level. Plants. 2019; 8(4):109. https://doi.org/10.3390/plants8040109
Chicago/Turabian StylePoidevin, Laetitia, Dilek Unal, Borja Belda-Palazón, and Alejandro Ferrando. 2019. "Polyamines as Quality Control Metabolites Operating at the Post-Transcriptional Level" Plants 8, no. 4: 109. https://doi.org/10.3390/plants8040109
APA StylePoidevin, L., Unal, D., Belda-Palazón, B., & Ferrando, A. (2019). Polyamines as Quality Control Metabolites Operating at the Post-Transcriptional Level. Plants, 8(4), 109. https://doi.org/10.3390/plants8040109




