Long-Term Tillage and Crop Rotation Regimes Reshape Soil-Borne Oomycete Communities in Soybean, Corn, and Wheat Production Systems
Abstract
1. Introduction
2. Results
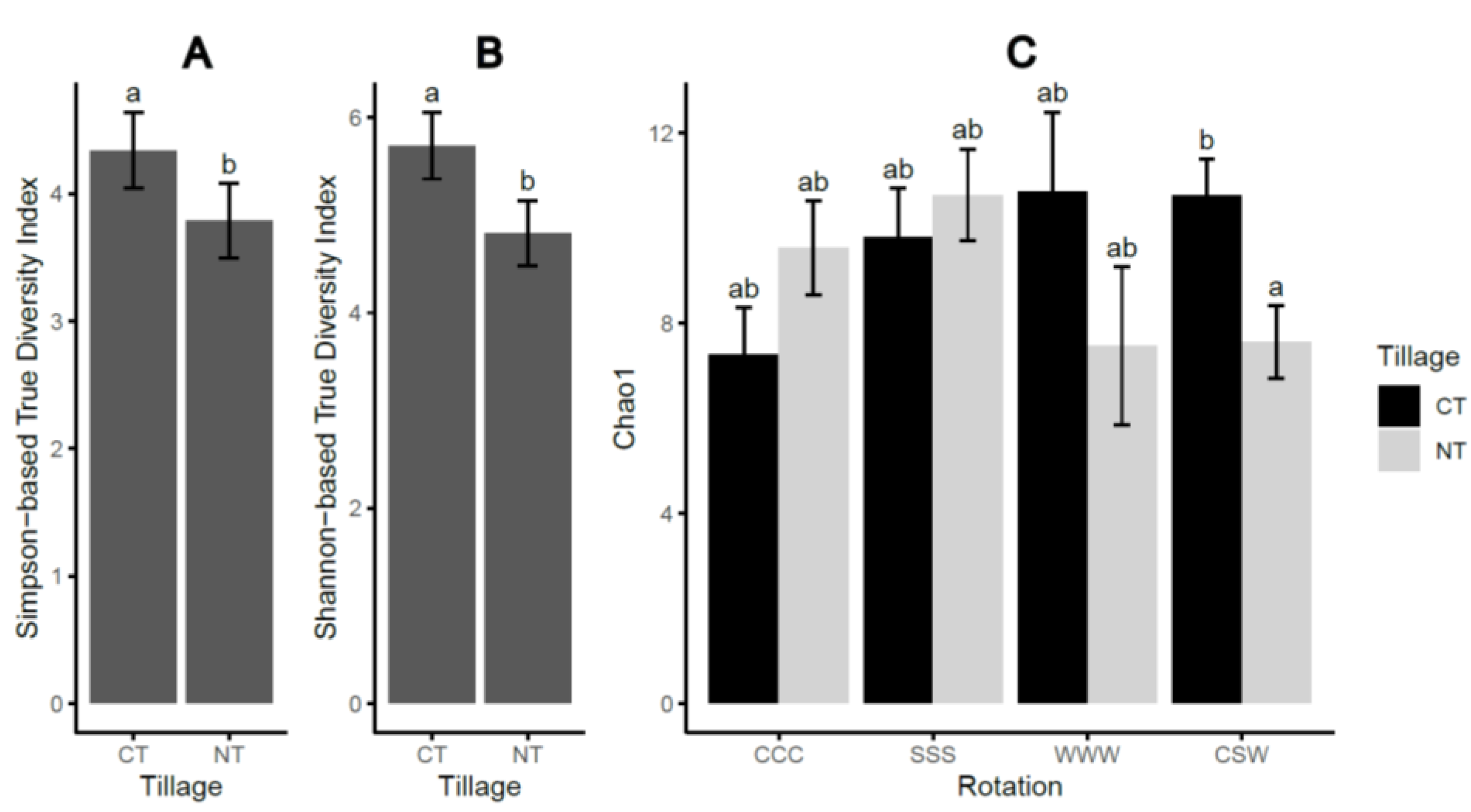
2.1. Soil-Borne Oomycete Community Diversity and Compositional Structure
2.2. Crop Yield and Soybean Seedling Emergence
3. Discussion
4. Materials and Methods
4.1. Study Site and Experimental Design
4.2. Soil Sampling
4.3. DNA Extraction
4.4. Sequencing Library Preparation and Illumina MiSeq Sequencing
4.5. Metabarcoding Data Processing and Analysis
4.6. Soybean Seedling Vitality Experiment
4.7. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Thines, M.; Voglmayr, H.; Göker, M. Taxonomy and Phylogeny of the Downy Mildews (Peronosporaceae). In Oomycete Genetics and Genomics; Wiley-VCH: Weinheim, Germany, 2009; pp. 47–75. [Google Scholar]
- Fry, W. Phytophthora infestans: The plant (and R gene) destroyer. Mol. Plant Pathol. 2008, 9, 385–402. [Google Scholar] [CrossRef] [PubMed]
- Phillips, A.J.; Anderson, V.L.; Robertson, E.J.; Secombes, C.J.; van West, P. New insights into animal pathogenic oomycetes. Trends Microbiol. 2008, 16, 13–19. [Google Scholar] [CrossRef] [PubMed]
- Cahill, D.M.; Rookes, J.E.; Wilson, B.A.; Gibson, L.; McDougall, K.L. Phytophthora cinnamomi and Australia’s biodiversity: Impacts, predictions and progress towards control. Aust. J. Bot. 2008, 56, 279–310. [Google Scholar] [CrossRef]
- Cohen, Y.; Coffey, M.D. Systemic fungicides and the control of oomycetes. Annu. Rev. Phytopathol. 1986, 24, 311–338. [Google Scholar] [CrossRef]
- Dorrance, A.; Grünwald, N.J. Phytophthora sojae: Diversity among and within Populations. In Oomycete Genetics and Genomics; Wiley-VCH: Weinheim, Germany, 2009; pp. 197–212. [Google Scholar]
- Cook, R.; Sitton, J.; Haglund, W. Influence of soil treatments on growth and yield of wheat and implications for control of Pythium root rot. Phytopathology 1987, 77, 1192–1198. [Google Scholar] [CrossRef]
- Cook, R.J.; Haglund, W.A. Wheat yield depression associated with conservation tillage caused by root pathogens in the soil not phytotoxins from the straw. Soil Biol. Biochem. 1991, 23, 1125–1132. [Google Scholar] [CrossRef]
- Panno, S.; Davino, S.; Caruso, A.G.; Bertacca, S.; Crnogorac, A.; Mandić, A.; Noris, E.; Matić, S. A Review of the Most Common and Economically Important Diseases That Undermine the Cultivation of Tomato Crop in the Mediterranean Basin. Agronomy 2021, 11, 2188. [Google Scholar] [CrossRef]
- Niu, F.; Xing, Y.; Jia, N.; Ding, K.; Xie, D.; Chen, H.; Chi, D. Effectiveness of entomopathogenic fungal strains against poplar/willow weevil (Cryptorhynchus lapathi L.) larvae. J. For. Res. 2022, 33, 1691–1702. [Google Scholar] [CrossRef]
- Bongiorno, G.; Postma, J.; Bünemann, E.K.; Brussaard, L.; de Goede, R.G.M.; Mäder, P.; Tamm, L.; Thuerig, B. Soil suppressiveness to Pythium ultimum in ten European long-term field experiments and its relation with soil parameters. Soil Biol. Biochem. 2019, 133, 174–187. [Google Scholar] [CrossRef]
- Peters, R.D.; Sturz, A.V.; Carter, M.R.; Sanderson, J.B. Developing disease-suppressive soils through crop rotation and tillage management practices. Soil Tillage Res. 2003, 72, 181–192. [Google Scholar] [CrossRef]
- Yuan, X.; Wang, B.; Hong, S.; Xiong, W.; Shen, Z.; Ruan, Y.; Li, R.; Shen, Q.; Dini-Andreote, F. Promoting soil microbial-mediated suppressiveness against Fusarium wilt disease by the enrichment of specific fungal taxa via crop rotation. Biol. Fertil. Soils 2021, 57, 1137–1153. [Google Scholar] [CrossRef]
- Santín-Montanyá, M.I.; Sombrero Sacristán, A. The effects of soil tillage techniques on weed flora in high input barley systems in northern Spain. Can. J. Plant Sci. 2020, 100, 245–252. [Google Scholar] [CrossRef]
- Bogunovic, I.; Pereira, P.; Kisic, I.; Sajko, K.; Sraka, M. Tillage management impacts on soil compaction, erosion and crop yield in Stagnosols (Croatia). Catena 2018, 160, 376–384. [Google Scholar] [CrossRef]
- Abdollahi, L.; Schjønning, P.; Elmholt, S.; Munkholm, L.J. The effects of organic matter application and intensive tillage and traffic on soil structure formation and stability. Soil Tillage Res. 2014, 136, 28–37. [Google Scholar] [CrossRef]
- Srour, A.Y.; Ammar, H.A.; Subedi, A.; Pimentel, M.; Cook, R.L.; Bond, J.; Fakhoury, A.M. Microbial Communities Associated With Long-Term Tillage and Fertility Treatments in a Corn-Soybean Cropping System. Front Microbiol 2020, 11, 1363. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Song, D.; Liang, S.; Dang, P.; Qin, X.; Liao, Y.; Siddique, K.H.M. Effect of no-tillage on soil bacterial and fungal community diversity: A meta-analysis. Soil Tillage Res. 2020, 204, 104721. [Google Scholar] [CrossRef]
- Sipilä Timo, P.; Yrjälä, K.; Alakukku, L.; Palojärvi, A. Cross-Site Soil Microbial Communities under Tillage Regimes: Fungistasis and Microbial Biomarkers. Appl. Environ. Microbiol. 2012, 78, 8191–8201. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Gumpertz, M.L.; Hu, S.; Ristaino, J.B. Effect of prior tillage and soil fertility amendments on dispersal of Phytophthora capsici and infection of pepper. Eur. J. Plant Pathol. 2008, 120, 273–287. [Google Scholar] [CrossRef]
- Peralta, A.L.; Sun, Y.; McDaniel, M.D.; Lennon, J.T. Crop rotational diversity increases disease suppressive capacity of soil microbiomes. Ecosphere 2018, 9, e02235. [Google Scholar] [CrossRef]
- Pedersen, E.A. The Effect of Crop Rotation on Development of the Septoria Disease Complex on Spring Wheat in Saskatchewan. Can. J. Plant Pathol. 1992, 14, 152–158. [Google Scholar] [CrossRef]
- Larkin, R.P.; Halloran, J.M. Management Effects of Disease-Suppressive Rotation Crops on Potato Yield and Soilborne Disease and Their Economic Implications in Potato Production. Am. J. Potato Res. 2014, 91, 429–439. [Google Scholar] [CrossRef]
- Lyu, J.; Jin, L.; Jin, N.; Xie, J.; Xiao, X.; Hu, L.; Tang, Z.; Wu, Y.; Niu, L.; Yu, J. Effects of Different Vegetable Rotations on Fungal Community Structure in Continuous Tomato Cropping Matrix in Greenhouse. Front. Microbiol. 2020, 11, 829. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Li, L.; Sui, P.; Liu, X.; Zhang, T.; Li, X. Crop rotation to diversify the soil microbiome in the semi-arid area of Inner Mongolia, China. Arch. Agron. Soil Sci. 2022, 69, 1161–1176. [Google Scholar] [CrossRef]
- Blakney, A.J.C.; Bainard, L.D.; St-Arnaud, M.; Hijri, M. Soil chemistry and soil history significantly structure oomycete communities in Brassicaceae crop rotations. Appl. Environ. Microbiol. 2022, 89, e01314–e01322. [Google Scholar] [CrossRef]
- Hwang, S.; Ahmed, H.; Gossen, B.; Kutcher, H.; Brandt, S.; Strelkov, S.; Chang, K.; Turnbull, G. Effect of crop rotation on the soil pathogen population dynamics and canola seedling establishment. Plant Pathol. J. 2009, 8, 106–112. [Google Scholar] [CrossRef]
- Radmer, L.; Anderson, G.; Malvick, D.M.; Kurle, J.E.; Rendahl, A.; Mallik, A. Pythium, Phytophthora, and Phytopythium spp. Associated with Soybean in Minnesota, Their Relative Aggressiveness on Soybean and Corn, and Their Sensitivity to Seed Treatment Fungicides. Plant Dis. 2017, 101, 62–72. [Google Scholar] [CrossRef]
- Zitnick-Anderson, K.K.; Nelson, B.D. Identification and Pathogenicity of Pythium on Soybean in North Dakota. Plant Dis. 2015, 99, 31–38. [Google Scholar] [CrossRef]
- Rojas, J.A.; Jacobs, J.L.; Napieralski, S.; Karaj, B.; Bradley, C.A.; Chase, T.; Esker, P.D.; Giesler, L.J.; Jardine, D.J.; Malvick, D.K.; et al. Oomycete Species Associated with Soybean Seedlings in North America—Part I: Identification and Pathogenicity Characterization. Phytopathology 2017, 107, 280–292. [Google Scholar] [CrossRef]
- Abreo, E.; Vaz-Jauri, P.; Nuñez, L.; Stewart, S.; Mattos, N.; Dini, B.; Altier, N. Pathogenicity of Pythium spp. obtained from agricultural soils and symptomatic legume seedlings in Uruguay. Australas. Plant Dis. Notes 2017, 12, 35. [Google Scholar] [CrossRef]
- Broders, K.D.; Lipps, P.E.; Paul, P.A.; Dorrance, A.E. Characterization of Pythium spp. Associated with Corn and Soybean Seed and Seedling Disease in Ohio. Plant Dis. 2007, 91, 727–735. [Google Scholar] [CrossRef]
- Coffua, L.S.; Veterano, S.T.; Clipman, S.J.; Mena-Ali, J.I.; Blair, J.E. Characterization of Pythium spp. Associated with Asymptomatic Soybean in Southeastern Pennsylvania. Plant Dis. 2016, 100, 1870–1879. [Google Scholar] [CrossRef]
- Gijzen, M.; Qutob, D. Phytophthora sojae and Soybean. In Oomycete Genetics and Genomics; Wiley-VCH: Weinheim, Germany, 2009; pp. 303–329. [Google Scholar]
- Klepadlo, M.; Balk, C.S.; Vuong, T.D.; Dorrance, A.E.; Nguyen, H.T. Molecular characterization of genomic regions for resistance to Pythium ultimum var. ultimum in the soybean cultivar Magellan. Theor. Appl. Genet. 2019, 132, 405–417. [Google Scholar] [CrossRef]
- Molin, C.; Ribeiro, N.R.; Matsumoto, M.N.; Fernando Giasson, N.; Brollo, J.; Zanardo, B.; Pelissoni, M.; Capitanio, S.; Comín, T.; Deuner, C.C.; et al. Damping-off of soybean in southern Brazil can be associated with different species of Globisporangium spp. and Pythium spp. Plant Pathology 2021, 70, 1686–1694. [Google Scholar] [CrossRef]
- Noel, Z.A.; Chang, H.-X.; Chilvers, M.I. Variation in soybean rhizosphere oomycete communities from Michigan fields with contrasting disease pressures. Appl. Soil Ecol. 2020, 150, 103435. [Google Scholar] [CrossRef]
- Wei, L.; Xue, A.; Cober, E.; Babcock, C.; Zhang, J.; Zhang, S.; Li, W.; Wu, J.; Liu, L. Pathogenicity of Pythium species causing seed rot and damping-off in soybean under controlled conditions. Phytoprotection 2010, 91, 3–10. [Google Scholar] [CrossRef]
- Rojas, J.A.; Jacobs, J.L.; Napieralski, S.; Karaj, B.; Bradley, C.A.; Chase, T.; Esker, P.D.; Giesler, L.J.; Jardine, D.J.; Malvick, D.K.; et al. Oomycete Species Associated with Soybean Seedlings in North America—Part II: Diversity and Ecology in Relation to Environmental and Edaphic Factors. Phytopathology 2017, 107, 293–304. [Google Scholar] [CrossRef]
- Morrison, M.J.; Cober, E.R.; Gregorich, E.; Voldeng, H.D.; Ma, B.; Topp, G.C. Tillage and crop rotation effects on the yield of corn, soybean and wheat in eastern Canada. Can. J. Plant Sci. 2017, 98, 183–191. [Google Scholar] [CrossRef]
- Jalli, M.; Huusela, E.; Jalli, H.; Kauppi, K.; Niemi, M.; Himanen, S.; Jauhiainen, L. Effects of Crop Rotation on Spring Wheat Yield and Pest Occurrence in Different Tillage Systems: A Multi-Year Experiment in Finnish Growing Conditions. Front. Sustain. Food Syst. 2021, 5, 647335. [Google Scholar] [CrossRef]
- Sapp, M.; Tyborski, N.; Linstädter, A.; López Sánchez, A.; Mansfeldt, T.; Waldhoff, G.; Bareth, G.; Bonkowski, M.; Rose, L.E. Site-specific distribution of oak rhizosphere-associated oomycetes revealed by cytochrome c oxidase subunit II metabarcoding. Ecol. Evol. 2019, 9, 10567–10581. [Google Scholar] [CrossRef] [PubMed]
- Rossmann, S.; Lysøe, E.; Skogen, M.; Talgø, V.; Brurberg, M.B. DNA Metabarcoding Reveals Broad Presence of Plant Pathogenic Oomycetes in Soil From Internationally Traded Plants. Front. Microbiol. 2021, 12, 637068. [Google Scholar] [CrossRef]
- Martin, F.N.; Loper, J.E. Soilborne Plant Diseases Caused by Pythium spp.: Ecology, Epidemiology, and Prospects for Biological Control. Crit. Rev. Plant Sci. 1999, 18, 111–181. [Google Scholar] [CrossRef]
- Uzuhashi, S.; Kakishima, M.; Tojo, M. Phylogeny of the genus Pythium and description of new genera. Mycoscience 2010, 51, 337–365. [Google Scholar] [CrossRef]
- Nguyen, H.D.T.; Dodge, A.; Dadej, K.; Rintoul, T.L.; Ponomareva, E.; Martin, F.N.; de Cock, A.W.A.M.; Lévesque, C.A.; Redhead, S.A.; Spies, C.F.J. Whole genome sequencing and phylogenomic analysis show support for the splitting of genus Pythium. Mycologia 2022, 114, 501–515. [Google Scholar] [CrossRef]
- Gan, H.; Chai, Z.; Lou, B.; Li, J. Pythium heterothallicum new to China and its pathogenicity. Mycosystem 2010, 29, 494–501. [Google Scholar]
- Bakker, M.G.; Moorman, T.B.; Kaspar, T.C.; Manter, D.K. Isolation of Cultivation-Resistant Oomycetes, First Detected as Amplicon Sequences, from Roots of Herbicide-Terminated Winter Rye. Phytobiomes 2017, 1, 24–35. [Google Scholar] [CrossRef]
- Reyes-Tena, A.; Vallejo-González, R.; Santillán-Mendoza, R.; Rodríguez-Alvarado, G.; Larsen, J.; Fernández-Pavía, S.P. Pythium arrhenomanes causal agent of root rot on yellow maize in Mexico. Australas. Plant Dis. Notes 2018, 13, 6. [Google Scholar] [CrossRef]
- Chamswarng, C.; Cook, R. Identification and comparative pathogenicity of Pythium spp. from wheat roots and wheat-field soils in the Pacific Northwest. Phytopathology 1985, 75, 821–827. [Google Scholar] [CrossRef]
- Paulitz, T.C. Low Input No-till Cereal Production in the Pacific Northwest of the U.S.: The Challenges of Root Diseases. Eur. J. Plant Pathol. 2006, 115, 271–281. [Google Scholar] [CrossRef]
- Waller, J.M. Observations on Pythium root rot of wheat and barley. Plant Pathol. 1979, 28, 17–24. [Google Scholar] [CrossRef]
- Yin, C.; McLaughlin, K.; Paulitz, T.C.; Kroese, D.R.; Hagerty, C.H. Population Dynamics of Wheat Root Pathogens Under Different Tillage Systems in Northeast Oregon. Plant Dis. 2020, 104, 2649–2657. [Google Scholar] [CrossRef] [PubMed]
- Hoitink, H.; Boehm, M. Biocontrol within the context of soil microbial communities: A substrate-dependent phenomenon. Annu. Rev. Phytopathol. 1999, 37, 427. [Google Scholar] [CrossRef] [PubMed]
- Berendsen, R.L.; Pieterse, C.M.J.; Bakker, P.A.H.M. The rhizosphere microbiome and plant health. Trends Plant Sci. 2012, 17, 478–486. [Google Scholar] [CrossRef] [PubMed]
- Shittu, K.A.; Oyedele, D.J.; Babatunde, K.M. The effects of moisture content at tillage on soil strength in maize production. Egypt. J. Basic Appl. Sci. 2017, 4, 139–142. [Google Scholar] [CrossRef]
- Yang, T.; Siddique, K.H.M.; Liu, K. Cropping systems in agriculture and their impact on soil health-A review. Glob. Ecol. Conserv. 2020, 23, e01118. [Google Scholar] [CrossRef]
- Cook, R.; Sitton, J.; Waldher, J. Evidence for Pythium as a pathogen of direct-drilled wheat in the Pacific Northwest. Plant Dis. 1980, 64, 61. [Google Scholar] [CrossRef]
- Pankhurst, C.E.; McDonald, H.J.; Hawke, B.G. Influence of tillage and crop rotation on the epidemiology of Pythium infections of wheat in a red-brown earth of South Australia. Soil Biol. Biochem. 1995, 27, 1065–1073. [Google Scholar] [CrossRef]
- Paul, B. A new species of Pythium isolated from a vineyard in France. FEMS Microbiol. Lett. 2006, 263, 194–199. [Google Scholar] [CrossRef]
- Jiang, Y.N.; Haudenshield, J.S.; Hartman, G.L. Characterization of Pythium spp. from soil samples in Illinois. Can. J. Plant Pathol. 2012, 34, 448–454. [Google Scholar] [CrossRef]
- Broders, K.D.; Wallhead, M.W.; Austin, G.D.; Lipps, P.E.; Paul, P.A.; Mullen, R.W.; Dorrance, A.E. Association of Soil Chemical and Physical Properties with Pythium Species Diversity, Community Composition, and Disease Incidence. Phytopathology 2009, 99, 957–967. [Google Scholar] [CrossRef]
- Martin, F.; Hancock, J. Association of chemical and biological factors in soils suppressive to Pythium ultimum. Phytopathology 1986, 76, 1221–1231. [Google Scholar] [CrossRef]
- Rojas, J.A.; Witte, A.; Noel, Z.A.; Jacobs, J.L.; Chilvers, M.I. Diversity and Characterization of Oomycetes Associated with Corn Seedlings in Michigan. Phytobiomes 2019, 3, 224–234. [Google Scholar] [CrossRef]
- Bockus, W.; Shroyer, J. The impact of reduced tillage on soilborne plant pathogens. Annu. Rev. Phytopathol. 1998, 36, 485–500. [Google Scholar] [CrossRef] [PubMed]
- Asad, N.I.; Wang, X.-B.; Dozois, J.; Azarbad, H.; Constant, P.; Yergeau, E. Early season soil microbiome best predicts wheat grain quality. FEMS Microbiol. Ecol. 2023, 99, fiac144. [Google Scholar] [CrossRef] [PubMed]
- Sapkota, R.; Nicolaisen, M. Cropping history shapes fungal, oomycete and nematode communities in arable soils and affects cavity spot in carrot. Agric. Ecosyst. Environ. 2018, 257, 120–131. [Google Scholar] [CrossRef]
- Emmond, G.S.; Ledingham, R.J. Effects of crop rotation on some soil-borne pathogens of potato. Can. J. Plant Sci. 1972, 52, 605–611. [Google Scholar] [CrossRef]
- Frank, J.A.; Murphy, H. The effect of crop rotations on Rhizoctonia disease of potatoes. Am. Potato J. 1977, 54, 315–322. [Google Scholar] [CrossRef]
- Scholte, K.H. The effect of crop rotation and granular nematicides on the incidence ofRhizoctonia solani in potato. J. Potato Res. 1987, 30, 187–199. [Google Scholar] [CrossRef]
- Bargués-Ribera, M.; Gokhale, C.S. Eco-evolutionary agriculture: Host-pathogen dynamics in crop rotations. PLoS Comput. Biol. 2020, 16, e1007546. [Google Scholar] [CrossRef]
- Lipps, P.; Bruehl, G. Snow rot of winter wheat in Washington. Phytopathology 1978, 68, 120–121. [Google Scholar] [CrossRef]
- Spies, C.F.J.; Mazzola, M.; McLeod, A. Characterisation and detection of Pythium and Phytophthora species associated with grapevines in South Africa. Eur. J. Plant Pathol. 2011, 131, 103–119. [Google Scholar] [CrossRef]
- Masigol, H.; Khodaparast, S.A.; Mostowfizadeh-Ghalamfarsa, R.; Rojas-Jimenez, K.; Woodhouse, J.N.; Neubauer, D.; Grossart, H.-P. Taxonomical and functional diversity of Saprolegniales in Anzali lagoon, Iran. Aquat. Ecol. 2020, 54, 323–336. [Google Scholar] [CrossRef]
- Rizvi, S.; Yang, X. Fungi associated with soybean seedling disease in Iowa. Plant Dis. 1996, 80, 57–60. [Google Scholar] [CrossRef]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef] [PubMed]
- Bengtsson-Palme, J.; Ryberg, M.; Hartmann, M.; Branco, S.; Wang, Z.; Godhe, A.; De Wit, P.; Sánchez-García, M.; Ebersberger, I.; de Sousa, F.; et al. Improved software detection and extraction of ITS1 and ITS2 from ribosomal ITS sequences of fungi and other eukaryotes for analysis of environmental sequencing data. Methods Ecol. Evol. 2013, 4, 914–919. [Google Scholar] [CrossRef]
- Bokulich, N.A.; Kaehler, B.D.; Rideout, J.R.; Dillon, M.; Bolyen, E.; Knight, R.; Huttley, G.A.; Caporaso, J.G. Optimizing taxonomic classification of marker-gene amplicon sequences with QIIME 2’s q2-feature-classifier plugin. Microbiome 2018, 6, 90. [Google Scholar] [CrossRef]
- Zahariev, M.; Chen, W.; Visagie, C.M.; Lévesque, C.A. Cluster oligonucleotide signatures for rapid identification by sequencing. BMC Bioinform. 2018, 19, 395. [Google Scholar] [CrossRef]
- Chen, W.; Radford, D.; Hambleton, S. Towards Improved Detection and Identification of Rust Fungal Pathogens in Environmental Samples Using a Metabarcoding Approach. Phytopathology 2021, 112, 535–548. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef]
- Price, M.N.; Dehal, P.S.; Arkin, A.P. FastTree 2—Approximately maximum-likelihood trees for large alignments. PLoS ONE 2010, 5, e9490. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2022. [Google Scholar]
- Oksanen, J.; Simpson, G.; Blanchet, F.; Kindt, R.; Legendre, P.; Minchin, P.; O’Hara, R.; Solymos, P.; Stevens, M.; Szoecs, E.; et al. Vegan: Community Ecology Package. R Package Version 2.6-4. 2022. Available online: https://CRAN.R-project.org/package=vegan (accessed on 22 October 2022).
- Kindt, R. Tree Diversity Analysis. A Manual and Software for Common Statistical Methods for Ecological and Biodiversity Studies; World Agroforestry Centre (ICRAF): Nairobi, Kenya, 2020; ISBN 92-9059-179-X. [Google Scholar]
- Jost, L. Entropy and diversity. Oikos 2006, 113, 363–375. [Google Scholar] [CrossRef]
- Pinheiro, J.; Bates, D. Nlme: Linear and Nonlinear Mixed Effects Models. R Package Version 3.1-161. 2022. Available online: https://CRAN.R-project.org/package=nlme (accessed on 20 October 2022).
- Anderson, M.J. Permutational Multivariate Analysis of Variance (PERMANOVA). In Wiley StatsRef: Statistics Reference Online; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2014; pp. 1–15. [Google Scholar]
- Hervé, M. RVAideMemoire: Testing and Plotting Procedures for Biostatistics. R Package Version 0.9-81-2. 2022. Available online: https://CRAN.R-project.org/package=RVAideMemoire (accessed on 1 October 2022).
- Baselga, A.; Orme, C.D.L. betapart: An R package for the study of beta diversity. Methods Ecol. Evol. 2012, 3, 808–812. [Google Scholar] [CrossRef]




| DF | Simpson-TD 1 | Shannon-TD 2 | Chao1 | |
|---|---|---|---|---|
| Tillage (T) | 1 | 0.014 | 0.007 | 0.160 |
| Rotation (R) | 3 | 0.613 | 0.563 | 0.4292 |
| T × R | 3 | 0.217 | 0.041 | <0.001 |
| Species a | Relative Abundance (%) | Analysis of Variance (p-Value) b | Disease Note c | Known Hosts c,d | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CT | NT | T | R | T*R | R CT | R NT | ||||||||||
| No. ASV | CCC | SSS | WWW | CSW | CCC | SSS | WWW | CSW | ||||||||
| Globisporangium aff. hypogynum | 1 | 0.000 | 0.000 | 0.000 | 0.153 | 0.000 | 0.000 | 0.000 | 0.000 | 0.320 | 0.78 | 0.772 | 0.7817 | N/A | ||
| Globisporangium apiculatum | 20 | 0.664 | 16.294 | 7.970 | 16.522 | 0.000 | 0.000 | 0.000 | 0.002 | <0.001 | 0.036 | 0.031 | 0.041 | 0.761 | Grape | |
| Globisporangium attrantheridium | 49 | 57.290 | 33.493 | 40.079 | 41.743 | 38.209 | 46.590 | 47.258 | 59.105 | 0.230 | 0.448 | 0.021 | 0.066 | 0.138 | Cavity spot lesions | Daucus carota, Prunus, soybean |
| Globisporangium glomeratum | 11 | 2.273 | 2.341 | 1.381 | 2.454 | 1.770 | 2.658 | 3.176 | 2.208 | 0.96 | 0.833 | 0.936 | 0.988 | 0.734 | Soybean | |
| Globisporangium heterothallicum | 27 | 10.438 | 11.844 | 2.942 | 6.979 | 13.878 | 7.603 | 7.139 | 3.802 | 0.539 | 0.039 | 0.558 | N/A | 0.076 | Damping off | Pepper, corn, lentils, soybean, spinach, wheat |
| Globisporangium hypogynum | 3 | 0.000 | 0.930 | 0.000 | 0.000 | 1.115 | 2.277 | 0.000 | 0.728 | 0.055 | 0.075 | 0.902 | 0.061 | 0.381 | Root rots | Soybean |
| Globisporangium intermedium | 3 | 1.000 | 0.000 | 4.290 | 0.388 | 0.000 | 0.995 | 0.281 | 1.700 | 0.755 | 0.074 | 0.001 | 0.004 | 0.407 | Damping off, rots | Abutilon, antirrhinum, arabis, beet, begonia, carrot, cauliflower, corn, chamaecyparis, cherry laurel, chrysanthemum, cotoneaster, cucumber, erica, ferns, Fragaria vesca, godetia, hazel, hop, hyacinth, lettuce, leyland cypress, lupin, nasturtium, pea, pear, pelargonium, pepper, saintpaulia, soybean, strawberry, tomato, violet, yew |
| Globisporangium irregulare | 2 | 0.000 | 0.000 | 0.000 | 0.247 | 0.074 | 0.000 | 0.000 | 0.000 | 0.559 | 0.867 | 0.563 | 0.782 | 0.347 | Blight, damping off, root and other rots, etc. | Soybean, wheat, corn |
| Globisporangium iwayamae | 2 | 0.000 | 0.000 | 1.860 | 0.163 | 0.000 | 0.000 | 0.000 | 0.000 | 0.024 | 0.001 | 0.0007 | 0.002 | N/A | Rots | Poaceae, wheat |
| Globisporangium macrosporum | 2 | 0.000 | 0.000 | 0.000 | 0.181 | 0.055 | 1.220 | 0.797 | 0.546 | 0.029 | 0.658 | 0.738 | 0.782 | 0.682 | Damping off, root rot | Iris, corn, soybean |
| Globisporangium nunn | 1 | 0.000 | 0.000 | 0.000 | 0.000 | 1.042 | 0.000 | 0.000 | 0.000 | 0.333 | 0.349 | 0.339 | N/A | 0.3415 | Soybean | |
| Globisporangium orthogonon | 1 | 0.000 | 1.578 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.305 | 0.323 | 0.264 | 0.3061 | N/A | Corn | |
| Globisporangium pleroticum | 5 | 2.38 | 0.3683 | 0.000 | 0.193 | 0.952 | 1.170 | 0.000 | 1.760 | 0.7545 | 0.074 | 0.009 | 0.711 | 0.841 | Lupins, peas, soybean | |
| Globisporangium recalcitrans | 14 | 1.103 | 6.190 | 0.000 | 1.321 | 3.613 | 4.969 | 3.326 | 1.693 | 0.131 | 0.194 | 0.566 | 0.219 | 0.435 | Root rot, damping off | Beet, hebe |
| Globisporangium rostratifingens | 5 | 0.000 | 0.000 | 0.000 | 0.329 | 0.725 | 0.233 | 0.000 | 0.000 | 0.699 | 0.671 | 0.107 | 0.529 | 0.174 | Root rot | Pea, soybean, corn, wheat |
| Globisporangium selbyi | 8 | 2.200 | 2.223 | 3.130 | 2.350 | 0.591 | 0.774 | 2.410 | 2.290 | 0.702 | 0.859 | 0.996 | 0.943 | 0.750 | Lesions | Corn, soybean |
| Globisporangium sylvaticum | 25 | 4.970 | 2.344 | 9.556 | 3.796 | 19.620 | 12.984 | 3.839 | 7.282 | 0.006 | 0.330 | 0.086 | 0.334 | 0.063 | Root disease, rots | Apples, carrot, cherry laurel, cress, chrysanthemum, cucumber, garlic, lettuce, onion, pea, radish, rhododendron, spinach, strawberry, yew, wheat |
| Globisporangium ultimum | 11 | 2.346 | 14.656 | 1.834 | 6.589 | 2.490 | 2.820 | 0.000 | 1.348 | <0.001 | 0.008 | 0.011 | 0.002 | 0.991 | Blight, damping off, root rot, etc. | garlic, grape, hyacinth, lettuce, lily, lupin, melon, mustard, onion, parsley, pea, pear, pelargonium, pepper, poinsettia, primula, radish, rhododendron, rhubarb, soybean, spinach, strawberry, sweet pea, tomato, tulip, wallflower, yew |
| Phytophthora cf. inundata D0S1P25 | 1 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.237 | 0.000 | 0.000 | 0.180 | 0.093 | 0.152 | N/A | 0.092 | ||
| Pythium acanthicum | 1 | 0.000 | 0.000 | 0.000 | 0.073 | 0.000 | 0.000 | 0.000 | 0.000 | 0.315 | 0.724 | 0.763 | 0.711 | N/A | Downy mildew. Blight, damping off, rots, etc. | Soybean, corn |
| Pythium aff. monospermum | 3 | 1.956 | 0.000 | 10.579 | 0.949 | 2.768 | 0.774 | 14.025 | 2.828 | 0.263 | <0.001 | 0.794 | <0.001 | 0.073 | Grapevine | |
| Pythium arrhenomanes | 8 | 1.116 | 0.000 | 0.000 | 0.872 | 1.257 | 3.545 | 5.713 | 2.795 | 0.014 | 0.782 | 0.223 | 0.492 | 0.409 | Blight, root rot | Corn, rice, barley, wheat |
| Pythium monospermum | 12 | 2.109 | 0.872 | 1.683 | 1.187 | 1.274 | 1.928 | 3.108 | 2.005 | 0.367 | 0.857 | 0.725 | 0.476 | 0.962 | Downy mildew. Root necrosis, not known as a strong pathogen | Cherry, juniper, spinach |
| Pythium oligandrum | 1 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.200 | 0.338 | 0.763 | 0.772 | N/A | 0.762 | Damping off; root, stem, and fruit rots | Soybean, wheat |
| Pythium oopapillum | 2 | 0.000 | 0.158 | 0.000 | 1.536 | 0.000 | 0.000 | 0.000 | 0.404 | 0.35 | 0.48 | 0.931 | 0.678 | 0.761 | Root rot | Soybean |
| Pythium torulosum | 1 | 0.000 | 0.000 | 0.000 | 1.326 | 0.000 | 0.000 | 0.000 | 0.000 | 0.144 | 0.503 | 0.477 | 0.469 | N/A | Damping off, root rot | Pea, soybean, corn |
| Pythium volutum | 6 | 0.000 | 0.000 | 6.287 | 4.399 | 0.000 | 0.002 | 1.895 | 0.053 | 0.020 | 0.004 | 0.143 | 0.034 | 0.014 | Root rot | Barley, melon, morning glory, turfgrass, wheat |
| Saprolegnia anisospora | 1 | 0.000 | 0.000 | 1.997 | 0.000 | 0.000 | 0.238 | 0.000 | 0.000 | 0.704 | 0.024 | 0.01 | 0.006 | 0.379 | ||
| Saprolegnia torulosa | 1 | 0.000 | 0.175 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 1.093 | 0.142 | 0.384 | 0.181 | 0.306 | 0.265 | ||
| Wilsoniana amaranthi | 1 | 0.000 | 0.000 | 0.000 | 0.638 | 1.252 | 0.743 | 0.000 | 0.000 | 0.646 | 0.876 | 0.369 | 0.782 | 0.515 | White blister rust | Amaranth |
| Wilsoniana portulacae | 6 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 5.157 | 0.000 | 1.132 | 0.031 | 0.19 | 0.211 | N/A | N/A | White blister rust | Portulacaceae |
| Corn Yield | Soybean Yield | Wheat Yield | |
|---|---|---|---|
| Tillage | |||
| CT | 9177 b | 2257 | 1854 |
| NT | 10,806 a | 2652 | 1795 |
| Rotation | |||
| Monoculture | 9735 | 2404 | 1298 b |
| Rotation | 10,248 | 2446 | 2305 a |
| Analysis of variance (p-values) | |||
| Tillage (T) | 0.018 | 0.226 | 0.548 |
| Rotation (R) | 0.442 | 0.577 | <0.001 |
| T × R | 0.163 | 0.343 | 0.652 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© His Majesty the King in Right of Canada, as represented by the Minister of Agriculture and Agri-Food Canada 2023. Submitted for possible open access publication under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gahagan, A.C.; Shi, Y.; Radford, D.; Morrison, M.J.; Gregorich, E.; Aris-Brosou, S.; Chen, W. Long-Term Tillage and Crop Rotation Regimes Reshape Soil-Borne Oomycete Communities in Soybean, Corn, and Wheat Production Systems. Plants 2023, 12, 2338. https://doi.org/10.3390/plants12122338
Gahagan AC, Shi Y, Radford D, Morrison MJ, Gregorich E, Aris-Brosou S, Chen W. Long-Term Tillage and Crop Rotation Regimes Reshape Soil-Borne Oomycete Communities in Soybean, Corn, and Wheat Production Systems. Plants. 2023; 12(12):2338. https://doi.org/10.3390/plants12122338
Chicago/Turabian StyleGahagan, Alison Claire, Yichao Shi, Devon Radford, Malcolm J. Morrison, Edward Gregorich, Stéphane Aris-Brosou, and Wen Chen. 2023. "Long-Term Tillage and Crop Rotation Regimes Reshape Soil-Borne Oomycete Communities in Soybean, Corn, and Wheat Production Systems" Plants 12, no. 12: 2338. https://doi.org/10.3390/plants12122338
APA StyleGahagan, A. C., Shi, Y., Radford, D., Morrison, M. J., Gregorich, E., Aris-Brosou, S., & Chen, W. (2023). Long-Term Tillage and Crop Rotation Regimes Reshape Soil-Borne Oomycete Communities in Soybean, Corn, and Wheat Production Systems. Plants, 12(12), 2338. https://doi.org/10.3390/plants12122338






