The PROSCOOP10 Gene Encodes Two Extracellular Hydroxylated Peptides and Impacts Flowering Time in Arabidopsis
Abstract
1. Introduction
2. Results
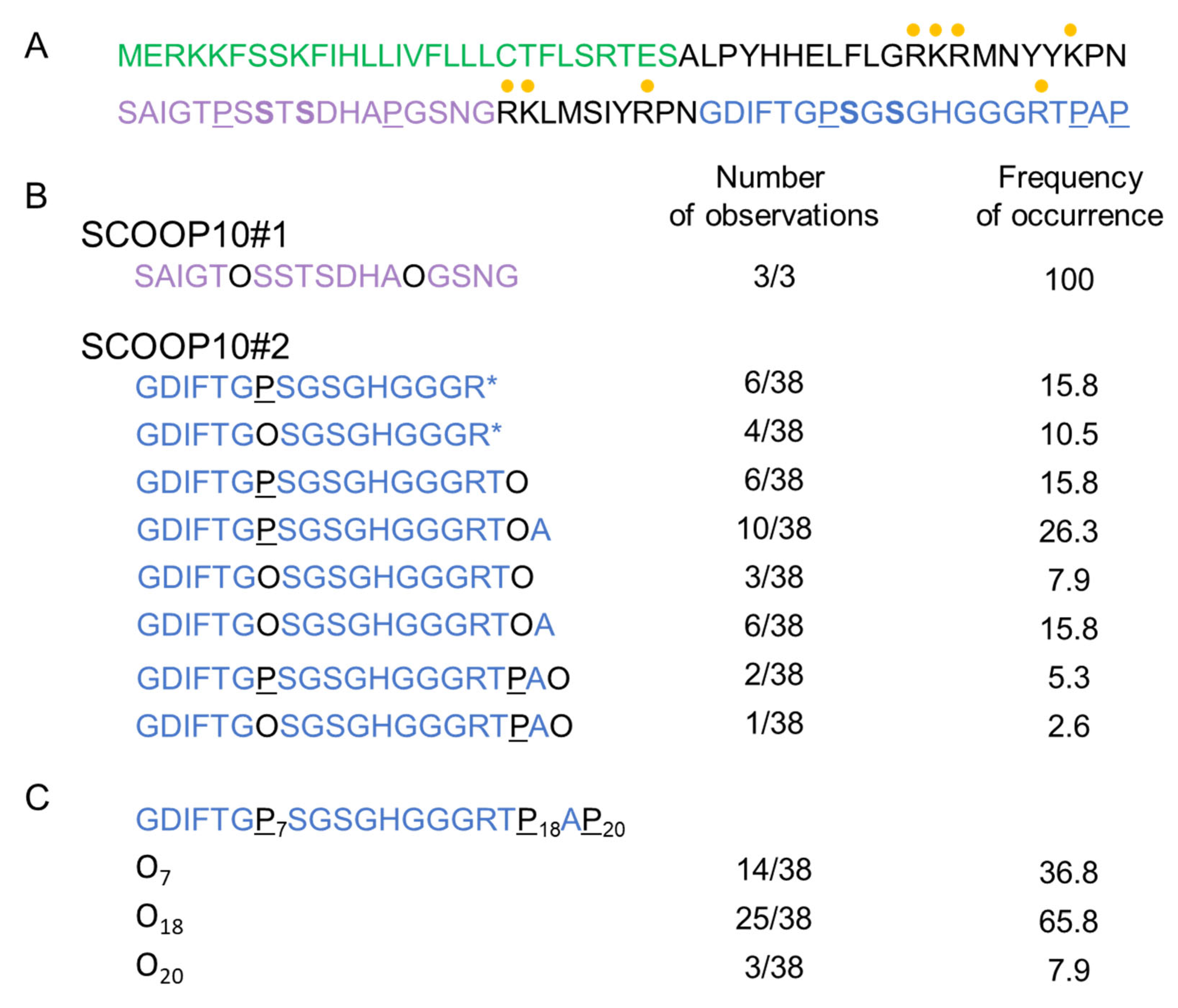
2.1. Identification of SCOOP10 Peptides in Extracellular Fluids by MS
2.2. SCOOP10#2 Is Unstructured in Solution but Cis-Trans Isomerization May Be Favoured in a Hydrophobic Environment
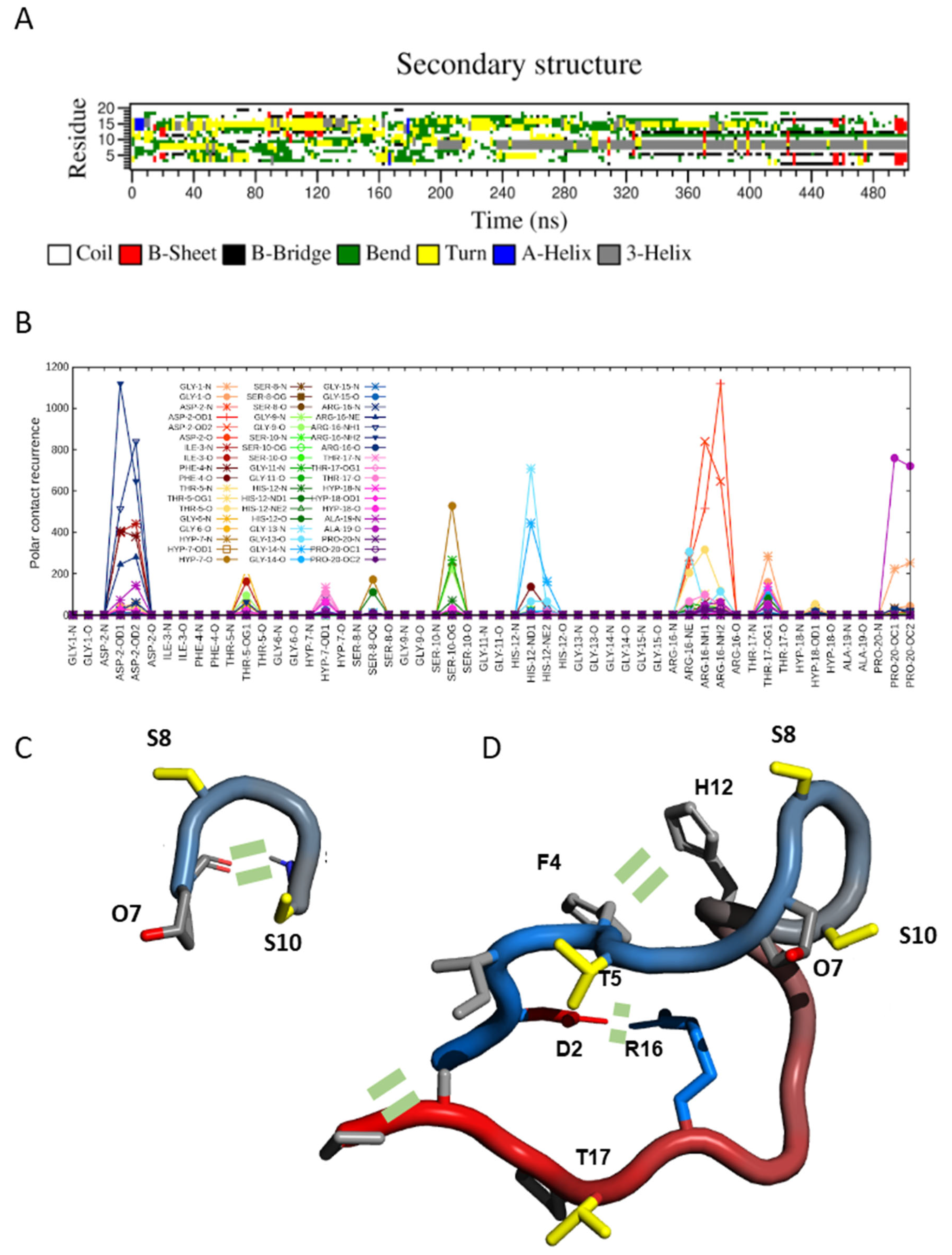
2.3. Transient Hairpin-like Structures Exposing S8 and S10 Might Have Functional Relevance
2.4. SCOOP10#1 Is Unstructured in Solution with Transient Head-to-Tail Contact
2.5. Native SCOOP10 Peptides Do Not Induce Strong ROS Production and Growth Inhibition
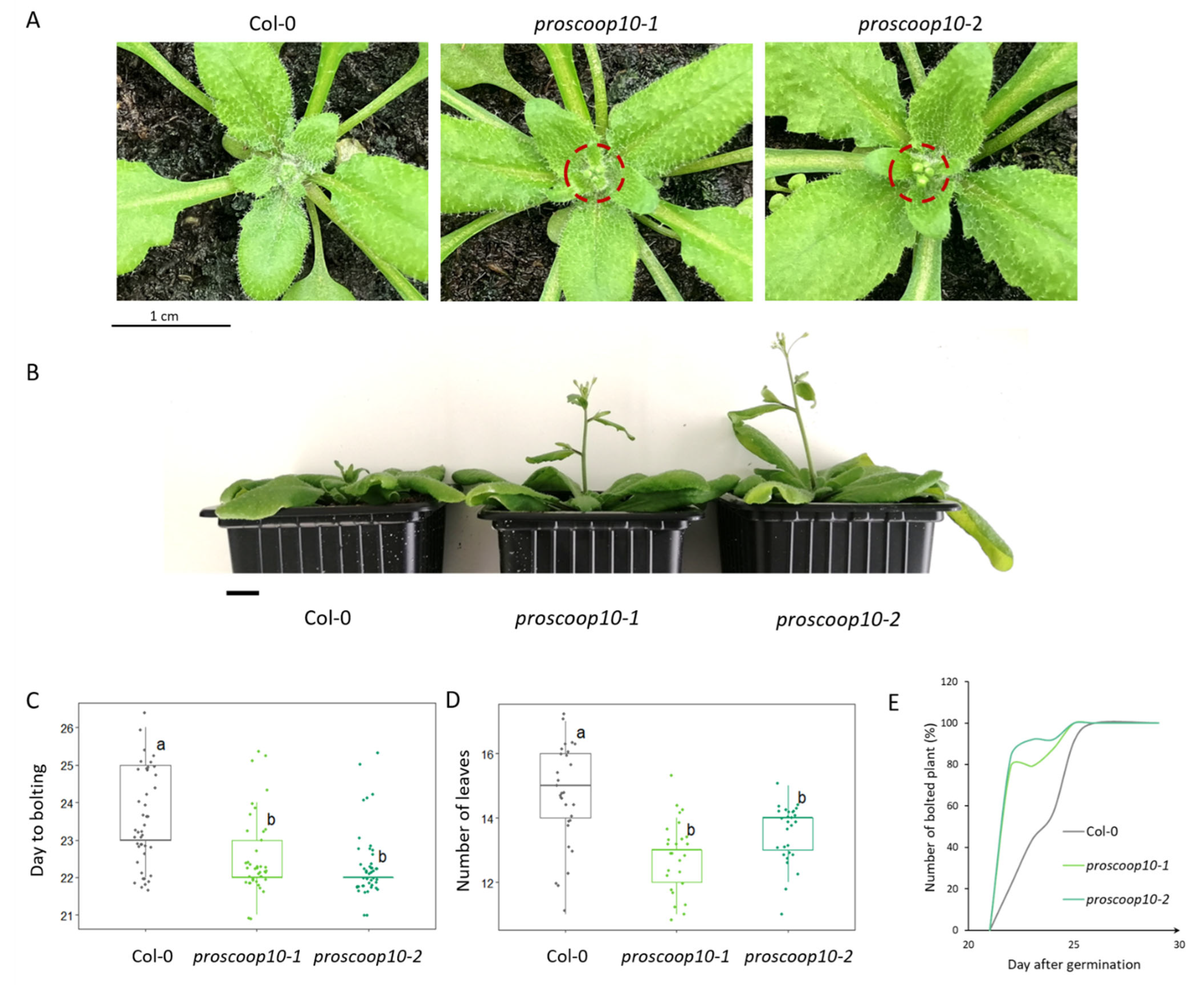
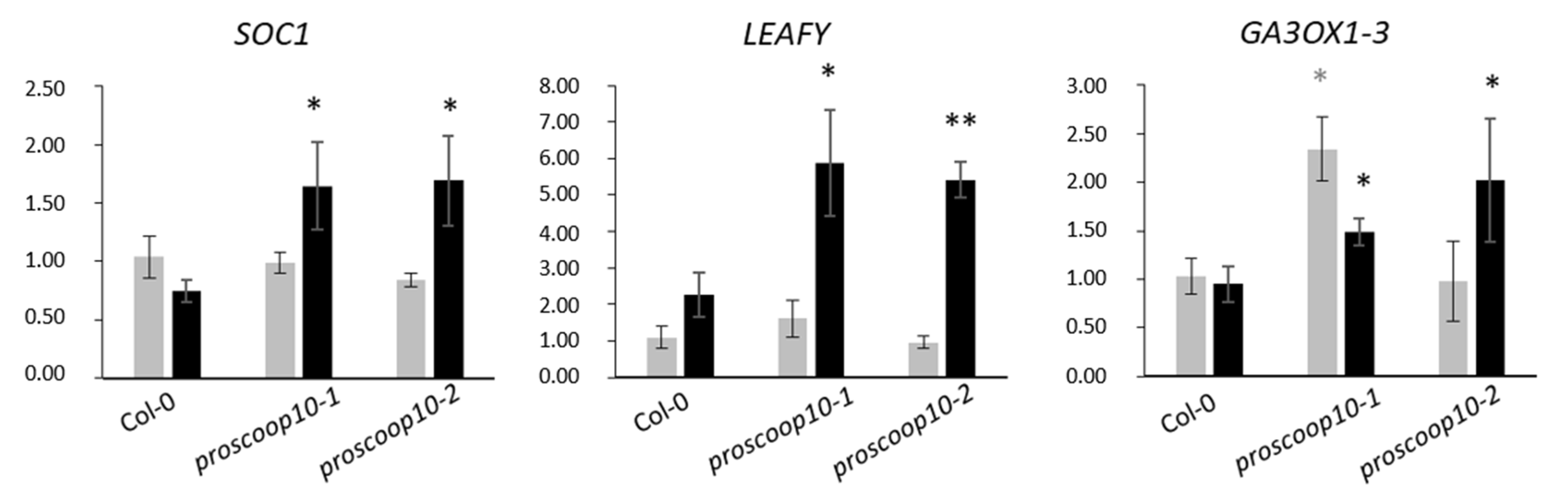
2.6. Mutations of PROSCOOP10 Impact Flowering Time
3. Discussion
3.1. Two Distinct Hydroxylated SCOOP10 Peptides Are Present in the Leaf Apoplasm
3.2. SCOOP10 Peptides Tend to Adopt a Hairpin Structure
3.3. PROSCOOP10 Delays the Floral Transition
4. Materials and Methods
4.1. Plant Material
4.2. Synthetic Peptides
4.3. Mass Spectrometry (MS) Analyses of Extracellular Fluids
4.4. Nuclear Magnetic Resonance (NMR) Analyses
4.5. Molecular Dynamics (MD) Simulations
4.6. Seedling Growth Inhibition Assay
4.7. ROS Assay
4.8. Gene Expression Analysis
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Murphy, E.; Smith, S.; De Smet, I. Small Signaling Peptides in Arabidopsis Development: How Cells Communicate over a Short Distance. Plant Cell 2012, 24, 3198–3217. [Google Scholar] [CrossRef] [PubMed]
- Matsubayashi, Y. Post-Translational Modifications in Secreted Peptide Hormones in Plants. Plant Cell Physiol. 2011, 52, 5–13. [Google Scholar] [CrossRef] [PubMed]
- Tavormina, P.; De Coninck, B.; Nikonorova, N.; De Smet, I.; Cammue, B.P.A. The Plant Peptidome: An Expanding Repertoire of Structural Features and Biological Functions. Plant Cell 2015, 27, 2095–2118. [Google Scholar] [CrossRef] [PubMed]
- Gully, K.; Pelletier, S.; Guillou, M.-C.; Ferrand, M.; Aligon, S.; Pokotylo, I.; Perrin, A.; Vergne, E.; Fagard, M.; Ruelland, E.; et al. The SCOOP12 Peptide Regulates Defense Response and Root Elongation in Arabidopsis thaliana. J. Exp. Bot. 2019, 70, 1349–1365. [Google Scholar] [CrossRef]
- Hou, S.; Liu, D.; Huang, S.; Luo, D.; Liu, Z.; Xiang, Q.; Wang, P.; Mu, R.; Han, Z.; Chen, S.; et al. The Arabidopsis MIK2 Receptor Elicits Immunity by Sensing a Conserved Signature from Phytocytokines and Microbes. Nat. Commun. 2021, 12, 1–15. [Google Scholar] [CrossRef]
- Rhodes, J.; Yang, H.; Moussu, S.; Boutrot, F.; Santiago, J.; Zipfel, C. Perception of a Divergent Family of Phytocytokines by the Arabidopsis Receptor Kinase MIK2. Nat. Commun. 2021, 12, 705. [Google Scholar] [CrossRef]
- Zhang, J.; Zhao, J.; Yang, Y.; Bao, Q.; Li, Y.; Wang, H.; Hou, S. EWR1 as a SCOOP Peptide Activates MIK2-Dependent Immunity in Arabidopsis. J. Plant Interact. 2022, 17, 562–568. [Google Scholar] [CrossRef]
- Yu, Z.; Xu, Y.; Zhu, L.; Zhang, L.; Liu, L.; Zhang, D.; Li, D.; Wu, C.; Huang, J.; Yang, G.; et al. The Brassicaceae-Specific Secreted Peptides, STMPs, Function in Plant Growth and Pathogen Defense. J. Integr. Plant Biol. 2019, 62, 403–420. [Google Scholar] [CrossRef]
- Guillou, M.-C.; Vergne, E.; Aligon, S.; Pelletier, S.; Simonneau, F.; Rolland, A.; Chabout, S.; Mouille, G.; Gully, K.; Grappin, P.; et al. The Peptide SCOOP12 Acts on Reactive Oxygen Species Homeostasis to Modulate Cell Division and Elongation in Arabidopsis Primary Root. J. Exp. Bot. 2022, 73, 6115–6132. [Google Scholar] [CrossRef]
- Zimmermann, P.; Hirsch-Hoffmann, M.; Hennig, L.; Gruissem, W. GENEVESTIGATOR. Arabidopsis Microarray Database and Analysis Toolbox. Plant Physiol. 2004, 136, 2621–2632. [Google Scholar] [CrossRef]
- Klepikova, A.V.; Kasianov, A.S.; Gerasimov, E.S.; Logacheva, M.D.; Penin, A.A. A High Resolution Map of the Arabidopsis thaliana Developmental Transcriptome Based on RNA-Seq Profiling. Plant J. 2016, 88, 1058–1070. [Google Scholar] [CrossRef] [PubMed]
- Clemente, H.S.; Jamet, E. WallProtDB, a Database Resource for Plant Cell Wall Proteomics. Plant Methods 2015, 11, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Clemente, H.S.; Kolkas, H.; Canut, H.; Jamet, E. Plant Cell Wall Proteomes: The Core of Conserved Protein Families and the Case of Non-Canonical Proteins. Int. J. Mol. Sci. 2022, 23, 4273. [Google Scholar] [CrossRef] [PubMed]
- Wishart, D.S.; Bigam, C.G.; Yao, J.; Abildgaard, F.; Dyson, H.J.; Oldfield, E.; Markley, J.L.; Sykes, B.D. 1H, 13C and 15N Chemical Shift Referencing in Biomolecular NMR. J. Biomol. NMR 1995, 6, 135–140. [Google Scholar] [CrossRef] [PubMed]
- Wishart, D.; Sykes, B. The 13C Chemical-Shift Index: A Simple Method for the Identification of Protein Secondary Structure Using 13C Chemical-Shift Data. J. Biomol. NMR 1994, 4, 171–180. [Google Scholar] [CrossRef]
- Wishart, D.S. Interpreting Protein Chemical Shift Data. Prog. Nucl. Magn. Reson. Spectrosc. 2011, 58, 62–87. [Google Scholar] [CrossRef]
- Nielsen, J.-T.; Mulder, F.-A.-A. POTENCI: Prediction of temperature, neighbor and pH-corrected chemical shifts for intrinsically disordered proteins. J. Biomol. NMR 2018, 70, 141–165. [Google Scholar] [CrossRef]
- Wang, Y. Probability-Based Protein Secondary Structure Identification Using Combined NMR Chemical-Shift Data. Protein Sci. 2002, 11, 852–861. [Google Scholar] [CrossRef]
- Dorman, D.E.; Bovey, F.A. Carbon-13 Magnetic Resonance Spectroscopy. Spectrum of Proline in Oligopeptides. J. Org. Chem. 1973, 38, 2379–2383. [Google Scholar] [CrossRef]
- Siemion, I.Z.; Wieland, T.; Pook, K.-H. Influence of the Distance of the Proline Carbonyl from the ß And Ɣ Carbon on The 13C Chemical Shifts. Angew. Chem. Int. Ed. Engl. 1975, 14, 702–703. [Google Scholar] [CrossRef]
- Wedemeyer, W.J.; Welker, E.; Scheraga, H.A. Proline Cis-Trans Isomerization and Protein Folding. Biochemistry 2002, 41, 14637–14644. [Google Scholar] [CrossRef] [PubMed]
- Gaggelli, E.; D’Amelio, N.; Gaggelli, N.; Valensin, G. Metal Ion Effects on the Cis/Trans Isomerization Equilibrium of Proline in Short-Chain Peptides: A Solution NMR Study. ChemBioChem 2001, 2, 524–529. [Google Scholar] [CrossRef] [PubMed]
- Pauwels, L.; De Clercq, R.; Goossens, J.; Iñigo, S.; Williams, C.; Ron, M.; Britt, A.; Goossens, A. A Dual SgRNA Approach for Functional Genomics in Arabidopsis thaliana. G3 Genes Genomes Genet. 2018, 8, 2603–2615. [Google Scholar]
- Roberts, I.; Smith, S.; De Rybel, B.; Van Den Broeke, J.; Smet, W.; De Cokere, S.; Mispelaere, M.; De Smet, I.; Beeckman, T. The CEP Family in Land Plants: Evolutionary Analyses, Expression Studies, and Role in Arabidopsis Shoot Development. J. Exp. Bot. 2013, 64, 5371–5381. [Google Scholar] [CrossRef]
- Vie, A.K.; Najafi, J.; Liu, B.; Winge, P.; Butenko, M.A.; Hornslien, K.S.; Kumpf, R.; Aalen, R.B.; Bones, A.M.; Brembu, T. The IDA/IDA-LIKE and PIP/PIP-LIKE Gene Families in Arabidopsis: Phylogenetic Relationship, Expression Patterns, and Transcriptional Effect of the PIPL3 Peptide. J. Exp. Bot. 2015, 66, 5351–5365. [Google Scholar] [CrossRef]
- Schardon, K.; Hohl, M.; Graff, L.; Pfannstiel, J.; Schulze, W.; Stintzi, A.; Schaller, A. Precursor Processing for Plant Peptide Hormone Maturation by Subtilisin-like Serine Proteinases. Science 2016, 354, 1594–1597. [Google Scholar] [CrossRef]
- Stintzi, A.; Schaller, A. Biogenesis of Post-Translationally Modified Peptide Signals for Plant Reproductive Development. Curr. Opin. Plant Biol. 2022, 69, 102274. [Google Scholar] [CrossRef]
- Casamitjana-Martinez, E.; Hofhuis, H.F.; Xu, J.; Liu, C.-M.; Heidstra, R.; Scheres, B. Root-Specific CLE19 Overexpression and the Sol1/2 Suppressors Implicate a CLV-like Pathway in the Control of Arabidopsis Root Meristem Maintenance. Curr. Biol. 2003, 13, 1435–1441. [Google Scholar] [CrossRef]
- Tamaki, T.; Betsuyaku, S.; Fujiwara, M.; Fukao, Y.; Fukuda, H.; Sawa, S. Suppressor of LLP1 1-Mediated C-Terminal Processing Is Critical for CLE19 Peptide Activity. Plant J. 2013, 76, 970–981. [Google Scholar] [CrossRef]
- Ohyama, K.; Ogawa, M.; Matsubayashi, Y. Identification of a Biologically Active, Small, Secreted Peptide in Arabidopsis by in Silico Gene Screening, Followed by LC-MS-Based Structure Analysis. Plant J. 2008, 55, 152–160. [Google Scholar] [CrossRef]
- Showalter, A.M.; Keppler, B.; Lichtenberg, J.; Gu, D.; Welch, L.R. A Bioinformatics Approach to the Identification, Classification, and Analysis of Hydroxyproline-Rich Glycoproteins. Plant Physiol. 2010, 153, 485–513. [Google Scholar] [CrossRef] [PubMed]
- Tan, L.; Leykam, J.F.; Kieliszewski, M.J. Glycosylation Motifs That Direct Arabinogalactan Addition to Arabinogalactan-Proteins. Plant Physiol. 2003, 132, 1362–1369. [Google Scholar] [CrossRef] [PubMed]
- Canut, H.; Albenne, C.; Jamet, E. Post-Translational Modifications of Plant Cell Wall Proteins and Peptides: A Survey from a Proteomics Point of View. Biochim. Biophys. Acta-Proteins Proteom. 2016, 1864, 983–990. [Google Scholar] [CrossRef] [PubMed]
- Duruflé, H.; Hervé, V.; Balliau, T.; Zivy, M.; Dunand, C.; Jamet, E. Proline Hydroxylation in Cell Wall Proteins: Is It Yet Possible to Define Rules? Front. Plant Sci. 2017, 8, 1802. [Google Scholar] [CrossRef] [PubMed]
- Royek, S.; Bayer, M.; Pfannstiel, J.; Pleiss, J.; Ingram, G.; Stintzi, A.; Schaller, A. Processing of a Plant Peptide Hormone Precursor Facilitated by Posttranslational Tyrosine Sulfation. Proc. Natl. Acad. Sci. USA 2022, 119, e2201195119. [Google Scholar] [CrossRef]
- Shinohara, H.; Matsubayashi, Y. Arabinosylated Glycopeptide Hormones: New Insights into CLAVATA3 Structure. Curr. Opin. Plant Biol. 2010, 13, 515–519. [Google Scholar] [CrossRef] [PubMed]
- Araya, T.; von Wirén, N.; Takahashi, H. CLE Peptides Regulate Lateral Root Development in Response to Nitrogen Nutritional Status of Plants. Plant Signal. Behav. 2014, 9, e29302. [Google Scholar] [CrossRef]
- Takahashi, F.; Suzuki, T.; Osakabe, Y.; Betsuyaku, S.; Kondo, Y.; Dohmae, N.; Fukuda, H.; Yamaguchi-Shinozaki, K.; Shinozaki, K. A Small Peptide Modulates Stomatal Control via Abscisic Acid in Long-Distance Signalling. Nature 2018, 556, 235–238. [Google Scholar] [CrossRef]
- Amano, Y.; Tsubouchi, H.; Shinohara, H.; Ogawa, M.; Matsubayashi, Y. Tyrosine-Sulfated Glycopeptide Involved in Cellular Proliferation and Expansion in Arabidopsis. Proc. Natl. Acad. Sci. USA 2007, 104, 18333–18338. [Google Scholar] [CrossRef]
- Pucheta-Martinez, E.; Saladino, G.; Morando, M.A.; Martinez-Torrecuadrada, J.; Lelli, M.; Sutto, L.; D’Amelio, N.; Gervasio, F.L. An Allosteric Cross-Talk Between the Activation Loop and the ATP Binding Site Regulates the Activation of Src Kinase. Sci. Rep. 2016, 6, 24235. [Google Scholar] [CrossRef]
- Sekhar, A.; Velyvis, A.; Zoltsman, G.; Rosenzweig, R.; Bouvignies, G.; Kay, L.E. Conserved Conformational Selection Mechanism of Hsp70 Chaperone-Substrate Interactions. eLife 2018, 7, e32764. [Google Scholar] [CrossRef] [PubMed]
- Parcy, F. Flowering: A Time for Integration. Int. J. Dev. Biol. 2005, 49, 585–593. [Google Scholar] [CrossRef] [PubMed]
- Moon, J.; Lee, H.; Kim, M.; Lee, I. Analysis of Flowering Pathway Integrators in Arabidopsis. Plant Cell Physiol. 2005, 46, 292–299. [Google Scholar] [CrossRef] [PubMed]
- Roux, F.; Touzet, P.; Cuguen, J.; Corre, V.L. How to Be Early Flowering: An Evolutionary Perspective. Trends Plant Sci. 2006, 11, 375–381. [Google Scholar] [CrossRef]
- Li, J.; Li, Y.; Chen, S.; An, L. Involvement of Brassinosteroid Signals in the Floral-Induction Network of Arabidopsis. J. Exp. Bot. 2010, 61, 4221–4230. [Google Scholar] [CrossRef]
- Simpson, G.G.; Dean, C. Arabidopsis, the Rosetta Stone of Flowering Time? Science 2002, 296, 285–289. [Google Scholar] [CrossRef]
- Blázquez, M.A.; Green, R.; Nilsson, O.; Sussman, M.R.; Weigel, D. Gibberellins Promote Flowering of Arabidopsis by Activating the LEAFY Promoter. Plant Cell 1998, 10, 791–800. [Google Scholar] [CrossRef]
- Klepikova, A.V.; Logacheva, M.D.; Dmitriev, S.E.; Penin, A.A. RNA-Seq Analysis of an Apical Meristem Time Series Reveals a Critical Point in Arabidopsis thaliana Flower Initiation. BMC Genom. 2015, 16, 1–16. [Google Scholar] [CrossRef]
- Gagnot, S.; Tamby, J.P.; Martin-Magniette, M.L.; Bitton, F.; Taconnat, L.; Balzergue, S.; Aubourg, S.; Renou, J.P.; Lecharny, A.; Brunaud, V. CATdb: A Public Access to Arabidopsis Transcriptome Data from the URGV-CATMA Platform. Nucleic Acids Res. 2008, 36 (Suppl. S1), D986–D990. [Google Scholar] [CrossRef][Green Version]
- Foyer, C.H.; Kerchev, P.I.; Hancock, R.D. The ABA-INSENSITIVE-4 (ABI4) Transcription Factor Links Redox, Hormone and Sugar Signaling Pathways. Plant Signal. Behav. 2012, 7, 276–281. [Google Scholar] [CrossRef]
- Hu, W.; Franklin, K.A.; Sharrock, R.A.; Jones, M.A.; Harmer, S.L.; Lagarias, J.C. Unanticipated Regulatory Roles for Arabidopsis phytochromes revealed by null mutant analysis. Proc. Natl. Acad. Sci. USA 2013, 110, 1542–1547. [Google Scholar] [CrossRef] [PubMed]
- Pavet, V.; Olmos, E.; Kiddle, G.; Mowla, S.; Kumar, S.; Antoniw, J.; Alvarez, M.E.; Foyer, C.H. Ascorbic Acid Deficiency Activates Cell Death and Disease Resistance Responses in Arabidopsis. Plant Physiol. 2005, 139, 1291–1303. [Google Scholar] [CrossRef]
- Kubota, A.; Ito, S.; Shim, J.S.; Johnson, R.S.; Song, Y.H.; Breton, G.; Goralogia, G.S.; Kwon, M.S.; Cintrón, D.L.; Koyama, T.; et al. TCP4-Dependent Induction of CONSTANS Transcription Requires GIGANTEA in Photoperiodic Flowering in Arabidopsis. PLoS Genet. 2017, 13, e1006856. [Google Scholar] [CrossRef] [PubMed]
- Nagpal, P.; Ellis, C.M.; Weber, H.; Ploense, S.E.; Barkawi, L.S.; Guilfoyle, T.J.; Hagen, G.; Alonso, J.M.; Cohen, J.D.; Farmer, E.E.; et al. Auxin Response Factors ARF6 and ARF8 Promote Jasmonic Acid Production and Flower Maturation. Development 2005, 132, 4107–4118. [Google Scholar] [CrossRef] [PubMed]
- Moon, J.; Zhu, L.; Shen, H.; Huq, E. PIF1 Directly and Indirectly Regulates Chlorophyll Biosynthesis to Optimize the Greening Process in Arabidopsis. Proc. Natl. Acad. Sci. USA 2008, 105, 9433–9438. [Google Scholar] [CrossRef]
- Pečenková, T.; Pejchar, P.; Moravec, T.; Drs, M.; Haluška, S.; Šantrůček, J.; Potocká, A.; Žárský, V.; Potocký, M. Immunity Functions of Arabidopsis Pathogenesis-Related 1 Are Coupled but Not Confined to Its C-Terminus Processing and Trafficking. Mol. Plant Pathol. 2022, 23, 664–678. [Google Scholar] [CrossRef]
- Wu, M.; Liu, D.; Abdul, W.; Upreti, S.; Liu, Y.; Song, G.; Wu, J.; Liu, B.; Gan, Y. PIL5 Represses Floral Transition in Arabidopsis under Long Day Conditions. Biochem. Biophys. Res. Commun. 2018, 499, 513–518. [Google Scholar] [CrossRef]
- Pak, H.; Guo, Y.; Chen, M.; Chen, K.; Li, Y.; Hua, S.; Shamsi, I.; Meng, H.; Shi, C.; Jiang, L. The Effect of Exogenous Methyl Jasmonate on the Flowering Time, Floral Organ Morphology, and Transcript Levels of a Group of Genes Implicated in the Development of Oilseed Rape Flowers (Brassica napus L.). Planta 2009, 231, 79–91. [Google Scholar] [CrossRef]
- Chávez Montes, R.A.; Ranocha, P.; Martinez, Y.; Minic, Z.; Jouanin, L.; Marquis, M.; Saulnier, L.; Fulton, L.M.; Cobbett, C.S.; Bitton, F.; et al. Cell Wall Modifications in Arabidopsis Plants with Altered α-l-Arabinofuranosidase Activity. Plant Physiol. 2008, 147, 63–77. [Google Scholar] [CrossRef]
- Charrier, A.; Vergne, E.; Dousset, N.; Richer, A.; Petiteau, A.; Chevreau, E. Efficient Targeted Mutagenesis in Apple and First Time Edition of Pear Using the CRISPR-Cas9 System. Front. Plant Sci. 2019, 10, 40. [Google Scholar] [CrossRef]
- Zhang, X.; Henriques, R.; Lin, S.S.; Niu, Q.W.; Chua, N.H. Agrobacterium-Mediated Transformation of Arabidopsis thaliana Using the Floral Dip Method. Nat. Protoc. 2006, 1, 641–646. [Google Scholar] [CrossRef] [PubMed]
- Boudart, G.; Jamet, E.; Rossignol, M.; Lafitte, C.; Borderies, G.; Jauneau, A.; Esquerré-Tugayé, M.-T.; Pont-Lezica, R. Cell Wall Proteins in Apoplastic Fluids Of Arabidopsis thaliana rosettes: Identification by Mass Spectrometry and Bioinformatics. Proteomics 2004, 5, 212–221. [Google Scholar] [CrossRef] [PubMed]
- Balliau, T.; Blein-Nicolas, M.; Zivy, M. Evaluation of Optimized Tube-Gel Methods of Sample Preparation for Large-Scale Plant Proteomics. Proteomes 2018, 6, 6. [Google Scholar] [CrossRef] [PubMed]
- Duruflé, H.; Ranocha, P.; Balliau, T.; Dunand, C.; Jamet, E. Transcriptomic and Cell Wall Proteomic Datasets of Rosettes and Floral Stems from Five Arabidopsis thaliana Ecotypes Grown at Optimal or Sub-Optimal Temperature. Data Br. 2019, 27, 104581. [Google Scholar] [CrossRef] [PubMed]
- Langella, O.; Valot, B.; Balliau, T.; Blein-Nicolas, M.; Bonhomme, L.; Zivy, M. X!TandemPipeline: A Tool to Manage Sequence Redundancy for Protein Inference and Phosphosite Identification. J. Proteome Res. 2016, 16, 494–503. [Google Scholar] [CrossRef]
- Lee, W.; Tonelli, M.; Markley, J.L. NMRFAM-SPARKY: Enhanced Software for Biomolecular NMR Spectroscopy. Development 2015, 132, 4107–4118. [Google Scholar] [CrossRef]
- Lee, W.; Yu, W.; Kim, S.; Chang, I.; Lee, W.; Markley, J.L. PACSY, a Relational Database Management System for Protein Structure and Chemical Shift Analysis. J. Biomol. NMR 2012, 54, 169–179. [Google Scholar] [CrossRef]
- Yang, J.; Yan, R.; Roy, A.; Xu, D.; Poisson, J.; Zhang, Y. The I-TASSER Suite: Protein Structure and Function Prediction. Nat. Methods 2014, 12, 7–8. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera? A Visualization System for Exploratory Research and Analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef]
- Krieger, E.; Vriend, G. YASARA View—Molecular Graphics for All Devices—From Smartphones to Workstations. Bioinformatics 2014, 30, 2981–2982. [Google Scholar] [CrossRef]
- Abraham, M.J.; Murtola, T.; Schulz, R.; Páll, S.; Smith, J.C.; Hess, B.; Lindahl, E. GROMACS: High Performance Molecular Simulations through Multi-Level Parallelism from Laptops to Supercomputers. SoftwareX 2015, 1–2, 19–25. [Google Scholar] [CrossRef]
- Lindorff-Larsen, K.; Piana, S.; Palmo, K.; Maragakis, P.; Klepeis, J.L.; Dror, R.O.; Shaw, D.E. Improved Side-Chain Torsion Potentials for the Amber Ff99SB Protein Force Field. Proteins Struct. Funct. Bioinform. 2010, 78, 1950–1958. [Google Scholar] [CrossRef] [PubMed]
- Pokotylo, I.; Hodges, M.; Kravets, V.; Ruelland, E. A Ménage à Trois: Salicylic Acid, Growth Inhibition, and Immunity. Trends Plant Sci. 2021, 27, 460–471. [Google Scholar] [CrossRef] [PubMed]
- DeLano, W.L. Pymol: An Open-Source Molecular Graphics Tool. CCP4 Newsl. Protein Crystallogr. 2002, 40, 82–92. [Google Scholar]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD: Visual Molecular Dynamics. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef] [PubMed]
- Janert, P. Gnuplot in Action: Understanding Data with Graphs; Manning Publications: New York, NY, USA, 2009. [Google Scholar]
- Lemkul, J. From Proteins to Perturbed Hamiltonians: A Suite of Tutorials for the GROMACS. Living J. Comput. Mol. Sci. 2019, 1, 5068. [Google Scholar]
- Francisco, R.-M.; Herrera-León, C.; Antonietti, V.; Sonnet, P.; Sarazin, C.; D’Amelio, N. Antimicrobial Peptide K11 Selectively Recognizes Bacterial Biomimetic Membranes and Acts by Twisting Their Bilayers. Pharmaceuticals 2020, 14, 1. [Google Scholar]
- Vandesompele, J.; De Preter, K.; Pattyn, F.; Poppe, B.; Van Roy, N.; De Paepe, A.; Speleman, F. Accurate Normalization of Real-Time Quantitative RT-PCR Data by Geometric Averaging of Multiple Internal Control Genes. Genome Biol. 2002, 3, 1–12. [Google Scholar] [CrossRef] [PubMed]



| Condition | Phenotype | PROSCOOP10 Transcription | Reference |
|---|---|---|---|
| 10th vs. 9th day in SAM wild-type | Wild-type | Down-regulated | [48] |
| pif1-5vs. wild-type | Early-flowering | Down-regulated | [55,57] |
| abi4vtc2vs. wild-type | Early-flowering | Down-regulated | [50] |
| phyABCDEvs. wild-type | Early-flowering | Down-regulated | [51] |
| Brassinolide treatment in wild-type | Early-flowering | Down-regulated | [45] |
| MeJA treatment in wild-type | Early-flowering | Down-regulated | [58] |
| 35S::ARAF1 vs. wild-type | Late-flowering | Up-regulated | [59] |
| vtc2vs. wild-type | Late-flowering | Up-regulated | [52] |
| tcp4vs. wild-type | Late-flowering | Up-regulated | [53] |
| ga1-3vs. wild-type | No flowering | Up-regulated | [47] |
| arf6-2/arf8-3vs. wild-type | Immature flowers | Up-regulated | [54] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Guillou, M.-C.; Balliau, T.; Vergne, E.; Canut, H.; Chourré, J.; Herrera-León, C.; Ramos-Martín, F.; Ahmadi-Afzadi, M.; D’Amelio, N.; Ruelland, E.; et al. The PROSCOOP10 Gene Encodes Two Extracellular Hydroxylated Peptides and Impacts Flowering Time in Arabidopsis. Plants 2022, 11, 3554. https://doi.org/10.3390/plants11243554
Guillou M-C, Balliau T, Vergne E, Canut H, Chourré J, Herrera-León C, Ramos-Martín F, Ahmadi-Afzadi M, D’Amelio N, Ruelland E, et al. The PROSCOOP10 Gene Encodes Two Extracellular Hydroxylated Peptides and Impacts Flowering Time in Arabidopsis. Plants. 2022; 11(24):3554. https://doi.org/10.3390/plants11243554
Chicago/Turabian StyleGuillou, Marie-Charlotte, Thierry Balliau, Emilie Vergne, Hervé Canut, Josiane Chourré, Claudia Herrera-León, Francisco Ramos-Martín, Masoud Ahmadi-Afzadi, Nicola D’Amelio, Eric Ruelland, and et al. 2022. "The PROSCOOP10 Gene Encodes Two Extracellular Hydroxylated Peptides and Impacts Flowering Time in Arabidopsis" Plants 11, no. 24: 3554. https://doi.org/10.3390/plants11243554
APA StyleGuillou, M.-C., Balliau, T., Vergne, E., Canut, H., Chourré, J., Herrera-León, C., Ramos-Martín, F., Ahmadi-Afzadi, M., D’Amelio, N., Ruelland, E., Zivy, M., Renou, J.-P., Jamet, E., & Aubourg, S. (2022). The PROSCOOP10 Gene Encodes Two Extracellular Hydroxylated Peptides and Impacts Flowering Time in Arabidopsis. Plants, 11(24), 3554. https://doi.org/10.3390/plants11243554








