Genome-Wide Association Study Reveals Marker–Trait Associations for Early Vegetative Stage Salinity Tolerance in Rice
Abstract
1. Introduction
2. Results
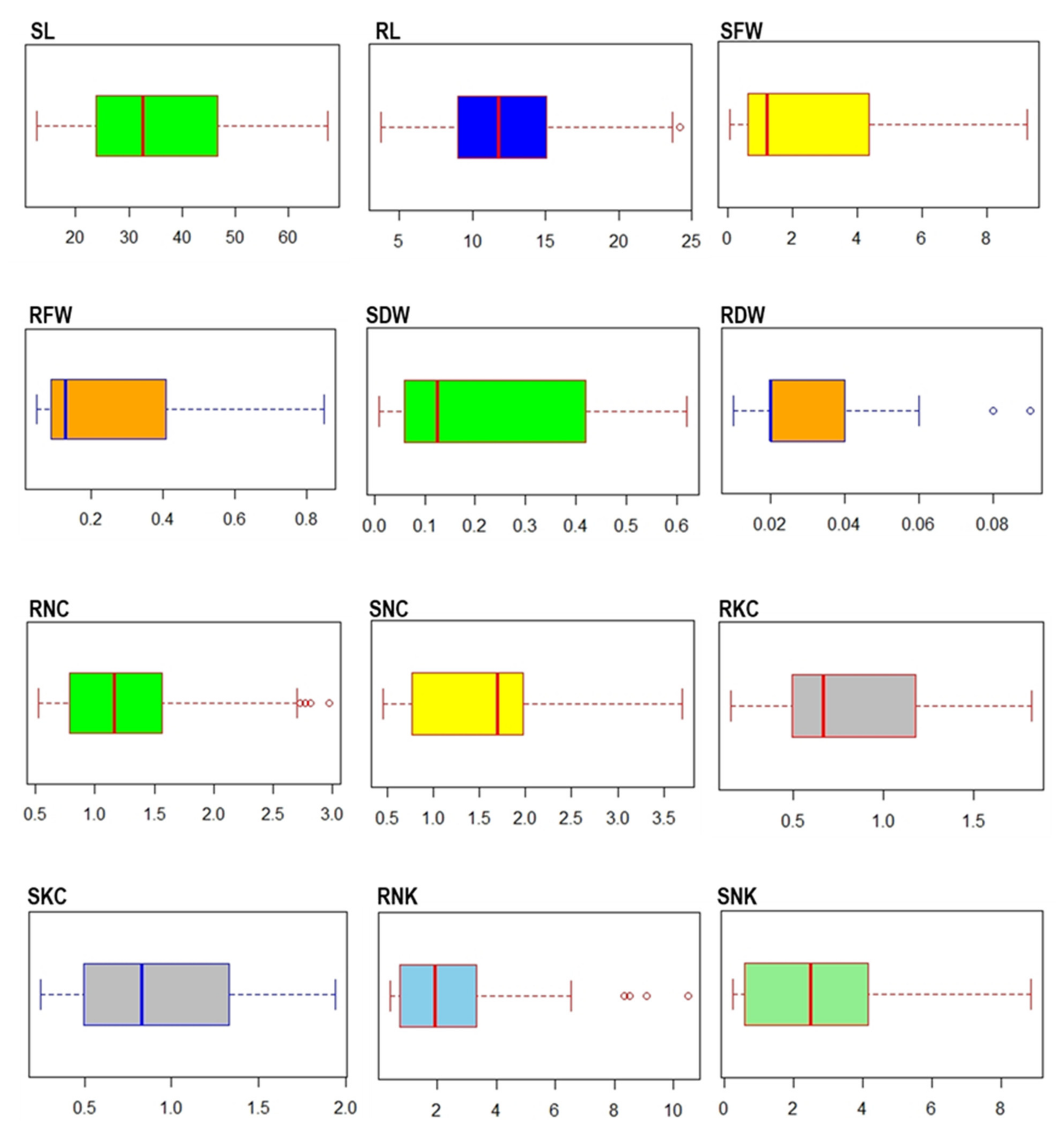
2.1. Phenotypic Evaluation
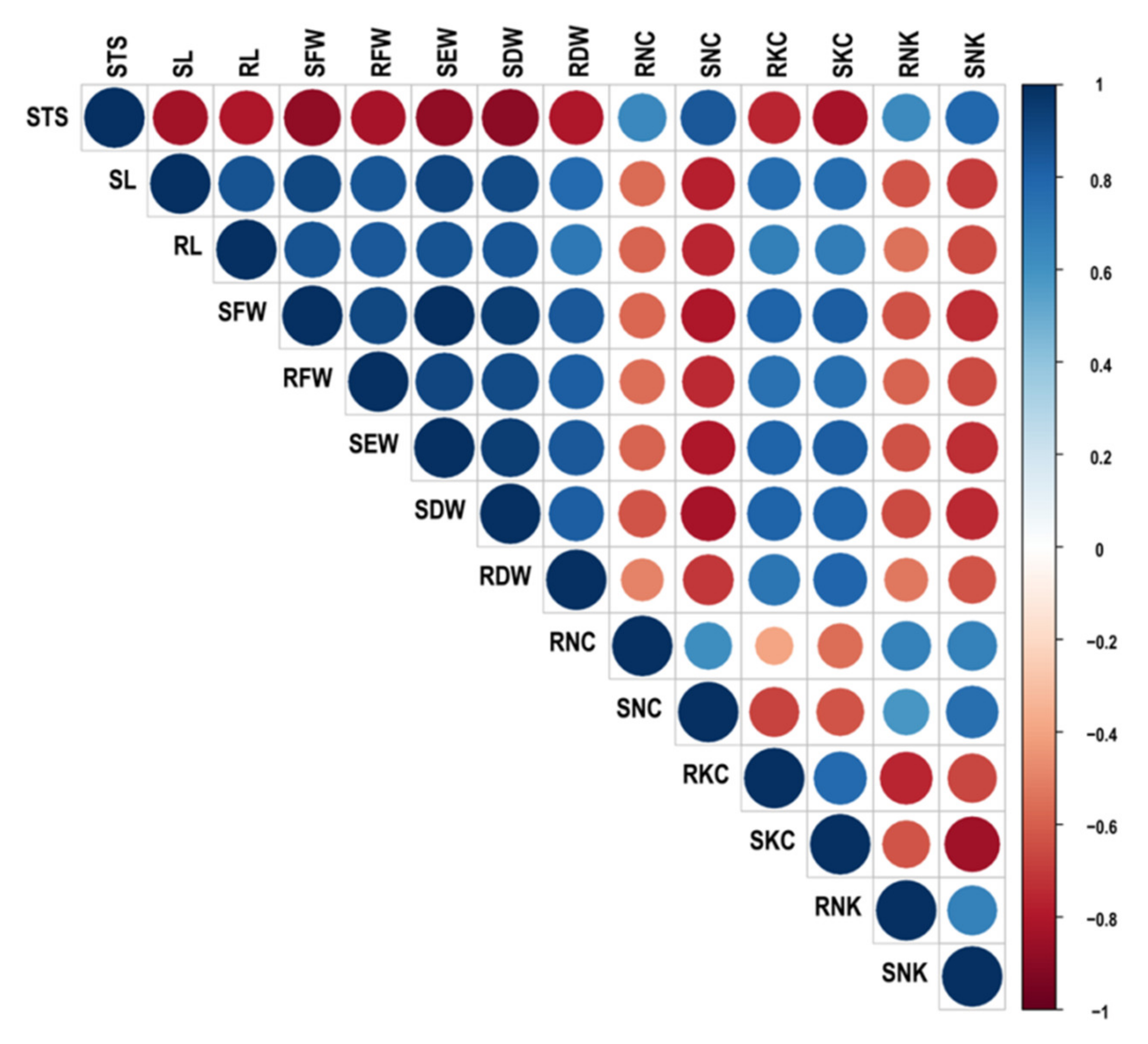
2.2. Correlation among Traits Related to Salt Stress
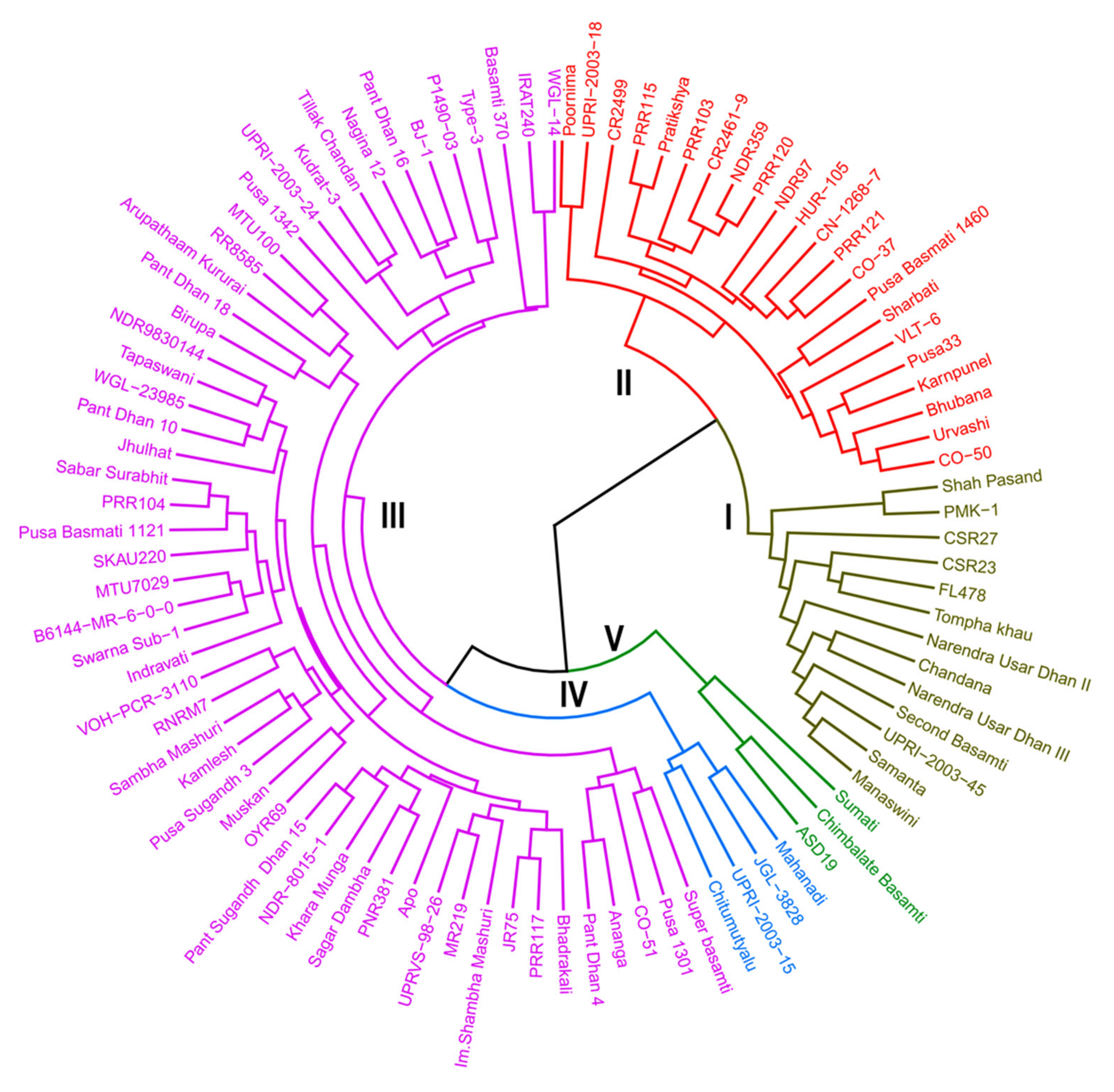
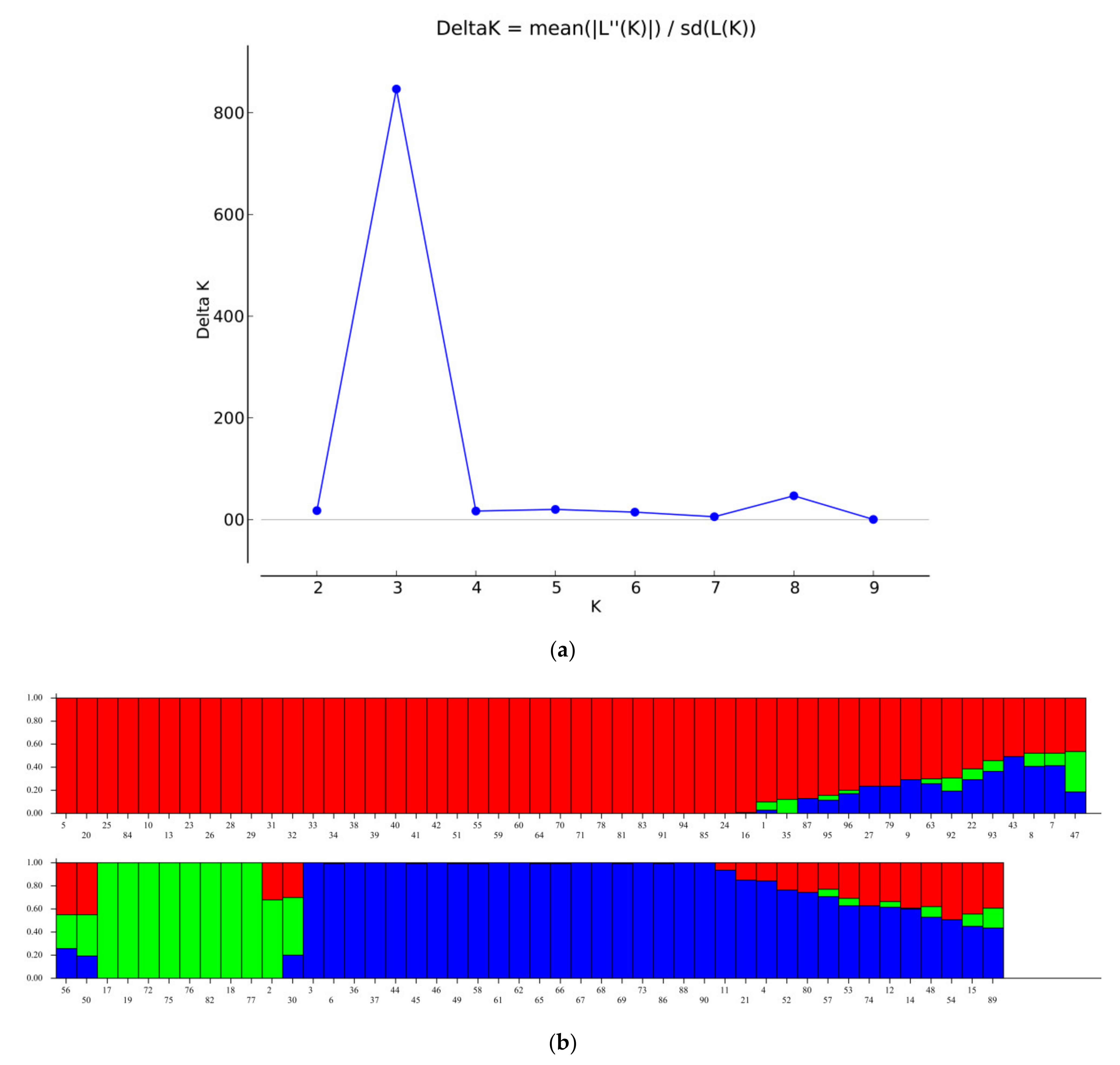
2.3. Population Structure
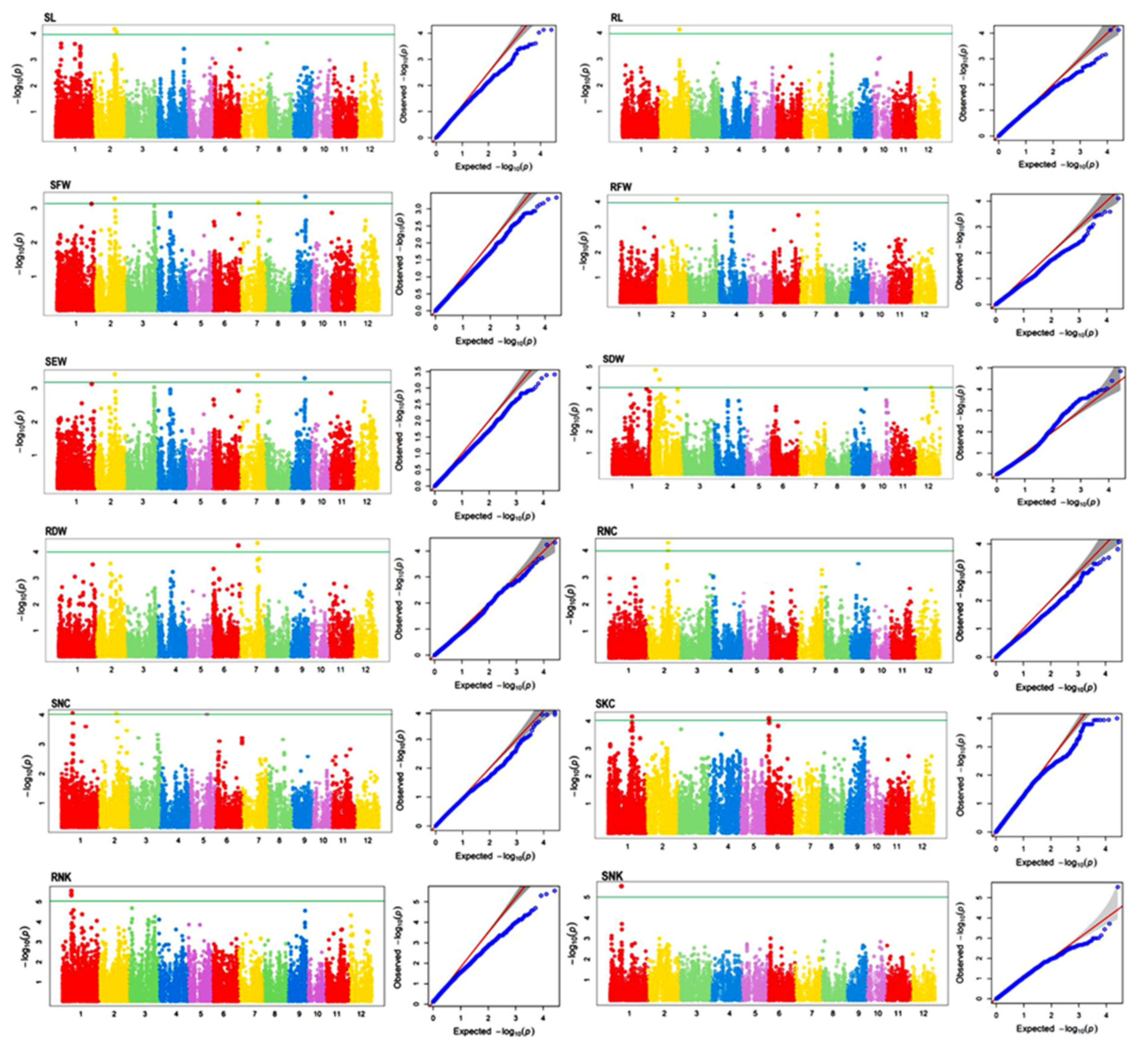
2.4. Genome-Wide Association Study for Traits Associated with Salinity Tolerance
3. Discussion
4. Material and Methods
4.1. Plant Materials
4.2. Evaluation for Seedling Stage Salinity Tolerance
4.3. Measurement of Morpho-Physiological Characters
4.4. Estimation of Na+ and K+ Ion Concentration
4.5. Data Analysis
4.6. DNA Isolation and SNP Genotyping
4.7. Population Structure Analysis
4.8. Genome-Wide Association Study
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- FAOSTAT. FAO Rice Market Monitor; Food and Agriculture Organization of the United Nations: Rome, Italy, 2017. [Google Scholar]
- Turral, H.; Burke, J.; Faurès, J.M. Climate Change, Water and Food Security; Food and Agriculture Organization of the United Nations (FAO): Rome, Italy, 2011; p. 36. [Google Scholar]
- Bouman, B.A.; Barker, R.; Humphreys, E.; Tuong, T.P.; Atlin, G.; Bennett, J.; Dawe, D.; Dittert, K.; Dobermann, A.; Facon, T.; et al. Rice: Feeding the Billions. In Water for Food, Water for Life: A Comprehensive Assessment of Water Management in Agriculture; IWMI: Colombo, Sri Lanka, 2007; pp. 515–549. Available online: https://cgspace.cgiar.org/handle/10568/17345 (accessed on 12 February 2021).
- Munns, R.; Tester, M. Mechanisms of salinity tolerance. Annu. Rev. Plant Biol. 2008, 59, 651–681. [Google Scholar] [CrossRef] [PubMed]
- Provin, T.; Pitt, J.L. Managing Soil Salinity. Texas FARMER Collection. 2001. Available online: https://oaktrust.library.tamu.edu/bitstream/handle/1969.1/86985/pdf_1397.pdf?sequence=1 (accessed on 12 February 2021).
- Lutts, S.; Kinet, J.M.; Bouharmont, J. Changes in plant response to NaCl during develop-ment of rice (Oryza sativa L.) varieties differing in salinity resistance. J. Exp. Bot. 1995, 46, 1843–1852. [Google Scholar] [CrossRef]
- García, A.; Rizzo, C.; Ud-Din, J.; Bartos, S.; Senadhira, D.; Flowers, T.; Yeo, A. Sodium and potassium transport to the xylem are inherited independently in rice, and the mechanism of sodium: Potassium selectivity differs between rice and wheat. Plant Cell Environ. 1997, 20, 1167–1174. [Google Scholar] [CrossRef]
- Zeng, L.; Shannon, M.C. Salinity Effects on Seedling Growth and Yield Components of Rice. Crop. Sci. 2000, 40, 996–1003. [Google Scholar] [CrossRef]
- Tester, M.; Davenport, R. Na+ tolerance and Na+ transport in higher plants. Ann. Bot. 2003, 91, 503–527. [Google Scholar] [CrossRef]
- Horie, T.; Karahara, I.; Katsuhara, M. Salinity tolerance mechanisms in glycophytes: An overview with the central focus on rice plants. Rice 2012, 5, 1–8. [Google Scholar] [CrossRef]
- Todaka, D.; Nakashima, K.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Toward understand-ing transcriptional regulatory networks in abiotic stress responses and tolerance in rice. Rice 2012, 5, 1–9. [Google Scholar] [CrossRef]
- Grieve, C.M.; Grattan, S.R.; Maas, E.V. Plant salt tolerance. ASCE Man. Rep. Eng. Pract. 2012, 71, 405–459. [Google Scholar]
- Umali, D.L. Irrigation-Induced Salinity: A Growing Problem for Development and the Environment; World Bank Publications: Washington, DC, USA, 1993; p. 125. [Google Scholar]
- Gregoria, G.B.; Senadhira, D.; Mendoza, R.D. Screening Rice for Salinity Tolerance. 1997. Available online: http://www.knowledgebank.irri.org/ricebreedingcourse/documents/Screening_manual.pdf (accessed on 13 March 2021).
- Gregorio, G.B.; Senadhira, D. Genetic analysis of salinity tolerance in rice (Oryza sativa L.). Theor. Appl. Genet. 1993, 86, 333–338. [Google Scholar] [CrossRef]
- Walia, H.; Wilson, C.; Condamine, P.; Liu, X.; Ismail, A.M.; Zeng, L.; Wanamaker, S.I.; Mandal, J.; Xu, J.; Cui, X.; et al. Comparative transcriptional profiling of two con-trasting rice genotypes under salinity stress during the vegetative growth stage. Plant Physiol. 2005, 139, 822–835. [Google Scholar] [CrossRef]
- Bonilla, P.; Dvorak, J.; Mackell, D.; Deal, K.; Gregorio, G. RFLP and SSLP Mapping of Salinity Tolerance Genes in Chromosome 1 of Rice (Oryza sativa L.) Using Recombinant Inbred Lines. In Philippine Agricultural Scientist (Philippines); ISSN National Centre of the Philippines: Manila, Philippines, 2002; Available online: https://pas.cafs.uplb.edu.ph/ (accessed on 13 March 2021).
- Lee, K.-S.; Choi, W.-Y.; Ko, J.-C.; Kim, T.-S.; Gregorio, G.B. Salinity tolerance of japonica and indica rice (Oryza sativa L.) at the seedling stage. Planta 2003, 216, 1043–1046. [Google Scholar] [CrossRef]
- Koyama, M.L.; Levesley, A.; Koebner, R.M.; Flowers, T.J.; Yeo, A.R. Quantitative Trait Loci for Component Physiological Traits Determining Salt Tolerance in Rice. Plant Physiol. 2001, 125, 406–422. [Google Scholar] [CrossRef] [PubMed]
- Lin, H.X.; Zhu, M.Z.; Yano, M.; Gao, J.P.; Liang, Z.W.; Su, W.A.; Hu, X.H.; Ren, Z.H.; Chao, D.Y. QTLs for Na+ and K+ uptake of the shoots and roots controlling rice salt tolerance. Theor. Appl. Genet. 2004, 108, 253–260. [Google Scholar] [CrossRef] [PubMed]
- Yao, M.Z.; Wang, J.F.; Chen, H.U.; Zhai, H.Q.; Zhang, H.S. Inheritance and QTL map-ping of salt tolerance in rice. Rice Sci. 2005, 12, 25–32. [Google Scholar]
- Lee, S.Y.; Ahn, J.H.; Cha, Y.S.; Yun, D.W.; Lee, M.C.; Ko, J.C.; Lee, K.S.; Eun, M.Y. Mapping QTLs related to salinity tolerance of rice at the young seedling stage. Plant Breed. 2007, 126, 43–46. [Google Scholar] [CrossRef]
- Sabouri, H.; Sabouri, A. New evidence of QTLs attributed to salinity tolerance in rice. Afr. J. Biotechnol. 2008, 7, 24. [Google Scholar]
- Thomson, M.J.; de Ocampo, M.; Egdane, J.; Rahman, M.A.; Sajise, A.G.; Adorada, D.L.; Tumimbang-Raiz, E.; Blumwald, E.; Seraj, Z.I.; Singh, R.K.; et al. Charac-terizing the Saltol quantitative trait locus for salinity tolerance in rice. Rice 2010, 3, 148–160. [Google Scholar] [CrossRef]
- Islam, M.R.; Hassan, L.; Salam, M.A.; Collard, B.C.; Singh, R.K.; Gregorio, G.B. QTL mapping for salinity tolerance at seedling stage in rice. Emir. J. Food Agric. 2011, 15, 137–146. [Google Scholar] [CrossRef]
- Cheng, L.; Wang, Y.; Meng, L.; Hu, X.; Cui, Y.; Sun, Y.; Zhu, L.; Ali, J.; Xu, J.; Li, Z. Identification of salt-tolerant QTLs with strong genetic background effect using two sets of reciprocal introgression lines in rice. Genome 2012, 55, 45–55. [Google Scholar] [CrossRef]
- Chen, T.; Zhu, Y.; Chen, K.; Shen, C.; Zhao, X.; Shabala, S.; Shabala, L.; Meinke, H.; Venkataraman, G.; Chen, Z.; et al. Identification of new QTL for salt tolerance from rice variety Pokkali. J. Agron. Crop. Sci. 2020, 206, 202–213. [Google Scholar] [CrossRef]
- Yadav, A.K.; Kumar, A.; Grover, N.; Ellur, R.K.; Krishnan, S.G.; Bollinedi, H.; Bhowmick, P.K.; Vinod, K.K.; Nagarajan, M.; Krishnamurthy, S.L.; et al. Marker aided introgression of ‘Saltol’, a major QTL for seedling stage salinity tolerance into an elite Basmati rice variety ‘Pusa Basmati 1509’. Sci. Rep. 2020, 10, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Cardon, L.R.; Bell, J.I. Association study designs for complex diseases. Nat. Rev. Genet. 2001, 2, 91–99. [Google Scholar] [CrossRef]
- Wang, J.; McClean, P.E.; Lee, R.; Goos, R.J.; Helms, T. Association mapping of iron deficiency chlorosis loci in soybean (Glycine max L. Merr.) advanced breeding lines. Theor. Appl. Genet. 2008, 116, 777–787. [Google Scholar] [CrossRef]
- Mackay, I.; Powell, W. Methods for linkage disequilibrium mapping in crops. Trends Plant Sci. 2007, 12, 57–63. [Google Scholar] [CrossRef]
- Zhu, C.; Gore, M.; Buckler, E.S.; Yu, J. Status and Prospects of Association Mapping in Plants. Plant Genome 2008, 1. [Google Scholar] [CrossRef]
- Yu, J.; Buckler, E.S. Genetic association mapping and genome organization of maize. Curr. Opin. Biotechnol. 2006, 17, 155–160. [Google Scholar] [CrossRef] [PubMed]
- Myles, S.; Peiffer, J.; Brown, P.J.; Ersoz, E.S.; Zhang, Z.; Costich, D.E.; Buckler, E.S. As-sociation mapping: Critical considerations shift from genotyping to experimental design. Plant Cell 2009, 21, 2194–2202. [Google Scholar] [CrossRef]
- Negrão, S.; Courtois, B.; Ahmadi, N.; Abreu, I.; Saibo, N.; Oliveira, M. Recent Updates on Salinity Stress in Rice: From Physiological to Molecular Responses. Crit. Rev. Plant Sci. 2011, 30, 329–377. [Google Scholar] [CrossRef]
- Shi, Y.; Gao, L.; Wu, Z.; Zhang, X.; Wang, M.; Zhang, C.; Zhang, F.; Zhou, Y.; Li, Z. Ge-nome-wide association study of salt tolerance at the seed germination stage in rice. BMC Plant Biol. 2017, 17, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Patishtan, J.; Hartley, T.N.; de Carvalho, R.F.; Maathuis, F.J. Genome-wide associa-tion studies to identify rice salt-tolerance markers. Plant Cell Environ. 2018, 41, 970–982. [Google Scholar] [CrossRef]
- Lekklar, C.; Pongpanich, M.; Suriya-Arunroj, D.; Chinpongpanich, A.; Tsai, H.; Comai, L.; Chadchawan, S.; Buaboocha, T. Genome-wide association study for salinity tolerance at the flowering stage in a panel of rice accessions from Thailand. BMC Genom. 2019, 20, 76. [Google Scholar] [CrossRef] [PubMed]
- Naveed, S.A.; Zhang, F.; Zhang, J.; Zheng, T.-Q.; Meng, L.-J.; Pang, Y.-L.; Xu, J.-L.; Li, Z.-K. Identification of QTN and candidate genes for Salinity Tolerance at the Germination and Seedling Stages in Rice by Genome-Wide Association Analyses. Sci. Rep. 2018, 8, 1–11. [Google Scholar] [CrossRef] [PubMed]
- An, H.; Liu, K.; Wang, B.; Tian, Y.; Ge, Y.; Zhang, Y.; Tang, W.; Chen, G.; Yu, J.; Wu, W.; et al. Genome-wide association study identifies QTLs conferring salt tolerance in rice. Plant Breed. 2020, 139, 73–82. [Google Scholar] [CrossRef]
- Wang, Z.; Wang, J.; Bao, Y.; Wu, Y.; Zhang, H. Quantitative trait loci controlling rice seed germination under salt stress. Euphytica 2010, 178, 297–307. [Google Scholar] [CrossRef]
- Tian, L.; Tan, L.; Liu, F.; Cai, H.; Sun, C. Identification of quantitative trait loci associated with salt tolerance at seedling stage from Oryza rufipogon. J. Genet. Genom. 2011, 38, 593–601. [Google Scholar] [CrossRef]
- Ouyang, S.; He, S.; Liu, P.; Zhang, W.; Zhang, J.; Chen, S. The role of tocopherol cyclase in salt stress tolerance of rice (Oryza sativa). Sci. China Life Sci. 2011, 54, 181–188. [Google Scholar] [CrossRef]
- Cao, H.; Guo, S.; Xu, Y.; Jiang, K.; Jones, A.M.; Chong, K. Reduced expression of a gene encoding a Golgi localized monosaccharide transporter (OsGMST1) confers hypersensitivity to salt in rice (Oryza sativa L.). J. Exp. Bot. 2011, 62, 4595–4604. [Google Scholar] [CrossRef] [PubMed]
- Luo, D.; Niu, X.; Yu, J.; Yan, J.; Gou, X.; Lu, B.-R.; Liu, Y. Rice choline monooxygenase (OsCMO) protein functions in enhancing glycine betaine biosynthesis in transgenic tobacco but does not accumulate in rice (Oryza sativa L. ssp. japonica). Plant Cell Rep. 2012, 31, 1625–1635. [Google Scholar] [CrossRef]
- Pundir, P.; Devi, A.; Krishnamurthy, S.L.; Sharma, P.C.; Vinaykumar, N.M. QTLs in salt rice variety CSR10 reveals salinity tolerance at reproductive stage. Acta Physiol. Plant. 2021, 43, 1–15. [Google Scholar] [CrossRef]
- Nayyeripasand, L.; Garoosi, G.A.; Ahmadikhah, A. Genome-wide association study (GWAS) to identify salt-tolerance QTLs carrying novel candidate genes in rice during early vegetative stage. Rice 2021, 14, 1–21. [Google Scholar] [CrossRef]
- Jahan, N.; Zhang, Y.; Lv, Y.; Song, M.; Zhao, C.; Hu, H.; Cui, Y.; Wang, Z.; Yang, S.; Zhang, A.; et al. QTL analysis for rice salinity tolerance and fine mapping of a candidate locus qSL7 for shoot length under salt stress. Plant Growth Regul. 2019, 90, 307–319. [Google Scholar] [CrossRef]
- Asif, M.A.; Schilling, R.K.; Tilbrook, J.; Brien, C.; Dowling, K.; Rabie, H.; Short, L.; Trittermann, C.; Garcia, A.; Barrett-Lennard, E.G.; et al. Mapping of novel salt tolerance QTL in an Excalibur × Kukri doubled haploid wheat population. Theor. Appl. Genet. 2018, 131, 2179–2196. [Google Scholar] [CrossRef]
- Ismail, A.M.; Heuer, S.; Thomson, M.J.; Wissuwa, M. Genetic and genomic approaches to develop rice germplasm for problem soils. Plant Mol. Biol. 2007, 65, 547–570. [Google Scholar] [CrossRef]
- Singh, R.K.; Flowers, T.J. The Physiology and Molecular Biology of the Effects of Salinity on Rice. In Handbook of Plant and Crop Stress, 3rd ed.; Pessarakli, M., Ed.; Taylor and Francis: Boca Raton, FL, USA, 2010; pp. 901–942. [Google Scholar]
- Akbar, M.; Yabuno, T.; Nakao, S. Breeding for Saline-resistant Varieties of Rice: I. Variabil-ity for Salt Tolerance among Some Rice Varietles. Jpn. J. Breed. 1972, 22, 277–284. [Google Scholar] [CrossRef]
- Flowers, T.J.; Yeo, A.R. Variability in the resistance of sodium chloride salinity within rice (Oryza sativa L.) varieties. New Phytol. 1981, 88, 363–373. [Google Scholar] [CrossRef]
- Misra, A.N.; Sahu, S.; Misra, M.; Singh, P.; Meera, I.; Das, N.; Kar, M.; Sahu, P. Sodium chloride induced changes in leaf growth, and pigment and protein contents in two rice cultivars. Biol. Plant. 1997, 39, 257–262. [Google Scholar] [CrossRef]
- An, P.; Inanaga, S.; Li, X.J.; Eneji, A.E.; Zhu, N.W. Interactive Effects of Salinity and Air Humidity on Two Tomato Cultivars Differing in Salt Tolerance. J. Plant Nutr. 2005, 28, 459–473. [Google Scholar] [CrossRef]
- Mazher, A.M.A.; El-Quesni, E.M.F.; Farahat, M.M. Responses of ornamental and woody trees to salinity. World J. Agric. Sci. 2007, 3, 386–395. [Google Scholar]
- Munns, R. Comparative physiology of salt and water stress. Plant Cell Environ. 2002, 25, 239–250. [Google Scholar] [CrossRef]
- Gale, J.; Zeroni, M. Cultivation of Plants in Brackish Water in Controlled Environment Agriculture. 1984. Available online: https://agris.fao.org/agris-search/search.do?recordID=US19860060131 (accessed on 12 February 2021).
- Ren, Z.H.; Gao, J.P.; Li, L.G.; Cai, X.L.; Huang, W.; Chao, D.Y.; Zhu, M.Z.; Wang, Z.Y.; Luan, S.; Lin, H.X. A rice quantitative trait locus for salt tolerance encodes a sodium trans-porter. Nat. Genet. 2005, 37, 1141–1146. [Google Scholar] [CrossRef]
- Pushparajan, N.; Krishnasamy, V.; Babu, R.C.; Kannanbabu, J.R. Association mapping of salinity tolerance in rice using molecular markers. Int. J. Biol. Stress Manag. 2011, 2, 307–312. [Google Scholar]
- Wang, Z.; Chen, Z.; Cheng, J.; Lai, Y.; Wang, J.; Bao, Y.; Huang, J.; Zhang, H. QTL Analysis of Na+ and K+ Concentrations in Roots and Shoots under Different Levels of NaCl Stress in Rice (Oryza sativa L.). PLoS ONE 2012, 7, e51202. [Google Scholar] [CrossRef]
- De Leon, T.B.; Linscombe, S.; Gregorio, G.; Subudhi, P.K. Genetic variation in Southern USA rice genotypes for seedling salinity tolerance. Front. Plant Sci. 2015, 6, 374. [Google Scholar] [CrossRef]
- Apse, M.P.; Aharon, G.S.; Blumwald, E. Salt Tolerance Conferred by Overexpression of a Vacuolar Na+/H+ Antiport in Arabidopsis. Science 1999, 285, 1256–1258. [Google Scholar] [CrossRef]
- Babu, N.N.; Vinod, K.K.; Krishnamurthy, S.L.; Krishnan, S.G.; Yadav, A.; Bhowmick, P.K.; Nagarajan, M.; Singh, N.K.; Prabhu, K.V.; Singh, A.K. Microsatellite based linkage disequilibrium analyses reveal Saltol haplotype fragmentation and identify novel QTLs for seedling stage salinity tolerance in rice (Oryza sativa L.). J. Plant Biochem. Biotechnol. 2016, 26, 310–320. [Google Scholar] [CrossRef]
- Zhao, K.; Tung, C.-W.; Eizenga, G.C.; Wright, M.H.; Ali, M.L.; Price, A.H.; Norton, G.J.; Islam, M.R.; Reynolds, A.R.; Mezey, J.G.; et al. Genome-wide association mapping reveals a rich genetic architecture of complex traits in Oryza sativa. Nat. Commun. 2011, 2, 467. [Google Scholar] [CrossRef]
- Zhang, P.; Liu, X.; Tong, H.; Lu, Y.; Li, J. Association Mapping for Important Agronomic Traits in Core Collection of Rice (Oryza sativa L.) with SSR Markers. PLoS ONE 2014, 9, e111508. [Google Scholar] [CrossRef]
- Ya-Fang, Z.; Yu-Yin, M.; Zong-Xiang, C.; Jie, Z.; Tian-Xiao, C.; Qian-Qian, L.; Xue-Biao, P.; Shi-Min, Z. Genome-Wide Association Studies Reveal New Genetic Targets for Five Panicle Traits of International Rice Varieties. Rice Sci. 2015, 22, 217–226. [Google Scholar] [CrossRef]
- Wang, Y.; Zheng, Y.; Cai, Q.; Liao, C.; Mao, X.; Xie, H.; Zhu, Y.; Lian, L.; Luo, X.; Xie, H.; et al. Population structure and association analysis of yield and grain quality traits in hybrid rice primal parental lines. Euphytica 2016, 212, 261–273. [Google Scholar] [CrossRef]
- Kaler, A.S.; Purcell, L.C. Estimation of a significance threshold for genome-wide association studies. BMC Genom. 2019, 20, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Wallace, J.G.; Zhang, X.; Beyene, Y.; Semagn, K.; Olsen, M.; Prasanna, B.M.; Buckler, E.S. Genome-wide Association for Plant Height and Flowering Time across 15 Tropical Maize Populations under Managed Drought Stress and Well-Watered Conditions in Sub-Saharan Africa. Crop. Sci. 2016, 56, 2365–2378. [Google Scholar] [CrossRef]
- Xu, Y.; Yang, T.; Zhou, Y.; Yin, S.; Li, P.; Liu, J.; Xu, S.; Yang, Z.; Xu, C. Genome-Wide Association Mapping of Starch Pasting Properties in Maize Using Single-Locus and Multi-Locus Models. Front. Plant Sci. 2018, 9, 1311. [Google Scholar] [CrossRef] [PubMed]
- Ward, B.P.; Brown-Guedira, G.; Kolb, F.L.; Van Sanford, D.A.; Tyagi, P.; Sneller, C.H.; Griffey, C.A. Genome-wide association studies for yield-related traits in soft red winter wheat grown in Virginia. PLoS ONE 2019, 14, e0208217. [Google Scholar] [CrossRef]
- Wei, W.; Mesquita, A.C.O.; Figueiró, A.D.A.; Wu, X.; Manjunatha, S.; Wickland, D.P.; Hudson, M.E.; Juliatti, F.C.; Clough, S.J. Genome-wide association mapping of resistance to a Brazilian isolate of Sclerotinia sclerotiorum in soybean genotypes mostly from Brazil. BMC Genom. 2017, 18, 849. [Google Scholar] [CrossRef] [PubMed]
- Jaiswal, V.; Bandyopadhyay, T.; Gahlaut, V.; Gupta, S.; Dhaka, A.; Ramchiary, N.; Prasad, M. Genome-wide association study (GWAS) delineates genomic loci for ten nutritional elements in foxtail millet (Setaria italica L.). J. Cereal Sci. 2019, 85, 48–55. [Google Scholar] [CrossRef]
- Zhou, Y.; Srinivasan, S.; Mirnezami, S.V.; Kusmec, A.; Fu, Q.; Attigala, L.; Fernandez, M.G.S.; Ganapathysubramanian, B.; Schnable, P.S. Semiautomated Feature Extraction from RGB Images for Sorghum Panicle Architecture GWAS. Plant Physiol. 2019, 179, 24–37. [Google Scholar] [CrossRef]
- Bollinedi, H.; Yadav, A.K.; Vinod, K.K.; Krishnan, S.G.; Bhowmick, P.K.; Nagarajan, M.; Neeraja, C.N.; Ellur, R.K.; Singh, A.K. Genome-Wide Association Study Reveals Novel Marker-Trait Associations (MTAs) Governing the Localization of Fe and Zn in the Rice Grain. Front. Genet. 2020, 11, 213. [Google Scholar] [CrossRef] [PubMed]
- Rohilla, M.; Singh, N.; Mazumder, A.; Sen, P.; Roy, P.; Chowdhury, D.; Singh, N.K.; Mondal, T.K. Genome-wide association studies using 50 K rice genic SNP chip unveil genetic architecture for anaerobic germination of deep-water rice population of Assam, India. Mol. Genet. Genom. 2020, 295, 1211–1226. [Google Scholar] [CrossRef] [PubMed]
- Biselli, C.; Volante, A.; Desiderio, F.; Tondelli, A.; Gianinetti, A.; Finocchiaro, F.; Taddei, F.; Gazza, L.; Sgrulletta, D.; Cattivelli, L.; et al. GWAS for Starch-Related Parameters in Japonica Rice (Oryza sativa L.). Plants 2019, 8, 292. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Lu, Q.; Zhai, R.; Zhang, M.; Xu, Q.; Yang, Y.; Wang, S.; Yuan, X.; Yu, H.; Wang, Y.; et al. Genome wide association mapping for grain shape traits in indica rice. Planta 2016, 244, 819–830. [Google Scholar] [CrossRef]
- Kim, S.-M.; Reinke, R.F. A novel resistance gene for bacterial blight in rice, Xa43(t) identified by GWAS, confirmed by QTL mapping using a bi-parental population. PLoS ONE 2019, 14, e0211775. [Google Scholar] [CrossRef]
- Gollan, P.J.; Bhave, M. Genome-wide analysis of genes encoding FK506-binding proteins in rice. Plant Mol. Biol. 2009, 72, 1–16. [Google Scholar] [CrossRef]
- Jain, M.; Nijhawan, A.; Arora, R.; Agarwal, P.; Ray, S.; Sharma, P.; Kapoor, S.; Tyagi, A.K.; Khurana, J.P. F-box proteins in rice. Genome-wide analysis, classification, tem-poral and spatial gene expression during panicle and seed development, and regulation by light and abiotic stress. Plant Physiol. 2007, 143, 1467–1483. [Google Scholar] [CrossRef]
- Zhao, Y.; Cai, M.; Zhang, X.; Li, Y.; Zhang, J.; Zhao, H.; Kong, F.; Zheng, Y.; Qiu, F. Genome-Wide Identification, Evolution and Expression Analysis of mTERF Gene Family in Maize. PLoS ONE 2014, 9, e94126. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, S. Routine Procedure for Growing Rice Plants in Culture Solution; Laboratory Manual for Physiological Studies of Rice, International Rice Research Institute: Manila, Philippines, 1997; p. 38. [Google Scholar]
- IRRI. Standard Evaluation System (SES) for Rice, 3rd ed.; International Rice Research Institute: Manila, Philippines, 2013. [Google Scholar]
- Johnson, S.C. Hierarchical clustering schemes. Psychometrika 1967, 32, 241–254. [Google Scholar] [CrossRef] [PubMed]
- Ward, J.H., Jr. Hierarchical grouping to optimize an objective function. J. Am. Stat. Assoc. 1963, 58, 236–244. [Google Scholar] [CrossRef]
- Doyle, J.J.; Doyle, J.L. Isolation of plant DNA from fresh tissue. Focus 1990, 12, 39–40. [Google Scholar]
- Singh, N.; Jayaswal, P.K.; Panda, K.; Mandal, P.; Kumar, V.; Singh, B.; Mishra, S.; Singh, Y.; Singh, R.; Rai, V.; et al. Single-copy gene based 50 K SNP chip for genetic studies and molecular breeding in rice. Sci. Rep. 2015, 5, 11600. [Google Scholar] [CrossRef]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multi-locus genotype data. Genetics 2000, 155, 945–959. [Google Scholar] [CrossRef]
- Alexander, D.H.; Novembre, J.; Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 2009, 19, 1655–1664. [Google Scholar] [CrossRef]
- Earl, D.A.; Vonholdt, B.M. STRUCTURE HARVESTER: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 2011, 4, 359–361. [Google Scholar] [CrossRef]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software structure: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.L. Development of an Iterative Usage of Fixed Effect and Random Effect Models for Powerful and Efficient Genome-Wide Association Studies. Master’s Thesis, Huazhong Agricultural University, Wuhan, China, 2015. [Google Scholar]
- Liu, X.; Huang, M.; Fan, B.; Buckler, E.S.; Zhang, Z. Iterative Usage of Fixed and Random Effect Models for Powerful and Efficient Genome-Wide Association Studies. PLoS Genet. 2016, 12, e1005767. [Google Scholar] [CrossRef] [PubMed]
| S.No | Traits | MTAs | SNP | Chr. | Position | p Value | R2 | Previous Report |
|---|---|---|---|---|---|---|---|---|
| 1 | SL | qSL2 | AX-95920196 | 2 | 23947447 | 6.74 × 10−5 | 15.11 | qPH2 [24]; qGP-2 [41] |
| 2 | RL | qRL2 | AX-95920196 | 2 | 23947447 | 7.54 × 10−5 | 19.26 | qPH2 [24]; qGP-2 [41] |
| 3 | SFW | qSFW2 | AX-95921620 | 2 | 23533590 | 5.18 × 10−4 | 14.64 | qPH2, qRKC2, qCHL2 [24]; qGP-2 [41] |
| qSFW7 | AX-95937657 | 7 | 20788892 | 6.79 × 10−4 | 13.98 | qRSW7 [42] | ||
| qSFW9 | AX-95931839 | 9 | 16483542 | 4.61 × 10−4 | 14.92 | qSNK9, qRNK9, qSES9 [24]; qGP-9 [41] | ||
| 4 | RFW | qRFW2 | AX-95921620 | 2 | 23533590 | 5.18 × 10−4 | 14.64 | qPH2, qRKC2, qCHL2 [24]; qGP-2 [41] |
| 5 | SEW | qSEW2 | AX-95921620 | 2 | 23533590 | 3.87 × 10−4 | 15.43 | qPH2, qRKC2, qCHL2 [24]; qGP-2 [41] |
| qSEW7 | AX-95937657 | 7 | 20788892 | 4.07 × 10−4 | 15.31 | qRSW7 [42] | ||
| qSEW9 | AX-95931839 | 9 | 16483542 | 5.06 × 10−4 | 14.77 | qSNK9, qRNK9 [24]; qGP-9 [41] | ||
| 6 | SDW | qSDW2.1 | AX-95920663 | 2 | 5664763 | 1.42 × 10−5 | 25.51 | - |
| qSDW2.2 | AX-95934798 | 2 | 10213902 | 4.05 × 10−5 | 22.52 | OsVTE1 [43]; OsGMST1 [44] | ||
| qSDW12.1 | AX-95939149 | 12 | 17404747 | 9.22 × 10−5 | 20.24 | qSES12, qSUR12, qCHL12 [24] | ||
| 7 | RDW | qRDW6 | AX-95956901 | 6 | 29729562 | 5.79 × 10−5 | 22.05 | OsCMO [45] |
| qRDW7 | AX-95929366 | 7 | 20782724 | 4.74 × 10−5 | 22.62 | qSES7.1, KR7.1 [46] | ||
| 8 | RNC | qRNC2.1 | AX-95921298 | 2 | 23260124 | 6.30 × 10−5 | 18.17 | qRKC2 [24]; qGP-2 [41] |
| qRNC2.2 | AX-95920628 | 2 | 22172032 | 9.45 × 10−5 | 15.15 | qRKC2 [24]; qGP-2 [41] | ||
| 9 | SNC | qSNC1 | AX-95940587 | 1 | 13758487 | 8.16 × 10−5 | 16.26 | qSKC, qSNK, qRNK [24] |
| qSNC2 | AX-95920537 | 2 | 22171212 | 8.35 × 10−5 | 16.64 | qRKC2 [24]; qGP-2 [41] | ||
| qSNC5 | AX-95927105 | 5 | 19697164 | 9.39 × 10−5 | 18.08 | - | ||
| 10 | SKC | qSKC1 | AX-95940587 | 1 | 13758487 | 7.86 × 10−5 | 16.26 | qSKC, qSNK, qRNK [24] |
| qSKC6 | AX-95937335 | 6 | 3429601 | 8.21 × 10−5 | 14.33 | qRFWn6.1, qRDWn6.1 [47] | ||
| 11 | RNK | qRNK1 | AX-95918556 | 1 | 11022718 | 6.26 × 10−6 | 25.11 | qRNK1, qSNC, qSKC, qRKC [24] |
| 12 | SNK | qSNK1 | AX-95940642 | 1 | 13322813 | 2.93 × 10−6 | 29.88 | qSKC, qSNK, qRNK [24] |
| Chr. | MSU-RAP ID | Position (bp) | Description or Putative Function |
|---|---|---|---|
| 2 | LOC_Os02g10580 | 5,565,956 | NB-ARC domain containing disease resistance protein |
| LOC_Os02g10590 | 5,569,801 | Peptidyl-prolyl cis-trans isomerase, FKBP-type | |
| LOC_Os02g10600 | 5,573,182 | OsFBA1—F-box and FBA domain containing protein | |
| LOC_Os02g10630 | 5,593,645 | GRAM and C2 domains containing protein | |
| LOC_Os02g10640 | 5,600,889 | 26S protease regulatory subunit, | |
| LOC_Os02g10650 | 5,604,236 | CRAL/TRIO domain containing protein | |
| LOC_Os02g10660 | 5,614,461 | Gycosyl hydrolases family 17 | |
| LOC_Os02g10690 | 5,623,352 | Targeting protein for Xklp2 | |
| LOC_Os02g10700 | 5,631,315 | OsFBL7—F-box domain and LRR containing protein | |
| LOC_Os02g10710 | 5,640,360 | hsp20/alpha crystallin family protein | |
| LOC_Os02g10750 | 5,672,334 | CBL-interacting protein kinase | |
| LOC_Os02g10760 | 5,688,698 | AP2 domain containing protein | |
| LOC_Os02g10770 | 5,697,834 | DEAD-box ATP-dependent RNA helicase 41 | |
| LOC_Os02g10780 | 5,706,636 | SPX domain containing protein | |
| LOC_Os02g10800 | 5,736,606 | Mitochondrial carrier protein | |
| LOC_Os02g10810 | 5,742,520 | Protein of unknown function domain containing protein | |
| LOC_Os02g10820 | 5,748,141 | Sel1 repeat domain containing protein | |
| LOC_Os02g10830 | 5,749,500 | Serine acetyltransferase protein | |
| 5 | LOC_Os05g33500 | 19,678,142 | mTERF domain containing protein |
| LOC_Os05g33510 | 19,681,732 | Peptide methionine sulfoxide reductase msrB | |
| LOC_Os05g33550 | 19,704,966 | Methyl-binding domain protein MBD | |
| LOC_Os05g33554 | 19,707,432 | Methyl-binding domain protein MBD | |
| LOC_Os05g33570 | 19,737,857 | Pyruvate, phosphate dikinase, chloroplast precursor | |
| LOC_Os05g33590 | 19,744,851 | Cytochrome P450, putative, expressed | |
| LOC_Os05g33600 | 19,758,913 | Cytochrome P450 72A1, putative, expressed | |
| LOC_Os05g33630 | 19,785,962 | Inosine-uridine preferring nucleoside hydrolase family protein | |
| LOC_Os05g33644 | 19,805,637 | Inosine-uridine preferring nucleoside hydrolase family protein | |
| LOC_Os05g33690 | 19,828,422 | Receptor-like protein kinase precursor | |
| LOC_Os05g33700 | 19,834,008 | 4F5 protein family protein | |
| LOC_Os05g33710 | 19,846,592 | WD domain, G-beta repeat domain containing protein | |
| LOC_Os05g33730 | 19,868,419 | Gibberellin receptor GID1L2 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yadav, A.K.; Kumar, A.; Grover, N.; Ellur, R.K.; Bollinedi, H.; Krishnan, S.G.; Bhowmick, P.K.; Vinod, K.K.; Nagarajan, M.; Singh, A.K. Genome-Wide Association Study Reveals Marker–Trait Associations for Early Vegetative Stage Salinity Tolerance in Rice. Plants 2021, 10, 559. https://doi.org/10.3390/plants10030559
Yadav AK, Kumar A, Grover N, Ellur RK, Bollinedi H, Krishnan SG, Bhowmick PK, Vinod KK, Nagarajan M, Singh AK. Genome-Wide Association Study Reveals Marker–Trait Associations for Early Vegetative Stage Salinity Tolerance in Rice. Plants. 2021; 10(3):559. https://doi.org/10.3390/plants10030559
Chicago/Turabian StyleYadav, Ashutosh Kumar, Aruna Kumar, Nitasha Grover, Ranjith Kumar Ellur, Haritha Bollinedi, Subbaiyan Gopala Krishnan, Prolay Kumar Bhowmick, Kunnummal Kurungara Vinod, Mariappan Nagarajan, and Ashok Kumar Singh. 2021. "Genome-Wide Association Study Reveals Marker–Trait Associations for Early Vegetative Stage Salinity Tolerance in Rice" Plants 10, no. 3: 559. https://doi.org/10.3390/plants10030559
APA StyleYadav, A. K., Kumar, A., Grover, N., Ellur, R. K., Bollinedi, H., Krishnan, S. G., Bhowmick, P. K., Vinod, K. K., Nagarajan, M., & Singh, A. K. (2021). Genome-Wide Association Study Reveals Marker–Trait Associations for Early Vegetative Stage Salinity Tolerance in Rice. Plants, 10(3), 559. https://doi.org/10.3390/plants10030559








