Space—Time Surveillance of COVID-19 Seasonal Clusters: A Case of Sweden
Abstract
:1. Introduction
2. Materials and Methods
2.1. Data
2.1.1. COVID-19 Data
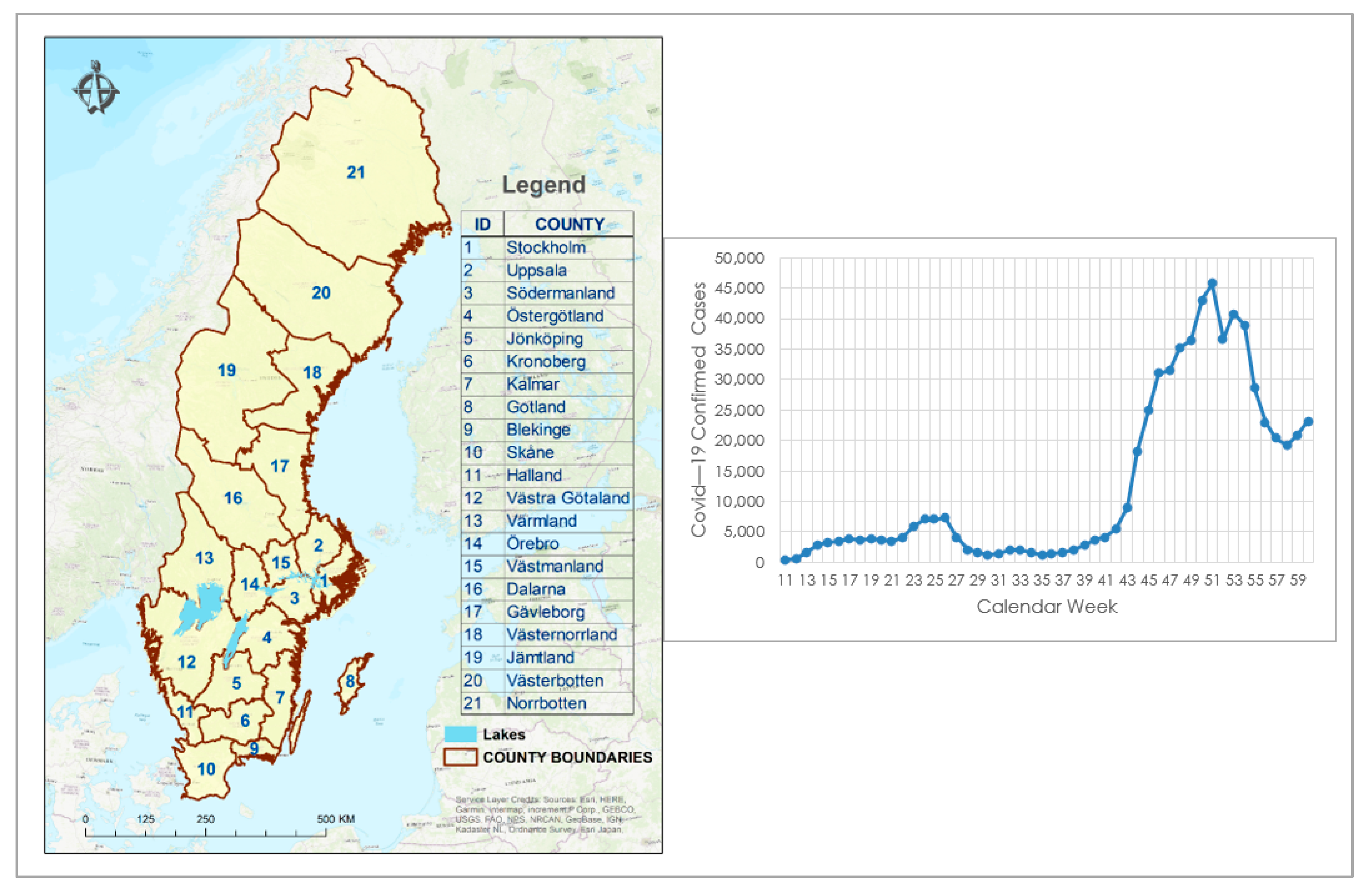
2.1.2. Municipality Boundaries
2.2. Methods
Retrospective and Prospective Space–Time Permutation
3. Results
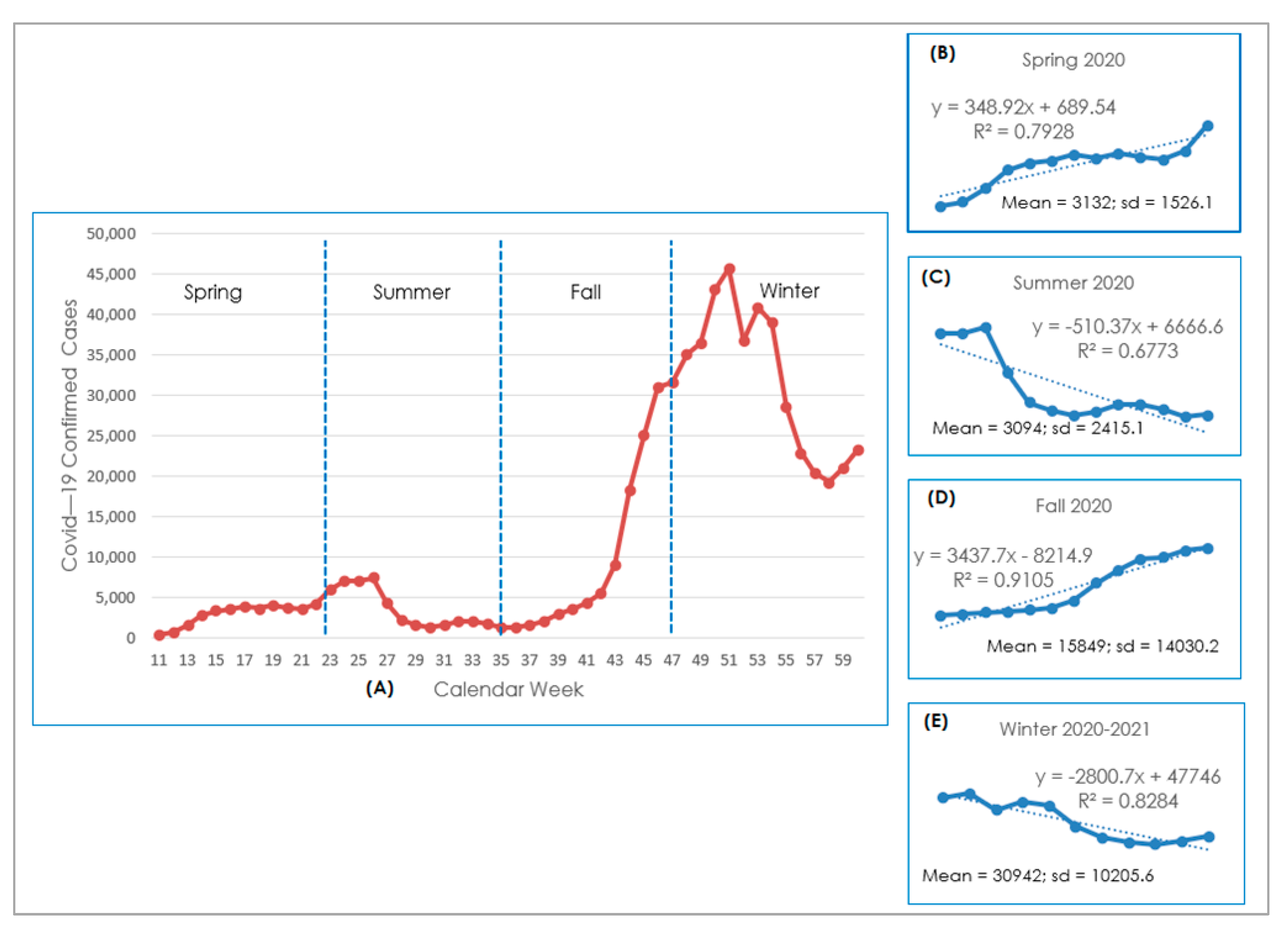
3.1. COVID-19 Reported Cases Profile
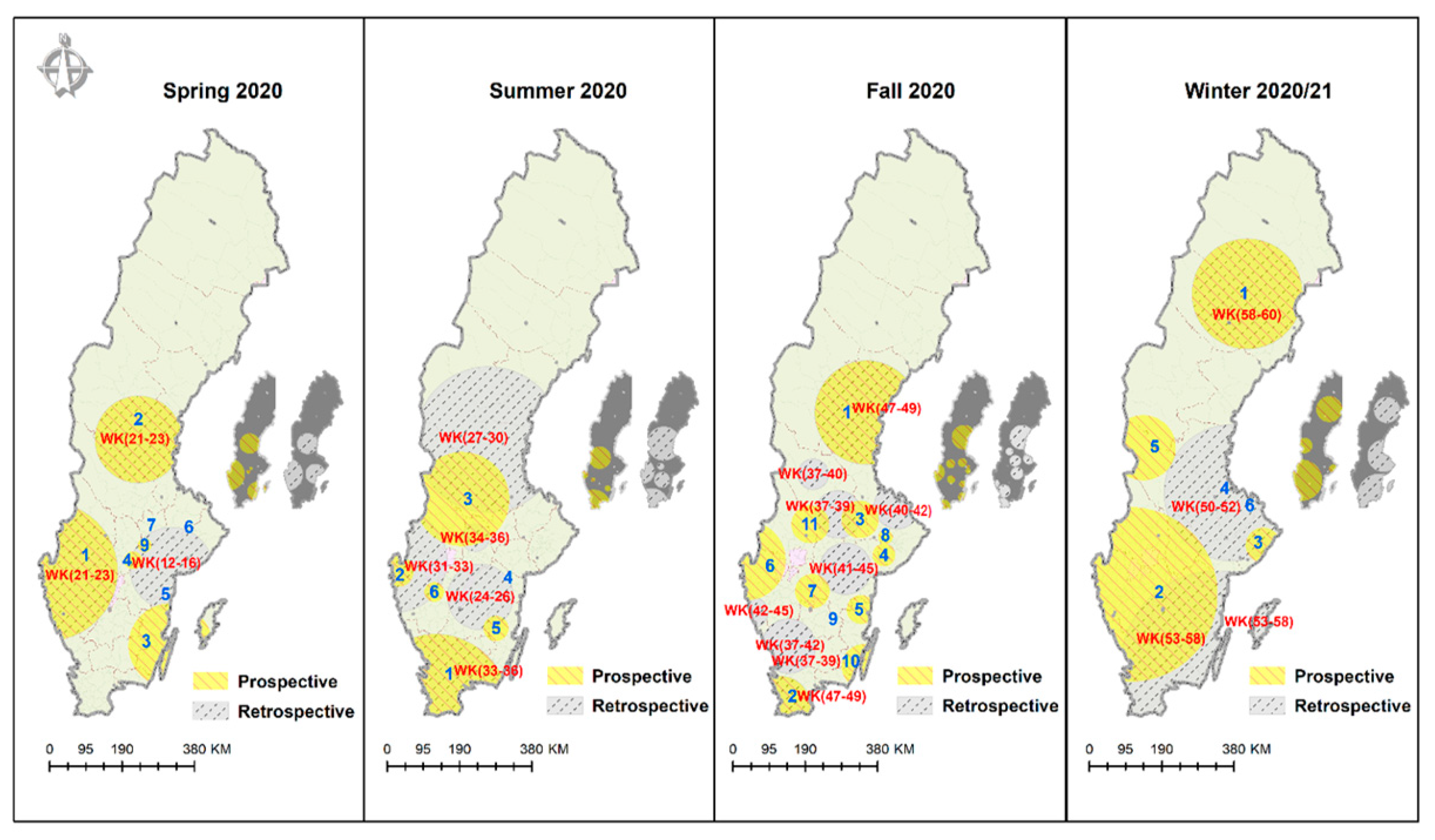
3.2. Municipality-Level Results—Spring 2020 (Week 11–Week 23)
3.3. Municipality-Level Results—Summer 2020 (Week 24–Week 36)
3.4. Municipality-Level Results—Fall 2020 (Week 37–Week 49)
3.5. Municipality-Level Results—Winter 2020/21 (Week 50–Week 60)
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Li, Q.; Guan, X.; Wu, P.; Wang, X.; Zhou, L.; Tong, Y.; Ren, R.; Leung, K.S.; Lau, E.H.; Wong, J.Y. Early transmission dynamics in Wuhan, China, of novel coronavirus–infected pneumonia. N. Engl. J. Med. 2020, 382, 1199–1207. [Google Scholar] [CrossRef] [PubMed]
- Spinelli, A.; Pellino, G. COVID-19 pandemic: Perspectives on an unfolding crisis. J. Br. Surg. 2020, 107, 785–787. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fauci, A.S.; Lane, H.C.; Redfield, R.R. COVID-19—Navigating the uncharted. Mass Med. Soc. 2020, 382, 1268–1269. [Google Scholar] [CrossRef] [PubMed]
- Verity, R.; Okell, L.C.; Dorigatti, I.; Winskill, P.; Whittaker, C.; Imai, N.; Cuomo-Dannenburg, G.; Thompson, H.; Walker, P.G.; Fu, H. Estimates of the severity of coronavirus disease 2019: A model-based analysis. Lancet Infect. Dis. 2020, 20, 669–677. [Google Scholar] [CrossRef]
- Worldometer. COVID-19 Corona Virus Pandemic. Available online: https://www.worldometers.info/coronavirus/ (accessed on 22 April 2022).
- WHO. WHO Coronavirus (COVID-19) Dashboard. Available online: https://covid19.who.int/ (accessed on 30 March 2021).
- Njenga, M.K.; Dawa, J.; Nanyingi, M.; Gachohi, J.; Ngere, I.; Letko, M.; Otieno, C.; Gunn, B.M.; Osoro, E. Why is there low morbidity and mortality of COVID-19 in Africa? Am. J. Trop. Med. Hyg. 2020, 103, 564–569. [Google Scholar] [CrossRef]
- Lalaoui, R.; Bakour, S.; Raoult, D.; Verger, P.; Sokhna, C.; Devaux, C.; Pradines, B.; Rolain, J.-M. What could explain the late emergence of COVID-19 in Africa? New Microbes New Infect. 2020, 38, 100760. [Google Scholar] [CrossRef]
- Mariam, S.H. The Severe Acute Respiratory Syndrome Coronavirus-2 (SARS-CoV-2) Pandemic: Are Africa’s Prevalence and Mortality Rates Relatively Low? Adv. Virol. 2022, 2022, 3387784. [Google Scholar] [CrossRef]
- Shephard, R.J. Immune changes induced by exercise in an adverse environment. Can. J. Physiol. Pharmacol. 1998, 76, 539–546. [Google Scholar] [CrossRef]
- Adedokun, K.A.; Olarinmoye, A.O.; Mustapha, J.O.; Kamorudeen, R.T. A close look at the biology of SARS-CoV-2, and the potential influence of weather conditions and seasons on COVID-19 case spread. Infect. Dis. Poverty 2020, 9, 77. [Google Scholar] [CrossRef]
- Choi, Y.W.; Tuel, A.; Eltahir, E.A. On the environmental determinants of COVID-19 seasonality. Geohealth 2021, 5, e2021GH000413. [Google Scholar] [CrossRef]
- Zoran, M.A.; Savastru, R.S.; Savastru, D.M.; Tautan, M.N.; Baschir, L.A.; Tenciu, D.V. Exploring the linkage between seasonality of environmental factors and COVID-19 waves in Madrid, Spain. Process Saf. Environ. Prot. 2021, 152, 583–600. [Google Scholar] [CrossRef]
- Byun, W.S.; Heo, S.W.; Jo, G.; Kim, J.W.; Kim, S.; Lee, S.; Park, H.E.; Baek, J.-H. Is coronavirus disease (COVID-19) seasonal? A critical analysis of empirical and epidemiological studies at global and local scales. Environ. Res. 2021, 196, 110972. [Google Scholar] [CrossRef] [PubMed]
- Socialstyrelsen. Statistics on COVID-19. Available online: https://www.socialstyrelsen.se/en/statistics-and-data/statistics/statistics-on-covid-19/ (accessed on 30 March 2021).
- Emborg, H.-D.; Carnahan, A.; Bragstad, K.; Trebbien, R.; Brytting, M.; Hungnes, O.; Byström, E.; Vestergaard, L.S. Abrupt termination of the 2019/20 influenza season following preventive measures against COVID-19 in Denmark, Norway and Sweden. Eurosurveillance 2021, 26, 2001160. [Google Scholar] [CrossRef] [PubMed]
- Panneer, S.; Kantamaneni, K.; Akkayasamy, V.S.; Susairaj, A.X.; Panda, P.K.; Acharya, S.S.; Rice, L.; Liyanage, C.; Pushparaj, R.R.B. The Great Lockdown in the Wake of COVID-19 and Its Implications: Lessons for Low and Middle-Income Countries. Int. J. Environ. Res. Public Health 2022, 19, 610. [Google Scholar] [CrossRef] [PubMed]
- Drefahl, S.; Wallace, M.; Mussino, E.; Aradhya, S.; Kolk, M.; Brandén, M.; Malmberg, B.; Andersson, G. A population-based cohort study of socio-demographic risk factors for COVID-19 deaths in Sweden. Nat. Commun. 2020, 11, 5097. [Google Scholar] [CrossRef]
- Gémes, K.; Talbäck, M.; Modig, K.; Ahlbom, A.; Berglund, A.; Feychting, M.; Matthews, A.A. Burden and prevalence of prognostic factors for severe COVID-19 in Sweden. Eur. J. Epidemiol. 2020, 35, 401–409. [Google Scholar] [CrossRef]
- Openshaw, S.; Charlton, M.; Craft, A.W.; Birch, J. Investigation of leukaemia clusters by use of a geographical analysis machine. Lancet 1988, 331, 272–273. [Google Scholar] [CrossRef]
- Pullan, R.L.; Sturrock, H.J.; Magalhaes, R.J.S.; Clements, A.C.; Brooker, S.J. Spatial parasite ecology and epidemiology: A review of methods and applications. Parasitology 2012, 139, 1870–1887. [Google Scholar] [CrossRef] [Green Version]
- Kulldorff, M.; Heffernan, R.; Hartman, J.; Assunçao, R.; Mostashari, F. A space–time permutation scan statistic for disease outbreak detection. PLoS Med. 2005, 2, e59. [Google Scholar] [CrossRef] [Green Version]
- Jones, R.C.; Liberatore, M.; Fernandez, J.R.; Gerber, S.I. Use of a prospective space-time scan statistic to prioritize shigellosis case investigations in an urban jurisdiction. Public Health Rep. 2006, 121, 133–139. [Google Scholar] [CrossRef]
- Glatman-Freedman, A.; Kaufman, Z.; Kopel, E.; Bassal, R.; Taran, D.; Valinsky, L.; Agmon, V.; Shpriz, M.; Cohen, D.; Anis, E. Near real-time space-time cluster analysis for detection of enteric disease outbreaks in a community setting. J. Infect. 2016, 73, 99–106. [Google Scholar] [CrossRef] [PubMed]
- Kulldorff, M. Prospective time periodic geographical disease surveillance using a scan statistic. J. R. Stat. Soc. Ser. A (Stat. Soc.) 2001, 164, 61–72. [Google Scholar] [CrossRef]
- Yin, F.; Li, X.; Ma, J.; Feng, Z. The early warning system based on the prospective space-time permutation statistic. Wei Sheng Yan Jiu J. Hyg. Res. 2007, 36, 455–458. [Google Scholar]
- Tang, X.; Geater, A.; McNeil, E.; Deng, Q.; Dong, A.; Zhong, G. Spatial, temporal and spatio-temporal clusters of measles incidence at the county level in Guangxi, China during 2004–2014: Flexibly shaped scan statistics. BMC Infect. Dis. 2017, 17, 243. [Google Scholar] [CrossRef] [Green Version]
- Hohl, A.; Delmelle, E.M.; Desjardins, M.R.; Lan, Y. Daily surveillance of COVID-19 using the prospective space-time scan statistic in the United States. Spat. Spatio-Temporal Epidemiol. 2020, 34, 100354. [Google Scholar] [CrossRef]
- Desjardins, M.; Hohl, A.; Delmelle, E. Rapid surveillance of COVID-19 in the United States using a prospective space-time scan statistic: Detecting and evaluating emerging clusters. Appl. Geogr. 2020, 118, 102202. [Google Scholar] [CrossRef]
- Gomes, D.; Andrade, L.; Ribeiro, C.; Peixoto, M.; Lima, S.; Duque, A.; Cirilo, T.; Góes, M.; Lima, A.; Santos, M. Risk clusters of COVID-19 transmission in northeastern Brazil: Prospective space–time modelling. Epidemiol. Infect. 2020, 148, E188. [Google Scholar] [CrossRef]
- Masrur, A.; Yu, M.; Luo, W.; Dewan, A. Space-time patterns, change, and propagation of COVID-19 risk relative to the intervention scenarios in Bangladesh. Int. J. Environ. Res. Public Health 2020, 17, 5911. [Google Scholar] [CrossRef]
- Folkhälsomyndigheten. Confirmed Cases of COVID-19 in Sweden. Available online: https://www.folkhalsomyndigheten.se/smittskydd-beredskap/utbrott/aktuella-utbrott/covid-19/statistik-och-analyser/bekraftade-fall-i-sverige/ (accessed on 30 March 2021).
- Statistics Sweden. Open Data for DeSO—Demographic Statistical Areas. Available online: https://www.scb.se/en/services/open-data-api/open-geodata/deso--demographic-statistical-areas/ (accessed on 30 March 2021).
- Sulyok, M.; Walker, M.D. Mobility and COVID-19 mortality across Scandinavia: A modeling study. Travel Med. Infect. Dis. 2021, 41, 102039. [Google Scholar] [CrossRef]
- Modig, K.; Ahlbom, A.; Ebeling, M. Excess mortality from COVID-19: Weekly excess death rates by age and sex for Sweden and its most affected region. Eur. J. Public Health 2021, 31, 17–22. [Google Scholar] [CrossRef]
- Hansson, E.; Albin, M.; Rasmussen, M.; Jakobsson, K. Stora skillnader i överdödlighet våren 2020 utifrån födelseland (English: Large differences in excess mortality in the spring of 2020 based on country of birth). Läkartidningen 2020, 2920, 20113. [Google Scholar]
- Le, T.H.; Tran, T.P.T. Alert for COVID-19 second wave: A lesson from Vietnam. J. Glob. Health 2021, 11, 03012. [Google Scholar] [CrossRef] [PubMed]
- Dellicour, S.; Linard, C.; Van Goethem, N.; Da Re, D.; Artois, J.; Bihin, J.; Schaus, P.; Massonnet, F.; Van Oyen, H.; Vanwambeke, S.O. Investigating the drivers of the spatio-temporal heterogeneity in COVID-19 hospital incidence—Belgium as a study case. Int. J. Health Geogr. 2021, 20, 29. [Google Scholar] [CrossRef] [PubMed]
- Venter, Z.S.; Barton, D.N.; Gundersen, V.; Figari, H.; Nowell, M.S. Back to nature: Norwegians sustain increased recreational use of urban green space months after the COVID-19 outbreak. Landsc. Urban Plan. 2021, 214, 104175. [Google Scholar] [CrossRef]
- Huang, Z. Spatiotemporal Evolution Patterns of the COVID-19 Pandemic Using Space-Time Aggregation and Spatial Statistics: A Global Perspective. ISPRS Int. J. Geo-Inf. 2021, 10, 519. [Google Scholar] [CrossRef]
| Cluster | Duration | p | t-Statistic | Observed (O) | Expected (E) | O/E | # Muni | Diameter (KM) | |
|---|---|---|---|---|---|---|---|---|---|
| 1 | weeks 21–23 | <0.001 | 323.65 | 4170 | 2785.46 | 1.50 | 57 | 181.65 | SPRING 2020 |
| 2 | weeks 21–23 | <0.001 | 107.15 | 713 | 391.60 | 1.82 | 12 | 113.67 | |
| 3 | weeks 21–23 | <0.001 | 43.57 | 342 | 197.46 | 1.73 | 14 | 109.86 | |
| 4 | weeks 19–23 | <0.001 | 13.78 | 192 | 128.23 | 1.50 | 3 | 23.68 | |
| 5 | weeks 20–23 | <0.001 | 10.57 | 45 | 20.77 | 2.17 | 1 | 0.00 | |
| 6 | weeks 21–23 | <0.001 | 8.17 | 43 | 21.64 | 1.99 | 1 | 0.00 | |
| 7 | weeks 21–23 | <0.001 | 7.79 | 18 | 5.99 | 3.00 | 1 | 0.00 | |
| 8 | weeks 19–23 | 0.004 | 6.74 | 39 | 20.33 | 1.92 | 1 | 0.00 | |
| 9 | weeks 20–23 | 0.008 | 6.23 | 52 | 30.52 | 1.70 | 2 | 18.31 | |
| 1 | weeks 33–36 | <0.001 | 395.94 | 1616 | 740.03 | 2.18 | 45 | 190.81 | SUMMER 2020 |
| 2 | weeks 31–36 | <0.001 | 76.63 | 235 | 92.65 | 2.54 | 7 | 41.29 | |
| 3 | weeks 34–36 | <0.001 | 43.34 | 288 | 157.72 | 1.83 | 29 | 123.54 | |
| 4 | weeks 33–36 | <0.001 | 19.85 | 110 | 56.47 | 1.95 | 1 | 0.00 | |
| 5 | weeks 34–36 | <0.001 | 12.58 | 26 | 8.03 | 3.24 | 3 | 32.81 | |
| 6 | weeks 31–36 | 0.007 | 9.19 | 73 | 42.24 | 1.73 | 3 | 25.90 | |
| 1 | weeks 47–49 | <0.001 | 308.36 | 4978 | 3438.12 | 1.45 | 13 | 135.63 | FALL 2020 |
| 2 | weeks 47–49 | <0.001 | 168.98 | 14,072 | 12,063.93 | 1.17 | 22 | 77.46 | |
| 3 | weeks 47–49 | <0.001 | 51.7 | 4414 | 3778.33 | 1.17 | 10 | 49.24 | |
| 4 | weeks 47–49 | <0.001 | 39.97 | 6266 | 5593.95 | 1.12 | 8 | 30.01 | |
| 5 | weeks 47–49 | <0.001 | 34.92 | 567 | 390.74 | 1.45 | 2 | 37.89 | |
| 6 | weeks 47–49 | <0.001 | 30.18 | 2334 | 1980.22 | 1.18 | 17 | 104.62 | |
| 7 | weeks 45–49 | <0.001 | 14.32 | 2860 | 2585.05 | 1.11 | 12 | 45.29 | |
| 8 | weeks 46–49 | <0.001 | 12.29 | 4637 | 4310.98 | 1.08 | 6 | 19.11 | |
| 9 | weeks 47–49 | <0.001 | 11.11 | 289 | 216.13 | 1.34 | 1 | 0 | |
| 10 | weeks 47–49 | <0.001 | 9.07 | 1146 | 1008.12 | 1.14 | 6 | 58.28 | |
| 11 | weeks 47–49 | <0.001 | 5.95 | 463 | 392.74 | 1.18 | 8 | 49.86 | |
| 1 | weeks 58–60 | <0.001 | 894.56 | 2650 | 1025.94 | 2.58 | 14 | 143.97 | WINTER 2020/21 |
| 2 | weeks 56–60 | <0.001 | 444.40 | 47,387 | 41,581.47 | 1.14 | 128 | 228.23 | |
| 3 | weeks 58–60 | <0.001 | 36.40 | 9011 | 8234.66 | 1.09 | 12 | 45.92 | |
| 4 | weeks 58–60 | <0.001 | 30.27 | 1048 | 816.17 | 1.28 | 1 | 0 | |
| 5 | weeks 57–60 | <0.001 | 29.71 | 289 | 176.99 | 1.63 | 3 | 85.52 | |
| 6 | weeks 58–60 | <0.001 | 15.05 | 170 | 108.14 | 1.57 | 1 | 0 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Aturinde, A.; Mansourian, A. Space—Time Surveillance of COVID-19 Seasonal Clusters: A Case of Sweden. ISPRS Int. J. Geo-Inf. 2022, 11, 307. https://doi.org/10.3390/ijgi11050307
Aturinde A, Mansourian A. Space—Time Surveillance of COVID-19 Seasonal Clusters: A Case of Sweden. ISPRS International Journal of Geo-Information. 2022; 11(5):307. https://doi.org/10.3390/ijgi11050307
Chicago/Turabian StyleAturinde, Augustus, and Ali Mansourian. 2022. "Space—Time Surveillance of COVID-19 Seasonal Clusters: A Case of Sweden" ISPRS International Journal of Geo-Information 11, no. 5: 307. https://doi.org/10.3390/ijgi11050307
APA StyleAturinde, A., & Mansourian, A. (2022). Space—Time Surveillance of COVID-19 Seasonal Clusters: A Case of Sweden. ISPRS International Journal of Geo-Information, 11(5), 307. https://doi.org/10.3390/ijgi11050307







