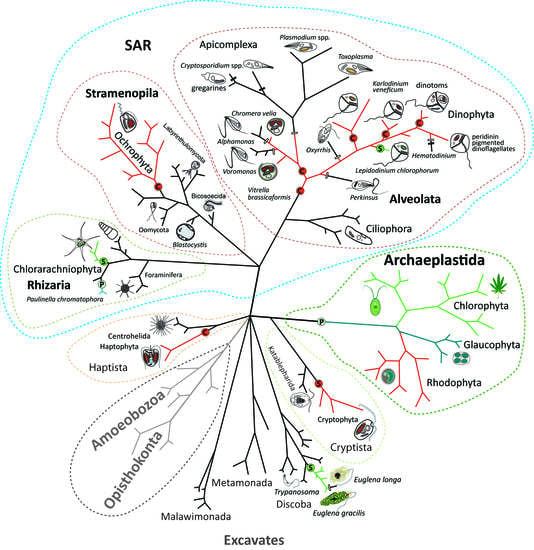
Endosymbiotic Evolution of Algae, Secondary Heterotrophy and Parasitism
Abstract
1. Introduction
2. Primary Endosymbioses
3. Complex Endosymbioses
4. Secondary Heterotrophy and Parasitism in Algae
5. Conclusions
Funding
Acknowledgments
Conflicts of Interest
References
- Shih, P.M. Cyanobacterial evolution: Fresh insight into ancient questions. Curr. Biol. 2015, 25, R193. [Google Scholar] [CrossRef] [PubMed]
- Shih, P.M.; Matzke, J. Primary endosymbiosis events date to the later Proterozoic with cross-calibrated phylogenetic dating of duplicated ATPase proteins. Proc. Natl. Acad. Sci USA 2013, 110, 12355–12360. [Google Scholar] [CrossRef] [PubMed]
- Mereschkowksy, C. Ober Natur und Ursprung der Chromatophoren im Pflanzenreiche. Biol. Zentralbl. 1905, 25, 593–604. [Google Scholar]
- Martin, W.; Kowallik, K.V. Annotated English translation of of Mereschkowsky’s 1905 paper “Über natur und Ursprung der Chromatophoren im Pflanzenreiche”. Eur. J. Phycol. 1999, 34, 287–295. [Google Scholar] [CrossRef]
- Pallen, M.J. Time to recognize that mitochondria are bacteria? Trends Microbiol. 2011, 19, 58–61. [Google Scholar] [CrossRef] [PubMed]
- Oborník, M. In the beginning was the word: How terminology drives our understanding of endosymbiotic organelles. Microbial Cell 2019, 6, 134–141. [Google Scholar] [CrossRef] [PubMed]
- Cavalier-Smith, T.; Lee, J.J. Protozoa as hosts for endosymbioses and the conversion of symbionts into organelles. J. Protozool. 1985, 32, 376–379. [Google Scholar] [CrossRef]
- Gould, S.B.; Waller, R.F.; McFadden, G.I. Plastid evolution. Ann. Rev. Plant. Biol. 2008, 59, 491–517. [Google Scholar] [CrossRef]
- Keeling, P.J. The number, speed, and impact of plastid endosymbioses in eukaryotic evolution. Ann. Rev. Plant. Biol. 2013, 64, 583–607. [Google Scholar] [CrossRef]
- Archibald, J.M. Endosymbiosis and eukaryotic cell evolution. Curr. Biol. 2015, 25, R911–R921. [Google Scholar] [CrossRef]
- Gruber, A. What`s a name? Why organelles of endosymbiotic origin are implicitly called by their eukaryotic species name and how they can be distinguished from endosymbionts. Microbial Cell 2019, 6, 123–133. [Google Scholar] [CrossRef]
- Keeling, P.J. Diversity and evolutionary history of plastids and their hosts. Am. J. Bot. 2004, 91, 1481–1493. [Google Scholar] [CrossRef] [PubMed]
- Jackson, C.; Clayden, S.; Reyes-Prieto, A. The Glaucophyta: The blue-green plants in a nutshell. Acta Soc. Bot. Pol. 2015, 84, 149–165. [Google Scholar] [CrossRef]
- Maréchal, E. (Ed.) Plastids: Methods and protocols. Methods in molecular biology. In Primary Endosymbiosis: Emergence of Primary Chloroplasts and Chromatophore Two Independent Events; Humana Press: New York, NY, USA, 2018; Volume 1829. [Google Scholar]
- Larkum, A.W.D.; Kühl, M. Chlorophyll d: The puzzle resolved. Trends Plant. Sci. 2005, 10, P355–P357. [Google Scholar] [CrossRef] [PubMed]
- Martin, W.; Herrmann, R.G. Gene transfer from organelles to the nucleus: How much, what happens, and why? Plant. Phys. 1998, 118, 9–17. [Google Scholar] [CrossRef] [PubMed]
- Oborník, M.; Green, B. Mosaic origin of the heme biosynthesis pathway in photosynthetic eukaryotes. Mol. Biol. Evol. 2005, 22, 2343–2353. [Google Scholar] [CrossRef] [PubMed]
- Cihlář, J.; Füssy, Z.; Oborník, M. Evolution of tetrapyrrole pathway in eukaryotic phototrophs. Adv. Bot. Res. 2019, 90, 273–309. [Google Scholar]
- Cihlář, J.; Füssy, Z.; Horák, A.; Oborník, M. Evolution of the tetrapyrrole biosyntheses pathway in secondary algae: Conservation, redundancy, and replacement. PLoS ONE 2016, 11, e0166338. [Google Scholar] [CrossRef]
- Stiller, J.W.; Reel, D.C.; Johnson, J.C. A single origin of plastids revisited: Convergent evolution in organellar genome content. J. Phycol. 2003, 39, 95–105. [Google Scholar] [CrossRef]
- Cavalier-Smith, T. Principles of protein and lipid targeting in secondary symbiogenesis: Euglenoid, dinoflagellate, and sporozoan plastid origins and the eukaryote family tree. J. Eukaryot. Microbiol. 1999, 46, 347–366. [Google Scholar] [CrossRef]
- Marin, B.; Nowack, E.C.M.; Melkonian, M. A plastid in the making: Evidence for a second primary endosymbiosis. Protist 2005, 156, 425–432. [Google Scholar] [CrossRef] [PubMed]
- Yoon, H.S.; Reyes-Prieto, A.; Melkonian, M.; Bhattacharya, D. Minimal plastid genome evolution in the Paulinella endosymbiont. Curr. Biol. 2006, 16, R670–R672. [Google Scholar] [CrossRef] [PubMed]
- Delaye, L.; Valadez-Cano, C.; Pérez-Zamorano, B. How really ancient is Paulinella chromatophora? PLoS Curr. Tree of Life 2016, 1. [Google Scholar] [CrossRef] [PubMed]
- Delwiche, C.F.; Palmer, J.D. The Origin Plastids and Their Spread via Secondary Symbiosis Plant Systematics and Evolution; Springer: Berlin, Germany, 1997; pp. 53–86. (Suppl. 11). [Google Scholar]
- Delwiche, C.F. Tracing the thread of plastid diversity through the tapestry of life. Am. Nat. 1999, 154, S164–S177. [Google Scholar] [CrossRef] [PubMed]
- Falkowski, P.G.; Katz, M.E.; Knoll, A.H.; Quigg, A.; Raven, J.A.; Schofield, O.; Taylor, F.J. The evolution of modern phytoplankton. Science 2004, 305, 354–360. [Google Scholar] [CrossRef] [PubMed]
- Oborník, M. The birth of red complex plastids: One, three, or four times? Trends Parasitol. 2018, 34, 923–925. [Google Scholar] [CrossRef] [PubMed]
- Curtis, B.; Tanifuji, G.; Burki, F.; Gruber, A.; Irimia, M.; Maruyama, S.; Arias, M.; Ball, S.; Gile, G.; Hirakawa, Y.; et al. Algal genomes reveal evolutionary mosaicism and the fate of nucleomorphs. Nature 2012, 492, 59–65. [Google Scholar] [CrossRef]
- Vanclová, A.M.G.; Hadariová, L.; Hrdá, Š.; Hampl, V. Secondary plastids of euglenophytes. Adv. Bot. Res. 2017, 84, 321–358. [Google Scholar]
- Matsumoto, T.; Shinozaki, F.; Chikuni, T.; Yabuki, A.; Takishita, K.; Kawachi, M.; Nakayama, T.; Inouye, I.; Hashimoto, T.; Inagaki, Y. Green-colored plastids in the dinoflagellate genus Lepidodinium are of core chlorophyte origin. Protist 2011, 162, 268–276. [Google Scholar] [CrossRef]
- Waller, R.F.; Kořený, L. Plastid complexity on dinogflagellates: A picture of gains, losses, replacements and revisions. Adv. Bot. Res. 2017, 84, 105–143. [Google Scholar]
- Larkum, A.W.D.; Lockhart, P.J.; Howe, C.J. Shopping for plastids. Trends Plant. Sci. 2007, 12, 189–195. [Google Scholar] [CrossRef] [PubMed]
- Cavalier-Smith, T. Kingdom Chromista and its eight phyla: A new synthesis emphasising periplastid protein targeting, cyto-skeletal and periplastid evolution, and ancient divergences. Protoplasma 2018, 255, 297–357. [Google Scholar] [CrossRef] [PubMed]
- Bodył, A.; Stiller, J.W.; Mackiewicz, P. Chromalveolate plastids: Direct descents or multiple endosymbioses? Trends Ecol. Evol. 2009, 24, 119–121. [Google Scholar] [CrossRef] [PubMed]
- Stiller, J.W.; Schreiber, J.; Yue, J.; Guo, H.; Ding, Q.; Huang, J. The evolution of photosynthesis in chromist algae through serial endosymbioses. Nat. Com. 2014, 5, 5764. [Google Scholar] [CrossRef] [PubMed]
- Petersen, J.; Ludewig, A.K.; Michael, V.; Bunk, B.; Jarek, M.; Baurain, D.; Brinkmann, H. Chromera velia, endosymbioses and the rhodoplex hypothesis—Plastid evolution in cryptophytes, alveolates, stramenopiles, and haptophytes (CASH lineages). Genome Biol. Evol. 2014, 6, 666–684. [Google Scholar] [CrossRef] [PubMed]
- Dorrell, D.R.; Bowler, C. Secondary plastids of stramenopiles. Adv. Bot. Res. 2017, 84, 57–103. [Google Scholar]
- Moore, C.E.; Archibald, J.M. Nucleomorph genomes. Ann. Rev. Genet. 2009, 43, 251–264. [Google Scholar] [CrossRef]
- Oborník, M.; Modrý, D.; Lukeš, M.; Černotíková-Stříbrná, E.; Cihlář, J.; Tesařová, M.; Kotabová, E.; Vancová, E.; Prášil, O.; Lukeš, J. Morphology, ultrastructure and life cycle of Vitrella brassicaformis n. sp., n. gen., a novel chromerid from the Great Barrier Reef. Protist 2012, 163, 306–323. [Google Scholar] [CrossRef]
- Ševčíková, T.; Horák, A.; Klimeš, V.; Zbránková, V.; Demir-Hilton, E.; Sudek, S.; Jenkins, J.; Schmutz, J.; Přibyl, P.; Fousek, J.; et al. Updating algal evolutionary relationships through plastid genome sequencing: Did alveolate plastids emerge through endosymbiosis of an ochrophyte? Sci. Rep. 2015, 5, 10134. [Google Scholar] [CrossRef] [PubMed]
- Sobotka, R.; Esson, H.J.; Koník, P.; Trsková, E.; Moravcová, L.; Horák, A.; Dufková, P.; Oborník, M. Extensive gain and loss of photosystem I subunits in chromerid algae, photosynthetic relatives of apicomplexans. Sci. Rep. 2017, 7, 13214. [Google Scholar] [CrossRef]
- Oborník, M.; Janouškovec, J.; Chrudimský, T.; Lukeš, J. Evolution of the apicoplast and its host: From heterotrophy to autotrophy and back again. Int. J. Parasitol. 2009, 39, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Pospíšil, P. Molecular mechanisms of production and scavenging of reactive oxygen species by photosystem II. Biochim. Biophys. Acta 2012, 1817, 218–231. [Google Scholar] [CrossRef] [PubMed]
- Koblížek, M.; Zeng, Y.; Horák, A.; Oborník, M. Regressive evolution of photosynthesis in the Roseobacter clade. Adv. Bot. Res. 2013, 66, 385–405. [Google Scholar]
- Hadariová, L.; Vesteg, M.; Hampl, V.; Krajčovič, J. Reductive evolution of chloroplasts in non-photosynthetic plants, algae and protists. Curr. Genet. 2018, 64, 365–387. [Google Scholar] [CrossRef] [PubMed]
- Votýpka, J.; Nodrý, D.; Oborník, M.; Šlapeta, J.; Lukeš, J. Apicomplexa. In Handbook of the Protists; Archibald, J.M., Simpson, G.B., Slamovits, C.H., Eds.; Springer: Berlin, Germany, 2017; pp. 567–624. [Google Scholar]
- Těšitel, J. Functional biology of parasitic plants: A review. Plant. Ecol. Evol. 2017, 149, 5–20. [Google Scholar] [CrossRef]
- De la Cruz, V.F.; Gittleson, S.M. The genus Polytomella: A review of classification, morphology, life cycle, metabolism, and motility. Arch. Protistenkunde 1981, 124, 1–28. [Google Scholar] [CrossRef]
- Boucias, D.G.; Becnel, J.J.; White, S.E.; Bott, M. In vivo and in vitro development of the protist Helicosporidium sp. J. Eukaryot. Microbiol. 2001, 48, 460–470. [Google Scholar] [CrossRef] [PubMed]
- De Koning, A.P.; Keeling, P.J. The complete plastid genome sequence of the parasitic green alga Helicosporidium sp. is highly reduced and structured. BMC Biol. 2006, 4, 12. [Google Scholar] [CrossRef] [PubMed]
- Blouin, N.A.; Lane, C.E. Red algae provide fertile ground for exploring parasite evolution. Persp. Phycol. 2016, 2016. 3, 11–19. [Google Scholar] [CrossRef]
- Marin, B.; Palm, A.; Klingberg, M.; Melkonian, M. Phylogeny and taxonomic revision of plastid-containing euglenophytes based on SSU rDNA sequence comparisons and synapomorhic signatures in the SSU rRNA secondary structure. Protist 2003, 154, 99–145. [Google Scholar] [CrossRef]
- Lowe, C.D.; Keeling, P.J.; Martin, L.E.; Slamovits, C.H.; Watts, P.C.; Montagnes, D.J.S. Who is Oxyrrhis marina? Morphological and phylogenetic studies on an unusual dinoflagellate. J. Plankton Res. 2011, 33, 555–567. [Google Scholar] [CrossRef]
- Füssy, Z.; Oborník, M. Chromerids and their plastids. Adv. Bot. Res. 2017, 84, 187–218. [Google Scholar]
- Füssy, Z.; Oborník, M. Plastids: Methods and protocols methods in molecular biology. In Complex Endosymbioses I: From Primary to Complex Plastids, Multiple Independent Events; Maréchal, E., Ed.; Humana Press: New York, NY, USA, 2018; Volume 1829, pp. 17–35. [Google Scholar]
- Sato, S. The apicomplexan plastid and its evolution. Cell. Mol. Life Sci. 2011, 68, 1285–1296. [Google Scholar] [CrossRef] [PubMed]
- Toso, M.A.; Omoto, C.K. Gregarina niphandrodes may lack both a plastid genome and organelle. J. Euk. Microbiol. 2007, 54, 66–72. [Google Scholar] [CrossRef] [PubMed]
- Gornik, S.G.; Febrimarsa, A.M.C.; MacRae, J.I.; Ramprasad, A.; Rchiad, Z.; McConville, M.J.; Bacic, A.; McFadden, G.I.; Pain, A.; Waller, R.F. Endosymbiosis undone by stepwise elimination of the plastid in a parasitic dinoflagellate. Proc. Natl. Acad. Sci. USA 2015, 112, 5767–5772. [Google Scholar] [CrossRef] [PubMed]
- Burki, F. The convoluted evolution of eukaryotes with complex plastids. Adv. Bot. Res. 2017, 84, 1–30. [Google Scholar]
- Stoecker, D.K.; Hansen, P.J.; Caron, D.A.; Mitra, A. Mixotrophy in the marine plankton. Ann. Rev. Mar. Sci. 2016, 9, 311–335. [Google Scholar] [CrossRef]
- Rezic, T.; Filipović, J.; Santec, B. Photo-mixotrophic cultivation of algae Euglena gracilis for lipid production. Agr. Consp. Sci. 2013, 2013 78, 65–69. [Google Scholar]
- Heredia-Arroyo, T.; Wei, W.; Hu, B. Mixotrophic cultivation of Chlorella vulgaris and its potential application for the oil accumulation from non-sugar materials. Biomass Bioenerg. 2011, 35, 2245–2253. [Google Scholar] [CrossRef]
- Villanova, V.; Fortunato, A.E.; Singh, D.; Bo, D.D.; Conte, M.; Obata, T.; Jouhet, J.; Fernie, A.R.; Maréchal, E.; Falciatore, A.; et al. Investigating mixotrophic metabolism in the model diatom Phaeodactylum tricornutum. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2017, 372, 20160404. [Google Scholar] [CrossRef]
- Jeong, J.H.; Yoo, Y.D.; Kim, J.S.; Seong, K.A.; Kang, N.S.; Kim, T.H. Growth, feeding and ecological roles of the mixotrophic and heterotrophic dinoflagellates in marine planktonic food webs. Ocean Sci. J. 2010, 45, 65–91. [Google Scholar] [CrossRef]
- McFadden, G.I.; Reith, M.E.; Munholland, J.; Lang-Unnasch, N. Plastid in human parasites. Nature 1996, 381, 482. [Google Scholar] [CrossRef] [PubMed]
- McFadden, G.I.; Waller, R.F. Plastids in parasites of humans. Bioessays 1997, 19, 1033–1040. [Google Scholar] [CrossRef] [PubMed]
- Moore, R.B.; Oborník, M.; Janouškovec, J.; Chrudimský, T.; Vancová, M.; Green, D.; Wright, S.; Davies, N.; Bolch, C.; Heimann, K.; et al. A photosynthetic alveolate closely related to apicomplexan parasites. Nature 2008, 451, 959–963. [Google Scholar] [CrossRef] [PubMed]
- Woo, Y.; Ansari, H.; Otto, T.D.; Klinger, C.; Kolísko, M.; Saxena, A.; Shanmugam, D.; Tayyrov, A.; Veluchamy, A.; Ali, S.; et al. Chromerid genomes reveal the evolutionary path from photosynthetic algae to obligate intracellular parasites. elife 2015, 4, e06974. [Google Scholar] [CrossRef] [PubMed]
- Janouškovec, J.; Tikhonenkov, D.V.; Burki, F.; Howe, A.T.; Kolísko, M.; Mylnikov, A.P.; Keeling, P.J. Factors mediating plastid dependency and the origins of parasitism in apicomplexans and their close relatives. Proc. Natl. Acad. Sci. USA 2015, 112, 10200–10207. [Google Scholar] [CrossRef] [PubMed]
- Kořený, L.; Sobotka, R.; Janouškovec, J.; Keeling, P.J.; Oborník, M. Tetrapyrrole synthesis of photosynthetic chromerids is likely homologous to the unusual pathway of apicomplexan parasites. Plant. Cell 2011, 23, 3454–3462. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Okamoto, N.; McFadden, G.I. The mother of all parasites. Futur. Microbiol. 2008, 3, 391–395. [Google Scholar] [CrossRef]
- Janouškovec, J.; Horák, A.; Barott, K.L.; Rohwer, F.L.; Keeling, P.J. Global analysis of plastid diversity reveals apicomplexan-related lineages in coral reefs. Curr. Biol. 2012, 22, R518–R519. [Google Scholar] [CrossRef]
- Janouškovec, J.; Horák, A.; Barott, K.L.; Rohwer, F.L.; Keeling, P.J. Environmental distribution of coral-associated relatives of apicomplexan parasites. ISME J. 2013, 7, 444–447. [Google Scholar] [CrossRef]
- Cumbo, V.R.; Baird, A.H.; Moore, R.B.; Negri, A.P.; Neilan, B.A.; Salih, A.; van Oppen, M.J.; Marquis, C.P. Chromera velia is endosymbiotic in larvae of the reef corals Acropora digitifera and A. tenuis. Protist 2013, 164, 237–244. [Google Scholar] [CrossRef] [PubMed]
- Mohamed, A.R.; Cumbo, V.R.; Harii, A.; Shizato, C.; Chan, C.X.; Ragan, M.A.; Satoh, N.; Ball, E.E.; Miller, D.J. Deciphering the nature of the coral-Chromera association. ISME J. 2018, 12, 776–790. [Google Scholar] [CrossRef] [PubMed]
- Skovgaard, A.; Karpov, S.A.; Guillou, L. The parasitic dinoflagellates Blastodinium spp. Inhabiting the gut of marine, plaktonic copepods: Morphology, ecology, and unrecognized species diversity. Front. Microbiol. 2012, 3, 305. [Google Scholar] [CrossRef] [PubMed]
- Keeling, P.J.; McCutcheon, J.P. Endosymbiosis: The feeling is not mutual. J. Theor. Biol. 2017, 434, 75–79. [Google Scholar] [CrossRef] [PubMed]
- Kwong, W.K.; Del Campo, J.; Mathur, V.; Vermeij, M.J.A.; Keeling, P.J. A widespread coral-infecting apicomplexan with chlorophyll biosynthesis genes. Nature 2019, 568, 103–107. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oborník, M. Endosymbiotic Evolution of Algae, Secondary Heterotrophy and Parasitism. Biomolecules 2019, 9, 266. https://doi.org/10.3390/biom9070266
Oborník M. Endosymbiotic Evolution of Algae, Secondary Heterotrophy and Parasitism. Biomolecules. 2019; 9(7):266. https://doi.org/10.3390/biom9070266
Chicago/Turabian StyleOborník, Miroslav. 2019. "Endosymbiotic Evolution of Algae, Secondary Heterotrophy and Parasitism" Biomolecules 9, no. 7: 266. https://doi.org/10.3390/biom9070266
APA StyleOborník, M. (2019). Endosymbiotic Evolution of Algae, Secondary Heterotrophy and Parasitism. Biomolecules, 9(7), 266. https://doi.org/10.3390/biom9070266