Membrane Active Peptides and Their Biophysical Characterization
Abstract
1. Introduction
2. Antimicrobial Peptides
3. Cell-Penetrating Peptides
4. Antimicrobial Peptides and Cell-Penetrating Peptides
5. Characterization of Antimicrobial and Cell-Penetrating Peptides and Their Interactions with Membranes
6. Structural Analysis
6.1. X-ray and Neutron Scattering/Diffraction
6.2. Nuclear Magnetic Resonance Spectroscopy
6.3. Circular Dichroism Spectroscopy
6.4. Calorimetry Methods
6.5. Atomic Force Microscopy
6.6. Fourier Transform Infrared Spectroscopy
6.7. Dynamic Light Scattering
7. Live Imaging
7.1. Surface Plasmon Resonance
7.2. Cryoelectron Microscopy
7.3. Fluorescence Imaging: Microscopy and Spectroscopy
7.4. Flow Cytometry
8. Emerging Approaches
9. Computer Simulations
10. Biophysical Focus on Amphipathic Membrane Active Peptides
10.1. Melittin
10.2. pVEC
11. Future Perspectives: Characterizing Membrane Active Peptides for Better Peptide-Based Drugs
Funding
Conflicts of Interest
References
- Henriques, S.T.; Melo, M.N.; Castanho, M.A. How to address CPP and AMP translocation? Methods to detect and quantify peptide internalization in vitro and in vivo (Review). Mol. Membr. Biol. 2007, 24, 173–184. [Google Scholar] [CrossRef] [PubMed]
- Moreno, M.; Giralt, E. Three Valuable Peptides from Bee and Wasp Venoms for Therapeutic and Biotechnological Use: Melittin, Apamin and Mastoparan. Toxins 2015, 7, 1126–1150. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.W.; Lee, C.T.; Wang, T.C.; Kao, Y.C.; Yang, C.H.; Lin, Y.M.; Huang, K.S. The Development of Peptide-based Antimicrobial Agents against Dengue Virus. Curr. Protein Pept. Sci. 2018, 19, 1. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Yu, M.; He, Y.; Xiao, L.; Wang, F.; Song, C.; Sun, S.; Ling, C.; Xu, Z. Melittin prevents liver cancer cell metastasis through inhibition of the Rac1-dependent pathway. Hepatology 2008, 47, 1964–1973. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Li, X.; Wang, Z. APD3: The antimicrobial peptide database as a tool for research and education. Nucleic Acids Res. 2016, 44, D1087–D1093. [Google Scholar] [CrossRef] [PubMed]
- Agrawal, P.; Bhalla, S.; Usmani, S.S.; Singh, S.; Chaudhary, K.; Raghava, G.P.S.; Gautam, A. CPPsite 2.0: A repository of experimentally validated cell-penetrating peptides. Nucleic Acids Res. 2016, 44, D1098–D1103. [Google Scholar] [CrossRef] [PubMed]
- Brogden, K.A. Antimicrobial peptides: Pore formers or metabolic inhibitors in bacteria? Nat. Rev. Microbiol. 2005, 3, 238–250. [Google Scholar] [CrossRef] [PubMed]
- Dubos, R.J. Studies on a bactericidal agent extracted from a soil Bacillus. J. Exp. Med. 1939, 70, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Zasloff, M. Antimicrobial peptides of multicellular organisms. Nature 2002, 415, 389–395. [Google Scholar] [CrossRef] [PubMed]
- Lehrer, R.I.; Ganz, T.; Selsted, M.E. Defensins: Endogenous Antibiotic of Animal Cells. Cell 1991, 64, 229–230. [Google Scholar] [CrossRef]
- Schittek, B.; Hipfel, R.; Sauer, B.; Bauer, J.; Kalbacher, H.; Stevanovic, S.; Schirle, M.; Schroeder, K.; Blin, N.; Meier, F.; et al. Dermcidin: A novel human antibiotic peptide secreted by sweat glands. Nat. Immunol. 2001, 2, 1133–1137. [Google Scholar] [CrossRef] [PubMed]
- Johansson, J.; Gudmundsson, G.H.; Rottenberg, M.E.; Berndt, K.D.; Agerberth, B. Conformation-dependent antibacterial activity of the naturally occurring human peptide LL-37. J. Biol. Chem. 1998, 273, 3718–3724. [Google Scholar] [CrossRef] [PubMed]
- Stotz, H.U.; Thomson, J.G.; Wang, Y. Plant defensins: Defense, development and application. Plant Signal. Behav. 2009, 4, 1010–1012. [Google Scholar] [CrossRef] [PubMed]
- Carvalho Ade, O.; Gomes, V.M. Plant defensins—Prospects for the biological functions and biotechnological properties. Peptides 2009, 30, 1007–1020. [Google Scholar] [CrossRef] [PubMed]
- Gachomo, E.W.; Jimenez Lopez, J.C.; Kayodé, A.P.P.; Baba Moussa, L.; Kotchoni, S.O. Structural characterization of plant defensin protein superfamily. Mol. Biol. Rep. 2012, 39, 4461–4469. [Google Scholar] [CrossRef] [PubMed]
- Habermann, E. Bee and wasp venoms. Science 1972, 177, 314–322. [Google Scholar] [CrossRef] [PubMed]
- Tamang, D.G.; Saier, M.H. The cecropin superfamily of toxic peptides. J. Mol. Microbiol. Biotechnol. 2006, 11, 94–103. [Google Scholar] [CrossRef] [PubMed]
- Zasloff, M. Magainins, a Class of Antimicrobial Peptides from Xenopus Skin: Isolation, Characterization of Two Active Forms, and Partial cDNA Sequence of a Precursor. J. Occup. Environ. Med. 1987, 30, 470. [Google Scholar] [CrossRef]
- Shagaghi, N.; Palombo, E.A.; Clayton, A.H.A.; Bhave, M. Antimicrobial peptides: Biochemical determinants of activity and biophysical techniques of elucidating their functionality. World J. Microbiol. Biotechnol. 2018, 34, 62. [Google Scholar] [CrossRef] [PubMed]
- Torres, A.M.; Kuchel, P.W. The β-defensin-fold family of polypeptides. Toxicon 2004, 44, 581–588. [Google Scholar] [CrossRef] [PubMed]
- Guder, A.; Wiedemann, I.; Sahl, H. Posttranslationally Modified Bacteriocins—The Lantibiotics. Biopolymers 2000, 55, 62–73. [Google Scholar] [CrossRef]
- Paulmann, M.; Arnold, T.; Linke, D.; Özdirekcan, S.; Kopp, A.; Gutsmann, T.; Kalbacher, H.; Wanke, I.; Schuenemann, V.J.; Habeck, M.; et al. Structure-activity analysis of the dermcidin-derived peptide DCD-1L, an anionic antimicrobial peptide present in human sweat. J. Biol. Chem. 2012, 287, 8434–8443. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Rigby, K.; Lai, Y.; Nair, V.; Peschel, A.; Schittek, B.; Otto, M. Staphylococcus aureus mutant screen reveals interaction of the human antimicrobial peptide dermcidin with membrane phospholipids. Antimicrob. Agents Chemother. 2009, 53, 4200–4210. [Google Scholar] [CrossRef] [PubMed]
- Strub, J.M.; Goumon, Y.; Lugardon, K.; Capon, C.; Lopez, M.; Moniatte, M.; Van Dorsselaer, A.; Aunis, D.; Metz-Boutigue, M.H. Antibacterial activity of glycosylated and phosphorylated chromogranin A-derived peptide 173–194 from bovine adrenal medullary chromaffin granules. J. Biol. Chem. 1996, 271, 28533–28540. [Google Scholar] [CrossRef] [PubMed]
- Fjell, C.D.; Hiss, J.A.; Hancock, R.E.W.; Schneider, G. Designing antimicrobial peptides: Form follows function. Nat. Rev. Drug Discov. 2012, 16, 37–51. [Google Scholar] [CrossRef] [PubMed]
- Song, C.; Weichbrodt, C.; Salnikov, E.S.; Dynowski, M.; Forsberg, B.O.; Bechinger, B.; Steinem, C.; de Groot, B.L.; Zachariae, U.; Zeth, K. Crystal structure and functional mechanism of a human antimicrobial membrane channel. Proc. Natl. Acad. Sci. USA 2013, 110, 4586–4591. [Google Scholar] [CrossRef] [PubMed]
- Wommack, A.J.; Robson, S.A.; Wanniarachchi, Y.A.; Wan, A.; Turner, C.J.; Wagner, G.; Nolan, E.M. NMR solution structure and condition-dependent oligomerization of the antimicrobial peptide human defensin 5. Biochemistry 2012, 51, 9624–9637. [Google Scholar] [CrossRef] [PubMed]
- Schnell, N.; Entian, K.D.; Schneider, U.; Götz, F.; Zähner, H.; Kellner, R.; Jung, G. Prepeptide sequence of epidermin, a ribosomally synthesized antibiotic with four sulphide-rings. Nature 1988, 333, 276–278. [Google Scholar] [CrossRef] [PubMed]
- Splith, K.; Neundorf, I. Antimicrobial peptides with cell-penetrating peptide properties and vice versa. Eur. Biophys. J. 2011, 40, 387–397. [Google Scholar] [CrossRef] [PubMed]
- Fox, N.K.; Brenner, S.E.; Chandonia, J.M. SCOPe: Structural Classification of Proteins–extended, integrating SCOP and ASTRAL data and classification of new structures. Nucleic Acids Res. 2013, 42, 304–309. [Google Scholar] [CrossRef] [PubMed]
- Guilhelmelli, F.; Vilela, N.; Albuquerque, P.; Derengowski, L.S.; Silva-Pereira, I.; Kyaw, C.M. Antibiotic development challenges: The various mechanisms of action of antimicrobial peptides and of bacterial resistance. Front. Microbiol. 2013, 4, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Fanghänel, S.; Wadhwani, P.; Strandberg, E.; Verdurmen, W.P.R.; Bürck, J.; Ehni, S.; Mykhailiuk, P.K.; Afonin, S.; Gerthsen, D.; Komarov, I.V.; et al. Structure analysis and conformational transitions of the cell penetrating peptide transportan 10 in the membrane-bound state. PLoS ONE 2014, 9, e99653. [Google Scholar] [CrossRef] [PubMed]
- Chan, D.I.; Prenner, E.J.; Vogel, H.J. Tryptophan- and arginine-rich antimicrobial peptides: Structures and mechanisms of action. Biochim. Biophys. Acta Biomembr. 2006, 1758, 1184–1202. [Google Scholar] [CrossRef] [PubMed]
- Wang, G. Structures of human host defense cathelicidin LL-37 and its smallest antimicrobial peptide KR-12 in lipid micelles. J. Biol. Chem. 2008, 283, 32637–32643. [Google Scholar] [CrossRef] [PubMed]
- Oh, D.; Shin, S.Y.; Kang, J.H.; Hahm, K.S.; Kim, K.L.; Kim, Y. NMR structural characterization of cecropin A(1–8)—Magainin 2(1–12) and cecropin A(1–8)—Melittin(1–12) hybrid peptides. J. Pept. Res. 1999, 53, 578–589. [Google Scholar] [CrossRef] [PubMed]
- Lindberg, M.; Biverståhl, H.; Gräslund, A.; Mäler, L. Structure and positioning comparison of two variants of penetratin in two different membrane mimicking systems by NMR. Eur. J. Biochem. 2003, 270, 3055–3063. [Google Scholar] [CrossRef] [PubMed]
- Bárány-Wallje, E.; Andersson, A.; Gräslund, A.; Mäler, L. NMR solution structure and position of transportan in neutral phospholipid bicelles. FEBS Lett. 2004, 567, 265–269. [Google Scholar] [CrossRef] [PubMed]
- Kerkis, I.; Hayashi, M.A.F.; da Silva, A.R.B.; Pereira, A.; De Sá Júnior, P.L.; Zaharenko, A.J.; Rádis-Baptista, G.; Kerkis, A.; Yamane, T. State of the Art in the Studies on Crotamine, a Cell Penetrating Peptide from South American Rattlesnake. BioMed Res. Int. 2014, 2014, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Kumar, P.; Kizhakkedathu, J.; Straus, S. Antimicrobial Peptides: Diversity, Mechanism of Action and Strategies to Improve the Activity and Biocompatibility In Vivo. Biomolecules 2018, 8, 4. [Google Scholar] [CrossRef] [PubMed]
- Wimley, W.C. Describing the mechanism of antimicrobial peptide action with the interfacial activity model. ACS Chem. Biol. 2011, 5, 905–917. [Google Scholar] [CrossRef] [PubMed]
- Vácha, R.; Frenkel, D. Simulations suggest possible novel membrane pore structure. Langmuir 2014, 30, 1304–1310. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Harroun, T.A.; Weiss, T.M.; Ding, L.; Huang, H.W. Barrel-stave model or toroidal model? A case study on melittin pores. Biophys. J. 2001, 81, 1475–1485. [Google Scholar] [CrossRef]
- Rapaport, D.; Shai, Y. Aggregation and organization of pardaxin in phospholipid bilayers. A fluorescence energy transfer study. J. Biol. Chem. 1992, 267, 6502–6509. [Google Scholar] [PubMed]
- Jafari, M.; Mehrnejad, F.; Doustdar, F. Insight into the interactions, residue snorkeling, and membrane disordering potency of a single antimicrobial peptide into different lipid bilayers. PLoS ONE 2017, 12, e0187216. [Google Scholar] [CrossRef] [PubMed]
- Leveritt, J.M., III; Pino Angeles, A.; Lazaridis, T. The Structure of a Melittin-Stabilized Pore. Biophys. J. 2015, 108, 2424–2426. [Google Scholar] [CrossRef] [PubMed]
- Pino Angeles, A.; Lazaridis, T. Effects of Peptide Charge, Orientation, and Concentration on Melittin Transmembrane Pores. Biophys. J. 2018, 114, 2865–2874. [Google Scholar] [CrossRef] [PubMed]
- Uematsu, N.; Matsuzaki, K. Polar Angle as a Determinant of Amphipathic α-Helix-Lipid Interactions: A Model Peptide Study. Biophys. J. 2000, 79, 2075–2083. [Google Scholar] [CrossRef]
- Yeaman, M.R. Mechanisms of Antimicrobial Peptide Action and Resistance. Pharmacol. Rev. 2003, 55, 27–55. [Google Scholar] [CrossRef] [PubMed]
- Lee, T.H.; Hall, K.N.; Aguilar, M.I. Antimicrobial Peptide Structure and Mechanism of Action: A Focus on the Role of Membrane Structure. Curr. Top. Med. Chem. 2016, 16, 25–39. [Google Scholar] [CrossRef] [PubMed]
- Marrink, S.J.; de Vries, A.H.; Tieleman, D.P. Lipids on the move: Simulations of membrane pores, domains, stalks and curves. Biochim. Biophys. Acta Biomembr. 2009, 1788, 149–168. [Google Scholar] [CrossRef] [PubMed]
- Matsuzaki, K.; Yoneyama, S.; Miyajima, K. Pore formation and translocation of melittin. Biophys. J. 1997, 73, 831–838. [Google Scholar] [CrossRef]
- Matsuzaki, K.; Sugishita, K.I.; Ishibe, N.; Ueha, M.; Nakata, S.; Miyajima, K.; Epand, R.M. Relationship of membrane curvature to the formation of pores by magainin 2. Biochemistry 1998, 37, 11856–11863. [Google Scholar] [CrossRef] [PubMed]
- Shai, Y. Mode of action of membrane active antimicrobial peptides. Biopolymers 2002, 66, 236–248. [Google Scholar] [CrossRef] [PubMed]
- Fernandez, D.I.; Le Brun, A.P.; Whitwell, T.C.; Sani, M.A.; James, M.; Separovic, F. The antimicrobial peptide aurein 1.2 disrupts model membranes via the carpet mechanism. Phys. Chem. Chem. Phys. 2012, 14, 15739–15751. [Google Scholar] [CrossRef] [PubMed]
- Gazit, E.; Miller, I.R.; Biggin, P.C.; Sansom, M.S.P.; Shai, Y. Structure and orientation of the mammalian antibacterial peptide cecropin P1 within phospholipid membranes. J. Mol. Biol. 1996, 258, 860–870. [Google Scholar] [CrossRef] [PubMed]
- Marquette, A.; Bechinger, B. Biophysical Investigations Elucidating the Mechanisms of Action of Antimicrobial Peptides and Their Synergism. Biomolecules 2018, 8, 18. [Google Scholar] [CrossRef] [PubMed]
- Kai-Larsen, Y.; Gudmundsson, G.H.; Agerberth, B. A review of the innate immune defence of the human foetus and newborn, with the emphasis on antimicrobial peptides. Acta Paediatr. Int. J. Paediatr. 2014, 103, 1000–1008. [Google Scholar] [CrossRef] [PubMed]
- Cerrato, C.P.; Künnapuu, K.; Langel, Ü. Cell-penetrating peptides with intracellular organelle targeting. Expert Opin. Drug Deliv. 2017, 14, 245–255. [Google Scholar] [CrossRef] [PubMed]
- Brumfitt, W.; Salton, M.R.; Hamilton Miller, J.M. Nisin, alone and combined with peptidoglycan-modulating antibiotics: Activity against methicillin-resistant Staphylococcus aureus and vancomycin-resistant enterococci. J. Antimicrob. Chemother. 2002, 50, 731–734. [Google Scholar] [CrossRef] [PubMed]
- Otvos, L., Jr.; Insug, O.; Rogers, M.E.; Consolvo, P.J.; Condie, B.A.; Lovas, S.; Bulet, P.; Blaszczyk-Thurin, M. Interaction between Heat Shock Proteins and Antimicrobial Peptides. Biochemistry 2000, 39, 14150–14159. [Google Scholar] [CrossRef] [PubMed]
- Kragol, G.; Lovas, S.; Varadi, G.; Condie, B.A.; Hoffmann, R.; Otvos, L.J. The antibacterial peptide pyrrhocoricin inhibits the ATPase actions of DnaK and prevents chaperone-assisted protein folding. Biochemistry 2001, 40, 3016–3026. [Google Scholar] [CrossRef] [PubMed]
- Wu, W.K.K.; Wang, G.; Coffelt, S.B.; Betancourt, A.M.; Lee, C.W.; Fan, D.; Wu, K.; Yu, J.; Sung, J.J.Y.; Cho, C.H. Emerging roles of the host defense peptide LL-37 in human cancer and its potential therapeutic applications. Int. J. Cancer 2011, 127, 1741–1747. [Google Scholar] [CrossRef] [PubMed]
- Kai-Larsen, Y.; Agerberth, B. The role of the multifunctional peptide LL-37 in host defense. Front. Biosci. 2008, 13, 3760–3767. [Google Scholar] [CrossRef] [PubMed]
- Yang, D.; Chertov, O.; Bykovskaia, S.N.; Chen, Q.; Buffo, M.J.; Shogan, J.; Anderson, M.; Schroder, J.M.; Wang, J.M.; Howard, M.Z.; et al. β-Defensins-Linking Innate and Adaptive Immunity Through Dendritic and T Cell CCR6. Science 1999, 286, 525–527. [Google Scholar] [CrossRef] [PubMed]
- Lindgren, M.; Hallbrink, M.; Prochiantz, A.; Langel, U. Cell-penetrating peptides. Trends Pharmacol. Sci. 2000, 21, 99–103. [Google Scholar] [CrossRef]
- Guidotti, G.; Brambilla, L.; Rossi, D. Cell-Penetrating Peptides: From Basic Research to Clinics. Trends Pharmacol. Sci. 2017, 38, 406–424. [Google Scholar] [CrossRef] [PubMed]
- Morris, M.C.; Deshayes, S.; Heitz, F.; Divita, G. Cell-penetrating peptides: From molecular mechanisms to therapeutics. Biol. Cell 2008, 100, 201–217. [Google Scholar] [CrossRef] [PubMed]
- Shi, N.Q.; Qi, X.R.; Xiang, B.; Zhang, Y. A survey on “Trojan Horse” peptides: Opportunities, issues and controlled entry to “Troy”. J. Control. Release 2014, 194, 53–70. [Google Scholar] [CrossRef] [PubMed]
- Heitz, F.; Morris, M.C.; Divita, G. Twenty years of cell-penetrating peptides: from molecular mechanisms to therapeutics. Br. J. Pharmacol. 2009, 157, 195–206. [Google Scholar] [CrossRef] [PubMed]
- Islam, M.Z.; Sharmin, S.; Moniruzzaman, M.; Yamazaki, M. Elementary processes for the entry of cell-penetrating peptides into lipid bilayer vesicles and bacterial cells. Appl. Microbiol. Biotechnol. 2018, 102, 3879–3892. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, D.; Peng, X.; Leal, J.; Mohanty, R.P. Peptides as drug delivery vehicles across biological barriers. J. Pharm. Investig. 2018, 48, 89–111. [Google Scholar] [CrossRef] [PubMed]
- Frankel, A.D.; Pabo, C.O. Cellular uptake of the tat protein from human immunodeficiency virus. Cell 1988, 55, 1189–1193. [Google Scholar] [CrossRef]
- Derossi, D.; Chassaing, G.; Prochiantz, A. Trojan peptides: The penetratin system for intracellular delivery. Trends Cell Biol. 1998, 8, 84–87. [Google Scholar] [CrossRef]
- Schwarze, S.R.; Ho, A.; Vocero-Akbani, A.; Dowdy, S.F. In Vivo Protein Transduction: Delivery of a Biologically Active Protein into the Mouse. Science 1999, 285, 1569–1572. [Google Scholar] [CrossRef] [PubMed]
- Derossi, D.; Joliot, A.H.; Chassaing, G.; Prochiantz, A. The third helix of the Antennapedia homeodomain translocates through biological membranes. J. Biol. Chem. 1994, 269, 10444–10450. [Google Scholar] [PubMed]
- Thorén, P.E.G.; Persson, D.; Karlsson, M.; Nordén, B. The Antennapedia peptide penetratin translocates across lipid bilayers—The first direct observation. FEBS Lett. 2000, 482, 265–268. [Google Scholar] [CrossRef]
- Rittner, K.; Benavente, A.; Bompard-Sorlet, A.; Heitz, F.; Divita, G.; Brasseur, R.; Jacobs, E. New basic membrane-destabilizing peptides for plasmid-based gene delivery in vitro and in vivo. Mol. Ther. 2002, 5, 104–114. [Google Scholar] [CrossRef] [PubMed]
- Morris, M.C.; Depollier, J.; Mery, J.; Heitz, F.; Divita, G. A peptide carrier for the delivery of biologically active proteins into mammalian cells. Nat. Biotechnol. 2001, 19, 1173–1176. [Google Scholar] [CrossRef] [PubMed]
- Choi, K.M.; Kwon, I.C.; Ahn, H.J. Self-assembled amphiphilic DNA-cholesterol/DNA-peptide hybrid duplexes with liposome-like structure for doxorubicin delivery. Biomaterials 2013, 34, 4183–4190. [Google Scholar] [CrossRef] [PubMed]
- Pooga, M.; Hällbrink, M.; Zorko, M.; Langel, U. Cell penetration by transportan. FASEB J. 1998, 12, 67–77. [Google Scholar] [CrossRef] [PubMed]
- Islam, M.Z.; Ariyama, H.; Alam, J.M.; Yamazaki, M. Entry of cell-penetrating peptide transportan 10 into a single vesicle by translocating across lipid membrane and its induced pores. Biochemistry 2014, 53, 386–396. [Google Scholar] [CrossRef] [PubMed]
- Gautam, A.; Singh, H.; Tyagi, A.; Chaudhary, K.; Kumar, R.; Kapoor, P.; Raghava, G.P.S. CPPsite: A curated database of cell penetrating peptides. Database 2012, 2012. [Google Scholar] [CrossRef] [PubMed]
- The UniProt Consortium. UniProt: the universal protein knowledgebase. Nucleic Acids Res. 2017, 45, D158–D169. [Google Scholar] [CrossRef] [PubMed]
- Milletti, F. Cell-penetrating peptides: Classes, origin, and current landscape. Drug Discov. Today 2012, 17, 850–860. [Google Scholar] [CrossRef] [PubMed]
- Eiríksdóttir, E.; Konate, K.; Langel, Ü.; Divita, G.; Deshayes, S. Secondary structure of cell-penetrating peptides controls membrane interaction and insertion. Biochim. Biophys. Acta Biomembr. 2010, 1798, 1119–1128. [Google Scholar] [CrossRef] [PubMed]
- Czajlik, A.; Meskó, E.; Penke, B.; Perczel, A. Investigation of penetratin peptides. Part 1. The environment dependent conformational properties of penetratin and two of its derivatives. J. Pept. Sci. 2002, 8, 151–171. [Google Scholar] [CrossRef] [PubMed]
- Bárány-Wallje, E.; Andersson, A.; Gräslund, A.; Mäler, L. Dynamics of transportan in bicelles is surface charge dependent. J. Biomol. NMR 2006, 35, 137–147. [Google Scholar] [CrossRef] [PubMed]
- Bolhassani, A.; Jafarzade, B.S.; Mardani, G. In vitro and in vivo delivery of therapeutic proteins using cell penetrating peptides. Peptides 2017, 87, 50–63. [Google Scholar] [CrossRef] [PubMed]
- Dinca, A.; Chien, W.M.; Chin, M.T. Intracellular delivery of proteins with cell-penetrating peptides for therapeutic uses in human disease. Int. J. Mol. Sci. 2016, 17, 263. [Google Scholar] [CrossRef] [PubMed]
- Vivès, E.; Brodin, P.; Lebleu, B. A truncated HIV-1 Tat protein basic domain rapidly translocates through the plasma membrane and accumulates in the cell nucleus. J. Biol. Chem. 1997, 272, 16010–16017. [Google Scholar] [CrossRef] [PubMed]
- Rhee, M.; Davis, P. Mechanism of uptake of C105Y, a novel cell-penetrating peptide. J. Biol. Chem. 2006, 281, 1233–1240. [Google Scholar] [CrossRef] [PubMed]
- Suhorutsenko, J.; Oskolkov, N.; Arukuusk, P.; Kurrikoff, K.; Eriste, E.; Copolovici, D.M.; Langel, Ü. Cell-penetrating peptides, PepFects, show no evidence of toxicity and immunogenicity in vitro and in vivo. Bioconjug. Chem. 2011, 22, 2255–2262. [Google Scholar] [CrossRef] [PubMed]
- Akita, H.; Masuda, T.; Nishio, T.; Niikura, K.; Ijiro, K.; Harashima, H. Improving in vivo hepatic transfection activity by controlling intracellular trafficking: The function of GALA and maltotriose. Mol. Pharm. 2011, 8, 1436–1442. [Google Scholar] [CrossRef] [PubMed]
- Bechara, C.; Sagan, S. Cell-penetrating peptides: 20 years later, where do we stand? FEBS Lett. 2013, 587, 1693–1702. [Google Scholar] [CrossRef] [PubMed]
- Ezzat, K.; El Andaloussi, S.; Zaghloul, E.M.; Lehto, T.; Lindberg, S.; Moreno, P.M.D.; Viola, J.R.; Magdy, T.; Abdo, R.; Guterstam, P.; et al. PepFect 14, a novel cell-penetrating peptide for oligonucleotide delivery in solution and as solid formulation. Nucleic Acids Res. 2011, 39, 5284–5298. [Google Scholar] [CrossRef] [PubMed]
- Bolhassani, A. Potential efficacy of cell-penetrating peptides for nucleic acid and drug delivery in cancer. Biochim. Biophys. Acta Rev. Cancer 2011, 1816, 232–246. [Google Scholar] [CrossRef] [PubMed]
- Duchardt, F.; Fotin-Mleczek, M.; Schwarz, H.; Fischer, R.; Brock, R. A comprehensive model for the cellular uptake of cationic cell-penetrating peptides. Traffic 2007, 8, 848–866. [Google Scholar] [CrossRef] [PubMed]
- Guterstam, P.; Madani, F.; Hirose, H.; Takeuchi, T.; Futaki, S.; El Andaloussi, S.; Gräslund, A.; Langel, Ü. Elucidating cell-penetrating peptide mechanisms of action for membrane interaction, cellular uptake, and translocation utilizing the hydrophobic counter-anion pyrenebutyrate. Biochim. Biophys. Acta Biomembr. 2009, 1788, 2509–2517. [Google Scholar] [CrossRef] [PubMed]
- Trabulo, S.; Cardoso, A.L.; Mano, M.; de Lima, M.C.P. Cell-penetrating peptides-mechanisms of cellular uptake and generation of delivery systems. Pharmaceuticals 2010, 3, 961–993. [Google Scholar] [CrossRef] [PubMed]
- Madani, F.; Lindberg, S.; Langel, Ü.; Futaki, S.; Gräslund, A. Mechanisms of cellular uptake of cell-penetrating peptides. J. Biophys. 2011, 2011. [Google Scholar] [CrossRef] [PubMed]
- Derossi, D.; Calvet, S.; Trembleau, A.; Brunissen, A.; Chassaing, G.; Prochiantz, A. Cell internalization of the third helix of the Antennapedia Homeodomain is receptor-independent. J. Biol. Chem. 1996, 271, 18188–18193. [Google Scholar] [CrossRef] [PubMed]
- Matsuzaki, K.; Yoneyama, S.; Murase, O.; Miyajima, K. Transbilayer transport of ions and lipids coupled with mastoparan X translocation. Biochemistry 1996, 35, 8450–8456. [Google Scholar] [CrossRef] [PubMed]
- Pouny, Y.; Rapaport, D.; Shai, Y.; Mor, A.; Nicolas, P. Interaction of Antimicrobial Dermaseptin and its Fluorescently Labeled Analogs with Phospholipid Membranes. Biochemistry 1992, 31, 12416–12423. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.T.; Hung, W.C.; Chen, F.Y.; Huang, H.W. Many-body effect of antimicrobial peptides: On the correlation between lipid’s spontaneous curvature and pore formation. Biophys. J. 2005, 89, 4006–4016. [Google Scholar] [CrossRef] [PubMed]
- Henriques, S.T.; Melo, M.N.; Castanho, M.A.R.B. Cell-penetrating peptides and antimicrobial peptides: How different are they? Biochem. J. 2006, 399, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Nan, Y.H.; Park, I.S.; Hahm, K.S.; Shin, S.Y. Antimicrobial activity, bactericidal mechanism and LPS-neutralizing activity of the cell-penetrating peptide pVEC and its analogs. J. Pept. Sci. 2011, 17, 812–817. [Google Scholar] [CrossRef] [PubMed]
- Bahnsen, J.S.; Franzyk, H.; Sandberg-Schaal, A.; Nielsen, H.M. Antimicrobial and cell-penetrating properties of penetratin analogs: Effect of sequence and secondary structure. Biochim. Biophys. Acta Biomembr. 2013, 1828, 223–232. [Google Scholar] [CrossRef] [PubMed]
- Budagavi, D.P.; Chugh, A. Antibacterial properties of Latarcin 1 derived cell-penetrating peptides. Eur. J. Pharm. Sci. 2018, 115, 43–49. [Google Scholar] [CrossRef] [PubMed]
- Bobone, S.; Piazzon, A.; Orioni, B.; Pedersen, J.Z.; Nan, Y.H.; Hahm, K.S.; Shin, S.Y.; Stella, L. The thin line between cell-penetrating and antimicrobial peptides: The case of Pep-1 and Pep-1-K. J. Pept. Sci. 2011, 17, 335–341. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Plaza, J.G.; Morales-Nava, R.; Diener, C.; Schreiber, G.; Gonzalez, Z.D.; Ortiz, M.T.L.; Blake, I.O.; Pantoja, O.; Volkmer, R.; Klipp, E.; et al. Cell-penetrating peptides and cationic antibacterial peptides: Two sides of the same coin. J. Biol. Chem. 2014, 289, 14448–14457. [Google Scholar] [CrossRef] [PubMed]
- Nekhotiaeva, N.; Elmquist, A.; Rajarao, G.K.; Good, L.; Stockholm, N. Cell entry and antimicrobial properties of eukaryotic cell-penetrating peptides. FASEB J. 2003, 18, 394–396. [Google Scholar] [CrossRef] [PubMed]
- Rydberg, H.A.; Kunze, A.; Carlsson, N.; Altgärde, N.; Svedhem, S.; Nordén, B. Peptide-membrane interactions of arginine-tryptophan peptides probed using quartz crystal microbalance with dissipation monitoring. Eur. Biophys. J. 2014, 43, 241–253. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; He, K.; Ludtke, S.J.; Huang, H.W. X-ray diffraction study of lipid bilayer membranes interacting with amphiphilic helical peptides: Diphytanoyl phosphatidylcholine with alamethicin at low concentrations. Biophys. J. 1995, 68, 2361–2369. [Google Scholar] [CrossRef]
- Münster, C.; Spaar, A.; Bechinger, B.; Salditt, T. Magainin 2 in phospholipid bilayers: Peptide orientation and lipid chain ordering studied by X-ray diffraction. Biochim. Biophys. Acta Biomembr. 2002, 1562, 37–44. [Google Scholar] [CrossRef]
- Mishra, A.; Gordon, V.D.; Yang, L.; Coridan, R.; Wong, G.C.L. HIV TAT forms pores in membranes by inducing saddle-splay curvature: Potential role of bidentate hydrogen bonding. Angew. Chem. Int. Ed. Engl. 2008, 47, 2986–2989. [Google Scholar] [CrossRef] [PubMed]
- Lamazière, A.; Maniti, O.; Wolf, C.; Lambert, O.; Chassaing, G.; Trugnan, G.; Ayala Sanmartin, J. Lipid domain separation, bilayer thickening and pearling induced by the cell penetrating peptide penetratin. Biochim. Biophys. Acta Biomembr. 2010, 1798, 2223–2230. [Google Scholar] [CrossRef] [PubMed]
- He, K.; Ludtke, S.J.; Huang, H.W.; Worcester, D.L. Antimicrobial Peptide Pores in Membranes Detected by Neutron In-Plane Scattering. Biochemistry 1995, 34, 15614–15618. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Sa’adedin, F.; Deme, B.; Rao, P.; Bradshaw, J. Insertion of TAT peptide and perturbation of negatively charged model phospholipid bilayer revealed by neutron diffraction. Biochim. Biophys. Acta Biomembr. 2013, 1828, 1982–1988. [Google Scholar] [CrossRef] [PubMed]
- Cruz, J.; Mihailescu, M.; Wiedman, G.; Herman, K.; Searson, P.C.; Wimley, W.C.; Hristova, K. A membrane-translocating peptide penetrates into bilayers without significant bilayer perturbations. Biophys. J. 2013, 104, 2419–2428. [Google Scholar] [CrossRef] [PubMed]
- Shenkarev, Z.O.; Balandin, S.V.; Trunov, K.I.; Paramonov, A.S.; Sukhanov, S.V.; Barsukov, L.I.; Arseniev, A.S.; Ovchinnikova, T.V. Molecular mechanism of action of β-Hairpin antimicrobial peptide arenicin: Oligomeric structure in dodecylphosphocholine micelles and pore formation in planar lipid bilayers. Biochemistry 2011, 50, 6255–6265. [Google Scholar] [CrossRef] [PubMed]
- Usachev, K.S.; Efimov, S.V.; Kolosova, O.A.; Filippov, A.V.; Klochkov, V.V. High-resolution NMR structure of the antimicrobial peptide protegrin-2 in the presence of DPC micelles. J. Biomol. NMR 2015, 61, 227–234. [Google Scholar] [CrossRef] [PubMed]
- Afonin, S.; Kubyshkin, V.; Mykhailiuk, P.K.; Komarov, I.V.; Ulrich, A.S. Conformational Plasticity of the Cell-Penetrating Peptide SAP As Revealed by Solid-State 19F-NMR and Circular Dichroism Spectroscopies. J. Phys. Chem. B 2017, 121, 6479–6491. [Google Scholar] [CrossRef] [PubMed]
- Hsu, C.H.; Chen, C.; Jou, M.L.; Lee, A.Y.L.; Lin, Y.C.; Yu, Y.P.; Huang, W.T.; Wu, S.H. Structural and DNA-binding studies on the bovine antimicrobial peptide, indolicidin: Evidence for multiple conformations involved in binding to membranes and DNA. Nucleic Acids Res. 2005, 33, 4053–4064. [Google Scholar] [CrossRef] [PubMed]
- Avitabile, C.; D’Andrea, L.D.; Romanelli, A. Circular Dichroism studies on the interactions of antimicrobial peptides with bacterial cells. Sci. Rep. 2014, 4, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Deshayes, S.; Gerbal Chaloin, S.; Morris, M.C.; Aldrian Herrada, G.; Charnet, P.; Divita, G.; Heitz, F. On the mechanism of nonendosomial peptide-mediated cellular delivery of nucleic acids. Biochim. Biophys. Acta Biomembr. 2004, 1667, 141–147. [Google Scholar] [CrossRef] [PubMed]
- Crombez, L.; Aldrian Herrada, G.; Konate, K.; Nguyen, Q.N.; McMaster, G.K.; Brasseur, R.; Heitz, F.; Divita, G. A new potent secondary amphipathic cell-penetrating peptide for siRNA delivery into mammalian cells. Mol. Ther. 2009, 17, 95–103. [Google Scholar] [CrossRef] [PubMed]
- Bhunia, A.; Domadia, P.N.; Torres, J.; Hallock, K.J.; Ramamoorthy, A.; Bhattacharjya, S. NMR structure of pardaxin, a pore-forming antimicrobial peptide, in lipopolysaccharide micelles: Mechanism of outer membrane permeabilization. J. Biol. Chem. 2010, 285, 3883–3895. [Google Scholar] [CrossRef] [PubMed]
- Wieprecht, T.; Apostolov, O.; Beyermann, M.; Seelig, J. Membrane binding and pore formation of the antibacterial peptide PGLa: Thermodynamic and mechanistic aspects. Biochemistry 2000, 39, 442–452. [Google Scholar] [CrossRef] [PubMed]
- Québatte, G.; Kitas, E.; Seelig, J. riDOM, a cell penetrating peptide. Interaction with phospholipid bilayers. Biochim. Biophys. Acta Biomembr. 2014, 1838, 968–977. [Google Scholar] [CrossRef] [PubMed]
- Aguilera, T.A.; Olson, E.S.; Timmers, M.M.; Jiang, T.; Tsien, R.Y. Systemic in vivo distribution of activatable cell penetrating peptides is superior to that of cell penetrating peptides. Integr. Biol. 2009, 1, 371–381. [Google Scholar] [CrossRef] [PubMed]
- Prenner, E.J.; Lewis, R.N.A.H.; Kondejewski, L.H.; Hodges, R.S.; McElhaney, R.N. Differential scanning calorimetric study of the effect of the antimicrobial peptide gramicidin S on the thermotropic phase behavior of phosphatidylcholine, phosphatidylethanolamine and phosphatidylglycerol lipid bilayer membranes. Biochim. Biophys. Acta Biomembr. 1999, 1417, 211–223. [Google Scholar] [CrossRef]
- Wadhwani, P.; Epand, R.F.; Heidenreich, N.; Bürck, J.; Ulrich, A.S.; Epand, R.M. Membrane-active peptides and the clustering of anionic lipids. Biophys. J. 2012, 103, 265–274. [Google Scholar] [CrossRef] [PubMed]
- Meincken, M.; Holroyd, D.L.; Rautenbach, M. Atomic force microscopy study of the effect of antimicrobial peptides on the cell envelope of Escherichia coli. Antimicrob. Agents Chemother. 2005, 49, 4085–4092. [Google Scholar] [CrossRef] [PubMed]
- Fantner, G.E.; Barbero, R.J.; Gray, D.S.; Belcher, A.M. Kinetics of antimicrobial peptide activity measured on individual bacterial cells using high-speed atomic force microscopy. Nat. Nanotechnol. 2010, 5, 280–285. [Google Scholar] [CrossRef] [PubMed]
- Plénat, T.; Boichot, S.; Dosset, P.; Milhiet, P.E.; Le Grimellec, C. Coexistence of a two-states organization for a cell-penetrating peptide in lipid bilayer. Biophys. J. 2005, 89, 4300–4309. [Google Scholar] [CrossRef] [PubMed]
- Shaw, J.E.; Epand, R.F.; Hsu, J.C.Y.; Mo, G.C.H.; Epand, R.M.; Yip, C.M. Cationic peptide-induced remodelling of model membranes: Direct visualization by in situ atomic force microscopy. J. Struct. Biol. 2008, 162, 121–138. [Google Scholar] [CrossRef] [PubMed]
- Sychev, S.V.; Panteleev, P.V.; Ovchinnikova, T.V. Structural study of the β-hairpin marine antimicrobial peptide arenicin-2 in PC/PG lipid bilayers by fourier transform infrared spectroscopy. Russ. J. Bioorg. Chem. 2017, 43, 502–508. [Google Scholar] [CrossRef]
- Jackson, M.; Mantsch, H.H.; Spencer, J.H. Conformation of Magainin-2 and Related Peptides in Aqueous Solution and Membrane Environments Probed by Fourier Transform Infrared Spectroscopy. Biochemistry 1992, 31, 7289–7293. [Google Scholar] [CrossRef] [PubMed]
- Plénat, T.; Deshayes, S.; Boichot, S.; Milhiet, P.E.; Cole, R.B.; Heitz, F.; Le Grimellec, C. Interaction of primary amphipathic cell-penetrating peptides with phospholipid-supported monolayers. Langmuir 2004, 20, 9255–9261. [Google Scholar] [CrossRef] [PubMed]
- Maniti, O.; Alves, I.; Trugnan, G.; Ayala-Sanmartin, J. Distinct behaviour of the homeodomain derived cell penetrating peptide penetratin in interaction with different phospholipids. PLoS ONE 2010, 5. [Google Scholar] [CrossRef] [PubMed]
- Hall, K.; Lee, T.H.; Aguilar, M.I. The role of electrostatic interactions in the membrane binding of melittin. J. Mol. Recognit. 2011, 24, 108–118. [Google Scholar] [CrossRef] [PubMed]
- Brian Chia, C.S.; Gong, Y.; Bowie, J.H.; Zuegg, J.; Cooper, M.A. Membrane binding and perturbation studies of the antimicrobial peptides caerin, citropin, and maculatin. Biopolymers 2011, 96, 147–157. [Google Scholar] [CrossRef] [PubMed]
- Cascales, L.; Henriques, S.T.; Kerr, M.C.; Huang, Y.H.; Sweet, M.J.; Daly, N.L.; Craik, D.J. Identification and characterization of a new family of cell-penetrating peptides: Cyclic cell-penetrating peptides. J. Biol. Chem. 2011, 286, 36932–36943. [Google Scholar] [CrossRef] [PubMed]
- Anko, M.; Majhenc, J.; Kogej, K.; Sillard, R.; Langel, Ü.; Anderluh, G.; Zorko, M. Influence of stearyl and trifluoromethylquinoline modifications of the cell penetrating peptide TP10 on its interaction with a lipid membrane. Biochim. Biophys. Acta Biomembr. 2012, 1818, 915–924. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Chen, C.H.; Hu, D.; Ulmschneider, M.B.; Ulmschneider, J.P. Spontaneous formation of structurally diverse membrane channel architectures from a single antimicrobial peptide. Nat. Commun. 2016, 7, 13535. [Google Scholar] [CrossRef] [PubMed]
- Domingues, T.M.; Perez, K.R.; Miranda, A.; Riske, K.A. Comparative study of the mechanism of action of the antimicrobial peptide gomesin and its linear analogue: The role of the β-hairpin structure. Biochim. Biophys. Acta Biomembr. 2015, 1848, 2414–2421. [Google Scholar] [CrossRef] [PubMed]
- Regberg, J.; Vasconcelos, L.; Madani, F.; Langel, Ü.; Hällbrink, M. pH-responsive PepFect cell-penetrating peptides. Int. J. Pharm. 2016, 501, 32–38. [Google Scholar] [CrossRef] [PubMed]
- Cerrato, C.P.; Pirisinu, M.; Vlachos, E.N.; Langel, Ü. Novel cell-penetrating peptide targeting mitochondria. FASEB J. 2015, 29, 4589–4599. [Google Scholar] [CrossRef] [PubMed]
- Sochacki, K.A.; Barns, K.J.; Bucki, R.; Weisshaar, J.C. Real-time attack on single Escherichia coli cells by the human antimicrobial peptide LL-37. Proc. Natl. Acad. Sci. USA 2011, 108, E77–E81. [Google Scholar] [CrossRef] [PubMed]
- Den Hertog, A.L.; van Marle, J.; van Veen, H.A.; Van’t Hof, W.; Bolscher, J.G.M.; Veerman, E.C.I.; Nieuw Amerongen, A.V. Candidacidal effects of two antimicrobial peptides: histatin 5 causes small membrane defects, but LL-37 causes massive disruption of the cell membrane. Biochem. J. 2005, 388, 689–695. [Google Scholar] [CrossRef] [PubMed]
- Elmquist, A.; Langel, Ü. In vitro uptake and stability study of pVEC and its all-D analog. Biol. Chem. 2003, 384, 387–393. [Google Scholar] [CrossRef] [PubMed]
- Balzeau, J.; Peterson, A.; Eyer, J. The vimentin-tubulin binding site peptide (Vim-TBS.58-81) crosses the plasma membrane and enters the nuclei of human glioma cells. Int. J. Pharm. 2012, 423, 77–83. [Google Scholar] [CrossRef] [PubMed]
- Schlamadinger, D.E.; Wang, Y.; McCammon, J.A.; Kim, J.E. Spectroscopic and computational study of melittin, cecropin A, and the hybrid peptide CM15. J. Phys. Chem. B 2012, 116, 10600–10608. [Google Scholar] [CrossRef] [PubMed]
- Bocchinfuso, G.; Palleschi, A.; Orioni, B.; Grande, G.; Formaggio, F.; Toniolo, C.; Park, Y.; Hahm, K.S.; Stella, L. Different mechanisms of action of antimicrobial peptides: Insights from fluorescence spectroscopy experiments and molecular dynamics simulations. J. Pept. Sci. 2009, 15, 550–558. [Google Scholar] [CrossRef] [PubMed]
- Orioni, B.; Bocchinfuso, G.; Kim, J.Y.; Palleschi, A.; Grande, G.; Bobone, S.; Park, Y.; Kim, J.I.; Hahm, K.-S.; Stella, L. Membrane perturbation by the antimicrobial peptide PMAP-23: A fluorescence and molecular dynamics study. Biochim. Biophys. Acta Biomembr. 2009, 1788, 1523–1533. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.S.; Cha, H.J. High-throughput and facile assay of antimicrobial peptides using pH-controlled fluorescence resonance energy transfer. Antimicrob. Agents Chemother. 2006, 50, 3330–3335. [Google Scholar] [CrossRef] [PubMed]
- Yandek, L.E.; Pokorny, A.; Almeida, P.F.F. Small changes in the primary structure of transportan 10 alter the thermodynamics and kinetics of its interaction with phospholipid vesicles. Biochemistry 2008, 47, 3051–3060. [Google Scholar] [CrossRef] [PubMed]
- Henriques, S.T.; Castanho, M.A.R. Consequences of nonlytic membrane perturbation to the translocation of the cell penetrating peptide pep-1 in lipidic vesicles. Biochemistry 2004, 43, 9716–9724. [Google Scholar] [CrossRef] [PubMed]
- Han, M.; Mei, Y.; Khant, H.; Ludtke, S.J. Characterization of antibiotic peptide pores using cryo-EM and comparison to neutron scattering. Biophys. J. 2009, 97, 164–172. [Google Scholar] [CrossRef] [PubMed]
- Gordon-Grossman, M.; Zimmermann, H.; Wolf, S.G.; Shai, Y.; Goldfarb, D. Investigation of model membrane disruption mechanism by melittin using pulse electron paramagnetic resonance spectroscopy and cryogenic transmission electron microscopy. J. Phys. Chem. B 2012, 116, 179–188. [Google Scholar] [CrossRef] [PubMed]
- Freire, J.M.; Gaspar, D.; De La Torre, B.G.; Veiga, A.S.; Andreu, D.; Castanho, M.A.R.B. Monitoring antibacterial permeabilization in real time using time-resolved flow cytometry. Biochim. Biophys. Acta Biomembr. 2015, 1848, 554–560. [Google Scholar] [CrossRef] [PubMed]
- O’Brien-Simpson, N.M.; Pantarat, N.; Attard, T.J.; Walsh, K.A.; Reynolds, E.C. A rapid and quantitative flow cytometry method for the analysis of membrane disruptive antimicrobial activity. PLoS ONE 2016, 11. [Google Scholar] [CrossRef] [PubMed]
- Illien, F.; Rodriguez, N.; Amoura, M.; Joliot, A.; Pallerla, M.; Cribier, S.; Burlina, F.; Sagan, S. Quantitative fluorescence spectroscopy and flow cytometry analyses of cell-penetrating peptides internalization pathways: Optimization, pitfalls, comparison with mass spectrometry quantification. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef] [PubMed]
- Berman, H.M.; Westbrook, J.; Feng, Z.; Gilliland, G.; Bhat, T.N.; Weissig, H.; Shindyalov, I.N.; Bourne, P.E. The Protein Data Bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Perrin, B.S.; Tian, Y.; Fu, R.; Grant, C.V.; Chekmenev, E.Y.; Wieczorek, W.E.; Dao, A.E.; Hayden, R.M.; Burzynski, C.M.; Venable, R.M.; et al. High-resolution structures and orientations of antimicrobial peptides piscidin 1 and piscidin 3 in fluid bilayers reveal tilting, kinking, and bilayer immersion. J. Am. Chem. Soc. 2014, 136, 3491–3504. [Google Scholar] [CrossRef] [PubMed]
- Mani, R.; Tang, M.; Wu, X.; Buffy, J.J.; Waring, A.J.; Sherman, M.A.; Hong, M. Membrane-bound dimer structure of a β-hairpin antimicrobial peptide from rotational-echo double-resonance solid-state NMR. Biochemistry 2006, 45, 8341–8349. [Google Scholar] [CrossRef] [PubMed]
- Bera, S.; Kar, R.K.; Mondal, S.; Pahan, K.; Bhunia, A. Structural Elucidation of the Cell-Penetrating Penetratin Peptide in Model Membranes at the Atomic Level: Probing Hydrophobic Interactions in the Blood-Brain Barrier. Biochemistry 2016, 55, 4982–4996. [Google Scholar] [CrossRef] [PubMed]
- Grégoire, C.; Péloponèse, J.M.; Esquieu, D.; Opi, S.; Campbell, G.; Solomiac, M.; Lebrun, E.; Lebreton, J.; Loret, E.P. Homonuclear 1H-NMR assignment and structural characterization of human immunodeficiency virus type 1 Tat Mal protein. Biopolym. Biospectroscopy Sect. 2001, 62, 324–335. [Google Scholar] [CrossRef] [PubMed]
- Nicastro, G.; Franzoni, L.; De Chiara, C.; Mancin, A.C.; Giglio, J.R.; Spisni, A. Solution structure of crotamine, a Na+ channel affecting toxin from Crotalus durissus terrificus venom. Eur. J. Biochem. 2003, 270, 1969–1979. [Google Scholar] [CrossRef] [PubMed]
- Coronado, M.A.; Gabdulkhakov, A.; Georgieva, D.; Sankaran, B.; Murakami, M.T.; Arni, R.K.; Betzel, C. Structure of the polypeptide crotamine from the Brazilian rattlesnake Crotalus durissus terrificus. Acta Crystallogr. D. Biol. Crystallogr. 2013, 69, 1958–1964. [Google Scholar] [CrossRef] [PubMed]
- Terwilliger, T.C.; Eisenberg, D. The structure of melittin. I. Structure determination and partial refinement. J. Biol. Chem. 1982, 257, 6010–6015. [Google Scholar] [PubMed]
- Hill, C.; Yee, J.; Selsted, M.; Eisenberg, D. Crystal structure of defensin HNP-3, an amphiphilic dimer: Mechanisms of membrane permeabilization. Science 1991, 251, 1481–1485. [Google Scholar] [CrossRef] [PubMed]
- Latal, A.; Degovics, G.; Epand, R.F.; Epand, R.M.; Lohner, K. Structural aspects of the interaction of peptidyl-glycylleucine-carboxyamide, a highly potent antimicrobial peptide from frog skin, with lipids. Eur. J. Biochem. 1997, 248, 938–946. [Google Scholar] [CrossRef] [PubMed]
- Epand, R.M.; Vogel, H.J. Diversity of antimicrobial peptides and their mechanism of action. Biochim. Biophys. Acta 1999, 1462, 11–28. [Google Scholar] [CrossRef]
- Hristova, K.; Wimley, W.C.; Mishra, V.K.; Anantharamiah, G.M.; Segrest, J.P.; White, S.H. An amphipathic α-helix at a membrane interface: A structural study using a novel X-ray diffraction method. J. Mol. Biol. 1999, 290, 99–117. [Google Scholar] [CrossRef] [PubMed]
- Qian, S.; Wang, C.; Yang, L.; Huang, H.W. Structure of the Alamethicin Pore Reconstructed by X-ray Diffraction Analysis. Biophys. J. 2008, 94, 3512–3522. [Google Scholar] [CrossRef] [PubMed]
- Pan, J.; Tieleman, D.P.; Nagle, J.F.; Kučerka, N.; Tristram Nagle, S. Alamethicin in lipid bilayers: Combined use of X-ray scattering and MD simulations. Biochim. Biophys. Acta Biomembr. 2009, 1788, 1387–1397. [Google Scholar] [CrossRef] [PubMed]
- Su, C.J.; Wu, S.S.; Jeng, U.S.; Lee, M.T.; Su, A.C.; Liao, K.F.; Lin, W.Y.; Huang, Y.S.; Chen, C.Y. Peptide-induced bilayer thinning structure of unilamellar vesicles and the related binding behavior as revealed by X-ray scattering. Biochim. Biophys. Acta Biomembr. 2013, 1828, 528–534. [Google Scholar] [CrossRef] [PubMed]
- Neale, C.; Huang, K.; García, A.; Tristram-Nagle, S. Penetration of HIV-1 Tat47–57 into PC/PE Bilayers Assessed by MD Simulation and X-ray Scattering. Membranes 2015, 5, 473–494. [Google Scholar] [CrossRef] [PubMed]
- Akabori, K.; Huang, K.; Treece, B.W.; Jablin, M.S.; Maranville, B.; Woll, A.; Nagle, J.F.; Garcia, A.E.; Tristram-Nagle, S. HIV-1 Tat membrane interaction probed using X-ray and neutron scattering, CD spectroscopy and MD simulations. Biochim. Biophys. Acta 2015, 1838, 3078–3087. [Google Scholar] [CrossRef] [PubMed]
- Heberle, F.A.; Myles, D.A.A.; Katsaras, J. Biomembranes research using thermal and cold neutrons. Chem. Phys. Lipids 2015, 192, 41–50. [Google Scholar] [CrossRef] [PubMed]
- He, K.; Ludtke, S.J.; Worcester, D.L.; Huang, H.W. Neutron scattering in the plane of membranes: Structure of Alamethicin pores. Biophys. J. 1996, 70, 2659–2666. [Google Scholar] [CrossRef]
- Choi, D.; Moon, J.H.; Kim, H.; Sung, B.J.; Kim, M.W.; Tae, G.Y.; Satija, S.K.; Akgun, B.; Yu, C.J.; Lee, H.W.; et al. Insertion mechanism of cell-penetrating peptides into supported phospholipid membranes revealed by X-ray and neutron reflection. Soft Matter 2012, 8, 8294. [Google Scholar] [CrossRef]
- Chen, X.; Liu, S.; Deme, B.; Cristiglio, V.; Marquardt, D.; Weller, R.; Rao, P.; Wang, Y.; Bradshaw, J. Efficient internalization of TAT peptide in zwitterionic DOPC phospholipid membrane revealed by neutron diffraction. Biochim. Biophys. Acta Biomembr. 2017, 1859, 910–916. [Google Scholar] [CrossRef] [PubMed]
- Luchini, A.; Delhom, R.; Demé, B.; Laux, V.; Moulin, M.; Haertlein, M.; Pichlerc, H.; Strohmeier, G.A.; Wacklinf, H.; Fragneto, G. The impact of deuteration on natural and synthetic lipids: A neutron diffraction study. Colloids Surf. B Biointerfaces 2018, 168, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Wider, G. Structure determination of biological macromolecules in solution using nuclear magnetic resonance spectroscopy. Biotechniques 2000, 29, 1278–1294. [Google Scholar] [PubMed]
- Naito, A.; Kawamura, I.; Javkhlantugs, N. Recent Solid-State NMR Studies of Membrane-Bound Peptides and Proteins. In Annual Reports on NMR Spectroscopy, 1st ed.; Elsevier: New York, NY, USA, 2015; Volume 86, pp. 333–411. [Google Scholar]
- Bechinger, B.; Salnikov, E.S. The membrane interactions of antimicrobial peptides revealed by solid-state NMR spectroscopy. Chem. Phys. Lipids 2012, 165, 282–301. [Google Scholar] [CrossRef] [PubMed]
- Jeong, J.H.; Kim, J.S.; Choi, S.S.; Kim, Y. NMR Structural Studies of Antimicrobial Peptides: LPcin Analogs. Biophys. J. 2016, 110, 423–430. [Google Scholar] [CrossRef] [PubMed]
- Damberg, P.; Jarvet, J.; Gräslund, A.J. Micellar systems as solvents in peptide and protein structure determination. Methods Enzymol. 2001, 339, 271–285. [Google Scholar] [PubMed]
- Dürr, U.H.N.; Gildenberg, M.; Ramamoorthy, A. The magic of bicelles lights up membrane protein structure. Chem. Rev. 2012, 112, 6054–6074. [Google Scholar] [CrossRef] [PubMed]
- Trexler, A.J.; Rhoades, E. α-Synuclein binds large unilamellar vesicles as an extended helix. Biophysics 2010, 48, 2304–2306. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, N.; Mishra, A.; Lai, G.H.; Wong, G.C.L. Arginine-rich cell-penetrating peptides. FEBS Lett. 2010, 584, 1806–1813. [Google Scholar] [CrossRef] [PubMed]
- Mishra, A.; Lai, G.H.; Schmidt, N.W.; Sun, V.Z.; Rodriguez, A.R.; Tong, R.; Tang, L.; Cheng, J.; Deming, T.J.; Kamei, D.T.; et al. Translocation of HIV TAT peptide and analogues induced by multiplexed membrane and cytoskeletal interactions. Proc. Natl. Acad. Sci. USA 2011, 108, 16883–16888. [Google Scholar] [CrossRef] [PubMed]
- Naito, A.; Matsumori, N.; Ramamoorthy, A. Dynamic membrane interactions of antibacterial and antifungal biomolecules, and amyloid peptides, revealed by solid-state NMR spectroscopy. Biochim. Biophys. Acta Gen. Subj. 2018, 1862, 307–323. [Google Scholar] [CrossRef] [PubMed]
- Ramamoorthy, A. Beyond NMR Spectra of Antimicrobial Peptides: Dynamical Images at Atomic Resolution and Functional Insights. Solid State Nucl. Magn. Reson. 2010, 35, 201–207. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Liu, L.; Maltseva, S.; Lorigan, G.A.; Dabney-Smith, C. Investigating the interaction between peptides of the amphipathic helix of Hcf106 and the phospholipid bilayer by solid-state NMR spectroscopy. Biochim. Biophys. Acta (BBA) Biomembr. 2014, 1838, 1301–1315. [Google Scholar] [CrossRef] [PubMed]
- Watts, A. Solid-state NMR approaches for studying the interaction of peptides and proteins with membranes. Biochim. Biophys. Acta Rev. Biomembr. 1998, 1376, 297–318. [Google Scholar] [CrossRef]
- Booth, V.; Warschawski, D.E.; Santisteban, N.P.; Laadhari, M.; Marcotte, I. Recent progress on the application of 2H solid-state NMR to probe the interaction of antimicrobial peptides with intact bacteria. BBA Proteins Proteom. 2017, 1865, 1500–1511. [Google Scholar] [CrossRef] [PubMed]
- Auger, M. Biological Membrane Structure by Solid-State NMR. Curr. Opin. Mol. Biol. 2000, 4, 119–124. [Google Scholar]
- Fillion, M.; Noël, M.; Lorin, A.; Voyer, N.; Auger, M. Investigation of the mechanism of action of novel amphipathic peptides: Insights from solid-state NMR studies of oriented lipid bilayers. Biochim. Biophys. Acta Biomembr. 2014, 1838, 2173–2179. [Google Scholar] [CrossRef] [PubMed]
- Bechinger, B.; Zasloff, M.; Opella, S.J. Structure and Dynamics of the Antibiotic Peptide PGLa in Membranes by Solution and Solid-State Nuclear Magnetic Resonance Spectroscopy. Protein Sci. 1993, 2, 2077–2089. [Google Scholar] [CrossRef] [PubMed]
- Gopal, R.; Park, J.S.; Seo, C.H.; Park, Y. Applications of circular dichroism for structural analysis of gelatin and antimicrobial peptides. Int. J. Mol. Sci. 2012, 13, 3229–3244. [Google Scholar] [CrossRef] [PubMed]
- Bürck, J.; Wadhwani, P.; Fanghänel, S.; Ulrich, A.S. Oriented Circular Dichroism: A Method to Characterize Membrane-Active Peptides in Oriented Lipid Bilayers. Acc. Chem. Res. 2016, 49, 184–192. [Google Scholar] [CrossRef] [PubMed]
- Lee, M. Advances in Physics: X Biophysical characterization of peptide—Membrane interactions. Adv. Phys. X 2018, 3, 144–163. [Google Scholar]
- Moffitt, W. Optical Rotatory Dispersion of Helical Polymers. J. Chem. Phys. 1956, 25, 467–478. [Google Scholar] [CrossRef]
- Wu, Y.; Huang, H.W.; Olah, G.A. Method of Oriented Circular-Dichroism. Biophys. J. 1990, 57, 797–806. [Google Scholar] [CrossRef]
- Lee, M.T.; Sun, T.L.; Hung, W.C.; Huang, H.W. Process of inducing pores in membranes by melittin. Proc. Natl. Acad. Sci. USA 2013, 110, 14243–14248. [Google Scholar] [CrossRef] [PubMed]
- Kocourková, L.; Novotná, P.; Šťovíčková-Habartová, L.; Čujová, S.; Čeřovský, V.; Urbanová, M.; Setnička, V. Vibrational and electronic circular dichroism as powerful tools for the conformational analysis of cationic antimicrobial peptides. Monatshefte Chem. Chem. Mon. 2016, 147, 1439–1445. [Google Scholar] [CrossRef]
- Galdiero, S.; Falanga, A.; Cantisani, M.; Vitiello, M.; Morelli, G.; Galdiero, M. Peptide-Lipid Interactions: Experiments and Applications. Int. J. Mol. Sci. 2013, 14, 18758–18789. [Google Scholar] [CrossRef] [PubMed]
- Domingues, T.M.; Mattei, B.; Seelig, J.; Perez, K.R.; Miranda, A.; Riske, K.A. Interaction of the antimicrobial peptide gomesin with model membranes: A calorimetric study. Langmuir 2013, 29, 8609–8618. [Google Scholar] [CrossRef] [PubMed]
- Doyle, M.L. Characterization of binding interactions by isothermal titration calorimetry. Curr. Opin. Biotechnol. 1997, 8, 31–35. [Google Scholar] [CrossRef]
- Rajarathnam, K.; Rösgen, J. Isothermal titration calorimetry of membrane proteins—Progress and challenges. Biochim. Biophys. Acta Biomembr. 2014, 1838, 69–77. [Google Scholar] [CrossRef] [PubMed]
- Copolovici, D.M.; Langel, K.; Eriste, E.; Langel, Ü. Cell-penetrating peptides: Design, synthesis, and applications. ACS Nano 2014, 8, 1972–1994. [Google Scholar] [CrossRef] [PubMed]
- Kabiri, M.; Unsworth, L.D. Application of isothermal titration calorimetry for characterizing thermodynamic parameters of biomolecular interactions: Peptide self-assembly and protein adsorption case studies. Biomacromolecules 2014, 15, 3463–3473. [Google Scholar] [CrossRef] [PubMed]
- Binder, H.; Lindblom, G. Charge-dependent translocation of the Trojan peptide penetratin across lipid membranes. Biophys. J. 2003, 85, 982–995. [Google Scholar] [CrossRef]
- Abraham, T.; Lewis, R.N.; Hodges, R.S.; McElhaney, R.N. Isothermal titration calorimetry studies of the binding of the antimicrobial peptide gramicidin S to phospholipid bilayer membranes. Biochemistry 2005, 44, 11279–11285. [Google Scholar] [CrossRef] [PubMed]
- Mattei, B.; Miranda, A.; Perez, K.R.; Riske, K.A. Structure-Activity relationship of the antimicrobial peptide gomesin: The role of peptide hydrophobicity in its interaction with model membranes. Langmuir 2014, 30, 3513–3521. [Google Scholar] [CrossRef] [PubMed]
- Wieprecht, T.; Beyermann, M.; Seelig, J. Binding of antibacterial magainin peptides to electrically neutral membranes: Thermodynamics and structure. Biochemistry 1999, 38, 10377–10387. [Google Scholar] [CrossRef] [PubMed]
- Saravanan, R.; Li, X.; Lim, K.; Mohanram, H.; Peng, L.; Mishra, B.; Basu, A.; Lee, J.M.; Bhattacharjya, S.; Leong, S.S.J. Design of short membrane selective antimicrobial peptides containing tryptophan and arginine residues for improved activity, salt-resistance, and biocompatibility. Biotechnol. Bioeng. 2014, 111, 37–49. [Google Scholar] [CrossRef] [PubMed]
- Lohner, K.; Prenner, E.J. Differential scanning calorimetry and X-ray diffraction studies of the specificity of the interaction of antimicrobial peptides with membrane-mimetic systems. Biochim. Biophys. Acta Biomembr. 1999, 1462, 141–156. [Google Scholar] [CrossRef]
- Arias, M.; Prenner, E.J.; Vogel, H.J. Calorimetry Methods to Study Membrane Interactions and Perturbations Induced by Antimicrobial Host Defense Peptides. In Antimicrobial Peptides. Methods in Molecular Biology; Hansen, P., Ed.; Humana Press: New York, NY, USA, 2017; pp. 119–140. [Google Scholar]
- Joanne, P.; Galanth, C.; Goasdoué, N.; Nicolas, P.; Sagan, S.; Lavielle, S.; Chassaing, G.; El Amri, C.; Alves, I.D. Lipid reorganization induced by membrane-active peptides probed using differential scanning calorimetry. Biochim. Biophys. Acta Biomembr. 2009, 1788, 1772–1781. [Google Scholar] [CrossRef] [PubMed]
- Oreopoulos, J.; Epand, R.F.; Epand, R.M.; Yip, C.M. Peptide-induced domain formation in supported lipid bilayers: Direct evidence by combined atomic force and polarized total internal reflection fluorescence microscopy. Biophys. J. 2010, 98, 815–823. [Google Scholar] [CrossRef] [PubMed]
- Mularski, A.; Wilksch, J.J.; Wang, H.; Hossain, M.A.; Wade, J.D.; Separovic, F.; Strugnell, R.A.; Gee, M.L. Atomic Force Microscopy Reveals the Mechanobiology of Lytic Peptide Action on Bacteria. Langmuir 2015, 31, 6164–6171. [Google Scholar] [CrossRef] [PubMed]
- Tamm, L.K.; Tatulian, S. Infrared spectroscopy of proteins and peptides in lipid bilayers. Q. Rev. Biophys. 1997, 30, 365–429. [Google Scholar] [CrossRef] [PubMed]
- Haris, P.I. Probing protein-protein interaction in biomembranes using Fourier transform infrared spectroscopy. Biochim. Biophys. Acta Biomembr. 2013, 1828, 2265–2271. [Google Scholar] [CrossRef] [PubMed]
- Coe, J.; Kupitz, C.; Basu, S.; Conrad, C.E.; Roy-Chowdhury, S.; Fromme, R.; Fromme, P. Crystallization of photosystem II for time-resolved structural studies using an X-ray free electron laser. Methods Enzymol. 2015, 557, 459–482. [Google Scholar] [PubMed]
- Lusvarghi, S.; Ghirlando, R.; Davison, J.R.; Bewley, C.A. Chemical and Biophysical Approaches for Complete Characterization of Lectin–Carbohydrate Interactions. Methods Enzymol. 2018, 598, 3–35. [Google Scholar] [PubMed]
- Stetefeld, J.; McKenna, S.A.; Patel, T.R. Dynamic light scattering: A practical guide and applications in biomedical sciences. Biophys. Rev. 2016, 8, 409–427. [Google Scholar] [CrossRef] [PubMed]
- Yennawar, N.H.; Fecko, J.A.; Showalter, S.A.; Bevilacqua, P.C. A High-Throughput Biological Calorimetry Core: Steps to Startup, Run, and Maintain a Multiuser Facility. Methods Enzymol. 2016, 567, 435–460. [Google Scholar] [PubMed]
- Domingues, M.M.; Santiago, P.S.; Castanho, M.A.R.B.; Santos, N.C. What can light scattering spectroscopy do for membrane-active peptide studies? J. Pept. Sci. 2008, 14, 394–400. [Google Scholar] [CrossRef] [PubMed]
- Ross Hallett, F. Particle size analysis by dynamic light scattering. Food Res. Int. 1994, 27, 195–198. [Google Scholar] [CrossRef]
- Moniruzzaman, M.; Islam, M.Z.; Sharmin, S.; Dohra, H.; Yamazaki, M. Entry of a Six-Residue Antimicrobial Peptide Derived from Lactoferricin B into Single Vesicles and Escherichia coli Cells without Damaging their Membranes. Biochemistry 2017, 56, 4419–4431. [Google Scholar] [CrossRef] [PubMed]
- Gee, M.L.; Burton, M.; Grevis-James, A.; Hossain, M.A.; McArthur, S.; Palombo, E.A.; Wade, J.D.; Clayton, A.H.A. Imaging the action of antimicrobial peptides on living bacterial cells. Sci. Rep. 2013, 3, 1557. [Google Scholar] [CrossRef] [PubMed]
- Hall, K.; Mozsolits, H.; Aguilar, M.I. Surface plasmon resonance analysis of antimicrobial peptide-membrane interactions: Affinity and mechanism of action. Lett. Pept. Sci. 2003, 10, 475–485. [Google Scholar] [CrossRef]
- Niu, L.; Wohland, T.; Knoll, W.; Köper, I. Interaction of a synthetic antimicrobial peptide with a model bilayer platform mimicking bacterial membranes. Biointerphases 2017, 12, 04E404. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Peinado, C.; Dias, S.A.; Domingues, M.M.; Benfield, A.H.; Freire, J.M.; Rádis-Baptista, G.; Gaspar, D.; Castanho, M.A.R.B.; Craik, D.J.; Henriques, S.T.; et al. Mechanisms of bacterial membrane permeabilization by crotalicidin (Ctn) and its fragment Ctn(15–34), antimicrobial peptides from rattlesnake venom. J. Biol. Chem. 2018, 293, 1536–1549. [Google Scholar] [CrossRef] [PubMed]
- Signorelli, S.; Santini, S.; Yamada, T.; Bizzarri, A.R.; Beattie, C.W.; Cannistraro, S. Binding of Amphipathic Cell Penetrating Peptide p28 to Wild Type and Mutated p53 as studied by Raman, Atomic Force and Surface Plasmon Resonance spectroscopies. Biochim. Biophys. Acta Gen. Subj. 2017, 1861, 910–921. [Google Scholar] [CrossRef] [PubMed]
- Deslouches, B.; Hasek, M.L.; Craigo, J.K.; Steckbeck, J.D.; Montelaro, R.C. Comparative functional properties of engineered cationic antimicrobial peptides consisting exclusively of tryptophan and either lysine or arginine. J. Med. Microbiol. 2016, 65, 554–565. [Google Scholar] [CrossRef] [PubMed]
- Dubochet, J.; Chang, J.J.; Freeman, R.; Lapault, J.; McDowall, A.W. Frozen Aquaous Suspensions. Ultramicroscopy 1982, 10, 55–62. [Google Scholar] [CrossRef]
- Bonomi, M.; Pellarin, R.; Vendruscolo, M. Simultaneous Determination of Protein Structure and Dynamics Using Cryo-Electron Microscopy. Biophys. J. 2018, 114, 1604–1613. [Google Scholar] [CrossRef] [PubMed]
- Bai, X.C.; McMullan, G.; Scheres, S.H.W. How cryo-EM is revolutionizing structural biology. Trends Biochem. Sci. 2015, 40, 49–57. [Google Scholar] [CrossRef] [PubMed]
- Nogales, E.; Scheres, S.H.W. Cryo-EM: A Unique Tool for the Visualization of Macromolecular Complexity. Mol. Cell 2016, 58, 677–689. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y. Single-particle cryo-EM at crystallographic resolution. Cell 2015, 161, 450–457. [Google Scholar] [CrossRef] [PubMed]
- Milne, J.L.S.; Borgnia, M.J.; Bartesaghi, A.; Tran, E.E.H.; Earl, L.A.; Schauder, D.M.; Lengyel, J.; Pierson, J.; Patwardhan, A.; Subramaniam, S. Cryo-electron microscopy: A primer for the nonmicroscopist. FEBS J. 2014, 280, 28–45. [Google Scholar] [CrossRef] [PubMed]
- El Jastimi, R.; Edwards, K.; Lafleur, M. Characterization of permeability and morphological perturbations induced by nisin on phosphatidylcholine membranes. Biophys. J. 1999, 77, 842–852. [Google Scholar] [CrossRef]
- Lambert, O.; Cavusoglu, N.; Gallay, J.; Vincent, M.; Rigaud, J.L.; Henry, J.P.; Ayala-Sanmartin, J. Novel Organization and Properties of Annexin 2-Membrane Complexes. J. Biol. Chem. 2004, 279, 10872–10882. [Google Scholar] [CrossRef] [PubMed]
- Boge, L.; Bysell, H.; Ringstad, L.; Wennman, D.; Umerska, A.; Cassisa, V.; Eriksson, J.; Joly-Guillou, M.L.; Edwards, K.; Andersson, M. Lipid-Based Liquid Crystals as Carriers for Antimicrobial Peptides: Phase Behavior and Antimicrobial Effect. Langmuir 2016, 32, 4217–4228. [Google Scholar] [CrossRef] [PubMed]
- Han, M.; Mei, Y.; Khant, H.; Ludtke, S.J. Pore formation by antimicrobial peptide, magainin 2, in phospholipd vesicles detected and visualized by cryo-electron microscopy. Microsc. Microanal. 2008, 14, 1610–1611. [Google Scholar] [CrossRef]
- Florin, T.; Maracci, C.; Graf, M.; Karki, P.; Klepacki, D.; Berninghausen, O.; Beckmann, R.; Vázquez-Laslop, N.; Wilson, D.N.; Rodnina, M.V.; et al. An antimicrobial peptide that inhibits translation by trapping release factors on the ribosome an antimicrobial peptide that inhibits translation by trapping release factors on the ribosome. Nat. Struct. Mol. 2017, 24, 752–757. [Google Scholar] [CrossRef] [PubMed]
- Cardoso, M.H.; Oshiro, K.G.; Rezende, S.B.; Cândido, E.S.; Franco, O.L. The Structure/Function Relationship in Antimicrobial Peptides: What Can We Obtain from Structural Data? In Advances in Protein Chemistry and Structural Biology; Donev, R., Ed.; Elsevier: New York, NY, USA, 2018; pp. 359–384. [Google Scholar]
- Matos, P.M.; Gonçalves, S.; Santos, N.C. Interaction of peptides with biomembranes assessed by potential-sensitive fluorescent probes. J. Pept. Sci. 2008, 14, 1084–1095. [Google Scholar] [CrossRef] [PubMed]
- Tiriveedhi, V.; Butko, P. A fluorescence spectroscopy study on the interactions of the TAT-PTD peptide with model lipid membranes. Biochemistry 2007, 46, 3888–3895. [Google Scholar] [CrossRef] [PubMed]
- Imura, Y.; Choda, N.; Matsuzaki, K. Magainin 2 in action: Distinct modes of membrane permeabilization in living bacterial and mammalian cells. Biophys. J. 2008, 95, 5757–5765. [Google Scholar] [CrossRef] [PubMed]
- Zamotaiev, O.M.; Postupalenko, V.Y.; Shvadchak, V.V.; Pivovarenko, V.G.; Klymchenko, A.S.; Mély, Y. Monitoring penetratin interactions with lipid membranes and cell internalization using a new hydration-sensitive fluorescent probe. Org. Biomol. Chem. 2014, 12, 7036–7044. [Google Scholar] [CrossRef] [PubMed]
- Deleu, M.; Crowet, J.M.; Nasir, M.N.; Lins, L. Complementary biophysical tools to investigate lipid specificity in the interaction between bioactive molecules and the plasma membrane: A review. Biochim. Biophys. Acta Biomembr. 2014, 1838, 3171–3190. [Google Scholar] [CrossRef] [PubMed]
- Munishkina, L.A.; Fink, A.L. Fluorescence as a method to reveal structures and membrane-interactions of amyloidogenic proteins. Biochim. Biophys. Acta Biomembr. 2007, 1768, 1862–1885. [Google Scholar] [CrossRef] [PubMed]
- Rezgui, R.; Blumer, K.; Yeoh-Tan, G.; Trexler, A.J.; Magzoub, M. Precise quantification of cellular uptake of cell-penetrating peptides using fluorescence-activated cell sorting and fluorescence correlation spectroscopy. Biochim. Biophys. Acta Biomembr. 2016, 1858, 1499–1506. [Google Scholar] [CrossRef] [PubMed]
- Eriste, E.; Kurrikoff, K.; Suhorutšenko, J.; Oskolkov, N.; Copolovici, D.M.; Jones, S.; Laakkonen, P.; Howl, J.; Langel, Ü. Peptide-based glioma-targeted drug delivery vector gHoPe2. Bioconjug. Chem. 2013, 24, 305–313. [Google Scholar] [CrossRef] [PubMed]
- Hedegaard, S.F.; Derbas, M.S.; Lind, T.K.; Kasimova, M.R.; Christensen, M.V.; Michaelsen, M.H.; Campbell, R.A.; Jorgensen, L.; Franzyk, H.; Cárdenas, M.; et al. Fluorophore labeling of a cell-penetrating peptide significantly alters the mode and degree of biomembrane interaction. Sci. Rep. 2018, 8. [Google Scholar] [CrossRef] [PubMed]
- Brown, W.; Wittner, C. Flow cytometry: Principles and clinical applications in hematology. Clin. Chem. 2000, 46, 1221–1229. [Google Scholar] [PubMed]
- Welsh, J.A.; Holloway, J.A.; Wilkinson, J.S.; Englyst, N.A. Extracellular Vesicle Flow Cytometry Analysis and Standardization. Front. Cell Dev. Biol. 2017, 5, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Adan, A.; Alizada, G.; Kiraz, Y.; Baran, Y.; Nalbant, A. Flow cytometry: Basic principles and applications. Crit. Rev. Biotechnol. 2017, 37, 163–176. [Google Scholar] [CrossRef] [PubMed]
- Léonard, L.; Chibane, L.B.; Bouhedda, B.O.; Degraeve, P.; Oulahal, N. Recent advances on multi-parameter flow cytometry to characterize antimicrobial treatments. Front. Microbiol. 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Hong, J.; Guan, W.; Jin, G.; Zhao, H.; Jiang, X.; Dai, J. Mechanism of tachyplesin I injury to bacterial membranes and intracellular enzymes, determined by laser confocal scanning microscopy and flow cytometry. Microbiol. Res. 2015, 170, 69–77. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.; Hwang, J.S.; Lee, J.; Kim, J.I.; Lee, D.G. Scolopendin 2, a cationic antimicrobial peptide from centipede, and its membrane-active mechanism. Biochim. Biophys. Acta Biomembr. 2015, 1848, 634–642. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; O’Brien-Simpson, N.M.; Tailhades, J.; Pantarat, N.; Dawson, R.M.; Otvos, L.; Reynolds, E.C.; Separovic, F.; Hossain, M.A.; Wade, J.D. Multimerization of a Proline-Rich Antimicrobial Peptide, Chex-Arg20, Alters Its Mechanism of Interaction with the Escherichia coli Membrane. Chem. Biol. 2015, 22, 1250–1258. [Google Scholar] [CrossRef] [PubMed]
- Barteneva, N.S.; Fasler-Kan, E.; Vorobjev, I.A. Imaging Flow Cytometry: Coping with Heterogeneity in Biological Systems. J. Histochem. Cytochem. 2012, 60, 723–733. [Google Scholar] [CrossRef] [PubMed]
- Yason, J.A.; Ajjampur, S.S.R.; Tan, K.S.W. Blastocystis isolate B exhibits multiple modes of resistance against antimicrobial peptide LL-37. Infect. Immun. 2016, 84, 2220–2232. [Google Scholar] [CrossRef] [PubMed]
- Skovbakke, S.L.; Heegaard, P.M.H.; Larsen, C.J.; Franzyk, H.; Forsman, H.; Dahlgren, C. The proteolytically stable peptidomimetic Pam-(Lys-βNSpe)6-NH2 selectively inhibits human neutrophil activation via formyl peptide receptor 2. Biochem. Pharmacol. 2015, 93, 182–195. [Google Scholar] [CrossRef] [PubMed]
- Sekiya, Y.; Sakashita, S.; Shimizu, K.; Usui, K.; Kawano, R. Channel current analysis estimates the pore-formation and the penetration of transmembrane peptides. Analyst 2018, 143. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, H.; Kawano, R. Channel Current Analysis for Pore-forming Properties of an Antimicrobial Peptide, Magainin 1, Using the Droplet Contact Method. Anal. Sci. 2016, 32, 57–60. [Google Scholar] [CrossRef] [PubMed]
- Valko, K.L.; Ivanova-Berndt, G.; Beswick, P.; Kindey, M.; Ko, D. Application of biomimetic HPLC to estimate lipophilicity, protein and phospholipid binding of potential peptide therapeutics. ADMET DMPK 2018, 6, 162. [Google Scholar] [CrossRef]
- Yang, Z.; Choi, H.; Weisshaar, J.C. Melittin-Induced Permeabilization, Re-sealing, and Re-permeabilization of E. coli Membranes. Biophys. J. 2018, 114, 368–379. [Google Scholar] [CrossRef] [PubMed]
- Ulmschneider, J.P.; Ulmschneider, M.B. Molecular Dynamics Simulations Are Redefining Our View of Peptides Interacting with Biological Membranes. Acc. Chem. Res. 2018, 51, 1106–1116. [Google Scholar] [CrossRef] [PubMed]
- Jeschke, G. Ensemble models of proteins and protein domains based on distance distribution restraints. Proteins Struct. Funct. Bioinform. 2016, 84, 544–560. [Google Scholar] [CrossRef] [PubMed]
- Kabelka, I.; Vácha, R. Optimal conditions for opening of membrane pore by amphiphilic peptides. J. Chem. Phys. 2015, 143, 243115. [Google Scholar] [CrossRef] [PubMed]
- Alaybeyoglu, B.; Akbulut Sariyar, B.; Ozkirimli, E. pVEC hydrophobic N-Terminus is critical for antibacterial activity. J. Pept. Sci. 2018, 24, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Gumbart, J.C.; Ulmschneider, M.B.; Hazel, A.; White, S.H.; Ulmschneider, J.P. Computed Free Energies of Peptide Insertion into Bilayers are Independent of Computational Method. J. Membr. Biol. 2018, 112, 10512–10528. [Google Scholar]
- Hu, Y.; Patel, S. Thermodynamics of cell-penetrating HIV1 TAT peptide insertion into PC/PS/CHOL model bilayers through transmembrane pores: The roles of cholesterol and anionic lipids. Soft Matter 2016, 12, 6716–6727. [Google Scholar] [CrossRef] [PubMed]
- Nelson, N.; Schwartz, D.K. Single-Molecule Resolution of Antimicrobial Peptide Interactions with Supported Lipid A Bilayers. Biophys. J. 2018, 114, 2606–2616. [Google Scholar] [CrossRef] [PubMed]
- Hong, C.; Tieleman, D.P.; Wang, Y. Microsecond Molecular Dynamics Simulations of Lipid Mixing. ACS Publ. 2014, 30, 11993–12001. [Google Scholar] [CrossRef] [PubMed]
- Marrink, S.J.; Risselada, H.J.; Yefimov, S.; Tieleman, D.P.; de Vries, A.H. The MARTINI force field: Coarse grained model for biomolecular simulations. J. Phys. Chem. B 2007, 111, 7812–7824. [Google Scholar] [CrossRef] [PubMed]
- Balatti, G.E.; Ambroggio, E.E.; Fidelio, G.D.; Martini, M.F.; Pickholz, M. Differential Interaction of Antimicrobial Peptides with Lipid Structures Studied by Coarse-Grained Molecular Dynamics Simulations. Molecules 2017, 22, 1775. [Google Scholar] [CrossRef] [PubMed]
- Lipkin, R.; Lazaridis, T. Computational studies of peptide-induced membrane pore formation. Philos. Trans. R. Soc. B Biol. Sci. 2017, 372, 20160213–20160219. [Google Scholar] [CrossRef] [PubMed]
- Shearer, J.; Khalid, S. Communication between the leaflets of asymmetric membranes revealed from coarse-grain molecular dynamics simulations. Sci. Rep. 2018, 8, 1805. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Hong, G.; Johnson, G.R.; Pachter, R.; Cheung, M.S. Biophysical properties of membrane-active peptides based on micelle modeling: A case study of cell-penetrating and antimicrobial peptides. J. Phys. Chem. B 2010, 114, 13726–13735. [Google Scholar] [CrossRef] [PubMed]
- Herce, H.D.; Garcia, A.E. Molecular dynamics simulations suggest a mechanism for translocation of the HIV-1 TAT peptide across lipid membranes. Proc. Natl. Acad. Sci. USA 2007, 104, 20805–20810. [Google Scholar] [CrossRef] [PubMed]
- Pimthon, J.; Willumeit, R.; Lendlein, A.; Hofmann, D. Membrane association and selectivity of the antimicrobial peptide NK-2: A molecular dynamics simulation study. J. Pept. Sci. 2009, 15, 654–667. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Liu, S.; Lakshminarayanan, R.; Bai, Y.; Pervushin, K.; Verma, C.; Beuerman, R.W. Molecular simulations suggest how a branched antimicrobial peptide perturbs a bacterial membrane and enhances permeability. Biochim. Biophys. Acta Biomembr. 2013, 1828, 1112–1121. [Google Scholar] [CrossRef] [PubMed]
- Islami, M.; Mehrnejad, F.; Doustdar, F.; Alimohammadi, M.; Khadem-Maaref, M.; Mir-Derikvand, M.; Taghdir, M. Study of orientation and penetration of LAH4 into lipid bilayer membranes: PH and composition dependence. Chem. Biol. Drug Des. 2014, 84, 242–252. [Google Scholar] [CrossRef] [PubMed]
- Sikorska, E.; Dawgul, M.; Greber, K.; Iłowska, E.; Pogorzelska, A.; Kamysz, W. Self-assembly and interactions of short antimicrobial cationic lipopeptides with membrane lipids: ITC, FTIR and molecular dynamics studies. Biochim. Biophys. Acta Biomembr. 2014, 1838, 2625–2634. [Google Scholar] [CrossRef] [PubMed]
- Tsai, C.W.; Hsu, N.Y.; Wang, C.H.; Lu, C.Y.; Chang, Y.; Tsai, H.H.G.; Ruaan, R.C. Coupling Molecular Dynamics Simulations with Experiments for the Rational Design of Indolicidin-Analogous Antimicrobial Peptides. J. Mol. Biol. 2009, 392, 837–854. [Google Scholar] [CrossRef] [PubMed]
- Ulmschneider, M.B.; Ulmschneider, J.P.; Schiller, N.; Wallace, B.A.; von Heijne, G.; White, S.H. Spontaneous transmembrane helix insertion thermodynamically mimics translocon-guided insertion. Nat. Commun. 2014, 5, 317. [Google Scholar] [CrossRef] [PubMed]
- Saini, A.; Jaswal, R.R.; Negi, R.; Nandel, F.S. Insights on the structural characteristics of Vim-TBS (58–81) peptide for future applications as a cell penetrating peptide. Biosci. Trends 2013, 7, 209–220. [Google Scholar] [CrossRef] [PubMed]
- Dunkin, C.M.; Pokorny, A.; Almeida, P.F.; Lee, H.-S. Molecular Dynamics Studies of Transportan 10 (Tp10) Interacting with a POPC Lipid Bilayer. J. Phys. Chem. B 2010, 14, 384–399. [Google Scholar] [CrossRef] [PubMed]
- Tsutsumi, A.; Javkhlantugs, N.; Kira, A.; Umeyama, M.; Kawamura, I.; Nishimura, K.; Ueda, K.; Naito, A. Structure and orientation of bovine lactoferrampin in the mimetic bacterial membrane as revealed by solid-state NMR and molecular dynamics simulation. Biophys. J. 2012, 103, 1735–1743. [Google Scholar] [CrossRef] [PubMed]
- Ulmschneider, J.P.; Smith, J.C.; Ulmschneider, M.B.; Ulrich, A.S.; Strandberg, E. Reorientation and dimerization of the membrane-bound antimicrobial peptide PGLa from microsecond all-atom MD simulations. Biophys. J. 2012, 103, 472–482. [Google Scholar] [CrossRef] [PubMed]
- Isralewitz, B.; Baudry, J.; Gullingsrud, J.; Kosztin, D.; Schulten, K. Steered Molecular Dynamics Investigations of Protein Function. J. Mol. Graph. Model. 2001, 19, 13–25. [Google Scholar] [CrossRef]
- Alaybeyoglu, B.; Sariyar Akbulut, B.; Ozkirimli, E. Insights into membrane translocation of the cell-penetrating peptide pVEC from molecular dynamics calculations. J. Biomol. Struct. Dyn. 2016, 34, 2387–2398. [Google Scholar] [CrossRef] [PubMed]
- Petkov, P.; Marinova, R.; Kochev, V.; Ilieva, N.; Lilkova, E.; Litov, L. Computational study of solution behavior of magainin 2 monomers. J. Biomol. Struct. Dyn. 2018, 11, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Lorenzo, A.C.; Bisch, P.M. Analyzing different parameters of steered molecular dynamics for small membrane interacting molecules. J. Mol. Graph. Model. 2005, 24, 59–71. [Google Scholar] [CrossRef] [PubMed]
- Akdag, I.O.; Ozkirimli, E. The Uptake Mechanism of the Cell-Penetrating pVEC Peptide. J. Chem. 2013, 2013, 1–9. [Google Scholar] [CrossRef]
- Lyu, Y.; Xiang, N.; Zhu, X.; Narsimhan, G. Potential of mean force for insertion of antimicrobial peptide melittin into a pore in mixed DOPC/DOPG lipid bilayer by molecular dynamics simulation. J. Chem. Phys. 2017, 146, 155101. [Google Scholar] [CrossRef] [PubMed]
- Almeida, P.F. And Yet It Moves. Biophys. J. 2017, 113, 759–761. [Google Scholar] [CrossRef] [PubMed]
- Faust, J.E.; Yang, P.Y.; Huang, H.W. Action of Antimicrobial Peptides on Bacterial and Lipid Membranes: A Direct Comparison. Biophys. J. 2017, 112, 1663–1672. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Navarro, M.; Teixidó, M.; Giralt, E. Jumping Hurdles: Peptides Able to Overcome Biological Barriers. Acc. Chem. Res. 2017, 50, 1847–1854. [Google Scholar] [CrossRef] [PubMed]
- Elmquist, A.; Hansen, M.; Langel, Ü. Structure-activity relationship study of the cell-penetrating peptide pVEC. BBA Biomembr. 2006, 1758, 721–729. [Google Scholar] [CrossRef] [PubMed]
- Elmquist, A.; Lindgren, M.; Bartfai, T.; Langel, Ü. Ve-cadherin-derived cell-penetrating peptide, pVEC with carrier functions. Exp. Cell Res. 2001, 269, 237–244. [Google Scholar] [CrossRef] [PubMed]
- Fawell, S.; Seery, J.; Daikh, Y.; Moore, C.; Chen, L.L.; Pepinsky, B.; Barsoum, J. Tat-mediated delivery of heterologous proteins into cells. Proc. Natl. Acad. Sci. USA 1994, 91, 664–668. [Google Scholar] [CrossRef] [PubMed]
- Vazdar, M.; Heyda, J.; Mason, P.E.; Tesei, G.; Allolio, C.; Lund, M.; Jungwirth, P. Arginine “Magic”: Guanidinium Like-Charge Ion Pairing from Aqueous Salts to Cell Penetrating Peptides. Acc. Chem. Res. 2018, 51, 1455–1464. [Google Scholar] [CrossRef] [PubMed]
- Sun, D.; Forsman, J.; Lund, M.; Woodward, C.E. Effect of arginine-rich cell penetrating peptides on membrane pore formation and life-times: A molecular simulation study. Phys. Chem. Chem. Phys. 2014, 16, 20785–20795. [Google Scholar] [CrossRef] [PubMed]
- Huang, K.; García, A.E. Free energy of translocating an arginine-rich cell-penetrating peptide across a lipid bilayer suggests pore formation. Biophys. J. 2013, 104, 412–420. [Google Scholar] [CrossRef] [PubMed]
- MacCallum, J.L.; Bennett, W.F.D.; Tieleman, D.P. Distribution of amino acids in a lipid bilayer from computer simulations. Biophys. J. 2008, 94, 3393–3404. [Google Scholar] [CrossRef] [PubMed]
- MacCallum, J.L.; Bennett, W.F.D.; Tieleman, D.P. Transfer of arginine into lipid bilayers is nonadditive. Biophys. J. 2011, 101, 110–117. [Google Scholar] [CrossRef] [PubMed]
- Su, Y.; Waring, A.J.; Ruchala, P.; Hong, M. Membrane-bound dynamic structure of an arginine-rich cell-penetrating peptide, the protein transduction domain of HIV tat, from solid-state NMR. Biochemistry 2010, 49, 6009–6020. [Google Scholar] [CrossRef] [PubMed]
- Mishra, V.K.; Palgunachari, M.N.; Segrest, J.P.; Anantharamaiah, G.M. Interactions of synthetic peptide analogs of the class A amphipathic helix with lipids. Evidence for the snorkel hypothesis. J. Biol. Chem. 1994, 269, 7185–7191. [Google Scholar] [PubMed]
- Strandberg, E.; Killian, J.A. Snorkeling of lysine side chains in transmembrane helices: How easy can it get? FEBS Lett. 2003, 544, 69–73. [Google Scholar] [CrossRef]
- Ulmschneider, M.B.; Ulmschneider, J.P.; Freites, J.A.; Heijne, G.; Tobias, D.J.; White, S.H. Transmembrane helices containing a charged arginine are thermodynamically stable. Eur. Biophys. J. 2017, 46, 627–637. [Google Scholar] [CrossRef] [PubMed]
- Lupas, A.; Van Dyke, M.; Stock, J. Predicting coiled coils from protein sequences. Science 1991, 252, 1162–1164. [Google Scholar] [CrossRef] [PubMed]
- Schrauber, H.; Eisenhaber, F.; Argos, P. Rotamers: To be or not to be? An Analysis of Amino Acid Side-chain Conformations in Globular Proteins. J. Mol. Biol. 1993, 230, 592–612. [Google Scholar] [CrossRef] [PubMed]
- Fukuda, M.; Gotoh, I.; Gotoh, Y.; Nishida, E. Cytoplasmic Localization of Mitogen-activated Protein Kinase Kinase Directed by Its NH2-terminal, Leucine-rich Short Amino Acid Sequence, Which Acts as a Nuclear Export Signal. J. Biol. Chem. 1996, 271, 20024–20028. [Google Scholar] [CrossRef] [PubMed]
- Kutay, U.; Güttinger, S. Leucine-rich nuclear-export signals: Born to be weak. Trends Cell Biol. 2005, 15, 121–124. [Google Scholar] [CrossRef] [PubMed]
- Landschulz, W.H.; Johnson, P.F.; McKnight, S.L. The leucine zipper: A hypothetical structure common to a new class of DNA binding proteins. Science 1988, 240, 1759–1764. [Google Scholar] [CrossRef] [PubMed]
- Chou, P.Y.; Fasman, G.D. Structural and functional role of leucine residues in proteins. Biophys. J. 1973, 74, 263–281. [Google Scholar] [CrossRef]
- Ulmschneider, J.P. Charged Antimicrobial Peptides can Translocate across Membranes without Forming Channel-like Pores. Biophys. J. 2017, 113, 73–81. [Google Scholar] [CrossRef] [PubMed]
- Gurezka, R.; Laage, R.; Brosig, B.; Langosch, D. A Heptad Motif of Leucine Residues Found in Membrane Proteins Can Drive Self-assembly of Artificial Transmembrane Segments. J. Biol. Chem. 1999, 274, 9265–9270. [Google Scholar] [CrossRef] [PubMed]
- Vasconcelos, L.; Madani, F.; Arukuusk, P.; Pärnaste, L.; Gräslund, A.; Langel, Ü. Effects of cargo molecules on membrane perturbation caused by transportan10 based cell-penetrating peptides. Biochim. Biophys. Acta 2014, 1838, 3118–3129. [Google Scholar] [CrossRef] [PubMed]
- Srimanee, A.; Arvanitidou, M.; Kim, K.; Hällbrink, M.; Langel, Ü. Cell-penetrating peptides for siRNA delivery to glioblastomas. Peptides 2018, 104, 62–69. [Google Scholar] [CrossRef] [PubMed]
- MacCallum, J.L.; Bennett, W.F.D.; Tieleman, D.P. Partitioning of Amino Acid Side Chains into Lipid Bilayers: Results from Computer Simulations and Comparison to Experiment. J. Gen. Physiol. 2007, 129, 371–377. [Google Scholar] [CrossRef] [PubMed]
- Jaud, S.; Fernández-Vidal, M.; Nilsson, I.; Meindl-Beinker, N.M.; Hübner, N.C.; Tobias, D.J.; von Heijne, G.; White, S.H. Insertion of short transmembrane helices by the Sec61 translocon. Proc. Natl. Acad. Sci. USA 2009, 106, 11588–11593. [Google Scholar] [CrossRef] [PubMed]
- Ulmschneider, J.P.; Smith, J.C.; White, S.H.; Ulmschneider, M.B. In Silico Partitioning and Transmembrane Insertion of Hydrophobic Peptides under Equilibrium Conditions. J. Am. Chem. Soc. 2011, 133, 15487–15495. [Google Scholar] [CrossRef] [PubMed]
- Marks, J.R.; Placone, J.; Hristova, K.; Wimley, W.C. Spontaneous membrane-translocating peptides by orthogonal high-throughput screening. J. Am. Chem. Soc. 2011, 133, 8995–9004. [Google Scholar] [CrossRef] [PubMed]
- Fuselier, T.; Wimley, W.C. Spontaneous Membrane Translocating Peptides: The Role of Leucine-Arginine Consensus Motifs. Biophys. J. 2017, 113, 835–846. [Google Scholar] [CrossRef] [PubMed]
- Alaybeyoglu, B.; Uluocak, B.G.; Akbulut, B.S.; Ozkirimli, E. The effect of a β-lactamase inhibitor peptide on bacterial membrane structure and integrity: A comparative study. J. Pept. Sci. 2017, 23, 374–383. [Google Scholar] [CrossRef] [PubMed]
- Gajski, G.; Garaj-Vrhovac, V. Melittin: A lytic peptide with anticancer properties. Environ. Toxicol. Pharmacol. 2013, 36, 697–705. [Google Scholar] [CrossRef] [PubMed]
- Jamasbi, E.; Batinovic, S.; Sharples, R.A.; Sani, M.-A.; Robins-Browne, R.M.; Wade, J.D.; Separovic, F.; Hossain, M.A. Melittin peptides exhibit different activity on different cells and model membranes. Amino Acids 2014, 46, 2759–2766. [Google Scholar] [CrossRef] [PubMed]
- Raghuraman, H.; Chattopadhyay, A. Melittin: A Membrane-active Peptide with Diverse Functions. Biosci. Rep. 2006, 27, 189–223. [Google Scholar] [CrossRef] [PubMed]
- Terwilliger, T.C.; Eisenberg, D. The structure of melittin. II. Interpretation of the structure. J. Biol. Chem. 1982, 257, 6016–6022. [Google Scholar] [PubMed]
- Asthana, N.; Yadav, S.P.; Ghosh, J.K. Dissection of Antibacterial and Toxic Activity of Melittin A Leucine zipper motif plays a crucial role in determining its hemolytic activity but not antibacterial activity. J. Biol. Chem. 2004, 279, 55042–55050. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, R.M.; Srivastava, S.; Singh, M.; Bajpai, V.K.; Ghosh, J.K. Consequences of Alteration in Leucine Zipper Sequence of Melittin in Its Neutralization of Lipopolysaccharide-induced Proinflammatory Response in Macrophage Cells and Interaction with Lipopolysaccharide. J. Biol. Chem. 2012, 287, 1980–1995. [Google Scholar] [CrossRef] [PubMed]
- Lauterwein, J.; Brown, L.R.; Wuethrich, K. High-resolution 1h-nmr studies of monomeric melittin in aqueous solution. Biochim. Biophys. Acta 1980, 622, 219–230. [Google Scholar] [CrossRef]
- Bazzo, R.; Tappin, M.J.; Pastore, A.; Harvey, T.S.; Carver, J.A.; Campbell, I.D. The structure of melittin. Eur. J. Biochem. 1988, 173, 139–146. [Google Scholar] [CrossRef] [PubMed]
- Hewish, D.R.; Barnham, K.J.; Werkmeister, J.A.; Kirkpatrick, A.; Bartone, N.; Liu, S.T.; Norton, R.S.; Curtain, C.; Rivetta, D.E. Structure and activity of D-Pro14 melittin. J. Protein Chem. 2002, 21, 243–253. [Google Scholar] [CrossRef] [PubMed]
- Ramirez, L.; Shekhtman, A.; Pande, J. Nuclear Magnetic Resonance-Based Structural Characterization and Backbone Dynamics of Recombinant Bee Venom Melittin. Biochemistry 2018, 57, 2775–2785. [Google Scholar] [CrossRef] [PubMed]
- Oren, Z.; Shai, Y. Selective lysis of bacteria but not mammalian cells by diastereomers of melittin: Structure-function study. Biochemistry 1997, 36, 1826–1835. [Google Scholar] [CrossRef] [PubMed]
- Tosteson, M.T.; Tosteson, D.C. The sting. Melittin forms channels in lipid bilayers. Biophys. J. 1981, 36, 109–116. [Google Scholar] [CrossRef]
- Wu, Y.; Ma, L.; Cheley, S.; Bayley, H.; Cui, Q.; Chapman, E.R. Permeation of Styryl Dyes through Nanometer-Scale Pores in Membranes. Biochemistry 2011, 50, 7493–7502. [Google Scholar] [CrossRef] [PubMed]
- Krauson, A.J.; He, J.; Wimley, W.C. Determining the mechanism of membrane permeabilizing peptides: Identification of potent, equilibrium pore-formers. Biochim. Biophys. Acta 2012, 1818, 1625–1632. [Google Scholar] [CrossRef] [PubMed]
- Wiedman, G.; Herman, K.; Searson, P.; Wimley, W.C.; Hristova, K. The electrical response of bilayers to the bee venom toxin melittin: Evidence for transient bilayer permeabilization. Biochim. Biophys. Acta 2013, 1828, 1357–1364. [Google Scholar] [CrossRef] [PubMed]
- Wimley, W.C. How Does Melittin Permeabilize Membranes? Biophys. J. 2018, 114, 251–253. [Google Scholar] [CrossRef] [PubMed]
- Nepal, B.; Leveritt, J., III; Lazaridis, T. Membrane Curvature Sensing by Amphipathic Helices: Insights from Implicit Membrane Modeling. Biophys. J. 2018, 114, 2128–2141. [Google Scholar] [CrossRef] [PubMed]
- Koller, D.; Lohner, K. The role of spontaneous lipid curvature in the interaction of interfacially active peptides with membranes. BBA Biomembr. 2014, 1838, 2250–2259. [Google Scholar] [CrossRef] [PubMed]
- Sun, D.; Forsman, J.; Woodward, C.E. Molecular Simulations of Melittin-Induced Membrane Pores. J. Phys. Chem. B. 2017, 121, 10209–10214. [Google Scholar] [CrossRef] [PubMed]
- Gong, Z.; Ikonomova, S.P.; Karlsson, A.J. Secondary structure of cell-penetrating peptides during interaction with fungal cells. Protein Sci. 2018, 27, 702–713. [Google Scholar] [CrossRef] [PubMed]
- Stalmans, S.; Bracke, N.; Wynendaele, E.; Gevaert, B.; Peremans, K.; Burvenich, C.; Polis, I.; De Spiegeleer, B. Cell-Penetrating Peptides Selectively Cross the Blood-Brain Barrier In Vivo. PLoS ONE 2015, 10, e0139652. [Google Scholar] [CrossRef] [PubMed]
- Silva, N.C.; Sarmento, B.; Pintado, M. The importance of antimicrobial peptides and their potential for therapeutic use in ophthalmology. Int. J. Antimicrob. Agents 2013, 41, 5–10. [Google Scholar] [CrossRef] [PubMed]
- Craik, D.J.; Fairlie, D.P.; Liras, S.; Price, D. The Future of Peptide-based Drugs. Chem. Biol. Drug Des. 2013, 81, 136–147. [Google Scholar] [CrossRef] [PubMed]
- Fonseca, S.B.; Pereira, M.P.; Kelley, S.O. Recent advances in the use of cell-penetrating peptides for medical and biological applications. Adv. Drug Deliv. Rev. 2009, 61, 953–964. [Google Scholar] [CrossRef] [PubMed]
- Haney, E.F.; Hancock, R.E.W. Peptide design for antimicrobial and immunomodulatory applications. Biopolymers 2013, 100, 572–583. [Google Scholar] [CrossRef] [PubMed]
- Reissmann, S. Cell penetration: Scope and limitations by the application of cell-penetrating peptides. J. Pept. Sci. 2014, 20, 760–784. [Google Scholar] [CrossRef] [PubMed]
- Olmez, E.O.; Akbulut, B.S. Protein-peptide interactions revolutionize drug development. In Binding Protein; InTech: Rijeka Croatia, 2012. [Google Scholar]
- Vlieghe, P.; Lisowski, V.; Martinez, J.; Khrestchatisky, M. Synthetic therapeutic peptides: Science and market. Drug Discov. Today 2010, 15, 40–56. [Google Scholar] [CrossRef] [PubMed]
- Fox, J.L. Antimicrobial peptides stage a comeback. Nat. Biotechnol. 2013, 31, 379–382. [Google Scholar] [CrossRef] [PubMed]
- Uhlig, T.; Kyprianou, T.; Martinelli, F.G.; Oppici, C.A.; Heiligers, D.; Hills, D.; Calvo, X.R.; Verhaert, P. The emergence of peptides in the pharmaceutical business: From exploration to exploitation. EuPA Open Proteom. 2014, 4, 58–69. [Google Scholar] [CrossRef]
- Sato, A.K.; Viswanathan, M.; Kent, R.B.; Wood, C.R. Therapeutic peptides: Technological advances driving peptides into development. Curr. Opin. Biotechnol. 2006, 17, 638–642. [Google Scholar] [CrossRef] [PubMed]
- McGregor, D.P. Discovering and improving novel peptide therapeutics. Curr. Opin. Pharmacol. 2008, 8, 616–619. [Google Scholar] [CrossRef] [PubMed]
- Lalatsa, A.; Schatzlein, A.G.; Uchegbu, I.F. Strategies to deliver peptide drugs to the brain. Mol. Pharm. 2014, 11, 1081–1093. [Google Scholar] [CrossRef] [PubMed]
- Liskamp, R.M.J.; Rijkers, D.T.S.; Kruijtzer, J.A.W.; Kemmink, J. Peptides and Proteins as a Continuing Exciting Source of Inspiration for Peptidomimetics. ChemBioChem 2011, 12, 1626–1653. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Hu, X.; Xiang, D. Nanoparticle drug delivery systems: An excellent carrier for tumor peptide vaccines. Drug Deliv. 2018, 25, 1319–1327. [Google Scholar] [CrossRef] [PubMed]
- Spicer, C.D.; Jumeaux, C.; Gupta, B.; Stevens, M.M. Peptide and protein nanoparticle conjugates: Versatile platforms for biomedical applications. Chem. Soc. Rev. 2018, 47, 3574–3620. [Google Scholar] [CrossRef] [PubMed]
- Misra, S.K.; Ye, M.; Kim, S.; Pan, D. Highly efficient anti-cancer therapy using scorpion “NanoVenin”. Chem. Commun. 2014, 50, 13220–13223. [Google Scholar] [CrossRef] [PubMed]
- Alberici, L.; Roth, L.; Sugahara, K.N.; Agemy, L.; Kotamraju, V.R.; Teesalu, T.; Bordignon, C.; Traversari, C.; Rizzardi, G.-P.; Ruoslahti, E. De novo design of a tumor-penetrating peptide. Cancer Res. 2013, 73, 804–812. [Google Scholar] [CrossRef] [PubMed]
- Uchida, Y.; Raina, D.; Kharbanda, S.; Kufe, D. Inhibition of the MUC1-C oncoprotein is synergistic with cytotoxic agents in the treatment of breast cancer cells. Cancer Biol. Ther. 2013, 14, 127–134. [Google Scholar] [CrossRef] [PubMed]
- Duong, H.H.P.; Yung, L.Y.L. Synergistic co-delivery of doxorubicin and paclitaxel using multi-functional micelles for cancer treatment. Int. J. Pharm. 2013, 454, 486–495. [Google Scholar] [CrossRef] [PubMed]
- Ouahab, A.; Cheraga, N.; Onoja, V.; Shen, Y.; Tu, J. Novel pH-sensitive charge-reversal cell penetrating peptide conjugated PEG-PLA micelles for docetaxel delivery: In vitro study. Int. J. Pharm. 2014, 466, 233–245. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Cheetham, A.G.; Lock, L.L.; Cui, H. Cellular uptake and cytotoxicity of drug-peptide conjugates regulated by conjugation site. Bioconjug. Chem. 2013, 24, 604–613. [Google Scholar] [CrossRef] [PubMed]
- Rakesh, K.P.; Suhas, R.; Gowda, D.C. Anti-inflammatory and Antioxidant Peptide-Conjugates: Modulation of Activity by Charged and Hydrophobic Residues. Int. J. Pept. Res. Ther. 2017, 1–8. [Google Scholar] [CrossRef]
- Cheng, Y.; Chen, Z.; Liao, T.; Lin, C.; Shen, H.C.; Wang, Y.; Chang, C.; Liu, R.; Chen, R.P.; Tu, P. An intranasally delivered peptide drug ameliorates cognitive decline in Alzheimer transgenic mice. EMBO Mol. Med. 2017, 9, 703–715. [Google Scholar] [CrossRef] [PubMed]
- Alaybeyoglu, B.; Akbulut, B.S.; Ozkirimli, E. A novel chimeric peptide with antimicrobial activity. J. Pept. Sci. 2015, 21, 294–301. [Google Scholar] [CrossRef] [PubMed]
- Baig, M.H.; Ahmad, K.; Saeed, M.; Alharbi, A.M.; Barreto, G.E.; Ashraf, G.M.; Choi, I. Peptide based therapeutics and their use for the treatment of neurodegenerative and other diseases. Biomed. Pharmacother. 2018, 103, 574–581. [Google Scholar] [CrossRef] [PubMed]
- Kumai, T.; Kobayashi, H.; Harabuchi, Y.; Celis, E. Peptide vaccines in cancer-old concept revisited. Curr. Opin. Immunol. 2017, 45, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Clifton, G.T.; Mittendorf, E.A.; Peoples, G.E. Adjuvant HER2/neu peptide cancer vaccines in breast cancer. Immunotherapy 2015, 7, 1159–1168. [Google Scholar] [CrossRef] [PubMed]
- Chamani, R.; Ranji, P.; Hadji, M.; Nahvijou, A.; Esmati, E.; Alizadeh, A.M. Application of E75 peptide vaccine in breast cancer patients: A systematic review and meta-analysis. Eur. J. Pharmacol. 2018, 831, 87–93. [Google Scholar] [CrossRef] [PubMed]
- Khanna, P.; Blais, N.; Gaudreau, P.-O.; Corrales-Rodriguez, L. Immunotherapy Comes of Age in Lung Cancer. Clin. Lung Cancer 2017, 18, 13–22. [Google Scholar] [CrossRef] [PubMed]
- Rosendahl-Huber, S.K.; Camps, M.G.M.; Jacobi, R.H.J.; Mouthaan, J.; van Dijken, H.; van Beek, J.; Ossendorp, F.; de Jonge, J. Synthetic Long Peptide Influenza Vaccine Containing Conserved T and B Cell Epitopes Reduces Viral Load in Lungs of Mice and Ferrets. PLoS ONE 2015, 10, e0127969. [Google Scholar] [CrossRef] [PubMed]
- Kenter, G.G.; Welters, M.J.P.; Valentijn, A.R.P.M.; Löwik, M.J.G.; der Meer, D.M.A.; Vloon, A.P.G.; Essahsah, F.; Fathers, L.M.; Offringa, R.; Drijfhout, J.W.; et al. Vaccination against HPV-16 oncoproteins for vulvar intraepithelial neoplasia. N. Engl. J. Med. 2009, 361, 1838–1847. [Google Scholar] [CrossRef] [PubMed]
- Van Poelgeest, M.I.E.; Welters, M.J.P.; Vermeij, R.; Stynenbosch, L.F.M.; Loof, N.M.; der Meer, D.M.A.; Löwik, M.J.G.; Hamming, I.L.E.; van Esch, E.M.G.; Hellebrekers, B.W.J.; et al. Vaccination against Oncoproteins of HPV16 for Noninvasive Vulvar/Vaginal Lesions: Lesion Clearance Is Related to the Strength of the T-Cell Response. Clin. Cancer Res. 2016, 22, 2342–2350. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.Y.; Wang, P.N.; Chiu, M.J.; Finstad, C.L.; Lin, F.; Lynn, S.; Tai, Y.H.; De Fang, X.; Zhao, K.; Hung, C.H.; et al. UB-311, a novel UBITh®amyloid β peptide vaccine for mild Alzheimer’s disease. Alzheimers Dement. 2017, 3, 262–272. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Joshi, M.D.; Singhania, S.; Ramsey, K.H.; Murthy, A.K. Peptide Vaccine: Progress and Challenges. Vaccines 2014, 2, 515–536. [Google Scholar] [CrossRef] [PubMed]
- Middleton, G.; Silcocks, P.; Cox, T.; Valle, J.; Wadsley, J.; Propper, D.; Coxon, F.; Ross, P.; Madhusudan, S.; Roques, T.; et al. Gemcitabine and capecitabine with or without telomerase peptide vaccine GV1001 in patients with locally advanced or metastatic pancreatic cancer (TeloVac): An open-label, randomised, phase 3 trial. Lancet Oncol. 2014, 15, 829–840. [Google Scholar] [CrossRef]
- Staff, C.; Mozaffari, F.; Frödin, J.-E.; Mellstedt, H.; Liljefors, M. Telomerase (GV1001) vaccination together with gemcitabine in advanced pancreatic cancer patients. Int. J. Oncol. 2014, 45, 1293–1303. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.; Seo, E.-H.; Lee, S.-H.; Kim, B.-J. The Telomerase-Derived Anticancer Peptide Vaccine GV1001 as an Extracellular Heat Shock Protein-Mediated Cell-Penetrating Peptide. Int. J. Mol. Sci. 2016, 17, 2054. [Google Scholar] [CrossRef] [PubMed]
- Moon, K.T.; Yoo, T.K.; Kwon, S.Y.; Ha, J.Y.; Byun, S.S.; Kim, J.H.; Chung, J. Il; Kim, T.H.; Moon, H.S.; Kim, S.J.; et al. A randomised, placebo-controlled, multicentre, Phase 2 clinical trial to evaluate the efficacy and safety of GV1001 in patients with benign prostatic hyperplasia. BJU Int. 2018, 59, 245. [Google Scholar] [CrossRef] [PubMed]
- National Institute of Health. U.S. National library of Medicine ClinicalTrials.gov Homepage. Available online: www.clinicaltrials.gov (accessed on 12 June 2018).
- Lyon, J.G.; Mokarram, N.; Saxena, T.; Carroll, S.L.; Bellamkonda, R.V. Engineering challenges for brain tumor immunotherapy. Adv. Drug Deliv. Rev. 2017, 114, 19–32. [Google Scholar] [CrossRef] [PubMed]
- Stutz, K.; Müller, A.T.; Hiss, J.A.; Schneider, P.; Blatter, M.; Pfeiffer, B.; Posselt, G.; Kanfer, G.; Kornmann, B.; Wrede, P.; et al. Peptide-Membrane Interaction between Targeting and Lysis. ACS Chem. Biol. 2017, 12, 2254–2259. [Google Scholar] [CrossRef] [PubMed]
- Müller, A.T.; Gabernet, G.; Hiss, J.A.; Schneider, G. modlAMP: Python for antimicrobial peptides. Bioinformatics 2017, 33, 2753–2755. [Google Scholar] [CrossRef] [PubMed]
- Kalafatovic, D.; Giralt, E. Cell-Penetrating Peptides: Design Strategies beyond Primary Structure and Amphipathicity. Molecules 2017, 22, 1929. [Google Scholar] [CrossRef] [PubMed]
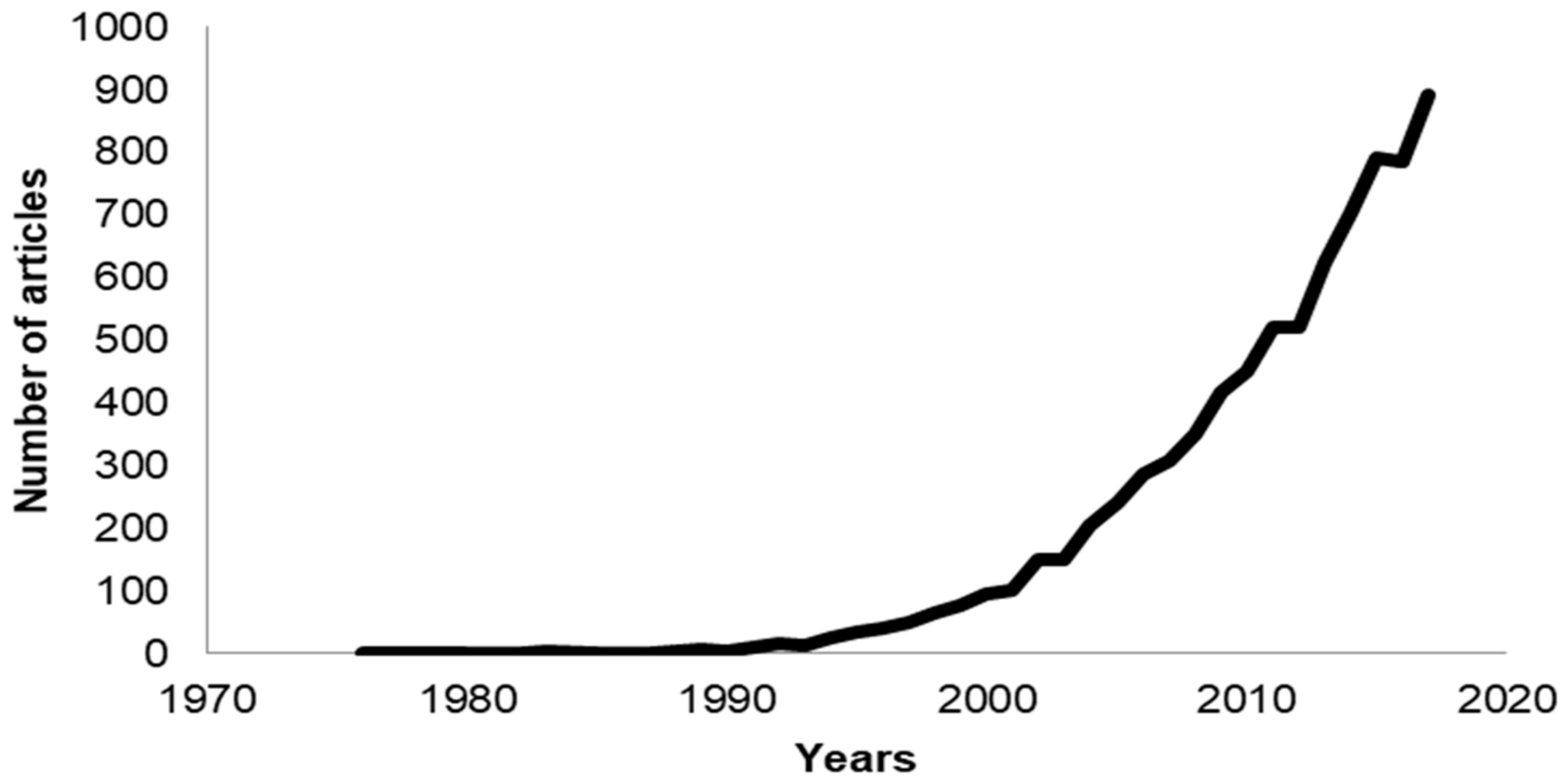
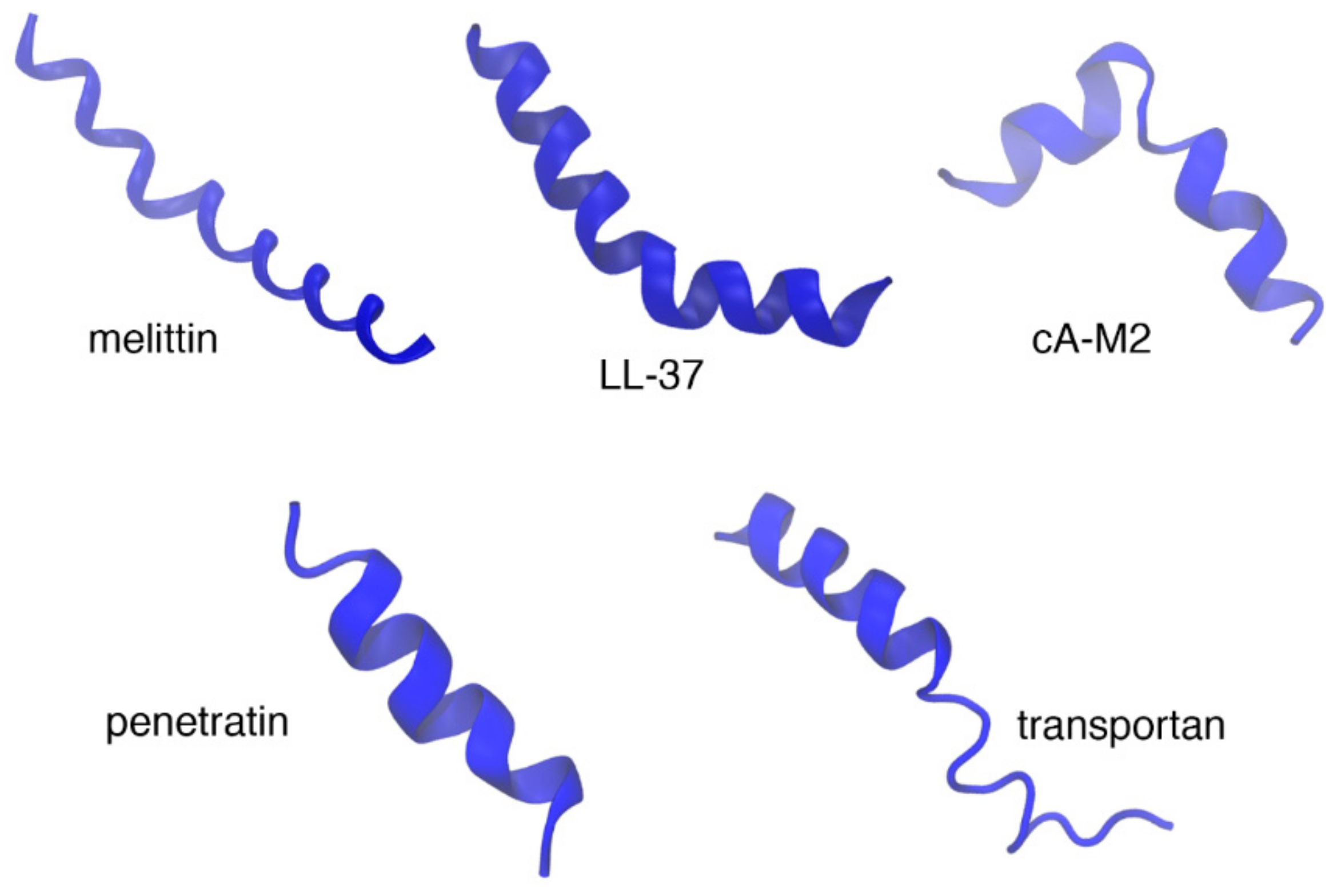
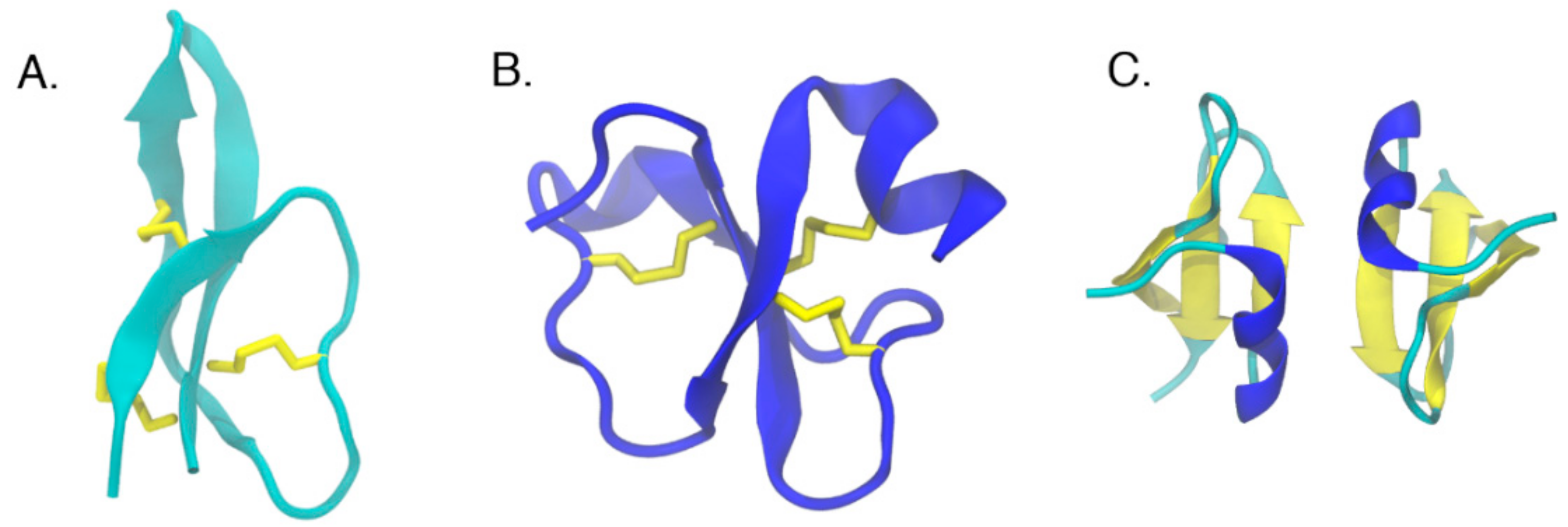
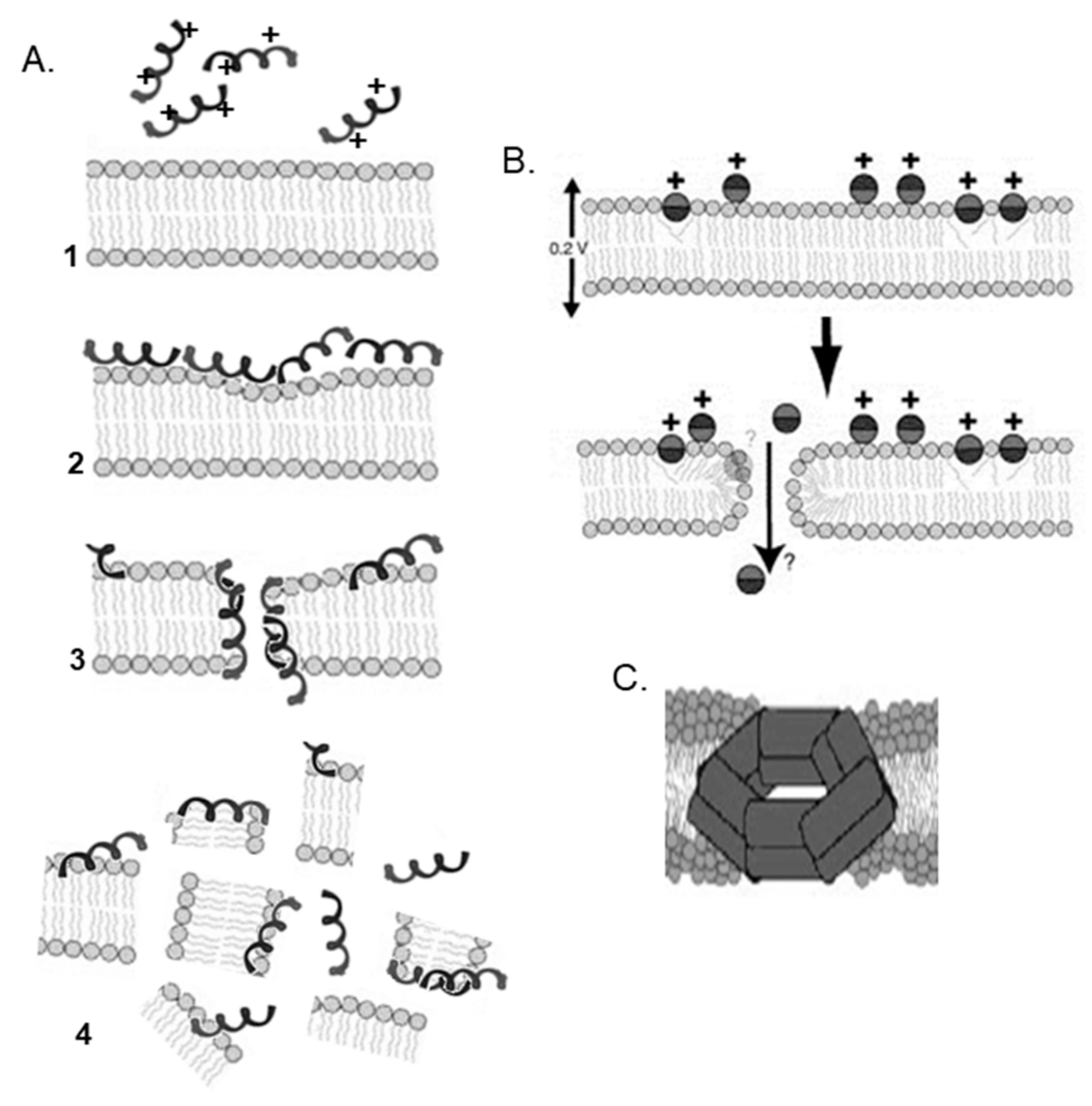
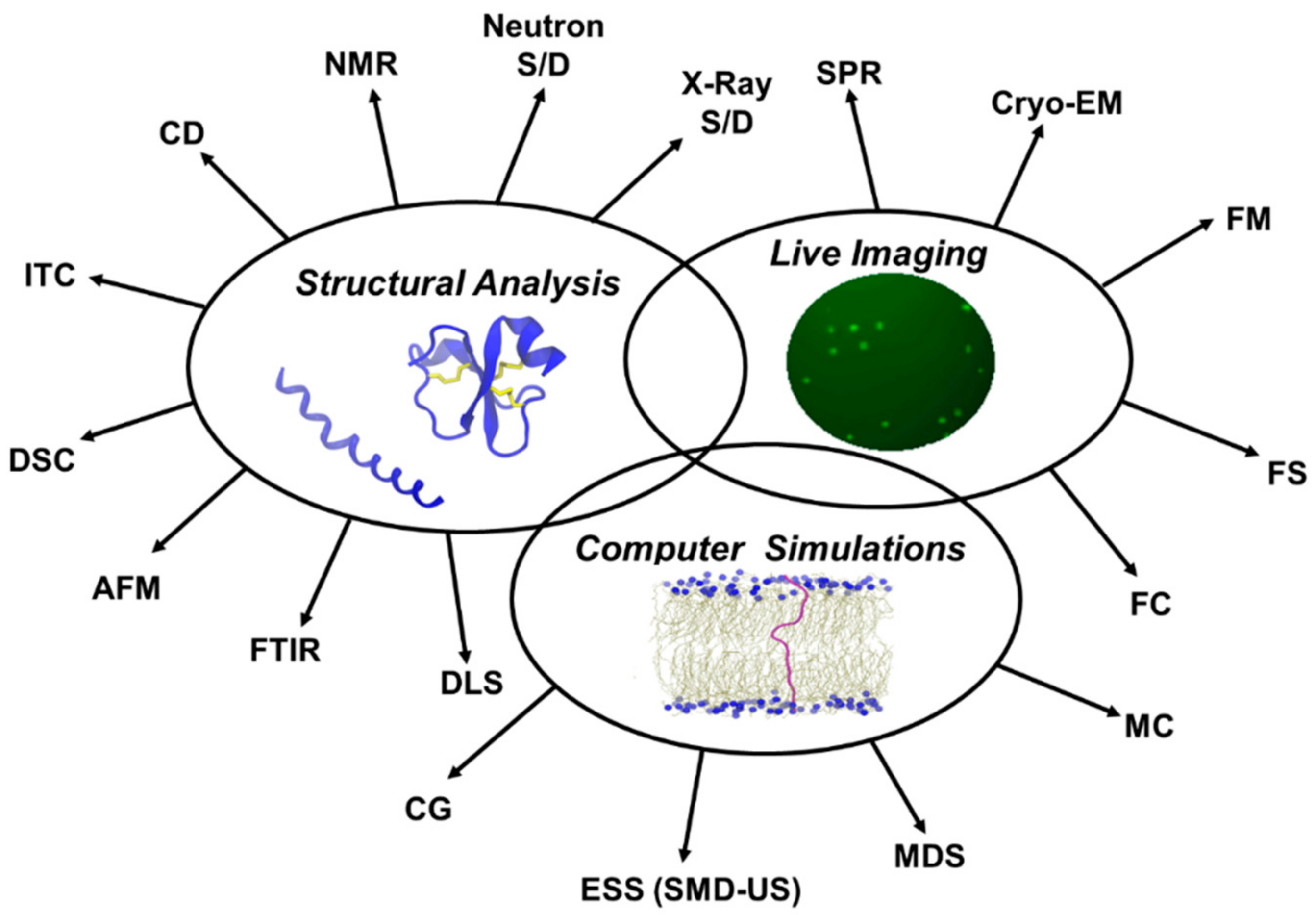
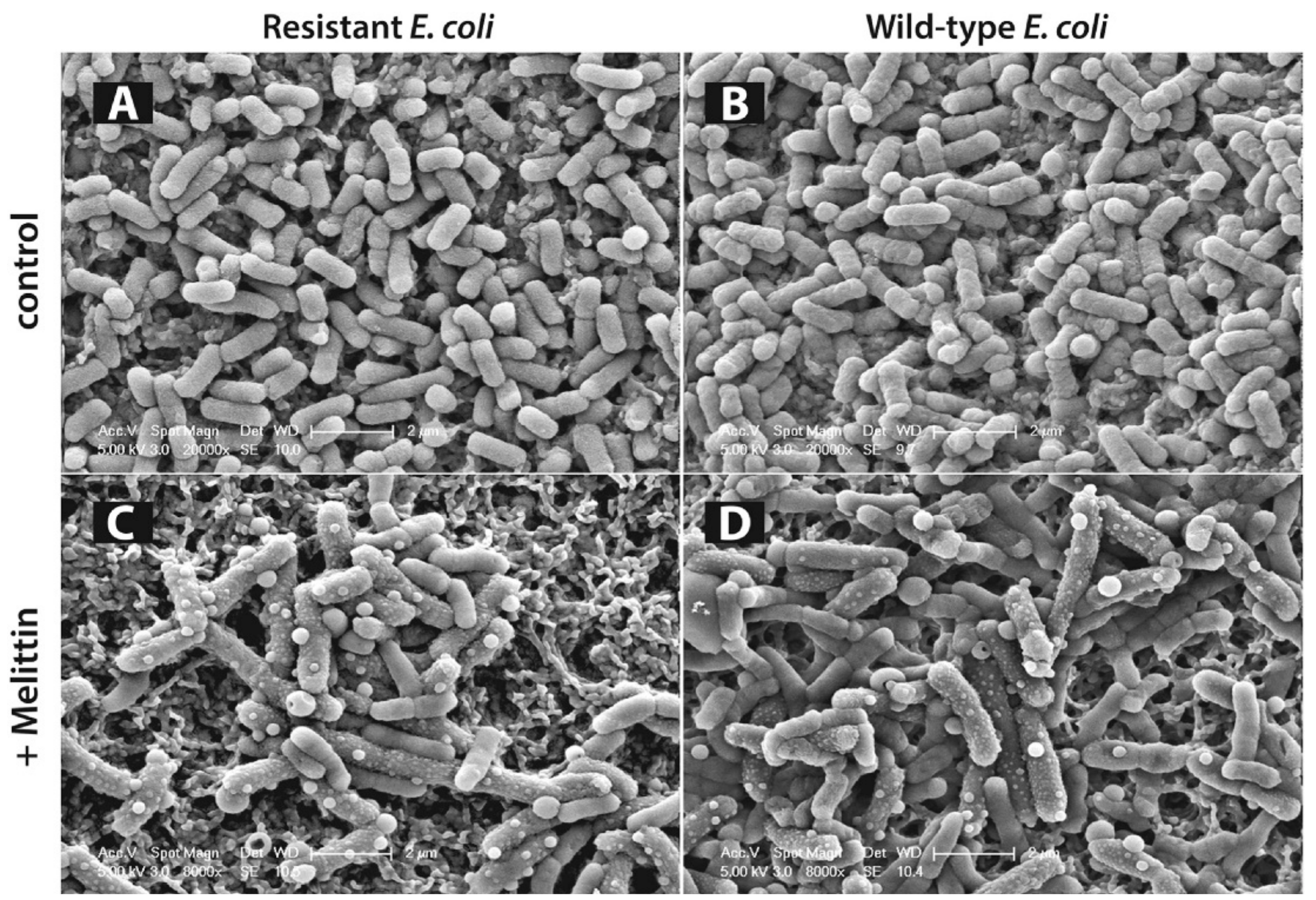
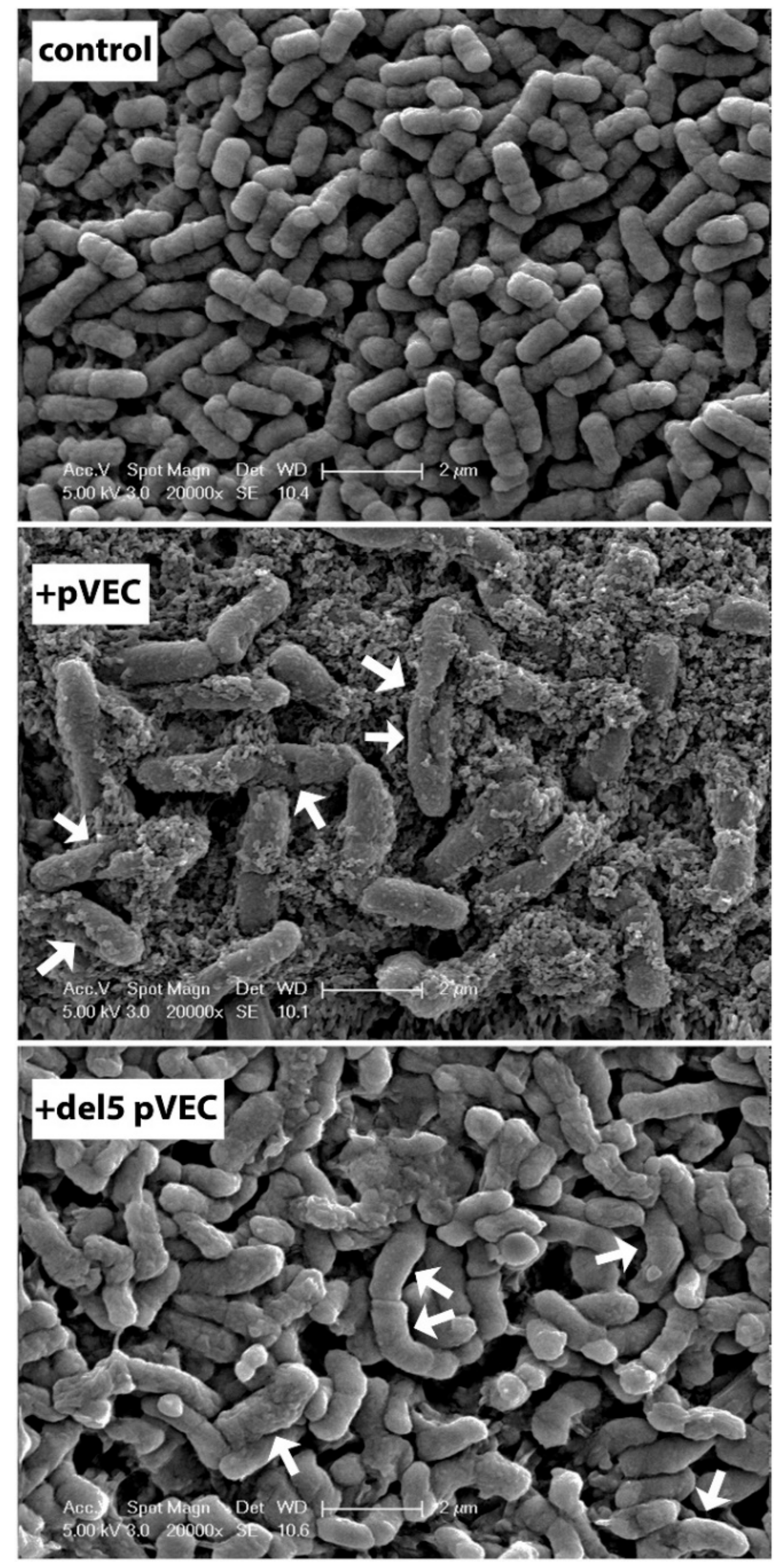
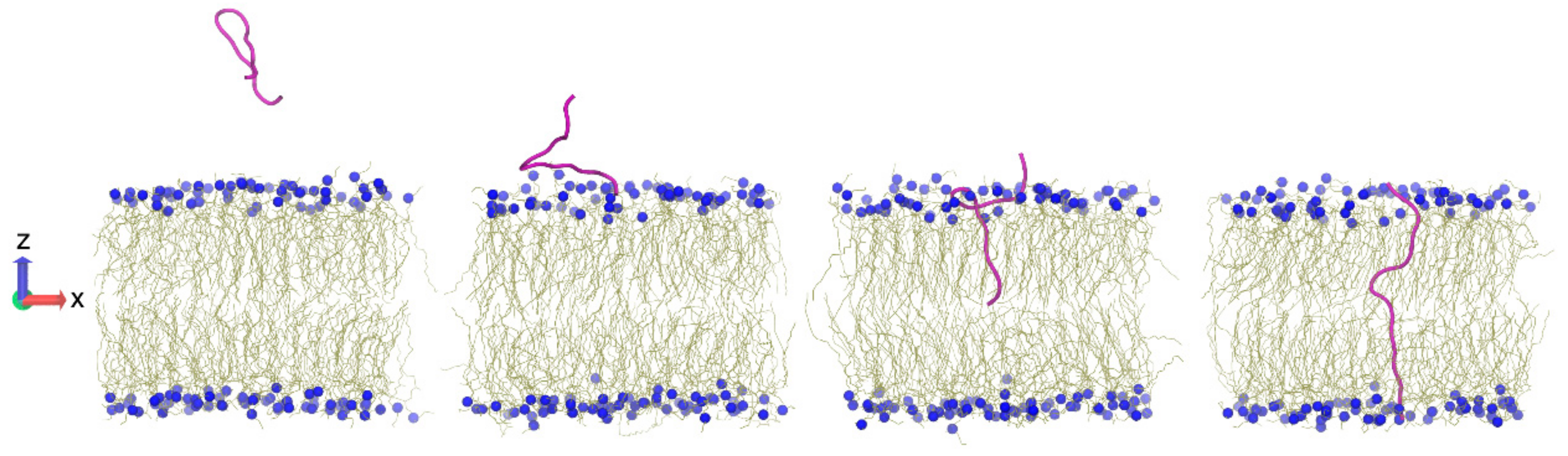
| Method | Application | Advantages/Disadvantages | AMP/CPP | References |
|---|---|---|---|---|
| X-ray Diffraction/Scattering | 3D Structure of the peptides Peptide–membrane interaction | Least expensive, high resolution, lower sample amounts, label-free. Large crystalline structures are needed, not suitable to study the structural dynamics of peptides in biological membranes, inability to image pores dynamically and in real-time. | Alamethicin Magainin 2 TAT Penetratin | [113] [114] [115] [116] |
| Neutron Diffraction/Scattering | 3D Structure of the peptide Peptide induced pore formation | High penetration ability of neutrons, label-free. Nuclear reactor is required, expensive, large amount of samples. | Melittin Alamethicin TAT TP-2 | [42] [117] [118] [119] |
| Nuclear magnetic resonance (NMR) spectroscopy | Structure of the peptides Orientation Membrane-peptide dynamics | High accuracy, label-free. Specialized expertise, expensive, need large amounts of peptides | Arenicin-2 Protegrin-2 Transportan 10 SAP | [120] [121] [32] [122] |
| Circular dichroism (CD) spectroscopy | Secondary structure of peptides and their changes in different environments | Quick estimation of secondary structure, cheap, simple Less accurate than NMR | Indolicidin Cecropin MPG CADY | [123] [124] [125] [126] |
| Isothermal titration calorimetry (ITC) | Thermodynamic properties of binding reaction Peptide induced lipid phase transitions Peptide aggregation at the membrane surface Peptide conformational changes | Complete and basic thermodynamic characterization, label-free Large sample amounts | Pardaxin PGLa riDOM r9 CPP | [127] [128] [129] [130] |
| Differential scanning calorimetry (DSC) | Changes in the phase transition temperature Thermodynamics of lipid phase transitions | Qualitative and quantitative measurement, provide information about physical and chemical change, label-free | Gramicidin KIGAKI MAP SAP | [131] [132] |
| Atomic force microscopy (AFM) | Structural changes on the membrane | Possibility of visualization of live cells in situ Do not provide sufficient topographical contrast, lacks chemical specificity, long image acquisition time, inability to image pores dynamically and in real-time | PGLa CM15 P(α) TAT | [133] [134] [135] [136] |
| Fourier transform infrared (FTIR) spectroscopy | Conformation and orientation of membrane-associated peptides and lipids Analysis of static and dynamic structure of peptides embedded in biological membranes | Possibility of working in a wide range of environment, requires less time and sample, inexpensive compared to X-ray diffraction, NMR, CD spectroscopy Difficulties in determination secondary structure and orientation | Arenicin 2 Magainin 2 P(α) Penetratin | [137] [138] [139] [140] |
| Surface plasmon resonance (SPR) | Real-time measurement of the change in adsorbed mass at the sensor surface as the peptides bind to selected biomimetic surfaces Kinetics of binding | High-surface sensitivity, real-time interactions analysis Expensive | Melittin Citropin 1.1 MCoTI-II Transportan | [141] [142] [143] [144] |
| Dynamic light scattering (DLS) | Measurement of size and size distribution of particles in a suspension Gives information about folding and conformation | Label-free, fast and noninvasive Needs low particle concentration | Maculatin Gomesin PepFect 3 (PF3) mtCPP-1 | [145] [146] [147] [148] |
| Fluorescence microscopy | Visualizing the membrane interaction and intracellular distribution of fluorescent-labelled peptides on biomimetic membranes or live cells Effect of peptides on membrane integrity | Inexpensive, wide-range of applications | LL-37 Histatin 5 pVEC Vim-TBS.58-81 | [149] [150] [151] [152] |
| Fluorescence spectroscopy | Kinetics of killing the live cell in real time Size and nature of transmembrane pores | Real-time interactions, quick, accurate Quenching of the contaminants | Cecropin A Trichogin GA IV TP-2 riDOM | [153] [154] [119] [129] |
| Förster resonance energy transfer (FRET) | Quantification of membrane disruption in live cells Determination of structural transition upon peptide–lipid interaction Peptide translocation across the bilayer | Highly sensitive Depend on concentration of fluorophores and distance between the donor and acceptor, require careful calibrations | PMAP-23 Halocidin 18-mer Transportan 10 Pep-1 | [155] [156] [157] [158] |
| Cryoelectron microscopy (cryo-EM) | Determination of 3 D structure Visualization of peptide–membrane interactions | High resolution, no need for crystal structure, require less sample and purity Proper equipment and expertise | Magainin 2 Melittin Penetratin | [159] [160] [116] |
| Flow cytometry | Kinetics of permeabilization of live cells and cell killing Cell viability and metabolic activity Membrane structure and its integrity Membrane potential | Statistically relevant and quantitative signal of fluorescence, very quick, sensitive detection Expensive equipment | PepR Melittin Vim-TBS.58-81 Penetratin | [161] [162] [152] [163] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Avci, F.G.; Sariyar Akbulut, B.; Ozkirimli, E. Membrane Active Peptides and Their Biophysical Characterization. Biomolecules 2018, 8, 77. https://doi.org/10.3390/biom8030077
Avci FG, Sariyar Akbulut B, Ozkirimli E. Membrane Active Peptides and Their Biophysical Characterization. Biomolecules. 2018; 8(3):77. https://doi.org/10.3390/biom8030077
Chicago/Turabian StyleAvci, Fatma Gizem, Berna Sariyar Akbulut, and Elif Ozkirimli. 2018. "Membrane Active Peptides and Their Biophysical Characterization" Biomolecules 8, no. 3: 77. https://doi.org/10.3390/biom8030077
APA StyleAvci, F. G., Sariyar Akbulut, B., & Ozkirimli, E. (2018). Membrane Active Peptides and Their Biophysical Characterization. Biomolecules, 8(3), 77. https://doi.org/10.3390/biom8030077





