Functions of the Proteasome on Chromatin
Abstract
:1. Introduction
2. A Primer on Chromatin
3. The Proteasome and Chromatin: A Caveat
4. The Form of the Proteasome That Associates with Chromatin
5. Proteasome Location and Recruitment
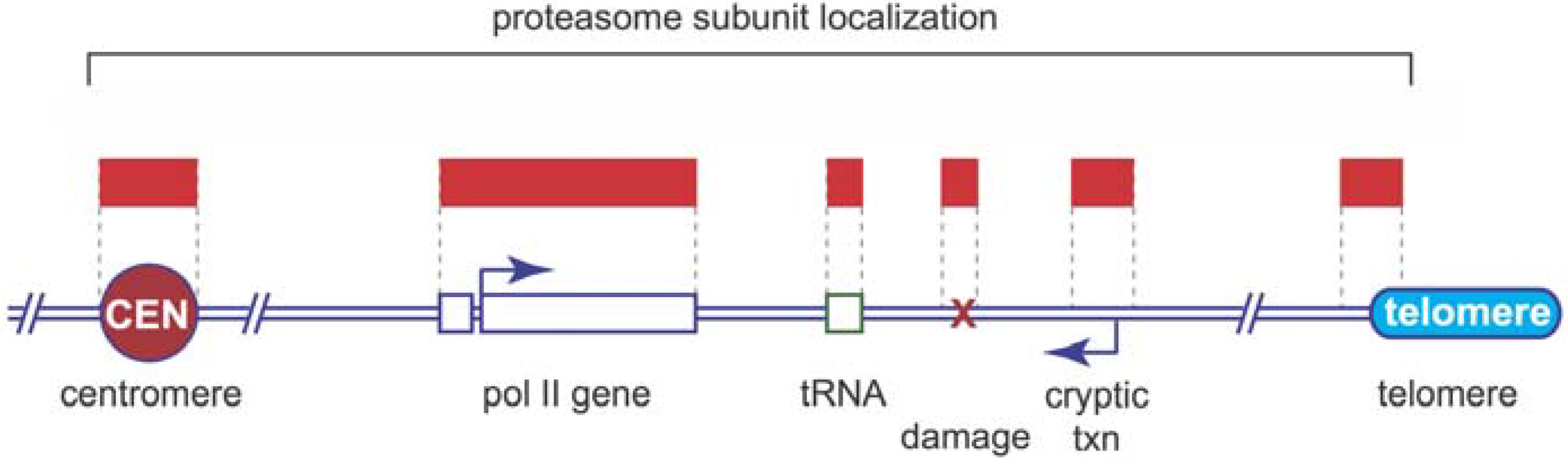
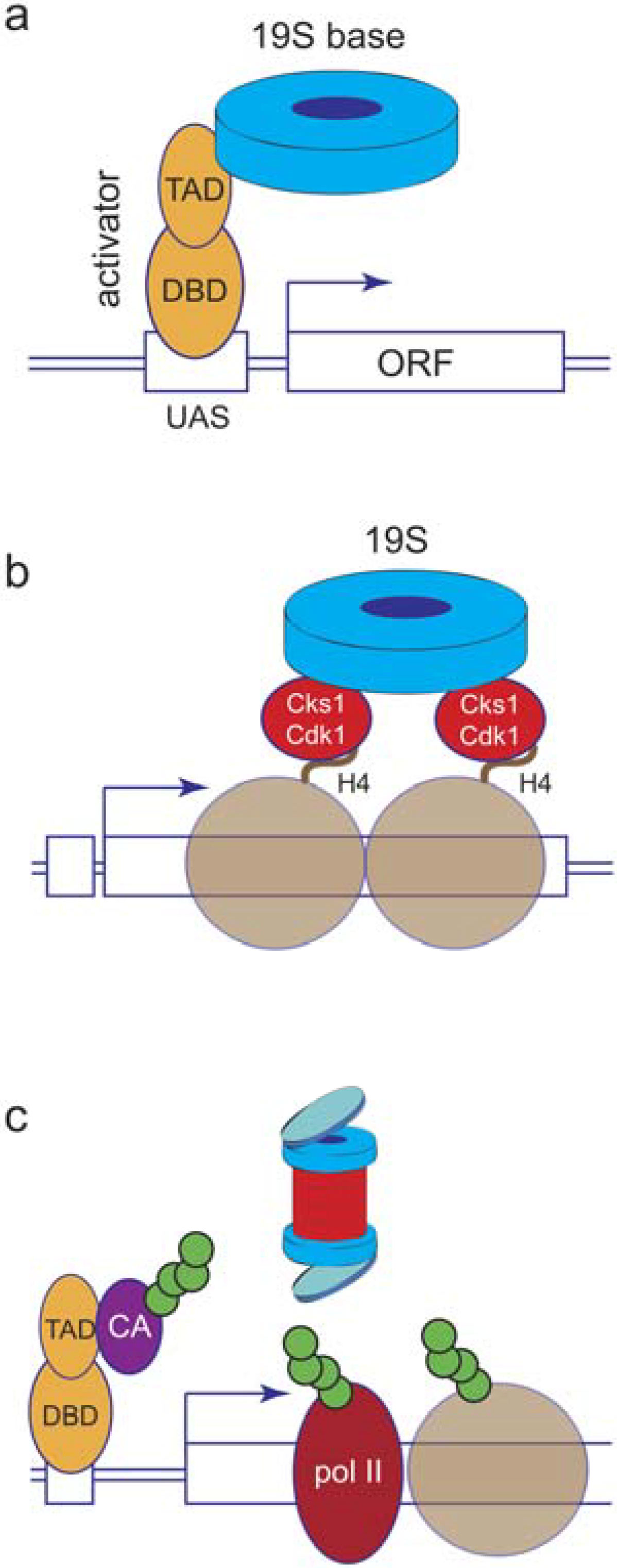
6. Proteolytic or Non-Proteolytic Functions of the Proteasome?
7. Functions of the Proteasome in mRNA-Type Transcription
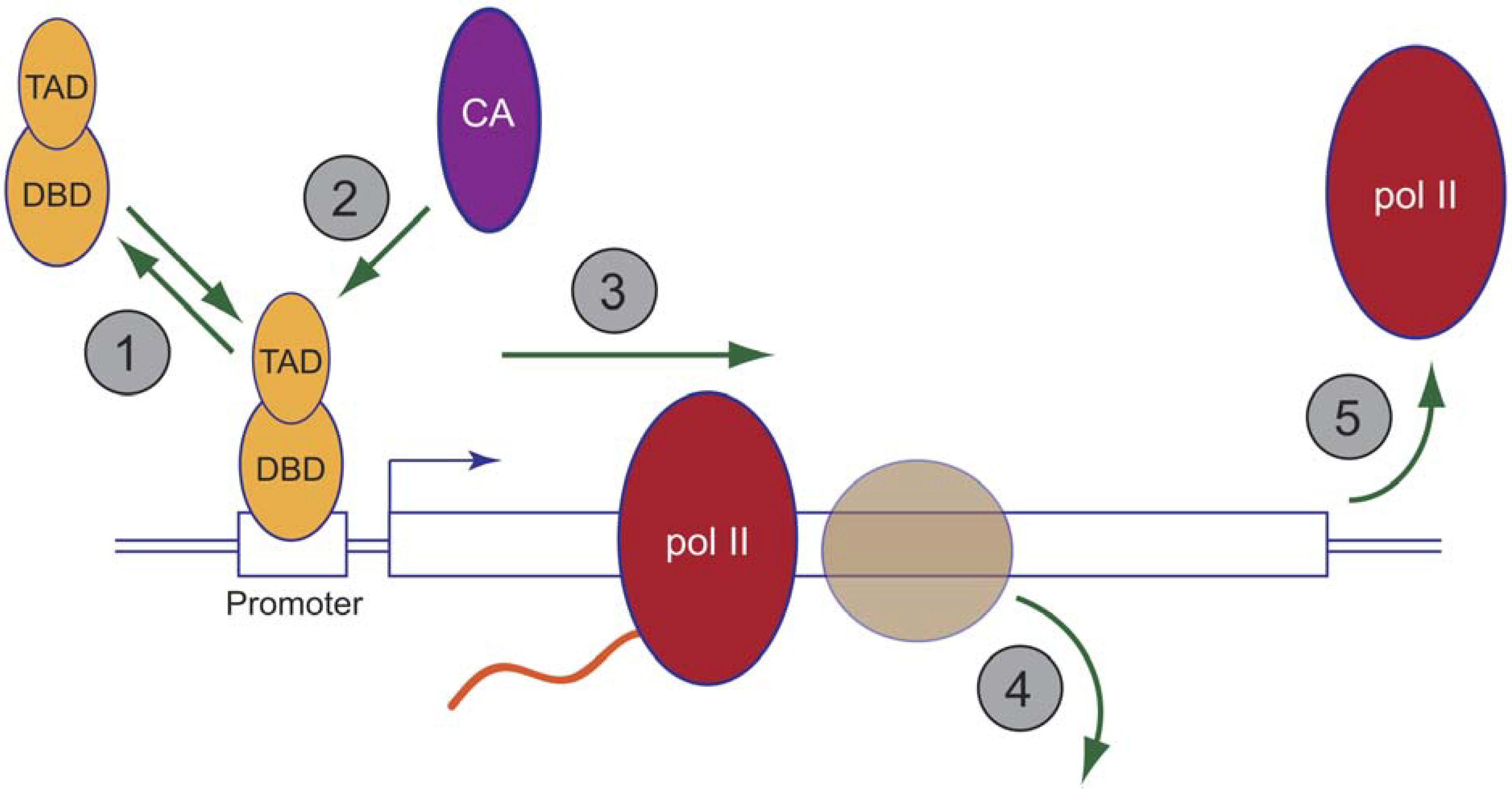
8. The Proteasome and Chromatin Modifiers
9. Cryptic Transcription
10. Conclusions and Perspectives
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Geng, F.; Wenzel, S.; Tansey, W.P. Ubiquitin and proteasomes in transcription. Annu. Rev. Biochem. 2012, 81, 177–201. [Google Scholar] [CrossRef] [PubMed]
- Muratani, M.; Tansey, W.P. How the ubiquitin-proteasome system controls transcription. Nat. Rev. Mol. Cell Biol. 2003, 4, 192–201. [Google Scholar] [CrossRef] [PubMed]
- Van Cuijk, L.; Vermeulen, W.; Marteijn, J.A. Ubiquitin at work: The ubiquitous regulation of the damage recognition step of NER. Exp. Cell Res. 2014. [Google Scholar] [CrossRef]
- Yao, T.; Ndoja, A. Regulation of gene expression by the ubiquitin-proteasome system. Semin. Cell Dev. Biol. 2012, 23, 523–529. [Google Scholar] [CrossRef] [PubMed]
- Fuchs, G.; Oren, M. Writing and reading H2B monoubiquitylation. Biochim. Biophys. Acta 2014, 1839, 694–701. [Google Scholar] [CrossRef] [PubMed]
- Annunziato, A.T. DNA packaging: Nucleosomes and chromatin. Nat. Educ. 2008, 1, 26. [Google Scholar]
- Margueron, R.; Reinberg, D. Chromatin structure and the inheritance of epigenetic information. Nat. Rev. Genet. 2010, 11, 285–296. [Google Scholar] [CrossRef] [PubMed]
- Gardner, K.E.; Allis, C.D.; Strahl, B.D. Operating on chromatin, a colorful language where context matters. J. Mol. Biol. 2011, 409, 36–46. [Google Scholar] [CrossRef] [PubMed]
- Swygert, S.G.; Peterson, C.L. Chromatin dynamics: Interplay between remodeling enzymes and histone modifications. Biochim. Biophys. Acta 2014, 1839, 728–736. [Google Scholar] [CrossRef] [PubMed]
- Bannister, A.J.; Kouzarides, T. Regulation of chromatin by histone modifications. Cell Res. 2011, 21, 381–395. [Google Scholar] [CrossRef] [PubMed]
- Shogren-Knaak, M.; Ishii, H.; Sun, J.M.; Pazin, M.J.; Davie, J.R.; Peterson, C.L. Histone H4-K16 acetylation controls chromatin structure and protein interactions. Science 2006, 311, 844–847. [Google Scholar] [CrossRef] [PubMed]
- Hargreaves, D.C.; Horng, T.; Medzhitov, R. Control of inducible gene expression by signal-dependent transcriptional elongation. Cell 2009, 138, 129–145. [Google Scholar] [CrossRef] [PubMed]
- Johnson, D.G.; Dent, S.Y. Chromatin: Receiver and quarterback for cellular signals. Cell 2013, 152, 685–689. [Google Scholar] [CrossRef] [PubMed]
- Biterge, B.; Schneider, R. Histone variants: Key players of chromatin. Cell Tissue Res. 2014, 356, 457–466. [Google Scholar] [CrossRef] [PubMed]
- Goldknopf, I.L.; Taylor, C.W.; Baum, R.M.; Yeoman, L.C.; Olson, M.O.; Prestayko, A.W.; Busch, H. Isolation and characterization of protein A24, a “histone-like” non-histone chromosomal protein. J. Biol. Chem. 1975, 250, 7182–7187. [Google Scholar] [PubMed]
- Levinger, L.; Varshavsky, A. Selective arrangement of ubiquitinated and D1 protein-containing nucleosomes within the drosophila genome. Cell 1982, 28, 375–385. [Google Scholar] [CrossRef] [PubMed]
- Domae, N.; Harmon, F.R.; Busch, R.K.; Spohn, W.; Subrahmanyam, C.S.; Busch, H. Donut-shaped “miniparticles” in nuclei of human and rat cells. Life Sci. 1982, 30, 469–477. [Google Scholar] [CrossRef] [PubMed]
- Swaffield, J.C.; Bromberg, J.F.; Johnston, S.A. Alterations in a yeast protein resembling HIV Tat-binding protein relieve requirement for an acidic activation domain in GAL4. Nature 1992, 357, 698–700. [Google Scholar] [CrossRef] [PubMed]
- Enenkel, C. Proteasome dynamics. Biochim. Biophys. Acta 2014, 1843, 39–46. [Google Scholar] [CrossRef] [PubMed]
- Palmer, A.; Rivett, A.J.; Thomson, S.; Hendil, K.B.; Butcher, G.W.; Fuertes, G.; Knecht, E. Subpopulations of proteasomes in rat liver nuclei, microsomes and cytosol. Biochem. J. 1996, 316, 401–407. [Google Scholar] [PubMed]
- Geng, F.; Tansey, W.P. Similar temporal and spatial recruitment of native 19S and 20S proteasome subunits to transcriptionally active chromatin. Proc. Natl. Acad. Sci. USA 2012, 109, 6060–6065. [Google Scholar] [CrossRef] [PubMed]
- Ezhkova, E.; Tansey, W.P. Proteasomal atpases link ubiquitylation of histone H2B to methylation of histone H3. Mol. Cell 2004, 13, 435–442. [Google Scholar] [CrossRef] [PubMed]
- Auld, K.L.; Brown, C.R.; Casolari, J.M.; Komili, S.; Silver, P.A. Genomic association of the proteasome demonstrates overlapping gene regulatory activity with transcription factor substrates. Mol. Cell 2006, 21, 861–871. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, F.; Delahodde, A.; Kodadek, T.; Johnston, S.A. Recruitment of a 19S proteasome subcomplex to an activated promoter. Science 2002, 296, 548–550. [Google Scholar] [CrossRef] [PubMed]
- Kitagawa, T.; Ishii, K.; Takeda, K.; Matsumoto, T. The 19S proteasome subunit Rpt3 regulates distribution of CENP-A by associating with centromeric chromatin. Nat. Commun. 2014, 5, 3597. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Krogan, N.J.; Lam, M.H.; Fillingham, J.; Keogh, M.C.; Gebbia, M.; Li, J.; Datta, N.; Cagney, G.; Buratowski, S.; Emili, A.; et al. Proteasome involvement in the repair of DNA double-strand breaks. Mol. Cell 2004, 16, 1027–1034. [Google Scholar] [CrossRef] [PubMed]
- Catic, A.; Suh, C.Y.; Hill, C.T.; Daheron, L.; Henkel, T.; Orford, K.W.; Dombkowski, D.M.; Liu, T.; Liu, X.S.; Scadden, D.T. Genome-wide map of nuclear protein degradation shows Ncor1 turnover as a key to mitochondrial gene regulation. Cell 2013, 155, 1380–1395. [Google Scholar] [CrossRef] [PubMed]
- Voges, D.; Zwickl, P.; Baumeister, W. The 26S proteasome: A molecular machine designed for controlled proteolysis. Annu. Rev. Biochem. 1999, 68, 1015–1068. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, M.; Haas, W.; Crosas, B.; Santamaria, P.G.; Gygi, S.P.; Walz, T.; Finley, D. The heat repeat protein Blm10 regulates the yeast proteasome by capping the core particle. Nat. Struct. Mol. Biol. 2005, 12, 294–303. [Google Scholar] [CrossRef] [PubMed]
- Pick, E.; Berman, T.S. Formation of alternative proteasomes: Same lady, different cap? FEBS Lett. 2013, 587, 389–393. [Google Scholar] [CrossRef]
- Nathan, J.A.; Spinnenhirn, V.; Schmidtke, G.; Basler, M.; Groettrup, M.; Goldberg, A.L. Immuno- and constitutive proteasomes do not differ in their abilities to degrade ubiquitinated proteins. Cell 2013, 152, 1184–1194. [Google Scholar] [CrossRef] [PubMed]
- Furey, T.S. ChIP-seq and beyond: New and improved methodologies to detect and characterize protein-DNA interactions. Nat. Rev. Genet. 2012, 13, 840–852. [Google Scholar] [CrossRef] [PubMed]
- Sikder, D.; Johnston, S.A.; Kodadek, T. Widespread, but non-identical, association of proteasomal 19 and 20S proteins with yeast chromatin. J. Biol. Chem. 2006, 281, 27346–27355. [Google Scholar] [CrossRef] [PubMed]
- Von der Lehr, N.; Johansson, S.; Wu, S.; Bahram, F.; Castell, A.; Cetinkaya, C.; Hydbring, P.; Weidung, I.; Nakayama, K.; Nakayama, K.I.; et al. The F-box protein Skp2 participates in c-Myc proteosomal degradation and acts as a cofactor for c-Myc-regulated transcription. Mol. Cell 2003, 11, 1189–1200. [Google Scholar] [CrossRef] [PubMed]
- Szutorisz, H.; Georgiou, A.; Tora, L.; Dillon, N. The proteasome restricts permissive transcription at tissue-specific gene loci in embryonic stem cells. Cell 2006, 127, 1375–1388. [Google Scholar] [CrossRef] [PubMed]
- Malik, S.; Shukla, A.; Sen, P.; Bhaumik, S.R. The 19S proteasome subcomplex establishes a specific protein interaction network at the promoter for stimulated transcriptional initiation in vivo. J. Biol. Chem. 2009, 284, 35714–35724. [Google Scholar] [CrossRef] [PubMed]
- Uprety, B.; Lahudkar, S.; Malik, S.; Bhaumik, S.R. The 19S proteasome subcomplex promotes the targeting of NuA4 HAT to the promoters of ribosomal protein genes to facilitate the recruitment of TFIID for transcriptional initiation in vivo. Nucleic Acids Res. 2012, 40, 1969–1983. [Google Scholar] [CrossRef] [PubMed]
- Satoh, T.; Ishizuka, T.; Yoshino, S.; Tomaru, T.; Nakajima, Y.; Shibusawa, N.; Hashimoto, K.; Yamada, M.; Mori, M. Roles of proteasomal 19S regulatory particles in promoter loading of thyroid hormone receptor. Biochem. Biophys. Res. Commun. 2009, 386, 697–702. [Google Scholar] [CrossRef] [PubMed]
- Schwarz, T.; Sohn, C.; Kaiser, B.; Jensen, E.D.; Mansky, K.C. The 19S proteasomal lid subunit POH1 enhances the transcriptional activation by Mitf in osteoclasts. J. Cell. Biochem. 2010, 109, 967–974. [Google Scholar] [PubMed]
- Ferry, C.; Gianni, M.; Lalevee, S.; Bruck, N.; Plassat, J.L.; Raska, I., Jr.; Garattini, E.; Rochette-Egly, C. Sug-1 plays proteolytic and non-proteolytic roles in the control of retinoic acid target genes via its interaction with Src-3. J. Biol. Chem. 2009, 284, 8127–8135. [Google Scholar] [CrossRef] [PubMed]
- Higazi, A.; Abed, M.; Chen, J.; Li, Q. Promoter context determines the role of proteasome in ligand-dependent occupancy of retinoic acid responsive elements. Epigenetics 2011, 6, 202–211. [Google Scholar] [CrossRef] [PubMed]
- Gillette, T.G.; Gonzalez, F.; Delahodde, A.; Johnston, S.A.; Kodadek, T. Physical and functional association of RNA polymerase II and the proteasome. Proc. Natl. Acad. Sci. USA 2004, 101, 5904–5909. [Google Scholar] [CrossRef] [PubMed]
- Lassot, I.; Latreille, D.; Rousset, E.; Sourisseau, M.; Linares, L.K.; Chable-Bessia, C.; Coux, O.; Benkirane, M.; Kiernan, R.E. The proteasome regulates HIV-1 transcription by both proteolytic and nonproteolytic mechanisms. Mol. Cell 2007, 25, 369–383. [Google Scholar] [CrossRef] [PubMed]
- Venkatasubrahmanyam, S.; Hwang, W.W.; Meneghini, M.D.; Tong, A.H.; Madhani, H.D. Genome-wide, as opposed to local, antisilencing is mediated redundantly by the euchromatic factors Set1 and H2A. Z. Proc. Natl. Acad. Sci. USA 2007, 104, 16609–16614. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.; Gonzalez, F.; Rothermel, B.; Sun, L.; Johnston, S.A.; Kodadek, T. The Gal4 activation domain binds Sug2 protein, a proteasome component, in vivo and in vitro. J. Biol. Chem. 2001, 276, 30956–30963. [Google Scholar] [CrossRef] [PubMed]
- Chaves, S.; Baskerville, C.; Yu, V.; Reed, S.I. Cks1, Cdk1, and the 19S proteasome collaborate to regulate gene induction-dependent nucleosome eviction in yeast. Mol. Cell. Biol. 2010, 30, 5284–5294. [Google Scholar] [CrossRef] [PubMed]
- Qian, M.X.; Pang, Y.; Liu, C.H.; Haratake, K.; Du, B.Y.; Ji, D.Y.; Wang, G.F.; Zhu, Q.Q.; Song, W.; Yu, Y.; et al. Acetylation-mediated proteasomal degradation of core histones during DNA repair and spermatogenesis. Cell 2013, 153, 1012–1024. [Google Scholar] [CrossRef] [PubMed]
- Wilcox, A.J.; Laney, J.D. A ubiquitin-selective AAA-ATPase mediates transcriptional switching by remodelling a repressor-promoter DNA complex. Nat. Cell Biol. 2009, 11, 1481–1486. [Google Scholar] [CrossRef] [PubMed]
- Verma, R.; Oania, R.; Fang, R.; Smith, G.T.; Deshaies, R.J. Cdc48/p97 mediates UV-dependent turnover of RNA pol II. Mol. Cell 2011, 41, 82–92. [Google Scholar] [CrossRef] [PubMed]
- Ndoja, A.; Cohen, R.E.; Yao, T. Ubiquitin signals proteolysis-independent stripping of transcription factors. Mol. Cell 2014, 53, 893–903. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Johnston, S.A.; Kodadek, T. Physical association of the APIS complex and general transcription factors. Biochem. Biophys. Res. Commun. 2002, 296, 991–999. [Google Scholar] [CrossRef] [PubMed]
- Russell, S.J.; Johnston, S.A. Evidence that proteolysis of Gal4 cannot explain the transcriptional effects of proteasome ATPase mutations. J. Biol. Chem. 2001, 276, 9825–9831. [Google Scholar] [CrossRef] [PubMed]
- Nalley, K.; Johnston, S.A.; Kodadek, T. Proteolytic turnover of the Gal4 transcription factor is not required for function in vivo. Nature 2006, 442, 1054–1057. [Google Scholar] [CrossRef] [PubMed]
- Collins, G.A.; Lipford, J.R.; Deshaies, R.J.; Tansey, W.P. Gal4 turnover and transcription activation. Nature 2009. [Google Scholar] [CrossRef]
- Collins, G.A.; Gomez, T.A.; Deshaies, R.J.; Tansey, W.P. Combined chemical and genetic approach to inhibit proteolysis by the proteasome. Yeast 2010, 27, 965–974. [Google Scholar] [CrossRef] [PubMed]
- Collins, G.A.; Tansey, W.P. The proteasome: A utility tool for transcription? Curr. Opin. Genet. Dev. 2006, 16, 197–202. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Muratani, M.; Tansey, W.P.; Ptashne, M. Proteolytic instability and the action of nonclassical transcriptional activators. Curr. Biol. 2010, 20, 868–871. [Google Scholar] [CrossRef] [PubMed]
- Navon, A.; Goldberg, A.L. Proteins are unfolded on the surface of the ATPase ring before transport into the proteasome. Mol. Cell 2001, 8, 1339–1349. [Google Scholar] [CrossRef] [PubMed]
- Nishiyama, A.; Tachibana, K.; Igarashi, Y.; Yasuda, H.; Tanahashi, N.; Tanaka, K.; Ohsumi, K.; Kishimoto, T. A nonproteolytic function of the proteasome is required for the dissociation of Cdc2 and cyclin B at the end of M phase. Genes Dev. 2000, 14, 2344–2357. [Google Scholar] [CrossRef] [PubMed]
- Kwak, H.; Lis, J.T. Control of transcriptional elongation. Annu. Rev. Genet. 2013, 47, 483–508. [Google Scholar] [CrossRef] [PubMed]
- Swaffield, J.C.; Melcher, K.; Johnston, S.A. A highly conserved ATPase protein as a mediator between acidic activation domains and the TATA-binding protein. Nature 1995, 374, 88–91. [Google Scholar] [CrossRef] [PubMed]
- Xu, Q.; Singer, R.A.; Johnston, G.C. Sug1 modulates yeast transcription activation by Cdc68. Mol. Cell. Biol. 1995, 15, 6025–6035. [Google Scholar] [PubMed]
- Russell, S.J.; Sathyanarayana, U.G.; Johnston, S.A. Isolation and characterization of Sug2. A novel ATPase family component of the yeast 26S proteasome. J. Biol. Chem. 1996, 271, 32810–32817. [Google Scholar] [CrossRef] [PubMed]
- Ferdous, A.; Sikder, D.; Gillette, T.; Nalley, K.; Kodadek, T.; Johnston, S.A. The role of the proteasomal ATPases and activator monoubiquitylation in regulating Gal4 binding to promoters. Genes Dev. 2007, 21, 112–123. [Google Scholar] [CrossRef] [PubMed]
- Muratani, M.; Kung, C.; Shokat, K.M.; Tansey, W.P. The F box protein Dsg1/Mdm30 is a transcriptional coactivator that stimulates Gal4 turnover and cotranscriptional mRNA processing. Cell 2005, 120, 887–899. [Google Scholar] [CrossRef] [PubMed]
- Stavreva, D.A.; Muller, W.G.; Hager, G.L.; Smith, C.L.; McNally, J.G. Rapid glucocorticoid receptor exchange at a promoter is coupled to transcription and regulated by chaperones and proteasomes. Mol. Cell. Biol. 2004, 24, 2682–2697. [Google Scholar] [CrossRef] [PubMed]
- Koutelou, E.; Hirsch, C.L.; Dent, S.Y. Multiple faces of the SAGA complex. Curr. Opin. Cell Biol. 2010, 22, 374–382. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.; Ezhkova, E.; Li, B.; Pattenden, S.G.; Tansey, W.P.; Workman, J.L. The proteasome regulatory particle alters the SAGA coactivator to enhance its interactions with transcriptional activators. Cell 2005, 123, 423–436. [Google Scholar] [CrossRef] [PubMed]
- Geng, F.G.; Tansey, W.P.; Department of Cell and Developmental Biology, Vanderbilt University School of Medicine, Nashville, TN, USA. Unpublished work. 2014.
- Ferdous, A.; Gonzalez, F.; Sun, L.; Kodadek, T.; Johnston, S.A. The 19S regulatory particle of the proteasome is required for efficient transcription elongation by RNA polymerase II. Mol. Cell 2001, 7, 981–991. [Google Scholar] [CrossRef] [PubMed]
- Pan, Y.R.; Sun, M.; Wohlschlegel, J.; Reed, S.I. Cks1 enhances transcription efficiency at the GAL1 locus by linking the Paf1 complex to the 19S proteasome. Eukaryot. Cell 2013, 12, 1192–1201. [Google Scholar] [CrossRef] [PubMed]
- Ransom, M.; Williams, S.K.; Dechassa, M.L.; Das, C.; Linger, J.; Adkins, M.; Liu, C.; Bartholomew, B.; Tyler, J.K. Fact and the proteasome promote promoter chromatin disassembly and transcriptional initiation. J. Biol. Chem. 2009, 284, 23461–23471. [Google Scholar] [CrossRef] [PubMed]
- Sheldon, K.E.; Mauger, D.M.; Arndt, K.M. A requirement for the Saccharomyces cerevisiae Paf1 complex in snoRNA 3' end formation. Mol. Cell 2005, 20, 225–236. [Google Scholar] [CrossRef] [PubMed]
- Penheiter, K.L.; Washburn, T.M.; Porter, S.E.; Hoffman, M.G.; Jaehning, J.A. A posttranscriptional role for the yeast Paf1-RNA polymerase II complex is revealed by identification of primary targets. Mol. Cell 2005, 20, 213–223. [Google Scholar] [CrossRef] [PubMed]
- Fatyol, K.; Grummt, I. Proteasomal ATPases are associated with RDNa: The ubiquitin proteasome system plays a direct role in RNA polymerase I transcription. Biochim. Biophys. Acta 2008, 1779, 850–859. [Google Scholar] [CrossRef] [PubMed]
- Lim, S.; Kwak, J.; Kim, M.; Lee, D. Separation of a functional deubiquitylating module from the SAGA complex by the proteasome regulatory particle. Nat. Commun. 2013, 4, 2641. [Google Scholar] [PubMed]
- Kohler, A.; Schneider, M.; Cabal, G.G.; Nehrbass, U.; Hurt, E. Yeast Ataxin-7 links histone deubiquitination with gene gating and mRNA export. Nat. Cell Biol. 2008, 10, 707–715. [Google Scholar] [CrossRef] [PubMed]
- Holzl, H.; Kapelari, B.; Kellermann, J.; Seemuller, E.; Sumegi, M.; Udvardy, A.; Medalia, O.; Sperling, J.; Muller, S.A.; Engel, A.; et al. The regulatory complex of drosophila melanogaster 26S proteasomes. Subunit composition and localization of a deubiquitylating enzyme. J. Cell Biol. 2000, 150, 119–130. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yao, T.; Song, L.; Jin, J.; Cai, Y.; Takahashi, H.; Swanson, S.K.; Washburn, M.P.; Florens, L.; Conaway, R.C.; Cohen, R.E.; et al. Distinct modes of regulation of the Uch37 deubiquitinating enzyme in the proteasome and in the Ino80 chromatin-remodeling complex. Mol. Cell 2008, 31, 909–917. [Google Scholar] [CrossRef] [PubMed]
- Cai, Y.; Jin, J.; Yao, T.; Gottschalk, A.J.; Swanson, S.K.; Wu, S.; Shi, Y.; Washburn, M.P.; Florens, L.; Conaway, R.C.; et al. Yy1 functions with Ino80 to activate transcription. Nat. Struct. Mol. Biol. 2007, 14, 872–874. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Shi, Y.; Mulligan, P.; Gay, F.; Landry, J.; Liu, H.; Lu, J.; Qi, H.H.; Wang, W.; Nickoloff, J.A.; et al. A YY1-Ino80 complex regulates genomic stability through homologous recombination-based repair. Nat. Struct. Mol. Biol. 2007, 14, 1165–1172. [Google Scholar] [CrossRef] [PubMed]
- Berretta, J.; Morillon, A. Pervasive transcription constitutes a new level of eukaryotic genome regulation. EMBO Rep. 2009, 10, 973–982. [Google Scholar] [CrossRef] [PubMed]
- Kaplan, C.D.; Laprade, L.; Winston, F. Transcription elongation factors repress transcription initiation from cryptic sites. Science 2003, 301, 1096–1099. [Google Scholar] [CrossRef] [PubMed]
- Cheung, V.; Chua, G.; Batada, N.N.; Landry, C.R.; Michnick, S.W.; Hughes, T.R.; Winston, F. Chromatin- and transcription-related factors repress transcription from within coding regions throughout the Saccharomyces cerevisiae genome. PLoS Biol. 2008, 6, e277. [Google Scholar] [PubMed]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
McCann, T.S.; Tansey, W.P. Functions of the Proteasome on Chromatin. Biomolecules 2014, 4, 1026-1044. https://doi.org/10.3390/biom4041026
McCann TS, Tansey WP. Functions of the Proteasome on Chromatin. Biomolecules. 2014; 4(4):1026-1044. https://doi.org/10.3390/biom4041026
Chicago/Turabian StyleMcCann, Tyler S., and William P. Tansey. 2014. "Functions of the Proteasome on Chromatin" Biomolecules 4, no. 4: 1026-1044. https://doi.org/10.3390/biom4041026



