Genetic Fingerprinting of Wheat and Its Progenitors by Mitochondrial Gene orf256
Abstract
:1. Introduction
2. Experimental Section
2.1. DNA Primer
| Primer Name | Sequence 5'  3' 3' |
|---|---|
| 1.5: −94 | CCA TAT TCA CGC AAC TGA T |
| 2.5: −215 | CTA CGA GAT CAC CTT CAC G |
| 3.5: −190 | CTG AGC CTT TAC GAG CAG G |
| 4.5: 35 | GCA GGT TTA CTG CTT TC |
| 5.5':253 | CTGAGCCTTTACGAGCAGG |
| 6.C:785 | TCA GAA TTA CTG AGC TAC |
| 7.C:477 | GGA ACG AAG CGC TTC ATC GA |
| 8.C: 219 | GCT TGG GGA TCC TGA ATC |
| 9.C:482 | GAG ATG CTG TTT CCC ACA AC |
| 10.C:980 | ATA GAG AGT CCC AAT ATC C |
| 11.C:1469 | GCT GTC ACT AGA ACG GAC C |
2.2. Growth of Wheat Shoots
2.3. Freeze drying of Wheat Shoots
2.4. Isolation of Mitochondria
2.5. Isolation of DNA
| Species | Description | Ploidy level | Notes |
|---|---|---|---|
| Triticum aestivum (Ta) | Hexaploid wheat | AABBDD | |
| Triticum timopheevii (Tt) | wheat progenitor | GGAA | Possible source of B (G = B) genome |
| Triticum turgidum (Ttu) | wheat progenitor | BBAA | S = B |
| Triticum monococcum monocuccom (Tm) | wheat progenitor | AA | Source of A genome |
| Triticum monocuccum bioeticum (Tb) | wheat progenitor | AA | Source of A genome |
| Aegilops speltoides (Asp) | wheat progenitor | SS | S = B |
| Aegilops bicornis (Ab) | wheat progenitor | SS | |
| Aegilops searsii (Ase) | wheat progenitor | SS | |
| Aegilops taushii (At) | wheat progenitor | DD | Source of D genome |
| Aegilops sharonensis (Ash) | wheat progenitor | SS | |
| Secale cereale (Sc) | wheat relative |
2.6. PCR Amplification
2.7. Sequencing of the PCR Amplified Fragments
2.8. Sequence Alignment and Phylogenetic Relationship
3. Results and Discussion
3.1. PCR
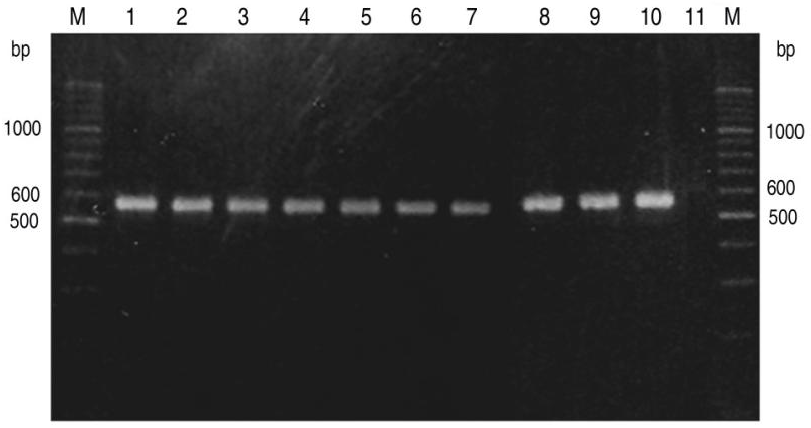

3.2. Sequencing and Bioinformatic Analysis of PCR Products
| Sequence obtained | Species | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Ttu | Tt | Ase | Ab | Ash | Asp | Tm | Tb | At | Sc | ||
| Primers | 5: −94/C:482 | 576 | 576 | 576 | 576 | 576 | 576 | 576 | 576 | 576 | 576 |
| 5:253/C:482 | 230 | 230 | 230 | 230 | 230 | 230 | 230 | 230 | 230 | 230 | |
| Number of base substitutions | 49 | 49 | 50 | 47 | 47 | 47 | 25 | 22 | 21 | - | |

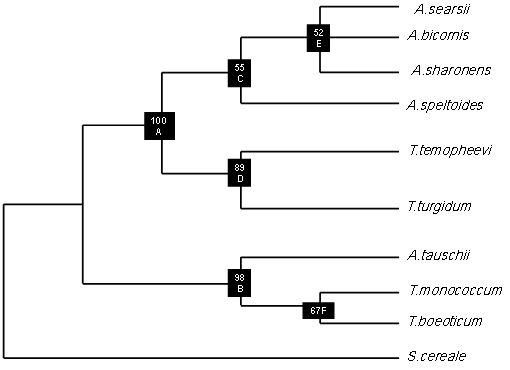
3.3. Phylogenetic Analysis
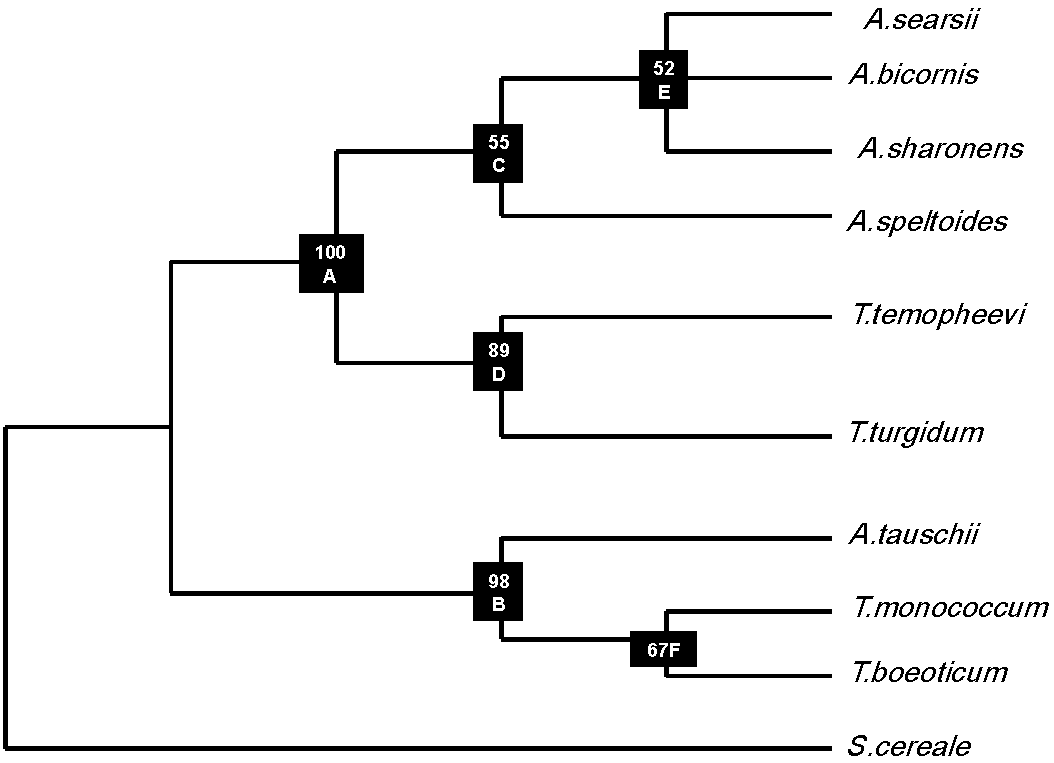
3.4. Distance Matrix
| Species | Ttu | Tt | Ase | Ab | Ash | Asp | Tm | Tb | At | Sc |
|---|---|---|---|---|---|---|---|---|---|---|
| T.turgidum | 0.00 | |||||||||
| T.temopheevii | 0.20 | 0.00 | ||||||||
| A.searsii | 3.06 | 2.85 | 0.00 | |||||||
| A.bicornis | 2.85 | 2.65 | 0.20 | 0.00 | ||||||
| A.sharonensis | 3.06 | 2.85 | 0.40 | 0.20 | 0.00 | |||||
| A.speltoides | 2.44 | 2.23 | 1.41 | 1.21 | 1.41 | 0.00 | ||||
| T.monococcum | 32.85 | 33.16 | 33.16 | 32.85 | 32.54 | 33.16 | 0.00 | |||
| T.boeoticum | 32.85 | 33.16 | 32.85 | 32.54 | 32.23 | 33.16 | 0.40 | 0.00 | ||
| A.tauschii | 33.47 | 33.78 | 33.47 | 33.16 | 32.85 | 33.78 | 1.82 | 1.41 | 0.00 | |
| S.cereale | 32.23 | 32.54 | 32.23 | 31.93 | 31.62 | 32.54 | 4.53 | 4.11 | 4.11 | 0.00 |
4. Conclusions
Acknowledgments
References
- Cox, T.S. Deepening the wheat gene pool. J. Crop Prod. 1998, 1. [Google Scholar] [CrossRef]
- Dvorak, J.; DiTerlizzi, P.; Zhang, H.B.; Resta, P. The evolution of polyploid wheats: Identification of the A genome donor species. Genome 1993, 36, 21–31. [Google Scholar]
- Wheat Genetic Resource Center. Available online: www.ksu.edu/wgrc (accessed on 19 May 2010).
- Huang, S.; Sirikhachornkit, A.; Su, X.; Faris, J.; Gill, B.; Haselkorn, R.; Gornicki, P. Phylogenetic analysis of the acetyl-CoA carboxylase and 3-phosphoglycerate kinase loci in wheat and other grasses. Plant Mol. Biol. 2002, 48, 805–820. [Google Scholar]
- McFadden, E.S.; Sears, E.R. The origin of Triticum speltoides and its free-threshing hexaploid relatives. J. Hered. 1946, 37, 81–89. [Google Scholar]
- Kimber, G.; Sears, E.R. Evolution in the Genus Triticum and the Origin of Cultivated Whea. In Wheat and Wheat Improvement; Heyne, E.G., Ed.; American Society of Agronomy: Madison, WI, USA, 1987; pp. 154–164. [Google Scholar]
- Friebe, B.; Gill, B.S. Chromosome Banding and Genome Analysis in Diploid and Cultivated Polyploid Wheats. In Methods of Genome Analysis in Plants; CRC Press: Boca Raton, FL, USA, 1996; pp. 39–60. [Google Scholar]
- Sasanuma, T.; Miyashita, N.T.; Tsunewaki, K. Wheat phylogeny determined by RFLP analysis of nuclear DNA. 3. Intra- and interspecific variations of five Aegilops Sitopsis species. Theor. Appl. Genet. 1996, 92, 928–934. [Google Scholar] [CrossRef]
- Wolfe, K.H.; Gouy, M.; Yang, Y.W.; Sharp, P.M.; Li, W.H. Date of the monocot-dicot divergence estimated from chloroplast DNA sequence data. Proc. Natl. Acad. Sci. USA 1989, 86, 6201–6205. [Google Scholar]
- Doeble, J.; Durbin, M.; Golenberg, E.M.; Clegg, M.T.; Ma, D.P. Evolutionary analysis of the large subunit of carboxylase (rbcL) nucleotide sequence among the grasses (Gramineae). Evolution 1990, 4, 1097–1108. [Google Scholar]
- Clark, L.G.; Zhang, W.; Wendel, J.F. A phylogeny of the grass family (Poaceae) based on sequence data. Syst. Bot. 1995, 20, 436–460. [Google Scholar]
- Gaut, B.S.; Doeble, J. DNA sequence evidence for the segmental allotetraploid origin of maize (duplicated lociychromosomal evolution). Proc. Natl. Acad. Sci. USA 1997, 94, 6809–6814. [Google Scholar]
- Hedgcoth, C.; El-Shehawi, A.M.; Wei, P.; Clardson, M.; Tamalis, D. A chimeric open reading frame associated with cytoplasmic male sterility in alloplasmic wheat with Triticum temopheevii mitochondria is present in several Triticum and Aegilops species, barley, and rye. Curr. Genet. 2002, 41, 357–365. [Google Scholar] [CrossRef]
- Tsunewaki, K. Plasmon Analysis as the Counterpart of Genome Analysis. In Methods of Genome Analysis in Plants; CRC Press: Boca Raton, FL, USA, 1996; pp. 271–299. [Google Scholar]
- Dvorak, J.; Zhang, H.B. Variation in repeated nucleotide sequences sheds light on the phylogeny of the wheat B and G genomes. Proc. Natl. Acad. Sci. USA 1990, 87, 9640–9644. [Google Scholar]
- Tsunewaki, K.; Ogihara, Y. The Molecular Basis of Genetic Diversity among Cytoplasms of TRITICUM and AEGILOPS Species. II. on the Origin of Polyploid Wheat Cytoplasms as Suggested by Chloroplast DNA Restriction Fragment Patterns. Genetics 1983, 104, 155–171. [Google Scholar]
- Wang, G.Z.; Miyashita, N.T.; Tsunewaki, K. Plasmon analyses of Triticum (wheat) and Aegilops: PCR-single-strand conformational polymorphism (PCR-SSCP) analyses of organellar DNAs. Proc. Natl. Acad. Sci. USA 1997, 94, 14570–14577. [Google Scholar] [CrossRef]
- Gaut, B.S. Molecular clocks and nucleotide substitution rates in higher plants. Evol. Biol. 1998, 30, 93–120. [Google Scholar]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTALW: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucl. Acids Res. 1994, 22, 4673–4680. [Google Scholar]
- Faris, J.; Sirikhachornkit, A.; Haselkorn, R.; Gill, B.; Gornicki, P. Chromosome mapping and phylogenetic analysis of the cytosolic acetyl-CoA carboxylase loci in wheat. Mol. Biol. Evol. 2001, 18, 1720–1733. [Google Scholar]
- Feuillet, C.; Penger, A.; Gellner, K.; Mast, A.; Keller, B. Molecular evolution of receptor-like kinase genes in hexaploid wheat. Independent evolution of orthologs after polyploidization and mechanisms of local rearrangements at paralogous loci. Plant Physiol. 2001, 125, 1304–1313. [Google Scholar] [CrossRef]
- Rathburn, H.; Hedgcoth, C. A chimeric open reading frame in the 5' flanking region of coxI mitochondrial DNA from cytoplasmic male-sterile wheat. Plant Mol. Biol. 1991, 16, 909–912. [Google Scholar]
- Rathburn, H.; Song, J.; Hedgcoth, C. Cytoplasmic male sterility and fertility restoration in wheat are not associated with rearrangements of mitochondrial DNA in the gene regions for cob, coxII, or coxI. Plant Mol. Biol. 1993, 21, 195–201. [Google Scholar] [CrossRef]
- Macrogen Company. Available online: http://www.macrogen.com (accessed on 13 June 2010).
- Speakman, J.B.; Krueger, W. A comparison of methods to surface sterilize wheat seeds. Trans. Br. Mycol. Soc. 1983, 80, 374–376. [Google Scholar]
- Song, J.; Hedgcoth, C. A chimeric gene (orf256) is expressed as protein only in cytoplasmic male-sterile lines of wheat. Plant Mol. Biol. 1994, 26, 535–539. [Google Scholar] [CrossRef]
- CLUSTALW2 Software. Available online: http://www.ebi.ac.uk/Tools/msa/clustalw2/ (accessed on 22 April 2011).
- PHLIP software. Available online: http://bioweb2.pasteur.fr/phylogeny (accessed on 15 June 2011).
- Huang, S.; Sirikhachornkit, A.; Su, X.; Faris, J.; Gill, B.; Haselkorn, R.; Gornicki, P. Genes encoding plastid acetyl-CoA carboxylase and 3-phosphoglycerate kinase of Triticum/Aegilops complex and the evolutionary history of polyploid wheat. Proc. Natl. Aca. Sci. USA 2002, 99, 8133–8138. [Google Scholar]
- Wei, S.; Peng, Z.; Zhou, Y.; Yang, Z.; Wu, K.; Ouyang, Z. Nucleotide diversity and molecular evolution of the WAG-2 gene in common wheat (Triticum aestivum L) and its relatives. Genet. Mol. Biol. 2011, 34, 606–616. [Google Scholar]
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/3.0/).
Share and Cite
El-Shehawi, A.M.; Fahmi, A.I.; Sayed, S.M.; Elseehy, M.M. Genetic Fingerprinting of Wheat and Its Progenitors by Mitochondrial Gene orf256. Biomolecules 2012, 2, 228-239. https://doi.org/10.3390/biom2020228
El-Shehawi AM, Fahmi AI, Sayed SM, Elseehy MM. Genetic Fingerprinting of Wheat and Its Progenitors by Mitochondrial Gene orf256. Biomolecules. 2012; 2(2):228-239. https://doi.org/10.3390/biom2020228
Chicago/Turabian StyleEl-Shehawi, Ahmed M., Abdelmeguid I. Fahmi, Samy M. Sayed, and Mona M. Elseehy. 2012. "Genetic Fingerprinting of Wheat and Its Progenitors by Mitochondrial Gene orf256" Biomolecules 2, no. 2: 228-239. https://doi.org/10.3390/biom2020228
APA StyleEl-Shehawi, A. M., Fahmi, A. I., Sayed, S. M., & Elseehy, M. M. (2012). Genetic Fingerprinting of Wheat and Its Progenitors by Mitochondrial Gene orf256. Biomolecules, 2(2), 228-239. https://doi.org/10.3390/biom2020228