Robust Recombinant Expression of Human Placental Ribonuclease Inhibitor in Insect Cells
Abstract
:1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Transfection, Virus Propagation and Protein Expression
2.3. Purification of RI
2.4. Temperature Dependent RNase Activity Measurements
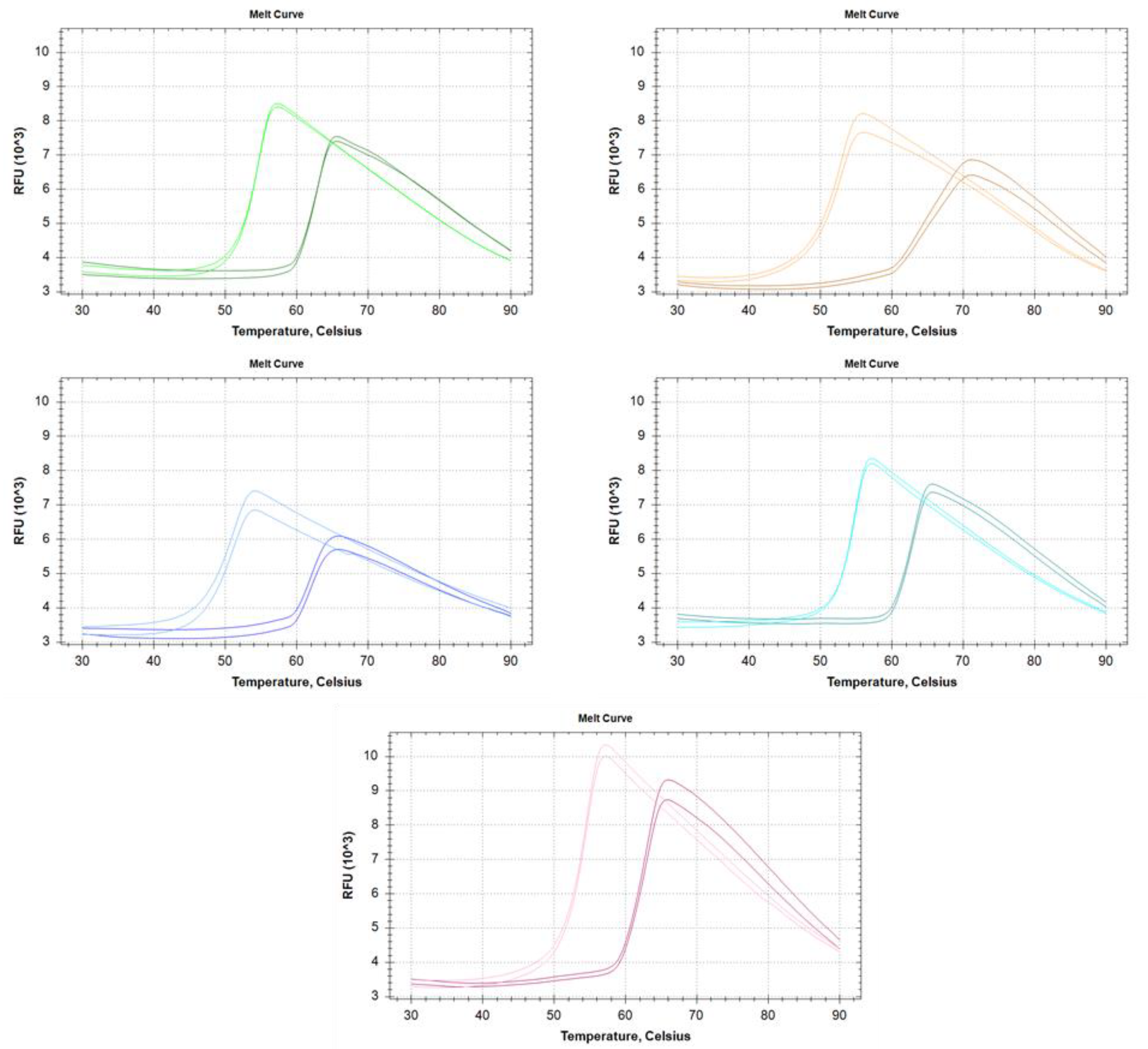
2.5. DSF Measurements
3. Results and Discussion
3.1. Design of Expression Plasmid
3.2. Expression of RI
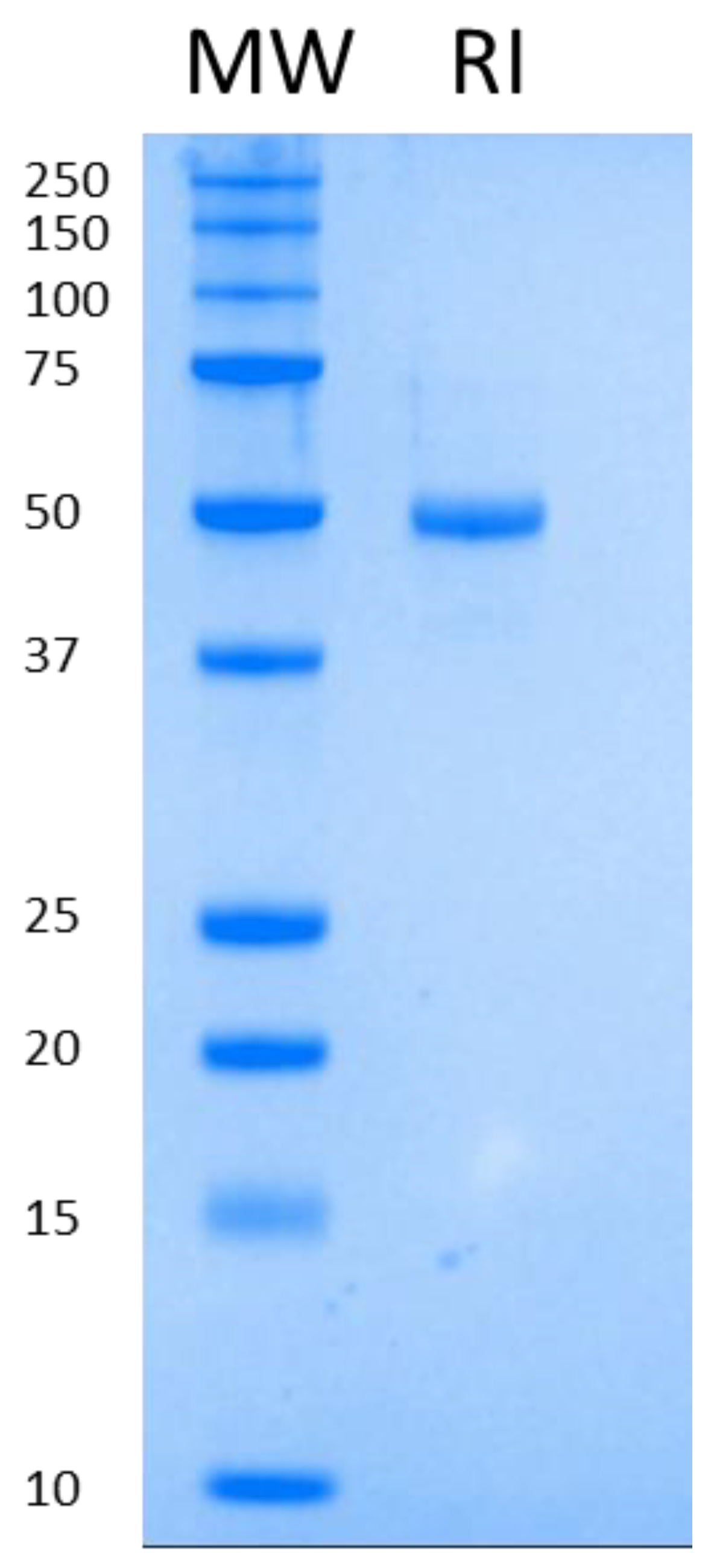
3.3. Purification
3.4. Heat Stability of RI
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kobe, B.; Deisenhofer, J. Crystal Structure of Porcine Ribonuclease Inhibitor, a Protein with Leucine-Rich Repeats. Nature 1993, 366, 751–756. [Google Scholar] [CrossRef] [PubMed]
- Leland, P.A.; Schultz, L.W.; Kim, B.M.; Raines, R.T. Ribonuclease A Variants with Potent Cytotoxic Activity. Proc. Natl. Acad. Sci. USA 1998, 95, 10407–10412. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Klink, T.A.; Vicentini, A.M.; Hofsteenge, J.; Raines, R.T. High-Level Soluble Production and Characterization of Porcine Ribonuclease Inhibitor. Protein. Expr. Purif. 2001, 22, 174–179. [Google Scholar] [CrossRef] [PubMed]
- Guo, W.; Cao, L.; Jia, Z.; Wu, G.; Li, T.; Lu, F.; Lu, Z. High Level Soluble Production of Functional Ribonuclease Inhibitor in Escherichia Coli by Fusing It to Soluble Partners. Protein. Expr. Purif. 2011, 77, 185–192. [Google Scholar] [CrossRef]
- Siurkus, J.; Neubauer, P. Reducing Conditions Are the Key for Efficient Production of Active Ribonuclease Inhibitor in Escherichia Coli. Microb. Cell Fact. 2011, 10, 31. [Google Scholar] [CrossRef] [Green Version]
- Vicentini, A.M.; Kieffer, B.; Matthies, R.; Meyhack, B.; Hemmings, B.A.; Stone, S.R.; Hofsteenge, J. Protein Chemical and Kinetic Characterization of Recombinant Porcine Ribonuclease Inhibitor Expressed in Saccharomyces Cerevisiae. Biochemistry 1990, 29, 8827–8834. [Google Scholar] [CrossRef]
- Park, J.-H.; Hwang, I.-S.; Kim, K.-I.; Lee, J.-M.; Park, Y.-M.; Park, C.-H.; Chung, I.S. Functional Expression of Recombinant Human Ribonuclease/Angiogenin Inhibitor in Stably Transformed Drosophila Melanogaster S2 Cells. Cytotechnology 2008, 57, 93–99. [Google Scholar] [CrossRef] [Green Version]
- Fominaya, J.M.; Hofsteenge, J. Inactivation of Ribonuclease Inhibitor by Thiol-Disulfide Exchange. J. Biol. Chem. 1992, 267, 24655–24660. [Google Scholar] [CrossRef]
- Kim, B.M.; Schultz, L.W.; Raines, R.T. Variants of Ribonuclease Inhibitor That Resist Oxidation. Protein. Sci. 1999, 8, 430–434. [Google Scholar] [CrossRef] [Green Version]
- Blackburn, P.; Wilson, G.; Moore, S. Ribonuclease Inhibitor from Human Placenta. Purification and Properties. J. Biol. Chem. 1977, 252, 5904–5910. [Google Scholar] [CrossRef]
- Lee, F.S.; Vallee, B.L. Expression of Human Placental Ribonuclease Inhibitor in Escherichia Coli. Biochem. Biophys. Res. Commun. 1989, 160, 115–120. [Google Scholar] [CrossRef]
- Panula-Perälä, J.; Siurkus, J.; Vasala, A.; Wilmanowski, R.; Casteleijn, M.G.; Neubauer, P. Enzyme Controlled Glucose Auto-Delivery for High Cell Density Cultivations in Microplates and Shake Flasks. Microb. Cell Fact. 2008, 7, 31. [Google Scholar] [CrossRef] [Green Version]
- Threat Assessment Brief: Emergence of SARS-CoV-2 B.1.617 Variants in India and Situation in the EU/EEA. Available online: https://www.ecdc.europa.eu/en/publications-data/threat-assessment-emergence-sars-cov-2-b1617-variants (accessed on 3 December 2021).
- Threat Assessment Brief: Implications of the Emergence and Spread of the SARS-CoV-2 B.1.1. 529 Variant of Concern (Omicron) for the EU/EEA. Available online: https://www.ecdc.europa.eu/en/publications-data/threat-assessment-brief-emergence-sars-cov-2-variant-b.1.1.529 (accessed on 3 December 2021).
- Jarvis, D.L. Baculovirus-Insect Cell Expression Systems. Methods Enzymol. 2009, 463, 191–222. [Google Scholar] [CrossRef] [PubMed]
- Ellis, E.L.; Delbrück, M. The Growth of Bacteriophage. J. Gen. Physiol. 1939, 22, 365–384. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hopkins, R.; Esposito, D. A Rapid Method for Titrating Baculovirus Stocks Using the Sf-9 Easy Titer Cell Line. Biotechniques 2009, 47, 785–788. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pantoliano, M.W.; Petrella, E.C.; Kwasnoski, J.D.; Lobanov, V.S.; Myslik, J.; Graf, E.; Carver, T.; Asel, E.; Springer, B.A.; Lane, P.; et al. High-Density Miniaturized Thermal Shift Assays as a General Strategy for Drug Discovery. J. Biomol. Screen 2001, 6, 429–440. [Google Scholar] [CrossRef] [PubMed]
- Peleg, Y.; Prabahar, V.; Bednarczyk, D.; Unger, T. Harnessing the Profinity EXactTM System for Expression and Purification of Heterologous Proteins in E. Coli. Methods Mol. Biol. 2017, 1586, 33–43. [Google Scholar] [CrossRef]
- Ruan, B.; Fisher, K.E.; Alexander, P.A.; Doroshko, V.; Bryan, P.N. Engineering Subtilisin into a Fluoride-Triggered Processing Protease Useful for One-Step Protein Purification. Biochemistry 2004, 43, 14539–14546. [Google Scholar] [CrossRef] [PubMed]
- Somogyi, M.; Szimler, T.; Baksa, A.; Végh, B.M.; Bakos, T.; Paréj, K.; Ádám, C.; Zsigmond, Á.; Megyeri, M.; Flachner, B.; et al. A Versatile Modular Vector Set for Optimizing Protein Expression among Bacterial, Yeast, Insect and Mammalian Hosts. PLoS ONE 2019, 14, e0227110. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hitchman, R.B.; Possee, R.D.; Crombie, A.T.; Chambers, A.; Ho, K.; Siaterli, E.; Lissina, O.; Sternard, H.; Novy, R.; Loomis, K.; et al. Genetic Modification of a Baculovirus Vector for Increased Expression in Insect Cells. Cell Biol. Toxicol. 2010, 26, 57–68. [Google Scholar] [CrossRef]
- Rieffel, S.; Roest, S.; Klopp, J.; Carnal, S.; Marti, S.; Gerhartz, B.; Shrestha, B. Insect Cell Culture in Reagent Bottles. MethodsX 2014, 1, 155–161. [Google Scholar] [CrossRef]
- Lomax, J.E.; Bianchetti, C.M.; Chang, A.; Phillips, G.N.; Fox, B.G.; Raines, R.T. Functional Evolution of Ribonuclease Inhibitor: Insights from Birds and Reptiles. J. Mol. Biol. 2014, 426, 3041–3056. [Google Scholar] [CrossRef] [Green Version]
- Sahin, U.; Muik, A.; Derhovanessian, E.; Vogler, I.; Kranz, L.M.; Vormehr, M.; Baum, A.; Pascal, K.; Quandt, J.; Maurus, D.; et al. COVID-19 Vaccine BNT162b1 Elicits Human Antibody and TH1 T Cell Responses. Nature 2020, 586, 594–599. [Google Scholar] [CrossRef]
- Schoenmaker, L.; Witzigmann, D.; Kulkarni, J.A.; Verbeke, R.; Kersten, G.; Jiskoot, W.; Crommelin, D.J.A. MRNA-Lipid Nanoparticle COVID-19 Vaccines: Structure and Stability. Int. J. Pharm. 2021, 601, 120586. [Google Scholar] [CrossRef]
- Borah, P.; Deb, P.K.; Al-Shar’i, N.A.; Dahabiyeh, L.A.; Venugopala, K.N.; Singh, V.; Shinu, P.; Hussain, S.; Deka, S.; Chandrasekaran, B.; et al. Perspectives on RNA Vaccine Candidates for COVID-19. Front. Mol. Biosci. 2021, 8, 30. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Kream, R.M.; Stefano, G.B. An Evidence Based Perspective on MRNA-SARS-CoV-2 Vaccine Development. Med. Sci. Monit. 2020, 26, e924700-1–e924700-8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huysmans, H.; Temmerman, J.D.; Zhong, Z.; Cafferty, S.M.; Combes, F.; Haesebrouck, F.; Sanders, N.N. Improving the Repeatability and Efficacy of Intradermal Electroporated Self-Replicating MRNA. Mol. Ther.—Nucleic Acids 2019, 17, 388–395. [Google Scholar] [CrossRef] [Green Version]
- Brito, L.A.; Singh, M. Acceptable Levels of Endotoxin in Vaccine Formulations during Preclinical Research. J. Pharm. Sci. 2011, 100, 34–37. [Google Scholar] [CrossRef]
- Whitley, J.; Zwolinski, C.; Denis, C.; Maughan, M.; Hayles, L.; Clarke, D.; Snare, M.; Liao, H.; Chiou, S.; Marmura, T.; et al. Development of MRNA Manufacturing for Vaccines and Therapeutics: MRNA Platform Requirements and Development of a Scalable Production Process to Support Early Phase Clinical Trials. Transl. Res. 2021, in press. [Google Scholar] [CrossRef] [PubMed]
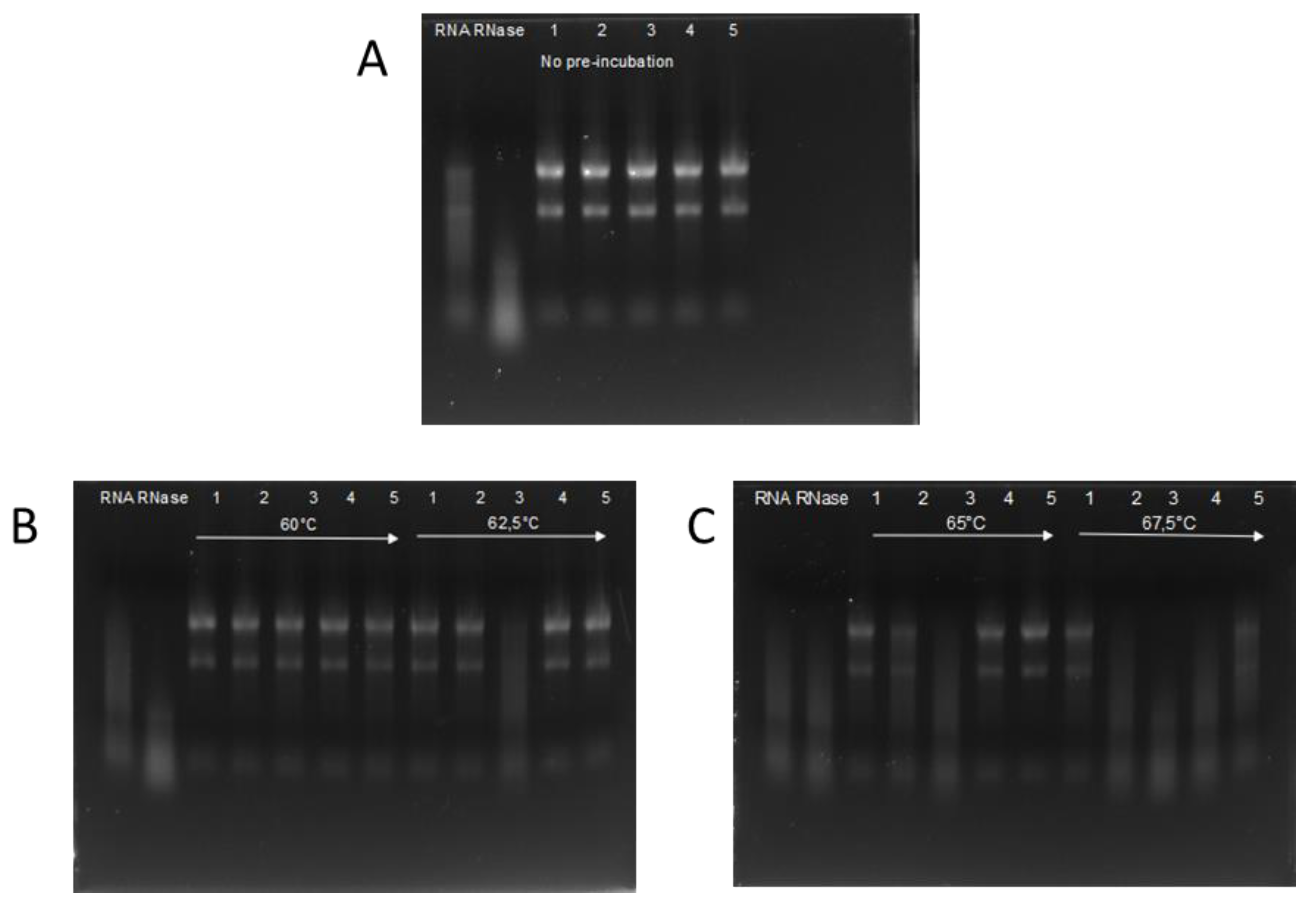
| 60 °C | 62.5 °C | 65 °C | 67.5 °C | |
|---|---|---|---|---|
| Insect RI | + | + | + | partial |
| ‘RI I’ | + | + | partial | − |
| ‘RI II’ | + | − | − | − |
| ‘RI III’ | + | + | + | − |
| ‘RI IV’ | + | + | + | − |
| - | Melting Temperature (°C) | |
|---|---|---|
| - RNase A | + RNase A | |
| Insect RI | 54.1 | 63.2 |
| ‘RI I’ | 52.2 | 63.3 |
| ‘RI II’ | 50.5 | 62.9 |
| ‘RI III’ | 54.5 | 64.1 |
| ‘RI IV’ | 54.6 | 63.2 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Flachner, B.; Dobi, K.; Benedek, A.; Cseh, S.; Lőrincz, Z.; Hajdú, I. Robust Recombinant Expression of Human Placental Ribonuclease Inhibitor in Insect Cells. Biomolecules 2022, 12, 273. https://doi.org/10.3390/biom12020273
Flachner B, Dobi K, Benedek A, Cseh S, Lőrincz Z, Hajdú I. Robust Recombinant Expression of Human Placental Ribonuclease Inhibitor in Insect Cells. Biomolecules. 2022; 12(2):273. https://doi.org/10.3390/biom12020273
Chicago/Turabian StyleFlachner, Beáta, Krisztina Dobi, Anett Benedek, Sándor Cseh, Zsolt Lőrincz, and István Hajdú. 2022. "Robust Recombinant Expression of Human Placental Ribonuclease Inhibitor in Insect Cells" Biomolecules 12, no. 2: 273. https://doi.org/10.3390/biom12020273
APA StyleFlachner, B., Dobi, K., Benedek, A., Cseh, S., Lőrincz, Z., & Hajdú, I. (2022). Robust Recombinant Expression of Human Placental Ribonuclease Inhibitor in Insect Cells. Biomolecules, 12(2), 273. https://doi.org/10.3390/biom12020273






