1. Introduction
Lipolytic enzymes which catalyze cleavage of the ester bonds in triacylglycerols are widely used in biotechnology including the food industry, fine chemical synthesis, production of detergents, and biofuel [
1,
2]. Lipases and esterases from extremophilic microorganisms (thermophilic, psychrophilic, halophilic, etc.) frequently possess unique structural and functional characteristics which enable them to operate in harsh environmental conditions [
3]. These properties provide a foundation for the ongoing search for new representatives of the family and numerous studies devoted to their characterization.
In particular, many cold-active lipolytic enzymes demonstrate high catalytic efficiency and activity at low temperatures [
4,
5]. They can also catalyze reactions in the presence of high salt concentration and organic solvents. However, low thermal stability usually limits their broad industrial application.
Similar to other lipolytic enzymes, cold-active lipases and esterases possess common α/β hydrolase fold which consists of a catalytic and a cap domain [
2,
6]. The former includes a β-sheet of 5–11 β-strands surrounded by α-helices and contains a catalytic triad of Ser-His-Asp/Glu residues. In comparison with more thermostable analogues, the molecules of cold-active enzymes typically demonstrate smaller hydrophobic core, lower number of stabilizing interactions, the presence of long extended loops and greater number of exposed hydrophobic residues at the protein surface [
7,
8].
Being a habitat for diverse cold-adapted microorganisms, Siberian permafrost is a unique environment which is characterized by the presence of permanently frozen ground, limited accessibility of organic matter, low water activity and other factors [
9,
10,
11]. Previously, we have described several lipolytic enzymes from the permafrost bacterium
Psychrobacter cryohalolentis K5
T [
12,
13,
14]. Furthermore, we performed screening of the metagenomic DNA library obtained from the permafrost-derived microcosm that resulted in production and characterization of new esterases PMGL2 and PMGL3 belonging to the bacterial hormone sensitive lipase (bHSL, EC 3.1.1.79) family [
15,
16]. The PMGL3 esterase demonstrated low thermal stability with rapid inactivation upon incubation at 40 °C, which is typical for the cold-active enzymes. In particular, the enzyme was shown to possess a dimeric state in solution, with a tendency to oligomerize upon heating [
17].
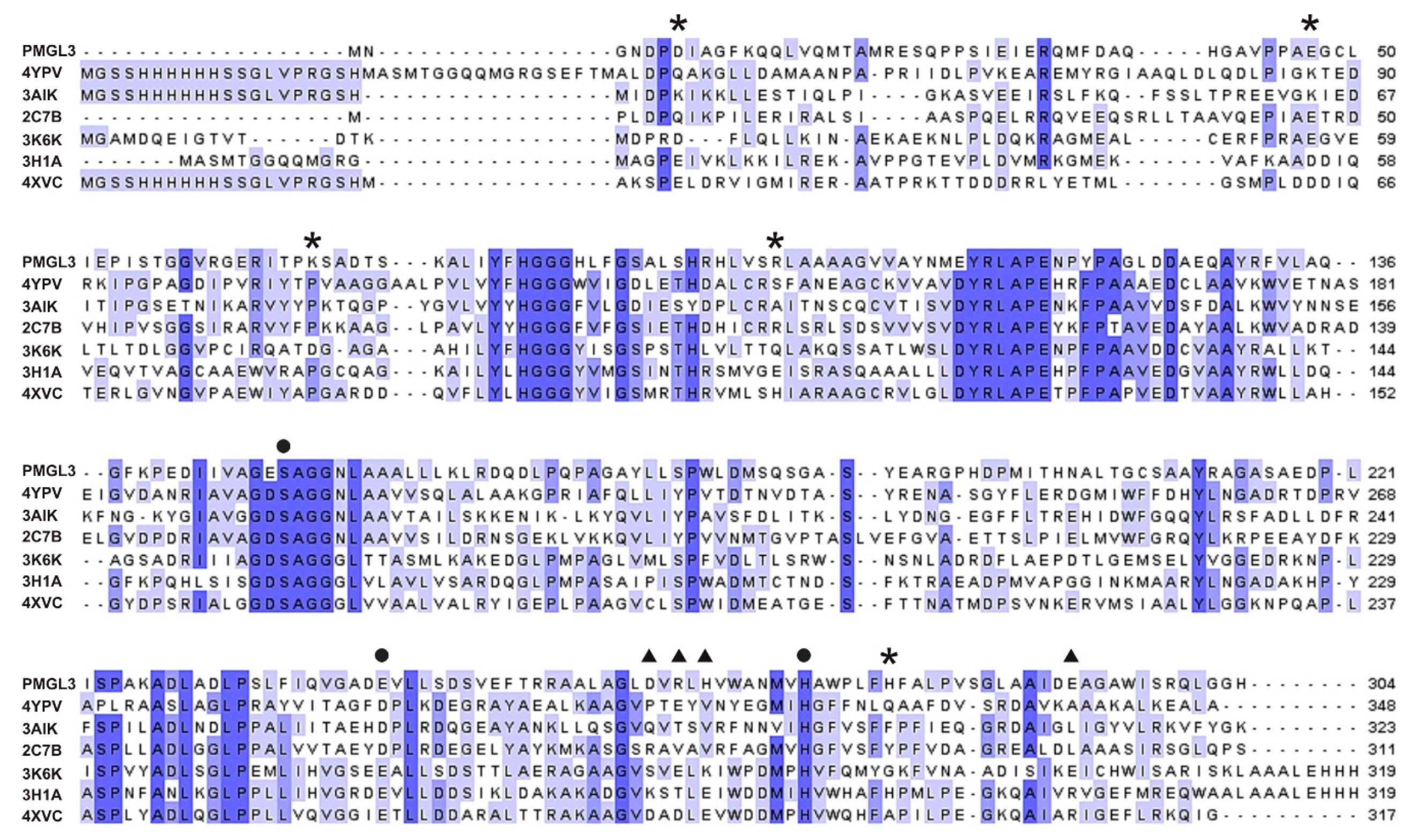
Here, we comprehensively studied the biochemical properties of the wild type PMGL3 and determined its 3D structure. Surprisingly, it has a tetrameric architecture in a crystal, with a subunit’s fold common to the members of the GDSAG motif subfamily of the bHSL family in spite of the low amino acid similarity with structurally characterized esterases. Moreover, the crystal tetramer of PMGL3 has a unique architecture compared to other known tetramers of bHSLs. In order to study an effect of tetramerization on biochemical properties of the enzyme and to identify structural motifs important for tetramerization, we carefully analyzed dimeric and tetrameric interfaces and performed site-directed mutagenesis of the residues possibly stabilizing tetrameric contacts through formation of salt bridges. Our results demonstrated that mutations of non-conserved D7 and K67 residues mostly affected the tetramerization and the thermal stability of PMGL3.
4. Discussion
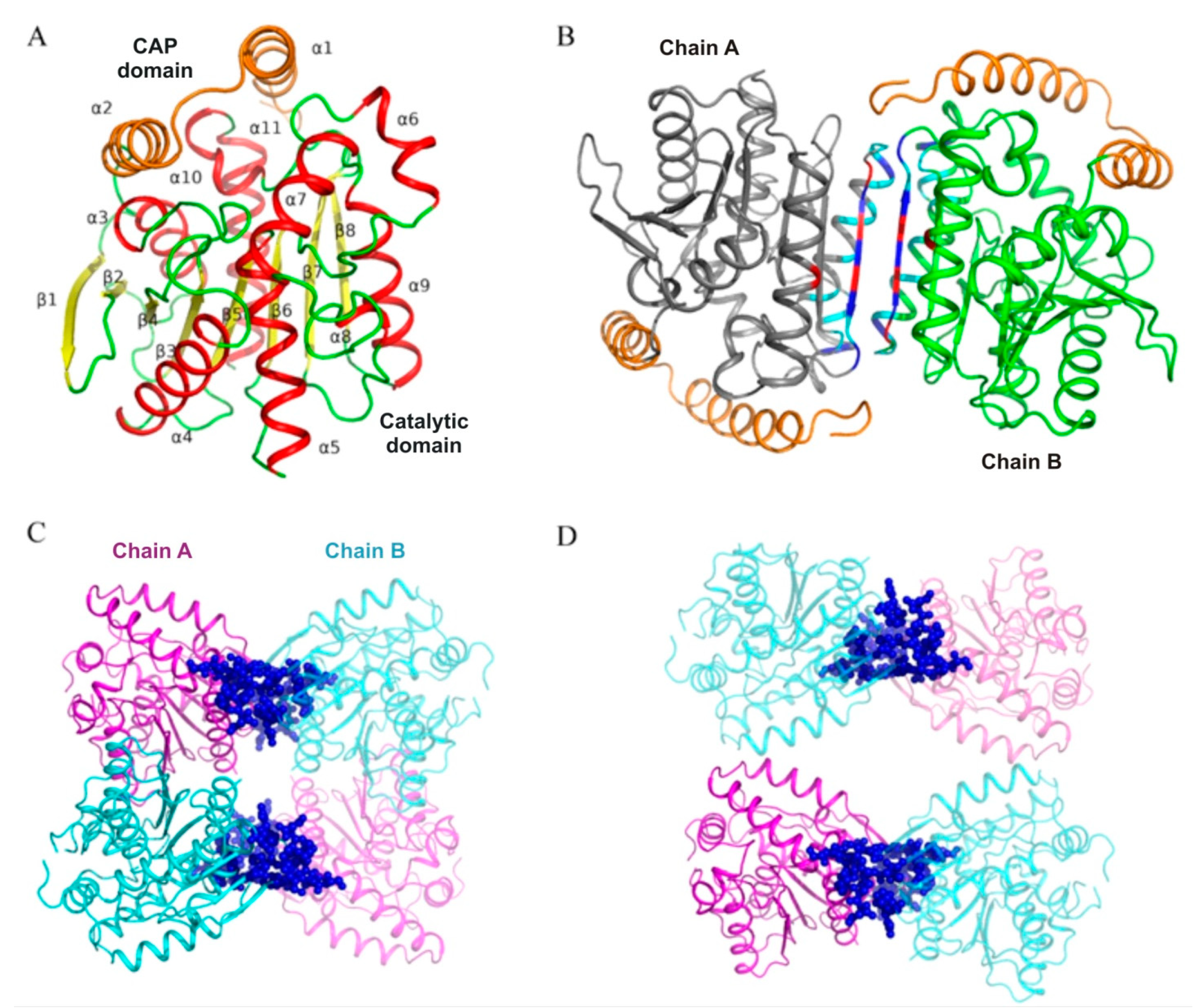
In this work, we performed structural and biochemical characterization of the PMGL3 esterase, a novel member of the GDSAG motif subfamily of the bHSL family. The protein was obtained from the metagenomic DNA library constructed from the permafrost-derived microcosm. The PMGL3 demonstrated a typical fold for the bHSL family with however a unique tetrameric architecture. In many bHSLs, a β8 strand of the adjacent subunits is an important determinant of the dimeric interface. A major difference between GDSAG and GT/XSAG bHSL subfamilies is the mutual orientation of β8 strands within the dimeric interface. They are antiparallel in case of GDSAG subfamily [
39,
40,
41] and about to perpendicular for GT/XSAG [
35,
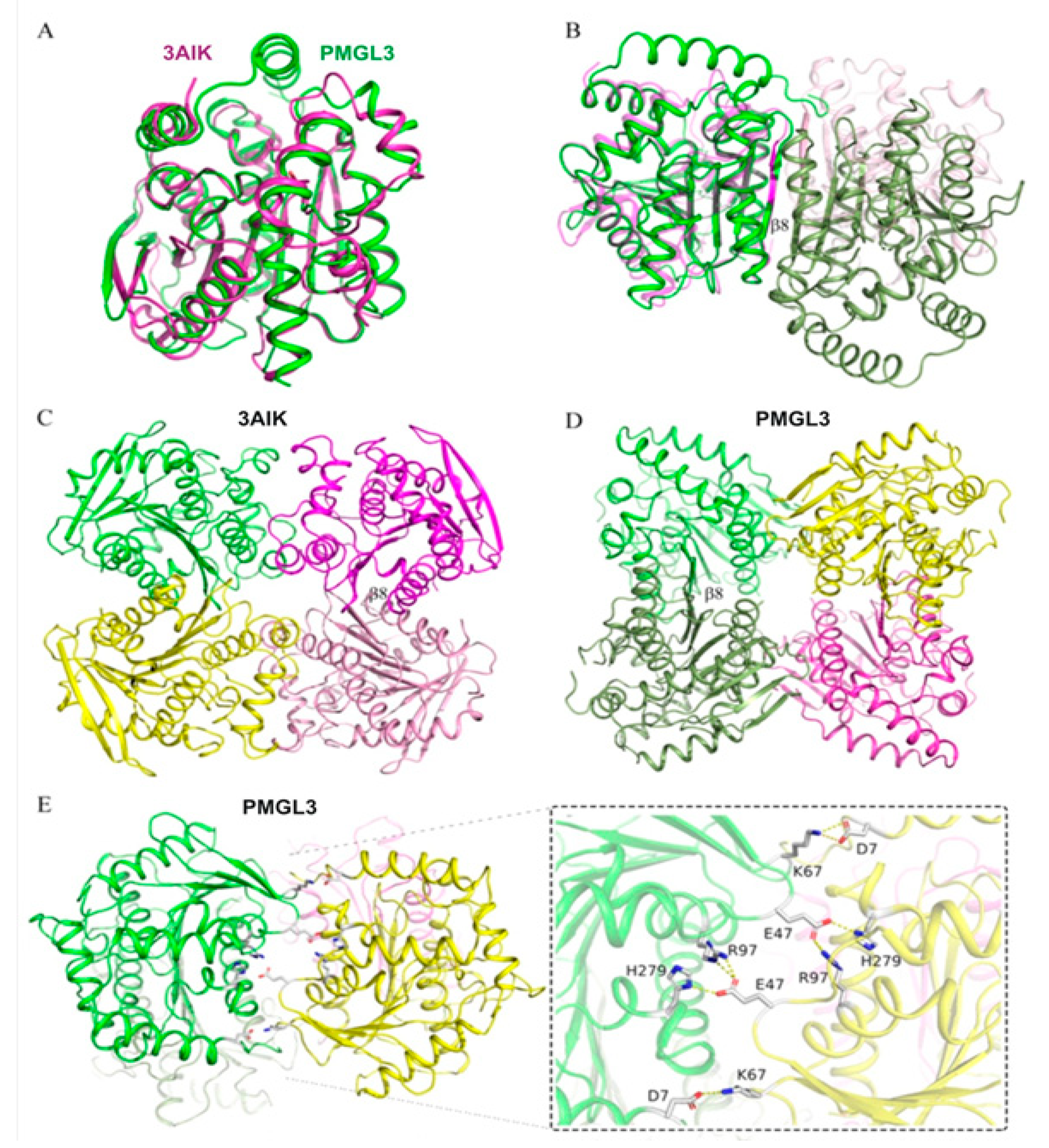
36]. According to our results, we speculate that bHSLs of GDSAG subfamily with dimeric and/or tetrameric organization could be further subdivided based on details of their dimeric interface. In the structures of most members of this subfamily (e.g., 3ZWQ, 1JKM, 3AIK, 2C7B, etc.) the β8-β8* strands are not strictly antiparallel, but intercrosses at angle of about 40° (
Figure 7A). In other cases (PMGL3, 3K6K), these strands are antiparallel and lie almost in the same plane (
Figure 7B). This difference leads to completely different dimeric or tetrameric (if any) architectures. Noteworthy, structures of some bHSLs revealed that oligomeric forms could be organized completely without involvement of β8 strands (e.g., 4C88 and 6AAE). Further widening of structural “universe” of bHSLs could shed light on this issue.
Previously, we have shown that PMGL3 is a cold-active esterase with low thermal stability [
16,
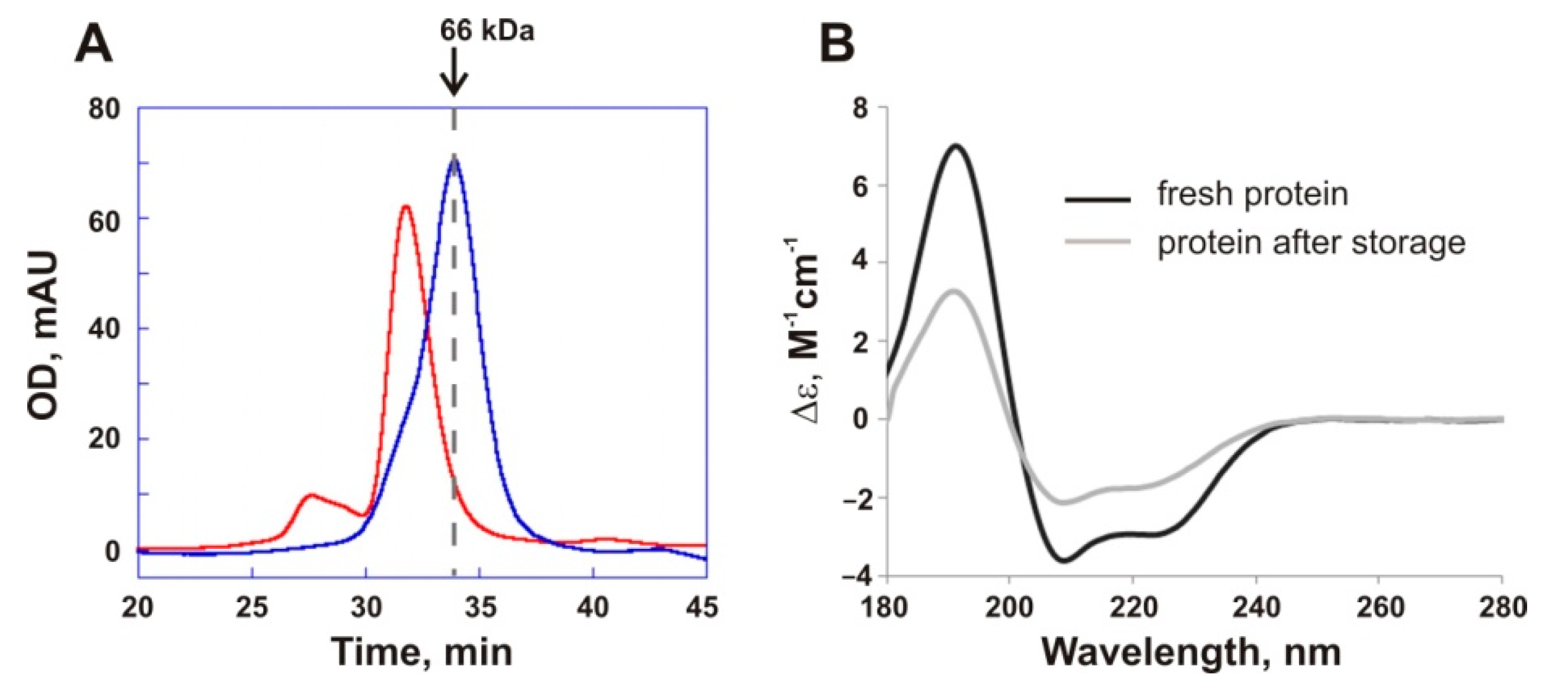
17]. It exhibited increased potential for oligomerization and dramatic loss of activity upon incubation even at 40 °C. In the current work, we demonstrated that prolonged storage of PMGL3 at 4 °C also leads to enzyme inactivation which is accompanied with the loss of the native secondary structure content and formation of oligomers. Limited thermal stability is a characteristic feature of the cold-active enzymes [
7,
8]. Several strategies have been proposed to increase their half-life based on the directed evolution or rational engineering approaches [
49,
50]. The extension of knowledge about 3D structures of cold-active enzymes can promote such studies due to preliminary identification of the sites for potential modifications.
In our earlier work, two cysteine residues of PMGL3, C49 and C207, were mutated in order to improve enzyme properties. The former mutation led to accelerated thermoinactivation of the enzyme, and the latter one increased thermal stability of the mutant protein and reduced its tendency to oligomerization [
17]. In the current paper, elucidation of the PMGL3 3D structure enabled us to explain this effect from a structural point of view. A slight decrease in temperature stability for C49V mutant could be a result of poor packing of the hydrophobic valine side chain in surrounding residues. Despite V49 can be modelled in this position without steric clashes, we could speculate that its placement is energetically less favorable than of the cysteine residue. In case of C207F substitution, the reason of improved thermostability seems to be clearer as large phenolic side chain of F207 might strengthen a hydrophobic core nearby, containing, in particular, residues L84, F85, W179 and L204.
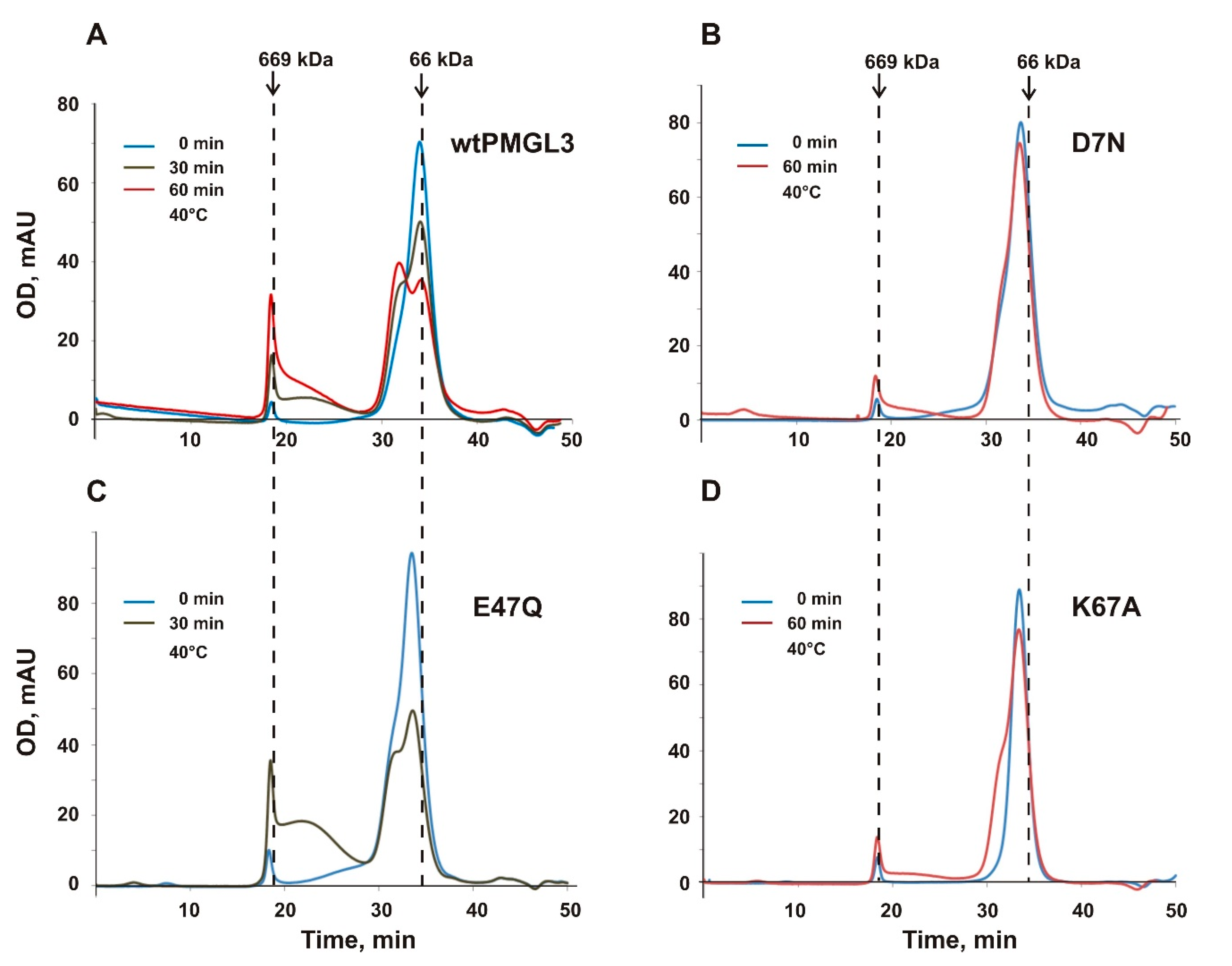
The availability of the 3D structure also allowed us to identify residues participating in the formation of tetrameric interface, which accompanies thermal unfolding and inactivation of the enzyme. We have focused on mutagenesis of the charged residues, which stabilize the tetrameric interface via the salt bridges, which are the strongest among polar interactions. However, no one of the studied single mutations was sufficient to completely prevent tetramerization of PMGL3 upon heating, pointing to the effect of other (e.g., hydrophobic) stabilizing interactions that lead to the high stability of the tetramer. Significant decrease of the tetramer formation after incubation for 60 min at 40 °C was observed for D7N and K67A mutants while for E47Q a substantial amount of tetramer formed even after 30 min heating. In accordance with that, the inactivation half-time of the E47Q mutant was by 28% lower than that of the wild type PMGL3 while the half-lives of the D7N and K67A mutants exceeded it by 40 and 60% correspondingly.
Single mutations D7N, E47Q, and K67A did not significantly changed substrate affinity; however, kcat of E47Q and K67A was reduced. It should be mentioned that two of the mutations, R97Q and H279A led to inactivation of the enzyme. Based on structural analysis, both R97 and H279 are disposed close to each other and participate in a network of hydrogen bonds, which, presumably, supports the native conformation of the enzyme.
Formation of oligomers is a common feature of thermophilic and hyperthermophilic enzymes, which is known to contribute to their high thermal stability [
51]. Thus, thermophilic esterases PestE from
Pyrobaculum calidifontis VA1 [
39] and EstE1 from metagenomic library [
40] are tetramers in a crystal. Moreover, strengthening of oligomeric interfaces, in particular, by introduction of the salt bridges is considered an efficient bioengineering approach to enzyme stabilization [
52,
53]. This approach was successfully used to increase thermal stability of malate dehydrogenase [
54], PyrR [
55], αE7 carboxylesterase [
56] and other proteins. However, if the partially unfolded proteins form oligomers then electrostatic interactions can lead to stabilization of aggregates and enzyme inactivation [
57]. Based on the presence of interfaces to form tetramers in PMGL3 molecule, we supposed that oligomers of partially unfolded enzyme can be stabilized by these interfaces and demonstrated that mutations that disrupt tetrameric interactions can increase thermal stability of the enzyme. The obtained results can be used further to increase the thermal stability of PMGL3 and other esterases for their biotechnological application.