An Economic Dilemma between Molecular Weapon Systems May Explain an Arachno-Atypical Venom in Wasp Spiders (Argiope bruennichi)
Abstract
1. Introduction
2. Materials and Methods
2.1. Collection of Specimens and Sample Preparation for Transcriptomics and Proteomics
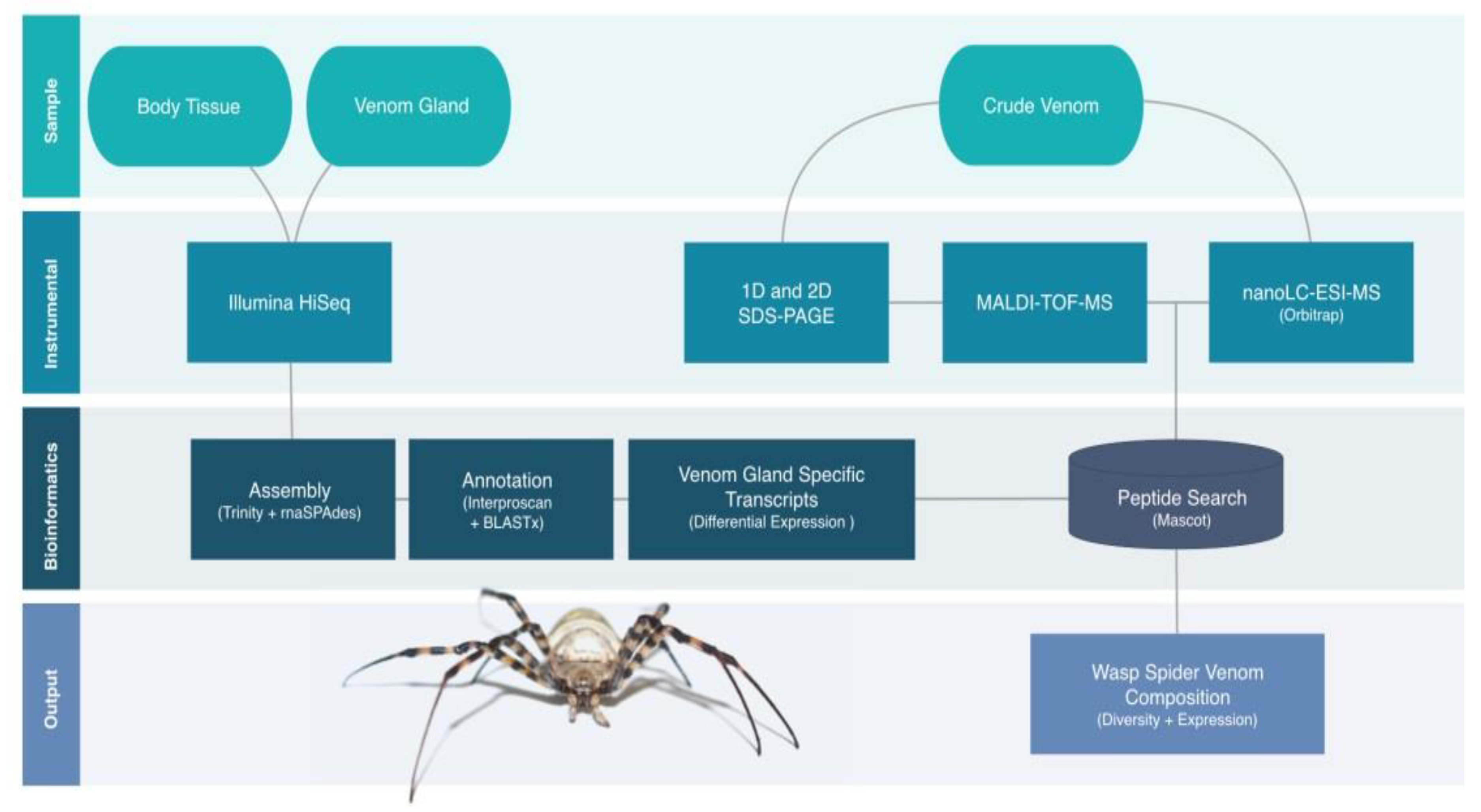
2.2. Proteotranscriptomics Overall Workflow
2.3. Transcriptomics of Venom Gland and Body Tissue
2.3.1. RNA Extraction and Sequencing
2.3.2. Transcriptome Assembly, Annotation and Quantification
2.4. Venom Proteomics
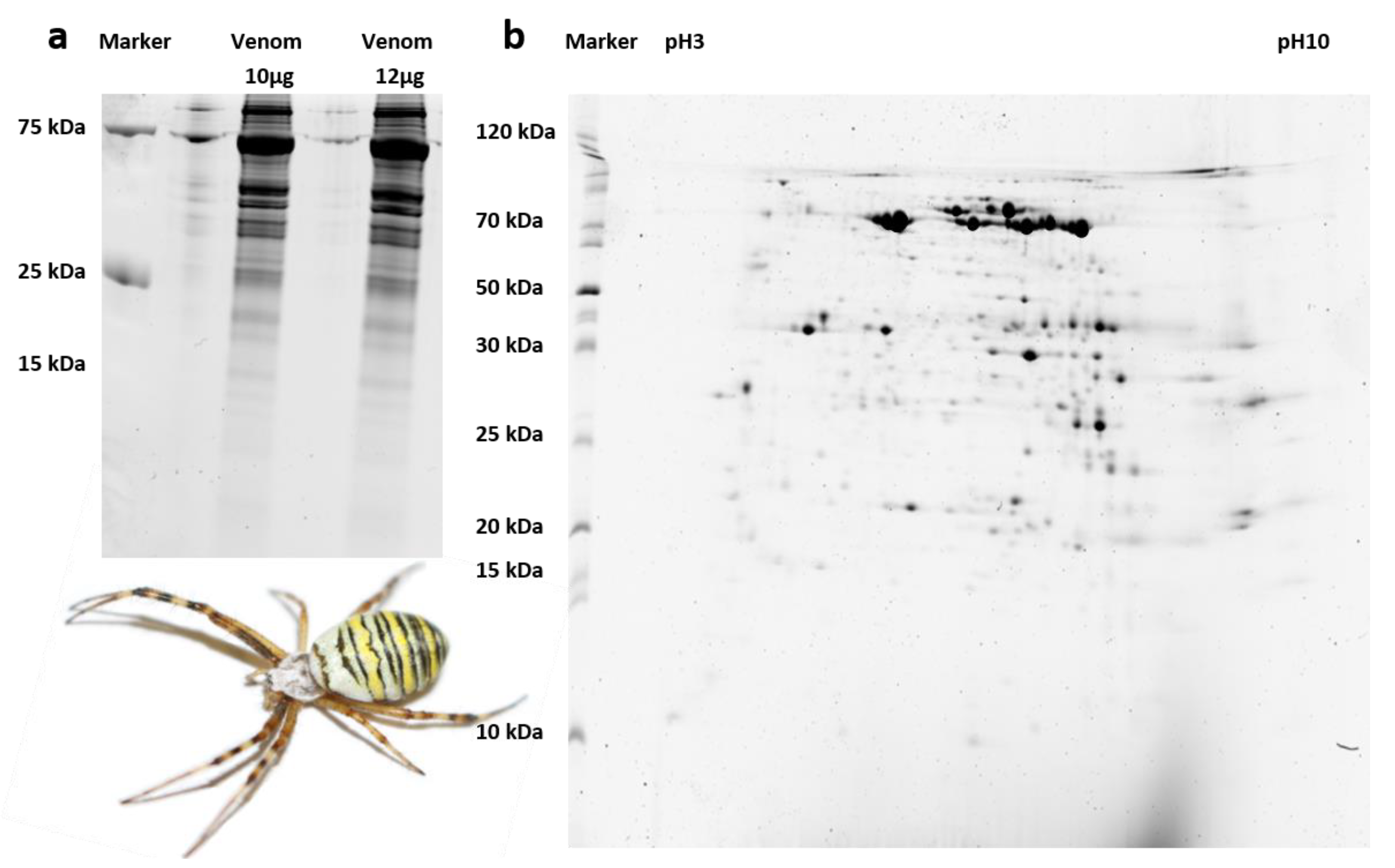
2.4.1. Fractionation of Venom Proteins by PAGE
2.4.2. MALDI-TOF-MS
2.4.3. NanoLC-ESI-MS
2.5. Reanalysis of Araneus Ventricosus Venom
3. Results
3.1. The A. bruennichi Venom Gland and Body Tissue Yield High-Quality Transcriptome Libraries
3.2. Only Large A. bruennichi Venom Proteins Are Detected by SDS-PAGE and MALDI-TOF-MS
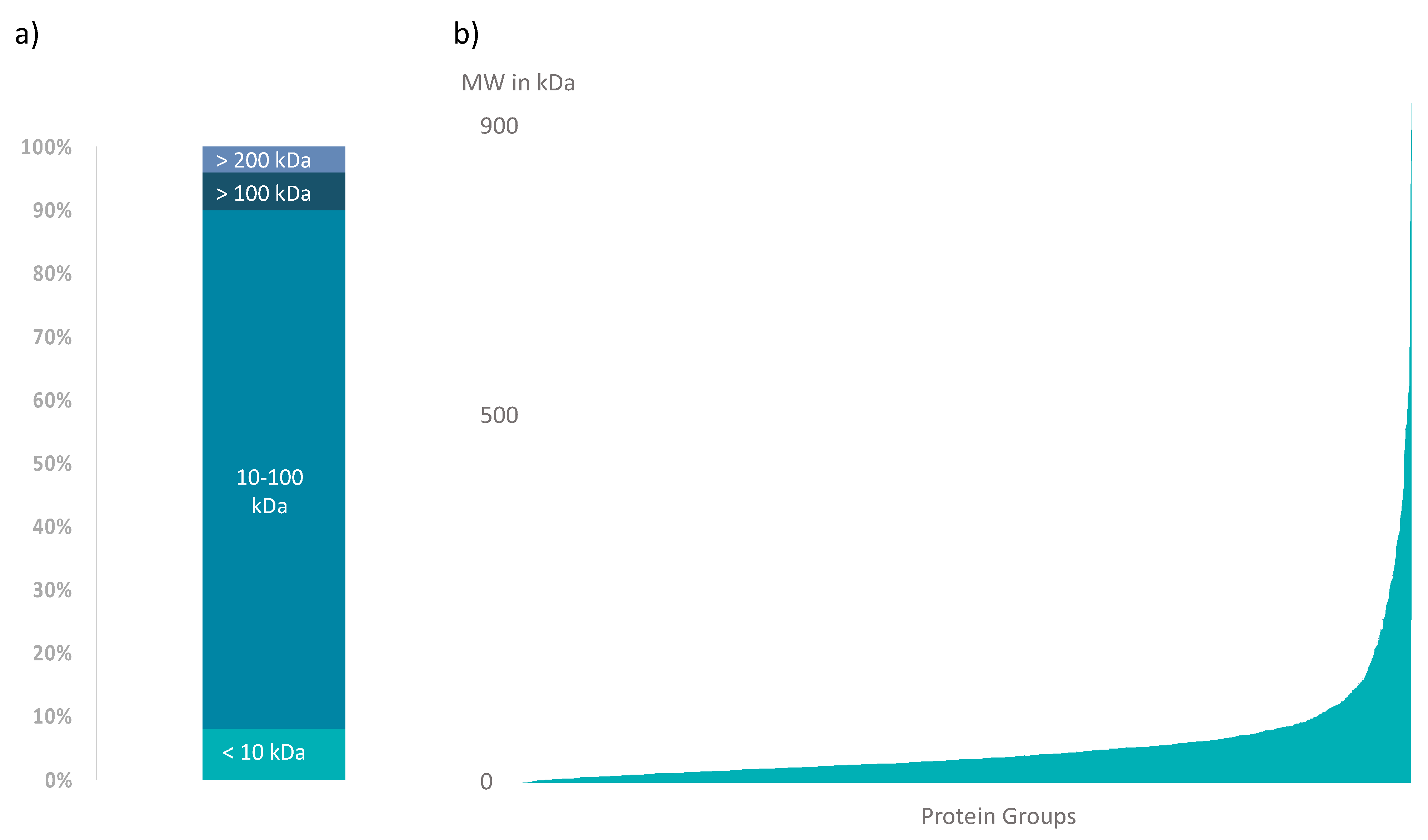
3.3. Further A. bruennichi Venom Proteins Are Revealed by High-Resolution NanoLC-ESI-MS
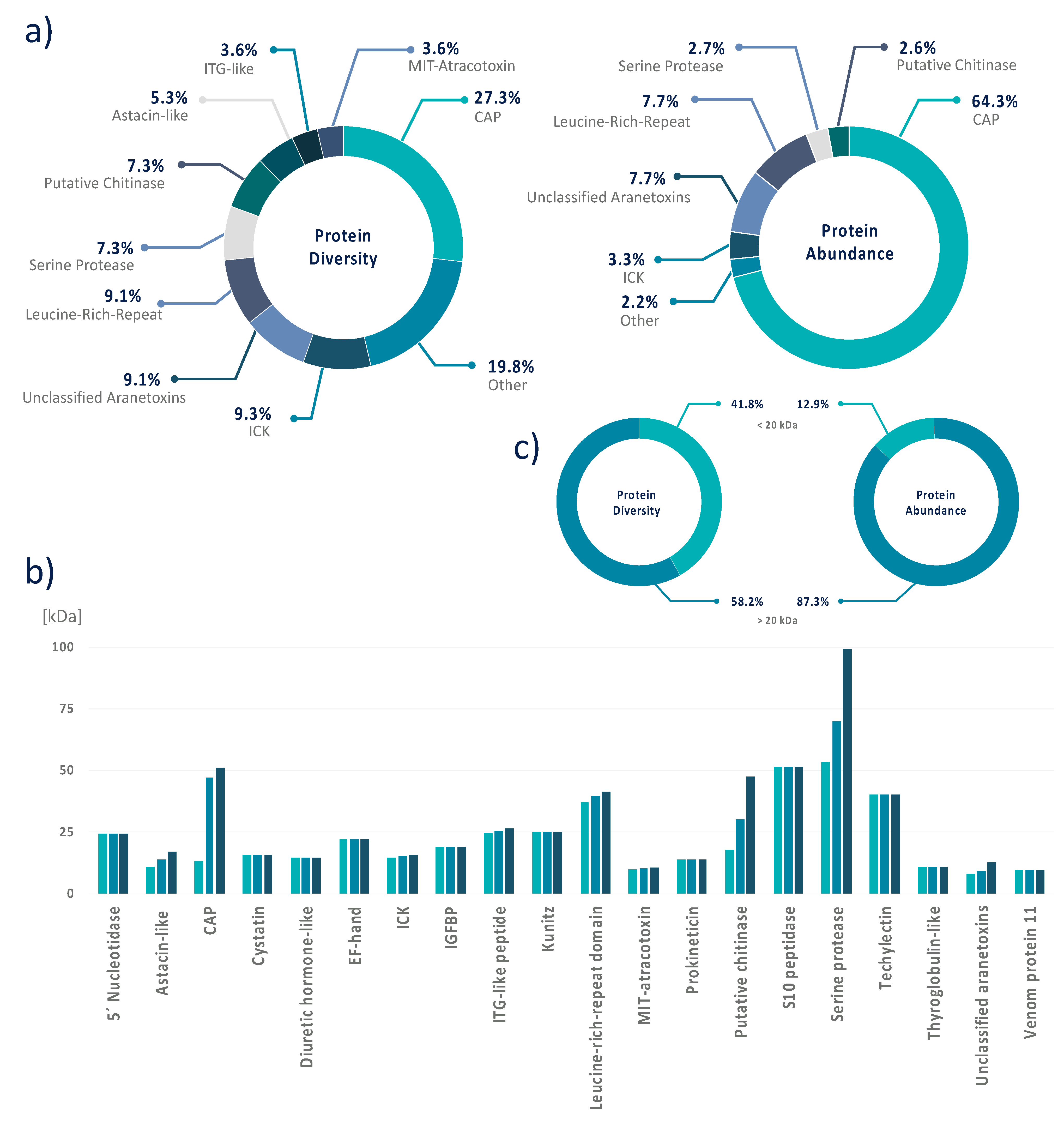
3.4. Data Integration Reveals That A. bruennichi Venom Is Atypical for Spiders
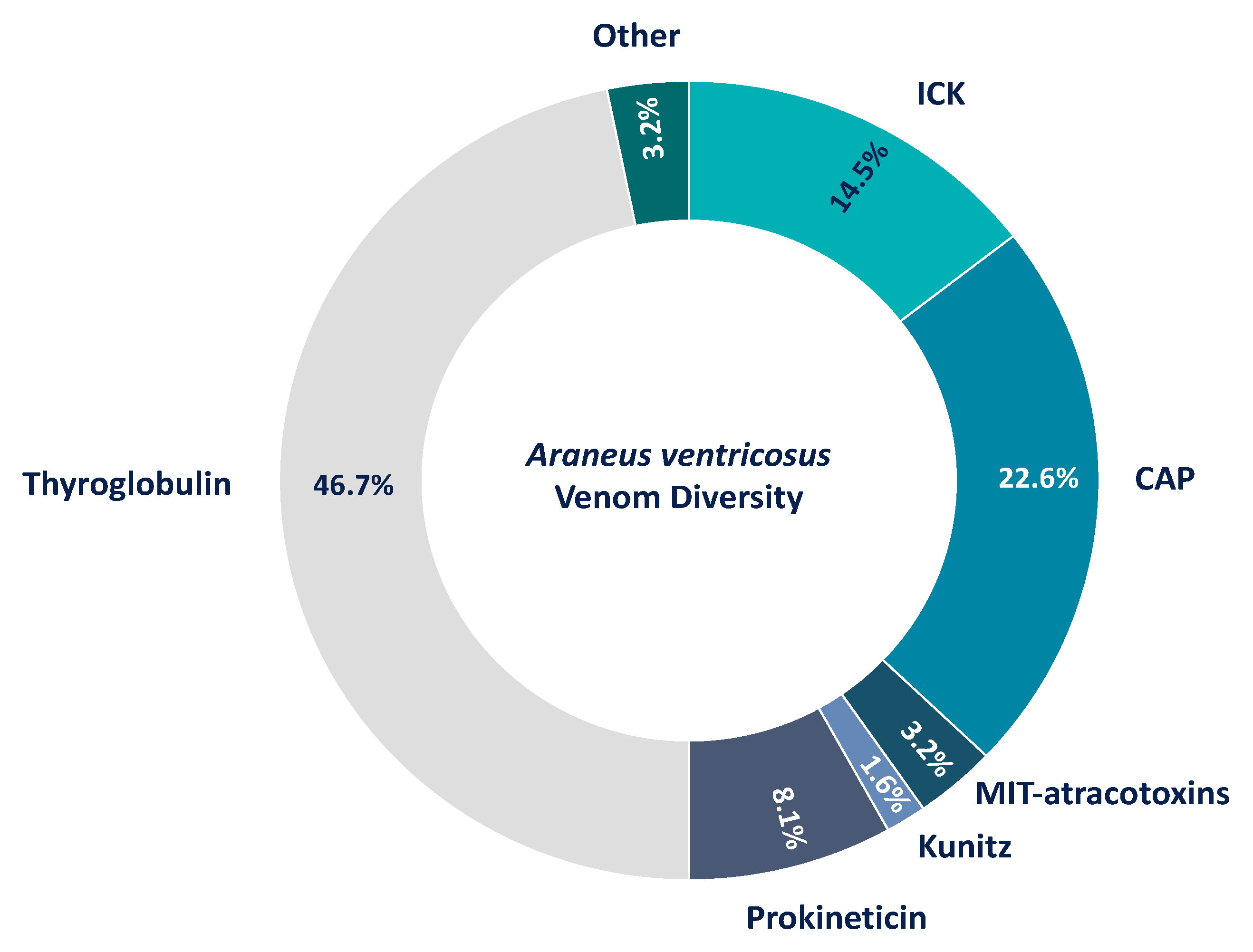
3.5. The Venom of Araneus Ventricosus Has Similarity to Argiope bruennichi Venom
4. Discussion
4.1. The Importance of CAP Superfamily Proteins in Wasp Spider Venom
4.2. Wasp Spider Venom Contains Potential New Toxin Classes Similar to Arthropod Neuropeptides
4.3. The Potential Ecological Role of Atypical Wasp Spider Venom
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- World Spider Catalog, Version 21. Available online: http://wsc.nmbe.ch (accessed on 16 February 2020).
- Garrison, N.L.; Rodriguez, J.; Agnarsson, I.; Coddington, J.A.; Griswold, C.E.; Hamilton, C.A.; Hedin, M.; Kocot, K.M.; Ledford, J.M.; Bond, J.E. Spider phylogenomics: Untangling the spider tree of life. PeerJ 2016. [Google Scholar] [CrossRef] [PubMed]
- Saez, N.J.; Senff, S.; Jensen, J.E.; Er, S.Y.; Herzig, V.; Rash, L.D.; King, G.F. Spider-venom peptides as therapeutics. Toxins 2010, 2, 2851–2871. [Google Scholar] [CrossRef]
- Riechert, S.E. Thoughts on the Ecological Significance of Spiders. BioScience 1974, 24, 352–356. [Google Scholar] [CrossRef]
- Fry, B.G.; Roelants, K.; Champagne, D.E.; Scheib, H.; Tyndall, J.D.A.; King, G.F.; Nevaleinen, T.J.; Norman, J.A.; Lewis, R.J.; Norton, R.S.; et al. The Toxicogenomic Multiverse: Convergent Recruitment of Proteins into Animal Venoms. Annu. Rev. Genom. Hum. Genet. 2009, 10, 483–511. [Google Scholar] [CrossRef] [PubMed]
- Nelsen, D.R.; Nisani, Z.; Cooper, A.M.; Fox, G.A.; Gren, E.C.K.; Corbit, A.G.; Hayes, W.K. Poisons, toxungens, and venoms: Redefining and classifying toxic biological secretions and the organisms that employ them. Biol. Rev. Camb. Philos. Soc. 2014, 89, 450–465. [Google Scholar] [CrossRef] [PubMed]
- Holford, M.; Daly, M.; King, G.F.; Norton, R.S. Venoms to the rescue. Science 2018, 361, 842–844. [Google Scholar] [CrossRef]
- Duterte, S.; Lewis, R.J. Use of Venom Peptides to Probe Ion Channel Structure and Function. J. Biol. Chem. 2010, 285, 13315–13320. [Google Scholar] [CrossRef]
- Herzig, V.; Cristofori-Armstrong, B.; Israel, M.R.; Nixon, S.A.; Vetter, I.; King, G.F. Animal toxins—Nature’s evolutionary refined toolkit for basic research and drug discovery. Biochem. Pharmacol. 2020, 114096. [Google Scholar] [CrossRef]
- Dongol, Y.; Cardoso, F.C.; Lewis, R.J. Spider Knottin Pharmacology at Voltage-Gated Sodium Channels and Their Potential to Modulate Pain Pathways. Toxins 2019, 11, 626. [Google Scholar] [CrossRef]
- Saez, N.J.; Herzig, V. Versatile spider venom peptides and their medical and agricultural applications. Toxicon 2019, 158, 109–126. [Google Scholar] [CrossRef]
- Pineda, S.; Chin, Y.K.Y.; Undheim, E.A.B.; Senff, S.; Mobli, M.; Dauly, C.; Nicholson, G.; Kaas, Q.; Guo, S.; Herzig, V.; et al. Structural Venomics Reveals Evolution of a Complex Venom by Duplication and Diversification of an Ancient Peptide-Encoding Gene. Proc. Natl. Acad. Sci. USA 2020, 117, 11399–11408. [Google Scholar] [CrossRef] [PubMed]
- Langenegger, N.; Nentwig, W.; Kuhn-Nentwig, L. Spider Venom: Components, Modes of Action, and Novel Strategies in Transcriptomic and Proteomic Analyses. Toxins 2019, 11, 611. [Google Scholar] [CrossRef]
- Pineda, S.S.; Chaumeil, P.A.; Kunert, A.; Kaas, Q.; Thang, M.W.C.; Le, L.; Nuhn, M.; Herzig, V.; Saez, N.J.; Cristofori-Armstrong, B.; et al. ArachnoServer 3.0: An online resource for automated discovery, analysis and annotation of spider toxins. Bioinformatics 2018, 34, 1074–1076. [Google Scholar] [CrossRef]
- Lüddecke, T.; Vilcinskas, A.; Lemke, S. Phylogeny-Guided Selection of Priority Groups for Venom Bioprospecting: Harvesting Toxin Sequences in Tarantulas as a case Study. Toxins 2019, 11, 488. [Google Scholar] [CrossRef] [PubMed]
- Osteen, J.D.; Herzig, V.; Gilchrist, J.; Emrick, J.J.; Zhang, C.; Wang, X.; Castro, J.; Garcia-Caraballo, S.; Grundy, L.; Rychkov, G.Y.; et al. Selective spider toxins reveal a role for Nav1.1 in mechanical pain. Nature 2016, 534, 494–499. [Google Scholar] [CrossRef] [PubMed]
- Chasagnon, I.R.; McCarthy, C.A.; Chin, Y.K.Y.; Pineda, S.S.; Keramidas, A.; Mobli, M.; Pham, V.; De Silva, T.M.; Lynch, J.W.; Widdop, R.E.; et al. Potent neuroprotection after stroke afforded by a double-knot spider-venom peptide that inhibits acid-sensing ion channel 1a. Proc. Natl. Acad. Sci. USA 2017, 114, 3750–3755. [Google Scholar] [CrossRef]
- Richards, K.L.; Milligan, C.J.; Richardson, R.J.; Jancovski, N.; Grunnet, M.; Jacobsen, L.H.; Undheim, E.A.B.; Mobli, M.; Chow, C.Y.; Herzig, V.; et al. Selective Nav1.1 activation rescues Dravet syndrome mice from seizures and premature death. Proc. Natl. Acad. Sci. USA 2018, 115, 8077–8085. [Google Scholar] [CrossRef]
- Wu, T.; Wang, M.; Wu, W.; Luo, Q.; Jiang, L.; Tao, H.; Deng, M. Spider venom peptides as potential drug candidates due to their anticancer and antinociceptive sctivities. J. Venom. Anim. Toxins Incl. Trop. Dis. 2019, 25. [Google Scholar] [CrossRef]
- Pallaghy, P.K.; Nielsen, K.J.; Craik, D.J.; Norton, R.S. A common structural motif incorporating a cysteine knot and a triple-stranded beta-sheet in toxic and inhibitory polypeptides. Protein Sci. 1994, 3, 1833–1839. [Google Scholar] [CrossRef]
- Norton, R.S.; Pallaghy, P.K. The cysteine knot structure of ion channel toxins and related polypeptides. Toxicon 1998, 36, 1573–1583. [Google Scholar] [CrossRef]
- Von Reumont, B.M.; Undheim, E.A.B.; Jauss, R.T.; Jenner, R.T. Venomics of Remipede Crustaceans Reveals Novel peptide Diversity and Illuminates the Venoms Biological Role. Toxins 2017, 9, 243. [Google Scholar] [CrossRef] [PubMed]
- Drukewitz, S.H.; Fuhrmann, N.; Undheim, E.A.B.; Blanke, A.; Giribaldi, J.; Mary, R.; Laconde, G.; Duterte, S.; von Reumont, B.M. A Dipteran´s Novel Sucker Punch: Evolution of Arthropod Atypical Venom with a neurotoxic Component in Robber Flies (Asilidae, Diptera). Toxins 2018, 10, 29. [Google Scholar] [CrossRef] [PubMed]
- Drukewitz, S.H.; Bokelmann, L.; Undheim, E.A.B.; von Reumont, B.M. Toxins from scratch? Diverse, multimodal gene origins in the predatory robber fly Dasypogon diadema indicate a dynamic venom evolution in dipteran insects. Gigascience 2019, 8, giz081. [Google Scholar] [CrossRef] [PubMed]
- Özbek, R.; Wielsch, N.; Vogel, H.; Lochnit, G.; Förster, F.; Vilcinskas, A.; von Reumont, B.M. Proteo-Transcriptomic Characterization of the venom from the Endoparasitoid Wasp Pimpla turionellae with Aspects on its Biology and Evolution. Toxins 2019, 11, 721. [Google Scholar] [CrossRef] [PubMed]
- The UniProt Consortium. UniProt: A worldwide hub of protein knowledge. Nucleic Acids Res. 2019, 47, D506–D515. [Google Scholar] [CrossRef]
- Friedel, T.; Nentwig, W. Immobilizing and lethal effects of spider venoms on the cockroach and and the common mealbeetle. Toxicon 1989, 27, 305–3016. [Google Scholar] [CrossRef]
- Hardy, M.C.; Daly, N.L.; Mobli, M.; Morales, R.A.; King, G.F. Isolation of an orally active insecticidal toxin from the venom of an Australian tarantula. PLoS ONE 2013. [Google Scholar] [CrossRef]
- Von Reumont, B.M. Studying Smaller and Neglected Organisms in Modern Evolutionary Venomics Implementing RNAseq (Transcriptomics)—A Critical Guide. Toxins 2018, 10, 29. [Google Scholar] [CrossRef]
- Oldrati, V.; Arrell, M.; Violette, A.; Perret, F.; Sprüngli, X.; Wolfender, J.L.; Stöcklin, R. Advances in Venomics. Mol. Biosyst. 2016, 12, 3530–3543. [Google Scholar] [CrossRef]
- Ul-Saban, S.; Rodriguez-Roman, E.; Reitzel, A.M.; Adams, R.M.M.; Herzig, V.; Nobile, C.J.; Saviola, A.J.; Trim, S.A.; Stiers, E.E.; Moschos, S.A.; et al. The emerging field of venom-microbiomics for exploring venom as a microenvironment, and the corresponding Initiative for Venom Associated Microbes and Parasites (iVAMP). Toxicon:X 2019, 4, 100016. [Google Scholar]
- Von Reumont, B.M.; Campbell, L.I.; Jenner, R.A. Quo vadis venomics? A roadmap to neglected venomous invertebrates. Toxins 2014, 6, 3488–3551. [Google Scholar] [CrossRef] [PubMed]
- Herzig, V.; King, G.F.; Undheim, E.A.B. Can we resolve the taxonomic bias in spider venom research? Toxicon:X 2019, 1, 100005. [Google Scholar] [CrossRef] [PubMed]
- Bond, J.E.; Garrison, N.L.; Hamilton, C.; Godwin, R.L.; Hedin, M.; Agnarsson, I. Phylogenomics resolves a spider backbone phylogeny and rejects a prevailing paradigm for orb web evolution. Curr. Biol. 2014, 24, 1765–1771. [Google Scholar] [CrossRef] [PubMed]
- Fernandez, R.; Hormiga, G.; Giribet, G. Phylogenomic analysis of spiders reveals nonmonophyly of orb weavers. Curr. Biol. 2014, 24, 1772–1777. [Google Scholar] [CrossRef] [PubMed]
- Duan, Z.; Cao, R.; Jiang, L.; Liang, S. A combined de novo protein sequencing and cDNA approach to the venomic analysis of Chinese spider Araneus ventricosus. J. Proteom. 2013, 78, 416–427. [Google Scholar] [CrossRef] [PubMed]
- Bush, A.A.; Yu, D.W.; Herberstein, M.E. Function of bright coloration in the wasp spider Argiope bruennichi (Araneae: Araneidae). Proc. Biol. Sci. 2008, 275, 1337–1342. [Google Scholar] [CrossRef] [PubMed]
- Zimmer, S.M.; Welke, K.W.; Schneider, J.M. Determinants of natural mating success in the cannibalistic orb-web spider Argiope bruennichi. PLoS ONE 2012, 7, e31389. [Google Scholar] [CrossRef] [PubMed]
- Chinta, S.P.; Goller, S.; Lux, S.; Uhl, G.; Schulz, S. The sex pheromone of the wasp spider Argiope bruennichi. Angew. Chem. Int. Ed. 2010, 49, 2033–2036. [Google Scholar] [CrossRef]
- Cory, A.L.; Schneider, J.M. Effects of social information on life history and mating tactics in the orb-web spider Argiope bruennichi. Ecol. Evol. 2017, 8, 344–355. [Google Scholar] [CrossRef]
- Cory, A.L.; Schneider, J.M. Mate availability does not influence mating strategies in males of the sexually cannibalistic spider Argiope bruennichi. PeerJ 2018. [Google Scholar] [CrossRef]
- Krehenwinkel, H.; Tautz, D. Northern range expansion of European populations of the wasp spider Argiope bruennichi is associated with global warming-correlated genetic admixture and population-specific temperature adaptions. Mol. Ecol. 2013, 2, 2232–2248. [Google Scholar] [CrossRef] [PubMed]
- Krehenwinkel, H.; Rödder, D.; Tautz, D. Eco-genomic analysis of the poleward range expansion of the wasp spider Argiope bruennichi shows rapid adaption and genomic admixture. Glob. Chang. Biol. 2015, 21, 4320–4332. [Google Scholar] [CrossRef] [PubMed]
- Krehenwinkel, H.; Pekar, S. An Analysis of factors Affecting Genotyping Success from Museum Specimens reveals an Increase of Genetic and Morphological variation during a Historical Range Expansion of a European Spider. PLoS ONE 2015, 10. [Google Scholar] [CrossRef] [PubMed]
- Sheffer, M.M.; Uhl, G.; Prost, S.; Lueders, T.; Ulrich, T.; Bengtsson, M.M. Tissue and Population-Level Microbiome Analysis of the Wasp Spider Argiope bruennichi identified a Novel bacterial Symbiont. Microorganisms 2019, 8, 8. [Google Scholar] [CrossRef]
- Welke, K.W.; Schneider, J.W. Males of the orb-web spider Argiope bruennichi sacrifice themselves to unrelated females. Biol. Lett. 2010, 6, 585–588. [Google Scholar] [CrossRef]
- Zhang, P.; Fang, H.Y.; Pan, W.J.; Pan, H.C. The complete mitochondrial genome of the wasp spider (Argiope bruennichi). Mitochondrial DNA A DNA Mapp. Seq. Anal. 2016, 27, 996–997. [Google Scholar] [CrossRef]
- Eisner, T.; Dean, J. Ploy and counterploy in predator-prey interactions: Orb-weaving spiders versus bombardier beetles. Proc. Natl. Acad. Sci. USA 1976, 73, 1365–1367. [Google Scholar] [CrossRef]
- Smith, J.; Undheim, E.A.B. True Lies: Using Proteomics to Assess the Accuracy of Transcriptome-Based Venomics in Centipedes Uncovers False Positives and Reveals Startling Intraspecific Variation in Scolopendra subspinipes. Toxins 2018, 10, 96. [Google Scholar] [CrossRef]
- Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data; Babraham Institute: Babraham, UK, 2019; Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 16 February 2020).
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Song, L.; Florea, L. Rcorrector: Efficient and accurate error correction for Illumina-seq reads. Gigascience 2015. [Google Scholar] [CrossRef]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zheng, Q.; et al. Full-length transcriptome assembly from RNA-seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef]
- Bushmanova, E.; Antipov, D.; Lapidus, A.; Prjibelski, A.D. rnaSPAdes: A de novo transcriptome assembler and its application to RNA-Seq data. Gigascience 2019, 8, giz100. [Google Scholar] [CrossRef] [PubMed]
- Haas, B.J.; Papanicolaou, A.; Yassour, M.; Grabherr, M.; Blood, P.D.; Bowden, J.; Couger, M.B.; Eccles, D.; Lieber, M.; MacManes, M.D.; et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat. Protoc. 2013, 8, 1494–1512. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Paggi, J.M.; Park, C.; Bennett, C.; Salzberg, S.L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 2019, 37, 907–915. [Google Scholar] [CrossRef] [PubMed]
- Pertea, M.; Pertea, G.M.; Antonescu, C.M.; Chang, T.C.; Mendell, J.T.; Salzberg, S.L. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 2015, 33, 290–295. [Google Scholar] [CrossRef] [PubMed]
- Pertea, M.; Kim, D.; Pertea, G.M.; Leek, J.T.; Salzberg, S.L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 2016, 11, 1650–1667. [Google Scholar] [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. The sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef]
- Camacho, C.; Coullouris, G.; Avagyan, V.; Ma, N.; Papadopoulus, J.; Bealer, K.; Madden, T.L. Blast+: Architecture and applications. BMC Bioinform. 2009, 10. [Google Scholar] [CrossRef]
- Jones, P.; Binns, D.; Chang, H.Y.; Fraser, M.; Li, W.; McAnulla, C.; McWilliam, H.; Maslen, J.; Mitchell, A.; Nuka, G.; et al. InterProScan 5: Genome-scale protein function classification. Bioinformatics 2014, 30, 1236–1240. [Google Scholar] [CrossRef]
- Ankenbrand, M.J.; Weber, L.; Becker, D.; Förster, F.; Bemm, F. TBro: Visualization and management of de novo transcriptomes. Database 2016, baw146. [Google Scholar] [CrossRef]
- Wessel, D.; Flugge, U.I. A method for the quantitative recovery of protein in dilute solution in the presence of detergents and lipids. Anal. Biochem. 1984, 138, 141–143. [Google Scholar] [CrossRef]
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef] [PubMed]
- Diniz, M.R.V.; Paiva, A.L.B.; Guerra-Duarte, C.; Nishiyama, M.Y.; Mudadu, M.A.; Oliveira, U.; Borges, M.H.; Yates, J.R.; Junqueira-de-Azevedo, I.L. An overview of Phoneutria nigriventer spider venom using combined transcriptomic and proteomic approaches. PLoS ONE 2018, 13. [Google Scholar] [CrossRef]
- Undheim, E.A.B.; Sunagar, K.; Herzig, V.; Kely, L.; Low, D.H.; Jackson, T.N.; Jones, A.; Kurniawan, N.; King, G.F.; Ali, S.A.; et al. A proteomics and transcriptomics investigation of the venom from the barychelid spider Trittame loki. Toxins 2013, 5, 2488–2503. [Google Scholar] [CrossRef] [PubMed]
- Kuhn-Nentwig, L.; Langenegger, N.; Heller, M.; Koua, D.; Nentwig, W. The Dual Prey-Inactivation Strategy of Spiders—In-Depth Venomic Analysis of Cupiennius salei. Toxins 2019, 11, 167. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Tang, C.J.; Wang, F.; Jiang, L.; Xiong, X.; Wang, M.; Rong, M.; Liu, Z.; Liang, S. Transcriptome analysis of the venom glands of the Chinese wolf spider Lycosa singoriensis. Zoology 2010, 113, 10–18. [Google Scholar] [CrossRef]
- Hu, Z.; Chen, B.; Xiao, Z.; Zhou, X.; Liu, Z. Transcriptomic Analysis of the Spider Venom Gland Reveals Venom Diversity and Species Consanguinity. Toxins 2019, 11, 68. [Google Scholar] [CrossRef] [PubMed]
- Cheng, T.C.; Long, R.W.; Wu, Y.Q.; Guo, Y.B.; Liu, D.L.; Peng, L.; Li, D.Q.; Yang, D.W.; Liu, F.X.; Xia, Q.Y. Identification and characterization of toxins in the venom gland of the Chinese bird spider, Haplopelma hainanum, by transcriptomic analysis. Insect Sci. 2016, 23, 487–499. [Google Scholar] [CrossRef]
- Estrada-Gomez, S.; Cardoso, F.C.; Vargas-Munoz, L.J.; Quintana-Castillo, J.C.; Arenas Gomez, C.M.; Pineda, S.S.; Saldarriaga-Cordoba, M.M. Venomic, Transcriptomic, and Bioactivity Analyses of Pamphobeteus verdolaga venom reveal Complex Disuldide-Rich Peptides That Modulate Calcium Channels. Toxins 2019, 11, 496. [Google Scholar] [CrossRef]
- Brown, R.L.; Haley, T.L.; West, K.A.; Crabb, J.W. Pseudechetoxin: A peptide blocker of cyclic nucleotide-gated ion channels. Proc. Natl. Acad. Sci. USA 1999, 96, 754–759. [Google Scholar] [CrossRef]
- Yamazaki, Y.; Brown, R.L.; Morita, T. Purification and cloning of toxins from elapid venoms that target cyclic nucleotide-gated ion channels. Biochemistry 2002, 41, 11331–11337. [Google Scholar] [CrossRef] [PubMed]
- Ito, N.; Mita, M.; Takahashi, Y.; Matsushima, A.; Watanabe, Y.G.; Hirano, S.; Odani, S. Novel cysteine-rich secretory protein in the buccal gland secretion of the parasitic lamprey Lethenteron japonicum. Biochem. Biophys. Res. Commun. 2007, 358, 35–40. [Google Scholar] [CrossRef] [PubMed]
- Ribeiro, J.M.C.; Andersen, J.; Silva-Neto, M.A.C.; Pham, V.M.; Garfield, M.K.; Valenzuela, J.G. Exploring the sialome of the blood-sucking bug Rhodnius prolixus. Insect Biochem. Mol. Biol. 2004, 34, 61–79. [Google Scholar] [CrossRef] [PubMed]
- Caruso, B.; Bonadonna, P.; Bovo, C.; Melloni, N.; Lombardo, C.; Senna, G.; Lippi, G. Wasp venom allergy screening with recombinant allergen testing. Diagnostic performance of rPol d 5 and and rVes v 5 for differentiating sensitization to Vespula and Polistes subspecies. Clin. Chim. Acta 2016, 453, 170–173. [Google Scholar] [CrossRef] [PubMed]
- Fang, K.S.Y.; Vitale, M.; Fehlner, P.; King, T.P. cDNA cloning and primary structure of a white-face hornet venom allergen, antigen. Proc. Natl. Acad. Sci. USA 1988, 85, 895–899. [Google Scholar] [CrossRef]
- Milne, T.J.; Abbenante, G.; Tyndall, J.D.A.; Halliday, J.; Lewis, R.J. Isolation and characterization of of a cone snail protease with homology to CRISP proteins of the pathogenesis-related protein superfamily. J. Biol. Chem. 2003, 278, 31105–31110. [Google Scholar] [CrossRef]
- Undheim, E.A.B.; Grimm, L.L.; Low, C.F.; Morgenstern, D.; Herzig, V.; Zobel-Thropp, P.A.; Pineda, S.S.; Habib, R.; Dziemborowicz, S.; Fry, B.G.; et al. Weaponization of a Hormone: Convergent recruitment of Hyperglycemic Hormone into the Venom of Arthropod Predators. Structure 2015, 23, 1283–1292. [Google Scholar] [CrossRef]
- Kuhn-Nentwig, L.; Largiader, C.R.; Streitberger, K.; Chandru, S.; Baumann, T.; Kampfer, U.; Schaller, J.; Schurch, S.; Nentwig, W. Purification, cDNA structure and biological significance of a single insulin-like growth factor-binding domain protein (SIBD-1) identified in the hemocytes of the spider Cupiennius salei. Insect Biochem. Mol. Biol. 2011, 41, 891–901. [Google Scholar] [CrossRef]
- Esteves, E.; Maruyama, S.R.; Kawahara, R.; Fujita, A.; Martins, L.A.; Righi, A.A.; Costa, F.B.; Palmisano, G.; Labruna, M.B.; Sa-Nunes, A.; et al. Analysis of the Salivary Gland Transcriptome of Unfed and Partially Fed Amblyomma sculptum Ticks and descriptive Proteome of the Sliva. Front. Cell. Infect. Microbiol. 2017, 7, 476. [Google Scholar] [CrossRef]
- Rokyta, D.R.; Ward, M. Venom-gland transcriptomics and venom proteomics of the black-back scorpion (Hadrurus spadix) reveal detectability challenges and an unexplored realm of animal toxin diversity. Toxicon 2017, 128, 23–37. [Google Scholar] [CrossRef]
- Santibanez-Lopez, C.E.; Cid-Uribe, J.I.; Batista, C.V.; Ortiz, E.; Possani, L.D. Venom Gland Transcriptomic and Proteomic Analyses of the Enigmatic Scorpion Superstitionia donensis (Scorpiones: Superstitioniidae), with Insights on the Evolution of its Venom Components. Toxins 2016, 8, 367. [Google Scholar] [CrossRef] [PubMed]
- Ward, M.J.; Ellsworth, S.A.; Rokyta, D.R. Venom-gland transcriptomics and proteomics of the Hentz striped scorpion (Centruroides hentzi; Buthidae) reveal high toxin diversity in a harmless member of a lethal family. Toxicon 2017, 142, 14–29. [Google Scholar] [CrossRef] [PubMed]
- Kono, N.; Nakamura, H.; Ohtoshi, R.; Moran, D.A.P.; Shinohara, A.; Yoshida, Y.; Fujiwara, M.; Mori, M.; Tomita, M.; Arakawa, K. Orb-weaving spider Araneus ventricosus genome elucidates the spidroin gene catalogue. Sci. Rep. 2019, 9, 8380. [Google Scholar] [CrossRef]
- Arvidson, R.; Kaiser, M.; Lee, S.S.; Urenda, J.P.; Dail, C.; Mohammed, H.; Nolan, C.; Pan, S.; Stajich, J.E.; Libersat, F.; et al. Parasitoid Jewel Wasp Mounts Multipronged Neurochemical Attack to Hijack a Host brain. Mol. Cell Proteom. 2019, 18, 99–114. [Google Scholar] [CrossRef] [PubMed]
- Yap, M.K.K.; Misuan, N. Exendin-4 from Heloderma suspectum venom: From discovery to its latest application as type II diabetes combatant. Basic Clin. Pharmacol. Toxicol. 2019, 124, 513–527. [Google Scholar] [CrossRef] [PubMed]
- Gaudino, G.; Fasolo, A.; Merlo, G.; Lazarus, L.H.; Renda, T.; D.´este, L.; Vandesande, F. Active peptides from amphibian skin are also amphibian neuropeptides. Peptides 1985, 6, 209–213. [Google Scholar] [CrossRef]
- Roelants, K.; Fry, B.G.; Norman, J.A.; Clynen, E.; Schoofs, L.; Bossuyt, F. Identical skin toxins by convergent molecular adaptions in frogs. Curr. Biol. 2010, 20, 125–130. [Google Scholar] [CrossRef]
- Roelants, K.; Fry, B.G.; Lumen, Y.; Stijlemans, B.; Brys, L.; Kok, P.; Clynen, E.; Schoofs, L.; Cornelis, P.; Bossuyt, F. Origin and Functional Diversification of an Amphibian Defense Peptide Arsenal. PLoS Genet. 2013, 9, e1003662. [Google Scholar] [CrossRef]
- Lüddecke, T.; Schulz, S.; Steinfartz, S.; Vences, M. A salamander´s toxic arsenal: Review of skin poison diversity and function in true salamanders, genus Salamandra. Sci. Nat. 2018, 105, 56. [Google Scholar] [CrossRef]
- Daltry, J.C.; Wüster, W. Diet and snake venom evolution. Nature 1996, 379, 537–540. [Google Scholar] [CrossRef]
- Li, M.; Fry, B.G.; Kini, M. Eggs-Only Diet: Its Implications for the Toxin Profile Changes and Ecology of the Marbled Sea Snake (Aipysurus eydouxii). J. Mol. Evol. 2005, 60, 81–89. [Google Scholar] [CrossRef] [PubMed]
- Fry, B.G.; Wüster, W.; Kini, M.; Brusic, V.; Khan, A.; Venkataraman, D.; Rooney, A.P. Molecular evolution and phylogeny of elapid snake venom three-finger toxins. J. Mol. Evol. 2006, 57, 110–129. [Google Scholar] [CrossRef] [PubMed]
- Lyons, K.; Dugon, M.M.; Healy, K. Diet Breadth Mediates the Prey Specificity of Venom Potency in Snakes. Toxins 2020, 12, 74. [Google Scholar] [CrossRef] [PubMed]
- Phuong, M.A.; Mahardika, G.N.; Alfaro, M.E. Dietary breadth is positively correlated with venom complexity in cone snails. BMC Genom. 2016, 17, 401. [Google Scholar] [CrossRef] [PubMed]
- Szymkowiak, P.; Tryjanowski, P.; Winiecki, A.; Grobelny, S.; Konwerski, S. Habitat differences in the food composition of the wasp-like spider Argiope bruennichi (Scop.) (Aranei: Araneidae) in Poland. Belg. J. Zool. 2005, 135, 33–37. [Google Scholar]
- Dean, J.; Aneshansley, D.J.; Edgerton, H.E.; Eisner, T. Defensive spray of the bombardier beetle: A biological pulse jet. Science 1990, 248, 1219–1221. [Google Scholar] [CrossRef]
- Nyffeler, M.; Benz, G. Eine Notiz zum Beutefangverhalten der Radnetzspinne Argiope bruennichi (Scopoli) (Araneae, Araneidae). Revue Suisse Zool. 1982, 89, 23–25. [Google Scholar] [CrossRef]
- Harwood, R.H. Predatory behaviour of Argiope aurantia (Lucas). Am. Midl. Nat. 1974, 91, 130–139. [Google Scholar] [CrossRef]
- Robinson, M.H. Predatory behaviour of Argiope argentata (Fabricius). Am. Zool. 1969, 9, 161–174. [Google Scholar] [CrossRef]
- Nisani, Z.; Boskovic, D.S.; Dunbar, S.G.; Kelln, W.; Hayes, W.K. Investigating the chemical profile of regenerated scorpion (Parabuthus transvaalicus) venom in relation to metabolic cost and toxicity. Toxicon 2012, 60, 315–323. [Google Scholar] [CrossRef]
- Blennerhassett, R.A.; Bell-Anderson, K.; Shine, R.; Brown, G.P. The cost of chemical defence: The impact of toxin depletion on growth and behaviour of cane toads (Rhinella marina). Proc. Biol. Sci. 2019, 286, 20190867. [Google Scholar] [CrossRef] [PubMed]
- Saggiomo, S.L.; Zelenka, C.; Seymor, J. Relationship between food and venom production in the estuarine stonefish Synanceia horrida. Toxicon 2017, 125, 19–23. [Google Scholar] [CrossRef] [PubMed]
- Walker, A.A.; Mayhew, M.L.; Jin, J.; Herzig, V.; Undheim, E.A.B.; Sombke, A.; Fry, B.G.; Merrit, D.J.; King, G.F. The assassin bug Pristesancus plagipennis produces two distinct venoms in separate gland lumens. Nat. Commun. 2018, 9, 755. [Google Scholar] [CrossRef] [PubMed]
- Guehrs, K.H.; Schlott, B.; Grosse, F.; Weisshart, K. Environmental conditions impinge on dragline silk protein composition. Insect Mol. Biol. 2008, 17, 553–564. [Google Scholar] [CrossRef] [PubMed]
- Zobel-Thropp, P.A.; Mullins, J.; Kristensen, C.; Kronmiller, B.A.; David, C.L.; Breci, L.A.; Binford, G.J. Not so dangerous After All? Venom Composition and Potency of the Pholcid (Daddy Long-leg) Spider Physocyclus mexicanus. Front. Ecol. Evol. 2019, 7, 256. [Google Scholar] [CrossRef]
| Spot ID | Class | Score | kDa | Peptides | Coverage (%) | ppm |
|---|---|---|---|---|---|---|
| 8501 | CAP | 136.0 | 48.9 | 14 | 21.1 | 13.81 |
| 7309 | CAP | 70.7 | 28.3 | 6 | 18.3 | 7.42 |
| 7501 | CAP | 182.0 | 50.0 | 18 | 35.3 | 23.55 |
| 6502 | CAP | 85.9 | 50.5 | 10 | 24.8 | 22.95 |
| 6502 | CAP | 124.0 | 45.7 | 15 | 33.7 | 23.73 |
| 6502 | CAP | 89.2 | 48.1 | 11 | 25.3 | 23.57 |
| Protein Class | Matched Peptides | MW (kDa) | Calc. pI | Mascot Score | Coverage (%) |
|---|---|---|---|---|---|
| 5′ Nucleotidase | 4 | 24.3 | 4.54 | 228 | 17 |
| Astacin-like metalloprotease | 2 | 10.9 | 4.79 | 149 | 15 |
| Astacin-like metalloprotease | 2 | 14.4 | 5.87 | 76 | 13 |
| Astacin-like metalloprotease | 3 | 17.0 | 7.93 | 50 | 24 |
| CAP | 23 | 51.2 | 7.77 | 3697 | 57 |
| CAP | 18 | 50.8 | 8.19 | 2333 | 52 |
| CAP | 14 | 50.0 | 7.65 | 2164 | 40 |
| CAP | 9 | 28.3 | 7.66 | 1918 | 59 |
| CAP | 16 | 48.9 | 8.44 | 1904 | 46 |
| CAP | 8 | 50.5 | 7.97 | 1661 | 26 |
| CAP | 15 | 51.0 | 8.50 | 1127 | 40 |
| CAP | 14 | 48.0 | 8.07 | 1020 | 46 |
| CAP | 11 | 28.9 | 9.09 | 944 | 44 |
| CAP | 5 | 17.0 | 5.14 | 604 | 32 |
| CAP | 6 | 13.2 | 8.79 | 311 | 49 |
| CAP | 3 | 25.1 | 6.84 | 173 | 27 |
| Cystatin | 2 | 15.6 | 7.33 | 48 | 22 |
| Diuretic hormone-like | 2 | 14.6 | 9.55 | 267 | 23 |
| EF-hand | 7 | 22.3 | 5.25 | 112 | 33 |
| ICK | 8 | 15.0 | 6.15 | 1076 | 40 |
| ICK | 2 | 14.7 | 4.68 | 209 | 14 |
| ICK | 4 | 15.7 | 6.76 | 59 | 24 |
| IGFBP | 2 | 18.8 | 4.92 | 63 | 16 |
| ITG-like peptide | 10 | 26.6 | 4.78 | 1666 | 51 |
| ITG-like peptide | 8 | 24.7 | 4.96 | 673 | 38 |
| Kunitz | 2 | 25.1 | 7.46 | 38 | 8 |
| Leucine-rich-repeat domain | 20 | 39.7 | 4.93 | 4279 | 71 |
| Leucine-rich-repeat domain | 10 | 41.0 | 5.29 | 771 | 34 |
| Leucine-rich-repeat domain | 9 | 36.9 | 5.74 | 428 | 37 |
| Leucine-rich-repeat domain | 7 | 39.2 | 5.11 | 215 | 27 |
| Leucine-rich-repeat domain | 5 | 41.2 | 5.50 | 117 | 23 |
| MIT-atracotoxin | 4 | 10.7 | 4,94 | 448 | 56 |
| MIT-atracotoxin | 5 | 9.8 | 5.50 | 140 | 66 |
| Prokineticin | 4 | 13.8 | 7.97 | 636 | 33 |
| Putative chitinase | 12 | 35.5 | 7.30 | 1159 | 45 |
| Putative chitinase | 8 | 30.1 | 6.49 | 219 | 39 |
| Putative chitinase | 2 | 18.0 | 5.19 | 112 | 11 |
| Putative chitinase | 3 | 47.4 | 11.15 | 53 | 13 |
| S10 peptidase | 4 | 51.4 | 8.07 | 118 | 15 |
| Serine protease | 8 | 53.2 | 6.40 | 608 | 24 |
| Serine protease | 8 | 53.2 | 6.64 | 469 | 23 |
| Serine protease | 5 | 86.2 | 6.54 | 135 | 9 |
| Serine protease | 3 | 99.1 | 6.27 | 100 | 4 |
| Serine protease | 3 | 55.2 | 6.13 | 54 | 7 |
| Techylectin | 3 | 40.3 | 7.17 | 88 | 7 |
| Thyroglobulin-like | 3 | 10.9 | 7.56 | 134 | 31 |
| Unclassified aranetoxins | 3 | 8.3 | 8.07 | 630 | 55 |
| Unclassified aranetoxins | 5 | 12.9 | 9.57 | 220 | 32 |
| Unclassified aranetoxins | 4 | 8.2 | 8.18 | 206 | 39 |
| Unclassified aranetoxins | 4 | 8.3 | 8.18 | 202 | 39 |
| Unclassified aranetoxins | 2 | 8.4 | 7.71 | 56 | 39 |
| Venom protein 11 | 2 | 9.5 | 8.00 | 234 | 35 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lüddecke, T.; von Reumont, B.M.; Förster, F.; Billion, A.; Timm, T.; Lochnit, G.; Vilcinskas, A.; Lemke, S. An Economic Dilemma between Molecular Weapon Systems May Explain an Arachno-Atypical Venom in Wasp Spiders (Argiope bruennichi). Biomolecules 2020, 10, 978. https://doi.org/10.3390/biom10070978
Lüddecke T, von Reumont BM, Förster F, Billion A, Timm T, Lochnit G, Vilcinskas A, Lemke S. An Economic Dilemma between Molecular Weapon Systems May Explain an Arachno-Atypical Venom in Wasp Spiders (Argiope bruennichi). Biomolecules. 2020; 10(7):978. https://doi.org/10.3390/biom10070978
Chicago/Turabian StyleLüddecke, Tim, Björn M. von Reumont, Frank Förster, André Billion, Thomas Timm, Günter Lochnit, Andreas Vilcinskas, and Sarah Lemke. 2020. "An Economic Dilemma between Molecular Weapon Systems May Explain an Arachno-Atypical Venom in Wasp Spiders (Argiope bruennichi)" Biomolecules 10, no. 7: 978. https://doi.org/10.3390/biom10070978
APA StyleLüddecke, T., von Reumont, B. M., Förster, F., Billion, A., Timm, T., Lochnit, G., Vilcinskas, A., & Lemke, S. (2020). An Economic Dilemma between Molecular Weapon Systems May Explain an Arachno-Atypical Venom in Wasp Spiders (Argiope bruennichi). Biomolecules, 10(7), 978. https://doi.org/10.3390/biom10070978









