Structure of the ALS Mutation Target Annexin A11 Reveals a Stabilising N-Terminal Segment
Abstract
1. Introduction
2. Materials and Methods
2.1. Cloning of Full-Length and N-Terminally Truncated Forms of AnxA11
2.2. Expression and Purification of Full-Length and N-Terminally Truncated Forms of AnxA11
2.3. Circular Dichroism Spectroscopy
2.4. Small-Angle X-Ray Scattering
2.5. Protein Crystallisation, Data Collection, and Structure Determination
3. Results and Discussion
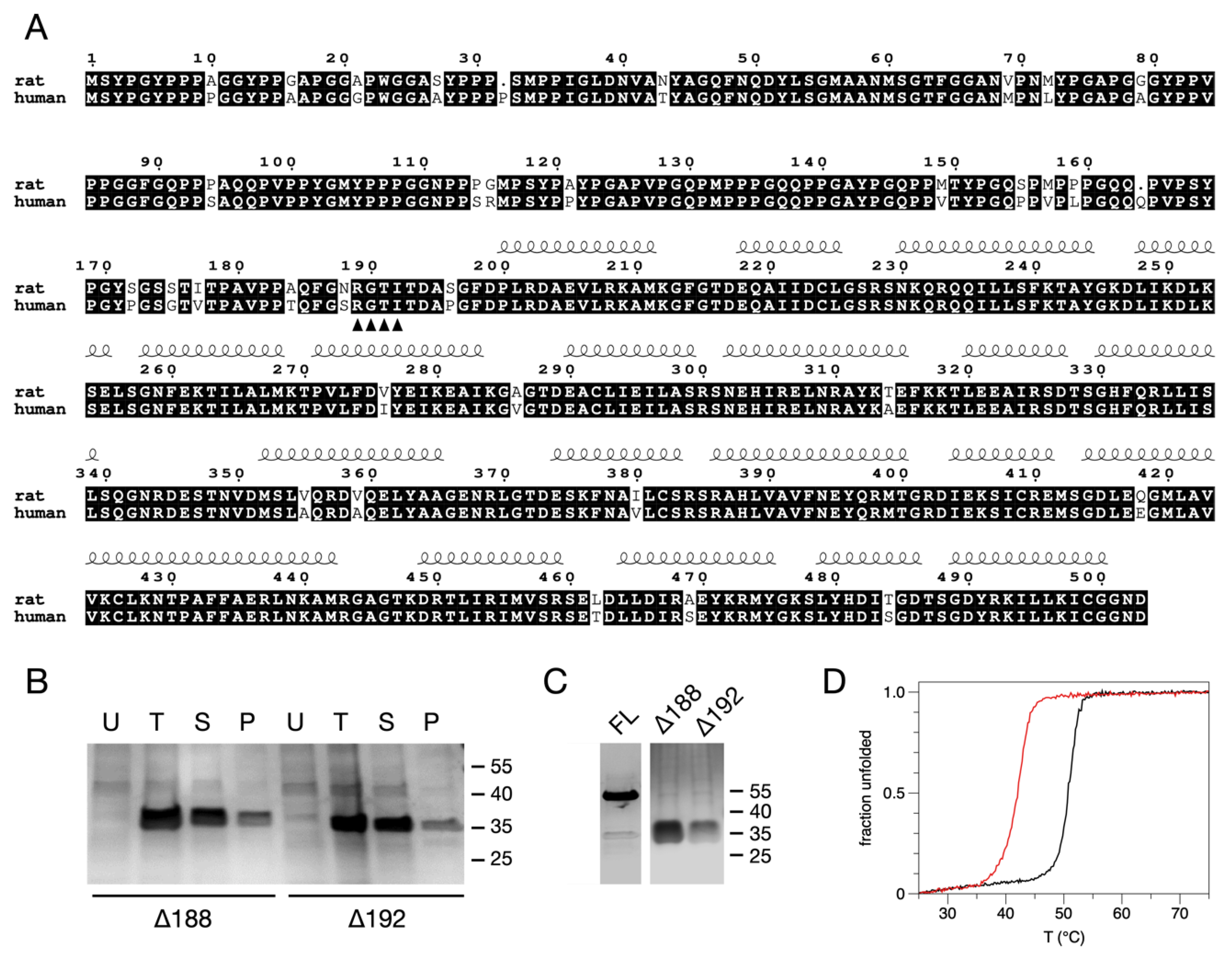
3.1. Solubility Assays, Large-Scale Expression, and Purification of the N-Terminally Truncated Forms of AnxA11
3.2. Thermal Stability of the N-Terminally Truncated Δ188AnxA11 and Δ192AnxA11
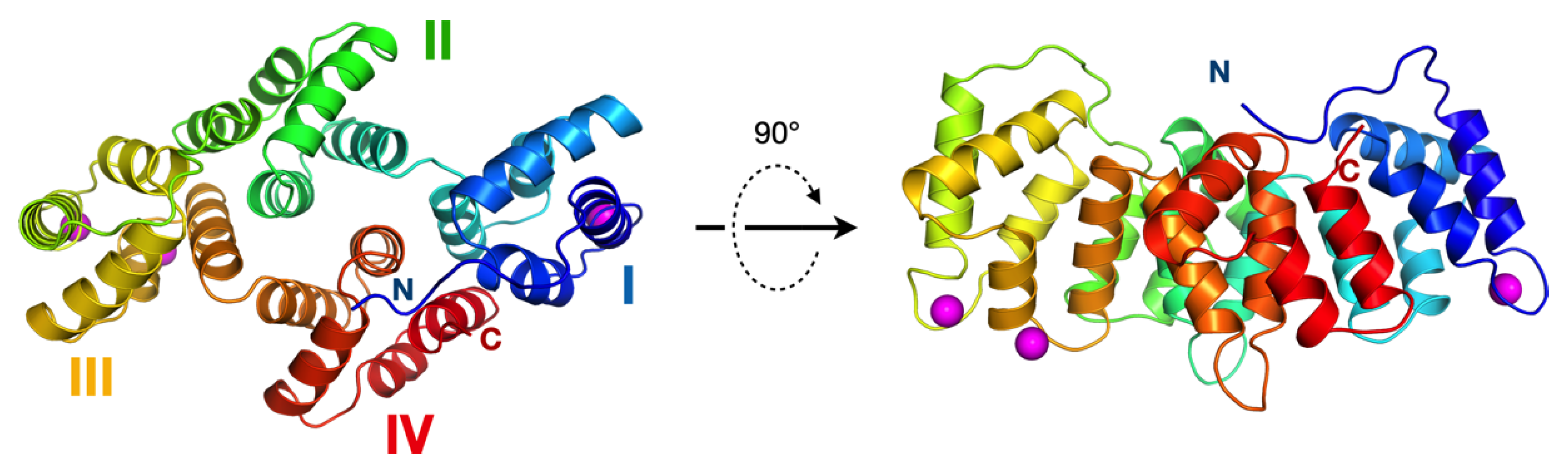
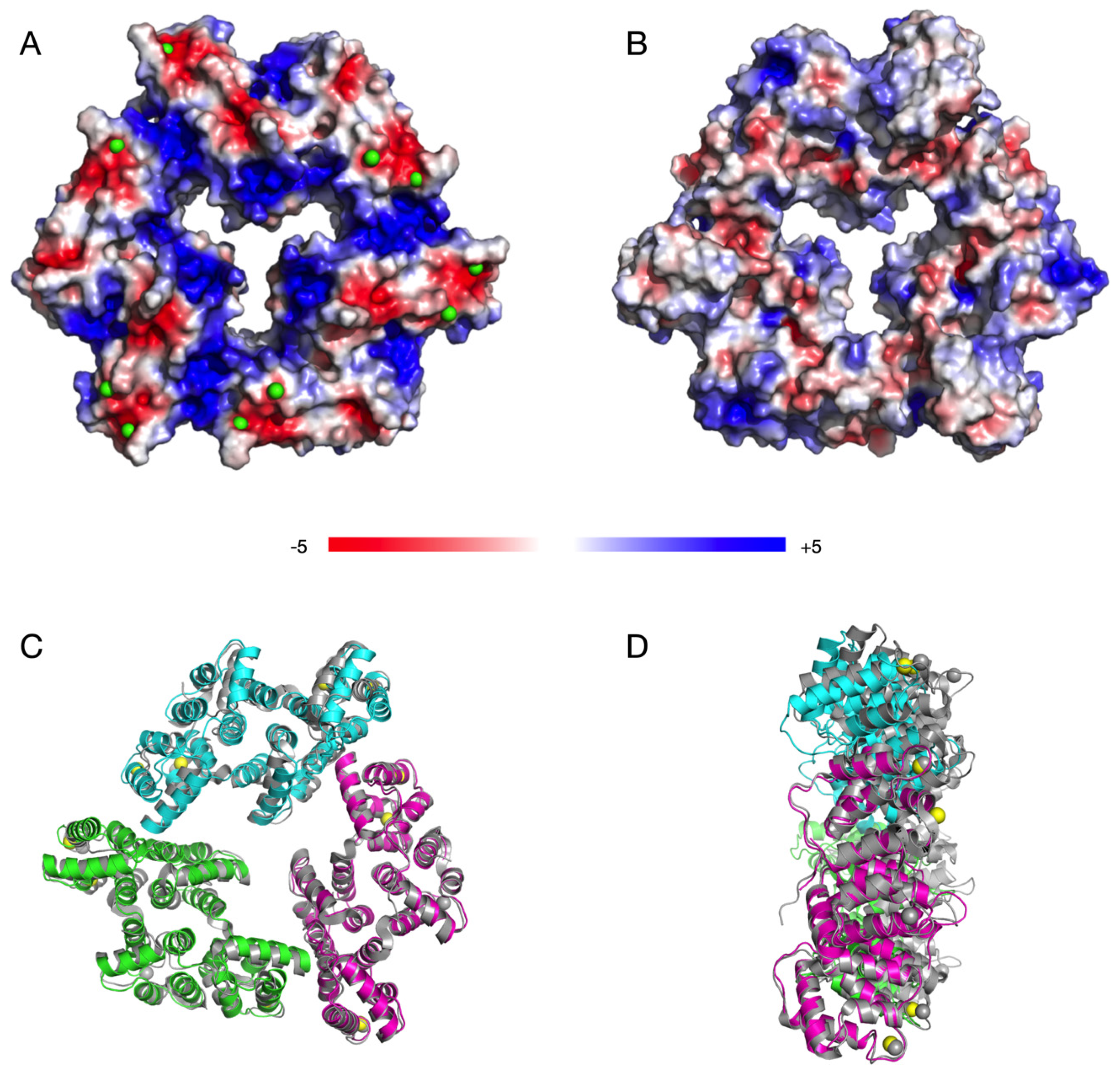
3.3. The Crystal Structure of AnxA11
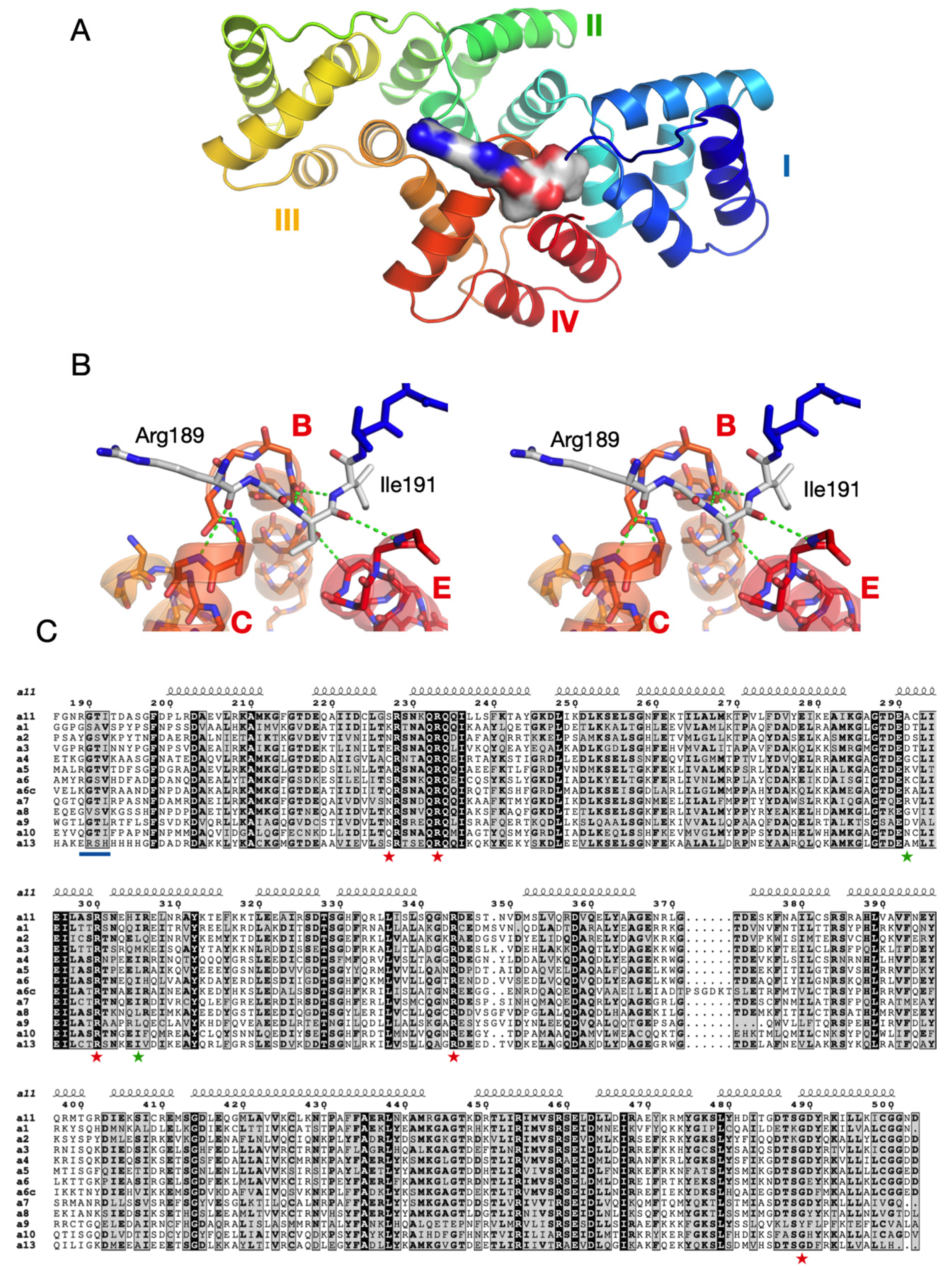
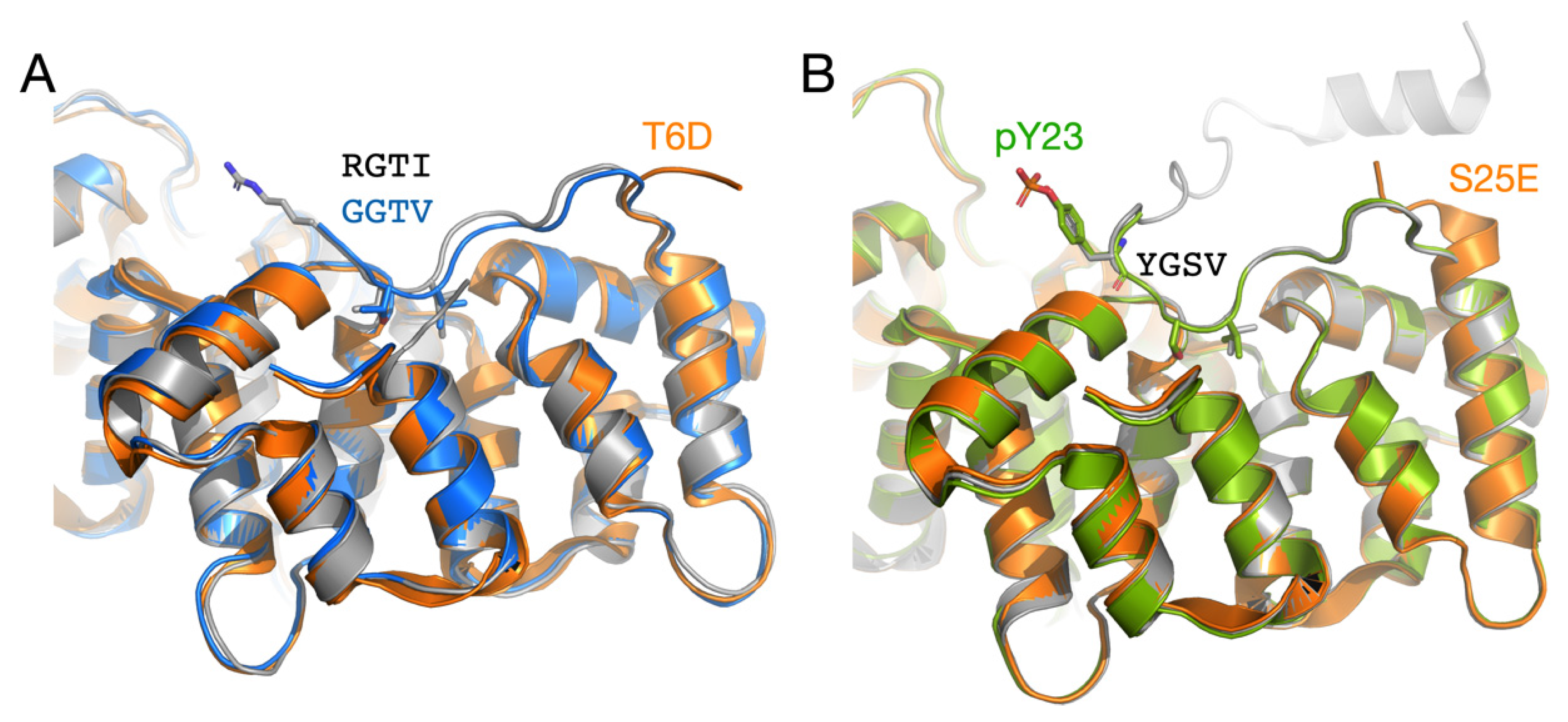
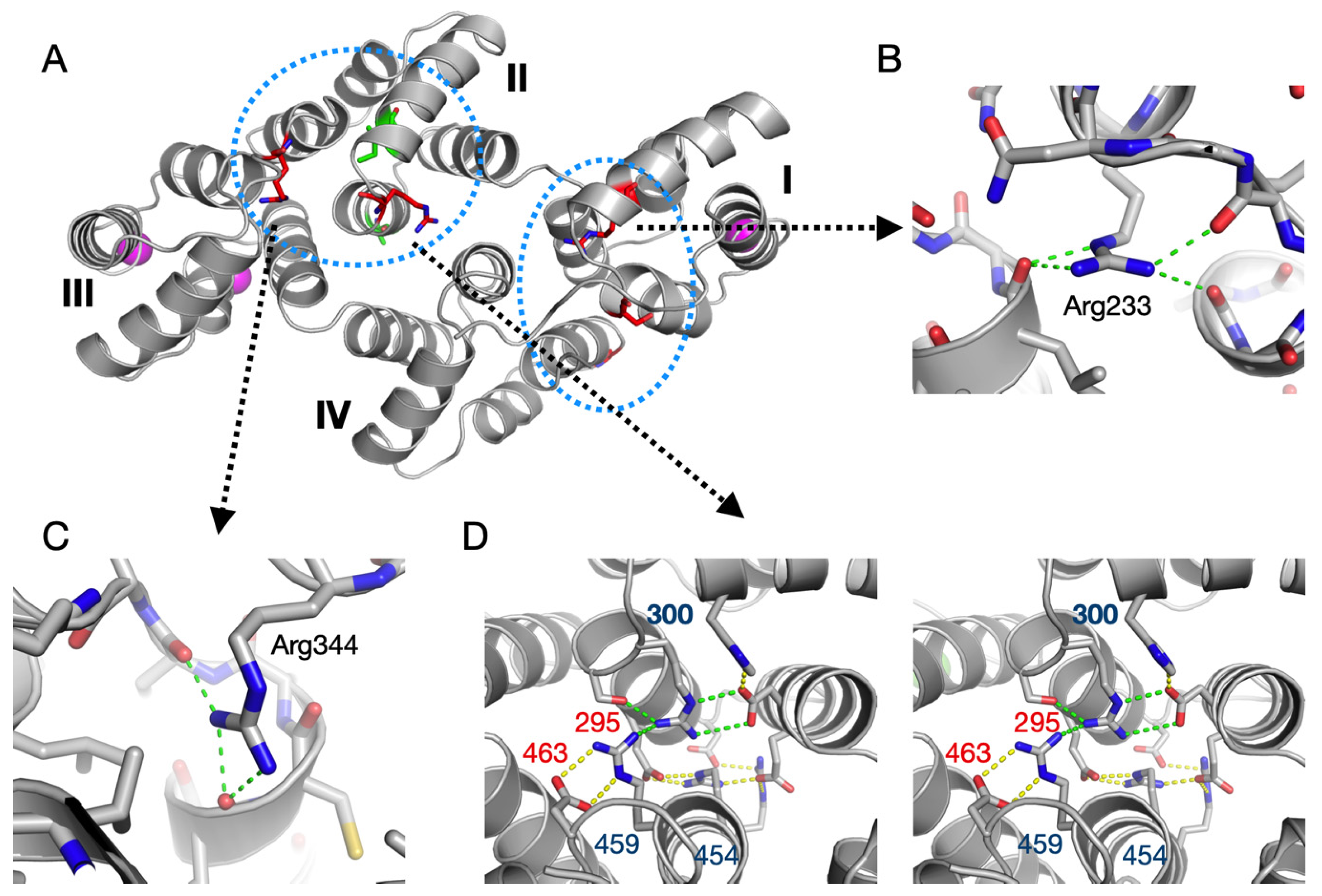
3.4. Structure-Based Analysis of ALS Disease Mutations
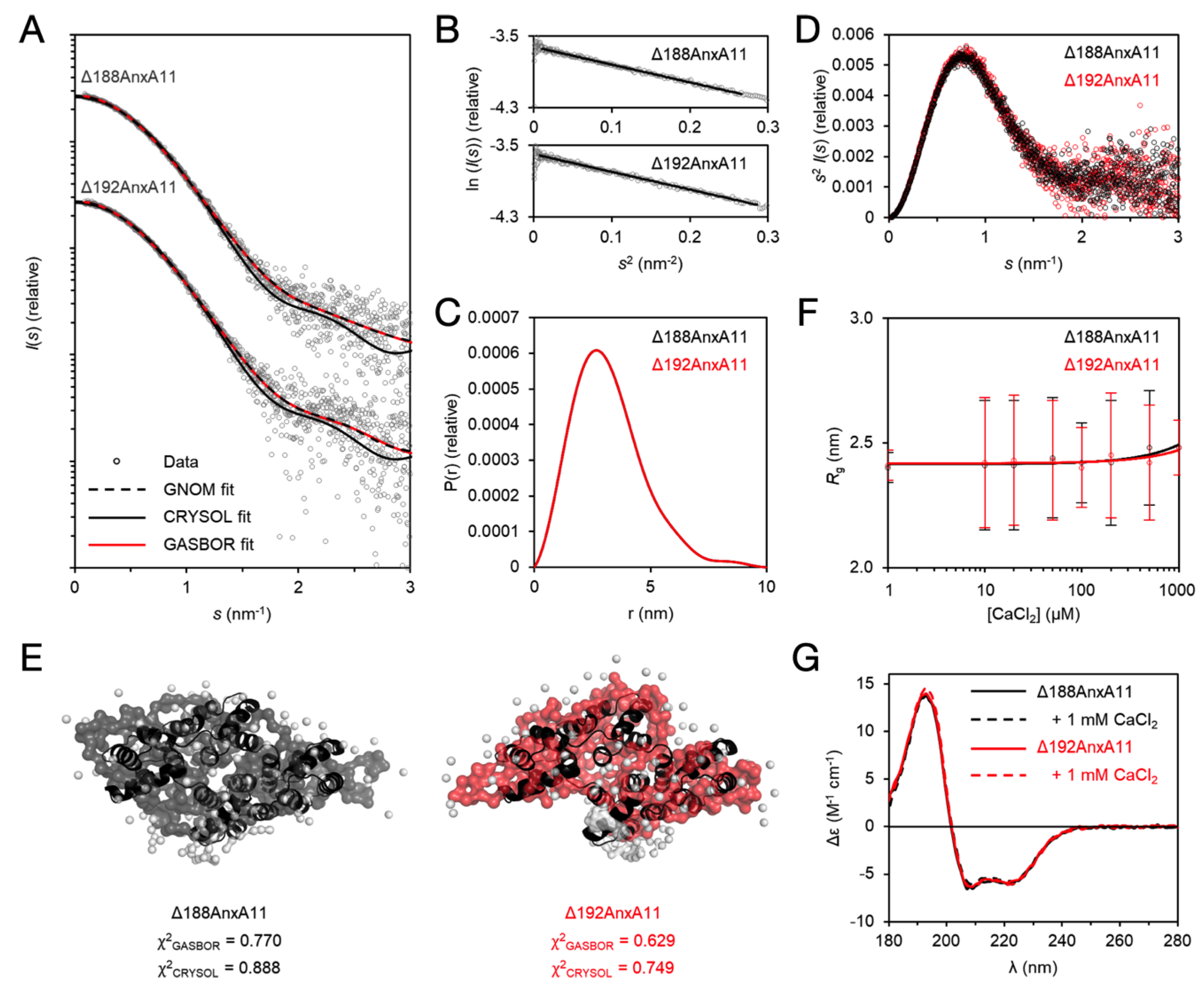
3.5. The Solution Behaviour of the AnxA11 Core
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Gerke, V.; Moss, S.E. Annexins: From structure to function. Physiol. Rev. 2002, 82, 331–371. [Google Scholar] [CrossRef] [PubMed]
- Gerke, V.; Creutz, C.E.; Moss, S.E. Annexins: Linking Ca2+ signalling to membrane dynamics. Nat. Rev. Mol. Cell Biol. 2005, 6, 449–461. [Google Scholar] [CrossRef] [PubMed]
- Moss, S.E.; Morgan, R.O. The annexins. Genome Biol. 2004, 5, 219. [Google Scholar] [CrossRef]
- Liemann, S.; Huber, R. Three-dimensional structure of annexins. Cell Mol. Life Sci. 1997, 53, 516–521. [Google Scholar] [CrossRef] [PubMed]
- Morgan, R.O.; Jenkins, N.A.; Gilbert, D.J.; Copeland, N.G.; Balsara, B.R.; Testa, J.R.; Fernandez, M.P. Novel human and mouse annexin A10 are linked to the genome duplications during early chordate evolution. Genomics 1999, 60, 40–49. [Google Scholar] [CrossRef] [PubMed]
- Rosengarth, A.; Luecke, H. A calcium-driven conformational switch of the N-terminal and core domains of annexin A1. J. Mol. Biol. 2003, 326, 1317–1325. [Google Scholar] [CrossRef]
- Boye, T.L.; Jeppesen, J.C.; Maeda, K.; Pezeshkian, W.; Solovyeva, V.; Nylandsted, J.; Simonsen, A.C. Annexins induce curvature on free-edge membranes displaying distinct morphologies. Sci. Rep. 2018, 8, 10309. [Google Scholar] [CrossRef]
- Liao, Y.C.; Fernandopulle, M.S.; Wang, G.; Choi, H.; Hao, L.; Drerup, C.M.; Patel, R.; Qamar, S.; Nixon-Abell, J.; Shen, Y.; et al. RNA Granules Hitchhike on Lysosomes for Long-Distance Transport, Using Annexin A11 as a Molecular Tether. Cell 2019, 179, 147–164.e20. [Google Scholar] [CrossRef]
- Tokumitsu, H.; Mizutani, A.; Muramatsu, M.; Yokota, T.; Arai, K.; Hidaka, H. Molecular cloning of rabbit CAP-50, a calcyclin-associated annexin protein. Biochem. Biophys. Res. Commun. 1992, 186, 1227–1235. [Google Scholar] [CrossRef]
- Burns, A.L.; Magendzo, K.; Shirvan, A.; Srivastava, M.; Rojas, E.; Alijani, M.R.; Pollard, H.B. Calcium channel activity of purified human synexin and structure of the human synexin gene. Proc. Natl. Acad. Sci. USA 1989, 86, 3798–3802. [Google Scholar] [CrossRef]
- Creutz, C.E.; Snyder, S.L.; Husted, L.D.; Beggerly, L.K.; Fox, J.W. Pattern of repeating aromatic residues in synexin. Similarity to the cytoplasmic domain of synaptophysin. Biochem. Biophys. Res. Commun. 1988, 152, 1298–1303. [Google Scholar] [CrossRef]
- Creutz, C.E. Novel protein ligands of the annexin A7 N-terminal region suggest pro-beta helices engage one another with high specificity. Gen. Physiol. Biophys. 2009, 28, F7–F13. [Google Scholar]
- Lecona, E.; Turnay, J.; Olmo, N.; Guzman-Aranguez, A.; Morgan, R.O.; Fernandez, M.P.; Lizarbe, M.A. Structural and functional characterization of recombinant mouse annexin A11: Influence of calcium binding. Biochem. J. 2003, 373, 437–449. [Google Scholar] [CrossRef] [PubMed]
- Satoh, H.; Shibata, H.; Nakano, Y.; Kitaura, Y.; Maki, M. ALG-2 interacts with the amino-terminal domain of annexin XI in a Ca(2+)-dependent manner. Biochem. Biophys. Res. Commun. 2002, 291, 1166–1172. [Google Scholar] [CrossRef] [PubMed]
- Mizutani, A.; Tokumitsu, H.; Kobayashi, R.; Hidaka, H. Phosphorylation of annexin XI (CAP-50) in SR-3Y1 cells. J. Biol. Chem. 1993, 268, 15517–15522. [Google Scholar]
- Sudo, T.; Hidaka, H. Regulation of calcyclin (S100A6) binding by alternative splicing in the N-terminal regulatory domain of annexin XI isoforms. J. Biol. Chem. 1998, 273, 6351–6357. [Google Scholar] [CrossRef]
- Tokumitsu, H.; Mizutani, A.; Hidaka, H. Calcyclin-binding site located on the NH2-terminal domain of rabbit CAP-50 (annexin XI): Functional expression of CAP-50 in Escherichia coli. Arch. Biochem. Biophys. 1993, 303, 302–306. [Google Scholar] [CrossRef]
- Tomas, A.; Moss, S.E. Calcium- and cell cycle-dependent association of annexin 11 with the nuclear envelope. J. Biol. Chem. 2003, 278, 20210–20216. [Google Scholar] [CrossRef]
- Naka, M.; Qing, Z.X.; Sasaki, T.; Kise, H.; Tawara, I.; Hamaguchi, S.; Tanaka, T. Purification and characterization of a novel calcium-binding protein, S100C, from porcine heart. Biochim. Biophys. Acta 1994, 1223, 348–353. [Google Scholar] [CrossRef]
- Powell, M.A.; Glenney, J.R. Regulation of calpactin I phospholipid binding by calpactin I light-chain binding and phosphorylation by p60v-src. Biochem. J. 1987, 247, 321–328. [Google Scholar] [CrossRef]
- Consortium, T.U. UniProt: A worldwide hub of protein knowledge. Nucleic Acids Res. 2019, 47, D506–D515. [Google Scholar] [CrossRef] [PubMed]
- Mizutani, A.; Usuda, N.; Tokumitsu, H.; Minami, H.; Yasui, K.; Kobayashi, R.; Hidaka, H. CAP-50, a newly identified annexin, localizes in nuclei of cultured fibroblast 3Y1 cells. J. Biol. Chem. 1992, 267, 13498–13504. [Google Scholar] [PubMed]
- Mizutani, A.; Watanabe, N.; Kitao, T.; Tokumitsu, H.; Hidaka, H. The long amino-terminal tail domain of annexin XI is necessary for its nuclear localization. Arch. Biochem. Biophys. 1995, 318, 157–165. [Google Scholar] [CrossRef] [PubMed]
- Tomas, A.; Futter, C.; Moss, S.E. Annexin 11 is required for midbody formation and completion of the terminal phase of cytokinesis. J. Cell Biol. 2004, 165, 813–822. [Google Scholar] [CrossRef] [PubMed]
- Duncan, R.; Carpenter, B.; Main, L.C.; Telfer, C.; Murray, G.I. Characterisation and protein expression profiling of annexins in colorectal cancer. Br. J. Cancer 2008, 98, 426–433. [Google Scholar] [CrossRef]
- Fernandez-Madrid, F.; Tang, N.; Alansari, H.; Granda, J.L.; Tait, L.; Amirikia, K.C.; Moroianu, M.; Wang, X.; Karvonen, R.L. Autoantibodies to Annexin XI-A and Other Autoantigens in the Diagnosis of Breast Cancer. Cancer Res. 2004, 64, 5089–5096. [Google Scholar] [CrossRef]
- Hua, K.; Li, Y.; Zhao, Q.; Fan, L.; Tan, B.; Gu, J. Downregulation of Annexin A11 (ANXA11) Inhibits Cell Proliferation, Invasion, and Migration via the AKT/GSK-3beta Pathway in Gastric Cancer. Med. Sci. Monit. 2018, 24, 149–160. [Google Scholar] [CrossRef]
- Song, J.; Shih Ie, M.; Chan, D.W.; Zhang, Z. Suppression of annexin A11 in ovarian cancer: Implications in chemoresistance. Neoplasia 2009, 11, 605–614, 1 p following 614. [Google Scholar] [CrossRef]
- Wang, J.; Guo, C.; Liu, S.; Qi, H.; Yin, Y.; Liang, R.; Sun, M.Z.; Greenaway, F.T. Annexin A11 in disease. Clin. Chim. Acta 2014, 431, 164–168. [Google Scholar] [CrossRef]
- Xin, W.; Rhodes, D.R.; Ingold, C.; Chinnaiyan, A.M.; Rubin, M.A. Dysregulation of the annexin family protein family is associated with prostate cancer progression. Am. J. Pathol. 2003, 162, 255–261. [Google Scholar] [CrossRef]
- Brenner, D.; Weishaupt, J.H. Update on amyotrophic lateral sclerosis genetics. Curr. Opin. Neurol. 2019, 32, 735–739. [Google Scholar] [CrossRef]
- Fernandopulle, M.; Wang, G.; Nixon-Abell, J.; Qamar, S.; Balaji, V.; Morihara, R.; St George-Hyslop, P.H. Inherited and Sporadic Amyotrophic Lateral Sclerosis and Fronto-Temporal Lobar Degenerations arising from Pathological Condensates of Phase Separating Proteins. Hum. Mol. Genet. 2019, 28, R187–R196. [Google Scholar] [CrossRef] [PubMed]
- Fratta, P.; Birsa, N.; Tosolini, A.P.; Schiavo, G. Travelling Together: A Unifying Pathomechanism for ALS. Trends Neurosci. 2020, 43, 1–2. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Wu, C.; He, J.; Zhang, N.; Fan, D. Two rare variants of the ANXA11 gene identified in Chinese patients with amyotrophic lateral sclerosis. Neurobiol. Aging 2019, 74, 235.e9–235.e12. [Google Scholar] [CrossRef] [PubMed]
- Smith, B.N.; Topp, S.D.; Fallini, C.; Shibata, H.; Chen, H.J.; Troakes, C.; King, A.; Ticozzi, N.; Kenna, K.P.; Soragia-Gkazi, A.; et al. Mutations in the vesicular trafficking protein annexin A11 are associated with amyotrophic lateral sclerosis. Sci. Transl. Med. 2017, 9. [Google Scholar] [CrossRef] [PubMed]
- Tsai, P.C.; Liao, Y.C.; Jih, K.Y.; Soong, B.W.; Lin, K.P.; Lee, Y.C. Genetic analysis of ANXA11 variants in a Han Chinese cohort with amyotrophic lateral sclerosis in Taiwan. Neurobiol. Aging 2018, 72, 188.e1–188.e2. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.; Liu, Q.; Liu, K.; Shen, D.; Tai, H.; Shu, S.; Ding, Q.; Fu, H.; Liu, S.; Wang, Z.; et al. ANXA11 mutations prevail in Chinese ALS patients with and without cognitive dementia. Neurol. Genet. 2018, 4, e237. [Google Scholar] [CrossRef]
- Chomczynski, P.; Sacchi, N. Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal. Biochem. 1987, 162, 156–159. [Google Scholar] [CrossRef]
- Miles, A.J.; Wallace, B.A. CDtoolX, a downloadable software package for processing and analyses of circular dichroism spectroscopic data. Protein Sci. 2018, 27, 1717–1722. [Google Scholar] [CrossRef]
- Blanchet, C.E.; Spilotros, A.; Schwemmer, F.; Graewert, M.A.; Kikhney, A.; Jeffries, C.M.; Franke, D.; Mark, D.; Zengerle, R.; Cipriani, F.; et al. Versatile sample environments and automation for biological solution X-ray scattering experiments at the P12 beamline (PETRA III, DESY). J. Appl. Crystallogr. 2015, 48, 431–443. [Google Scholar] [CrossRef]
- Franke, D.; Petoukhov, M.V.; Konarev, P.V.; Panjkovich, A.; Tuukkanen, A.; Mertens, H.D.T.; Kikhney, A.G.; Hajizadeh, N.R.; Franklin, J.M.; Jeffries, C.M.; et al. ATSAS 2.8: A comprehensive data analysis suite for small-angle scattering from macromolecular solutions. J. Appl. Crystallogr. 2017, 50, 1212–1225. [Google Scholar] [CrossRef] [PubMed]
- Svergun, D.I. Determination of the regularization parameter in indirect-transform methods using perceptual criteria. J. Appl. Crystallogr. 1992, 25, 495–503. [Google Scholar] [CrossRef]
- Svergun, D.; Barberato, C.; Koch, M.H.J. CRYSOL—A Program to Evaluate X-ray Solution Scattering of Biological Macromolecules from Atomic Coordinates. J. Appl. Crystallogr. 1995, 28, 768–773. [Google Scholar] [CrossRef]
- Svergun, D.I.; Petoukhov, M.V.; Koch, M.H. Determination of domain structure of proteins from X-ray solution scattering. Biophys. J. 2001, 80, 2946–2953. [Google Scholar] [CrossRef]
- Kozin, M.B.; Svergun, D.I. Automated matching of high-and low-resolution structural models. J. Appl. Crystallogr. 2001, 34, 33–41. [Google Scholar] [CrossRef]
- Cianci, M.; Bourenkov, G.; Pompidor, G.; Karpics, I.; Kallio, J.; Bento, I.; Roessle, M.; Cipriani, F.; Fiedler, S.; Schneider, T.R. P13, the EMBL macromolecular crystallography beamline at the low-emittance PETRA III ring for high- and low-energy phasing with variable beam focusing. J. Synchrotron Radiat. 2017, 24, 323–332. [Google Scholar] [CrossRef] [PubMed]
- Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 2010, 66, 125–132. [Google Scholar] [CrossRef]
- McCoy, A.J.; Grosse-Kunstleve, R.W.; Adams, P.D.; Winn, M.D.; Storoni, L.C.; Read, R.J. Phaser crystallographic software. J. Appl. Crystallogr. 2007, 40, 658–674. [Google Scholar] [CrossRef]
- Sutton, R.B.; Sprang, S.R. Three dimensional structure of annexin IV. In Annexins: Molecular Structure to Cellular Function; Seaton, B.A., Ed.; Landes Biosciences: Georgetown, TX, USA, 1996. [Google Scholar]
- Afonine, P.V.; Grosse-Kunstleve, R.W.; Echols, N.; Headd, J.J.; Moriarty, N.W.; Mustyakimov, M.; Terwilliger, T.C.; Urzhumtsev, A.; Zwart, P.H.; Adams, P.D. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D Biol. Crystallogr. 2012, 68, 352–367. [Google Scholar] [CrossRef]
- Emsley, P.; Lohkamp, B.; Scott, W.G.; Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 2010, 66, 486–501. [Google Scholar] [CrossRef]
- Davis, I.W.; Murray, L.W.; Richardson, J.S.; Richardson, D.C. MOLPROBITY: Structure validation and all-atom contact analysis for nucleic acids and their complexes. Nucleic Acids Res. 2004, 32, W615–W619. [Google Scholar] [CrossRef] [PubMed]
- DeLano, W.L. Pymol: An open-source molecular graphics tool. CCP4 Newsl. Protein Crystallogr. 2002, 40, 82–92. [Google Scholar]
- Unni, S.; Huang, Y.; Hanson, R.M.; Tobias, M.; Krishnan, S.; Li, W.W.; Nielsen, J.E.; Baker, N.A. Web servers and services for electrostatics calculations with APBS and PDB2PQR. J. Comput. Chem. 2011, 32, 1488–1491. [Google Scholar] [CrossRef] [PubMed]
- Aukrust, I.; Evensen, L.; Hollås, H.; Berven, F.; Atkinson, R.A.; Travé, G.; Flatmark, T.; Vedeler, A. Engineering, biophysical characterisation and binding properties of a soluble mutant form of annexin A2 domain IV that adopts a partially folded conformation. J. Mol. Biol. 2006, 363, 469–481. [Google Scholar] [CrossRef]
- Ecsédi, P.; Kiss, B.; Gógl, G.; Radnai, L.; Buday, L.; Koprivanacz, K.; Liliom, K.; Leveles, I.; Vértessy, B.; Jeszenői, N.; et al. Regulation of the Equilibrium between Closed and Open Conformations of Annexin A2 by N-Terminal Phosphorylation and S100A4-Binding. Structure 2017, 25, 1195–1207.e5. [Google Scholar] [CrossRef]
- Bances, P.; Fernandez, M.R.; Rodriguez-Garcia, M.I.; Morgan, R.O.; Fernandez, M.P. Annexin A11 (ANXA11) gene structure as the progenitor of paralogous annexins and source of orthologous cDNA isoforms. Genomics 2000, 69, 95–103. [Google Scholar] [CrossRef]
- Iglesias, J.M.; Morgan, R.O.; Jenkins, N.A.; Copeland, N.G.; Gilbert, D.J.; Fernandez, M.P. Comparative genetics and evolution of annexin A13 as the founder gene of vertebrate annexins. Mol. Biol. Evol. 2002, 19, 608–618. [Google Scholar] [CrossRef]
- Fernandez, M.P.; Jenkins, N.A.; Gilbert, D.J.; Copeland, N.G.; Morgan, R.O. Sequence and Chromosomal Localization of Mouse Annexin XI. Genomics 1996, 37, 366–374. [Google Scholar] [CrossRef]
- Hornbeck, P.V.; Chabra, I.; Kornhauser, J.M.; Skrzypek, E.; Zhang, B. PhosphoSite: A bioinformatics resource dedicated to physiological protein phosphorylation. Proteomics 2004, 4, 1551–1561. [Google Scholar] [CrossRef]
- Kaetzel, M.A.; Mo, Y.D.; Mealy, T.R.; Campos, B.; Bergsma-Schutter, W.; Brisson, A.; Dedman, J.R.; Seaton, B.A. Phosphorylation mutants elucidate the mechanism of annexin IV-mediated membrane aggregation. Biochemistry 2001, 40, 4192–4199. [Google Scholar] [CrossRef]
- Freye-Minks, C.; Kretsinger, R.H.; Creutz, C.E. Structural and dynamic changes in human annexin VI induced by a phosphorylation-mimicking mutation, T356D. Biochemistry 2003, 42, 620–630. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Mao, Y.; Yang, J.; Zhang, T.; Zhao, L.; Yu, K.; Zheng, M.; Jiang, H.; Yang, H. Characterizing the binding of annexin V to a lipid bilayer using molecular dynamics simulations. Proteins 2014, 82, 312–322. [Google Scholar] [CrossRef] [PubMed]
- Hong, S.; Na, S.; Kim, O.H.; Jeong, S.; Oh, B.C.; Ha, N.C. High-resolution structures of annexin A5 in a two-dimensional array. J. Struct. Biol. 2020, 209, 107401. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.C.; Chipot, C.; Scheuring, S. Annexin-V stabilizes membrane defects by inducing lipid phase transition. Nat. Commun. 2020, 11, 230. [Google Scholar] [CrossRef]
- Mo, Y.; Campos, B.; Mealy, T.R.; Commodore, L.; Head, J.F.; Dedman, J.R.; Seaton, B.A. Interfacial basic cluster in annexin V couples phospholipid binding and trimer formation on membrane surfaces. J. Biol. Chem. 2003, 278, 2437–2443. [Google Scholar] [CrossRef]
- Concha, N.O.; Head, J.F.; Kaetzel, M.A.; Dedman, J.R.; Seaton, B.A. Annexin V forms calcium-dependent trimeric units on phospholipid vesicles. FEBS Lett. 1992, 314, 159–162. [Google Scholar] [CrossRef]
- Crosby, K.C.; Postma, M.; Hink, M.A.; Zeelenberg, C.H.; Adjobo-Hermans, M.J.; Gadella, T.W. Quantitative analysis of self-association and mobility of annexin A4 at the plasma membrane. Biophys. J. 2013, 104, 1875–1885. [Google Scholar] [CrossRef]
- Ravanat, C.; Torbet, J.; Freyssinet, J.M. A neutron solution scattering study of the structure of annexin-V and its binding to lipid vesicles. J. Mol. Biol. 1992, 226, 1271–1278. [Google Scholar] [CrossRef]
- Aukrust, I.; Hollås, H.; Strand, E.; Evensen, L.; Travé, G.; Flatmark, T.; Vedeler, A. The mRNA-binding site of annexin A2 resides in helices C-D of its domain IV. J. Mol. Biol. 2007, 368, 1367–1378. [Google Scholar] [CrossRef]
- Solbak, S.M.Ø.; Abdurakhmanov, E.; Vedeler, A.; Danielson, U.H. Characterization of interactions between hepatitis C virus NS5B polymerase, annexin A2 and RNA—Effects on NS5B catalysis and allosteric inhibition. Virol. J. 2017, 14, 236. [Google Scholar] [CrossRef]
- Pandurangan, A.P.; Ochoa-Montaño, B.; Ascher, D.B.; Blundell, T.L. SDM: A server for predicting effects of mutations on protein stability. Nucleic Acids Res. 2017, 45, W229–W235. [Google Scholar] [CrossRef] [PubMed]
- Gomes, E.; Shorter, J. The molecular language of membraneless organelles. J. Biol. Chem. 2019, 294, 7115–7127. [Google Scholar] [CrossRef]
- Vernon, R.M.; Chong, P.A.; Tsang, B.; Kim, T.H.; Bah, A.; Farber, P.; Lin, H.; Forman-Kay, J.D. Pi-Pi contacts are an overlooked protein feature relevant to phase separation. Elife 2018, 7. [Google Scholar] [CrossRef] [PubMed]
- Misaki, Y.; Van Venrooij, W.J.; Pruijn, G.J. Prevalence and characteristics of anti-56K/annexin XI autoantibodies in systemic autoimmune diseases. J. Rheumatol. 1995, 22, 97–102. [Google Scholar] [PubMed]
- Sjolin, C.; Movitz, C.; Lundqvist, H.; Dahlgren, C. Translocation of annexin XI to neutrophil subcellular organelles. Biochim. Biophys. Acta 1997, 1326, 149–156. [Google Scholar] [CrossRef]
- Li, H.R.; Chiang, W.C.; Chou, P.C.; Wang, W.J.; Huang, J.R. TAR DNA-binding protein 43 (TDP-43) liquid-liquid phase separation is mediated by just a few aromatic residues. J. Biol. Chem. 2018, 293, 6090–6098. [Google Scholar] [CrossRef]
- Nguyen, H.P.; Van Broeckhoven, C.; van der Zee, J. ALS Genes in the Genomic Era and their Implications for FTD. Trends Genet. 2018, 34, 404–423. [Google Scholar] [CrossRef]
- Wei, Q.; Chen, X.; Chen, Y.; Ou, R.; Cao, B.; Hou, Y.; Zhang, L.; Shang, H.F. Unique characteristics of the genetics epidemiology of amyotrophic lateral sclerosis in China. Sci. China Life Sci. 2019, 62, 517–525. [Google Scholar] [CrossRef]
| Data Collection Parameters | ||
| Instrument | P12, PETRAIII, DESY | |
| Wavelength (nm) | 0.124 | |
| Angular range (nm−1) | 0.022–7.33 | |
| Exposure time (s) | 0.045 | |
| Exposure temperature (°C) | 10 | |
| Protein | AnxA11Δ188 | AnxA11Δ192 |
| Concentration range (mg/mL) | 0.77–2.96 | 0.41–1.18 |
| Structural Parameters | ||
| I0 (relative) [from Guinier] | 0.02759 | 0.0274 |
| Rg (nm) [from Guinier] | 2.44 ± 0.13 | 2.43 ± 0.20 |
| I0 (relative) [from P(r)] | 0.02692 | 0.02748 |
| Rg (nm) [from P(r)] | 2.500 ± 0.016 | 2.503 ± 0.019 |
| Dmax (nm) [from GNOM] | 10.00 | 9.97 |
| VPorod (nm3) [from GNOM] | 58.03 | 59.53 |
| Molecular Mass Determination | ||
| Molecular mass Mr (kDa) [from I0 using Guinier] 1 | 34.3 | 34.1 |
| Molecular mass Mr (kDa) [from I0 using P(r)] 1 | 33.5 | 34.2 |
| Molecular mass Mr (kDa) [from VPorod] | 34.1 | 35.0 |
| Molecular mass Mr (kDa) [from absolute scale] | 34.8 | 34.8 |
| Theoretical Mr from sequence (kDa) | 36.8 | 36.4 |
| Software | ||
| Primary data reduction & processing | PRIMUS | |
| Ab initio modelling | GASBOR | |
| χ2 | 0.770 | 0.629 |
| Fitting of theoretical scattering curve to data | CRYSOL | |
| χ2 | 0.888 | 0.749 |
| Data Collection | |
| Construct | ∆188AnxA11 |
| Wavelength (Å) | 0.9763 |
| Space group | P1 |
| Unit cell parameters | a = 39.1 Å, b = 86.7 Å, c = 87.3 Å ⍺ = 114.0°, β = 102.0°, γ = 97.2° |
| Resolution range (Å) | 50–2.30 (2.36–2.30) 1 |
| Completeness (%) | 95.4 (95.0) |
| Reflections total/unique | 141110/42245 (9818/3046) |
| ⟨I/σI⟩ | 5.2 (1.1) |
| Rsym (%) | 17.5 (164.0) |
| Rmeas (%) | 20.9 (196.6) |
| CC1/2 (%) | 99.2 (47.6) |
| redundancy | 3.3 (3.2) |
| Data Processing | |
| Rcryst/Rfree (%) | 22.8 (26.6) |
| Rmsd bond length (Å) | 0.002 |
| Rmsd bond angle (°) | 0.4 |
| MolProbity score (percentile) | 1.58 (98th) |
| Ramachandran favoured/disallowed (%) | 97.2/0.3 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lillebostad, P.A.G.; Raasakka, A.; Hjellbrekke, S.J.; Patil, S.; Røstbø, T.; Hollås, H.; Sakya, S.A.; Szigetvari, P.D.; Vedeler, A.; Kursula, P. Structure of the ALS Mutation Target Annexin A11 Reveals a Stabilising N-Terminal Segment. Biomolecules 2020, 10, 660. https://doi.org/10.3390/biom10040660
Lillebostad PAG, Raasakka A, Hjellbrekke SJ, Patil S, Røstbø T, Hollås H, Sakya SA, Szigetvari PD, Vedeler A, Kursula P. Structure of the ALS Mutation Target Annexin A11 Reveals a Stabilising N-Terminal Segment. Biomolecules. 2020; 10(4):660. https://doi.org/10.3390/biom10040660
Chicago/Turabian StyleLillebostad, Peder A. G., Arne Raasakka, Silje J. Hjellbrekke, Sudarshan Patil, Trude Røstbø, Hanne Hollås, Siri A. Sakya, Peter D. Szigetvari, Anni Vedeler, and Petri Kursula. 2020. "Structure of the ALS Mutation Target Annexin A11 Reveals a Stabilising N-Terminal Segment" Biomolecules 10, no. 4: 660. https://doi.org/10.3390/biom10040660
APA StyleLillebostad, P. A. G., Raasakka, A., Hjellbrekke, S. J., Patil, S., Røstbø, T., Hollås, H., Sakya, S. A., Szigetvari, P. D., Vedeler, A., & Kursula, P. (2020). Structure of the ALS Mutation Target Annexin A11 Reveals a Stabilising N-Terminal Segment. Biomolecules, 10(4), 660. https://doi.org/10.3390/biom10040660






