Characterization of Lipid Profiles after Dietary Intake of Polyunsaturated Fatty Acids Using Integrated Untargeted and Targeted Lipidomics
Abstract
1. Introduction
2. Materials and Methods
2.1. Standard Chemicals
2.2. Animals
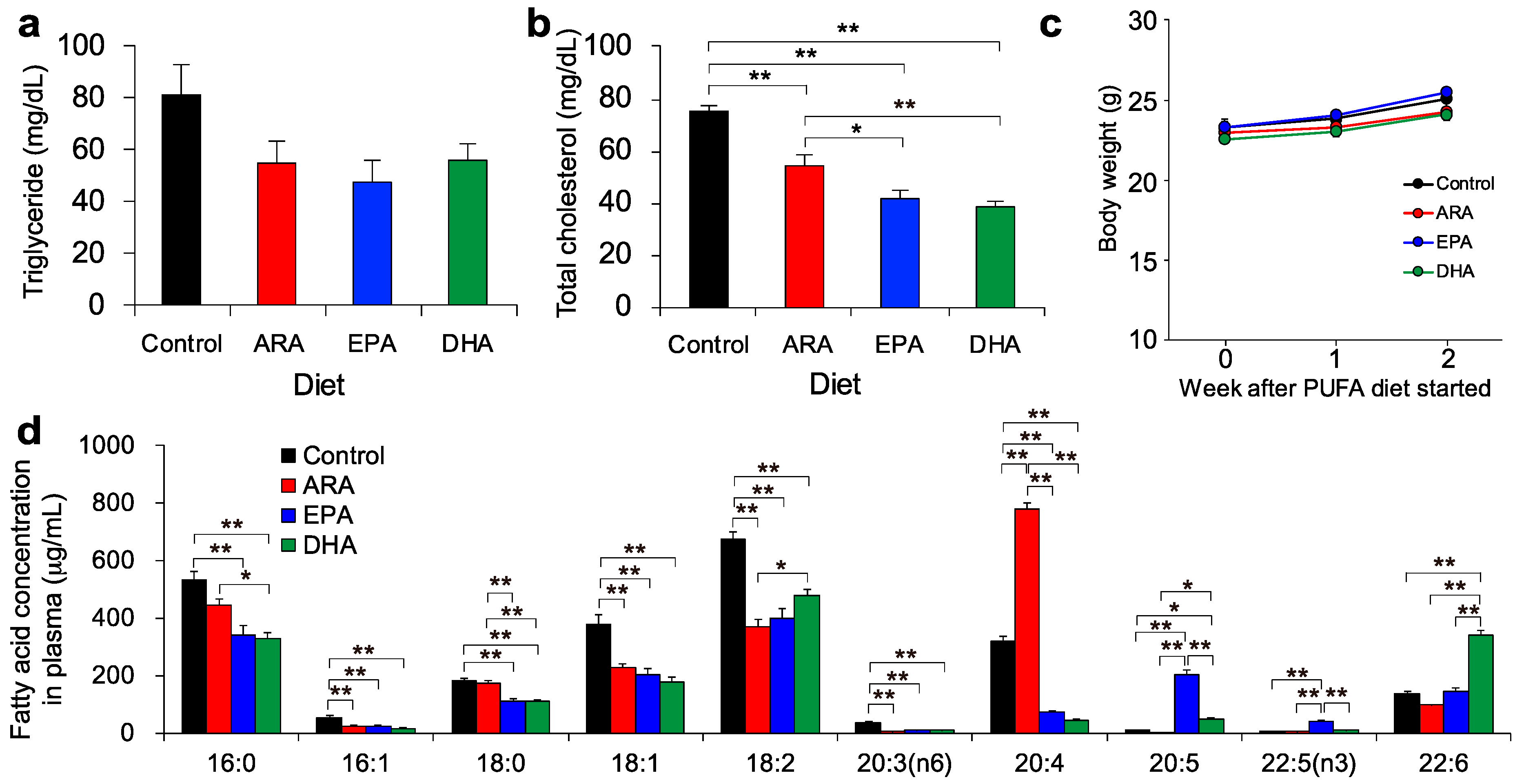
2.3. Determination of Plasma Triglyceride, Cholesterol, and Fatty Acid Composition
2.4. Untargeted LC-MS/MS-Based Lipidomics
2.5. Targeted LC-MS/MS-Based Lipidomics
2.6. Data Analysis
3. Results
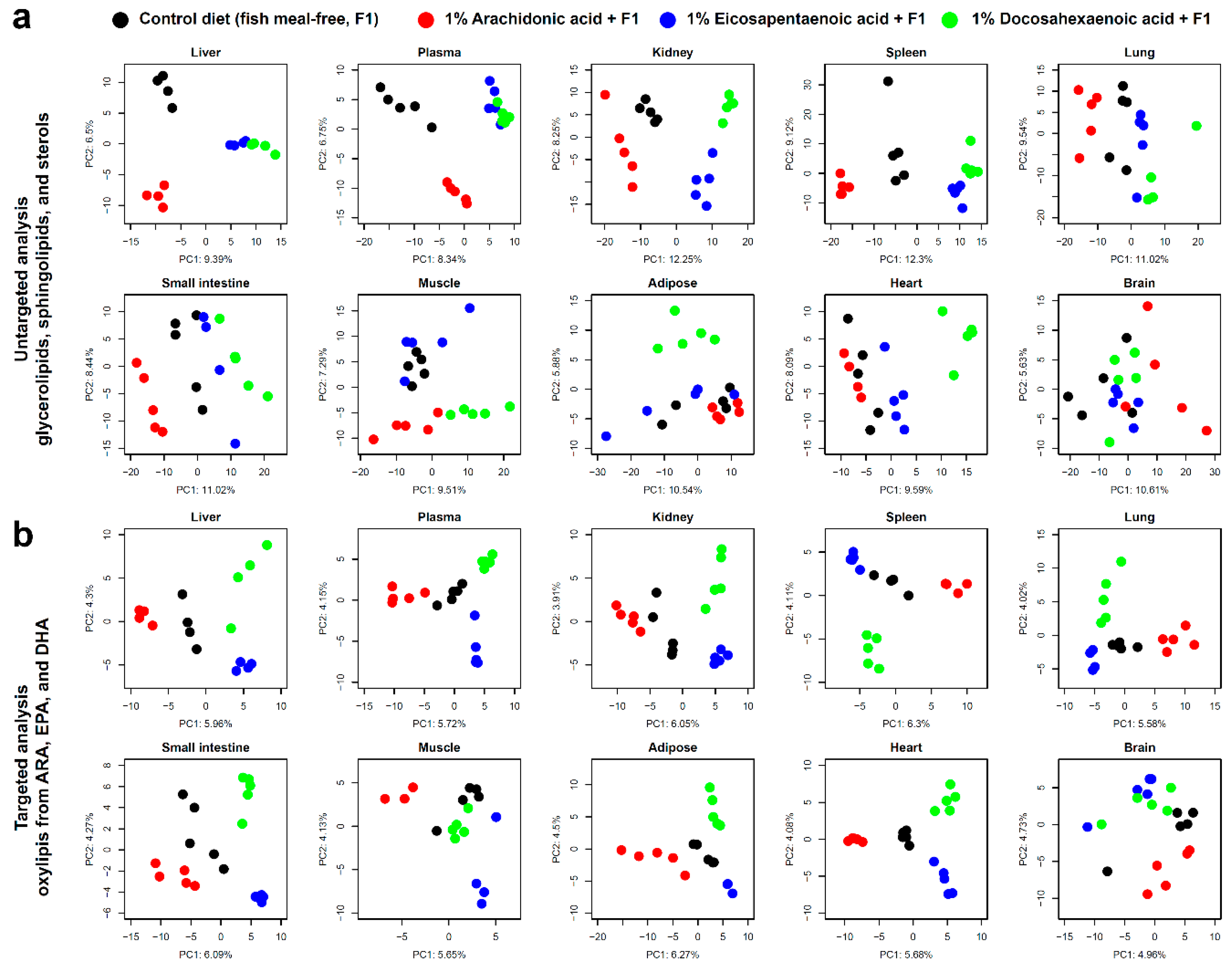
3.1. Lipid Profiles after Dietary Intake of ARA, EPA, or DHA
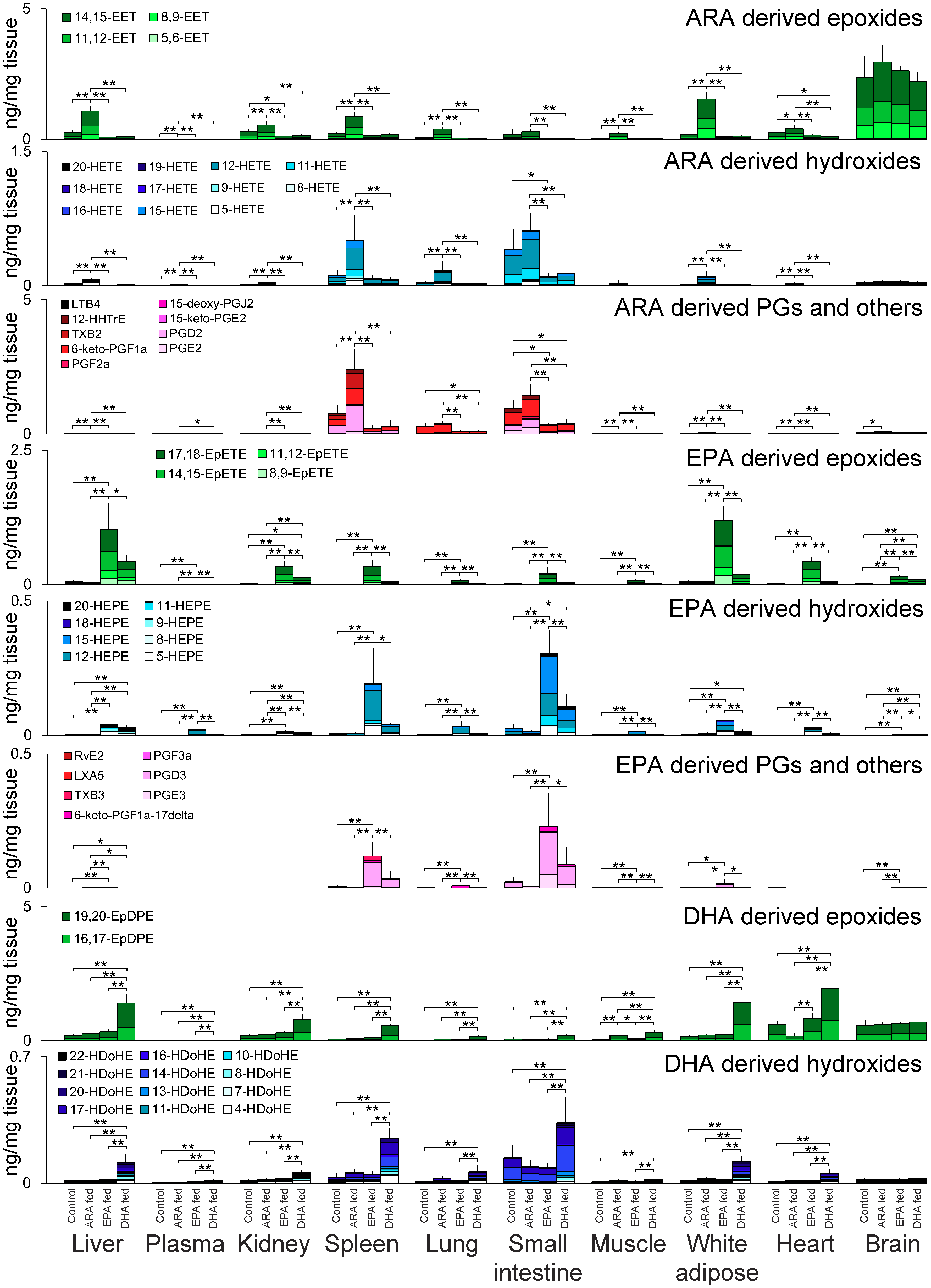
3.2. Oxylipin Profiles in Targeted Lipidomics
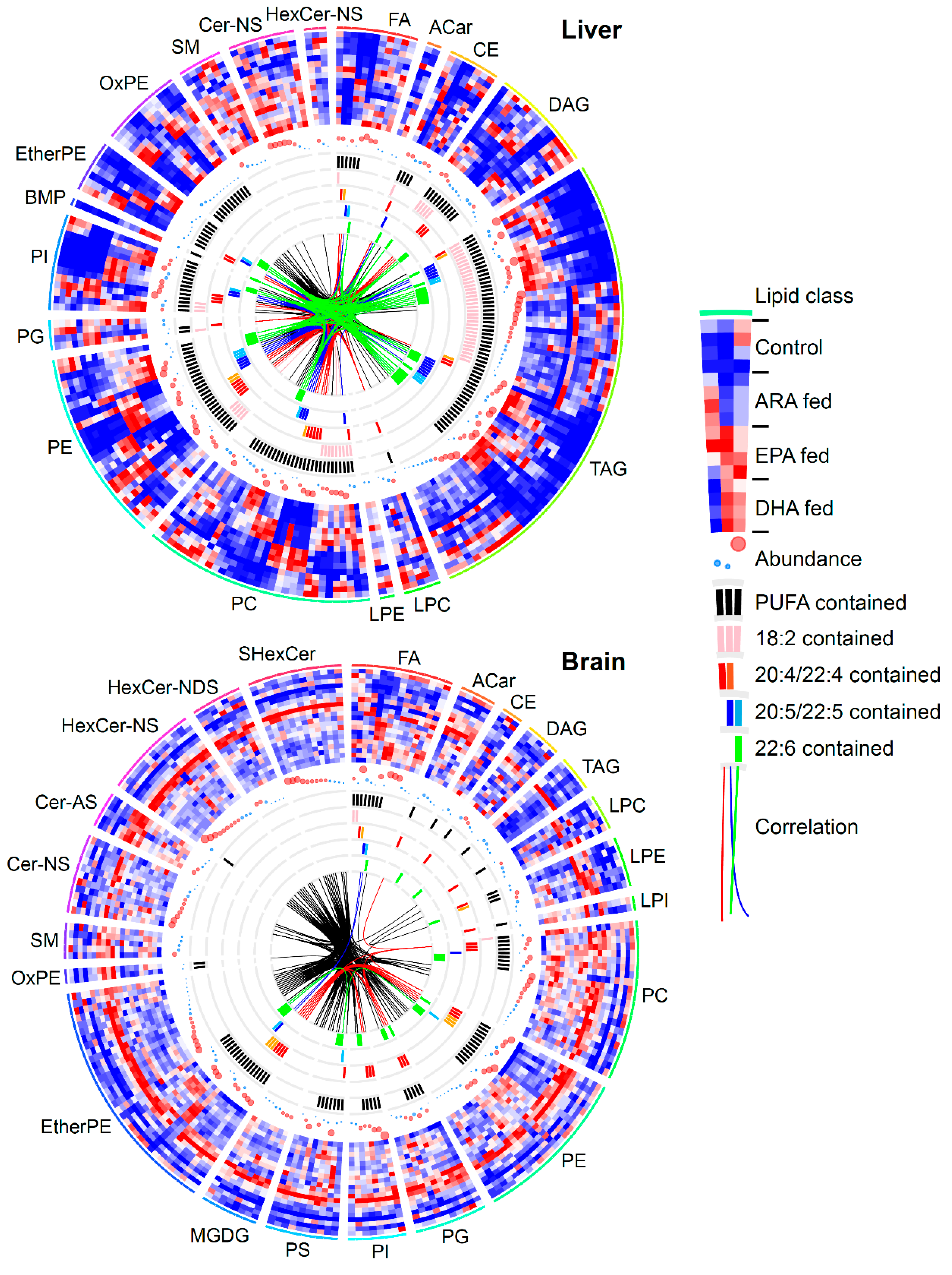
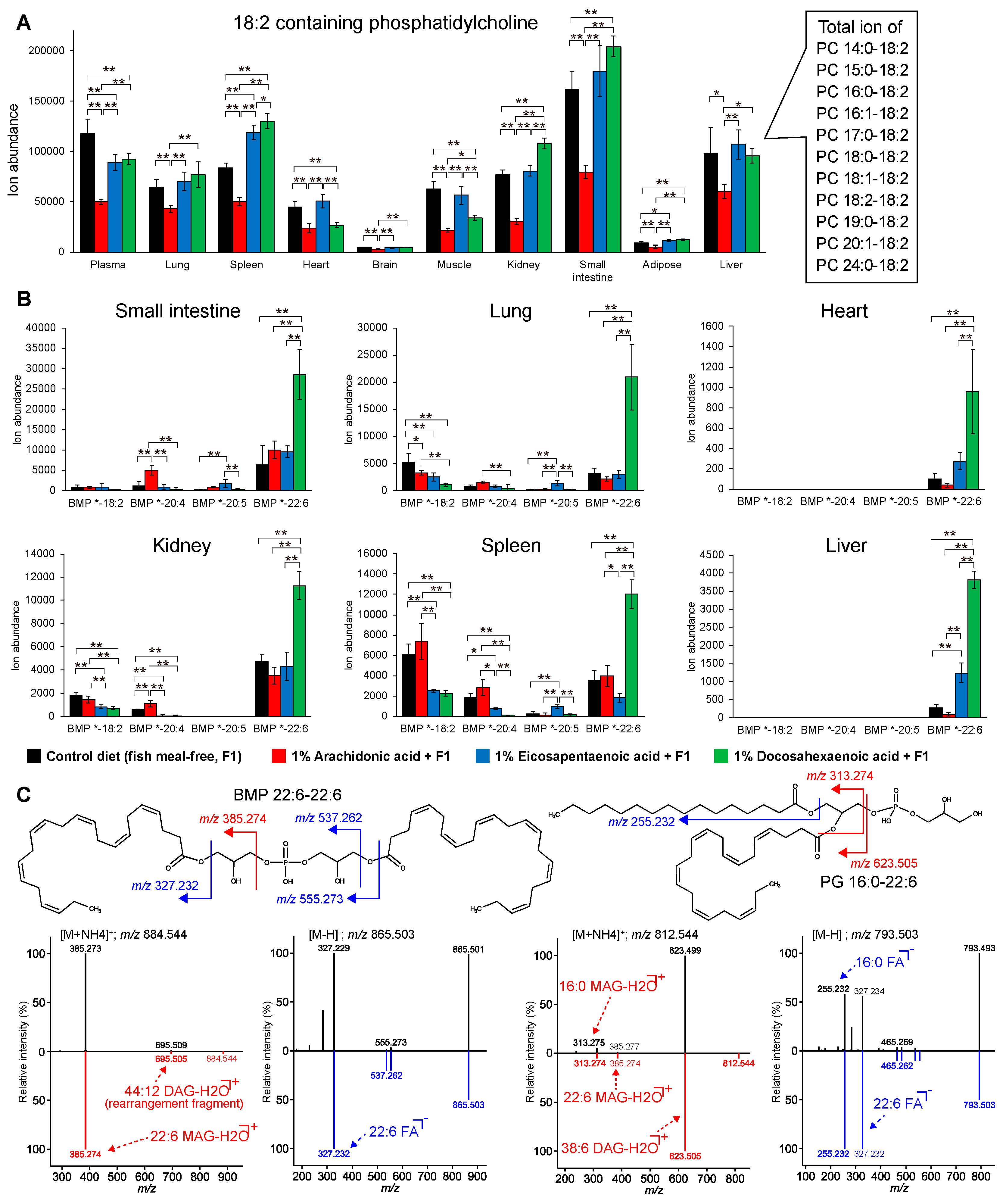
3.3. Lipidomic Signatures in Untargeted Analyses
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Saini, R.K.; Keum, Y.S. Omega-3 and omega-6 polyunsaturated fatty acids: Dietary sources, metabolism, and significance—A review. Life Sci. 2018, 203, 255–267. [Google Scholar] [CrossRef] [PubMed]
- Zárate, R.; el Jaber-Vazdekis, N.; Tejera, N.; Pérez, J.A.; Rodríguez, C. Significance of long chain polyunsaturated fatty acids in human health. Clin. Transl. Med. 2017, 6, 25. [Google Scholar] [CrossRef] [PubMed]
- Wathes, D.C.; Abayasekara, D.R.; Aitken, R.J. Polyunsaturated fatty acids in male and female reproduction. Biol. Reprod. 2007, 77, 190–201. [Google Scholar] [CrossRef] [PubMed]
- Ostermann, A.I.; Schebb, N.H. Effects of omega-3 fatty acid supplementation on the pattern of oxylipins: A short review about the modulation of hydroxy-, dihydroxy-, and epoxy-fatty acids. Food Funct. 2017, 8, 2355–2367. [Google Scholar] [CrossRef] [PubMed]
- Ostermann, A.I.; Waindok, P.; Schmidt, M.J.; Chiu, C.Y.; Smyl, C.; Rohwer, N.; Weylandt, K.H.; Schebb, N.H. Modulation of the endogenous omega-3 fatty acid and oxylipin profile in vivo—A comparison of the fat-1 transgenic mouse with C57BL/6 wildtype mice on an omega-3 fatty acid enriched diet. PLoS ONE 2017, 12, e0184470. [Google Scholar] [CrossRef] [PubMed]
- Wijendran, V.; Downs, I.; Srigley, C.T.; Kothapalli, K.S.; Park, W.J.; Blank, B.S.; Zimmer, J.P.; Butt, C.M.; Salem, N.; Brenna, T. Dietary arachidonic acid and docosahexaenoic acid regulate liver fatty acid desaturase (FADS) alternative transcript expression in suckling piglets. Prostaglandins Leukot. Essent. Fat. Acids 2013, 89, 345–350. [Google Scholar] [CrossRef] [PubMed]
- Arterburn, L.M.; Hall, E.B.; Oken, H. Distribution, interconversion, and dose response of n-3 fatty acids in humans. Am. J. Clin. Nutr. 2006, 83, 1467S–1476S. [Google Scholar] [CrossRef]
- Balvers, M.G.J.; Verhoeckx, K.C.M.; Bijlsma, S.; Rubingh, C.M.; Meijerink, J.; Wortelboer, H.M.; Witkamp, R.F. Fish oil and inflammatory status alter the n-3 to n-6 balance of the endocannabinoid and oxylipin metabolomes in mouse plasma and tissues. Metabolomics 2012, 8, 1130–1147. [Google Scholar] [CrossRef]
- Wood, J.T.; Williams, J.S.; Pandarinathan, L.; Janero, D.R.; Lammi-Keefe, C.J.; Makriyannis, A. Dietary docosahexaenoic acid supplementation alters select physiological endocannabinoid-system metabolites in brain and plasma. J. Lipid Res. 2010, 51, 1416–1423. [Google Scholar] [CrossRef]
- Harayama, T.; Riezman, H. Understanding the diversity of membrane lipid composition. Nat. Rev. Mol. Cell Biol. 2018, 19, 281–296. [Google Scholar] [CrossRef]
- Olzmann, J.A.; Carvalho, P. Dynamics and functions of lipid droplets. Nat. Rev. Mol. Cell Biol. 2019, 20, 137–155. [Google Scholar] [CrossRef] [PubMed]
- Arita, M. Mediator lipidomics in acute inflammation and resolution. J. Biochem. 2012, 152, 313–319. [Google Scholar] [CrossRef] [PubMed]
- Tsugawa, H.; Ikeda, K.; Arita, M. The importance of bioinformatics for connecting data-driven lipidomics and biological insights. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2017, 1862, 762–765. [Google Scholar] [CrossRef] [PubMed]
- Burla, B.; Arita, M.; Arita, M.; Bendt, A.K.; Cazenave-Gassiot, A.; Dennis, E.A.; Ekroos, K.; Han, X.; Ikeda, K.; Liebisch, G.; et al. MS-based lipidomics of human blood plasma: A community-initiated position paper to develop accepted guidelines. J. Lipid Res. 2018, 59, 2001–2017. [Google Scholar] [CrossRef] [PubMed]
- Tsugawa, H.; Satoh, A.; Uchino, H.; Cajka, T.; Arita, M.; Arita, M. Mass Spectrometry Data Repository Enhances Novel Metabolite Discoveries with Advances in Computational Metabolomics. Metabolites 2019, 9, 119. [Google Scholar] [CrossRef] [PubMed]
- Tsugawa, H.; Ikeda, K.; Tanaka, W.; Senoo, Y.; Arita, M.; Arita, M. Comprehensive identification of sphingolipid species by in silico retention time and tandem mass spectral library. J. Cheminformatics 2017, 9, 1–12. [Google Scholar] [CrossRef]
- Tsugawa, H.; Cajka, T.; Kind, T.; Ma, Y.; Higgins, B.; Ikeda, K.; Kanazawa, M.; VanderGheynst, J.; Fiehn, O.; Arita, M. MS-DIAL: Data-independent MS/MS deconvolution for comprehensive metabolome analysis. Nat. Methods 2015, 12, 523–526. [Google Scholar] [CrossRef]
- Tsugawa, H.; Nakabayashi, R.; Mori, T.; Yamada, Y.; Takahashi, M.; Rai, A.; Sugiyama, R.; Yamamoto, H.; Nakaya, T.; Yamazaki, M.; et al. A cheminformatics approach to characterize metabolomes in stable-isotope-labeled organisms. Nat. Methods 2019, 16, 295–298. [Google Scholar] [CrossRef]
- Hansen, J.B.; Olsen, J.O.; Wilsgard, L.; Lyngmo, V.; Svensson, B. Comparative effects of prolonged intake of highly purified fish oils as ethyl ester or triglyceride on lipids, haemostasis and platelet function in normolipaemic men. Eur. J. Clin. Nutr. 1993, 47, 497–507. [Google Scholar]
- Neubronner, J.; Schuchardt, J.P.; Kressel, G.; Merkel, M.; von Schacky, C.; Hahn, A. Enhanced increase of omega-3 index in response to long-term n-3 fatty acid supplementation from triacylglycerides versus ethyl esters. Eur. J. Clin. Nutr. 2011, 65, 247–254. [Google Scholar] [CrossRef]
- Schuchardt, J.P.; Hahn, A. Bioavailability of long-chain omega-3 fatty acids. Prostaglandins Leukot. Essent. Fat. Acids 2013, 89, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Hishikawa, D.; Valentine, W.J.; Iizuka-Hishikawa, Y.; Shindou, H.; Shimizu, T. Metabolism and functions of docosahexaenoic acid-containing membrane glycerophospholipids. FEBS Lett. 2017, 591, 2730–2744. [Google Scholar] [CrossRef] [PubMed]
- Spector, A.A.; Norris, A.W. Action of epoxyeicosatrienoic acids on cellular function. Am. J. Physiol. Physiol. 2007, 292, C996–C1012. [Google Scholar] [CrossRef] [PubMed]
- Nagatake, T.; Shiogama, Y.; Inoue, A.; Kikuta, J.; Honda, T.; Tiwari, P.; Kishi, T.; Yanagisawa, A.; Isobe, Y.; Matsumoto, N.; et al. The 17,18-epoxyeicosatetraenoic acid–G protein–coupled receptor 40 axis ameliorates contact hypersensitivity by inhibiting neutrophil mobility in mice and cynomolgus macaques. J. Allergy Clin. Immunol. 2018, 142, 470–484. [Google Scholar] [CrossRef]
- Kunisawa, J.; Arita, M.; Hayasaka, T.; Harada, T.; Iwamoto, R.; Nagasawa, R.; Shikata, S.; Nagatake, T.; Suzuki, H.; Hashimoto, E.; et al. Dietary ω3 fatty acid exerts anti-allergic effect through the conversion to 17,18-epoxyeicosatetraenoic acid in the gut. Sci. Rep. 2015, 5, 9750. [Google Scholar] [CrossRef]
- Mochimaru, T.; Fukunaga, K.; Miyata, J.; Matsusaka, M.; Kabata, H.; Ueda, S.; Suzuki, Y.; Goto, T.; Urabe, D.; Inoue, M.; et al. 12-OH-17,18-Epoxyeicosatetraenoic acid alleviates eosinophilic airway inflammation in murine lungs. Allergy 2018, 73, 369–378. [Google Scholar] [CrossRef]
- Kubota, T.; Arita, M.; Isobe, Y.; Iwamoto, R.; Goto, T.; Yoshioka, T.; Urabe, D.; Inoue, M.; Arai, H. Eicosapentaenoic acid is converted via ω-3 epoxygenation to the anti-inflammatory metabolite 12-hydroxy-17,18-epoxyeicosatetraenoic acid. FASEB J. 2014, 28, 586–593. [Google Scholar] [CrossRef]
- Dasilva, G.; Pazos, M.; García-Egido, E.; Pérez-Jiménez, J.; Torres, J.L.; Giralt, M.; Nogués, M.R.; Medina, I. Lipidomics to analyze the influence of diets with different EPA:DHA ratios in the progression of Metabolic Syndrome using SHROB rats as a model. Food Chem. 2016, 205, 196–203. [Google Scholar] [CrossRef]
- Grønn, M.; Christensen, E.; Hagve, T.A.; Christophersen, B.O. Peroxisomal retroconversion of docosahexaenoic acid (22:6(n-3)) to eicosapentaenoic acid (20:5(n-3)) studied in isolated rat liver cells. Biochim. Biophys. Acta (BBA) Lipids Lipid Metab. 1991, 1081, 85–91. [Google Scholar] [CrossRef]
- Zhang, J.Y.; Kothapalli, K.S.D.; Brenna, J.T. Desaturase and elongase limiting endogenous long chain polyunsaturated fatty acid biosynthesis. Curr. Opin. Clin. Nutr. Metab. Care 2016, 19, 103–110. [Google Scholar] [CrossRef]
- Schmidt-Schultz, T.; Althaus, H.H. Monogalactosyl Diglyceride, a Marker for Myelination, Activates Oligodendroglial Protein Kinase C. J. Neurochem. 1994, 62, 1578–1585. [Google Scholar] [CrossRef] [PubMed]
- Murray, R.K.; Narasimhan, R.; Levine, M.; Pinteric, L.; Shirley, M.; Lingwood, C.; Schacter, H. Galactoglycerolipids of Mammalian Testis, Spermatozoa, and Nervous Tissue; ACS Symposium Series; American Chemical Society: Washington, DC, USA, 1980; pp. 105–125. [Google Scholar]
- Påhlsson, P.; Spitalnik, S.L.; Spitalnik, P.F.; Fantini, J.; Rakotonirainy, O.; Ghardashkhani, S.; Lindberg, J.; Konradsson, P.; Larson, G. Characterization of galactosyl glycerolipids in the HT29 human colon carcinoma cell line. Arch. Biochem. Biophys. 2001, 396, 187–198. [Google Scholar] [CrossRef] [PubMed]
- Jain, M.; Ngoy, S.; Sheth, S.A.; Swanson, R.A.; Rhee, E.P.; Liao, R.; Clish, C.B.; Mootha, V.K.; Nilsson, R. A systematic survey of lipids across mouse tissues. Am. J. Physiol. Metab. 2014, 306, E854–E868. [Google Scholar] [CrossRef] [PubMed]
- Yen, C.L.E.; Stone, S.J.; Koliwad, S.; Harris, C.; Farese, R.V. Thematic Review Series: Glycerolipids. DGAT enzymes and triacylglycerol biosynthesis. J. Lipid Res. 2008, 49, 2283–2301. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.; Jin, X.; Tsueng, G.; Afrasiabi, C.; Su, A.I. BioGPS: Building your own mash-up of gene annotations and expression profiles. Nucleic Acids Res. 2016, 44, D313–D316. [Google Scholar] [CrossRef] [PubMed]
- Hashidate-Yoshida, T.; Harayama, T.; Hishikawa, D.; Morimoto, R.; Hamano, F.; Tokuoka, S.M.; Eto, M.; Tamura-Nakano, M.; Yanobu-Takanashi, R.; Mukumoto, Y.; et al. Fatty acyl-chain remodeling by LPCAT3 enriches arachidonate in phospholipid membranes and regulates triglyceride transport. Elife 2015, 4, e06328. [Google Scholar] [CrossRef]
- Adam, O.; Wolfram, T.G. Impact of linoleic acid intake on arachidonic acid formation and eicosanoid biosynthesis in humans. Prostaglandins Leukot. Essent. Fat. Acids 2008, 75, 177–181. [Google Scholar] [CrossRef]
- Akgoc, Z.; Sena-Esteves, M.; Martin, D.R.; Han, X.; d’Azzo, A.; Seyfried, T.N. Bis(monoacylglycero)phosphate: A secondary storage lipid in the gangliosidoses. J. Lipid Res. 2015, 56, 1006–1013. [Google Scholar] [CrossRef]
- Cao, J.; Shen, W.; Chang, Z.; Shi, Y. ALCAT1 is a polyglycerophospholipid acyltransferase potently regulated by adenine nucleotide and thyroid status. Am. J. Physiol. Metab. 2008, 296, E647–E653. [Google Scholar] [CrossRef]
- Yamashita, A.; Hayashi, Y.; Nemoto-Sasaki, Y.; Ito, M.; Oka, S.; Tanikawa, T.; Waku, K.; Sugiura, T. Acyltransferases and transacylases that determine the fatty acid composition of glycerolipids and the metabolism of bioactive lipid mediators in mammalian cells and model organisms. Prog. Lipid Res. 2014, 53, 18–81. [Google Scholar] [CrossRef]
- Arnold, C.; Markovic, M.; Blossey, K.; Wallukat, G.; Fischer, R.; Dechend, R.; Konkel, A.; von Schacky, C.; Luft, F.C.; Muller, D.N.; et al. Arachidonic Acid-metabolizing Cytochrome P450 Enzymes Are Targets of ω-3 Fatty Acids. J. Biol. Chem. 2010, 285, 32720–32733. [Google Scholar] [CrossRef] [PubMed]
- Igarashi, M.; Chang, L.; Ma, K.; Rapoport, S.I. Kinetics of eicosapentaenoic acid in brain, heart and liver of conscious rats fed a high n-3 PUFA containing diet. Prostaglandins Leukot. Essent. Fat. Acids 2013, 89, 403–412. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.T.; Bazinet, R.P. β-oxidation and rapid metabolism, but not uptake regulate brain eicosapentaenoic acid levels. Prostaglandins Leukot. Essent. Fat. Acids 2015, 92, 33–40. [Google Scholar] [CrossRef] [PubMed]
- Scherer, M.; Schmitz, G. Metabolism, function and mass spectrometric analysis of bis(monoacylglycero)phosphate and cardiolipin. Chem. Phys. Lipids 2011, 164, 556–562. [Google Scholar] [CrossRef] [PubMed]
- Pribasnig, M.A.; Mrak, I.; Grabner, G.F.; Taschler, U.; Knittelfelder, O.; Scherz, B.; Eichmann, T.O.; Heier, C.; Grumet, L.; Kowaliuk, J.; et al. α/β hydrolase domain-containing 6 (ABHD6) degrades the late Endosomal/Lysosomal lipid Bis(Monoacylglycero)phosphate. J. Biol. Chem. 2015, 290, 29869–29881. [Google Scholar] [CrossRef]
- Matsuo, H.; Chevallier, J.; Mayran, N.; Le Blanc, I.; Ferguson, C.; Fauré, J.; Blanc, N.S.; Matile, S.; Dubochet, J.; Sadoul, R.; et al. Role of LBPA and Alix in Multivesicular Liposome Formation and Endosome Organization. Science 2004, 303, 531–534. [Google Scholar] [CrossRef]
- Kobayashi, T.; Stang, E.; Fang, K.S.; De Moerloose, P.; Parton, R.G.; Gruenberg, J. A lipid associated with the antiphospholipid syndrome regulates endosome structure and function. Nature 1998, 392, 193–197. [Google Scholar] [CrossRef]
- Kolter, T.; Sandhoff, K. Principles of lysosomal membrane digestion: Stimulation of sphingolipid degradation by sphingolipid activator proteins and anionic lysosomal lipids. Annu. Rev. Cell Dev. Biol. 2005, 21, 81–103. [Google Scholar] [CrossRef]
- Bissig, C.; Gruenberg, J. Lipid sorting and multivesicular endosome biogenesis. Cold Spring Harb. Perspect. Biol. 2013, 5, a016816. [Google Scholar] [CrossRef]
- Bouvier, J.; Zemski Berry, K.A.; Hullin-Matsuda, F.; Makino, A.; Michaud, S.; Geloën, A.; Murphy, R.C.; Kobayashi, T.; Lagarde, M.; Delton-Vandenbroucke, I. Selective decrease of bis(monoacylglycero)phosphate content in macrophages by high supplementation with docosahexaenoic acid. J. Lipid Res. 2009, 50, 243–255. [Google Scholar] [CrossRef]
- Liebisch, G.; Ahrends, R.; Arita, M.; Arita, M.; Bowden, J.A.; Ejsing, C.S.; Griffiths, W.J.; Holcapek, M.; Kofeler, H.; Mitchell, T.W.; et al. Lipidomics needs more standardization. Nat. Metab. 2019, 1, 745–747. [Google Scholar]

| Method | Class | Liver | Plasma | Kidney | Spleen | Lung | Small Intestine | Muscle | Adipose | Heart | Brain |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Untargeted | FA | 12 | 16 | 18 | 17 | 15 | 22 | 15 | 14 | 13 | 14 |
| Untargeted | ACar | 2 | 2 | 9 | 7 | 10 | 7 | 14 | 0 | 12 | 6 |
| Untargeted | MAG | 1 | 2 | 2 | 2 | 3 | 4 | 2 | 1 | 0 | 3 |
| Untargeted | DAG | 14 | 8 | 14 | 15 | 18 | 29 | 13 | 28 | 6 | 5 |
| Untargeted | TAG | 70 | 80 | 73 | 88 | 116 | 96 | 80 | 156 | 116 | 6 |
| Untargeted | CE | 7 | 10 | 4 | 6 | 2 | 5 | 3 | 0 | 3 | 1 |
| Untargeted | LPC | 7 | 13 | 8 | 7 | 8 | 7 | 7 | 1 | 7 | 6 |
| Untargeted | LPE | 4 | 6 | 10 | 9 | 8 | 8 | 6 | 0 | 4 | 7 |
| Untargeted | LPG | 0 | 0 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | 0 |
| Untargeted | LPS | 0 | 0 | 1 | 0 | 0 | 1 | 0 | 0 | 0 | 0 |
| Untargeted | LPI | 2 | 0 | 3 | 1 | 0 | 2 | 0 | 0 | 1 | 4 |
| Untargeted | PC | 33 | 30 | 25 | 43 | 45 | 38 | 47 | 6 | 33 | 22 |
| Untargeted | EtherPC | 0 | 1 | 18 | 17 | 9 | 3 | 4 | 0 | 7 | 2 |
| Untargeted | PE | 27 | 9 | 34 | 29 | 32 | 28 | 31 | 4 | 28 | 25 |
| Untargeted | EtherPE | 8 | 3 | 24 | 40 | 35 | 26 | 16 | 2 | 21 | 35 |
| Untargeted | PG | 5 | 1 | 13 | 23 | 22 | 9 | 7 | 1 | 8 | 11 |
| Untargeted | BMP | 1 | 0 | 3 | 6 | 7 | 7 | 0 | 0 | 1 | 0 |
| Untargeted | PS | 2 | 0 | 6 | 20 | 11 | 2 | 1 | 0 | 1 | 10 |
| Untargeted | PI | 15 | 14 | 18 | 25 | 18 | 16 | 15 | 1 | 12 | 10 |
| Untargeted | OxPC | 2 | 0 | 9 | 0 | 0 | 5 | 5 | 0 | 8 | 0 |
| Untargeted | OxPE | 13 | 0 | 29 | 11 | 2 | 16 | 10 | 0 | 18 | 2 |
| Untargeted | OxPG | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Untargeted | OxPI | 2 | 0 | 5 | 0 | 0 | 2 | 1 | 0 | 0 | 0 |
| Untargeted | OxPS | 0 | 0 | 7 | 2 | 2 | 1 | 0 | 0 | 1 | 0 |
| Untargeted | Cer-NS | 10 | 6 | 19 | 16 | 24 | 12 | 12 | 6 | 13 | 12 |
| Untargeted | Cer-NDS | 0 | 0 | 1 | 2 | 1 | 4 | 1 | 0 | 0 | 1 |
| Untargeted | Cer-NP | 0 | 0 | 4 | 0 | 4 | 8 | 0 | 0 | 0 | 1 |
| Untargeted | Cer-AS | 0 | 0 | 2 | 0 | 6 | 0 | 2 | 0 | 0 | 5 |
| Untargeted | Cer-AP | 0 | 0 | 0 | 0 | 1 | 5 | 0 | 0 | 0 | 0 |
| Untargeted | HexCer-NS | 4 | 4 | 8 | 11 | 12 | 5 | 4 | 2 | 2 | 13 |
| Untargeted | HexCer-AP | 0 | 0 | 4 | 0 | 0 | 15 | 0 | 0 | 0 | 0 |
| Untargeted | SHexCer | 0 | 0 | 5 | 0 | 2 | 0 | 2 | 0 | 0 | 14 |
| Untargeted | SM | 6 | 6 | 13 | 13 | 18 | 11 | 6 | 2 | 9 | 5 |
| Targeted | FFA | 17 | 17 | 17 | 16 | 17 | 17 | 17 | 17 | 17 | 16 |
| Targeted | LA-O | 7 | 7 | 5 | 6 | 6 | 7 | 7 | 7 | 7 | 7 |
| Targeted | ALA-O | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 2 | 2 |
| Targeted | GLA-O | 1 | 0 | 0 | 0 | 0 | 1 | 1 | 0 | 0 | 0 |
| Targeted | DGLA-O | 1 | 2 | 1 | 2 | 2 | 4 | 4 | 3 | 1 | 1 |
| Targeted | MA-O | 1 | 2 | 1 | 2 | 0 | 2 | 2 | 1 | 1 | 1 |
| Targeted | ARA-O | 26 | 23 | 28 | 27 | 30 | 29 | 27 | 30 | 28 | 27 |
| Targeted | EPA-O | 16 | 13 | 13 | 15 | 14 | 19 | 13 | 19 | 14 | 13 |
| Targeted | DHA-O | 15 | 17 | 15 | 16 | 16 | 17 | 18 | 16 | 15 | 15 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Naoe, S.; Tsugawa, H.; Takahashi, M.; Ikeda, K.; Arita, M. Characterization of Lipid Profiles after Dietary Intake of Polyunsaturated Fatty Acids Using Integrated Untargeted and Targeted Lipidomics. Metabolites 2019, 9, 241. https://doi.org/10.3390/metabo9100241
Naoe S, Tsugawa H, Takahashi M, Ikeda K, Arita M. Characterization of Lipid Profiles after Dietary Intake of Polyunsaturated Fatty Acids Using Integrated Untargeted and Targeted Lipidomics. Metabolites. 2019; 9(10):241. https://doi.org/10.3390/metabo9100241
Chicago/Turabian StyleNaoe, Satoko, Hiroshi Tsugawa, Mikiko Takahashi, Kazutaka Ikeda, and Makoto Arita. 2019. "Characterization of Lipid Profiles after Dietary Intake of Polyunsaturated Fatty Acids Using Integrated Untargeted and Targeted Lipidomics" Metabolites 9, no. 10: 241. https://doi.org/10.3390/metabo9100241
APA StyleNaoe, S., Tsugawa, H., Takahashi, M., Ikeda, K., & Arita, M. (2019). Characterization of Lipid Profiles after Dietary Intake of Polyunsaturated Fatty Acids Using Integrated Untargeted and Targeted Lipidomics. Metabolites, 9(10), 241. https://doi.org/10.3390/metabo9100241





