Comparative Rice Bran Metabolomics across Diverse Cultivars and Functional Rice Gene–Bran Metabolite Relationships
Abstract
1. Introduction
2. Results
2.1. Classification of Rice Bran Metabolite Profiles and Metabolic Pathways by Non-Targeted Metabolomics
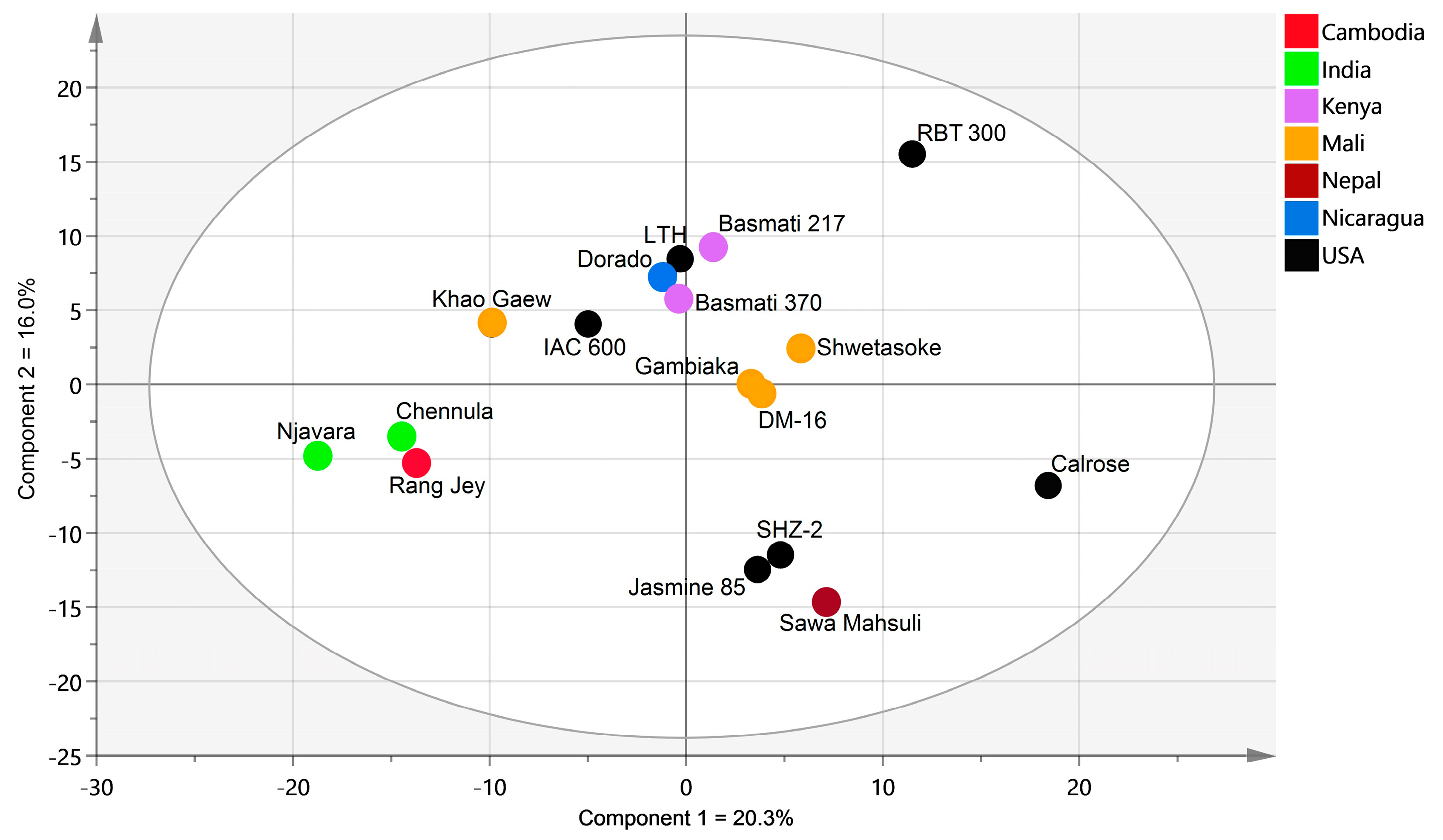
2.2. Global Rice Bran Metabolome Variation for 17 Cultivars
2.3. Comparison of Rice Bran Metabolites across 17 Cultivars
2.3.1. Cultivar Variation in Rice Bran Amino Acids
2.3.2. Cultivar Variation in Rice Bran Carbohydrates
2.3.3. Cultivar Variation in Rice Bran Cofactors and Vitamins
2.3.4. Cultivar Variation in Rice Bran Lipids
2.3.5. Cultivar Variation in Rice Bran Nucleotides
2.3.6. Cultivar Variation in Rice Bran Secondary Metabolites
2.4. Integrating Rice Biosynthetic Genes with Rice Bran Metabolites
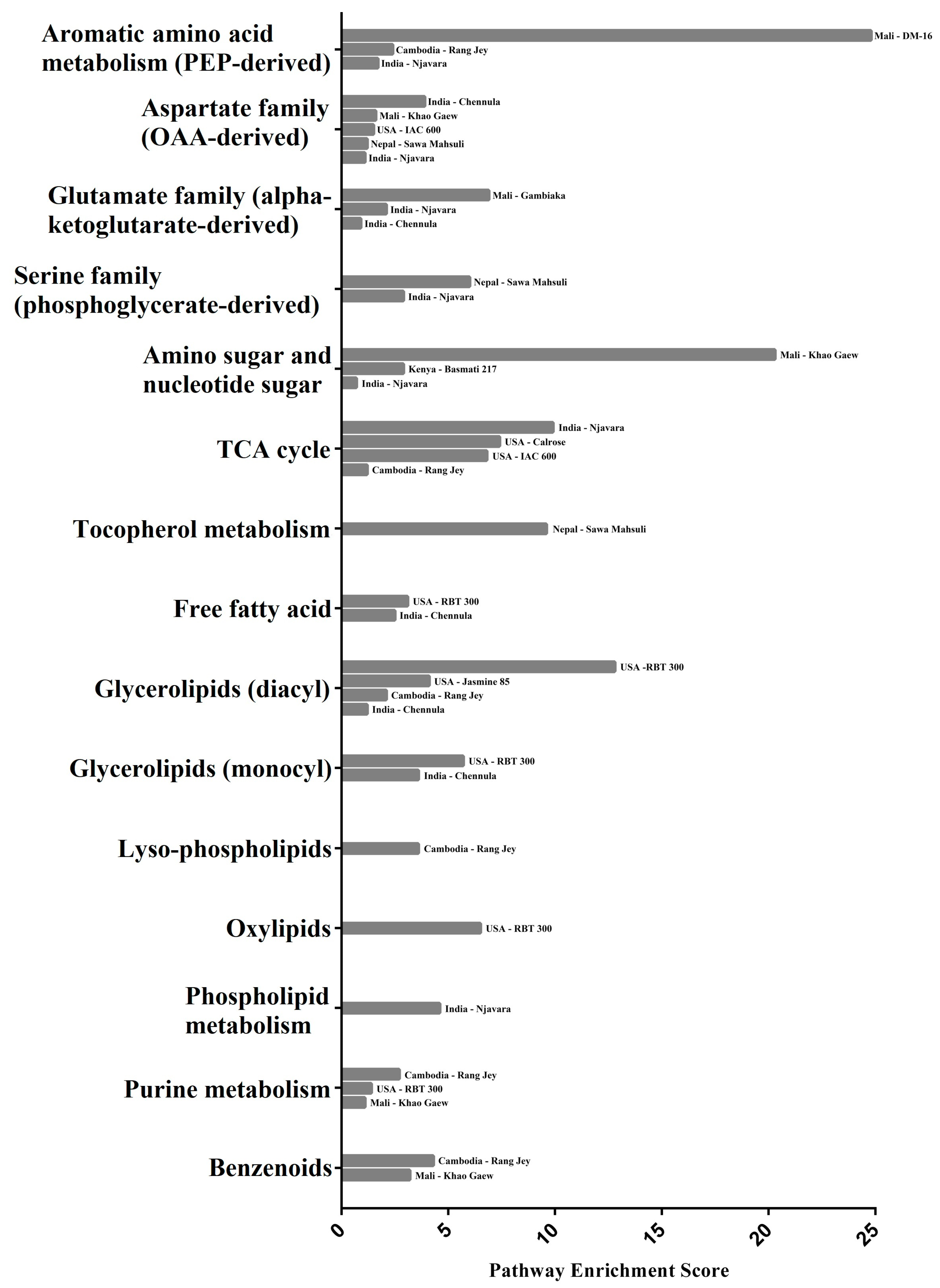
2.5. Integration of Rice Bran Metabolites, Metabolic Pathways, and Rice Genes
2.5.1. Amino Acid Metabolic Pathway Enrichment Scores and Gene Associations
2.5.2. Carbohydrate Metabolic Pathway Enrichment Scores and Gene Associations
2.5.3. Cofactors and Vitamins Metabolic Pathway Enrichment Scores and Gene Associations
2.5.4. Lipids Metabolic Pathway Enrichment Scores and Gene Associations
2.5.5. Nucleotide Metabolic Pathway Enrichment Scores and Gene Associations
2.5.6. Secondary Metabolite Pathway Enrichment Scores and Gene Associations
3. Discussion
4. Materials and Methods
4.1. Rice Cultivars and the Heat Stabilization of Bran
4.2. Rice Bran Metabolite Extraction and Sample Preparation
4.3. UPLC-MS/MS Analysis
4.4. Metabolite Data Extraction and Compound Identification
4.5. Metabolic Pathway Analysis
4.6. Rice Biosynthetic Gene Identification for Selected Rice Bran Metabolites
4.7. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Sarla, N.; Swamy, B.P.M. Oryza glaberrima: A source for the improvement of Oryza sativa. Curr. Sci. 2005, 89, 955–963. [Google Scholar]
- Khush, G.S. What it will take to Feed 5.0 Billion Rice consumers in 2030. Plant Mol. Biol. 2005, 59, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Gani, A.; Wani, S.; Masoodi, F.; Hameed, G. Whole-grain cereal bioactive compounds and their health benefits: A review. J. Food Process. Technol. 2012, 3, 146–156. [Google Scholar] [CrossRef]
- Cosslett, T.L.; Cosslett, P.D. Sustainable Development of Rice and Water Resources in Mainland Southeast Asia and Mekong River Basin; Springer: Singapore, 2018. [Google Scholar]
- Sánchez, B.; Rasmussen, A.; Porter, J.R. Temperatures and the growth and development of maize and rice: A review. Glob. Chang. Biol. 2014, 20, 408–417. [Google Scholar] [CrossRef] [PubMed]
- Kusano, M.; Yang, Z.; Okazaki, Y.; Nakabayashi, R.; Fukushima, A.; Saito, K. Using metabolomic approaches to explore chemical diversity in rice. Mol. Plant 2015, 8, 58–67. [Google Scholar] [CrossRef] [PubMed]
- Food and Agricultur Organization of the United Nation. FAO Rice Market Monitor (RMM). Available online: http://www.fao.org/economic/est/publications/rice-publications/rice-market-monitor-rmm/en/ (accessed on 27 April 2018).
- Van Hoed, V.; Depaemelaere, G.; Ayala, J.V.; Santiwattana, P.; Verhe, R.; De Greyt, W. Influence of chemical refining on the major and minor components of rice brain oil. J. Am. Oil Chem.Soc. 2006, 83, 315–321. [Google Scholar] [CrossRef]
- Henderson, A.J.; Ollila, C.A.; Kumar, A.; Borresen, E.C.; Raina, K.; Agarwal, R.; Ryan, E.P. Chemopreventive properties of dietary rice bran: Current status and future prospects. Adv. Nutr. 2012, 3, 643–653. [Google Scholar] [CrossRef] [PubMed]
- Borresen, E.C.; Ryan, E.P. Rice bran: A food ingredient with global public health opportunities. Wheat Rice Dis. Prev. Health 2014, 2014, 301–310. [Google Scholar]
- Borresen, E.C.; Brown, D.G.; Harbison, G.; Taylor, L.; Fairbanks, A.; O’Malia, J.; Bazan, M.; Rao, S.; Bailey, S.M.; Wdowik, M.; et al. A Randomized Controlled Trial to Increase Navy Bean or Rice Bran Consumption in Colorectal Cancer Survivors. Nutr. Cancer 2016, 68, 1269–1280. [Google Scholar] [CrossRef] [PubMed]
- Alauddina, A.; Islam, J.; Shirakawaa, H.; Koseki, T.; Ardiansyah, A.; Komai, M. Rice Bran as a Functional Food: An Overview of the Conversion of Rice Bran into a Superfood/Functional Food; InTechOpen: London, UK, 2017. [Google Scholar]
- Zarei, I.; Brown, D.G.; Nealon, N.J.; Ryan, E.P. Rice Bran Metabolome Contains Amino Acids, Vitamins & Cofactors, and Phytochemicals with Medicinal and Nutritional Properties. Rice 2017, 10, 24. [Google Scholar] [CrossRef] [PubMed]
- Brown, D.G.; Borresen, E.C.; Brown, R.J.; Ryan, E.P. Heat-stabilised rice bran consumption by colorectal cancer survivors modulates stool metabolite profiles and metabolic networks: A randomised controlled trial. Br. J. Nutr. 2017, 117, 1244–1256. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; Godber, J.S. Purification and Identification of Components of γ-Oryzanol in Rice Bran Oil. J. Agric. Food Chem. 1999, 47, 2724–2728. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; Hua, N.; Godber, J.S. Antioxidant Activity of Tocopherols, Tocotrienols, and γ-Oryzanol Components from Rice Bran against Cholesterol Oxidation Accelerated by 2,2′-Azobis(2-methylpropionamidine) Dihydrochloride. J. Agric. Food Chem. 2001, 49, 2077–2081. [Google Scholar] [CrossRef] [PubMed]
- Stoggl, W.; Huck, C.; Wongyai, S.; Scherz, H.; Bonn, G. Simultaneous determination of carotenoids, tocopherols, and gamma-oryzanol in crude rice bran oil by liquid chromatography coupled to diode array and mass spectrometric detection employing silica C30 stationary phases. J. Sep. Sci. 2005, 28, 1712–1718. [Google Scholar] [CrossRef] [PubMed]
- Parrado, J.; Miramontes, E.; Jover, M.; Gutierrez, J.F.; Collantes de Terán, L.; Bautista, J. Preparation of a rice bran enzymatic extract with potential use as functional food. Food Chem. 2006, 98, 742–748. [Google Scholar] [CrossRef]
- Chen, F.; Wang, Z.; Zhao, G.; Liao, X.; Cai, T.; Guo, L.; Hu, X. Purification process of octacosanol extracts from rice bran wax by molecular distillation. J. Food Eng. 2007, 79, 63–68. [Google Scholar] [CrossRef]
- Sugihara, N.; Kanda, A.; Nakano, T.; Nakamura, T.; Igusa, H.; Hara, S. Novel Fractionation Method for Squalene and Phytosterols Contained in the Deodorization Distillate of Rice Bran Oil. J. Oleo Sci. 2010, 59, 65–70. [Google Scholar] [CrossRef] [PubMed]
- De Deckere, E.A.; Korver, O. Minor constituents of rice bran oil as functional foods. Nutr. Rev. 1996, 54, S120–S126. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Wen, W.; Zhang, R.; Wei, Z.; Deng, Y.; Xiao, J.; Zhang, M. Complex enzyme hydrolysis releases antioxidative phenolics from rice bran. Food Chem. 2017, 214, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Kinyuru, J.N.; Borresen, E.C.; Ryan, E.P. Nutritional and Safety Evaluation of Heat-Stabilized Rice Bran for Sup-plementary Feeding of Malnourished Children in Kenya. Int. J. Food Sci. Nutr. Diet 2015, 4, 226–232. [Google Scholar] [CrossRef]
- Sheflin, A.M.; Borresen, E.C.; Kirkwood, J.S.; Boot, C.M.; Whitney, A.K.; Lu, S.; Brown, R.J.; Broeckling, C.D.; Ryan, E.P.; Weir, T.L. Dietary supplementation with rice bran or navy bean alters gut bacterial metabolism in colorectal cancer survivors. Mol. Nutr. Food Res. 2017, 61. [Google Scholar] [CrossRef] [PubMed]
- Borresen, E.C.; Jenkins-Puccetti, N.; Schmitz, K.; Brown, D.G.; Pollack, A.; Fairbanks, A.; Wdowik, M.; Rao, S.; Nelson, T.L.; Luckasen, G.; et al. A Pilot Randomized Controlled Clinical Trial to Assess Tolerance and Efficacy of Navy Bean and Rice Bran Supplementation for Lowering Cholesterol in Children. Glob. Pediatr. Health 2017, 4. [Google Scholar] [CrossRef] [PubMed]
- Lai, M.-H.; Chen, Y.-T.; Chen, Y.-Y.; Chang, J.-H.; Cheng, H.-H. Effects of rice bran oil on the blood lipids profiles and insulin resistance in type 2 diabetes patients. J. Clin. Biochem. Nutr. 2012, 51, 15–18. [Google Scholar] [CrossRef] [PubMed]
- Choi, S.P.; Kim, S.P.; Kang, M.Y.; Nam, S.H.; Friedman, M. Protective Effects of Black Rice Bran against Chemically-Induced Inflammation of Mouse Skin. J. Agric. Food Chem. 2010, 58, 10007–10015. [Google Scholar] [CrossRef] [PubMed]
- Qureshi, A.A.; Bradlow, B.A.; Salser, W.A.; Brace, L.D. Novel tocotrienols of rice bran modulate cardiovascular disease risk parameters of hypercholesterolemic humans. J. Nutr. Biochem. 1997, 8, 290–298. [Google Scholar] [CrossRef]
- Verschoyle, R.D.; Greaves, P.; Cai, H.; Edwards, R.E.; Steward, W.P.; Gescher, A.J. Evaluation of the cancer chemopreventive efficacy of rice bran in genetic mouse models of breast, prostate and intestinal carcinogenesis. Br. J. Cancer 2007, 96, 248–254. [Google Scholar] [CrossRef] [PubMed]
- Badr El-Din, N.K.; Ali, D.A.; Othman, R.; Ghoneum, M. Abstract 5259: Prevention of hepatocarcinogenesis in rats by arabinoxylan rice bran, MGN-3/Biobran. Cancer Res. 2016, 76, 5259. [Google Scholar] [CrossRef]
- Yang, Z.; Nakabayashi, R.; Mori, T.; Takamatsu, S.; Kitanaka, S.; Saito, K. Metabolome Analysis of Oryza sativa (Rice) Using Liquid Chromatography-Mass Spectrometry for Characterizing Organ Specificity of Flavonoids with Anti-inflammatory and Anti-oxidant Activity. Chem. Pharm. Bull. 2016, 64, 952–956. [Google Scholar] [CrossRef] [PubMed]
- Mahender, A.; Anandan, A.; Pradhan, S.K.; Pandit, E. Rice grain nutritional traits and their enhancement using relevant genes and QTLs through advanced approaches. Springer Plus 2016, 5, 2086. [Google Scholar] [CrossRef] [PubMed]
- Park, H.Y.; Lee, K.W.; Choi, H.D. Rice bran constituents: Immunomodulatory and therapeutic activities. Food Funct. 2017, 8, 935–943. [Google Scholar] [CrossRef] [PubMed]
- Ryan, E.P. Bioactive food components and health properties of rice bran. J. Am. Vet. Med. Assoc. 2011, 238, 593–600. [Google Scholar] [CrossRef] [PubMed]
- Maruyama, K.; Urano, K.; Yoshiwara, K.; Morishita, Y.; Sakurai, N.; Suzuki, H.; Kojima, M.; Sakakibara, H.; Shibata, D.; Saito, K.; et al. Integrated Analysis of the Effects of Cold and Dehydration on Rice Metabolites, Phytohormones, and Gene Transcripts. Plant Physiol. 2014, 164, 1759. [Google Scholar] [CrossRef] [PubMed]
- Navarro-Reig, M.; Jaumot, J.; Pina, B.; Moyano, E.; Galceran, M.T.; Tauler, R. Metabolomic analysis of the effects of cadmium and copper treatment in Oryza sativa L. using untargeted liquid chromatography coupled to high resolution mass spectrometry and all-ion fragmentation. Metallomics 2017, 9, 660–675. [Google Scholar] [CrossRef] [PubMed]
- Masumoto, C.; Miyazawa, S.-I.; Ohkawa, H.; Fukuda, T.; Taniguchi, Y.; Murayama, S.; Kusano, M.; Saito, K.; Fukayama, H.; Miyao, M. Phosphoenolpyruvate carboxylase intrinsically located in the chloroplast of rice plays a crucial role in ammonium assimilation. Proc. Natl. Acad. Sci. USA 2010, 107, 5226–5231. [Google Scholar] [CrossRef] [PubMed]
- Okazaki, Y.; Otsuki, H.; Narisawa, T.; Kobayashi, M.; Sawai, S.; Kamide, Y.; Kusano, M.; Aoki, T.; Hirai, M.Y.; Saito, K. A new class of plant lipid is essential for protection against phosphorus depletion. Nat. Commun. 2013, 4, 1510. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Gong, L.; Guo, Z.; Wang, W.; Zhang, H.; Liu, X.; Yu, S.; Xiong, L.; Luo, J. A Novel Integrated Method for Large-Scale Detection, Identification, and Quantification of Widely Targeted Metabolites: Application in the Study of Rice Metabolomics. Mol. Plant 2013, 6, 1769–1780. [Google Scholar] [CrossRef] [PubMed]
- Mahdavi, V.; Farimani, M.M.; Fathi, F.; Ghassempour, A. A targeted metabolomics approach toward understanding metabolic variations in rice under pesticide stress. Anal. Biochem. 2015, 478, 65–72. [Google Scholar] [CrossRef] [PubMed]
- Kusano, M.; Fukushima, A.; Kobayashi, M.; Hayashi, N.; Jonsson, P.; Moritz, T.; Ebana, K.; Saito, K. Application of a metabolomic method combining one-dimensional and two-dimensional gas chromatography-time-of-flight/mass spectrometry to metabolic phenotyping of natural variants in rice. J. Chromatogr. B 2007, 855, 71–79. [Google Scholar] [CrossRef] [PubMed]
- Heuberger, A.L.; Lewis, M.R.; Chen, M.-H.; Brick, M.A.; Leach, J.E.; Ryan, E.P. Metabolomic and functional genomic analyses reveal varietal differences in bioactive compounds of cooked rice. PLoS ONE 2010, 5, e12915. [Google Scholar] [CrossRef] [PubMed]
- Hu, C.; Shi, J.; Quan, S.; Cui, B.; Kleessen, S.; Nikoloski, Z.; Tohge, T.; Alexander, D.; Guo, L.; Lin, H.; et al. Metabolic variation between japonica and indica rice cultivars as revealed by non-targeted metabolomics. Sci. Rep. 2014, 4, 5067. [Google Scholar] [CrossRef] [PubMed]
- Galland, M.; He, D.; Lounifi, I.; Arc, E.; Clement, G.; Balzergue, S.; Huguet, S.; Cueff, G.; Godin, B.; Collet, B.; et al. An Integrated “Multi-Omics” Comparison of Embryo and Endosperm Tissue-Specific Features and Their Impact on Rice Seed Quality. Front. Plant Sci. 2017, 8, 1984. [Google Scholar] [CrossRef] [PubMed]
- Concepcion, J.C.T.; Ouk, S.; Riedel, A.; Calingacion, M.; Zhao, D.; Ouk, M.; Garson, M.J.; Fitzgerald, M.A. Quality evaluation, fatty acid analysis and untargeted profiling of volatiles in Cambodian rice. Food Chem. 2018, 240, 1014–1021. [Google Scholar] [CrossRef] [PubMed]
- Daygon, V.D.; Calingacion, M.; Forster, L.C.; Voss, J.J.; Schwartz, B.D.; Ovenden, B.; Alonso, D.E.; McCouch, S.R.; Garson, M.J.; Fitzgerald, M.A. Metabolomics and genomics combine to unravel the pathway for the presence of fragrance in rice. Sci. Rep. 2017, 7, 8767. [Google Scholar] [CrossRef] [PubMed]
- Ryan, E.P.; Heuberger, A.L.; Weir, T.L.; Barnett, B.; Broeckling, C.D.; Prenni, J.E. Rice bran fermented with saccharomyces boulardii generates novel metabolite profiles with bioactivity. J. Agric. Food Chem. 2011, 59, 1862–1870. [Google Scholar] [CrossRef] [PubMed]
- Chaudhary, N.; Khurana, P. Vitamin E biosynthesis genes in rice: Molecular characterization, expression profiling and comparative phylogenetic analysis. Plant Sci. 2009, 177, 479–491. [Google Scholar] [CrossRef]
- Katherine, J.; Li, E.C.B.; NaNet, J.-P.; Gary, L.; Elizabeth, P.R. Navy Bean and Rice Bran Intake Alters the Plasma Metabolome of Children at Risk for Cardiovascular Disease. Front. Nutr. 2017, 4, 71. [Google Scholar] [CrossRef]
- Trehan, I.; Manary, M.J. Management of severe acute malnutrition in low-income and middle-income countries. Arch. Dis. Child. 2015, 100, 283–287. [Google Scholar] [CrossRef] [PubMed]
- Rutishauser-Perera, A. Tackling the Double Burden of Malnutrition in Low and Middle-Income Countries: Response of the International Community. Available online: https://www.ennonline.net/fex/53/tacklingthedoubleburden (accessed on 12 November 2016).
- Yates, C.R.; Miller, D.D.; Zeng, K.; Thompson, K.E. Anti-Inflammatory Quinic Acid Derivatives for Oral Administration. U.S. Application US20090234015A1, 14 February 2012. [Google Scholar]
- De-Miguel, F.F.; Trueta, C. Synaptic and extrasynaptic secretion of serotonin. Cell. Mol. Neurobiol. 2005, 25, 297–312. [Google Scholar] [CrossRef] [PubMed]
- Naylor, J.M.; Leibel, T.; Middleton, D.M. Effect of glutamine or glycine containing oral electrolyte solutions on mucosal morphology, clinical and biochemical findings, in calves with viral induced diarrhea. Can. J. Vet. Res. 1997, 61, 43–48. [Google Scholar] [PubMed]
- Lu, T.-C.; Ko, Y.-Z.; Huang, H.-W.; Hung, Y.-C.; Lin, Y.-C.; Peng, W.-H. Analgesic and anti-inflammatory activities of aqueous extract from Glycine tomentella root in mice. J. Ethnopharmacol. 2007, 113, 142–148. [Google Scholar] [CrossRef] [PubMed]
- Ligumsky, M.; Sestieri, M.; Okon, E.; Ginsburg, I. Antioxidants Inhibit Ethanol-Induced Gastric Injury in the Rat: Role of Manganese, Glycine, and Carotene. Scand. J. Gastroenterol. 1995, 30, 854–860. [Google Scholar] [CrossRef] [PubMed]
- Jain, M.; Nilsson, R.; Sharma, S.; Madhusudhan, N.; Kitami, T.; Souza, A.L.; Kafri, R.; Kirschner, M.W.; Clish, C.B.; Mootha, V.K. Metabolite profiling identifies a key role for glycine in rapid cancer cell proliferation. Science 2012, 336, 1040–1044. [Google Scholar] [CrossRef] [PubMed]
- Tastesen, H.S.; Keenan, A.H.; Madsen, L.; Kristiansen, K.; Liaset, B. Scallop protein with endogenous high taurine and glycine content prevents high-fat, high-sucrose-induced obesity and improves plasma lipid profile in male C57BL/6J mice. Amino Acids 2014, 46, 1659–1671. [Google Scholar] [CrossRef] [PubMed]
- He, M. Pipecolic acid in microbes: Biosynthetic routes and enzymes. J. Ind. Microbiol. Biotechnol. 2006, 33, 401–407. [Google Scholar] [CrossRef] [PubMed]
- Bendahan, D.; Mattei, J.; Ghattas, B.; Confort-Gouny, S.; Le Guern, M.E.; Cozzone, P. Citrulline/malate promotes aerobic energy production in human exercising muscle. Br. J. Sports Med. 2002, 36, 282–289. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Wu, Q.; Huang, J.; Chen, R. Effects of L-malate on physical stamina and activities of enzymes related to the malate-aspartate shuttle in liver of mice. Physiol. Res. 2007, 56, 213. [Google Scholar] [PubMed]
- Nealon, N.J.; Worcester, C.R.; Ryan, E.P. Lactobacillus paracasei metabolism of rice bran reveals metabolome associated with Salmonella Typhimurium growth reduction. J. Appl. Microbiol. 2017, 122, 1639–1656. [Google Scholar] [CrossRef] [PubMed]
- Forster, G.M.; Raina, K.; Kumar, A.; Kumar, S.; Agarwal, R.; Chen, M.H.; Bauer, J.E.; McClung, A.M.; Ryan, E.P. Rice varietal differences in bioactive bran components for inhibition of colorectal cancer cell growth. Food Chem. 2013, 141, 1545–1552. [Google Scholar] [CrossRef] [PubMed]
- Dietrich, M.; Traber, M.G.; Jacques, P.F.; Cross, C.E.; Hu, Y.; Block, G. Does γ-Tocopherol Play a Role in the Primary Prevention of Heart Disease and Cancer? A Review. J. Am. Coll. Nutr. 2006, 25, 292–299. [Google Scholar] [CrossRef] [PubMed]
- Meganathan, P.; Fu, J.Y. Biological Properties of Tocotrienols: Evidence in Human Studies. Int. J. Mol. Sci. 2016, 17, 1682. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Q.; Elson-Schwab, I.; Courtemanche, C.; Ames, B.N. γ-Tocopherol and its major metabolite, in contrast to α-tocopherol, inhibit cyclooxygenase activity in macrophages and epithelial cells. Proc. Nat. Acad. Sci. USA 2000, 97, 11494–11499. [Google Scholar] [CrossRef] [PubMed]
- Chatelain, E.; Boscoboinik, D.O.; Bartoli, G.-M.; Kagan, V.E.; Gey, F.K.; Packer, L.; Azzi, A. Inhibition of smooth muscle cell proliferation and protein kinase C activity by tocopherols and tocotrienols. Biochim. Biophys. Acta 1993, 1176, 83–89. [Google Scholar] [CrossRef]
- Simopoulos, A.P. Omega-3 fatty acids in health and disease and in growth and development. Am. J. Clin. Nutr. 1991, 54, 438–463. [Google Scholar] [CrossRef] [PubMed]
- Carta, G.; Murru, E.; Banni, S.; Manca, C. Palmitic Acid: Physiological Role, Metabolism and Nutritional Implications. Front. Physiol. 2017, 8, 902. [Google Scholar] [CrossRef] [PubMed]
- Shultz, T.; Chew, B.; Seaman, W.; Luedecke, L. Inhibitory effect of conjugated dienoic derivatives of linoleic acid and β-carotene on the in vitro growth of human cancer cells. Cancer Lett. 1992, 63, 125–133. [Google Scholar] [CrossRef]
- Lee, K.N.; Kritchevsky, D.; Parizaa, M.W. Conjugated linoleic acid and atherosclerosis in rabbits. Atherosclerosis 1994, 108, 19–25. [Google Scholar] [CrossRef]
- Barceló-Coblijn, G.; Murphy, E.J. Alpha-linolenic acid and its conversion to longer chain n − 3 fatty acids: Benefits for human health and a role in maintaining tissue n − 3 fatty acid levels. Prog. Lipid Res. 2009, 48, 355–374. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.Y.; Chapkin, R.S. Importance of dietary gamma-linolenic acid in human health and nutrition. J. Nutr. 1998, 128, 1411–1414. [Google Scholar] [CrossRef] [PubMed]
- Kosova, M.; Hrádková, I.; Mátlová, V.; Kadlec, D.; Šmidrkal, J.; Filip, V. Antimicrobial effect of 4-hydroxybenzoic acid ester with glycerol. J. Clin. Pharm. Ther. 2015, 40, 436–440. [Google Scholar] [CrossRef] [PubMed]
- Barreca, D.; Laganà, G.; Leuzzi, U.; Smeriglio, A.; Trombetta, D.; Bellocco, E. Evaluation of the nutraceutical, antioxidant and cytoprotective properties of ripe pistachio (Pistacia vera L., variety Bronte) hulls. Food Chem. 2016, 196, 493–502. [Google Scholar] [CrossRef] [PubMed]
- Rivas-San Vicente, M.; Plasencia, J. Salicylic acid beyond defence: Its role in plant growth and development. J. Exp. Bot. 2011, 62, 3321–3338. [Google Scholar] [CrossRef] [PubMed]
- Hawley, S.A.; Fullerton, M.D.; Ross, F.A.; Schertzer, J.D.; Chevtzoff, C.; Walker, K.J.; Peggie, M.W.; Zibrova, D.; Green, K.A.; Mustard, K.J. The ancient drug salicylate directly activates AMP-activated protein kinase. Science 2012, 336, 918–922. [Google Scholar] [CrossRef] [PubMed]
- Ittaman, S.V.; VanWormer, J.J.; Rezkalla, S.H. The Role of Aspirin in the Prevention of Cardiovascular Disease. Clin. Med. Res. 2014, 12, 147–154. [Google Scholar] [CrossRef] [PubMed]
- Christophersen, O.A. Radiation protection following nuclear power accidents: A survey of putative mechanisms involved in the radioprotective actions of taurine during and after radiation exposure. Microb. Ecol. Health Dis. 2012, 23. [Google Scholar] [CrossRef] [PubMed]
- Gürer, H.; Özgünes, H.; Saygin, E.; Ercal, N. Antioxidant effect of taurine against lead-induced oxidative stress. Arch. Environ. Contam. Toxicol. 2001, 41, 397–402. [Google Scholar] [CrossRef] [PubMed]
- Tsuboyama-Kasaoka, N.; Shozawa, C.; Sano, K.; Kamei, Y.; Kasaoka, S.; Hosokawa, Y.; Ezaki, O. Taurine (2-aminoethanesulfonic acid) deficiency creates a vicious circle promoting obesity. Endocrinology 2006, 147, 3276–3284. [Google Scholar] [CrossRef] [PubMed]
- El Idrissi, A.; Messing, J.; Scalia, J.; Trenkner, E. Prevention of Epileptic Seizures by Taurine; Springer: Basel, Switzerland, 2003; pp. 515–525. [Google Scholar]
- Huxtable, R. Physiological actions of taurine. Physiol. Rev. 1992, 72, 101–163. [Google Scholar] [CrossRef] [PubMed]
- Poulev, A.; Chen, M.-H.; Cherravuru, S.; Raskin, I.; Belanger, F.C. Variation in levels of the flavone tricin in bran from rice genotypes varying in pericarp color. J. Cereal Sci. 2018, 79, 226–232. [Google Scholar] [CrossRef]
- Chen, W.; Wang, W.; Peng, M.; Gong, L.; Gao, Y.; Wan, J.; Wang, S.; Shi, L.; Zhou, B.; Li, Z. Comparative and parallel genome-wide association studies for metabolic and agronomic traits in cereals. Nat. Commun. 2016, 7, 12767. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Gao, Y.; Xie, W.; Gong, L.; Lu, K.; Wang, W.; Li, Y.; Liu, X.; Zhang, H.; Dong, H.; et al. Genome-wide association analyses provide genetic and biochemical insights into natural variation in rice metabolism. Nat. Genet. 2014, 46, 714–721. [Google Scholar] [CrossRef] [PubMed]
- Matsuda, F.; Okazaki, Y.; Oikawa, A.; Kusano, M.; Nakabayashi, R.; Kikuchi, J.; Yonemaru, J.; Ebana, K.; Yano, M.; Saito, K. Dissection of genotype-phenotype associations in rice grains using metabolome quantitative trait loci analysis. Plant J. Cell Mol. Biol. 2012, 70, 624–636. [Google Scholar] [CrossRef] [PubMed]
- Sellamuthu, R.; Liu, G.F.; Ranganathan, C.B.; Serraj, R. Genetic analysis and validation of quantitative trait loci associated with reproductive-growth traits and grain yield under drought stress in a doubled haploid line population of rice (Oryza sativa L.). Field Crops Res. 2011, 124, 46–58. [Google Scholar] [CrossRef]
- Huang, C.Y.; Roessner, U.; Eickmeier, I.; Genc, Y.; Callahan, D.L.; Shirley, N.; Langridge, P.; Bacic, A. Metabolite Profiling Reveals Distinct Changes in Carbon and Nitrogen Metabolism in Phosphate-Deficient Barley Plants (Hordeum vulgare L.). Plant Cell Physiol. 2008, 49, 691–703. [Google Scholar] [CrossRef] [PubMed]
- Harrigan, G.G.; Stork, L.G.; Riordan, S.G.; Reynolds, T.L.; Ridley, W.P.; Masucci, J.D.; MacIsaac, S.; Halls, S.C.; Orth, R.; Smith, R.G.; et al. Impact of Genetics and Environment on Nutritional and Metabolite Components of Maize Grain. J. Agric. Food Chem. 2007, 55, 6177–6185. [Google Scholar] [CrossRef] [PubMed]
- Harrigan, G.G.; Stork, L.G.; Riordan, S.G.; Ridley, W.P.; MacIsaac, S.; Halls, S.C.; Orth, R.; Rau, D.; Smith, R.G.; Wen, L.; et al. Metabolite Analyses of Grain from Maize Hybrids Grown in the United States under Drought and Watered Conditions during the 2002 Field Season. J. Agric. Food Chem. 2007, 55, 6169–6176. [Google Scholar] [CrossRef] [PubMed]
- Roessner, U.; Luedemann, A.; Brust, D.; Fiehn, O.; Linke, T.; Willmitzer, L.; Fernie, A.R. Metabolic Profiling Allows Comprehensive Phenotyping of Genetically or Environmentally Modified Plant Systems. Plant Cell 2001, 13, 11. [Google Scholar] [CrossRef] [PubMed]
- Schauer, N.; Semel, Y.; Roessner, U.; Gur, A.; Balbo, I.; Carrari, F.; Pleban, T.; Perez-Melis, A.; Bruedigam, C.; Kopka, J. Comprehensive metabolic profiling and phenotyping of interspecific introgression lines for tomato improvement. Nat. Biotechnol. 2006, 24, 447–454. [Google Scholar] [CrossRef] [PubMed]
- Schauer, N.; Semel, Y.; Balbo, I.; Steinfath, M.; Repsilber, D.; Selbig, J.; Pleban, T.; Zamir, D.; Fernie, A.R. Mode of inheritance of primary metabolic traits in tomato. Plant Cell 2008, 20, 509–523. [Google Scholar] [CrossRef] [PubMed]
- Goodyear, A.; Kumar, A.; Ehrhart, E.J.; Swanson, K.S.; Grusak, M.A.; Leach, J.E.; Dow, S.W.; McClung, A.; Ryan, E.P. Dietary rice bran supplementation prevents Salmonella colonization differentially across varieties and by priming intestinal immunity. J. Funct. Foods 2015, 18, 653–664. [Google Scholar] [CrossRef]
- Mahesh, H.B.; Shirke, M.D.; Singh, S.; Rajamani, A.; Hittalmani, S.; Wang, G.-L.; Gowda, M. Indica rice genome assembly, annotation and mining of blast disease resistance genes. BMC Genom. 2016, 17, 242. [Google Scholar] [CrossRef] [PubMed]

| Rice Cultivar | Kenya—Basmati 217 | Kenya—Basmati 370 | Mali—Gambiaka * | Mali—Shwetasoke | Mali—DM-16 | Mali—Khao Gaew | Nicaragua—Dorado | Nepal—Sawa Mahsuli | India—Chennula | India—Njavara * | USA—Calrose | USA—RBT 300 | USA—Jasmine 85 | USA—IAC 600 | USA—LTH | USA—SHZ-2 | Cambodia —Rang Jey | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Chemical Class | ||||||||||||||||||
| Amino acids | 119 | 117 | 120 | 117 | 115 | 99 | 117 | 117 | 105 | 99 | 119 | 122 | 110 | 113 | 119 | 117 | 108 | |
| Carbohydrates | 51 | 52 | 53 | 53 | 52 | 50 | 51 | 50 | 51 | 50 | 48 | 49 | 50 | 53 | 53 | 50 | 51 | |
| Cofactors & vitamins | 27 | 27 | 27 | 27 | 26 | 24 | 27 | 27 | 27 | 26 | 27 | 23 | 27 | 27 | 27 | 26 | 27 | |
| Lipids | 166 | 165 | 165 | 166 | 159 | 164 | 161 | 155 | 155 | 151 | 165 | 161 | 146 | 163 | 165 | 150 | 156 | |
| Nucleotides | 37 | 38 | 35 | 35 | 34 | 29 | 37 | 38 | 30 | 29 | 35 | 33 | 37 | 36 | 35 | 36 | 36 | |
| Peptides | 0 | 2 | 11 | 11 | 11 | 7 | 3 | 11 | 1 | 1 | 11 | 1 | 11 | 10 | 8 | 11 | 2 | |
| Secondary metabolites | 19 | 21 | 19 | 19 | 20 | 19 | 16 | 20 | 19 | 22 | 16 | 17 | 17 | 20 | 22 | 17 | 21 | |
| Total | 419 | 422 | 430 | 428 | 417 | 392 | 412 | 418 | 388 | 378 | 421 | 406 | 398 | 422 | 429 | 407 | 401 | |
| Rice Bran Metabolites | Precursor | Biosynthesis Pathway | No. of Genes * | Gene ID | Gene(s) Name |
|---|---|---|---|---|---|
| Amino Acids | |||||
| Aromatic amino acids (phosphoenolpyruvate (PEP)-derived) | |||||
| l-quinate | Trans-5-O-caffeoyl-d-quinate | Caffeoylglucarate biosynthesis | 5 | GN7F-30156 | LOC_Os02g39170.1 |
| GN7F-16973 | LOC_Os02g39590.1 | ||||
| GN7F-15850 | LOC_Os06g47910.1 | ||||
| Serotonin | Tryptamine | Hydroxycinnamic acid Serotonin amides biosynthesis | 2 | GN7F-19639 | LOC_Os08g04560.1 |
| GN7F-25663 | LOC_Os08g04540.1 | ||||
| Tryptophan | l-serine | Tryptophan biosynthesis | 4+ (153) | GN7F-27027 | LOC_Os03g58260.1 |
| GN7F-24368 | LOC_Os03g58290.1 | ||||
| GN7F-25293 | LOC_Os06g42560.4 | ||||
| GN7F-19428 | LOC_Os08g04180.1 | ||||
| Tyrosine | l-phenylalanine, l-arogenate | Phenylalanine degradation V, tyrosine biosynthesis II & III | 3+ (153) | GN7F-27976 | LOC_Os06g35050.1 |
| GN7F-19057 | LOC_Os06g49505.1 | ||||
| GN7F-18001 | LOC_Os06g49520.1 | ||||
| Aspartate family (oxaloacetate (OAA)-derived) | |||||
| Asparagine | l-aspartate, 3-cyano-l-alanine | Asparagine biosynthesis I & II, cyanide detoxification I | 5+ (153) | GN7F-32447 | LOC_Os12g38630.1 |
| GN7F-23509 | LOC_Os06g15420.1 | ||||
| GN7F-23610 | LOC_Os03g18130.1 | ||||
| GN7F-15965 | LOC_Os02g42350.1 | ||||
| GN7F-23159 | LOC_Os02g42330.1 | ||||
| Aspartate | l-asparagine, 3-cyano-l-alanine, indole-3-acetyl-aspartate-N-β-d-glucose | Asparagine degradation I, cyanide detoxification I, Indole-3-acetate conjugate biosynthesis II | 3+ (153) | GN7F-15965 | LOC_Os02g42350.1 |
| GN7F-23159 | LOC_Os02g42330.1 | ||||
| GN7F-27949 | LOC_Os04g58600.2 | ||||
| Lysine | Meso-diaminopimelate | Lysine biosynthesis VI | 1+ (153) | GN7F-25633 | LOC_Os02g24354.1 |
| Methionine sulfoxide | An acyl-CoA, an aldehyde, a carboxylic ester | Not in pathway | 9+ (150) | GN7F-28729 | LOC_Os09g34190.1 |
| GN7F-19329 | LOC_Os04g47120.1 | ||||
| GN7F-31811 | LOC_Os01g12910.1 | ||||
| GN7F-31843 | LOC_Os07g27870.1 | ||||
| GN7F-32115 | LOC_Os04g35590.1 | ||||
| GN7F-32205 | LOC_Os07g27960.1 | ||||
| GN7F-32376 | LOC_Os02g32200.1 | ||||
| GN7F-32723 | LOC_Os01g12920.1 | ||||
| GN7F-32765 | LOC_Os01g65950.1 | ||||
| Threonine | O-phospho-l-homoserine, l-threonine 3-O-phosphate | Threonine biosynthesis from homoserine, l-threonine 3-O-phosphate | 3+ (153) | GN7F-18835 | LOC_Os01g49890.1 |
| GN7F-30196 | LOC_Os05g47640.1 | ||||
| GN7F-29436 | LOC_Os08g17784.1 | ||||
| Glutamate family (α-ketoglutarate-derived) | |||||
| Arginine | l-arginino-succinate | Arginine biosynthesis I & II, Citrulline-nitric oxide cycle | 3+ (153) | GN7F-20973 | LOC_Os03g19280.1 |
| GN7F-32460 | LOC_Os03g60976.1 | ||||
| GN7F-32707 | LOC_Os03g60992.1 | ||||
| Glutamate 1 | More than 35 precursors (top two: l-glutamine, 2-oxoglutarate) | More than 27 pathways (top two: 4-aminobenzoate biosynthesis, 4-aminobutyrate degradation) | 91+ (157) | GN7F-17849 | LOC_Os06g48620.1 |
| GN7F-25902 | LOC_Os04g52440.1 | ||||
| GN7F-28339 | LOC_Os08g10510.1 | ||||
| GN7F-27233 | LOC_Os02g02210.1 | ||||
| GN7F-19896 | LOC_Os04g52450.1 | ||||
| Glutamine | More than eight precursors (top three: l-glutamate, a dipeptide with proline at the C-terminal, a γ l-glutamyl-l-amino acid) | More than six pathways (top four: Ammonia assimilation cycle I & II, glutamine biosynthesis I & II) | 5+ (153) | GN7F-15709 | LOC_Os03g50490.1 |
| GN7F-15901 | LOC_Os04g56400.1 | ||||
| GN7F-22516 | LOC_Os03g12290.1 | ||||
| GN7F-26393 | LOC_Os10g31820.1 | ||||
| GN7F-27460 | LOC_Os02g50240.1 | ||||
| N-acetylglutamate | l-glutamate | Arginine biosynthesis II (acetyl cycle), ornithine biosynthesis | 6+ (150) | GN7F-20894 | LOC_Os03g17120.1 |
| GN7F-19328 | LOC_Os07g39690.1 | ||||
| GN7F-17187 | LOC_Os03g31690.1 | ||||
| GN7F-31311 | LOC_Os03g46200.1 | ||||
| GN7F-32148 | LOC_Os03g58010.1 | ||||
| GN7F-32821 | LOC_Os03g58030.1 | ||||
| Pyroglutamine | An (γ-l-glutamyl)-l-amino acid | γ-glutamylcyclotransferase | 2 | GN7F-31386 | LOC_Os03g63700.1 |
| GN7F-32110 | LOC_Os11g04420.4 | ||||
| Serine family (phosphoglycerate-derived) | |||||
| Glycine | l-cysteinyl-glycine | γ-glutamyl cycle, Phytochelatins biosynthesis | 8+ (151) | GN7F-30607 | LOC_Os01g05810.1 |
| GN7F-16957 | LOC_Os04g38450.1 | ||||
| GN7F-25454 | LOC_Os01g05820.1 | ||||
| GN7F-26619 | LOC_Os05g34290.1 | ||||
| GN7F-28075 | LOC_Os06g01260.1 | ||||
| GN7F-16002 | LOC_Os12g35890.1 | ||||
| GN7F-26733 | LOC_Os09g32290.2 | ||||
| GN7F-19516 | LOC_Os01g21380.1 | ||||
| Taurine 2 | In transport reactions | 4 | GN7F-31312 | LOC_Os09g29660.1 | |
| GN7F-31505 | LOC_Os05g31080.1 | ||||
| GN7F-32616 | LOC_Os03g20170.1 | ||||
| GN7F-32729 | LOC_Os09g29670.1 | ||||
| Lipids | |||||
| Free fatty acid | |||||
| α-linolenate | A phosphatidylcholine | No common pathways | 1+ (334) | GN7F-18386 | LOC_Os11g04940.1 |
| Laurate | Lauroyl-CoA, a dodecanoyl-[acyl-carrier protein] | Palmitate biosynthesis II (bacteria and plants), sporopollenin precursors biosynthesis | 1+ (285) | GN7F-19329 | LOC_Os04g47120.1 |
| Linoleate | Linoleoyl-CoA | No common pathways | 2+ (332) | GN7F-33079 | LOC_Os02g44654.2 |
| GN7F-16232 | LOC_Os04g47250.1 | ||||
| Myristate | Myristoyl-CoA | No common pathways | 0+ (325) | - | - |
| Palmitate | Palmitoyl-CoA, a palmitoyl-[acp], 1,2-dipalmitoyl-phosphatidylcholine, 1-palmitoyl-2-linoleoyl-phosphatidylcholine | Cutin biosynthesis, sporopollenin precursors biosynthesis, suberin monomers biosynthesis, palmitate biosynthesis II (bacteria and plants), phospholipid remodeling (phosphatidylcholine, yeast) | 6+ (333) | GN7F-19329 | LOC_Os04g47120.1 |
| GN7F-28729 | LOC_Os09g34190.1 | ||||
| GN7F-32532 | LOC_Os01g66240.1 | ||||
| GN7F-24830 | LOC_Os01g46250.1 | ||||
| GN7F-20654 | LOC_Os01g73740.1 | ||||
| GN7F-18158 | LOC_Os01g51360.1 | ||||
| Palmitoleate | A palmitoleoyl-[acyl-carrier protein] | No common pathways | 4+ (333) | GN7F-31362 | LOC_Os04g46710.1 |
| GN7F-31587 | LOC_Os04g46730.1 | ||||
| GN7F-31765 | LOC_Os02g44134.1 | ||||
| GN7F-32681 | LOC_Os02g44200.1 | ||||
| Oxylipins | |||||
| 9,10-epoxystearate | Oleate, a hydroperoxy-fatty-acyl-[lipid] | Cutin biosynthesis, poly-hydroxy fatty acid biosynthesis | 10+ (332) | GN7F-31064 | LOC_Os10g37070.1 |
| GN7F-30659 | LOC_Os08g05620.1 | ||||
| GN7F-18468 | LOC_Os01g24810.1 | ||||
| GN7F-27809 | LOC_Os10g05020.1 | ||||
| GN7F-16745 | LOC_Os04g03890.1 | ||||
| GN7F-21459 | LOC_Os10g37100.1 | ||||
| GN7F-19209 | LOC_Os04g33370.1 | ||||
| GN7F-18601 | LOC_Os02g01890.1 | ||||
| GN7F-29035 | LOC_Os08g05610.1 | ||||
| GN7F-15971 | LOC_Os06g46680.1 | ||||
| Phospholipid Metabolism | |||||
| Glycerophosphorylcholin (sn-glycero-3-phosphocholine) | A 1-acyl-sn-glycero-3-phosphocholine | No common pathways | 7 | GN7F-33055 | LOC_Os01g07960.3 |
| GN7F-29598 | LOC_Os04g57370.1 | ||||
| GN7F-18763 | LOC_Os04g09540.1 | ||||
| GN7F-31398 | LOC_Os01g42690.1 | ||||
| GN7F-32181 | LOC_Os04g57390.1 | ||||
| GN7F-32498 | LOC_Os05g51050.1 | ||||
| GN7F-32638 | LOC_Os08g42680.1 | ||||
| Rice Bran Metabolites | Precursor | Biosynthesis Pathway | No. of Genes * | Gene ID | Gene(s) Name |
|---|---|---|---|---|---|
| Carbohydrates | |||||
| Citric acid cycle (TCA) cycle | |||||
| α-ketoglutarate (2-oxoglutarate) 1 | l-glutamate, d-threo-isocitrate | Nine pathways (top four: Alanine degradation II, Glutamate degradation I, Arginine biosynthesis II, Ornithine biosynthesis) | 11+ (160) | GN7F-23833 | LOC_Os03g58040.1 |
| GN7F-16177 | LOC_Os02g43470.1 | ||||
| GN7F-28651 | LOC_Os04g45970.1 | ||||
| GN7F-23252 | LOC_Os05g03830.1 | ||||
| GN7F-26515 | LOC_Os07g27780.1 | ||||
| cis-aconitate | Citrate | Glutamine biosynthesis III glyoxylate cycle, TCA cycle II (plants and fungi) | 4+ (150) | GN7F-24702 | |
| GN7F-28499 | LOC_Os10g03960.1 | ||||
| GN7F-23691 | LOC_Os06g19960.1 | ||||
| GN7F-25422 | LOC_Os03g04410.1 | ||||
| GN7F-25644 | LOC_Os08g09200.1 | ||||
| Malate | Acetyl-CoA, Glyoxylate, Fumarate | Glycolate and glyoxylate degradation II, glyoxylate cycle, superpathway of glyoxylate cycle and fatty acid degradation, TCA cycle II (plants and fungi) | 2+ (150) | GN7F-24000 | LOC_Os04g40990.1 |
| GN7F-21211 | LOC_Os03g21950.1 | ||||
| Cofactors & vitamins | |||||
| Tocopherol metabolism | |||||
| β-tocopherol | δ-tocopherol, S-adenosyl-l-methionine | Vitamin E biosynthesis (tocopherols) | 4 | GN7F-31982 | LOC_Os10g41970.1 |
| GN7F-31239 | LOC_Os03g26200.1 | ||||
| GN7F-31334 | LOC_Os08g02600.1 | ||||
| GN7F-25500 | LOC_Os02g47310.1 | ||||
| γ-tocotrienol | 2,3-dimethyl-6-geranylgeranyl-1,4-benzoquinol | Vitamin E biosynthesis (tocopherols) | 1 | GN7F-24601 | LOC_Os02g17650.1 |
| Rice Bran Metabolites | Precursor | Biosynthesis Pathway | No. of Genes * | Gene ID | Gene(s) Name |
|---|---|---|---|---|---|
| Nucleotides | |||||
| Purine metabolism | |||||
| Adenine 1 | S-methyl-5′-thioadenosine, adenosine, N6-dimethylallyladenine, trans-zeatin, cis-zeatin, N1-ethyladenine, N1-methyladenine | S-methyl-5′-thioadenosine degradation I, adenine and adenosine salvage II, cytokinins degradation | 18 | GN7F-25353 | LOC_Os08g44370.1 |
| GN7F-26929 | LOC_Os09g39440.1 | ||||
| GN7F-32781 | LOC_Os05g33644.1 | ||||
| GN7F-32797 | LOC_Os05g33630.1 | ||||
| GN7F-19530 | LOC_Os06g37500.1 | ||||
| Adenosine | S-adenosyl-l-homocysteine, trans-zeatin riboside, isopentenyl adenosine | S-adenosyl-l-methionine cycle II, l-methionine degradation I (to l-homocysteine), cytokinins degradation | 3+ (8) | GN7F-20280 | LOC_Os02g12780.1 |
| GN7F-19530 | LOC_Os06g37500.1 | ||||
| GN7F-20388 | LOC_Os01g09260.1 | ||||
| Adenosine 5′-monophosphate 1 | Adenosine triphosphate (ATP) | More than 97 pathways (top four: Trans-zeatin biosynthesis, Adenosine nucleotides degradation I, 4-hydroxybenzoate biosynthesis I (eukaryotes), l-arginine biosynthesis I (via l-ornithine) | 299+ (24) | GN7F-23647 | LOC_Os02g46970.1 |
| GN7F-23504 | LOC_Os06g44620.1 | ||||
| GN7F-28551 | LOC_Os08g34790.1 | ||||
| GN7F-25622 | LOC_Os08g14760.1 | ||||
| GN7F-25996 | LOC_Os01g24030.1 | ||||
| Hypoxanthine | Inosine | Adenosine nucleotide degradation I | 5 | GN7F-25353 | LOC_Os08g44370.1 |
| GN7F-21573 | LOC_Os03g31170.1 | ||||
| GN7F-26929 | LOC_Os09g39440.1 | ||||
| GN7F-32781 | LOC_Os05g33644.1 | ||||
| GN7F-32797 | LOC_Os05g33630.1 | ||||
| Secondary metabolites | |||||
| Benzenoids | |||||
| Salicylate | Methylsalicylate | Unknown | 3 + (152) | GN7F-26541 | LOC_Os05g30760.1 |
| GN7F-21475 | LOC_Os01g37650.1 | ||||
| GN7F-28107 | LOC_Os01g25360.1 | ||||
| Cultivar | Grain Type | Bran Color | Country of Origin | Growing Location |
|---|---|---|---|---|
| Basmati 217 | Long | Brown | India | Kenya |
| Basmati 370 | Long | Brown | India | Kenya |
| Calrose | Medium | Brown | USA | California |
| Chennula | Long | Brown | India | India |
| DM-16 | Short | Brown | South America | Mali |
| Dorado | Long | Brown | Colombia | Nicaragua |
| Gambiaka | Long | Brown | Mali | Mali |
| IAC 600 | Medium | Purple | Brazil | Arkansas |
| Jasmine 85 | Long | Brown | Philippines | Arkansas |
| Khao Gaew | Long | Brown | Thailand | Mali |
| Li-Jiang-Xin-Tuan-Hei-Gu (LTH) | Medium | Red | China | Arkansas |
| Njavara | Long | Red | India | India |
| Rang Jey | Medium | Brown | Cambodia | Cambodia |
| RBT 300 * | Medium | Brown | USA | California |
| Sawa Mahsuli | Long | Brown | Nepal | Nepal |
| Shan-Huang-Zhan-2 (SHZ-2) | Long | Brown | China | Arkansas |
| Shwetasoke | Long | Brown | Mali | Mali |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zarei, I.; Luna, E.; Leach, J.E.; McClung, A.; Vilchez, S.; Koita, O.; Ryan, E.P. Comparative Rice Bran Metabolomics across Diverse Cultivars and Functional Rice Gene–Bran Metabolite Relationships. Metabolites 2018, 8, 63. https://doi.org/10.3390/metabo8040063
Zarei I, Luna E, Leach JE, McClung A, Vilchez S, Koita O, Ryan EP. Comparative Rice Bran Metabolomics across Diverse Cultivars and Functional Rice Gene–Bran Metabolite Relationships. Metabolites. 2018; 8(4):63. https://doi.org/10.3390/metabo8040063
Chicago/Turabian StyleZarei, Iman, Emily Luna, Jan E. Leach, Anna McClung, Samuel Vilchez, Ousmane Koita, and Elizabeth P. Ryan. 2018. "Comparative Rice Bran Metabolomics across Diverse Cultivars and Functional Rice Gene–Bran Metabolite Relationships" Metabolites 8, no. 4: 63. https://doi.org/10.3390/metabo8040063
APA StyleZarei, I., Luna, E., Leach, J. E., McClung, A., Vilchez, S., Koita, O., & Ryan, E. P. (2018). Comparative Rice Bran Metabolomics across Diverse Cultivars and Functional Rice Gene–Bran Metabolite Relationships. Metabolites, 8(4), 63. https://doi.org/10.3390/metabo8040063







