Distribution of Heparan Sulfate Oligosaccharides in Murine Mucopolysaccharidosis Type IIIA
Abstract
1. Introduction
2. Results
2.1. UA in Mouse Tissues and Urine
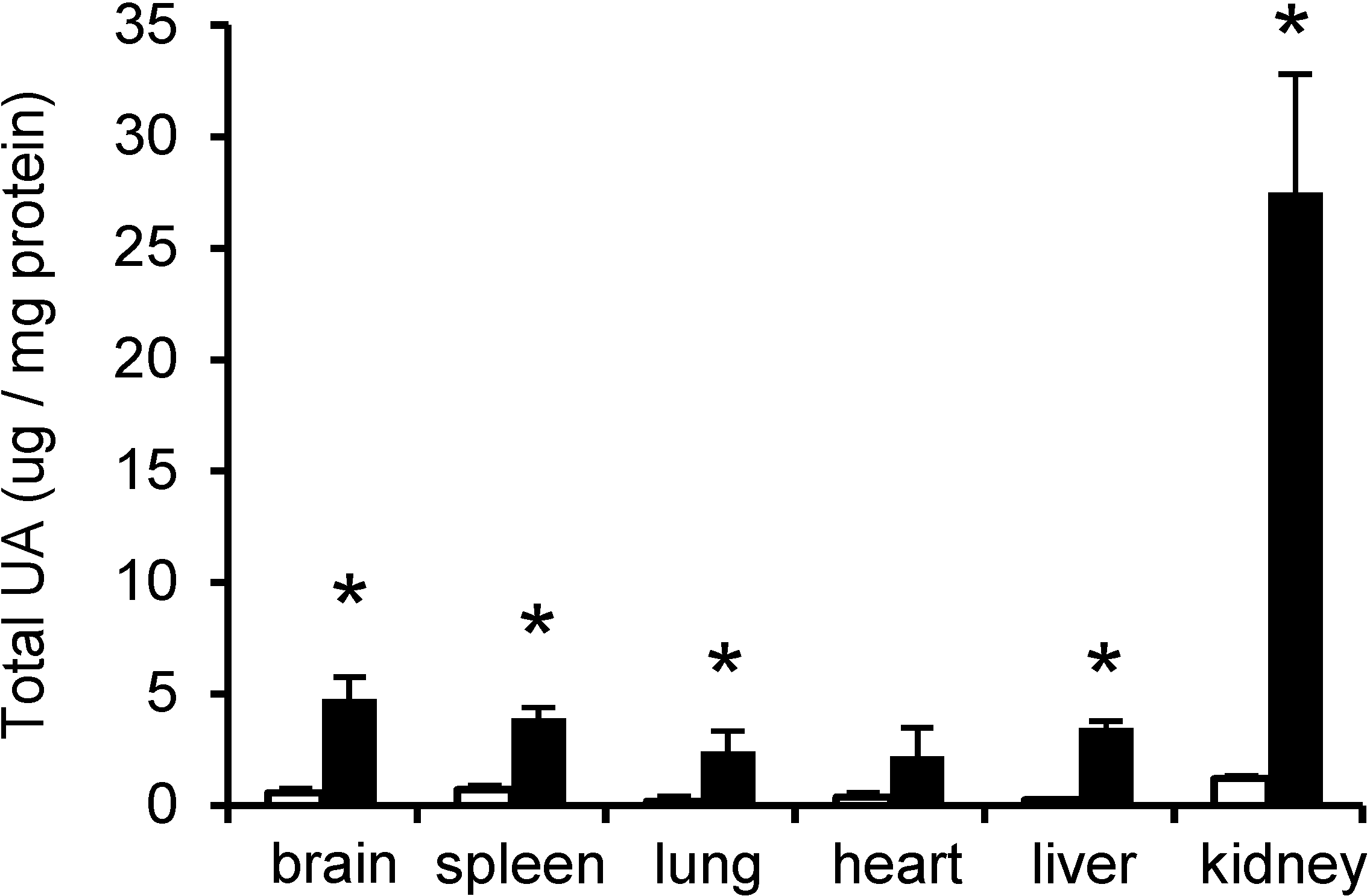
2.2. Distribution of Di- to Hexasaccharides in Mouse Tissues and Urine
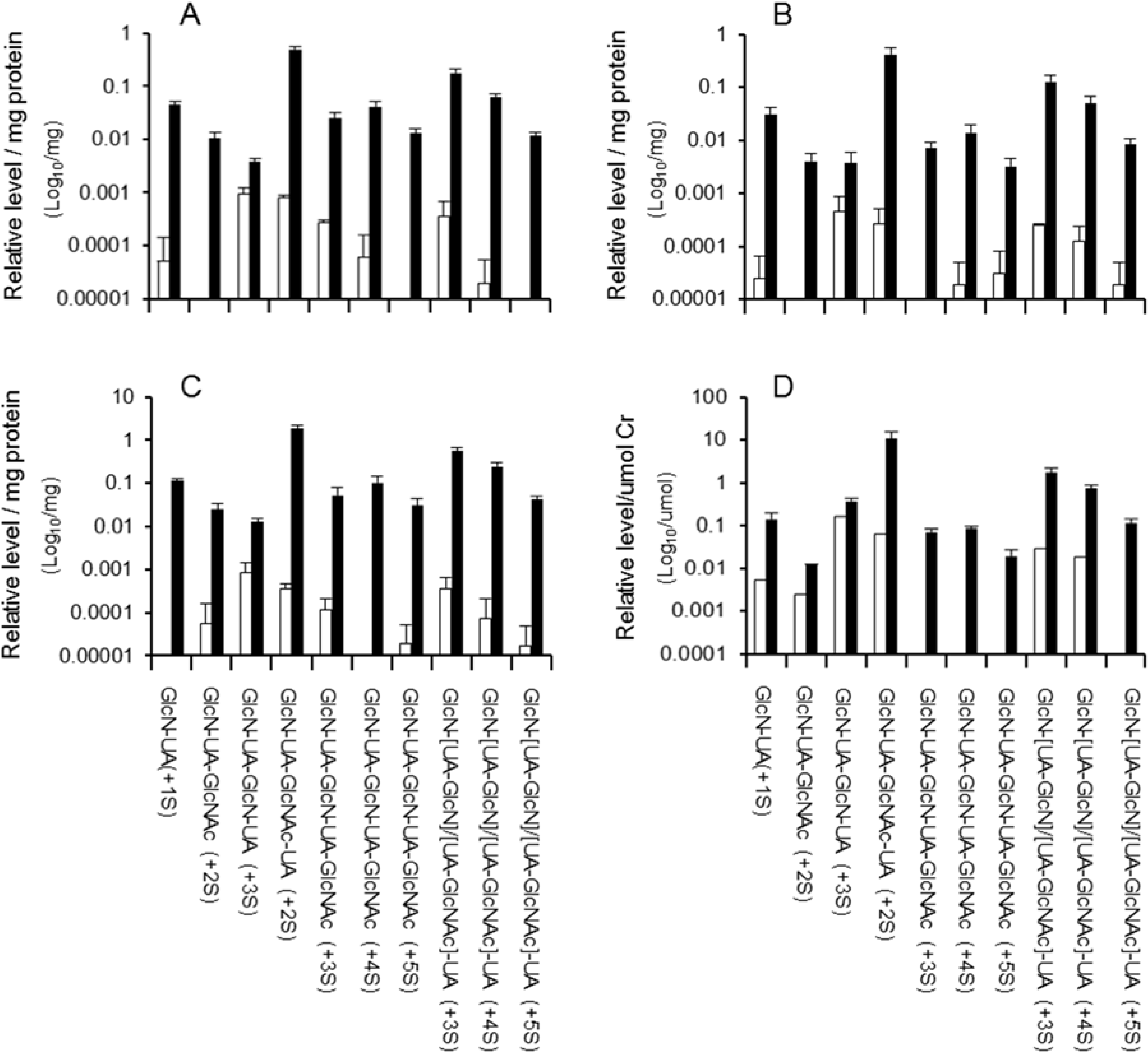
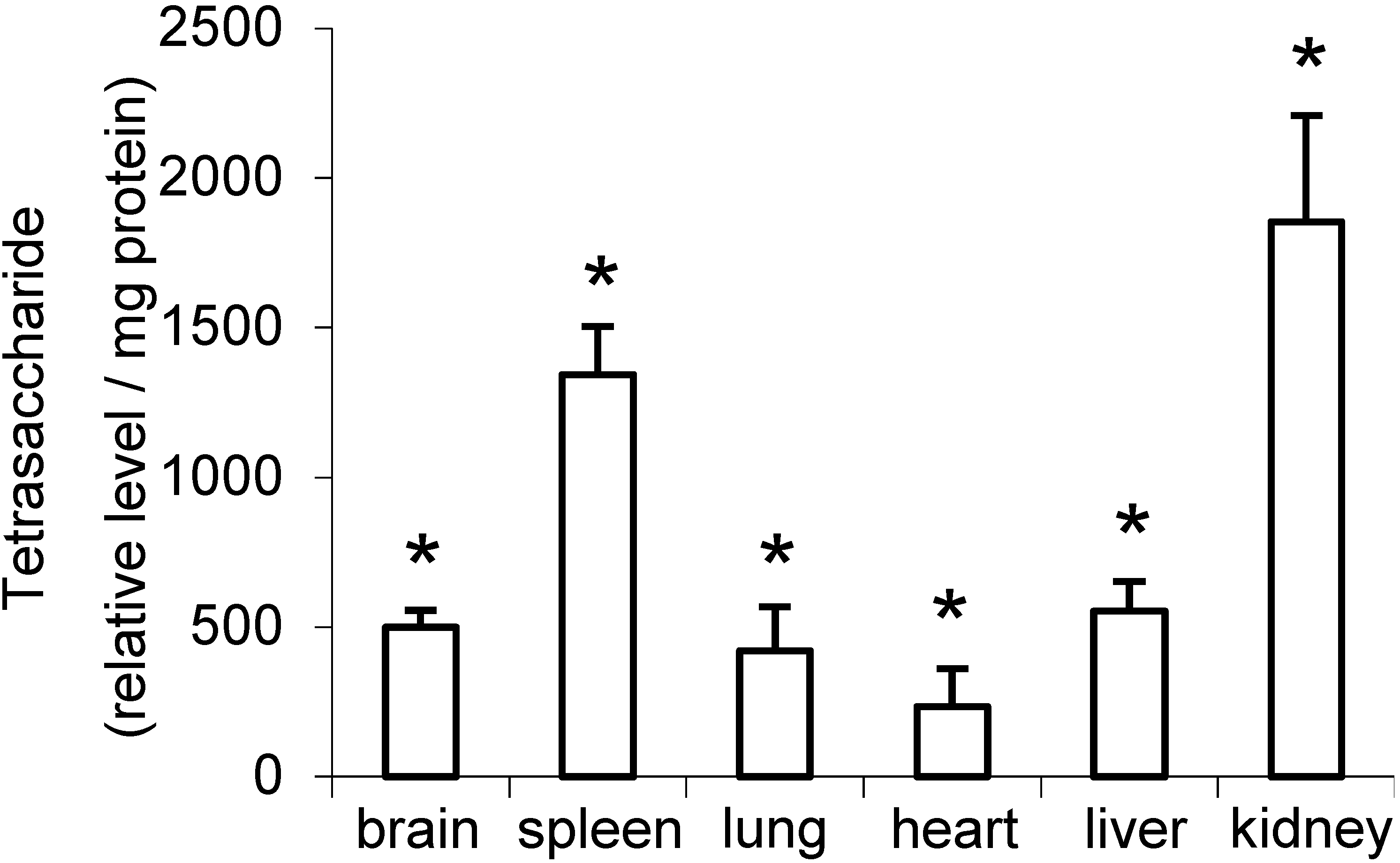
| Structure of oligosaccharide b | Brain c | Spleen c | Lung c | Heart c | Liver c | Kidney c | Average for all Tissues | Urine c |
|---|---|---|---|---|---|---|---|---|
| GlcN-UA (+S) | 5.0 | 4.8 | 4.6 | 5.2 | 6.0 | 3.7 | 5.0 | 2.1 |
| GlcN-UA-GlcNAc (+2S) | 1.1 | 0.7 | 0.6 | 0.6 | 0.9 | 0.8 | 0.8 | 0.9 |
| GlcN-UA-GlcN-UA (+3S) | 0.4 | 0.4 | 0.5 | 0.4 | 0.5 | 0.4 | 0.4 | 1.1 |
| GlcN-UA-GlcNAc-UA (+2S) | 56.1 | 75.2 | 62.7 | 58.2 | 65.6 | 61.3 | 63.2 | 64.4 |
| GlcN-UA-GlcN-UA-GlcNAc (+3 to+5S) | 8.9 | 2.5 | 3.7 | 5.2 | 4.2 | 6.1 | 5.0 | 3.4 |
| GlcN-[UA-GlcN]/[UA-GlcNAc]-UA (+3 to +5S) | 28.5 | 16.4 | 27.9 | 30.4 | 22.8 | 27.7 | 25.6 | 28.0 |
2.3. Correlation between HS Oligosaccharides and UA in MPS IIIA Mouse Tissues
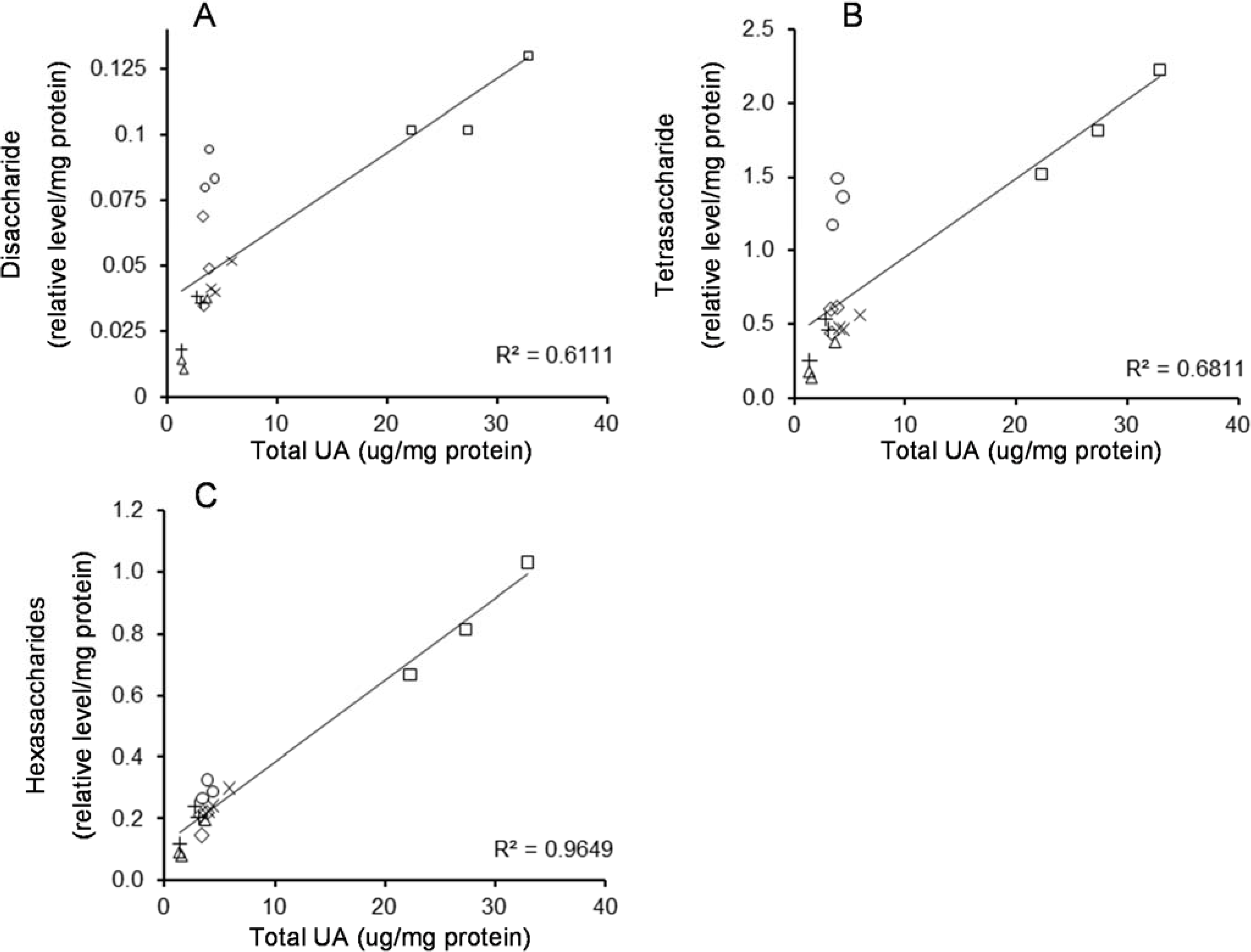
3. Experimental Section
3.1. Materials
3.2. Animals
3.3. Isolation of Oligosaccharides from Tissues and Urine
3.4. Total UA in Tissues and Urine
3.5. Derivatization of Oligosaccharides
3.6. HPLC ESI-MS/MS Analysis of di- to Hexasaccharides
3.7. Statistical Analysis
4. Discussion
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Neufeld, E.F.; Muenzer, J. The Mucopolysaccharidoses. In The Metabolic and Molecular Bases of Inherited Disease, 8th ed.; Scriver, C.R., Beaudet, A.L., Sly, W.S., Valle, D., Eds.; McGraw-Hill: New York, NY, USA, 2001; pp. 3421–3452. [Google Scholar]
- Blumenkrantz, N.; Asboe-Hansen, G. New method for quantitative determination of uronic acids. Anal. Biochem. 1973, 54, 484–489. [Google Scholar] [CrossRef]
- Roberts, A.L.K.; Thomas, B.J.; Wilkinson, A.S.; Fletcher, J.M.; Byers, S. Inhibition of glycosaminoglycan synthesis using rhodamine B in a mouse model of mucopolysaccharidosis type IIIA. Pediatr. Res. 2006, 60, 309–314. [Google Scholar] [CrossRef] [PubMed]
- Anson, D.S.; McIntyre, C.; Thomas, B.; Koldej, R.; Ranieri, E.; Roberts, A.; Clements, P.R.; Dunning, K.; Byers, S. Lentiviral-mediated gene correction of mucopolysaccharidosis type IIIA. Genet. Vaccines Therap. 2007. [Google Scholar] [CrossRef]
- McIntyre, C.; Roberts, A.L.D.; Ranieri, E.; Clements, P.R.; Byers, S.; Anson, D.S. Lentirviral-mediated gene therapy for murine mucopolysaccharidosis type IIIA. Mol. Genet. MeTab. 2008, 93, 411–418. [Google Scholar] [CrossRef] [PubMed]
- Ramsay, S.L.; Meikle, P.J.; Hopwood, J.J. Determination of monosaccharides and disaccharides in mucopolysaccharidoses patients by electrospray ionisation mass spectrometry. Mol. Genet. MeTab. 2003, 78, 193–204. [Google Scholar] [CrossRef]
- Fuller, M.; Rozaklis, T.; Ramsay, S.L.; Hopwood, J.J.; Meikle, P.J. Disease-Specific Markers for the Mucopolysaccharidoses. Pediatr. Res. 2004, 56, 733–738. [Google Scholar] [CrossRef] [PubMed]
- Fuller, M.; Meikle, P.J.; Hopwood, J.J. Glycosaminoglycan degradation fragments in mucopolysaccharidosis I. Glycobiology 2004, 14, 443–450. [Google Scholar] [CrossRef] [PubMed]
- Fuller, M.; Brooks, D.A.; Evangelista, M.; Hein, L.K.; Hopwood, J.J.; Meikle, P.J. Prediction of neuropathology in mucopolysaccharidosis I patients. Mol. Genet. MeTab. 2005, 84, 18–24. [Google Scholar] [CrossRef] [PubMed]
- Fuller, M.; Chau, A.; Nowak, R.C.; Hopwood, J.J.; Meikle, P.J. A defect in exo-degradative pathways provides insight into endo-degradation of heparan and dermatan sulfates. Glycobiology 2006, 16, 318–325. [Google Scholar] [CrossRef] [PubMed]
- Neilsen, T.C.; Rozek, T.; Hopwood, J.J.; Fuller, M. Determination of urinary oligosaccharides by high-performance liquid chromatography/electrospray ionization-tandem mass spectrometry: Application to Hunter syndrome. Anal. Biochem. 2010, 402, 113–120. [Google Scholar] [CrossRef] [PubMed]
- Oguma, T.; Tomatsu, S.; Montano, A.M.; Ozazaki, O. Analytical method for the determination of disaccharides derived from keratan, heparan and dermatan sulfates in human serum and plasma by high-performance liquid chromatograpy/turbo ionspray ionization tandem mass spectrometry. Anal. Biochem. 2007, 368, 79–86. [Google Scholar] [CrossRef] [PubMed]
- Tomatsu, S.; Montano, A.M.; Oguma, T.; Dung, V.C.; Oikawa, H.; Gutierrez, M.L.; Yamaguchi, S.; Suzuki, Y.; Fukushi, M.; Barrera, L.A.; et al. Validation of disaccharide compositions derived from dermatan sulfate and heparan sulfate in mucopolysaccharidosis and mucolipidoses II and III by tandem mass spectrometry. Mol. Genet. MeTab. 2010, 99, 124–131. [Google Scholar] [CrossRef] [PubMed]
- King, B.; Savas, P.; Fuller, M.; Hopwood, J.J.; Hemsley, K. Validation of a heparan sulfate-derived disaccharide as a marker of accumulation in murine mucopolysaccharidosis type IIIA. Mol. Genet. MeTab. 2006, 87, 107–112. [Google Scholar] [CrossRef] [PubMed]
- Savas, P.; Hemsley, K.M.; Hopwood, J.J. Intracerebral injection of sulfamidase delays neuropathology in murine MPS-IIIA. Mol. Genet. MeTab. 2004, 82, 273–285. [Google Scholar] [CrossRef] [PubMed]
- Hemsley, K.M.; Hopwood, J.J. Development of motor deficits in a murine model of mucopolysaccharidosis type IIIA (MPS-IIIA). Behav. Brain. Res. 2005, 158, 191–199. [Google Scholar] [CrossRef] [PubMed]
- Hemsley, K.M.; King, B.; Hopwood, J.J. Injection of recombinant human sulfamidase into the CSF via the cerebellomedullary cistern in MPS IIIA mice. Mol. Genet. MeTab. 2006, 90, 313–328. [Google Scholar] [CrossRef] [PubMed]
- Hemsley, K.M.; Beard, H.; King, B.M.; Hopwood, J.J. Effect of high dose, repeated intra-CSF injection of sulphamidase on neuropathology in MPS IIIA mice. Genes. Brain. Behav. 2008, 7, 740–753. [Google Scholar] [CrossRef]
- Hemsley, K.M.; Luck, A.J.; Crawley, A.C.; Hassiotis, S.; Beard, H.; King, B.; Rozek, T.; Rozaklis, T.; Fuller, M.; Hopwood, J.J. Examination of intravenous and intra-CSF protein delivery for treatment of neurological disease. Eur. J. Neurosci. 2009, 29, 1197–1214. [Google Scholar] [CrossRef]
- Mason, K.E.; Meikle, P.J.; Hopwood, J.J.; Fuller, M. Characterization of sulfated oligosaccharides in mucopolysaccharidosis type IIIA by electrospray ionization mass spectrometry. Anal. Chem. 2006, 78, 4534–4542. [Google Scholar] [CrossRef] [PubMed]
- Nakajima, M.; Irimura, T.; di Ferrante, N.; Nicolson, G.L. Metastatic melanoma cell heparanase. Characterization of heparan sulfate degradation fragments produced by B16 melanoma endoglucuronidase. J. Biol. Chem. 1984, 259, 2283–2290. [Google Scholar] [PubMed]
- Bame, K.J.; Robson, K. Heparanases produce distinct populations of heparan sulfate glycosaminoglycans in Chinese hamster ovary cells. J. Biol. Chem. 1997, 272, 2245–2251. [Google Scholar] [CrossRef] [PubMed]
- Hulett, M.D.; Freeman, C.; Hamdorf, B.J.; Baker, R.T.; Harris, M.J.; Parish, C.R. Cloning of mammalian heparanase, an important enzyme in tumor invasion and metastasis. Nat. Med. 1999, 5, 803–809. [Google Scholar] [CrossRef] [PubMed]
- Okada, Y.; Yamada, S.; Toyoshima, M.; Dong, J.; Nakajima, M.; Sugahara, K. Stuctural recognition by recombinant human heparanase that plays critical roles in tumor metastasis. J. Biol. Chem. 2002, 277, 42488–42495. [Google Scholar] [CrossRef] [PubMed]
- Bame, K.J. Heparanases: Endoglycosidases that degrade heparan sulfate proteoglycans. Glycobiology 2001, 11, 91R–98R. [Google Scholar] [CrossRef] [PubMed]
- Koliopanos, A.; Friess, H.; Kleeff, J.; Shi, X.; Liao, Q.; Pecker, I.; Vlodavsky, I.; Zimmermann, A.; Buchler, M.W. Heparanase expression in primary and metastatic pancreatic cancer. Cancer Res. 2001, 61, 4655–4659. [Google Scholar] [PubMed]
- Varchalama, E.; Rodolakis, A.; Strati, A.; Papageorgiou, T.; Valavanis, C.; Vorgias, G.; Lianidou, E.; Antsaklis, A. Quantitative analysis of heparanse gene expression in normal cervical, cervical intraepithelial neoplastic, and cervical carcinoma tissues. Int. J. Gynecol. Cancer 2009, 19, 1614–1619. [Google Scholar] [CrossRef] [PubMed]
- Levidiotis, V.; Freeman, C.; Tikellis, C.; Cooper, M.E.; Power, D.A. Heparanase is involved in the pathogenesis of proteinuria as a result of glomerulonephritis. J. Am. Soc. Nephrol. 2004, 15, 68–78. [Google Scholar] [CrossRef] [PubMed]
- Parish, C.R.; Freeman, C.; Hulett, M.D. Heparanase: A key enzyme involved in cell invasion. Biophys. Acta. 2001, 1471, M99–M108. [Google Scholar]
- Naparstek, Y.; Cohen, I.R.; Fuks, Z.; Vlodavsky, I. Activated T lymphocytes produce a matrix-degrading heparan sulphate endoglycosidase. Nature 1984, 310, 241–244. [Google Scholar] [CrossRef] [PubMed]
- Arfi, A.; Richard, M.; Gandolphe, C.; Bonnefont-Rousselot, D.; Thérond, P.; Schermann, D. Neuroinflammatory and oxidative stress phenomena in MPS IIIA mouse model: the positive effect of long-term aspirin treatment. Mol. Genet. MeTab. 2011, 103, 18–25. [Google Scholar] [CrossRef] [PubMed]
- Bhaumik, M.; Muller, V.J.; Rozaklis, T.; Johnson, L.; Dobrenis, K.; Bhattacharyya, R.; Wurzelmann, S.; Finamore, P.; Hopwood, J.J.; Walkley, S.U.; et al. A mouse model for mucopolysaccharidosis type III A (Sanfilippo syndrome). Glycobiology 1999, 9, 1389–1396. [Google Scholar] [PubMed]
- Li, H.H.; Yu, W.H.; Rozengurt, N.; Zhao, H.Z.; Lyons, K.M.; Anagnostaras, S.; Fanselow, M.S.; Suzuki, K.; Vanier, M.T.; Neufeld, E.F. Mouse model of Sanfilippo syndrome type B produced by targeted disruption of the gene encoding alpha-N-acetylglucosaminidase. PNAS 1999, 96, 14505–14510. [Google Scholar] [CrossRef] [PubMed]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mason, K.; Meikle, P.; Hopwood, J.; Fuller, M. Distribution of Heparan Sulfate Oligosaccharides in Murine Mucopolysaccharidosis Type IIIA. Metabolites 2014, 4, 1088-1100. https://doi.org/10.3390/metabo4041088
Mason K, Meikle P, Hopwood J, Fuller M. Distribution of Heparan Sulfate Oligosaccharides in Murine Mucopolysaccharidosis Type IIIA. Metabolites. 2014; 4(4):1088-1100. https://doi.org/10.3390/metabo4041088
Chicago/Turabian StyleMason, Kerryn, Peter Meikle, John Hopwood, and Maria Fuller. 2014. "Distribution of Heparan Sulfate Oligosaccharides in Murine Mucopolysaccharidosis Type IIIA" Metabolites 4, no. 4: 1088-1100. https://doi.org/10.3390/metabo4041088
APA StyleMason, K., Meikle, P., Hopwood, J., & Fuller, M. (2014). Distribution of Heparan Sulfate Oligosaccharides in Murine Mucopolysaccharidosis Type IIIA. Metabolites, 4(4), 1088-1100. https://doi.org/10.3390/metabo4041088






