The Integration of Metabolomics with Other Omics: Insights into Understanding Prostate Cancer
Abstract
1. Introduction
2. Metabolomics: The “Supra-Omic”
3. Integration of Metabolomics to Other Omic Platforms
4. Why Focus on Metabolomics for PCa Cancer Research?
5. Why Merge Metabolomics with Other Omics in PCa?
6. Clinical Applications of Metabolomics in PCa
7. Metabolomic Tools
8. Metabolomics and Genomics
| Reference | Experimental Condition | Sample/ n Samples | Analytical Tool for Metabolites | Altered Metabolites (+/−) | Dysregulated Metabolic Pathways | Main Findings |
|---|---|---|---|---|---|---|
| Hsu et al., 2021 [144] | Arginine starvation | Cell lines: CWR22Rv1, PC3, MDA-MB-231 | LC-MS Seahorse flux analysis | Arginine metabolites (−) α-ketoglutarate (−) | Oxidative phosphorylation DNA repair pathway Type I interferon response | Deficiency in arginine synthesis (defects in PCa), performed as arginine starvation resulted in cell death via epigenetic silencing and metabolite depletion. cGAS-STING activation contributed to cell death. |
| Cai et al., 2020 [217] | Citrate synthase (CS) down- regulation | 71 = adenocarcinoma 2 = leiomyo-sarcoma 1 = hyperplasia 6 = normal | UPHPLC-MS/MS Seahorse assay | Glyceraldehyde 3-phosphate (−) Citrate (−) | Lipid metabolism Mitochondrial function | CS expression: PCa > normal prostate. Decreased CS expression resulted in inhibited PCa proliferation, colony formation, migration, invasion, cell cycle in vitro, and low tumor growth in vivo. CS downregulation lowers lipid metabolism and mitochondrial function. |
| Kim et al., 2020 [145] | Withaferin (WA) treatment | 22Rv1 LNCaP, 22Rv1 (validation) Hi-MYC | Fluorometric assay | ATP citrase lyase, acetyl-coA carboxylase 1, fatty acid synthase, carnitine palmitoyltransferase (−) | Fatty acid synthesis | WA treatment in all cell lines downregulated mRNA and protein levels of key fatty acid synthesis enzymes. Suppression of a acetyl-coA carboxylase, expression of fatty acid synthase, and PCa cell survival from WA treatment → expression of c-MYC, not AKT. |
| Adams et al., 2018 [146] | Metabolite-PCa causality | 24,925 = GWAS metabolites 44,825 = GWAS PCa 27,904 control | Data mining and statistical analysis, no experimental tool | Lipids and lipoproteins Fatty acids and ratios Amino acids Fluids 35 metabolites association w/ PCa, 14 has no causality | Lipid metabolism Fatty acid metabolism Amino acid metabolism | 35 metabolites were associated w/ PCa, and 14 of those were found not to have causality w/ PCa progression. |
| Khodayari-Moez et al., 2018 [136] | AKT and MYC dysregulation | 60 = human PCa samples 16 = normal prostate | Data analysis, no experimental tool | Metabolites related to dysregulated metabolic pathways | D-glutamine and D-glutamate metabolism Fatty acid biosynthesis Fructose and mannose Metabolism Nitrogen metabolism Pyrimidine metabolism | Dysregulation of AKT1 and MYC alters non-glucose-mediated pathways and their downstream targets. MYC is one of the leading oncogenes in PCa development. |
| Heger et al., 2016 [128] | Sarcosine dehydro- genase (SDH) supplementation | PC3, LNCaP PCa murine xenograft (validation) | IEC | Glycine, serine, sarcosine (+) dimethylglycine and glycine-N-methyltransferase (slight +) | Sarcosine metabolism | SDH supplementation significantly increased levels of glycine, serine, and sarcosine, but slight increase in dimethylglycine and glycine-N-methyltransferase levels. PC-3 → 25, LNCaP → 32, overlapping → 18 differentially expressed genes. |
| Liu et al., 2015 [137] | Gene-metabolite association | 16 = benign 12 = PCa 14 = metasta- sized | Mathematical, no experimental tool, second-hand LC/GC-MS from Sreekumar et al. | 1353 genes 1489 metabolites | Non-applicable | Directed random walk global gene-metabolite graph (DRW-GM) = from integrated matched gene and matched metabolomic profiles →accurate evaluation of gene importance and pathway activities in PCa. Use of method in three independent datasets → accurate evaluation of risk pathways. |
| Shafi et al., 2015 [186] | Androgen receptor variant 7 (AR-V7) | LNCaP | Seahorse assay LC-MS | Glucose/fructose (−) 3-phosphoglycerate, 2-phosphoglycerate (−) Pyruvate (+) Citrate (−) α-ketoglutarate (+) Malate (−) Oxaloacetate (+) Glutamine (+) Citrate (−) | Glycolysis via extracellular acidification rate (ECAR) Glutamine metabolism via reductive carboxylation Tricarboxylic acid (TCA) cycle Glutaminolysis | AR-V7 stimulated growth, migration, and glycolysis measured by ECAR (extracellular acidification rate) similar to AR. AR → increase citrate, AR-V7 → reduce citrate mirroring metabolic shifts (castration-resistant PCa). AR-V7 is highly dependent on glutaminolysis and reductive carboxylation → produce metabolites consumed by TCA cycle. |
| Gilbert et al., 2014 [218] | SNPs of vitamin D-PCa association | 1275 = PCa 2062 = healthy controls | MS | 25-hydroxyvitamin-D (25(OH)D) 1,25-dihydroxyvitamin, (1,25(OH)2D) | 25(OH)D synthesis 25(OH)D metabolism | Vitamin D-binding protein SNPs were associated with prostate cancer. Low 25(OH)D metabolism score was associated with high grade. |
| Zecchini et al., 2014 [219] | Beta-arrestin 1 (ARB1) | C4-2 786-O | 1,2-13C2 glucose assay GC-MS | Succinate dehydrogenase Fumarate hydratase | Oxidative phosphorylation Aerobic glycolysis | ARB1 contributes to PCa metabolic shift via regulation of hypoxia-inducible factor 1A (HIF1A) transcription through regulation of succinate dehydrogenase and fumarate hydratase in normoxic conditions. ARB1 was directly linked in PCa as a promoter by altering metabolic pathways. Survival of PCa cells in harsh conditions due to ARB1. |
| Hong et al., 2013 [220] | Metabolic quantitative trait loci (mQTLs) via genome-wide association study (GWAS) | 214 = PCa 188 = control 489 = PCa (replication) | UPLC-MS w/ XCMS | Caprolactam Glycerolphosphocholine 2,6-dimethylheptanoylcarnitine Glycerolphosphocholine Bilirubin C9H14Ona Glycerophospho-N-palmitoyl ethanolamine Stearoylcarnitine Glycochenodeoxycholic acid 3-glucuronide | Fatty acid β-oxidation via acyl-CoA dehydrogenase | Seven genes (PYROXD2, FADS1, PON1, CYP4F2, UGT1A8, ACADL, and LIPC) and their variants contributed significantly to trait variance for one or more metabolites. Enrichment of 6 genes was associated w/ increased ACAD activity. mQTL SNPs and mQTL-harboring genes over-represented in GWAS → implications in PCa. |
| Poisson et al., 2012 [221] | Gene expression mapping | 402 = original 488 = replication | Statistical and mathematical, no experimental tool | Non-applicable | Non-applicable | Convert gene information to p-value weight via 4 enrichment tests and 4 weight functions. Used p weights on PCa metabolomic dataset. Disjoint pathways → higher capability to differentiate metabolites than enriched pathways. |
| Lu et al., 2011 [222] | Single-minded homolog 2 (SIM2) expression | PC3 LNCaP VCaP DU145 | LC-MS-MS | 38 dysregulated metabolites | PTEN signaling PI3K/AKT signaling Toll-like receptor signaling | Lenti-shRNA in PC3 → downregulates SIM2 gene and protein → affects key signaling and metabolic pathways. |
| Massie et al., 2011 [223] | AR regulatory effects | LNCaP | NMR 1,2-13C2 glucose assay GC-MS | Calcium/calmodulin-dependent protein kinase kinase 2 (CAMKK2) | Glycolysis via activating 5’ AMP-activated protein kinase (AMPK)- phosphofructokinase (PFK) signaling | AR regulates aerobic glycolysis and anabolism in PCa. CAMKK2, a direct AR target gene, regulates downstream metabolic processes. CAMKK2 is important in androgen-dependent and castration-resistant PCa. |
9. Metabolomics and Transcriptomics
| Reference | Experimental Condition | Sample/ n Samples | Analytical Tool for Metabolites | Altered Metabolites (+/−) | Dysregulated Metabolic Pathways | Main Findings |
|---|---|---|---|---|---|---|
| Imir et al., 2021 [147] | Perfluoroalkyl sulfonate (PFAS) exposure | RWPE-1 RWPE-kRAS | GC-MS | Acetyl-coA Pyruvate dehydrogenase complex (PDC) | Glycolysis via Warburg effect and transfer of acetyl group into mitochondria TCA cycle Threonine and 2-oxobutanoate degradation Phosphatidylethanol-amine biosynthesis Lysine degradation Pentose phosphate pathway (PPP) | PFAS exposure led to increase in xenograft tumor growth and altered metabolic phenotype of PCa, particularly those associated w/ glucose metabolism via the Warburg effect, involving the transfer of acetyl groups into mitochondria and TCA (pyruvate). PFAS increased PPAR signaling and histone acetylation in PCa. |
| Tilborg and Saccenti 2021 [224] | Gene expression-metabolic dysregulation relationships | 14 metabolic data sets, one of those is for PCa. 7 = tissue PCa 7 = tissue normal | Statistical, no experimental tool | Out of 72 metabolites investigated in PCa, 0 significantly differentially abundant metabolites were found (padj < 0.05) | No enriched or dysregulated pathways for PCa | Topological analysis of Gaussian networks → PCa more defined by genetic networks than metabolic ones. PCa-related metabolites were not significantly altered between controls and PCa samples. |
| Wang et al., 2021 [225] | Differential metabolites between PCa and BHP | 41 = PCa 38 = BPH | GC-MS GC/Q-TOF-MS Multivariate and univariate statistical analysis | 12 metabolites (+/−) including L-serine, myo-inositol, and decanoic acid | L-serine, myo-inositol, and decanoic acid metabolism | L-serine, myo-inositol, and decanoic acid → potential biomarkers for discriminating PCa from BHP. The 3 metabolites → increased area under the curve (AUC) of cPSA and tPSA from 0.542 and 0.592 to 0.781, respectively. |
| Gómez-Cebrián et al., 2020 [226] | Dysregulated PCa metabolic pathway mapping | 73 using serum and urine | NMR | 36 metabolites (+/−) including glucose, glycine, 1-methylnicotinamide | Energy metabolism Nucleotide synthesis | 36 metabolic pathways were dysregulated in PCa based on Gleason score (GS) (low-GS (GS < 7), high-GS PCa (GS ≥ 7) groups). Levels of glucose, glycine, and 1-methylnicotinamide → significantly altered between Gleason groups. |
| Chen et al., 2020 [148] | EMT-PCa and epithelial PCa differentiation | ARCaPE ARCaPM | LC-MS Glucose uptake assay | Aspartate (+) Glycolytic enzymes (+) except for glucose 2 transporter (−) TCA cycle: pyruvate dehydrogenase kinase 1/2, pyruvate dehydrogenase 2 (+) Succinate dehydrogenase A, aconitase 2 (−) Glutaminase 1/2 (+) | Glucose uptake Aspartate metabolism Glycolysis TCA cycle Glutamine–glutamate conversion | PCa cells undergoing epithelial-mesenchymal transition (EMT) showed low glucose consumption. Glucose metabolism in ARCaPE downregulated. Glucose metabolism in transcription factor- (TF) induced EMT models downregulated. ARCaPM cells showed increased aspartate metabolism. |
| Joshi et al., 2020 [149] | Carnitine palmitoyl transferase I (CPT1A) expression | LNCaP-C4-2 | UPHLC-MS | Acyl-carnitines Mitochondrial reactive oxygen species Superoxide dismutase 2 | ER stress Serine biosynthesis Lipid catabolism Androgen response | Upregulated pathways via transcriptomic analysis → ER stress, serine biosynthesis, lipid catabolism. Overexpressed (OE) of CPT1A showed increased SOD2 when subjected to low fatty acids and no androgen → better antioxidant defense w/ CPT1A OE. High lipid metabolism, low androgen response → worse progression-free survival. |
| Lee et al., 2020 [162] | Urine-enriched mRNA characteriza-tion | Urine: 20 = BPH 11 = PTT 20 = PCa 20 = normal 65 = PCa (validation) | UHPLC-HRMS | Alanine, aspartate, and glutamate (+) Glutamic-oxaloacetic transaminase 1 (+) | 14 metabolic pathways including aminoacyl-tRNA biosynthesis TCA cycle Pyruvate metabolism Amino acid pathways | Integrated gene expression-metabolite signature analysis → glutamate metabolism and TCA aberration contributed to PCa phenotype via GOT1-mediated redox balance. |
| Marin de Mas et al., 2019 [150] | Aldrin exposure analysis via gene-protein-reactions (GPR) associations | DU145 | Dataset processing, no experimental tool | 19 metabolites, both consuming and producing | Carnitine shuttle Prostaglandin biosynthesis | The application of novel stoichiometric gene–protein reaction (S-GPR) (imbedded in genome-scale metabolic models, GSMM) on the transcriptomic data of Aldrin-exposed DU145 PCa revealed increased metabolite use/production. Carnitine shuttle and prostaglandin biosynthesis → significantly altered in Aldrin-exposed DU145 PCa. |
| Andersen et al., 2018 [227] | Differential genes and metabolites | 158 tissue samples from 43 patients | HR-MAS MRS | 23 metabolites differentially expressed between high RSG and low RSG, including spermine, taurine, scyllo-inositol, and citrate | Immunity and ECM remodeling DNA repair pathway Type I interferon signaling | High RSG (≥16%) was associated w/ PCa biochemical recurrence (BCR). These high reactive stromata → upregulated genes and metabolites involved in immune functions and ECM remodeling. |
| Shao et al., 2018 [228] | Metabolomics-RNA-seq analysis | Tissue: 21 = PCa 21 = normal 50 = PCa and normal each (validation) | GC-MS | Fumarate Malate Branched-chain amino acid (+) Glutaminase, glutamate dehydrogenase ½ (+) Pyruvate dehydrogenase (+) | TCA cycle BCAA degradation Glutamine catabolism Pyruvate catabolism | Fumarate and malate levels → highly correlated w/ Gleason score, tumor stage, and expression of genes involved in BCAA degradation. BCAA degradation, glutamine catabolism, and pyruvate catabolism replenished TCA cycle metabolites. |
| Al Khadi et al., 2017 [229] | Peripheral and transitional zone differentiation | 20 PCa patients undergoing prostatectomy | Network-based integrative analysis, no experimental tool | 23 metabolites (+) including fatty acid synthase (FC = 2.9) and ELOVL fatty acid elongase 2 (FC = 2.8) | 15 KEGG pathways including de novo lipogenesis and fatty acid β-oxidation | RNA sequencing and high-throughput metabolic analyses (non-cancerous tissue, prostatectomy patients) → genes involved in de novo lipogenesis: peripheral > transitional. Peripheral zone induced lipo-rich priming → PCa oncogenesis. |
| Sandsmark et al., 2017 [230] | CWP, NCWP, EMT evaluation | 129 1519 samples (validation) | HR-MAS MRS MRSI | Citrate (−) Spermine (−) | TCA cycle | Increased NCWP activation via Wnt5a/Fzd2 Wnt activation mode → common in PCa. NCWP activation is associated w/ high EMT expression and high Gleason score. NCWP-EMT → significant predictor of PCa metastasis and biochemical recurrence. |
| Ren et al., 2016 [231] | Paired approach for altered pathways determination | 25 = PCa and adjacent non-cancerous tissues each 51 = PCa and 16 = BHP (validation) | LC-MS TOF-MS | Sphingosine (+) Sphingosine-1-phosphate receptor 2 (−) Choline, S-adenosylhomoserine, 5- methylthioadensine, S-adenosylmethionine, Nicotinamide mononucleotide, Nicotinamide adenine dinucleotide, and Nicotinamide adenine dinucleotide phosphate (+) Adenosine, uric acid (−) | Cysteine metabolism Methionine metabolism Nicotinamide adenine dinucleotide metabolism Hexosamine biosynthesis | Cysteine, methionine, and nicotinamide adenine dinucleotide metabolisms and hexosamine biosynthesis were aberrantly altered in PCT vs. ANT. Sphingosine was able to distinguish PCa from BHP cells for patients w/ low PSA levels. The loss of sphingosine-1-phosphate receptor 2 signaling → loss of TSG (oncogenic pathway). |
| Torrano et al., 2016 [232] | Peroxisome proliferator-activated receptor gamma coactivator 1-alpha (PGC1α) assessment | 150 = PCa 29 = control LNCaP DU145 PC3 | LCHR-MS Stable isotope 13C-U6-glucose labeling |
PGC1α (−) PGC1β Histone deacetylase 1 |
PGC1α pathway Estrogen-related receptor α (ERRα) pathway | PGC1α was a co-regulator and inhibits PCa progression and metastasis. Its deletion in murine prostate epithelium confirmed the finding. PGC1α dictates PCa oncogenic metabolic wiring, and its tumor-suppressive ability was mediated by the ERRα pathway. |
| Zhang et al., 2016 [233] | Angelica gigas Nakai (AGN) evaluation | 5 mice per group | UHPLC-MS-MS | 11 metabolites (+) including glutathione disulfide and taurine 11 metabolites (−) including lysine, tyrosine, and lactate | Methionine-cysteine metabolism Purine metabolism Citrate metabolism | Dosing w/ AGN → detectable decursinol, little decursin decursinol angelate. |
| Cerasuolo et al., 2015 [234] | Neuro- Endocrine transdifferen-tiation | LNCaP | H-NMR, Mathematical modeling | Creatinine + phosphor-creatinine (+) Glycine (+) Proline (+) Alanine (+) Fatty acids (+) Phospholipids (+) Glutathione (+) Glutamine (+) | Glucose oxidation Arginine and proline metabolism Glycine, serine, and threonine metabolism Glutamine and glutamate metabolism Glutathione metabolism | Hormone-deprived LNCaP cells were transdifferentiated to non-malignant neuroendocrine phenotype. Initially, LNCaP cells dwindled, neuroendocrine-type cells proliferated → later, neuroendocrine-type cells sustained LNCaP cells making them androgen-independent. |
| Meller et al., 2015 [235] | Metabolites analysis | 106 = PCa | GC-MS LC-MS MRM | Malignant vs. non-malignant: 156 metabolites (+) 17 metabolites (−) Gleason score: 11 metabolites (+) 4 metabolites (−) ERG translocation: 53 metabolites (+) 17 metabolites (−) | Fatty acid β-oxidation Sphingolipids metabolism Polyamines metabolism Cholesterol metabolism | Fatty acid β-oxidation and sphingolipids metabolism were dysregulated in PCa relative to non-malignant tumors. TMPRSS-ERG translocated was positively correlated (causality) w/ metabolites from PCa samples. Advanced PCA tumors exhibited increased cholesterol metabolism → energy storage. |
10. Metabolomics and Proteomics
| Reference | Experimental Condition | Sample/ n Samples | Analytical Tool for Metabolites | Altered Metabolites (+/−) | Dysregulated Metabolic Pathways | Main Findings |
|---|---|---|---|---|---|---|
| Kopylov et al., 2021 [239] | Schizophrenia-PCa association | 52 = PCa | Q-TOF MS UPLC | Cer(d18:1/14:0) 3Cholesta-3,5-dien-7-one 1α,25-dihydroxy-19-nor-22-oxavitamin D312:0 Cholesteryl ester24-hydroxy-cholesterol11-cis-RetinolElaidolinoleic acid14-hydroxy palmitic acid12-amino-dodecanoic acidL-Leucine | Sphingolipid metabolism 3 CholestanoidSteroid biosynthesisSteroid biosynthesis Bile acid biosynthesis Retinol metabolism Linoleic acid metabolismFatty acid biosynthesisFatty acid biosynthesis Valine, leucine and isoleucine degradation | Proteomic and metabolic data → input to approach employing systems biology and one-dimensional convolutional neural network (1DCNN) machine learning. Systems biology + 1DCNN → efficiently discriminate between: Unrelated pathologies = 0.90 (SCZ and oncophenotypes) Oncophenotypes/gender specific diseases = 0.93 (PCa). 1DCNN → high efficiency in PCa diagnosis. |
| Shen et al., 2021 [240] | Laser-capture-micro-dissection (LCM) androgen quantification | 16 = PCa | LC-SRM-MS | Androsterone 4 Androstenedione Dehydroepiandrosterone Testosterone | Interleukin signaling 4 IGF signaling NOTCH4 signaling Wnt signaling PDGF signaling Steroid metabolism ECM signaling, RAF/MAPK signaling by integrins | Coupled parallel LC-MS-based global proteomics and targeted metabolomics → ultrasensitive and robust quantification of androgen from low sample quantity. LC-MS-based method → robust and reliable protein quantification in LCM, including highly accurate profiling of stroma and epithelial LCM of PCa patients. |
| Teng et al., 2021 [151] | Mast cell (MC) and cancer-associated fibroblasts (CAF) profiling | PCa tissue from prostatectomy patients BPH-1 HMC-1 | SAMD14 (+) 5 | Immune signaling ECM processes | Transcriptomic profiling of MCs isolated from prostate tumor region → downregulated SAMD14. Proteomic profiling of HMC-1 → overexpression of SAMD14 → modified proteins associated w/ immune regulation and ECM processes. Add HMC-1-SAMD14+ medium to culture of (CAF + prostate epithelium) → reduced deposition and alignment of ECM generated by CAF; suppressed tumorigenic morphology of prostate epithelium. | |
| Blomme et al., 2020 [152] | Androgen receptor inhibitor (ARI)-based LNCaP characterization | LNCaP WT 6 LNCaP bicalut-res LNCaP apalut-res LNCaP enzalut-res | LTQ-OVMS FT-MS QEO-MS LC-MS | Metabolites associated w/ glucose metabolism (citrate, acetyl-coA) and lipid metabolism (+) for DECR1 overexpression Dihydroxyacetone phosphate and glycerol 3-phosphate (−) for DECR1 knockout | Glucose metabolism Fatty acid β-oxidation | 2,4-dienoyl-coA reductase (DECR1) knockout → induced ER stress, and stimulated CRPC cells to undergo ferroptosis. DECR1 deletion in vivo → inhibited lipid metabolism, and reduced CRPC tumor growth. |
| Felgueiras et al., 2020 [238] | PCa-normal prostate differentiation | Tissue: 8 = PCa 8 = normal | FT-IR | Polysaccharide and glycogen (−) Nucleic acid (+) | Lipid metabolism Protein phosphorylation | FT-IR (spectroscopic profiling) and antibody microarray (signaling proteins) → dysregulation in lipid metabolism and increased protein phosphorylation. |
| Li et al., 2020 [153] | FUN14-domain-containing protein-1 (FUNDC1) silencing | PC3 DU145 C42B | LC-MS UPHLC | AAA+ protease LonP1 Complex V (ATP synthase) TCA intermediates: pyruvate, cis-aconitase, α-ketoglutarate, succinate (−) Glutathione, ROS (+) | TCA cycle Oxidative phosphorylation | FUNDC1 affects cellular plasticity via sustaining oxidative phosphorylation, buffering ROS generation, and supporting cell proliferation. FUNDC1 expression → facilitated LonP1 proteostasis → preserved complex V function and decreased ROS generation. |
| Dougan et al., 2019 [154] | Peroxidasin (PXDN) knockdown | RWPE1 DU145 PC3 22Rv1 LNCaP | LC-MS-MS | Metabolites that prevent oxidative stress and promote nucleotide biosynthesis (−) (i.e., desirable to increase oxidative stress and decrease nucleotide biosynthesis → apoptosis of PCa cells) | Oxidative stress response Phagosome maturation Eukaryotic initiation factor 2 (eIF2) signaling Mitochondrial bioenergetics Gluconeogenesis I | Increased PXDN expression positively correlated w/ PCa progression. PXDN knockdown → increased oxidative stress and decreased nucleotide synthesis. PXDN knockdown → increased ROS → decreased cell viability, increased apoptosis. PXDN knockdown → decreased colony formation. |
11. Integrated Omic Analysis
| Reference | Experimental Condition | Sample/ n Samples | Analytical Tool | Altered Metabolites (+/−) | dysregulated Metabolic Pathways | Combined Modality/Main Findings |
|---|---|---|---|---|---|---|
| Kiebish et al., 2020 [100] | PCa prognostic markers identification | 382 pre-surgical serum samples from PCa patients 267 = training set (validation) 115 = testing set (validation) | MS-MS HILC-MS LC-MS GC-TOF-MS | 1-methyladenosine (+) | Cholesterol metabolism | Proteomics + Lipidomics + Metabolomics: Linear regression + Bayesian method + multi-omics → Tenascin C (TNC) and Apolipoprotein A1V (Apo-AIV), 1-Methyladenosine (1-MA), and phosphatidic acid (PA) 18:0–22:0, AUC = 0.78 (OR (95% CI) = 6.56 (2.98–14.40), P < 0.05) → high differentiating ability w/ and w/o BCR. |
| Oberhuber et al., 2020 [241] | Signal transducer and activator of transcription 3 (STAT3) expression | 84 = PCa from prostatectomy patients | LC-MS-MS LC-HRMS | Pyruvate dehydrogenase kinase 4 (+) | Oxidative phosphorylation TCA cycle Pyruvate oxidation | Transcriptomics + Proteomics + Metabolomics: High STAT3 expression → OXPHOS downregulated (Transcriptomics). High STAT3 expression → TCA cycle/OXPHOS downregulated (Proteomics). High PDK4 expression → inhibited PCa tumor growth. |
| Itkonen et al., 2019 [242] | Cyclin-dependent kinase 9 (CDK9) inhibition | LNCaP PC3 | Seahorse metabolic flux analysis | Acyl-carnitines (+) | Oxidative phosphorylation ATP synthesis AMP-activated protein kinase (AMPK) phosphorylation | Lipidomics + Fluxomics + Metabolomics: CDK9 inhibition → acute metabolic stress in PCa cells. CDK9 inhibition → downregulated oxidative phosphorylation, ATP depletion, and sustained AMPK phosphorylation. CDK9 inhibition → increased levels of acyl-carnitines |
| Gao et al., 2019 [243] | LASCPC-01 and LNCaP differentiation | LASCPC-01 LNCaP | GC-TOF-MS LC-MS | 25 metabolites altered from control Carnitine (−) | Glycolysis One-carbon metabolism | Transcriptomics + Lipidomics + Metabolomics: 62 genes upregulated in LSCPC-01, 112 genes upregulated in LNCaP (Transcriptomics). 25 genes significantly altered from control (Lipidomics + Metabolomics). LASCPC-01: high glycolytic rate, low-level triglycerides. LNCaP: high 1C metabolism rate, low carnitine. |
| Kregel et al., 2019 [244] | Bromodomain/ extraterminal (BET)- containing proteins (BRD2/3/4) inhibitor analysis | 22RV1 LNCaP VCaP PC3 DU145 | LC-MS | Polyunsaturated fatty acids (+) Thioredoxin-interacting protein Interferon regulatory transcription factor (−) | Cyclin-dependent kinase 9 inhibition CDK9 hyperphosporylation Polycomb repressive complex 2 activity | Proteomics + Lipidomics + Metabolomics: BET inhibitors: affected AR+ PCa (22RV1, LNCaP, VCaP) more than AR- PCa (PC3, DU145). BET inhibitors → disrupted AR and MYC signaling at concentrations: (BET) < (BET inhibitors) (Proteomics). |
| Zadra et al., 2019 [245] | Fatty acid synthase (FASN) suppression via IPI-9119 | LNCaP 22RV1 HeK293T RWPE-1 | UPLC-MS-MS LC-MS GC-MS 14C-labeling | 91 of the 418 metabolites modulated Malonyl-coA carnitine (+) Carnitine palmitoyltransferase 1 (−) | De novo fatty acid synthesis and neutral lipid accumulation ER stress response signaling Amino acid synthesis TCA cycle Carbohydrate metabolism Nucleotide metabolism | Lipidomics + Metabolomics: IPI-9119, a selective inhibitor of FASN altered the PCa metabolome by inhibiting fatty acid oxidation via accumulating malonyl-coA carnitine. Malonyl-coA carnitine accumulation → inhibited carnitine palmitoyltransferase 1 → FAO suppression. FA synthesis suppression → inhibited AR and AR-V7 expression. IPI-9119 → induced ER stress, inhibited AR/AR-V7 translation. |
| Murphy et al., 2018 [246] | PCa biomarker identification | 158 = PCa prostatectomy patients | LC-MS-MS Statistical modeling | 13 glycosylation metabolites (+) including tetraantennary tetrasialylated structures and A3G3S3 | Glycosylation | Genomics + Transcriptomics + Proteomics +Lipidomics + Metabolomics: Integration of data across 5 omic platforms from tissue and serum → single AUC value that better differentiates aggressive PCa from the indolent type compared to AUCs obtained from single omics. |
| Hansen et al., 2016 [247] | TMPRSS2-ERG expression | 129 = PCa samples from 41 patients 40 = PCa samples from 40 patients | HR-MAS-MRSI | Out of 23 metabolites, citrate and spermine (−) | TCA cycle Nucleic acid synthesis Citrate metabolism Polyamines metabolism | Transcriptomics + Metabolomics: ERGhigh = low citrate and spermine concentrations → increased PCa aggressiveness (Metabolomics). Metabolomic alterations for ERGhigh vs. ERGlow → more pronounced in low Gleason samples → implication: potential risk stratification tool. |
12. Metabolomic Profile of Prostate Cancer
12.1. Glycolysis
12.2. OXPHOS via the TCA cycle
12.3. De Novo Lipogenesis
12.4. Glycogenesis/Glycogenolysis
12.5. Pentose Phosphate Pathway
12.6. Amino Acid Metabolism
13. Conclusions and Future Perspectives
Author Contributions
Funding
Conflicts of Interest
References
- Trock, B.J. Application of metabolomics to prostate cancer. Urol. Oncol. Semin. Orig. Investig. 2011, 29, 572–581. [Google Scholar] [CrossRef] [PubMed]
- Nagana, G.G.; Raftery, D. Biomarker discovery and translation in metabolomics. Curr. Metab. 2013, 1, 227–240. [Google Scholar] [CrossRef] [PubMed]
- Patti, G.J.; Yanes, O.; Siuzdak, G. Metabolomics: The apogee of the omics trilogy. Nat. Rev. Mol. Cell Biol. 2012, 13, 263–269. [Google Scholar] [CrossRef] [PubMed]
- Johnson, C.H.; Ivanisevic, J.; Siuzdak, G. Metabolomics: Beyond biomarkers and towards mechanisms. Nat. Rev. Mol. Cell Biol. 2016, 17, 451–459. [Google Scholar] [CrossRef]
- Aderemi, A.V.; Ayeleso, A.O.; Oyedapo, O.O.; Mukwevho, E. Metabolomics: A Scoping Review of Its Role as a Tool for Disease Biomarker Discovery in Selected Non-Communicable Diseases. Metabolites 2021, 11, 418. [Google Scholar] [CrossRef]
- Aboud, O.A.; Weiss, R.H. New opportunities from the cancer metabolome. Clin. Chem. 2013, 59, 138–146. [Google Scholar] [CrossRef]
- European Bioinformatics Institute (EMBL-EBI). The Metabolome and Metabolic Reactions. Available online: https://www.ebi.ac.uk/training/online/courses/metabolomics-introduction/the-metabolome-and-metabolic-reactions/ (accessed on 12 January 2022).
- Weiss, R.H.; Kim, K. Metabolomics in the study of kidney diseases. Nat. Rev. Nephrol. 2012, 8, 22–33. [Google Scholar] [CrossRef]
- Lai, Z.; Kind, T.; Fiehn, O. Using accurate mass gas chromatography–mass spectrometry with the MINE database for epi-metabolite annotation. Anal. Chem. 2017, 89, 10171–10180. [Google Scholar] [CrossRef]
- Showalter, M.R.; Cajka, T.; Fiehn, O. Epimetabolites: Discovering metabolism beyond building and burning. Curr. Opin. Chem. Biol. 2017, 36, 70–76. [Google Scholar] [CrossRef]
- Mamani-Huanca, M.; Gradillas, A.; Gil de la Fuente, A.; López-Gonzálvez, Á.; Barbas, C. Unveiling the fragmentation mechanisms of modified amino acids as the key for their targeted identification. Anal. Chem. 2020, 92, 4848–4857. [Google Scholar] [CrossRef]
- Prodhan, M.A.I.; McClain, C.; Zhang, X. Comprehensive two-dimensional gas chromatography mass spectrometry-based metabolomics. In Cancer Metabolomics: Methods and Applications; Hu, S., Ed.; Springer Nature Switzerland AG: Cham, Switzerland, 2021; Volume 1280, pp. 57–67. [Google Scholar]
- Dueñas, M.E.; Lee, Y.J. Single-Cell Metabolomics by Mass Spectrometry Imaging. In Cancer Metabolomics: Methods and Applications; Hu, S., Ed.; Springer Nature Switzerland AG: Cham, Switzerland, 2021; Volume 1280, pp. 69–82. [Google Scholar]
- Liu, R.; Yang, Z. Single cell metabolomics using mass spectrometry: Techniques and data analysis. Anal. Chim. Acta 2020, 1143, 124–134. [Google Scholar] [CrossRef]
- Lin, X.; Lécuyer, L.; Liu, X.; Triba, M.; Deschasaux-Tanguy, M.; Demidem, A.; Liu, Z.; Palama, T.; Rossary, A.; Vasson, M.-P.; et al. Plasma Metabolomics for Discovery of Early Metabolic Markers of Prostate Cancer Based on Ultra-High-Performance Liquid Chromatography-High Resolution Mass Spectrometry. Cancers 2021, 13, 3140. [Google Scholar] [CrossRef]
- Xu, X. Capillary electrophoresis-mass spectrometry for cancer metabolomics. In Cancer Metabolomics: Methods and Applications; Hu, S., Ed.; Springer Nature Switzerland AG: Cham, Switzerland, 2021; Volume 1280, pp. 189–200. [Google Scholar]
- Zhang, X.; Wei, D.; Yap, Y.; Li, L.; Guo, S.; Chen, F. Mass spectrometry-based “omics” technologies in cancer diagnostics. Mass Spectrom. Rev. 2007, 26, 403–431. [Google Scholar] [CrossRef]
- Ganti, S.; Taylor, S.; Abu Aboud, O.; Yang, J.; Evans, C.; Osier, M.V.; Alexander, D.C.; Kim, K.; Weiss, R.H. Kidney Tumor Biomarkers Revealed by Simultaneous Multiple Matrix Metabolomics Analysis. Cancer Res. 2012, 72, 3471–3479. [Google Scholar] [CrossRef]
- López-López, Á.; López-Gonzálvez, Á.; Barker-Tejeda, T.C.; Barbas, C. A review of validated biomarkers obtained through metabolomics. Expert Rev. Mol. Diagn. 2018, 18, 557–575. [Google Scholar] [CrossRef]
- Kuehnbaum, N.L.; Gillen, J.B.; Kormendi, A.; Lam, K.P.; Dibattista, A.; Gibala, M.J.; Britz-McKibbin, P. Multiplexed separations for biomarker discovery in metabolomics: Elucidating adaptive responses to exercise training. Electrophoresis 2015, 36, 2226–2236. [Google Scholar] [CrossRef]
- Wellington, N.; Shanmuganathan, M.; de Souza, R.J.; Zulyniak, M.A.; Azab, S.; Bloomfield, J.; Mell, A.; Ly, R.; Desai, D.; Anand, S.S.; et al. Metabolic Trajectories Following Contrasting Prudent and Western Diets from Food Provisions: Identifying Robust Biomarkers of Short-Term Changes in Habitual Diet. Nutrients 2019, 11, 2407. [Google Scholar] [CrossRef]
- Rattray, N.J.W.; DeZiel, N.C.; Wallach, J.D.; Khan, S.A.; Vasiliou, V.; Ioannidis, J.P.A.; Johnson, C.H. Beyond genomics: Understanding exposotypes through metabolomics. Hum. Genom. 2018, 12, 4. [Google Scholar] [CrossRef]
- Gao, W.; Sun, H.-X.; Xiao, H.; Cui, G.; Hillwig, M.L.; Jackson, A.; Wang, X.; Shen, Y.; Zhao, N.; Zhang, L.; et al. Combining metabolomics and transcriptomics to characterize tanshinone biosynthesis in Salvia miltiorrhiza. BMC Genom. 2014, 15, 73. [Google Scholar] [CrossRef]
- Schmidt, D.R.; Patel, R.; Kirsch, D.G.; Lewis, C.A.; Vander Heiden, M.G.; Locasale, J.W. Metabolomics in cancer research and emerging applications in clinical oncology. CA Cancer J. Clin. 2021, 71, 333–358. [Google Scholar] [CrossRef]
- Wilson, I.D.; Nicholson, J.K.; Castro-Perez, J.; Granger, J.H.; Johnson, K.A.; Smith, A.B.W.; Plumb, R.S. High Resolution “Ultra Performance” Liquid Chromatography Coupled to oa-TOF Mass Spectrometry as a Tool for Differential Metabolic Pathway Profiling in Functional Genomic Studies. J. Proteome Res. 2005, 4, 591–598. [Google Scholar] [CrossRef]
- BioRender.com. Available online: https://biorender.com/ (accessed on 12 January 2022).
- Griffin, J.L.; Shockcor, J.P. Metabolic profiles of cancer cells. Nat. Cancer 2004, 4, 551–561. [Google Scholar] [CrossRef]
- Cheung, P.K.; Ma, M.H.; Tse, H.F.; Yeung, K.F.; Tsang, H.F.; Chu, M.K.M.; Kan, C.M.; Cho, W.C.S.; Ng, L.B.W.; Chan, L.W.C.; et al. The applications of metabolomics in the molecular diagnostics of cancer. Expert Rev. Mol. Diagn. 2019, 19, 785–793. [Google Scholar] [CrossRef]
- Pinto, R.C. Chemometrics methods and strategies in metabolomics. In Metabolomics: From Fundamentals to Clinical Applications; Sussulini, A., Ed.; Springer International Publishing AG: Cham, Switzerland, 2017; Volume 965, pp. 163–190. [Google Scholar]
- Vermeersch, K.A.; Styczynski, M.P. Applications of metabolomics in cancer research. J. Carcinog. 2013, 12, 9. [Google Scholar] [CrossRef] [PubMed]
- Pang, H.; Jia, W.; Hu, Z. Emerging Applications of Metabolomics in Clinical Pharmacology. Clin. Pharmacol. Ther. 2019, 106, 544–556. [Google Scholar] [CrossRef] [PubMed]
- Dona, A.C.; Kyriakides, M.; Scott, F.; Shephard, E.; Varshavi, D.; Veselkov, K.; Everett, J.R. A guide to the identification of metabolites in NMR-based metabonomics/metabolomics experiments. Comput. Struct. Biotechnol. J. 2016, 14, 135–153. [Google Scholar] [CrossRef]
- Lu, X.; Solmonson, A.; Lodi, A.; Nowinski, S.M.; Sentandreu, E.; Riley, C.L.; Mills, E.M.; Tiziani, S. The early metabolomic response of adipose tissue during acute cold exposure in mice. Sci. Rep. 2017, 7, 3455. [Google Scholar] [CrossRef]
- Psychogios, N.; Hau, D.D.; Peng, J.; Guo, A.C.; Mandal, R.; Bouatra, S.; Sinelnikov, I.; Krishnamurthy, R.; Eisner, R.; Gautam, B.; et al. The Human Serum Metabolome. PLoS ONE 2011, 6, e16957. [Google Scholar] [CrossRef] [PubMed]
- German, J.B.; Hammock, B.D.; Watkins, S.M. Metabolomics: Building on a century of biochemistry to guide human health. Metabolomics 2005, 1, 3–9. [Google Scholar] [CrossRef]
- Wishart, D.S. Current Progress in computational metabolomics. Brief. Bioinform. 2007, 8, 279–293. [Google Scholar] [CrossRef] [PubMed]
- Xia, J.; Broadhurst, D.I.; Wilson, M.; Wishart, D.S. Translational biomarker discovery in clinical metabolomics: An intro-ductory tutorial. Metabolomics 2013, 9, 280–299. [Google Scholar] [CrossRef] [PubMed]
- Wishart, D.S. Emerging applications of metabolomics in drug discovery and precision medicine. Nat. Rev. Drug Discov. 2016, 15, 473–484. [Google Scholar] [CrossRef] [PubMed]
- Wishart, D.S.; Mandal, R.; Stanislaus, A.; Ramirez-Gaona, M. Cancer Metabolomics and the Human Metabolome Database. Metabolites 2016, 6, 10. [Google Scholar] [CrossRef] [PubMed]
- National Institute of Health-National Human Genome Research Institute (NIH-NHGRI). The Human Genome Project. 2021. Available online: https://www.genome.gov/human-genome-project (accessed on 12 January 2022).
- Willyard, C. New human gene tally reignites debate. Nature 2018, 558, 354–355. [Google Scholar] [CrossRef]
- Yu, L.; Li, K.; Zhang, X. Next-generation metabolomics in lung cancer diagnosis, treatment and precision medicine: Mini review. Oncotarget 2017, 8, 115774–115786. [Google Scholar] [CrossRef]
- Milo, R.; Jorgensen, P.; Moran, U.; Weber, G.; Springer, M. BioNumbers—the database of key numbers in molecular and cell biology. Nucleic Acids Res. 2009, 38, D750–D753. [Google Scholar] [CrossRef]
- Zhang, H.; Egger, R.L.; Kelliher, T.; Morrow, D.; Fernandes, J.; Nan, G.-L.; Walbot, V. Transcriptomes and Proteomes Define Gene Expression Progression in Pre-meiotic Maize Anthers. G3 Genes Genomes Genet. 2014, 4, 993–1010. [Google Scholar] [CrossRef][Green Version]
- Zhang, J.; White, N.M.; Schmidt, H.K.; Fulton, R.S.; Tomlinson, C.; Warren, W.C.; Wilson, R.K.; Maher, C.A. INTEGRATE: Gene fusion discovery using whole genome and transcriptome data. Genome Res. 2015, 26, 108–118. [Google Scholar] [CrossRef]
- Mertins, P.; Cptac, N.; Mani, D.R.; Ruggles, K.V.; Gillette, M.A.; Clauser, K.R.; Wang, P.; Wang, X.; Qiao, J.W.; Cao, S.; et al. Proteogenomics connects somatic mutations to signalling in breast cancer. Nature 2016, 534, 55–62. [Google Scholar] [CrossRef]
- Dang, L.; Yen, K.; Attar, E.C. IDH mutations in cancer and progress toward development of targeted therapeutics. Ann. Oncol. 2016, 27, 599–608. [Google Scholar] [CrossRef]
- Pirozzi, C.J.; Yan, H. The implications of IDH mutations for cancer development and therapy. Nat. Rev. Clin. Oncol. 2021, 18, 645–661. [Google Scholar] [CrossRef] [PubMed]
- Schrimpe-Rutledge, A.C.; Codreanu, S.G.; Sherrod, S.D.; McLean, J.A. Untargeted Metabolomics Strategies—Challenges and Emerging Directions. J. Am. Soc. Mass Spectrom. 2016, 27, 1897–1905. [Google Scholar] [CrossRef] [PubMed]
- Cerrato, A.; Bedia, C.; Capriotti, A.L.; Cavaliere, C.; Gentile, V.; Maggi, M.; Montone, C.M.; Piovesana, S.; Sciarra, A.; Tauler, R.; et al. Untargeted metabolomics of prostate cancer zwitterionic and positively charged compounds in urine. Anal. Chim. Acta 2021, 1158, 338381. [Google Scholar] [CrossRef] [PubMed]
- Qi, S.-A.; Wu, Q.; Chen, Z.; Zhang, W.; Zhou, Y.; Mao, K.; Li, J.; Li, Y.; Chen, J.; Huang, Y.; et al. High-resolution metabolomic biomarkers for lung cancer diagnosis and prognosis. Sci. Rep. 2021, 11, 11805. [Google Scholar] [CrossRef]
- Jobard, E.; Dossus, L.; Baglietto, L.; Fornili, M.; Lécuyer, L.; Mancini, F.R.; Gunter, M.J.; Trédan, O.; Boutron-Ruault, M.-C.; Elena-Herrmann, B.; et al. Investigation of circulating metabolites associated with breast cancer risk by untargeted metabolomics: A case–control study nested within the French E3N cohort. Br. J. Cancer 2021, 124, 1734–1743. [Google Scholar] [CrossRef]
- Huan, T.; Forsberg, E.M.; Rinehart, D.; Johnson, C.H.; Ivanisevic, J.; Benton, H.P.; Fang, M.; Aisporna, A.; Hilmers, B.; Poole, F.L.; et al. Systems biology guided by XCMS Online metabolomics. Nat. Methods 2017, 14, 461–462. [Google Scholar] [CrossRef]
- Giera, M.; dos Santos, F.B.; Siuzdak, G. Metabolite-Induced Protein Expression Guided by Metabolomics and Systems Biology. Cell Metab. 2018, 27, 270–272. [Google Scholar] [CrossRef]
- Rosato, A.; Tenori, L.; Cascante, M.; De Atauri Carulla, P.R.; Dos Santos, V.A.P.M.; Saccenti, E. From correlation to causation: Analysis of metabolomics data using systems biology approaches. Metabolomics 2018, 14, 37. [Google Scholar] [CrossRef]
- Guijas, C.; Montenegro-Burke, J.R.; Warth, B.; Spilker, M.E.; Siuzdak, G. Metabolomics activity screening for identifying metabolites that modulate phenotype. Nat. Biotechnol. 2018, 36, 316–320. [Google Scholar] [CrossRef]
- Kell, D.B. Metabolomics and systems biology: Making sense of the soup. Curr. Opin. Microbiol. 2004, 7, 296–307. [Google Scholar] [CrossRef]
- Ruepp, S.U.; Tonge, R.P.; Shaw, J.; Wallis, N.; Pognan, F. Genomics and Proteomics Analysis of Acetaminophen Toxicity in Mouse Liver. Toxicol. Sci. 2002, 65, 135–150. [Google Scholar] [CrossRef]
- Nicholson, J.K.; Wilson, I.D.; Lindon, J.C. Pharmacometabonomics as an effector for personalized medicine. Pharmacogenomics 2011, 12, 103–111. [Google Scholar] [CrossRef]
- Dumas, M.-E.; Wilder, S.P.; Bihoreau, M.-T.; Barton, R.H.; Fearnside, J.F.; Argoud, K.; D’Amato, L.; Wallis, R.H.; Blancher, C.; Keun, H.C.; et al. Direct quantitative trait locus mapping of mammalian metabolic phenotypes in diabetic and normoglycemic rat models. Nat. Genet. 2007, 39, 666–672. [Google Scholar] [CrossRef]
- Golubnitschaja, O.; Costigliola, V.; EPMA. General Report & Recommendations in Predictive, Preventive and Personalised Medicine 2012: White Paper of the European Association for Predictive, Preventive and Personalised Medicine. EPMA J. 2012, 3, 14–53. [Google Scholar] [CrossRef]
- Golubnitschaja, O.; Baban, B.; Boniolo, G.; Wang, W.; Bubnov, R.; Kapalla, M.; Krapfenbauer, K.; Mozaffari, M.S.; Costigliola, V. Medicine in the early twenty-first century: Paradigm and anticipation-EPMA position paper 2016. EPMA J. 2016, 7, 23. [Google Scholar] [CrossRef]
- Golubnitschaja, O.; Filep, N.; Yeghiazaryan, K.; Blom, H.J.; Hofmann-Apitius, M.; Kuhn, W. Multi-omic approach decodes paradoxes of the triple-negative breast cancer: Lessons for predictive, preventive and personalised medicine. Amino Acids 2017, 50, 383–395. [Google Scholar] [CrossRef]
- Ibrahim, R.; Pasic, M.; Yousef, G.M. Omics for personalized medicine: Defining the current we swim in. Expert Rev. Mol. Diagn. 2016, 16, 719–722. [Google Scholar] [CrossRef]
- Chen, R.; Snyder, M. Promise of personalized omics to precision medicine. Wiley Interdiscip. Rev. Syst. Biol. Med. 2013, 5, 73–82. [Google Scholar] [CrossRef]
- Turanli, B.; Yildirim, E.; Gulfidan, G.; Arga, K.Y.; Sinha, R. Current State of “Omics” Biomarkers in Pancreatic Cancer. J. Pers. Med. 2021, 11, 127. [Google Scholar] [CrossRef]
- Subbannayya, Y.; Di Fiore, R.; Urru, S.A.M.; Calleja-Agius, J. The Role of Omics Approaches to Characterize Molecular Mechanisms of Rare Ovarian Cancers: Recent Advances and Future Perspectives. Biomedicines 2021, 9, 1481. [Google Scholar] [CrossRef]
- Nicholson, J.K.; Holmes, E.; Lindon, J.; Wilson, I.D. The challenges of modeling mammalian biocomplexity. Nat. Biotechnol. 2004, 22, 1268–1274. [Google Scholar] [CrossRef]
- Suhre, K.; Shin, S.Y.; Petersen, A.K.; Mohney, R.P.; Meredith, D.; Wagele, B.; Altmaier, E.; Gram, C.; Deloukas, P.; Erdmann, J.; et al. Human metabolic individuality in biomedical and pharmaceutical research. Nature 2011, 477, 54–60. [Google Scholar] [CrossRef]
- Moskowitz, J.E.; Doran, A.G.; Lei, Z.; Busi, S.B.; Hart, M.L.; Franklin, C.L.; Sumner, L.W.; Keane, T.M.; Amos-Landgraf, J.M. Integration of genomics, metagenomics, and metabolomics to identify interplay between susceptibility alleles and microbiota in adenoma initiation. BMC Cancer 2020, 20, 600. [Google Scholar] [CrossRef]
- Quanbeck, S.M.M.; Brachova, L.; Campbell, A.A.; Guan, X.; Perera, A.; He, K.; Rhee, S.Y.; Bais, P.; Dickerson, J.A.; Dixon, P.; et al. Metabolomics as a Hypothesis-Generating Functional Genomics Tool for the Annotation of Arabidopsis thaliana Genes of “Unknown Function”. Front. Plant Sci. 2012, 3, 15. [Google Scholar] [CrossRef] [PubMed]
- Gieger, C.; Geistlinger, L.; Altmaier, E.; de Angelis, M.H.; Kronenberg, F.; Meitinger, T.; Mewes, H.-W.; Wichmann, H.-E.; Weinberger, K.; Adamski, J.; et al. Genetics Meets Metabolomics: A Genome-Wide Association Study of Metabolite Profiles in Human Serum. PLoS Genet. 2008, 4, e1000282. [Google Scholar] [CrossRef] [PubMed]
- Nicholson, G.; Rantalainen, M.; Li, J.; Maher, A.D.; Malmodin, D.; Ahmadi, K.R.; Faber, J.H.; Barrett, A.; Min, J.L.; Rayner, N.W.; et al. A Genome-Wide Metabolic QTL Analysis in Europeans Implicates Two Loci Shaped by Recent Positive Selection. PLoS Genet. 2011, 7, e1002270. [Google Scholar] [CrossRef] [PubMed]
- Thevis, M.; Schänzer, W. Synthetic Anabolic Agents: Steroids and Nonsteroidal Selective Androgen Receptor Modulators. In Doping in Sports: Biochemical Principles, Effects and Analysis. Handbook of Experimental Pharmacology; Thieme, D., Hemmersbach, P., Eds.; Springer: Berlin/Heidelberg, Germany, 2009; Volume 195, pp. 99–126. [Google Scholar]
- Rading, A.; Anielski, P.; Thieme, D.; Keiler, A.M. Detection of the selective androgen receptor modulator GSK2881078 and metabolites in urine and hair after single oral administration. Drug Test. Anal. 2020, 13, 217–222. [Google Scholar] [CrossRef]
- Le Bail, J.-C.; Aubourg, L.; Habrioux, G. Effects of pinostrobin on estrogen metabolism and estrogen receptor transactivation. Cancer Lett. 2000, 156, 37–44. [Google Scholar] [CrossRef]
- Wang, C.-Y.; Chiao, C.-C.; Phan, N.N.; Li, C.-Y.; Sun, Z.-D.; Jiang, J.-Z.; Hung, J.-H.; Chen, Y.-L.; Yen, M.-C.; Weng, T.-Y.; et al. Gene signatures and potential therapeutic targets of amino acid metabolism in estrogen receptor-positive breast cancer. Am. J. Cancer Res. 2020, 10, 95–113. [Google Scholar]
- Layton, A.C.; Sanseverino, J.; Gregory, B.W.; Easter, J.P.; Sayler, G.S.; Schultz, T.W. In Vitro estrogen receptor binding of PCBs: Measured Aativity and detection of hydroxylated metabolites in a recombinant yeast assay. Toxicol. Appl. Pharmacol. 2002, 180, 157–163. [Google Scholar] [CrossRef]
- Ren, Z.; Liu, Q.S.; Sun, Z.; Xu, H.; Zhou, Q.; Jiang, G. Perfluorinated iodine alkanes induce tissue-specific expression of estrogen receptor and its phosphorylation. Sci. Total Environ. 2021, 787, 147722. [Google Scholar] [CrossRef]
- Raut, S.; Kumar, A.V.; Deshpande, S.; Khambata, K.; Balasinor, N.H. Sex hormones regulate lipid metabolism in adult Sertoli cells: A genome-wide study of estrogen and androgen receptor binding sites. J. Steroid Biochem. Mol. Biol. 2021, 211, 105898. [Google Scholar] [CrossRef]
- Eidelman, E.; Twum-Ampofo, J.; Ansari, J.; Siddiqui, M.M. The Metabolic Phenotype of Prostate Cancer. Front. Oncol. 2017, 7, 131. [Google Scholar] [CrossRef]
- Ladurner, A.G. Rheostat Control of Gene Expression by Metabolites. Mol. Cell 2006, 24, 1–11. [Google Scholar] [CrossRef]
- Lempp, M.; Farke, N.; Kuntz, M.; Freibert, S.A.; Lill, R.; Link, H. Systematic identification of metabolites controlling gene expression in E. coli. Nat. Commun. 2019, 10, 4463. [Google Scholar] [CrossRef]
- Van der Knaap, J.A.; Verrijzer, C.P. Undercover: Gene control by metabolites and metabolic enzymes. Genes Dev. 2016, 30, 2345–2369. [Google Scholar] [CrossRef]
- Commichau, F.; Gunka, K.; Landmann, J.J.; Stülke, J. Glutamate Metabolism in Bacillus subtilis: Gene Expression and Enzyme Activities Evolved to Avoid Futile Cycles and To Allow Rapid Responses to Perturbations of the System. J. Bacteriol. 2008, 190, 3557–3564. [Google Scholar] [CrossRef]
- Donohoe, D.R.; Bultman, S.J. Metaboloepigenetics: Interrelationships between energy metabolism and epigenetic control of gene expression. J. Cell. Physiol. 2012, 227, 3169–3177. [Google Scholar] [CrossRef]
- Winkler, W.C.; Nahvi, A.; Roth, A.; Collins, J.A.; Breaker, R.R. Control of gene expression by a natural metabolite-responsive ribozyme. Nature 2004, 428, 281–286. [Google Scholar] [CrossRef]
- Diskin, C.; Ryan, T.A.J.; O’Neill, L.A.J. Modification of Proteins by Metabolites in Immunity. Immunity 2020, 54, 19–31. [Google Scholar] [CrossRef]
- Sovova, Z.; Suttnar, J.; Dyr, J.E. Molecular dynamic simulations suggest that metabolite-induced post-translational modifi-cations alter the behavior of the fibrinogen coiled-coil domain. Metabolites 2021, 11, 307. [Google Scholar] [CrossRef] [PubMed]
- Martín, J.F.; Liras, P.; Sánchez, S. Modulation of gene expression in actinobacteria by translational modification of tran-scriptional factors and secondary metabolite biosynthetic enzymes. Front. Microbiol. 2021, 12, 630694. [Google Scholar] [CrossRef] [PubMed]
- Harachi, M.; Masui, K.; Cavenee, W.K.; Mischel, P.S.; Shibata, N. Protein Acetylation at the Interface of Genetics, Epigenetics and Environment in Cancer. Metabolites 2021, 11, 216. [Google Scholar] [CrossRef] [PubMed]
- Qin, W.; Yang, F.; Wang, C. Chemoproteomic profiling of protein–metabolite interactions. Curr. Opin. Chem. Biol. 2020, 54, 28–36. [Google Scholar] [CrossRef]
- Wilkinson, D.J.; Hossain, T.; Hill, D.S.; Phillips, B.E.; Crossland, H.; Williams, J.; Atherton, P.J. Effects of leucine and its metabolite β-hydroxy-β-methylbutyrate on human skeletal muscle protein metabolism. J. Physiol. 2013, 591, 2911–2923. [Google Scholar] [CrossRef]
- Zhao, T.; Liu, J.; Zeng, X.; Wang, W.; Li, S.; Zang, T.; Peng, J.; Yang, Y. Prediction and collection of protein–metabolite interactions. Brief. Bioinform. 2021, 22. [Google Scholar] [CrossRef]
- Yang, Q.; Vijayakumar, A.; Kahn, B.B. Metabolites as regulators of insulin sensitivity and metabolism. Nat. Rev. Mol. Cell Biol. 2018, 19, 654–672. [Google Scholar] [CrossRef]
- Menzies, K.J.; Zhang, H.; Katsyuba, E.; Auwerx, J. Protein acetylation in metabolism—Metabolites and cofactors. Nat. Rev. Endocrinol. 2015, 12, 43–60. [Google Scholar] [CrossRef]
- Brunt, V.E.; LaRocca, T.J.; Bazzoni, A.E.; Sapinsley, Z.J.; Miyamoto-Ditmon, J.; Gioscia-Ryan, R.A.; Neilson, A.P.; Link, C.D.; Seals, D.R. The gut microbiome–derived metabolite trimethylamine N-oxide modulates neuroinflammation and cognitive function with aging. GeroScience 2021, 43, 377–394. [Google Scholar] [CrossRef]
- Yamanaka, S.; Murai, H.; Saito, D.; Abe, G.; Tokunaga, E.; Iwasaki, T.; Takahashi, H.; Takeda, H.; Suzuki, T.; Shibata, N.; et al. Thalidomide and its metabolite 5-hydroxythalidomide induce teratogenicity via the cereblon neosubstrate PLZF. EMBO J. 2021, 40, e105375. [Google Scholar] [CrossRef]
- Cheng, L.; Wu, S.; Zhang, K.; Qing, Y.; Xu, T. A comprehensive overview of exosomes in ovarian cancer: Emerging biomarkers and therapeutic strategies. J. Ovarian Res. 2017, 10, 73. [Google Scholar] [CrossRef]
- Kiebish, M.A.; Cullen, J.; Mishra, P.; Ali, A.; Milliman, E.; Rodrigues, L.O.; Chen, E.Y.; Tolstikov, V.; Zhang, L.; Panagopoulos, K.; et al. Multi-omic serum biomarkers for prognosis of disease progression in prostate cancer. J. Transl. Med. 2020, 18, 10. [Google Scholar] [CrossRef]
- Lu, M.; Zhan, X. The crucial role of multiomic approach in cancer research and clinically relevant outcomes. EPMA J. 2018, 9, 77–102. [Google Scholar] [CrossRef]
- Kane, L.E.; Mellotte, G.S.; Conlon, K.C.; Ryan, B.M.; Maher, S.G. Multi-omic biomarkers as potential tools for the char-acterisation of pancreatic cystic lesions and cancer: Innovative patient data integration. Cancers 2021, 13, 769. [Google Scholar] [CrossRef]
- Knowles, M.A.; Hurst, C.D. Molecular biology of bladder cancer: New insights into pathogenesis and clinical diversity. Nat. Rev. Cancer 2014, 15, 25–41. [Google Scholar] [CrossRef]
- Sehgal, V.; Seviour, E.; Moss, T.J.; Mills, G.B.; Azencott, R.; Ram, P.T. Robust Selection Algorithm (RSA) for Multi-Omic Biomarker Discovery; Integration with Functional Network Analysis to Identify miRNA Regulated Pathways in Multiple Cancers. PLoS ONE 2015, 10, e0140072. [Google Scholar] [CrossRef]
- Chu, S.H.; Huang, M.; Kelly, R.S.; Benedetti, E.; Siddiqui, J.K.; Zeleznik, O.A. Integration of metabolomic and other omics data in population-based study designs: An epidemiological per-spective. Metabolites 2019, 9, 117. [Google Scholar] [CrossRef]
- Stäubert, C.; Bhuiyan, H.; Lindahl, A.; Broom, O.J.; Zhu, Y.; Islam, S.; Linnarsson, S.; Lehtiö, J.; Nordström, A. Rewired Metabolism in Drug-resistant Leukemia Cells. J. Biol. Chem. 2015, 290, 8348–8359. [Google Scholar] [CrossRef]
- Wishart, D.S.; Guo, A.; Oler, E.; Wang, F.; Anjum, A.; Peters, H.; Dizon, R.; Sayeeda, Z.; Tian, S.; Lee, B.L.; et al. HMDB 5.0: The Human Metabolome Database for 2022. Nucleic Acids Res. 2022, 50, D622–D631. [Google Scholar] [CrossRef]
- Saoi, M.; Britz-McKibbin, P. New Advances in Tissue Metabolomics: A Review. Metabolites 2021, 11, 672. [Google Scholar] [CrossRef]
- Lima, A.R.; De Bastos, M.L.; Carvalho, M.; de Pinho, P.G. Biomarker Discovery in Human Prostate Cancer: An Update in Metabolomics Studies. Transl. Oncol. 2016, 9, 357–370. [Google Scholar] [CrossRef]
- Lima, A.; Pinto, J.; Amaro, F.; Bastos, M.; Carvalho, M.; de Pinho, P.G. Advances and Perspectives in Prostate Cancer Biomarker Discovery in the Last 5 Years through Tissue and Urine Metabolomics. Metabolites 2021, 11, 181. [Google Scholar] [CrossRef]
- Gómez-Cebrián, N.; Rojas-Benedicto, A.; Albors-Vaquer, A.; López-Guerrero, J.A.; Pineda-Lucena, A.; Puchades-Carrasco, L. Metabolomics Contributions to the Discovery of Prostate Cancer Biomarkers. Metabolites 2019, 9, 48. [Google Scholar] [CrossRef]
- Salciccia, S.; Capriotti, A.; Laganà, A.; Fais, S.; Logozzi, M.; De Berardinis, E.; Busetto, G.; Di Pierro, G.; Ricciuti, G.; Del Giudice, F.; et al. Biomarkers in Prostate Cancer Diagnosis: From Current Knowledge to the Role of Metabolomics and Exosomes. Int. J. Mol. Sci. 2021, 22, 4367. [Google Scholar] [CrossRef]
- Ahmad, F.; Cherukuri, M.K.; Choyke, P.L. Metabolic reprogramming in prostate cancer. Br. J. Cancer 2021, 125, 1185–1196. [Google Scholar] [CrossRef]
- Lucarelli, G.; Rutigliano, M.; Galleggiante, V.; Giglio, A.; Palazzo, S.; Ferro, M.; Simone, C.; Bettocchi, C.; Battaglia, M.; Ditonno, P. Metabolomic profiling for the identification of novel diagnostic markers in prostate cancer. Expert Rev. Mol. Diagn. 2015, 15, 1211–1224. [Google Scholar] [CrossRef]
- Clish, C.B. Metabolomics: An emerging but powerful tool for precision medicine. Mol. Case Stud. 2015, 1, a000588. [Google Scholar] [CrossRef]
- Kdadra, M.; Höckner, S.; Leung, H.; Kremer, W.; Schiffer, E. Metabolomics Biomarkers of Prostate Cancer: A Systematic Review. Diagnostics 2019, 9, 21. [Google Scholar] [CrossRef] [PubMed]
- Kelly, R.S.; Heiden, M.G.V.; Giovannucci, E.L.; Mucci, L.A. Metabolomic Biomarkers of Prostate Cancer: Prediction, Diagnosis, Progression, Prognosis, and Recurrence. Cancer Epidemiol. Biomark. Prev. 2016, 25, 887–906. [Google Scholar] [CrossRef] [PubMed]
- Beyoğlu, D.; Idle, J. Metabolic Rewiring and the Characterization of Oncometabolites. Cancers 2021, 13, 2900. [Google Scholar] [CrossRef] [PubMed]
- Franko, A.; Shao, Y.; Heni, M.; Hennenlotter, J.; Hoene, M.; Hu, C.; Liu, X.; Zhao, X.; Wang, Q.; Birkenfeld, A.L.; et al. Human Prostate Cancer is Characterized by an Increase in Urea Cycle Metabolites. Cancers 2020, 12, 1814. [Google Scholar] [CrossRef]
- Vykoukal, J.; Fahrmann, J.F.; Gregg, J.R.; Tang, Z.; Basourakos, S.; Irajizad, E.; Park, S.; Yang, G.; Creighton, C.J.; Fleury, A.; et al. Caveolin-1-mediated sphingolipid oncometabolism underlies a metabolic vulnerability of prostate cancer. Nat. Commun. 2020, 11, 4279. [Google Scholar] [CrossRef]
- Sreekumar, A.; Poisson, L.M.; Rajendiran, T.M.; Khan, A.P.; Cao, Q.; Yu, J.; Laxman, B.; Mehra, R.; Lonigro, R.J.; Li, Y.; et al. Metabolomic profiles delineate potential role for sarcosine in prostate cancer progression. Nature 2009, 457, 910–914. [Google Scholar] [CrossRef]
- Jentzmik, F. Sarcosine in urine after digital rectal examination fails as a marker in prostate cancer detection and identi-fication of aggressive tumours. Eur. Urol. 2010, 58, 12–18. [Google Scholar] [CrossRef]
- Cao, D.-L.; Ye, D.-W.; Zhu, Y.; Zhang, H.-L.; Wang, Y.-X.; Yao, X.-D. Efforts to resolve the contradictions in early diagnosis of prostate cancer: A comparison of different algorithms of sarcosine in urine. Prostate Cancer Prostatic Dis. 2011, 14, 166–172. [Google Scholar] [CrossRef]
- Yang, B.; Zhang, C.; Cheng, S.; Li, G.; Griebel, J.; Neuhaus, J. Novel Metabolic Signatures of Prostate Cancer Revealed by 1H-NMR Metabolomics of Urine. Diagnostics 2021, 11, 149. [Google Scholar] [CrossRef]
- Song, Y.H.; Shiota, M.; Kuroiwa, K.; Naito, S.; Oda, Y. The important role of glycine N-methyltransferase in the carcino-genesis and progression of prostate cancer. Mod. Pathol. 2011, 24, 1272–1280. [Google Scholar] [CrossRef]
- Ottaviani, S.; Brooke, G.N.; O’Hanlon-Brown, C.; Waxman, J.; Ali, S.; Buluwela, L. Characterisation of the androgen regulation of glycine N-methyltransferase in prostate cancer cells. J. Mol. Endocrinol. 2013, 51, 301–312. [Google Scholar] [CrossRef]
- Huang, Y.-C. Haplotypes, loss of heterozygosity, and expression levels of glycine N-methyltransferase in prostate cancer. Clin. Cancer Res. 2007, 13, 1412–1420. [Google Scholar] [CrossRef]
- Heger, Z.; Rodrigo, M.A.M.; Michalek, P.; Polanska, H.; Masarik, M.; Vit, V.; Plevova, M.; Pacik, D.; Eckschlager, T.; Stiborova, M.; et al. Sarcosine Up-Regulates Expression of Genes Involved in Cell Cycle Progression of Metastatic Models of Prostate Cancer. PLoS ONE 2016, 11, e0165830. [Google Scholar] [CrossRef]
- De Vogel, S. Sarcosine and other metabolites along the choline oxidation pathway in relation to prostate cancer-A large nested case-control study within the JANUS cohort in Norway. Int. J. Cancer 2014, 134, 197–206. [Google Scholar] [CrossRef]
- Yousefi, M.; Qujeq, D.; Shafi, H.; Tilaki, K.H. Serum and Urine Levels of Sarcosine in Benign Prostatic Hyperplasia and Newly Diagnosed Prostate Cancer Patients. J. Kermanshah Univ. Med Sci. 2020, 24, e97000. [Google Scholar] [CrossRef]
- Dehghani, N.; Salehipour, M.; Javanmard, B. Evaluation of GNMT Gene Expression in Prostate Cancer Tissues using Real-Time PCR. J. Tolooebehdasht 2021, 19, 44–54. [Google Scholar] [CrossRef]
- Jendoubi, T. Approaches to Integrating Metabolomics and Multi-Omics Data: A Primer. Metabolites 2021, 11, 184. [Google Scholar] [CrossRef]
- Haukaas, T.H.; Euceda, L.R.; Giskeødegård, G.F.; Bathen, T.F. Metabolic Portraits of Breast Cancer by HR MAS MR Spectroscopy of Intact Tissue Samples. Metabolites 2017, 7, 18. [Google Scholar] [CrossRef]
- Ryan, D.; Robards, K. Metabolomics: The greatest omics of them all? Anal. Chem. 2006, 78, 7954–7958. [Google Scholar] [CrossRef]
- Buescher, J.M.; Driggers, E.M. Integration of omics: More than the sum of its parts. Cancer Metab. 2016, 4, 4. [Google Scholar] [CrossRef]
- Moez, E.K.; Pyne, S.; Dinu, I. Association between bivariate expression of key oncogenes and metabolic phenotypes of patients with prostate cancer. Comput. Biol. Med. 2018, 103, 55–63. [Google Scholar] [CrossRef]
- Liu, W.; Bai, X.; Liu, Y.; Wang, W.; Han, J.; Wang, Q.; Xu, Y.; Zhang, C.; Zhang, S.; Li, X.; et al. Topologically inferring pathway activity toward precise cancer classification via integrating genomic and metabolomic data: Prostate cancer as a case. Sci. Rep. 2015, 5, 13192. [Google Scholar] [CrossRef] [PubMed]
- Del Carratore, F. Integrated probabilistic annotation: A Bayesian-based annotation method for metabolomic profiles in-tegrating biochemical connections, isotope patterns, and adduct relationships. Anal. Chem. 2019, 91, 12799–12807. [Google Scholar] [CrossRef] [PubMed]
- McNaughton, A.D.; Bredeweg, E.L.; Manzer, J.; Zucker, J.; Munoz, N.M.; Burnet, M.C.; Nakayasu, E.S.; Pomraning, K.R.; Merkley, E.D.; Dai, Z.; et al. Bayesian Inference for Integrating Yarrowia lipolytica Multiomics Datasets with Metabolic Modeling. ACS Synth. Biol. 2021, 10, 2968–2981. [Google Scholar] [CrossRef]
- Sen, P.; Lamichhane, S.; Mathema, V.B.; McGlinchey, A.; Dickens, A.M.; Khoomrung, S.; Orešič, M. Deep learning meets metabolomics: A methodological perspective. Briefings Bioinform. 2020, 22, 1531–1542. [Google Scholar] [CrossRef]
- Kang, M.; Ko, E.; Mersha, T.B. A roadmap for multi-omics data integration using deep learning. Brief. Bioinform. 2021, 23, bbab454. [Google Scholar] [CrossRef]
- Zheng, H.; Hu, Y.; Dong, L.; Shu, Q.; Zhu, M.; Li, Y.; Chen, C.; Gao, H.; Yang, L. Predictive diagnosis of chronic obstructive pulmonary disease using serum metabolic biomarkers and least-squares support vector machine. J. Clin. Lab. Anal. 2020, 35, e23641. [Google Scholar] [CrossRef]
- Gagnebin, Y.; Pezzatti, J.; Lescuyer, P.; Boccard, J.; Ponte, B.; Rudaz, S. Combining the advantages of multilevel and orthogonal partial least squares data analysis for longitudinal metabolomics: Application to kidney transplantation. Anal. Chim. Acta 2019, 1099, 26–38. [Google Scholar] [CrossRef]
- Hsu, S.-C. Arginine starvation elicits chromatin leakage and cGAS-STING activation via epigenetic silencing of metabolic and DNA-repair genes. Theranostics 2021, 11, 7527–7545. [Google Scholar] [CrossRef]
- Kim, S.-H.; Hahm, E.-R.; Singh, K.B.; Shiva, S.; Stewart-Ornstein, J.; Singh, S.V. RNA-seq reveals novel mechanistic targets of withaferin A in prostate cancer cells. Carcinogenesis 2020, 41, 778–789. [Google Scholar] [CrossRef]
- Adams, C.D.; Richmond, R.; Ferreira, D.L.S.; Spiller, W.; Tan, V.; Zheng, J.; Würtz, P.; Donovan, J.; Hamdy, F.; Neal, D.; et al. Circulating Metabolic Biomarkers of Screen-Detected Prostate Cancer in the ProtecT Study. Cancer Epidemiol. Biomark. Prev. 2018, 28, 208–216. [Google Scholar] [CrossRef]
- Imir, O.B.; Kaminsky, A.Z.; Zuo, Q.-Y.; Liu, Y.-J.; Singh, R.; Spinella, M.J.; Irudayaraj, J.; Hu, W.-Y.; Prins, G.S.; Erdogan, Z.M. Per- and Polyfluoroalkyl Substance Exposure Combined with High-Fat Diet Supports Prostate Cancer Progression. Nutrients 2021, 13, 3902. [Google Scholar] [CrossRef]
- Chen, Y. Decreased glucose bioavailability and elevated aspartate metabolism in prostate cancer cells undergoing epi-thelial-mesenchymal transition. J. Cell. Physiol. 2020, 235, 5602–5612. [Google Scholar] [CrossRef]
- Joshi, M.; Kim, J.; D’Alessandro, A.; Monk, E.; Bruce, K.; Elajaili, H.; Nozik-Grayck, E.; Goodspeed, A.; Costello, J.C.; Schlaepfer, I.R. CPT1A Over-Expression Increases Reactive Oxygen Species in the Mitochondria and Promotes Antioxidant Defenses in Prostate Cancer. Cancers 2020, 12, 3431. [Google Scholar] [CrossRef]
- De Mas, I.M.; Torrents, L.; Bedia, C.; Nielsen, L.K.; Cascante, M.; Tauler, R. Stoichiometric gene-to-reaction associations enhance model-driven analysis performance: Metabolic response to chronic exposure to Aldrin in prostate cancer. BMC Genom. 2019, 20, 652. [Google Scholar] [CrossRef]
- Teng, L.K.H.; Pereira, B.; Keerthikumar, S.; Huang, C.; Niranjan, B.; Lee, S.; Richards, M.; Schittenhelm, R.; Furic, L.; Goode, D.; et al. Mast Cell-Derived SAMD14 Is a Novel Regulator of the Human Prostate Tumor Microenvironment. Cancers 2021, 13, 1237. [Google Scholar] [CrossRef]
- Blomme, A. 2,4-dienoyl-CoA reductase regulates lipid homeostasis in treatment-resistant prostate cancer. Nat. Commun. 2020, 11, 2508. [Google Scholar] [CrossRef]
- Li, J.; Agarwal, E.; Bertolini, I.; Seo, J.H.; Caino, M.C.; Ghosh, J.C.; Kossenkov, A.V.; Liu, Q.; Tang, H.-Y.; Goldman, A.R.; et al. The mitophagy effector FUNDC1 controls mitochondrial reprogramming and cellular plasticity in cancer cells. Sci. Signal. 2020, 13, eaaz8240. [Google Scholar] [CrossRef]
- Dougan, J.; Hawsawi, O.; Burton, L.J.; Edwards, G.; Jones, K.; Zou, J.; Odero-Marah, V.A. Proteomics-metabolomics combined approach identifies peroxidasin as a protector against metabolic and oxidative stress in prostate cancer. Int. J. Mol. Sci. 2019, 20, 3046. [Google Scholar] [CrossRef]
- Peppicelli, S.; Andreucci, E.; Ruzzolini, J.; Bianchini, F.; Calorini, L. FDG uptake in cancer: A continuing debate. Theranostics 2020, 10, 2944–2948. [Google Scholar] [CrossRef]
- Haj-Ahmad, T.A.; Abdalla, M.A.; Haj-Ahmad, Y. Potential Urinary miRNA Biomarker Candidates for the Accurate Detection of Prostate Cancer among Benign Prostatic Hyperplasia Patients. J. Cancer 2014, 5, 182–191. [Google Scholar] [CrossRef] [PubMed]
- Robertson, M.S.; Sakellis, C.G.; Hyun, H.; Jacene, H.A. Extraprostatic Uptake of 18F-Fluciclovine: Differentiation of Nonprostatic Neoplasms From Metastatic Prostate Cancer. Am. J. Roentgenol. 2020, 214, 641–648. [Google Scholar] [CrossRef] [PubMed]
- Kairemo, K.; Rasulova, N.; Partanen, K.; Joensuu, T. Preliminary clinical experience of trans-1-amino-3-(18)F-fluorocyclobutanecarboxylic acid (anti-(18)F-FACBC) PET/CT imaging in prostate cancer patients. BioMed Res. Int. 2014, 2014, 305182. [Google Scholar] [CrossRef] [PubMed]
- Saarinen, I. Correlation between 18F-1-amino-3-fluorocyclobutane-1-carboxylic acid (18F-fluciclovine) uptake and ex-pression of alanine-serine-cysteine-transporter 2 (ASCT2) and L-type amino acid transporter 1 (LAT1) in primary prostate cancer. EJNMMI Res. 2019, 9, 50. [Google Scholar] [CrossRef]
- Lieu, E.L.; Nguyen, T.; Rhyne, S.; Kim, J. Amino acids in cancer. Exp. Mol. Med. 2020, 52, 15–30. [Google Scholar]
- Dereziński, P.; Klupczynska, A.; Sawicki, W.; Pałka, J.A.; Kokot, Z.J. Amino Acid Profiles of Serum and Urine in Search for Prostate Cancer Biomarkers: A Pilot Study. Int. J. Med. Sci. 2017, 14, 1–12. [Google Scholar] [CrossRef]
- Lee, B.; Mahmud, I.; Marchica, J.; Dereziński, P.; Qi, F.; Wang, F.; Joshi, P.; Valerio, F.; Rivera, I.; Patel, V.; et al. Integrated RNA and metabolite profiling of urine liquid biopsies for prostate cancer biomarker discovery. Sci. Rep. 2020, 10, 3716. [Google Scholar] [CrossRef]
- Taavitsainen, S.; Engedal, N.; Cao, S.; Handle, F.; Erickson, A.; Prekovic, S.; Wetterskog, D.; Tolonen, T.; Vuorinen, E.M.; Kiviaho, A.; et al. Single-cell ATAC and RNA sequencing reveal pre-existing and persistent cells associated with prostate cancer relapse. Nat. Commun. 2021, 12, 5307. [Google Scholar] [CrossRef]
- Markley, J.L.; Brüschweiler, R.; Edison, A.S.; Eghbalnia, H.R.; Powers, R.; Raftery, D.; Wishart, D.S. The future of NMR-based metabolomics. Curr. Opin. Biotechnol. 2017, 43, 34–40. [Google Scholar] [CrossRef]
- Nagana Gowda, G.A.; Raftery, D. Can NMR solve some significant challenges in metabolomics? J. Magn. Reson. 2015, 260, 144–160. [Google Scholar] [CrossRef]
- Beale, D.J.; Pinu, F.R.; Kouremenos, K.A.; Poojary, M.M.; Narayana, V.K.; Boughton, B.A.; Kanojia, K.; Dayalan, S.; Jones, O.A.H.; Dias, D.A. Review of recent developments in GC–MS approaches to metabolomics-based research. Metabolomics 2018, 14, 152. [Google Scholar] [CrossRef]
- Tikunov, Y.M.; Verstappen, F.W.A.; Hall, R.D. Metabolomic Profiling of Natural Volatiles: Headspace Trapping: GC-MS. In Metabolomics: Methods and Protocols; Weckwerth, W., Ed.; Humana Press Inc.: Totowa, NJ, USA, 2007; Volume 358, pp. 39–56. [Google Scholar]
- Zhou, B.; Xiao, J.F.; Tuli, L.; Ressom, H.W. LC-MS-based metabolomics. Mol. BioSyst. 2012, 8, 470–481. [Google Scholar] [CrossRef]
- Sands, C.J.; Gómez-Romero, M.; Correia, G.; Chekmeneva, E.; Camuzeaux, S.; Izzi-Engbeaya, C.; Dhillo, W.S.; Takats, Z.; Lewis, M.R. Representing the Metabolome with High Fidelity: Range and Response as Quality Control Factors in LC-MS-Based Global Profiling. Anal. Chem. 2021, 93, 1924–1933. [Google Scholar] [CrossRef]
- Lima, C.; Muhamadali, H.; Goodacre, R. The Role of Raman Spectroscopy Within Quantitative Metabolomics. Annu. Rev. Anal. Chem. 2021, 14, 323–345. [Google Scholar] [CrossRef]
- Su, K.-Y.; Lee, W.-L. Fourier transform infrared spectroscopy as a cancer screening and diagnostic tool: A review and pro-spects. Cancers 2020, 12, 115. [Google Scholar] [CrossRef]
- Ge, Y.; Sun, M.; Salomé-Abarca, L.F.; Wang, M.; Choi, Y.H. Investigation of species and environmental effects on rhubarb roots metabolome using 1H NMR combined with high performance thin layer chromatography. Metabolomics 2018, 14, 137. [Google Scholar] [CrossRef]
- Xie, G.; Wang, L.; Chen, T.; Zhou, K.; Zhang, Z.; Li, J.; Sun, B.; Guo, Y.; Wang, X.; Wang, Y.; et al. A Metabolite Array Technology for Precision Medicine. Anal. Chem. 2021, 93, 5709–5717. [Google Scholar] [CrossRef]
- Shi, Y.; Han, J.J.; Tennakoon, J.B.; Mehta, F.F.; Merchant, F.; Burns, A.R.; Howe, M.K.; McDonnell, D.P.; Frigo, D.E. Androgens Promote Prostate Cancer Cell Growth through Induction of Autophagy. Mol. Endocrinol. 2013, 27, 280–295. [Google Scholar] [CrossRef]
- Singh, R.; Mills, I.G. The Interplay Between Prostate Cancer Genomics, Metabolism, and the Epigenome: Perspectives and Future Prospects. Front. Oncol. 2021, 11, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Rubin, M.A.; Demichelis, F. The Genomics of Prostate Cancer: A Historic Perspective. Cold Spring Harb. Perspect. Med. 2019, 9, a034942. [Google Scholar] [CrossRef] [PubMed]
- Zhang, E.; Zhang, M.; Shi, C.; Sun, L.; Shan, L.; Zhang, H.; Song, Y. An overview of advances in multi-omics analysis in prostate cancer. Life Sci. 2020, 260, 118376. [Google Scholar] [CrossRef] [PubMed]
- Feldman, B.J.; Feldman, D. The development of androgen-independent prostate cancer. Nat. Rev. Cancer 2001, 1, 34–45. [Google Scholar] [CrossRef] [PubMed]
- Xie, T.; Song, X.-L.; Wang, C.; Yu, Y.-Z.; Wang, J.-Q.; Chen, Z.-S.; Zhao, S.-C. The role of androgen therapy in prostate cancer: From testosterone replacement therapy to bipolar androgen therapy. Drug Discov. Today 2021, 26, 1293–1301. [Google Scholar] [CrossRef]
- Jääskeläinen, J.; Mongan, N.P.; Harland, S.; Hughes, I.A. Five novel androgen receptor gene mutations associated with complete androgen insensitivity syndrome. Hum. Mutat. 2006, 27, 291. [Google Scholar] [CrossRef]
- Hornig, N.C.; Holterhus, P.-M. Molecular basis of androgen insensitivity syndromes. Mol. Cell. Endocrinol. 2021, 523, 111146. [Google Scholar] [CrossRef]
- Helsen, C.; Dubois, V.; Verfaillie, A.; Young, J.; Trekels, M.; Vancraenenbroeck, R.; De Maeyer, M.; Claessens, F. Evidence for DNA-Binding Domain–Ligand-Binding Domain Communications in the Androgen Receptor. Mol. Cell. Biol. 2012, 32, 3033–3043. [Google Scholar] [CrossRef]
- Xia, F.; Xu, X.; Zhai, H.; Meng, Y.; Zhang, H.; Du, S.; Xu, H.; Wu, H.; Lu, Y. Castration-induced testosterone deficiency increases fasting glucose associated with hepatic and extra-hepatic insulin resistance in adult male rats. Reprod. Biol. Endocrinol. 2013, 11, 106. [Google Scholar] [CrossRef]
- Di Sebastiano, K.M.; Pinthus, J.H.; Duivenvoorden, W.C.M.; Mourtzakis, M. Glucose impairments and insulin resistance in prostate cancer: The role of obesity, nutrition and exercise. Obes. Rev. 2018, 19, 1008–1016. [Google Scholar] [CrossRef]
- Shafi, A.A.; McNair, C.M.; McCann, J.J.; Alshalalfa, M.; Shostak, A.; Severson, T.M.; Zhu, Y.; Bergman, A.; Gordon, N.; Mandigo, A.C.; et al. The circadian cryptochrome, CRY1, is a pro-tumorigenic factor that rhythmically modulates DNA repair. Nat. Commun. 2021, 12, 401. [Google Scholar] [CrossRef]
- Shafi, A.A.; Putluri, V.; Arnold, J.; Tsouko, E.; Maity, S.; Roberts, J.M.; Coarfa, C.; Frigo, D.; Putluri, N.; Sreekumar, A.; et al. Differential regulation of metabolic pathways by androgen receptor (AR) and its constitutively active splice variant, AR-V7, in prostate cancer cells. Oncotarget 2015, 6, 31997–32012. [Google Scholar] [CrossRef]
- Audet-Walsh, É.; Dufour, C.R.; Yee, T.; Zouanat, F.Z.; Yan, M.; Kalloghlian, G.; Giguère, V. Nuclear mTOR acts as a transcriptional integrator of the androgen signaling pathway in prostate cancer. Genes Dev. 2017, 31, 1228–1242. [Google Scholar] [CrossRef]
- Twum-Ampofo, J.; Fu, D.-X.; Passaniti, A.; Hussain, A.; Siddiqui, M.M. Metabolic targets for potential prostate cancer therapeutics. Curr. Opin. Oncol. 2016, 28, 241–247. [Google Scholar] [CrossRef]
- Bader, D.A.; Hartig, S.M.; Putluri, V.; Foley, C.; Hamilton, M.P.; Smith, E.A.; Saha, P.K.; Panigrahi, A.; Walker, C.; Zong, L.; et al. Mitochondrial pyruvate import is a metabolic vulnerability in androgen receptor-driven prostate cancer. Nat. Metab. 2019, 1, 70–85. [Google Scholar] [CrossRef]
- Flaig, T.W.; Salzmann-Sullivan, M.; Su, L.-J.; Zhang, Z.; Joshi, M.; Gijón, M.A.; Kim, J.; Arcaroli, J.J.; Van Bokhoven, A.; Lucia, M.S.; et al. Lipid catabolism inhibition sensitizes prostate cancer cells to antiandrogen blockade. Oncotarget 2017, 8, 56051–56065. [Google Scholar] [CrossRef]
- Álvarez-Garcia, V.; Tawil, Y.; Wise, H.M.; Leslie, N.R. Mechanisms of PTEN loss in cancer: It’s all about diversity. Semin. Cancer Biol. 2019, 59, 66–79. [Google Scholar] [CrossRef]
- Smith, R. Enzalutamide response in a panel of prostate cancer cell lines reveals a role for glucocorticoid receptor in en-zalutamide resistant disease. Sci. Rep. 2020, 10, 21750. [Google Scholar] [CrossRef] [PubMed]
- VanDeusen, H.R.; Ramroop, J.R.; Morel, K.L.; Bae, S.Y.; Sheahan, A.V.; Sychev, Z.; Lau, N.A.; Cheng, L.C.; Tan, V.M.; Li, Z.; et al. Targeting RET Kinase in Neuroendocrine Prostate Cancer. Mol. Cancer Res. 2020, 18, 1176–1188. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Li, S.; Wang, F.; Fan, C.; Wang, J. Identification of key pathways and genes in PTEN mutation prostate cancer by bioinformatics analysis. BMC Med. Genet. 2019, 20, 191. [Google Scholar] [CrossRef] [PubMed]
- Jamaspishvili, T.; Berman, D.M.; Ross, A.E.; Scher, H.I.; De Marzo, A.M.; Squire, J.A.; Lotan, T.L. Clinical implications of PTEN loss in prostate cancer. Nat. Rev. Urol. 2018, 15, 222–234. [Google Scholar] [CrossRef]
- Bastos, D.C. Genetic ablation of FASN attenuates the invasive potential of prostate cancer driven by Pten loss. J. Pathol. 2021, 253, 292–303. [Google Scholar] [CrossRef]
- Chen, M.; Zhang, J.; Sampieri, K.; Clohessy, J.G.; Mendez, L.; Gonzalez-Billalabeitia, E.; Liu, X.-S.; Lee, Y.-R.; Fung, J.; Katon, J.M.; et al. An aberrant SREBP-dependent lipogenic program promotes metastatic prostate cancer. Nat. Genet. 2018, 50, 206–218. [Google Scholar] [CrossRef]
- Maughan, B.L. p53 status in the primary tumor predicts efficacy of subsequent abiraterone and enzalutamide in castra-tion-resistant prostate cancer. Prostate Cancer Prostatic Dis. 2018, 21, 260–268. [Google Scholar] [CrossRef]
- Chappell, W.H. Roles of p53, NF-κB and the androgen receptor in controlling NGAL expression in prostate cancer cell lines. Adv. Biol. Regul. 2018, 69, 43–62. [Google Scholar] [CrossRef]
- Liu, C.; Zhu, Y.; Lou, W.; Nadiminty, N.; Chen, X.; Zhou, Q.; Shi, X.B.; White, R.W.D.; Gao, A.C. Functional p53 determines docetaxel sensitivity in prostate cancer cells. Prostate 2013, 73, 418–427. [Google Scholar] [CrossRef]
- Dong, J.-T. Prevalent mutations in prostate cancer. J. Cell. Biochem. 2006, 97, 433–447. [Google Scholar] [CrossRef]
- Aggarwal, M.; Saxena, R.; Sinclair, E.; Fu, Y.; Jacobs, A.; Dyba, M.; Wang, X.; Cruz, I.; Berry, D.; Kallakury, B.; et al. Reactivation of mutant p53 by a dietary-related compound phenethyl isothiocyanate inhibits tumor growth. Cell Death Differ. 2016, 23, 1615–1627. [Google Scholar] [CrossRef]
- Reina-Campos, M.; Linares, J.F.; Duran, A.; Cordes, T.; L’Hermitte, A.; Badur, M.G.; Bhangoo, M.S.; Thorson, P.K.; Richards, A.; Rooslid, T.; et al. Increased Serine and One-Carbon Pathway Metabolism by PKCλ/ι Deficiency Promotes Neuroendocrine Prostate Cancer. Cancer Cell 2019, 35, 385–400.e9. [Google Scholar] [CrossRef]
- Ben-Salem, S.; Venkadakrishnan, V.B. Novel insights in cell cycle dysregulation during prostate cancer progression. Endocr.-Relat. Cancer 2021, 28, R141–R155. [Google Scholar] [CrossRef]
- Wang, C.; Fang, H.; Zhang, J.; Gu, Y. Targeting “undruggable” c-Myc protein by synthetic lethality. Front. Med. 2021, 15, 541–550. [Google Scholar] [CrossRef]
- Miller, D.R.; Ingersoll, M.A.; Teply, B.A.; Lin, M.-F. Targeting treatment options for castration-resistant prostate cancer. Am. J. Clin. Exp. Urol. 2021, 9, 101–120. [Google Scholar]
- Dey, P.; Kimmelman, A.C.; DePinho, R.A. Metabolic Codependencies in the Tumor Microenvironment. Cancer Discov. 2021, 11, 1067–1081. [Google Scholar] [CrossRef]
- Mukha, A.; Kahya, U.; Dubrovska, A. Targeting glutamine metabolism and autophagy: The combination for prostate cancer radiosensitization. Autophagy 2021, 17, 3879–3881. [Google Scholar] [CrossRef]
- Clegg, N.J.; Couto, S.S.; Wongvipat, J.; Hieronymus, H.; Carver, B.S.; Taylor, B.S.; Ellwood-Yen, K.; Gerald, W.L.; Sander, C.; Sawyers, C.L. MYC Cooperates with AKT in Prostate Tumorigenesis and Alters Sensitivity to mTOR Inhibitors. PLoS ONE 2011, 6, e17449. [Google Scholar] [CrossRef]
- Goetzman, E.S.; Prochownik, E.V. The Role for Myc in Coordinating Glycolysis, Oxidative Phosphorylation, Glutaminolysis, and Fatty Acid Metabolism in Normal and Neoplastic Tissues. Front. Endocrinol. 2018, 9, 129. [Google Scholar] [CrossRef]
- Priolo, C.; Pyne, S.; Rose, J.; Regan, E.R.; Zadra, G.; Photopoulos, C.; Cacciatore, S.; Schultz, D.; Scaglia, N.; McDunn, J.; et al. AKT1 and MYC Induce Distinctive Metabolic Fingerprints in Human Prostate Cancer. Cancer Res. 2014, 74, 7198–7204. [Google Scholar] [CrossRef]
- Bai, S.; Cao, S.; Jin, L.; Kobelski, M.; Schouest, B.; Wang, X.; Ungerleider, N.; Baddoo, M.; Zhang, W.; Corey, E.; et al. A positive role of c-Myc in regulating androgen receptor and its splice variants in prostate cancer. Oncogene 2019, 38, 4977–4989. [Google Scholar] [CrossRef]
- Bernard, D.; Pourtier-Manzanedo, A.; Gil, J.; Beach, D.H. Myc confers androgen-independent prostate cancer cell growth. J. Clin. Investig. 2003, 112, 1724–1731. [Google Scholar] [CrossRef]
- Gao, L. Androgen receptor promotes ligand-independent prostate cancer progression through c-Myc upregulation. PLoS ONE 2013, 8, e63563. [Google Scholar]
- Barfeld, S.J.; Urbanucci, A.; Itkonen, H.M.; Fazli, L.; Hicks, J.L.; Thiede, B.; Rennie, P.S.; Yegnasubramanian, S.; DeMarzo, A.M.; Mills, I.G. c-Myc Antagonises the Transcriptional Activity of the Androgen Receptor in Prostate Cancer Affecting Key Gene Networks. EBioMedicine 2017, 18, 83–93. [Google Scholar] [CrossRef]
- Long, T.; Hicks, M.; Yu, H.-C.; Biggs, W.H.; Kirkness, E.F.; Menni, C.; Zierer, J.; Small, K.S.; Mangino, M.; Messier, H.; et al. Whole-genome sequencing identifies common-to-rare variants associated with human blood metabolites. Nat. Genet. 2017, 49, 568–578. [Google Scholar] [CrossRef] [PubMed]
- Cai, Z.; Deng, Y.; Ye, J.; Zhuo, Y.; Liu, Z.; Liang, Y.; Zhang, H.; Zhu, X.; Luo, Y.; Feng, Y.; et al. Aberrant Expression of Citrate Synthase is Linked to Disease Progression and Clinical Outcome in Prostate Cancer. Cancer Manag. Res. 2020, 12, 6149–6163. [Google Scholar] [CrossRef]
- Gilbert, R.; Bonilla, C.; Metcalfe, C.; Lewis, S.; Evans, D.M.; Fraser, W.D.; Kemp, J.P.; Donovan, J.L.; Hamdy, F.C.; Neal, D.E.; et al. Associations of vitamin D pathway genes with circulating 25-hydroxyvitamin-D, 1,25-dihydroxyvitamin-D, and prostate cancer: A nested case–control study. Cancer Causes Control. 2014, 26, 205–218. [Google Scholar] [CrossRef]
- Zecchini, V. Nuclear ARRB1 induces pseudohypoxia and cellular metabolism reprogramming in prostate cancer. EMBO J. 2014, 33, 1365–1382. [Google Scholar] [CrossRef]
- Hong, M.-G.; Karlsson, R.; Magnusson, P.K.E.; Lewis, M.R.; Isaacs, W.; Zheng, L.S.; Xu, J.; Grönberg, H.; Ingelsson, E.; Pawitan, Y.; et al. A Genome-Wide Assessment of Variability in Human Serum Metabolism. Hum. Mutat. 2013, 34, 515–524. [Google Scholar] [CrossRef] [PubMed]
- Poisson, L.M.; Sreekumar, A.; Chinnaiyan, A.M.; Ghosh, D. Pathway-directed weighted testing procedures for the inte-grative analysis of gene expression and metabolomic data. Genomics 2012, 99, 265–274. [Google Scholar] [CrossRef] [PubMed]
- Lu, B.; Asara, J.M.; Sanda, M.G.; Arredouani, M.S. The Role of the Transcription Factor SIM2 in Prostate Cancer. PLoS ONE 2011, 6, e28837. [Google Scholar] [CrossRef] [PubMed]
- Massie, C.E.; Lynch, A.; Ramos-Montoya, A.; Boren, J.; Stark, R.; Fazli, L.; Warren, A.; Scott, H.; Madhu, B.; Sharma, N.; et al. The androgen receptor fuels prostate cancer by regulating central metabolism and biosynthesis. EMBO J. 2011, 30, 2719–2733. [Google Scholar] [CrossRef]
- Van Tilborg, D.; Saccenti, E. Cancers in agreement? Exploring the cross-talk of cancer metabolomic and transcriptomic landscapes using publicly available data. Cancers 2021, 13, 393. [Google Scholar] [CrossRef]
- Wang, W.; He, Z.; Kong, Y.; Liu, Z.; Gong, L. GC-MS-based metabolomics reveals new biomarkers to assist the differentiation of prostate cancer and benign prostatic hyperplasia. Clin. Chim. Acta 2021, 519, 10–17. [Google Scholar] [CrossRef]
- Gómez-Cebrián, N.; García-Flores, M.; Rubio-Briones, J.; López-Guerrero, J.A.; Pineda-Lucena, A.; Puchades-Carrasco, L. Targeted Metabolomics Analyses Reveal Specific Metabolic Alterations in High-Grade Prostate Cancer Patients. J. Proteome Res. 2020, 19, 4082–4092. [Google Scholar] [CrossRef]
- Andersen, M.K.; Rise, K.; Giskeødegård, G.F.; Richardsen, E.; Bertilsson, H.; Størkersen, Ø.; Bathen, T.F.; Rye, M.; Tessem, M.-B. Integrative metabolic and transcriptomic profiling of prostate cancer tissue containing reactive stroma. Sci. Rep. 2018, 8, 14269. [Google Scholar] [CrossRef]
- Shao, Y.; Ye, G.; Ren, S.; Piao, H.-L.; Zhao, X.; Lu, X.; Wang, F.; Ma, W.; Li, J.; Yin, P.; et al. Metabolomics and transcriptomics profiles reveal the dysregulation of the tricarboxylic acid cycle and related mechanisms in prostate cancer. Int. J. Cancer 2018, 143, 396–407. [Google Scholar] [CrossRef]
- Al Kadhi, O. Increased transcriptional and metabolic capacity for lipid metabolism in the peripheral zone of the prostate may underpin its increased susceptibility to cancer. Oncotarget 2017, 8, 84902–84916. [Google Scholar] [CrossRef]
- Sandsmark, E.; Hansen, A.F.; Selnæs, K.M.; Bertilsson, H.; Bofin, A.M.; Wright, A.J.; Viset, T.; Richardsen, E.; Drabløs, F.; Bathen, T.F.; et al. A novel non-canonical Wnt signature for prostate cancer aggressiveness. Oncotarget 2017, 8, 9572–9586. [Google Scholar] [CrossRef]
- Ren, S.; Shao, Y.; Zhao, X.; Hong, C.S.; Wang, F.; Lu, X.; Li, J.; Ye, G.; Yan, M.; Zhuang, Z.; et al. Integration of Metabolomics and Transcriptomics Reveals Major Metabolic Pathways and Potential Biomarker Involved in Prostate Cancer. Mol. Cell. Proteom. 2016, 15, 154–163. [Google Scholar] [CrossRef]
- Torrano, V.; Valcarcel-Jimenez, L.; Cortazar, A.R.; Liu, X.; Urosevic, J.; Castillo-Martin, M.; Fernández-Ruiz, S.; Morciano, G.; Caro-Maldonado, A.; Guiu, M.; et al. The metabolic co-regulator PGC1α suppresses prostate cancer metastasis. Nature 2016, 18, 645–656. [Google Scholar] [CrossRef]
- Zhang, J.; Li, L.; Tang, S.; Zhang, Y.; Markiewski, M.; Xing, C.; Jiang, C.; Lu, J. Pyranocoumarin Tissue Distribution, Plasma Metabolome and Prostate Transcriptome Impacts of Sub-Chronic Exposure to Korean Angelica Supplement in Mice. Am. J. Chin. Med. 2016, 44, 321–353. [Google Scholar] [CrossRef]
- Cerasuolo, M.; Paris, D.; Iannotti, F.; Melck, D.; Verde, R.; Mazzarella, E.; Motta, A.; Ligresti, A. Neuroendocrine Transdifferentiation in Human Prostate Cancer Cells: An Integrated Approach. Cancer Res. 2015, 75, 2975–2986. [Google Scholar] [CrossRef]
- Meller, S. Integration of tissue metabolomics, transcriptomics and immunohistochemistry reveals ERG- and gleason score-specific metabolomic alterations in prostate cancer. Oncotarget 2016, 7, 1421–1438. [Google Scholar] [CrossRef]
- Kim, S.; Pevzner, P.A. MS-GF+ makes progress towards a universal database search tool for proteomics. Nat. Commun. 2014, 5, 5277. [Google Scholar] [CrossRef]
- Schroeder, M.; Jakovcevski, M.; Polacheck, T.; Lebow, M.; Drori, Y.; Engel, M.; Ben-Dor, S.; Chen, A. A Methyl-Balanced Diet Prevents CRF-Induced Prenatal Stress-Triggered Predisposition to Binge Eating-like Phenotype. Cell Metab. 2017, 25, 1269–1281.e6. [Google Scholar] [CrossRef]
- Felgueiras, J.; Silva, J.; Nunes, A.; Fernandes, I.; Patrício, A.; Maia, N.; Pelech, S.; Fardilha, M. Investigation of spectroscopic and proteomic alterations underlying prostate carcinogenesis. J. Proteom. 2020, 226, 103888. [Google Scholar] [CrossRef]
- Kopylov, A.T.; Petrovsky, D.V.; Stepanov, A.A.; Rudnev, V.R.; Malsagova, K.A.; Butkova, T.V.; Zakharova, N.V.; Kostyuk, G.P.; Kulikova, L.I.; Enikeev, D.V.; et al. Convolutional neural network in proteomics and metabolomics for determination of comorbidity between cancer and schizophrenia. J. Biomed. Inform. 2021, 122, 103890. [Google Scholar] [CrossRef]
- Shen, S.; Li, J.; Huo, S.; Ma, M.; Zhu, X.; Rasam, S.; Duan, X.; Qu, M.; Titus, M.A.; Qu, J. Parallel, High-Quality Proteomic and Targeted Metabolomic Quantification Using Laser Capture Microdissected Tissues. Anal. Chem. 2021, 93, 8711–8718. [Google Scholar] [CrossRef]
- Oberhuber, M.; Pecoraro, M.; Rusz, M.; Oberhuber, G.; Wieselberg, M.; Haslinger, P.; Gurnhofer, E.; Schlederer, M.; Limberger, T.; Lagger, S.; et al. STAT 3-dependent analysis reveals PDK 4 as independent predictor of recurrence in prostate cancer. Mol. Syst. Biol. 2020, 16, e9247. [Google Scholar] [CrossRef]
- Itkonen, H.M.; Poulose, N.; Walker, S.; Mills, I.G. CDK9 Inhibition Induces a Metabolic Switch that Renders Prostate Cancer Cells Dependent on Fatty Acid Oxidation. Neoplasia 2019, 21, 713–720. [Google Scholar] [CrossRef]
- Gao, B.; Lue, H.-W.; Podolak, J.; Fan, S.; Zhang, Y.; Serawat, A.; Alumkal, J.J.; Fiehn, O.; Thomas, G.V. Multi-Omics Analyses Detail Metabolic Reprogramming in Lipids, Carnitines, and Use of Glycolytic Intermediates between Prostate Small Cell Neuroendocrine Carcinoma and Prostate Adenocarcinoma. Metabolites 2019, 9, 82. [Google Scholar] [CrossRef] [PubMed]
- Kregel, S. Functional and mechanistic interrogation of BET bromodomain degraders for the treatment of metastatic cas-tration-resistant prostate cancer. Clin. Cancer Res. 2019, 25, 4038–4048. [Google Scholar] [CrossRef]
- Zadra, G.; Ribeiro, C.F.; Chetta, P.; Ho, Y.; Cacciatore, S.; Gao, X.; Syamala, S.; Bango, C.; Photopoulos, C.; Huang, Y.; et al. Inhibition of de novo lipogenesis targets androgen receptor signaling in castration-resistant prostate cancer. Proc. Natl. Acad. Sci. USA 2019, 116, 631–640. [Google Scholar] [CrossRef]
- Murphy, K.; Murphy, B.T.; Boyce, S.; Flynn, L.; Gilgunn, S.; O’Rourke, C.J.; Rooney, C.; Stöckmann, H.; Walsh, A.L.; Finn, S.; et al. Integrating biomarkers across omic platforms: An approach to improve stratification of patients with indolent and aggressive prostate cancer. Mol. Oncol. 2018, 12, 1513–1525. [Google Scholar] [CrossRef] [PubMed]
- Hansen, A.F.; Sandsmark, E.; Rye, M.B.; Wright, A.J.; Bertilsson, H.; Richardsen, E.; Viset, T.; Bofin, A.M.; Angelsen, A.; Selnæs, K.M.; et al. Presence of TMPRSS2-ERG is associated with alterations of the metabolic profile in human prostate cancer. Oncotarget 2016, 7, 42071–42085. [Google Scholar] [CrossRef] [PubMed]
- American Cancer Society. Key Statistics for Prostate Cancer. Available online: https://www.cancer.org/cancer/prostate-cancer/about/key-statistics.html (accessed on 28 December 2020).
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2020. CA Cancer J. Clin. 2020, 70, 7–30. [Google Scholar] [CrossRef] [PubMed]
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef]
- Zhang, A.; Sun, H.; Xu, H.; Qiu, S.; Wang, X. Cell metabolomics. OMICS A J. Integr. Biol. 2013, 17, 495–501. [Google Scholar] [CrossRef]
- Costello, L.C.; Franklin, R.B. Concepts of citrate production and secretion by prostate 1. Metabolic relationships. Prostate 1991, 18, 25–46. [Google Scholar] [CrossRef]
- Clos-Garcia, M.; Loizaga-Iriarte, A.; Zuñiga-Garcia, P.; Sánchez-Mosquera, P.; Cortazar, A.R.; González, E.; Torrano, V.; Alonso, C.; Pérez-Cormenzana, M.; Ugalde-Olano, A.; et al. Metabolic alterations in urine extracellular vesicles are associated to prostate cancer pathogenesis and progression. J. Extracell. Vesicles 2018, 7, 1470442. [Google Scholar] [CrossRef]
- Puhka, M. Metabolomic profiling of extracellular vesicles and alternative normalization methods reveal enriched metab-olites and strategies to study prostate cancer-related changes. Theranostics 2017, 7, 3824–3841. [Google Scholar] [CrossRef]
- Davalieva, K.; Kostovska, I.M.; Kiprijanovska, S.; Markoska, K.; Kubelka-Sabit, K.; Filipovski, V.; Stavridis, S.; Stankov, O.; Komina, S.; Petrusevska, G.; et al. Proteomics analysis of malignant and benign prostate tissue by 2D DIGE/MS reveals new insights into proteins involved in prostate cancer. Prostate 2015, 75, 1586–1600. [Google Scholar] [CrossRef]
- Giskeødegård, G.F.; Hansen, A.F.; Bertilsson, H.; Gonzalez, S.V.; Kristiansen, K.A.; Bruheim, P.; Mjøs, S.A.; Angelsen, A.; Bathen, T.F.; Tessem, M.-B. Metabolic markers in blood can separate prostate cancer from benign prostatic hyperplasia. Br. J. Cancer 2015, 113, 1712–1719. [Google Scholar] [CrossRef]
- Lin, H.-M.; Mahon, K.; Weir, J.M.; Mundra, P.A.; Spielman, C.; Briscoe, K.; Gurney, H.; Mallesara, G.; Marx, G.; Stockler, M.R.; et al. A distinct plasma lipid signature associated with poor prognosis in castration-resistant prostate cancer. Int. J. Cancer 2017, 141, 2112–2120. [Google Scholar] [CrossRef]
- Albanes, D.; Weinstein, S.J.; Mondul, A.M. Abstract 3987: Prospective serum metabolomic profiles of prostate cancer by size and extent of primary tumor. In Clinical Research (Excluding Clinical Trials); American Association for Cancer Research: Philadelphia, PA, USA, 2016; p. 3981. [Google Scholar] [CrossRef]
- Roberts, M.J.; Richards, R.S.; Chow, C.W.; Buck, M.; Yaxley, J.; Lavin, M.; Schirra, H.J.; Gardiner, R.A. Seminal plasma enables selection and monitoring of active surveillance candidates using nuclear magnetic resonance-based metabolomics: A preliminary investigation. Prostate Int. 2017, 5, 149–157. [Google Scholar] [CrossRef]
- Andras, I.; Crisan, N.; Vesa, S.; Rahota, R.; Romanciuc, F.; Lazar, A.; Socaciu, C.; Matei, D.-V.; De Cobelli, O.; Bocsan, I.-S.; et al. Serum metabolomics can predict the outcome of first systematic transrectal prostate biopsy in patients with PSA <10 ng/ml. Futur. Oncol. 2017, 13, 1793–1800. [Google Scholar] [CrossRef]
- Huang, J.; Mondul, A.M.; Weinstein, S.J.; Karoly, E.D.; Sampson, J.N.; Albanes, D. Prospective serum metabolomic profile of prostate cancer by size and extent of primary tumor. Oncotarget 2017, 8, 45190–45199. [Google Scholar] [CrossRef]
- Kühn, T. Higher plasma levels of lysophosphatidylcholine 18:0 are related to a lower risk of common cancers in a pro-spective metabolomics study. BMC Med. 2016, 14, 13. [Google Scholar] [CrossRef] [PubMed]
- Averna, T.A.; Kline, E.E.; Smith, A.Y.; Sillerud, L.O. A decrease in 1H nuclear magnetic resonance spectroscopically determined citrate in human seminal fluid accompanies the development of prostate adenocarcinoma. J. Urol. 2005, 173, 433–438. [Google Scholar] [CrossRef] [PubMed]
- Kumar, D.; Gupta, A.; Mandhani, A.; Sankhwar, S.N. NMR spectroscopy of filtered serum of prostate cancer: A new frontier in metabolomics. Prostate 2016, 76, 1106–1119. [Google Scholar] [CrossRef] [PubMed]
- Van der Mijn, J.C. Lactic acidosis in prostate cancer: Consider the Warburg effect. Case Rep. Oncol. 2017, 10, 1085–1091. [Google Scholar] [CrossRef] [PubMed]
- Sadeghi, R.N.; Karami-Tehrani, F.; Salami, S. Targeting prostate cancer cell metabolism: Impact of hexokinase and CPT-1 enzymes. Tumor Biol. 2014, 36, 2893–2905. [Google Scholar] [CrossRef]
- Warburg, O. On the Origin of Cancer Cells. Science 1956, 123, 309–314. [Google Scholar] [CrossRef]
- Kwon, H.; Oh, S.; Jin, X.; An, Y.J.; Park, S. Cancer metabolomics in basic science perspective. Arch. Pharmacal Res. 2015, 38, 372–380. [Google Scholar] [CrossRef]
- Koppenol, W.H.; Bounds, P.L.; Dang, C.V. Otto Warburg’s contributions to current concepts of cancer metabolism. Nat. Rev. Cancer 2011, 11, 325–337. [Google Scholar] [CrossRef]
- Wang, X.; Liu, H.; Ni, Y.; Shen, P.; Han, X. Lactate shuttle: From substance exchange to regulatory mechanism. Hum. Cell 2022, 35, 1–14. [Google Scholar] [CrossRef]
- Tessem, M.-B. Evaluation of lactate and alanine as metabolic biomarkers of prostate cancer using 1 H HR-MAS spec-troscopy of biopsy tissues. Magn. Reson. Med. 2008, 60, 510–516. [Google Scholar] [CrossRef]
- Lima, A.R.; Araújo, A.M.; Pinto, J.; Jerónimo, C.; Henrique, R.; Bastos, M.D.L.; Carvalho, M.; de Pinho, P.G. GC-MS-Based Endometabolome Analysis Differentiates Prostate Cancer from Normal Prostate Cells. Metabolites 2018, 8, 23. [Google Scholar] [CrossRef]
- Andersen, M.K.; Giskeødegård, G.F.; Tessem, M.-B. Metabolic alterations in tissues and biofluids of patients with prostate cancer. Curr. Opin. Endocr. Metab. Res. 2020, 10, 23–28. [Google Scholar] [CrossRef]
- Cascardo, F.; Anselmino, N.; Páez, A.; Labanca, E.; Sanchis, P.; Antico-Arciuch, V.; Navone, N.; Gueron, G.; Vázquez, E.; Cotignola, J. HO-1 Modulates Aerobic Glycolysis through LDH in Prostate Cancer Cells. Antioxidants 2021, 10, 966. [Google Scholar] [CrossRef]
- Baron, A.; Migita, T.; Tang, D.; Loda, M. Fatty acid synthase: A metabolic oncogene in prostate cancer? J. Cell. Biochem. 2004, 91, 47–53. [Google Scholar] [CrossRef]
- Singh, K.B.; Hahm, E.-R.; Kim, S.-H.; Wendell, S.G.; Singh, S.V. A novel metabolic function of Myc in regulation of fatty acid synthesis in prostate cancer. Oncogene 2021, 40, 592–602. [Google Scholar] [CrossRef]
- Suburu, J.; Chen, Y.Q. Lipids and prostate cancer. Prostaglandins Other Lipid Mediat. 2012, 98, 1–10. [Google Scholar] [CrossRef]
- Liu, Y. Fatty acid oxidation is a dominant bioenergetic pathway in prostate cancer. Prostate Cancer Prostatic Dis. 2006, 9, 230–234. [Google Scholar] [CrossRef]
- Giskeødegård, G.F.; Bertilsson, H.; Selnæs, K.M.; Wright, A.J.; Bathen, T.F.; Viset, T.; Halgunset, J.; Angelsen, A.; Gribbestad, I.S.; Tessem, M.-B. Spermine and Citrate as Metabolic Biomarkers for Assessing Prostate Cancer Aggressiveness. PLoS ONE 2013, 8, e62375. [Google Scholar] [CrossRef]
- Zabala-Letona, A.; Arruabarrena-Aristorena, A.; Martín-Martín, N.; Fernandez-Ruiz, S.; Sutherland, J.D.; Clasquin, M.; Tomas-Cortazar, J.; Jimenez, J.; Torres, I.; Quang, P.; et al. mTORC1-dependent AMD1 regulation sustains polyamine metabolism in prostate cancer. Nature 2017, 547, 109–113. [Google Scholar] [CrossRef]
- Lloyd, S.M.; Arnold, J.; Sreekumar, A. Metabolomic profiling of hormone-dependent cancers: A bird’s eye view. Trends Endocrinol. Metab. 2015, 26, 477–485. [Google Scholar] [CrossRef]
- Goodwin, A.C.; Jadallah, S.; Toubaji, A.; Lecksell, K.; Hicks, J.L.; Kowalski, J.; Bova, G.S.; De Marzo, A.M.; Netto, G.J.; Casero, R.A., Jr. Increased spermine oxidase expression in human prostate cancer and prostatic intraepithelial neoplasia tissues. Prostate 2008, 68, 766–772. [Google Scholar] [CrossRef]
- Sharma, U.; Jagannathan, N.R. Metabolism of prostate cancer by magnetic resonance spectroscopy (MRS). Biophys. Rev. 2020, 12, 1163–1173. [Google Scholar] [CrossRef]
- Madhu, B.; Shaw, G.L.; Warren, A.Y.; Neal, D.E.; Griffiths, J.R. Response of Degarelix treatment in human prostate cancer monitored by HR-MAS 1H NMR spectroscopy. Metabolomics 2016, 12, 120. [Google Scholar] [CrossRef]
- Awwad, H.M.; Geisel, J.; Obeid, R. The role of choline in prostate cancer. Clin. Biochem. 2012, 45, 1548–1553. [Google Scholar] [CrossRef]
- Wen, S.; He, Y.; Wang, L.; Zhang, J.; Quan, C.; Niu, Y.; Huang, H. Aberrant activation of super enhancer and choline metabolism drive antiandrogen therapy resistance in prostate cancer. Oncogene 2020, 39, 6556–6571. [Google Scholar] [CrossRef] [PubMed]
- Tayari, N.; Wright, A.J.; Heerschap, A. Absolute choline tissue concentration mapping for prostate cancer localization and characterization using 3D 1 H MRSI without water-signal suppression. Magn. Reson. Med. 2022, 87, 561–573. [Google Scholar] [CrossRef]
- Wang, M.; Zou, L.; Liang, J.; Wang, X.; Zhang, D.; Fang, Y.; Zhang, J.; Xiao, F.; Liu, M. The Urinary Sarcosine/Creatinine Ratio is a Potential Diagnostic and Prognostic Marker in Prostate Cancer. Med Sci. Monit. 2018, 24, 3034–3041. [Google Scholar] [CrossRef]
- Corbin, J.M.; Ruiz-Echevarría, M.J. One-Carbon Metabolism in Prostate Cancer: The Role of Androgen Signaling. Int. J. Mol. Sci. 2016, 17, 1208. [Google Scholar] [CrossRef]
- Valle, S.; Sharifi, N. Targeting Glucocorticoid Metabolism in Prostate Cancer. Endocrinology 2021, 162. [Google Scholar] [CrossRef] [PubMed]
- Puhr, M. The glucocorticoid receptor Is a key player for prostate cancer cell survival and a target for improved anti-androgen therapy. Clin. Cancer Res. 2018, 24, 927–938. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.; Chen, Q. The role of glucocorticoid receptor in prostate cancer progression: From bench to bedside. Int. Urol. Nephrol. 2017, 49, 369–380. [Google Scholar] [CrossRef]
- Moon, J.-S.; Hisata, S.; Park, M.-A.; DeNicola, G.M.; Ryter, S.W.; Nakahira, K.; Choi, A.M.K. mTORC1-Induced HK1-Dependent Glycolysis Regulates NLRP3 Inflammasome Activation. Cell Rep. 2015, 12, 102–115. [Google Scholar] [CrossRef]
- Mamouni, K.; Kallifatidis, G.; Lokeshwar, B. Targeting Mitochondrial Metabolism in Prostate Cancer with Triterpenoids. Int. J. Mol. Sci. 2021, 22, 2466. [Google Scholar] [CrossRef]
- Singh, K.K.; Desouki, M.M.; Franklin, R.B.; Costello, L.C. Mitochondrial aconitase and citrate metabolism in malignant and nonmalignant human prostate tissues. Mol. Cancer 2006, 5, 14. [Google Scholar] [CrossRef]
- Barron, E.G.; Huggins, C. The Metabolism of the Prostate: Transamination and Citric Acid. J. Urol. 1946, 55, 385–390. [Google Scholar] [CrossRef]
- Ding, C.; Zhang, J.; Li, B.; Ding, Z.; Cheng, W.; Gao, F.; Zhang, Y.; Xu, Y.; Zhang, S. Cornin protects SH-SY5Y cells against oxygen and glucose deprivation-induced autophagy through the PI3K/Akt/mTOR pathway. Mol. Med. Rep. 2017, 17, 87–92. [Google Scholar] [CrossRef]
- Costello, L.C.; Franklin, R.B. Zinc is decreased in prostate cancer: An established relationship of prostate cancer! JBIC J. Biol. Inorg. Chem. 2011, 16, 3–8. [Google Scholar] [CrossRef]
- Flavin, R.; Zadra, G.; Loda, M. Metabolic alterations and targeted therapies in prostate cancer. J. Pathol. 2010, 223, 284–295. [Google Scholar] [CrossRef]
- Strmiska, V.; Michalek, P.; Eckschlager, T.; Stiborova, M.; Adam, V.; Krizkova, S.; Heger, Z. Prostate cancer-specific hallmarks of amino acids metabolism: Towards a paradigm of precision medicine. Biochim. Biophys. Acta 2019, 1871, 248–258. [Google Scholar] [CrossRef]
- Soga, T. Cancer metabolism: Key players in metabolic reprogramming. Cancer Sci. 2013, 104, 275–281. [Google Scholar] [CrossRef]
- Vander Heiden, M.G.; Cantley, L.C.; Thompson, C.B. Understanding the Warburg Effect: The Metabolic Requirements of Cell Proliferation. Science 2009, 324, 1029–1033. [Google Scholar] [CrossRef] [PubMed]
- Lee, G.; Zheng, Y.; Cho, S.; Jang, C.; England, C.; Dempsey, J.M.; Yu, Y.; Liu, X.; He, L.; Cavaliere, P.M.; et al. Post-transcriptional Regulation of De Novo Lipogenesis by mTORC1-S6K1-SRPK2 Signaling. Cell 2017, 171, 1545–1558.e18. [Google Scholar] [CrossRef] [PubMed]
- Herzig, S.; Shaw, R.J. AMPK: Guardian of metabolism and mitochondrial homeostasis. Nat. Rev. Mol. Cell Biol. 2018, 19, 121–135. [Google Scholar] [CrossRef] [PubMed]
- Hardie, D.G.; Ross, F.A.; Hawley, S.A. AMPK: A nutrient and energy sensor that maintains energy homeostasis. Nat. Rev. Mol. Cell Biol. 2012, 13, 251–262. [Google Scholar] [CrossRef]
- Braglia, L.; Zavatti, M.; Vinceti, M.; Martelli, A.M.; Marmiroli, S. Deregulated PTEN/PI3K/AKT/mTOR signaling in prostate cancer: Still a potential druggable target? Biochim. Biophys. Acta (BBA)-Mol. Cell Res. 2020, 1867, 118731. [Google Scholar] [CrossRef]
- Hsieh, A.; Edlind, M. PI3K-AKT-mTOR signaling in prostate cancer progression and androgen deprivation therapy re-sistance. Asian J. Androl. 2014, 16, 378. [Google Scholar] [CrossRef]
- Chang, L.; Graham, P.H.; Ni, J.; Hao, J.; Bucci, J.; Cozzi, P.J.; Li, Y. Targeting PI3K/Akt/mTOR signaling pathway in the treatment of prostate cancer radioresistance. Crit. Rev. Oncol. 2015, 96, 507–517. [Google Scholar] [CrossRef]
- Fang, M.; Shen, Z.; Huang, S.; Zhao, L.; Chen, S.; Mak, T.W.; Wang, X. The ER UDPase ENTPD5 Promotes Protein N-Glycosylation, the Warburg Effect, and Proliferation in the PTEN Pathway. Cell 2010, 143, 711–724. [Google Scholar] [CrossRef]
- Wang, Y.; Romigh, T.; He, X.; Tan, M.-H.; Orloff, M.S.; Silverman, R.H.; Heston, W.D.; Eng, C. Differential regulation of PTEN expression by androgen receptor in prostate and breast cancers. Oncogene 2011, 30, 4327–4338. [Google Scholar] [CrossRef]
- Zhou, X.; Yang, X.; Sun, X.; Xu, X.; Li, X.; Guo, Y.; Wang, J.; Yao, L.; Wang, H.; Shen, L. Effect of PTEN loss on metabolic reprogramming in prostate cancer cells. Oncol. Lett. 2019, 17, 2856–2866. [Google Scholar] [CrossRef]
- Deng, Y.; Lu, J. Targeting hexokinase 2 in castration-resistant prostate cancer. Mol. Cell. Oncol. 2015, 2, e974465. [Google Scholar] [CrossRef]
- Clapé, C.; Fritz, V.; Henriquet, C.; Apparailly, F.; Fernandez, P.L.; Iborra, F.; Avancès, C.; Villalba, M.; Culine, S.; Fajas, L. miR-143 Interferes with ERK5 Signaling, and Abrogates Prostate Cancer Progression in Mice. PLoS ONE 2009, 4, e7542. [Google Scholar] [CrossRef]
- Xu, B.; Niu, X.; Zhang, X.; Tao, J.; Wu, D.; Wang, Z.; Li, P.; Zhang, W.; Wu, H.; Feng, N.; et al. miR-143 decreases prostate cancer cells proliferation and migration and enhances their sensitivity to docetaxel through suppression of KRAS. Mol. Cell. Biochem. 2011, 350, 207–213. [Google Scholar] [CrossRef]
- Chu, H.; Zhong, D.; Tang, J.; Li, J.; Xue, Y.; Tong, N.; Qin, C.; Yin, C.; Zhang, Z.; Wang, M. A functional variant in miR-143 promoter contributes to prostate cancer risk. Arch. Toxicol. 2014, 90, 403–414. [Google Scholar] [CrossRef]
- Wach, S.; Brandl, M.; Borchardt, H.; Weigelt, K.; Lukat, S.; Nolte, E.; Al-Janabi, O.; Hart, M.; Grässer, F.; Giedl, J.; et al. Exploring the MIR143-UPAR Axis for the Inhibition of Human Prostate Cancer Cells In Vitro and In Vivo. Mol. Ther.-Nucleic Acids 2019, 16, 272–283. [Google Scholar] [CrossRef]
- Ros, S.; Santos, C.R.; Moco, S.; Baenke, F.; Kelly, G.; Howell, M.; Zamboni, N.; Schulze, A. Functional Metabolic Screen Identifies 6-Phosphofructo-2-Kinase/Fructose-2,6-Biphosphatase 4 as an Important Regulator of Prostate Cancer Cell Survival. Cancer Discov. 2012, 2, 328–343. [Google Scholar] [CrossRef]
- Chiacchiera, F.; Simone, C. The AMPK-FoxO3A axis as a target for cancer treatment. Cell Cycle 2010, 9, 1091–1096. [Google Scholar] [CrossRef]
- Ahmadian, M.; Abbott, M.J.; Tang, T.; Hudak, C.S.; Kim, Y.; Bruss, M.; Hellerstein, M.K.; Lee, H.-Y.; Samuel, V.T.; Shulman, G.I.; et al. Desnutrin/ATGL Is Regulated by AMPK and Is Required for a Brown Adipose Phenotype. Cell Metab. 2011, 13, 739–748. [Google Scholar] [CrossRef]
- Egan, D.F.; Shackelford, D.B.; Mihaylova, M.M.; Gelino, S.; Kohnz, R.A.; Mair, W.; Vasquez, D.S.; Joshi, A.; Gwinn, D.M.; Taylor, R.; et al. Phosphorylation of ULK1 (hATG1) by AMP-Activated Protein Kinase Connects Energy Sensing to Mitophagy. Science 2011, 331, 456–461. [Google Scholar] [CrossRef]
- Vakifahmetoglu-Norberg, H.; Ouchida, A.T.; Norberg, E. The role of mitochondria in metabolism and cell death. Biochem. Biophys. Res. Commun. 2017, 482, 426–431. [Google Scholar] [CrossRef]
- Abrate, A.; Lughezzani, G.; Gadda, G.M.; Lista, G.; Kinzikeeva, E.; Fossati, N.; Larcher, A.; Dell’Oglio, P.; Mistretta, F.; Buffi, N.; et al. Clinical Use of [-2]proPSA (p2PSA) and Its Derivatives (% p2PSA and Prostate Health Index) for the Detection of Prostate Cancer: A Review of the Literature. Korean J. Urol. 2014, 55, 436–445. [Google Scholar] [CrossRef]
- Beattie, D.S. Bioenergetics and oxidative metabolism. In Textbook of Biochemistry with Clinical Correlations; Devlin, T.M., Ed.; Wiley-Liss: Hoboken, NJ, USA, 2006; pp. 529–580. [Google Scholar]
- Costello, L.C.; Franklin, R.B. Prostatic fluid electrolyte composition for the screening of prostate cancer: A potential solution to a major problem. Prostate Cancer Prostatic Dis. 2008, 12, 17–24. [Google Scholar] [CrossRef][Green Version]
- Pértega-Gomes, N.; Baltazar, F. Lactate transporters in the context of prostate cancer metabolism: What do we know? Int. J. Mol. Sci. 2014, 15, 18333–18348. [Google Scholar] [CrossRef]
- Dakubo, G.D.; Parr, R.L.; Costello, L.C.; Franklin, R.B.; Thayer, R.E. Altered metabolism and mitochondrial genome in prostate cancer. J. Clin. Pathol. 2006, 59, 10–16. [Google Scholar] [CrossRef]
- Kratochvilova, M.; Raudenska, M.; Heger, Z.; Richtera, L.; Cernei, N.; Adam, V.; Babula, P.; Novakova, M.; Masarik, M.; Gumulec, J. Amino Acid Profiling of Zinc Resistant Prostate Cancer Cell Lines: Associations with Cancer Progression. Prostate 2017, 77, 604–616. [Google Scholar] [CrossRef]
- Franz, M.-C.; Anderle, P.; Bürzle, M.; Suzuki, Y.; Freeman, M.; Hediger, M.; Kovacs, G. Zinc transporters in prostate cancer. Mol. Asp. Med. 2013, 34, 735–741. [Google Scholar] [CrossRef]
- Eide, D.J. The SLC39 family of metal ion transporters. Pflug. Arch. Eur. J. Physiol. 2004, 447, 796–800. [Google Scholar] [CrossRef]
- Palmiter, R.D.; Huang, L. Efflux and compartmentalization of zinc by members of the SLC30 family of solute carriers. Pflug. Arch. Eur. J. Physiol. 2004, 447, 744–751. [Google Scholar] [CrossRef] [PubMed]
- Gaither, L.A.; Eide, D.J. Functional Expression of the Human hZIP2 Zinc Transporter. J. Biol. Chem. 2000, 275, 5560–5564. [Google Scholar] [CrossRef] [PubMed]
- Singh, C.; Chhabra, G.; Patel, A.; Chang, H.; Ahmad, N. Dietary Phytochemicals in Zinc Homeostasis: A Strategy for Prostate Cancer Management. Nutrients 2021, 13, 1867. [Google Scholar] [CrossRef] [PubMed]
- Desouki, M.M.; Geradts, J.; Milon, B.; Franklin, R.B.; Costello, L.C. hZip2 and hZip3 zinc transporters are down regulated in human prostate adenocarcinomatous glands. Mol. Cancer 2007, 6, 37. [Google Scholar] [CrossRef]
- Rishi, I.; Baidouri, H.; Abbasi, J.A.; Bullard-Dillard, R.; Kajdacsy-Balla, A.; Pestaner, J.P.; Skacel, M.; Tubbs, R.; Bagasra, O. Prostate Cancer in African American Men Is Associated With Downregulation of Zinc Transporters. Appl. Immunohistochem. Mol. Morphol. 2003, 11, 253–260. [Google Scholar] [CrossRef]
- Chen, Q. The role of zinc transporter ZIP4 in prostate carcinoma. Urol. Oncol. Semin. Orig. Investig. 2012, 30, 906–911. [Google Scholar] [CrossRef]
- Kuliyev, E.; Zhang, C.; Sui, D.; Hu, J. Zinc transporter mutations linked to acrodermatitis enteropathica disrupt function and cause mistrafficking. J. Biol. Chem. 2021, 296, 100269. [Google Scholar] [CrossRef]
- Golovine, K.; Uzzo, R.G.; Makhov, P.; Crispen, P.L.; Kunkle, D.; Kolenko, V.M. Depletion of intracellular zinc increases expression of tumorigenic cytokines VEGF, IL-6 and IL-8 in prostate cancer cells via NF-κB-dependent pathway. Prostate 2008, 68, 1443–1449. [Google Scholar] [CrossRef]
- Golovine, K. Overexpression of the zinc uptake transporter hZIP1 inhibits nuclear factor-κB and reduces the malignant potential of prostate cancer cells in vitro and in vivo. Clin. Cancer Res. 2008, 14, 5376–5384. [Google Scholar] [CrossRef]
- Feng, P.; Li, T.-L.; Guan, Z.-X.; Franklin, R.B.; Costello, L.C. Direct effect of zinc on mitochondrial apoptogenesis in prostate cells. Prostate 2002, 52, 311–318. [Google Scholar] [CrossRef]
- Wickström, M.; Larsson, R.; Nygren, P.; Gullbo, J. Aminopeptidase N (CD13) as a target for cancer chemotherapy. Cancer Sci. 2011, 102, 501–508. [Google Scholar] [CrossRef]
- Uzzo, R.G.; Crispen, P.L.; Golovine, K.; Makhov, P.; Horwitz, E.M.; Kolenko, V.M. Diverse effects of zinc on NF- B and AP-1 transcription factors: Implications for prostate cancer progression. Carcinogenesis 2006, 27, 1980–1990. [Google Scholar] [CrossRef]
- Latonen, L.; Afyounian, E.; Jylhä, A.; Nättinen, J.; Aapola, U.; Annala, M.; Kivinummi, K.K.; Tammela, T.T.L.; Beuerman, R.W.; Uusitalo, H.; et al. Integrative proteomics in prostate cancer uncovers robustness against genomic and transcriptomic aberrations during disease progression. Nat. Commun. 2018, 9, 1176. [Google Scholar] [CrossRef]
- Zadra, G.; Loda, M. Metabolic Vulnerabilities of Prostate Cancer: Diagnostic and Therapeutic Opportunities. Cold Spring Harb. Perspect. Med. 2018, 8, a030569. [Google Scholar] [CrossRef] [PubMed]
- Yue, S.; Li, J.; Lee, S.-Y.; Lee, H.J.; Shao, T.; Song, B.; Cheng, L.; Masterson, T.A.; Liu, X.; Ratliff, T.L.; et al. Cholesteryl Ester Accumulation Induced by PTEN Loss and PI3K/AKT Activation Underlies Human Prostate Cancer Aggressiveness. Cell Metab. 2014, 19, 393–406. [Google Scholar] [CrossRef] [PubMed]
- Lloyd, M.D. α-Methylacyl-CoA racemase (AMACR): Metabolic enzyme, drug metabolizer and cancer marker P504S. Prog. Lipid Res. 2013, 52, 220–230. [Google Scholar] [CrossRef] [PubMed]
- Swinnen, J.V.; Roskams, T.; Joniau, S.; Van Poppel, H.; Oyen, R.; Baert, L.; Heyns, W.; Verhoeven, G. Overexpression of fatty acid synthase is an early and common event in the development of prostate cancer. Int. J. Cancer 2002, 98, 19–22. [Google Scholar] [CrossRef] [PubMed]
- Swanson, M.G.; Keshari, K.R.; Tabatabai, Z.L.; Simko, J.P.; Shinohara, K.; Carroll, P.R.; Zektzer, A.S.; Kurhanewicz, J. Quantification of choline- and ethanolamine-containing metabolites in human prostate tissues using1H HR-MAS total correlation spectroscopy. Magn. Reson. Med. 2008, 60, 33–40. [Google Scholar] [CrossRef] [PubMed]
- Van de Sande, T. High-level expression of fatty acid synthase in human prostate cancer tissues is linked to activation and nuclear localization of Akt/PKB. J. Pathol. 2005, 206, 214–219. [Google Scholar] [CrossRef]
- Swinnen, J.V. Androgens, lipogenesis and prostate cancer. J. Steroid Biochem. Mol. Biol. 2004, 92, 273–279. [Google Scholar] [CrossRef]
- Ettinger, S.L.; Sobel, R.; Whitmore, T.G.; Akbari, M.; Bradley, D.R.; Gleave, M.E.; Nelson, C.C. Dysregulation of Sterol Response Element-Binding Proteins and Downstream Effectors in Prostate Cancer during Progression to Androgen Independence. Cancer Res. 2004, 64, 2212–2221. [Google Scholar] [CrossRef]
- Guo, D.; Bell, E.; Mischel, P.; Chakravarti, A. Targeting SREBP-1-driven Lipid Metabolism to Treat Cancer. Curr. Pharm. Des. 2014, 20, 2619–2626. [Google Scholar] [CrossRef]
- Huang, W.-C.; Li, X.; Liu, J.; Lin, J.; Chung, L.W. Activation of Androgen Receptor, Lipogenesis, and Oxidative Stress Converged by SREBP-1 Is Responsible for Regulating Growth and Progression of Prostate Cancer Cells. Mol. Cancer Res. 2012, 10, 133–142. [Google Scholar] [CrossRef]
- De Piano, M. Exploring a role for fatty acid synthase in prostate cancer cell migration. Small GTPases 2021, 12, 265–272. [Google Scholar] [CrossRef]
- Roberts, M.J.; Schirra, H.J.; Lavin, M.; Gardiner, R.A. Metabolomics: A Novel Approach to Early and Noninvasive Prostate Cancer Detection. Korean J. Urol. 2011, 52, 79–89. [Google Scholar] [CrossRef]
- Jamnagerwalla, J.; Howard, L.E.; Allott, E.H.; Vidal, A.C.; Moreira, D.M.; Castro-Santamaria, R.; Andriole, G.L.; Freeman, M.R.; Freedland, S.J. Serum cholesterol and risk of high-grade prostate cancer: Results from the REDUCE study. Prostate Cancer Prostatic Dis. 2018, 21, 252–259. [Google Scholar] [CrossRef]
- Platz, E.A.; Till, C.; Goodman, P.J.; Parnes, H.L.; Figg, W.D.; Albanes, D.; Neuhouser, M.L.; Klein, E.A.; Thompson, I.M.; Kristal, A. Men with Low Serum Cholesterol Have a Lower Risk of High-Grade Prostate Cancer in the Placebo Arm of the Prostate Cancer Prevention Trial. Cancer Epidemiol. Biomark. Prev. 2009, 18, 2807–2813. [Google Scholar] [CrossRef]
- Pelton, K.; Freeman, M.R.; Solomon, K.R. Cholesterol and prostate cancer. Curr. Opin. Pharmacol. 2012, 12, 751–759. [Google Scholar] [CrossRef]
- Wang, X.; Sun, B.; Wei, L.; Jian, X.; Shan, K.; He, Q.; Huang, F.; Ge, X.; Gao, X.; Feng, N.; et al. Cholesterol and saturated fatty acids synergistically promote the malignant progression of prostate cancer. Neoplasia 2022, 24, 86–97. [Google Scholar] [CrossRef]
- Cheng, Y.; Meng, Y.; Li, S.; Cao, D.; Ben, S.; Qin, C.; Hua, L.; Cheng, G. Genetic variants in the cholesterol biosynthesis pathway genes and risk of prostate cancer. Gene 2021, 774, 145432. [Google Scholar] [CrossRef]
- Garrido, M.M.; Marta, J.C.; Ribeiro, R.M.; Pinheiro, L.C.; Guimarães, J.T. Serum lipids and prostate cancer. J. Clin. Lab. Anal. 2021, 35, e23705. [Google Scholar] [CrossRef]
- Xiong, K. The cholesterol esterification inhibitor avasimibe suppresses tumour proliferation and metastasis via the E2F-1 signalling pathway in prostate cancer. Cancer Cell Int. 2021, 21, 461. [Google Scholar] [CrossRef]
- Kim, S.; Kim, K.-H. Modulation of Cholesterol Metabolism Improves Response to Enzalutamide Treatment in Prostate Cancer. Curr. Dev. Nutr. 2021, 5, 269. [Google Scholar] [CrossRef]
- Pan, T.; Lin, S.-C.; Lee, Y.-C.; Yu, G.; Song, J.H.; Pan, J.; Titus, M.; Satcher, R.L.; Panaretakis, T.; Logothetis, C.; et al. Statins reduce castration-induced bone marrow adiposity and prostate cancer progression in bone. Oncogene 2021, 40, 4592–4603. [Google Scholar] [CrossRef] [PubMed]
- Kalogirou, C.; Linxweiler, J.; Schmucker, P.; Snaebjornsson, M.T.; Schmitz, W.; Wach, S.; Krebs, M.; Hartmann, E.; Puhr, M.; Müller, A.; et al. MiR-205-driven downregulation of cholesterol biosynthesis through SQLE-inhibition identifies therapeutic vulnerability in aggressive prostate cancer. Nat. Commun. 2021, 12, 5066. [Google Scholar] [CrossRef] [PubMed]
- Eckhardt, C.; Sbiera, I.; Krebs, M.; Sbiera, S.; Spahn, M.; Kneitz, B.; Joniau, S.; Fassnacht, M.; Kübler, H.; Weigand, I.; et al. High expression of Sterol-O-Acyl transferase 1 (SOAT1), an enzyme involved in cholesterol metabolism, is associated with earlier biochemical recurrence in high risk prostate cancer. Prostate Cancer Prostatic Dis. 2021, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Zabielska, J.; Sledzinski, T.; Stelmanska, E. Acyl-Coenzyme A: Cholesterol Acyltransferase Inhibition in Cancer Treatment. Anticancer Res. 2019, 39, 3385–3394. [Google Scholar] [CrossRef]
- Rogers, M.A. Acyl-CoA:cholesterol acyltransferases (ACATs/SOATs): Enzymes with multiple sterols as substrates and as activators. J. Steroid Biochem. Mol. Biol. 2015, 151, 102–107. [Google Scholar] [CrossRef]
- Yang, J.; Wang, L.; Jia, R. Role of de novo cholesterol synthesis enzymes in cancer. J. Cancer 2020, 11, 1761–1767. [Google Scholar] [CrossRef]
- Krycer, J.R.; Brown, A.J. Cholesterol accumulation in prostate cancer: A classic observation from a modern perspective. Biochim. Biophys. Acta 2013, 1835, 219–229. [Google Scholar] [CrossRef]
- Krycer, J.R.; Phan, L.; Brown, A.J. A key regulator of cholesterol homoeostasis, SREBP-2, can be targeted in prostate cancer cells with natural products. Biochem. J. 2012, 446, 191–201. [Google Scholar] [CrossRef]
- Ayyagari, V.N.; Wang, X.; Diaz-Sylvester, P.L.; Groesch, K.; Brard, L. Assessment of acyl-CoA cholesterol acyltransferase (ACAT-1) role in ovarian cancer progression—An in vitro study. PLoS ONE 2020, 15, e0228024. [Google Scholar] [CrossRef]
- Patel, D.; Ahmad, F.; Kambach, D.M.; Sun, Q.; Halim, A.S.; Kramp, T.; Camphausen, K.A.; Stommel, J.M. LXRβ controls glioblastoma cell growth, lipid balance, and immune modulation independently of ABCA1. Sci. Rep. 2019, 9, 15458. [Google Scholar] [CrossRef]
- Yi, X.; Li, Y.; Hu, X.; Wang, F.; Liu, T. Changes in phospholipid metabolism in exosomes of hormone-sensitive and hor-mone-resistant prostate cancer cells. J. Cancer 2021, 12, 2893–2902. [Google Scholar] [CrossRef]
- Chen, J.; Guccini, I.; Di Mitri, D.; Brina, D.; Revandkar, A.; Sarti, M.; Pasquini, E.; Alajati, A.; Pinton, S.; Losa, M.; et al. Compartmentalized activities of the pyruvate dehydrogenase complex sustain lipogenesis in prostate cancer. Nat. Genet. 2018, 50, 219–228. [Google Scholar] [CrossRef]
- Zhang, Z.; Hou, X.; Shao, C.; Li, J.; Cheng, J.-X.; Kuang, S.; Ahmad, N.; Ratliff, T.; Liu, X. Plk1 Inhibition Enhances the Efficacy of Androgen Signaling Blockade in Castration-Resistant Prostate Cancer. Cancer Res. 2014, 74, 6635–6647. [Google Scholar] [CrossRef]
- Creative Diagnostics (CD). PI3K-AKT Signaling Pathway. 2022. Available online: https://www.creative-diagnostics.com/PI3K-AKT-Signaling-Pathway.htm (accessed on 14 January 2022).
- Li, X.; Chen, Y.-T.; Hu, P.; Huang, W.-C. Fatostatin Displays High Antitumor Activity in Prostate Cancer by Blocking SREBP-Regulated Metabolic Pathways and Androgen Receptor Signaling. Mol. Cancer Ther. 2014, 13, 855–866. [Google Scholar] [CrossRef]
- Wei, S.; Machamer, C.E.; Espenshade, P.J. Fatostatin blocks ER exit of SCAP but inhibits cell growth in a SCAP-independent manner. J. Lipid Res. 2016, 57, 1564–1573. [Google Scholar] [CrossRef]
- Vettenranta, A.; Murtola, T.J.; Raitanen, J.; Raittinen, P.; Talala, K.; Taari, K.; Stenman, U.-H.; Tammela, T.L.J.; Auvinen, A. Outcomes of Screening for Prostate Cancer Among Men Who Use Statins. JAMA Oncol. 2021, 8, 61. [Google Scholar] [CrossRef]
- Jiménez-Vacas, J.M.; Herrero-Aguayo, V.; Montero-Hidalgo, A.J.; Sáez-Martínez, P.; Gómez-Gómez, E.; León-González, A.J.; Fuentes-Fayos, A.C.; Yubero-Serrano, E.M.; Requena-Tapia, M.J.; López, M.; et al. Clinical, Cellular, and Molecular Evidence of the Additive Antitumor Effects of Biguanides and Statins in Prostate Cancer. J. Clin. Endocrinol. Metab. 2021, 106, e696–e710. [Google Scholar] [CrossRef]
- Jeong, I.G.; Lim, B.; Yun, S.-C.; Lim, J.H.; Hong, J.H.; Kim, C.-S. Adjuvant Low-dose Statin Use after Radical Prostatectomy: The PRO-STAT Randomized Clinical Trial. Clin. Cancer Res. 2021, 27, 5004–5011. [Google Scholar] [CrossRef]
- Prabhu, N.; Kapur, N.; Catalona, W.; Leikin, R.; Helenowski, I.; Jovanovich, B.; Gurley, M.; Okwuosa, T.M.; Kuzel, T.M. Statin use and risk of prostate cancer biochemical recurrence after radical prostatectomy. Urol. Oncol. Semin. Orig. Investig. 2021, 39, 130.e9–130.e15. [Google Scholar] [CrossRef]
- Hamilton, R.J.; Ding, K.; Crook, J.M.; O’Callaghan, C.J.; Higano, C.S.; Dearnaley, D.P.; Horwitz, E.M.; Goldenberg, S.L.; Gospodarowicz, M.K.; Klotz, L. The Association Between Statin Use and Outcomes in Patients Initiating Androgen Deprivation Therapy. Eur. Urol. 2021, 79, 446–452. [Google Scholar] [CrossRef]
- Schnier, J.B.; Nishi, K.; Gumerlock, P.H.; Gorin, F.A.; Bradbury, E.M. Glycogen synthesis correlates with andro-gen-dependent growth arrest in prostate cancer. BMC Urol. 2005, 5, 6. [Google Scholar] [CrossRef]
- Pelletier, J.; Bellot, G.; Gounon, P.; Lacas-Gervais, S.; Pouysségur, J.; Mazure, N.M. Glycogen Synthesis is Induced in Hypoxia by the Hypoxia-Inducible Factor and Promotes Cancer Cell Survival. Front. Oncol. 2012, 2, 18. [Google Scholar] [CrossRef]
- Schwartz, N.B. Carbohydrates metabolism II: Special pathways and glycoconjugates. In Textbook of Biochemistry with Clinical Applications; Devlin, T.M., Ed.; Wiley-Liss: Hoboken, NJ, USA, 2006; pp. 637–660. [Google Scholar]
- Yang, H.C.; Wu, Y.H.; Yen, W.C.; Liu, H.Y.; Hwang, T.L.; Stern, A.; Chiu, D.T.Y. The redox role of G6PD in cell growth, cell death, and cancer. Cells 2019, 8, 1055. [Google Scholar] [CrossRef]
- Tsouko, E. Regulation of the pentose phosphate pathway by an androgen receptor–mTOR-mediated mechanism and its role in prostate cancer cell growth. Oncogenesis 2014, 3, e103. [Google Scholar] [CrossRef]
- Liu, X.; Olszewski, K.; Zhang, Y.; Lim, E.W.; Shi, J.; Zhang, X.; Zhang, J.; Lee, H.; Koppula, P.; Lei, G.; et al. Cystine transporter regulation of pentose phosphate pathway dependency and disulfide stress exposes a targetable metabolic vulnerability in cancer. Nat. Cell Biol. 2020, 22, 476–486. [Google Scholar] [CrossRef]
- Gillis, J.L.; Hinneh, J.A.; Ryan, N.K.; Irani, S.; Moldovan, M.; Quek, L.-E.; Shrestha, R.K.; Hanson, A.R.; Xie, J.; Hoy, A.J.; et al. A feedback loop between the androgen receptor and 6-phosphogluoconate dehydrogenase (6PGD) drives prostate cancer growth. ELife 2021, 10, e62592. [Google Scholar] [CrossRef]
- Ge, T.; Yang, J.; Zhou, S.; Wang, Y.; Li, Y.; Tong, X. The Role of the Pentose Phosphate Pathway in Diabetes and Cancer. Front. Endocrinol. 2020, 11, 365. [Google Scholar] [CrossRef]
- Tedeschi, P.M.; Markert, E.K.; Gounder, M.; Lin, H.; Dvorzhinski, D.; Dolfi, S.C.; Chan, L.L.-Y.; Qiu, J.; DiPaola, R.S.; Hirshfield, K.M.; et al. Contribution of serine, folate and glycine metabolism to the ATP, NADPH and purine requirements of cancer cells. Cell Death Dis. 2013, 4, e877. [Google Scholar] [CrossRef]
- Chaneton, B.; Hillmann, P.; Zheng, L.; Martin, A.C.L.; Maddocks, O.D.K.; Chokkathukalam, A.; Coyle, J.E.; Jankevics, A.; Holding, F.P.; Vousden, K.H.; et al. Serine is a natural ligand and allosteric activator of pyruvate kinase M2. Nature 2012, 491, 458–462. [Google Scholar] [CrossRef]
- Cardoso, H.; Figueira, M.; Madureira, P.; Socorro, S. PO-255 The pivotal role of glutaminolysis in prostate cancer cells and its regulation by androgens. ESMO Open 2018, 3, A120. [Google Scholar] [CrossRef]
- Cardoso, H.J.; Figueira, M.I.; Vaz, C.V.; Carvalho, T.M.A.; Brás, L.A.; Madureira, P.A.; Oliveira, P.J.; Sardão, V.A.; Socorro, S. Glutaminolysis is a metabolic route essential for survival and growth of prostate cancer cells and a target of 5α-dihydrotestosterone regulation. Cell. Oncol. 2021, 44, 385–403. [Google Scholar] [CrossRef] [PubMed]
- Koochekpour, S.; Majumdar, S.; Azabdaftari, G.; Attwood, K.; Scioneaux, R.; Subramani, D.; Manhardt, C.; Lorusso, G.D.; Willard, S.S.; Thompson, H.; et al. Serum Glutamate Levels Correlate with Gleason Score and Glutamate Blockade Decreases Proliferation, Migration, and Invasion and Induces Apoptosis in Prostate Cancer Cells. Clin. Cancer Res. 2012, 18, 5888–5901. [Google Scholar] [CrossRef] [PubMed]
- White, M.A.; Lin, C.; Rajapakshe, K.; Dong, J.; Shi, Y.; Tsouko, E.; Mukhopadhyay, R.; Jasso, D.; Dawood, W.; Coarfa, C.; et al. Glutamine Transporters Are Targets of Multiple Oncogenic Signaling Pathways in Prostate Cancer. Mol. Cancer Res. 2017, 15, 1017–1028. [Google Scholar] [CrossRef] [PubMed]
- Dorai, T.; Dorai, B.; Pinto, J.T.; Grasso, M.; Cooper, A.J.L. High Levels of Glutaminase II Pathway Enzymes in Normal and Cancerous Prostate Suggest a Role in ‘Glutamine Addiction’. Biomolecules 2019, 10, 2. [Google Scholar] [CrossRef] [PubMed]
- Zacharias, N.M.; McCullough, C.; Shanmugavelandy, S.; Lee, J.; Lee, Y.; Dutta, P.; McHenry, J.; Nguyen, L.; Norton, W.; Jones, L.W.; et al. Metabolic Differences in Glutamine Utilization Lead to Metabolic Vulnerabilities in Prostate Cancer. Sci. Rep. 2017, 7, 16159. [Google Scholar] [CrossRef] [PubMed]
- Iacobazzi, V.; Infantino, V. Citrate–new functions for an old metabolite. Biol. Chem. 2014, 395, 387–399. [Google Scholar] [CrossRef] [PubMed]
- Arora, H.; Panara, K.; Kuchakulla, M.; Kulandavelu, S.; Burnstein, K.L.; Schally, A.V.; Hare, J.M.; Ramasamy, R. Alterations of tumor microenvironment by nitric oxide impedes castration-resistant prostate cancer growth. Proc. Natl. Acad. Sci. USA 2018, 115, 11298–11303. [Google Scholar] [CrossRef]
- Bhowmick, R.; Girotti, A.W. Pro-survival and pro-growth effects of stress-induced nitric oxide in a prostate cancer photo-dynamic therapy model. Cancer Lett. 2014, 343, 115–122. [Google Scholar] [CrossRef]
- D’Aniello, C.; Patriarca, E.J.; Phang, J.M.; Minchiotti, G. Proline Metabolism in Tumor Growth and Metastatic Progression. Front. Oncol. 2020, 10, 776. [Google Scholar] [CrossRef]
- Coomes, M.W. Amino acid metabolism. In Textbook of Biochemistry with Clinical Applications; Devlin, T.M., Ed.; Wiley-Liss: Hoboken, NJ, USA, 2006; pp. 743–787. [Google Scholar]
- Donkena, K.V.; Yuan, H.; Young, C.Y. Vitamin Bs, one carbon metabolism and prostate cancer. Mini-Rev. Med. Chem. 2010, 10, 1385–1392. [Google Scholar] [CrossRef]
- Malviya, G.; Patel, R.; Salji, M.; Martinez, R.S.; Repiscak, P.; Mui, E.; Champion, S.; Mrowinska, A.; Johnson, E.; AlRasheedi, M.; et al. 18F-Fluciclovine PET metabolic imaging reveals prostate cancer tumour heterogeneity associated with disease resistance to androgen deprivation therapy. EJNMMI Res. 2020, 10, 143. [Google Scholar] [CrossRef]
- Saito, Y.; Soga, T. Amino acid transporters as emerging therapeutic targets in cancer. Cancer Sci. 2021, 112, 2958–2965. [Google Scholar] [CrossRef]
- Xu, M.; Sakamoto, S.; Matsushima, J.; Kimura, T.; Ueda, T.; Mizokami, A.; Kanai, Y.; Ichikawa, T. Up-Regulation of LAT1 during Antiandrogen Therapy Contributes to Progression in Prostate Cancer Cells. J. Urol. 2016, 195, 1588–1597. [Google Scholar] [CrossRef]
- Martinez, R.S.; Salji, M.J.; Rushworth, L.; Ntala, C.; Blanco, G.R.; Hedley, A.; Clark, W.; Peixoto, P.; Hervouet, E.; Renaude, E.; et al. SLFN5 Regulates LAT1-Mediated mTOR Activation in Castration-Resistant Prostate Cancer. Cancer Res. 2021, 81, 3664–3678. [Google Scholar] [CrossRef]

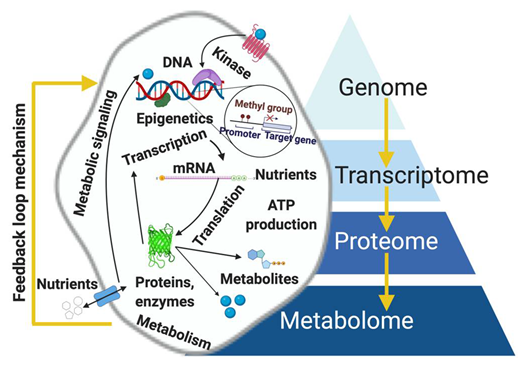
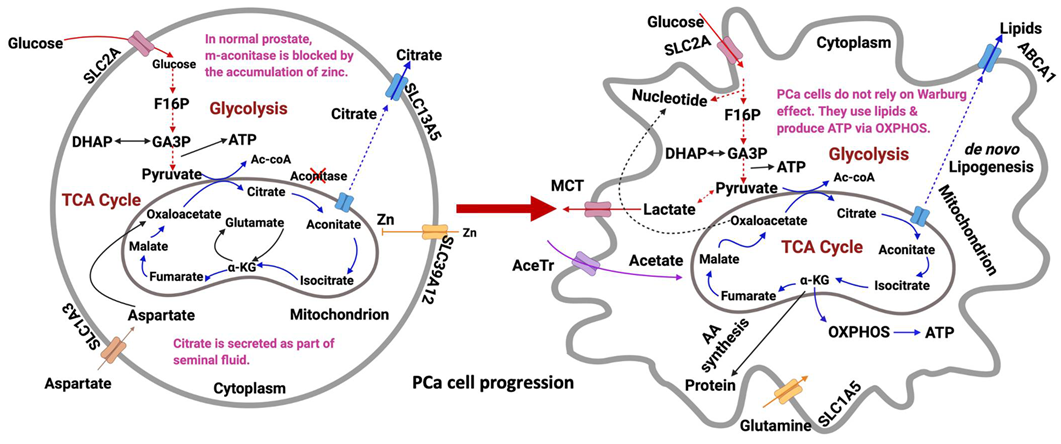

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Resurreccion, E.P.; Fong, K.-w. The Integration of Metabolomics with Other Omics: Insights into Understanding Prostate Cancer. Metabolites 2022, 12, 488. https://doi.org/10.3390/metabo12060488
Resurreccion EP, Fong K-w. The Integration of Metabolomics with Other Omics: Insights into Understanding Prostate Cancer. Metabolites. 2022; 12(6):488. https://doi.org/10.3390/metabo12060488
Chicago/Turabian StyleResurreccion, Eleazer P., and Ka-wing Fong. 2022. "The Integration of Metabolomics with Other Omics: Insights into Understanding Prostate Cancer" Metabolites 12, no. 6: 488. https://doi.org/10.3390/metabo12060488
APA StyleResurreccion, E. P., & Fong, K.-w. (2022). The Integration of Metabolomics with Other Omics: Insights into Understanding Prostate Cancer. Metabolites, 12(6), 488. https://doi.org/10.3390/metabo12060488






