Gut Microbiota and Serum Metabolome in Elite Cross-Country Skiers: A Controlled Study
Abstract
:1. Introduction
2. Results
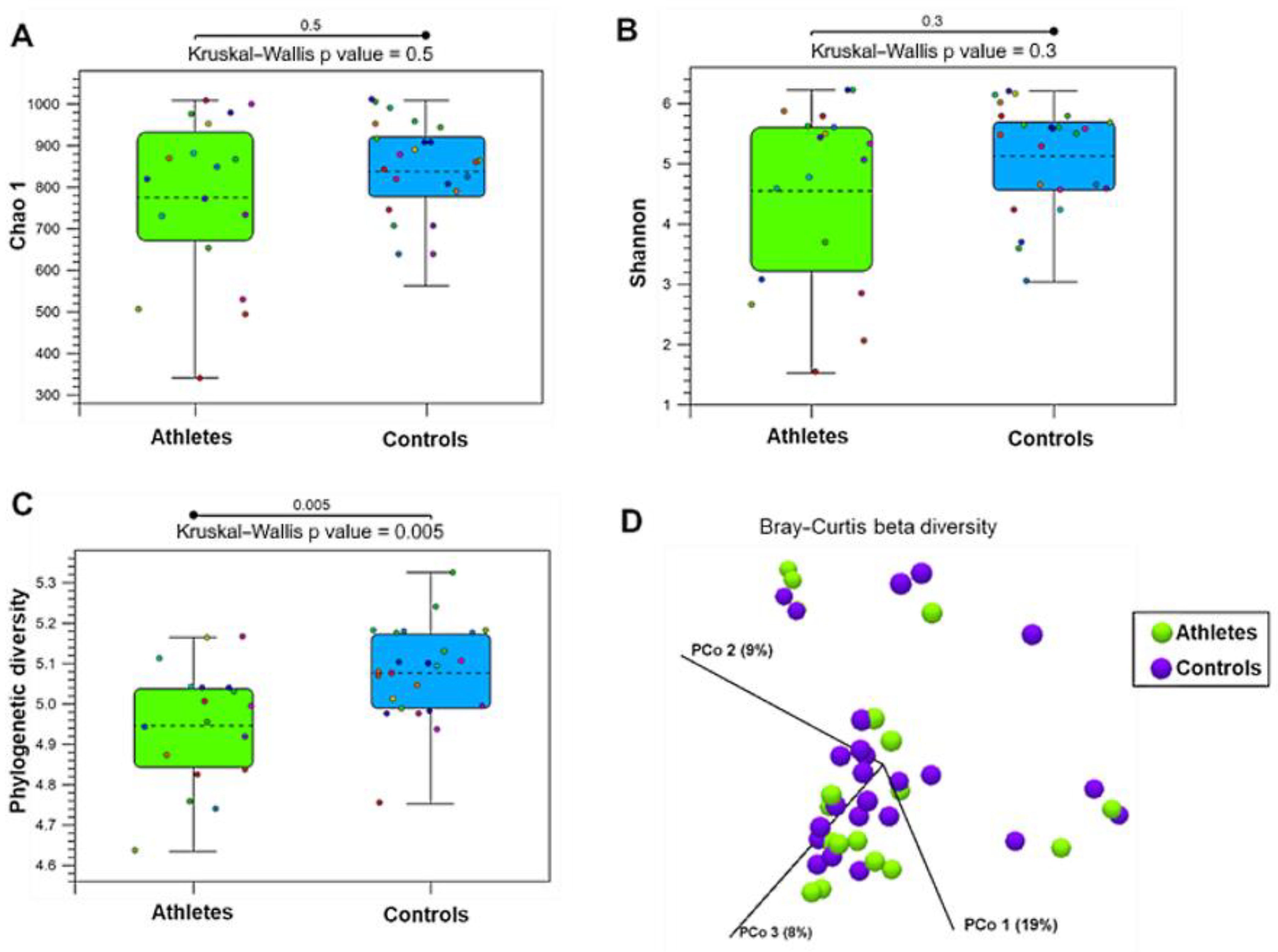
2.1. Gut Microbiota Diversity
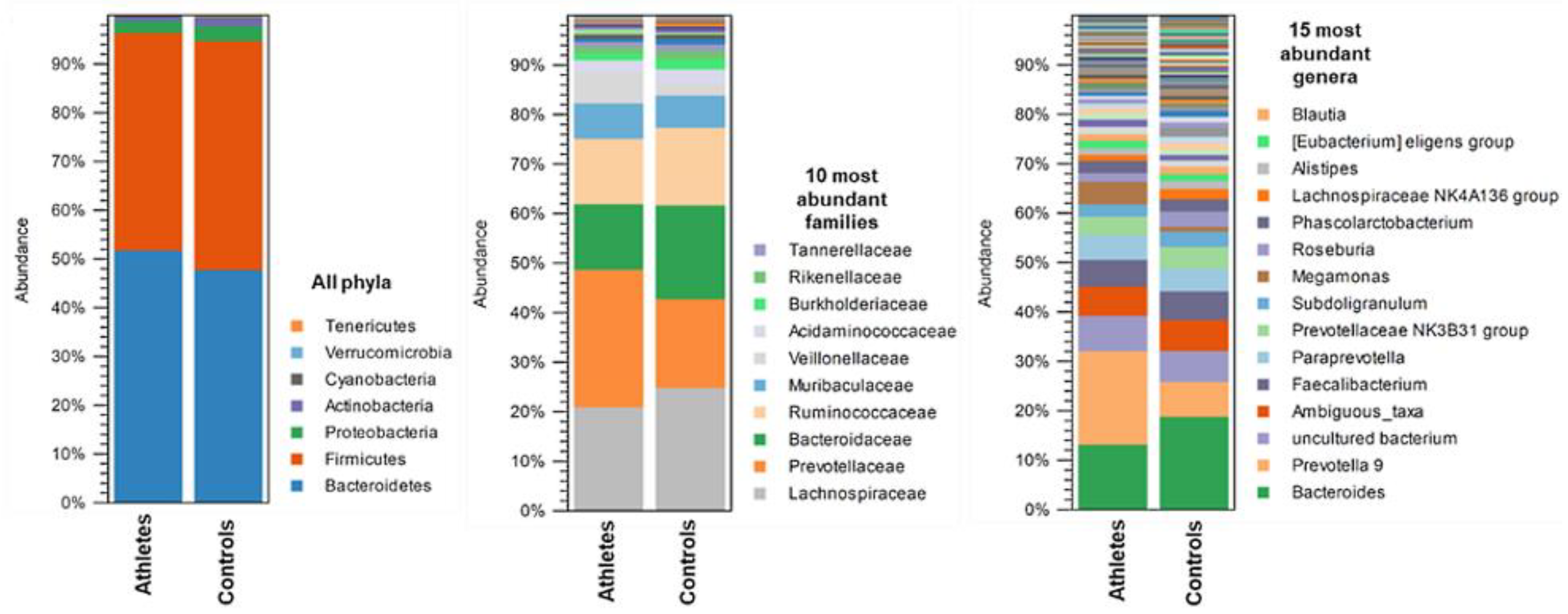
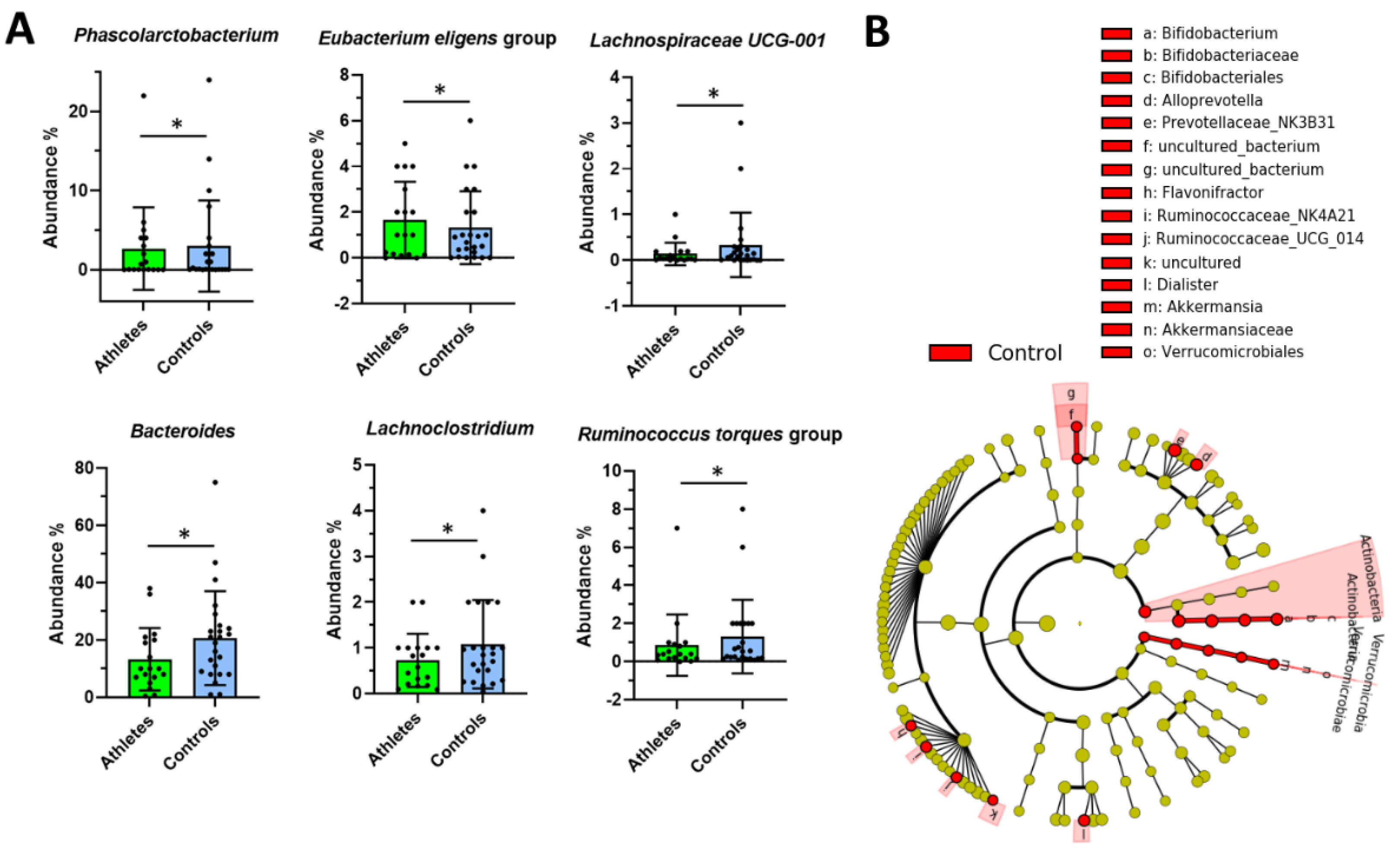
2.2. Gut Microbiota Composition
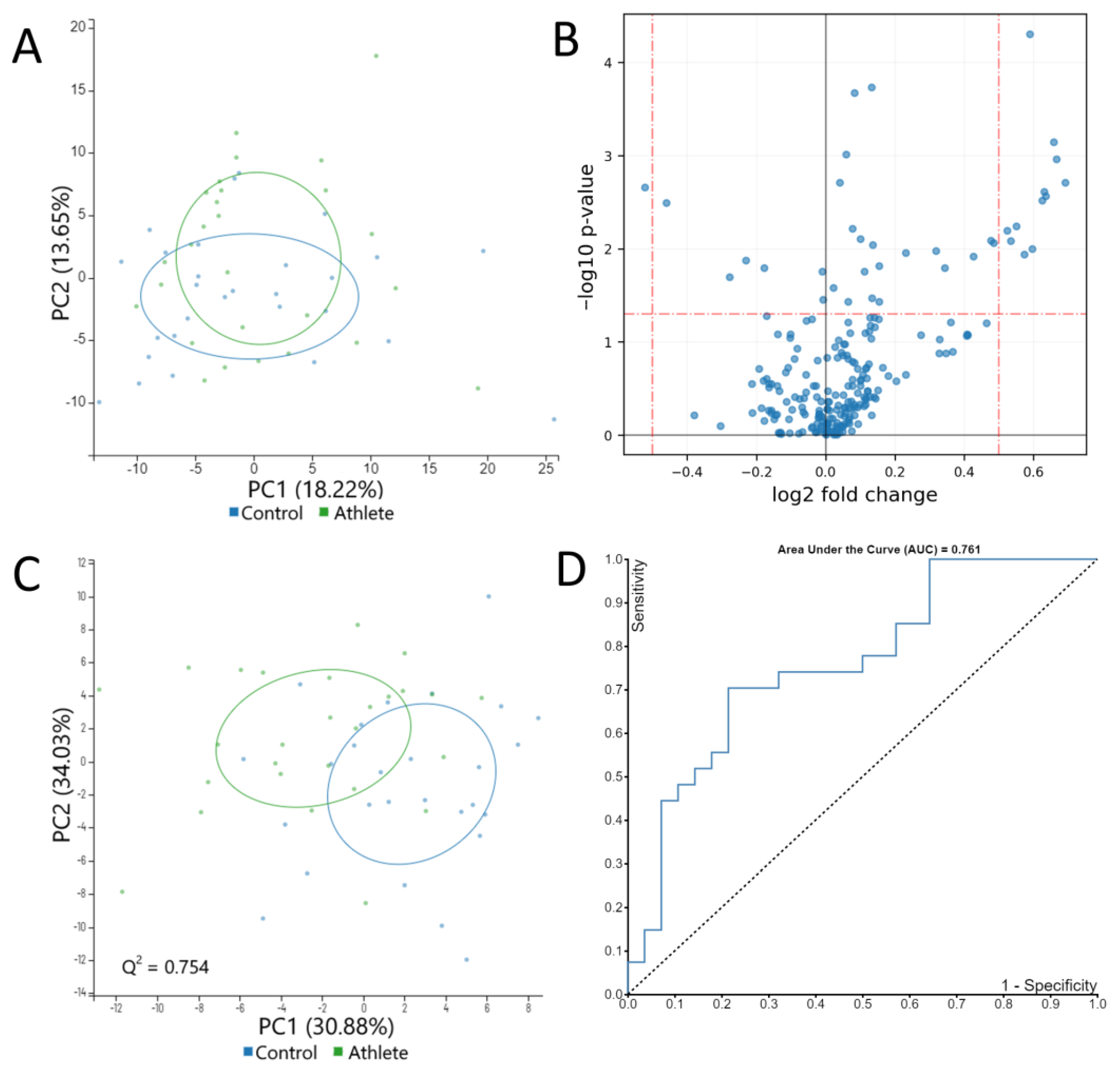
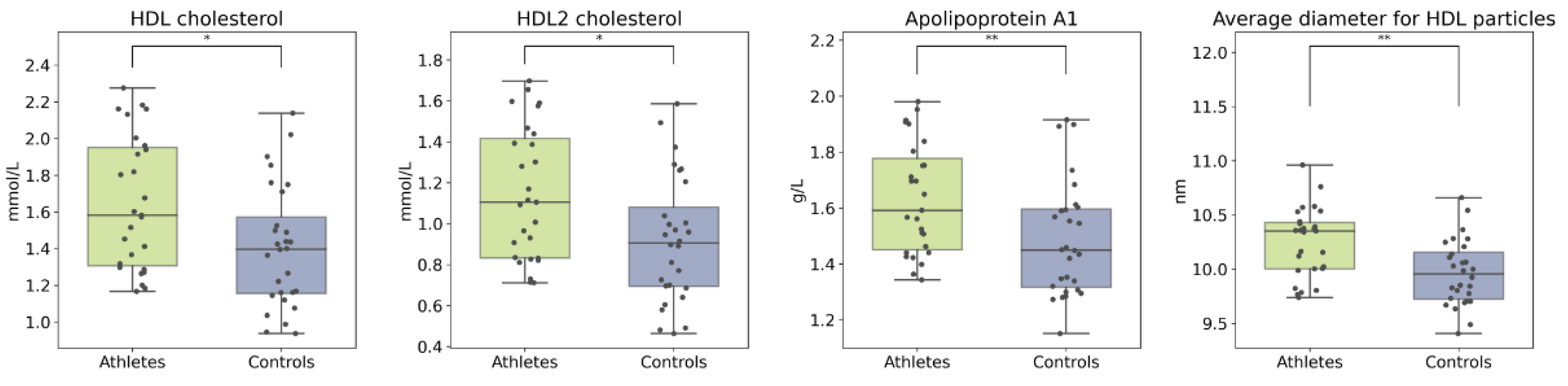
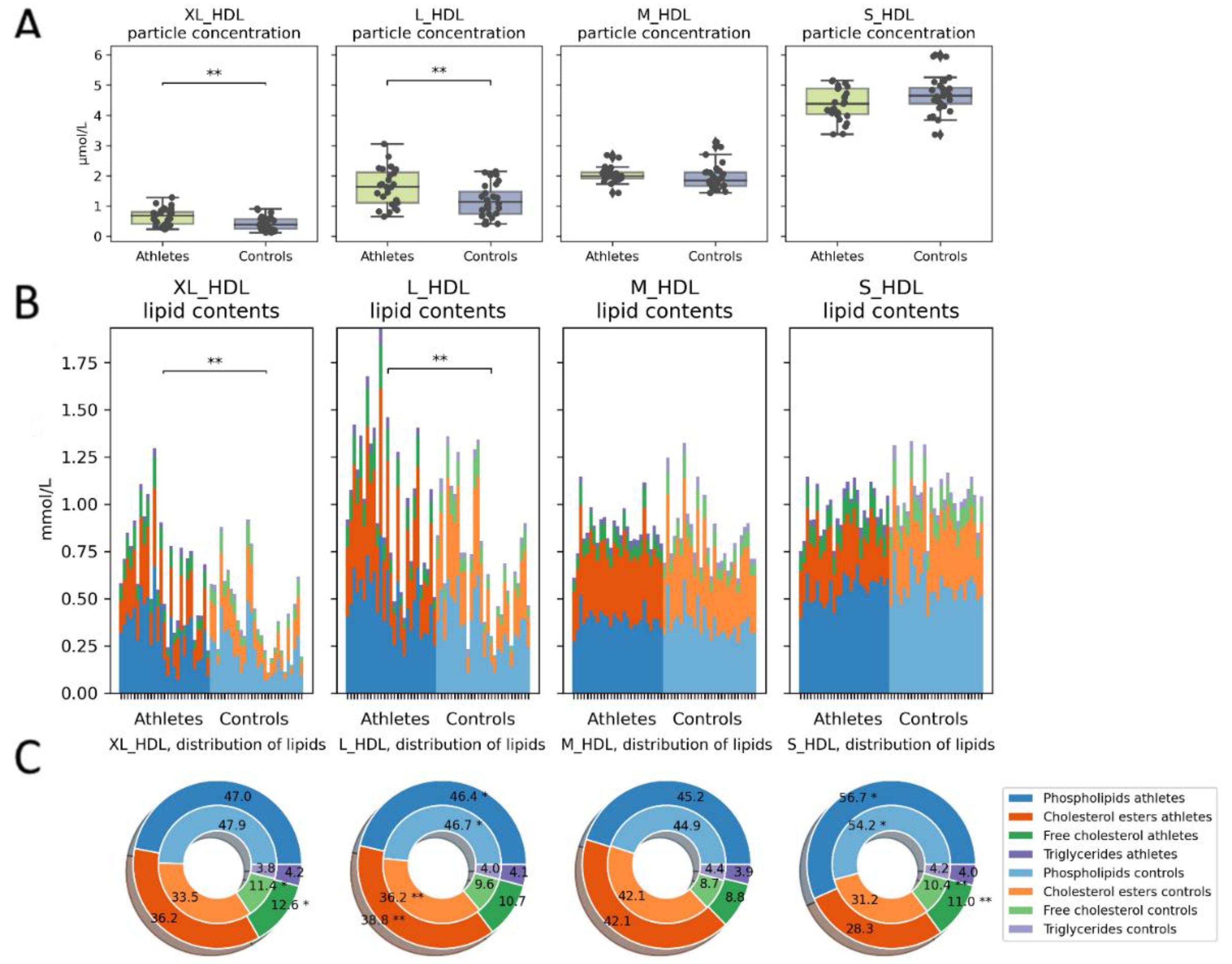
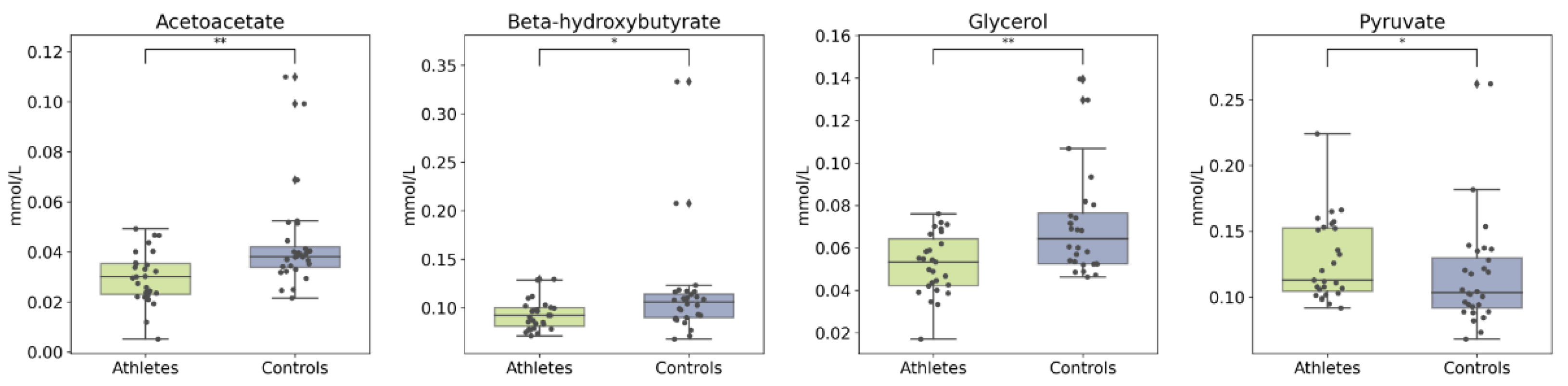
2.3. Serum Metabolome
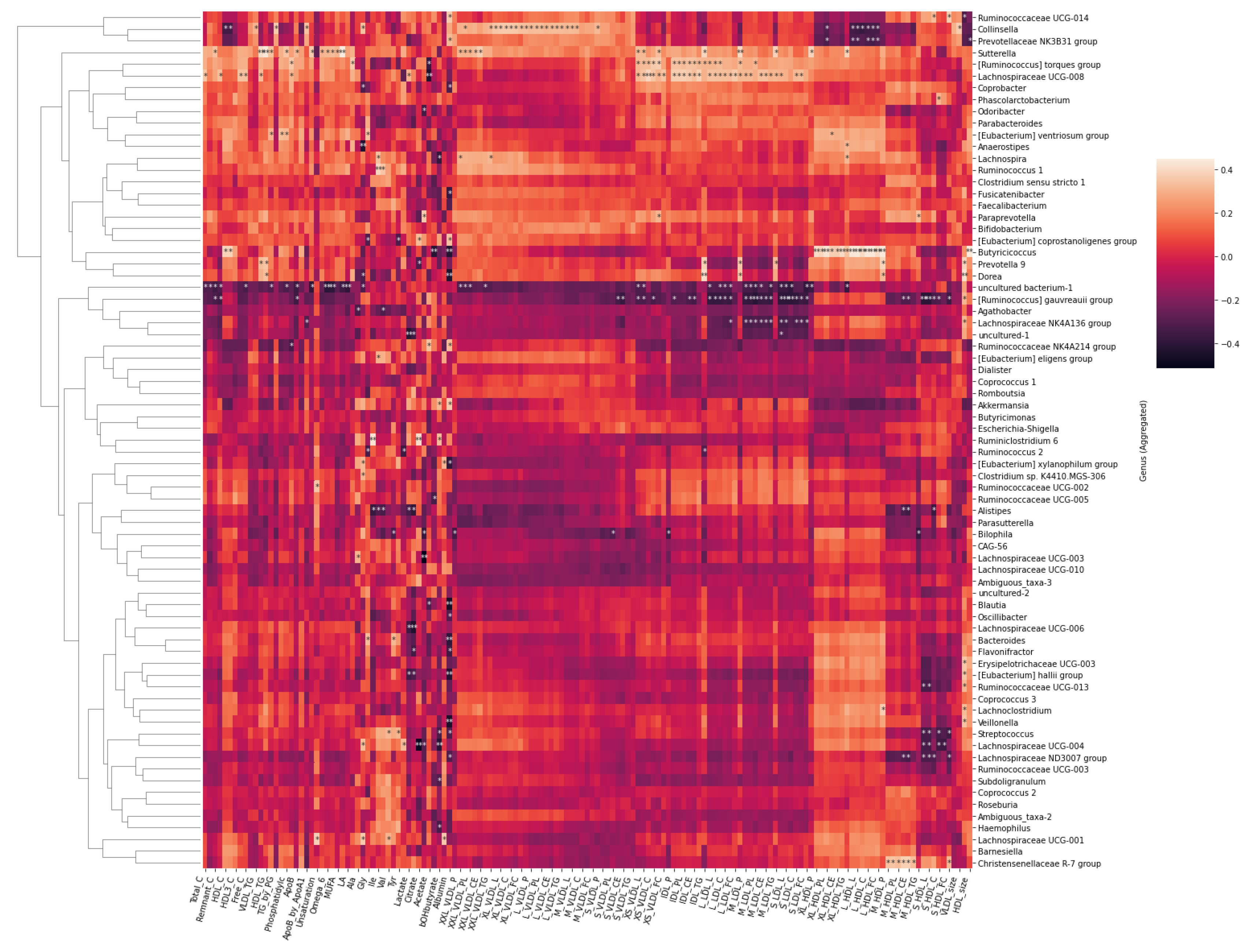
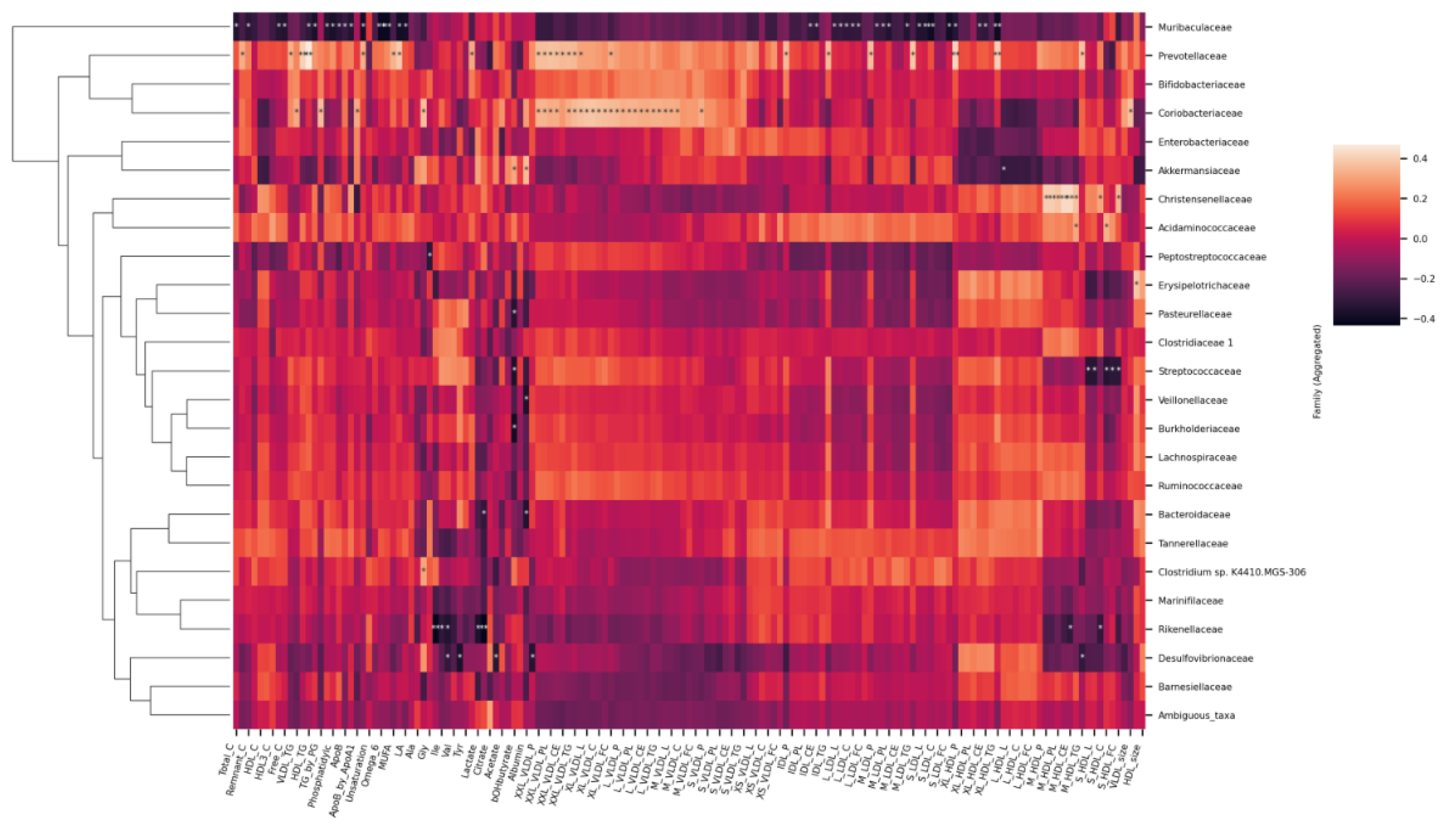
2.4. Associations between the Metabolites and Gut Microbiota
3. Discussion
4. Materials and Methods
4.1. Study Design and Population
4.2. Fecal Sample Collection and Microbial DNA Extraction
4.3. 16 S rRNA Gene Sequencing and Sequence Data Processing
4.4. Gut Microbiota Composition Analyses
4.5. Metabolite Analyses
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Qin, J.; Li, R.; Raes, J.; Arumugam, M.; Burgdorf, K.S.; Manichanh, C.; Nielsen, T.; Pons, N.; Levenez, F.; Yamada, T.; et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 2010, 464, 59–65. [Google Scholar] [CrossRef] [Green Version]
- Sender, R.; Fuchs, S.; Milo, R. Revised Estimates for the Number of Human and Bacteria Cells in the Body. PLOS Biol. 2016, 14, e1002533. [Google Scholar] [CrossRef] [Green Version]
- Salvi, P.S.; Cowles, R.A. Butyrate and the Intestinal Epithelium: Modulation of Proliferation and Inflammation in Homeostasis and Disease. Cells 2021, 10, 1775. [Google Scholar] [CrossRef] [PubMed]
- Chambers, E.S.; Byrne, C.S.; Aspey, K.; Chen, Y.; Khan, S.; Morrison, D.J.; Frost, G. Acute oral sodium propionate supplementation raises resting energy expenditure and lipid oxidation in fasted humans. Diabetes Obes. Metab. 2018, 20, 1034–1039. [Google Scholar] [CrossRef] [PubMed]
- Bongiovanni, T.; Yin, M.O.L.; Heaney, L. The Athlete and Gut Microbiome: Short-chain Fatty Acids as Potential Ergogenic Aids for Exercise and Training. Int. J. Sports Med. 2021, 42, 1143–1158. [Google Scholar] [CrossRef] [PubMed]
- Lensu, S.; Pekkala, S. Gut Microbiota, Microbial Metabolites and Human Physical Performance. Metabolites 2021, 11, 716. [Google Scholar] [CrossRef]
- Cox, A.J.; West, N.P.; Cripps, A.W. Obesity, inflammation, and the gut microbiota. Lancet Diabetes Endocrinol. 2015, 3, 207–215. [Google Scholar] [CrossRef]
- Mohr, A.E.; Jäger, R.; Carpenter, K.C.; Kerksick, C.M.; Purpura, M.; Townsend, J.R.; West, N.P.; Black, K.; Gleeson, M.; Pyne, D.B.; et al. The athletic gut microbiota. J. Int. Soc. Sports Nutr. 2020, 17, 1–33. [Google Scholar] [CrossRef]
- Dorelli, B.; Gallè, F.; de Vito, C.; Duranti, G.; Iachini, M.; Zaccarin, M.; Standoli, J.P.; Ceci, R.; Romano, F.; Liguori, G.; et al. Can Physical Activity Influence Human Gut Microbiota Composition Independently of Diet? A Systematic Review. Nutrients 2021, 13, 1890. [Google Scholar] [CrossRef]
- Makin, S. Do microbes affect athletic performance? Nature 2021, 592, S17–S19. [Google Scholar] [CrossRef]
- Barton, W.; Penney, N.C.; Cronin, O.; Garcia-Perez, I.; Molloy, M.G.; Holmes, E.; Shanahan, F.; Cotter, P.D.; O’Sullivan, O. The microbiome of professional athletes differs from that of more sedentary subjects in composition and particularly at the functional metabolic level. Gut 2018, 67, 625–633. [Google Scholar] [CrossRef]
- Flint, H.J.; Scott, K.P.; Duncan, S.H.; Louis, P.; Forano, E. Microbial degradation of complex carbohydrates in the gut. Gut Microbes 2012, 3, 289–306. [Google Scholar] [CrossRef] [Green Version]
- Valentino, T.R.; Vechetti, I.J.; Mobley, C.B.; Dungan, C.M.; Golden, L.; Goh, J.; McCarthy, J.J. Dysbiosis of the gut microbiome impairs mouse skeletal muscle adaptation to exercise. J. Physiol. 2021, 599, 4845–4863. [Google Scholar] [CrossRef]
- Zhao, X.; Zhang, Z.; Hu, B.; Huang, W.; Yuan, C.; Zou, L. Response of Gut Microbiota to Metabolite Changes Induced by Endurance Exercise. Front. Microbiol. 2018, 9, 765. [Google Scholar] [CrossRef]
- Morishima, S.; Aoi, W.; Kawamura, A.; Kawase, T.; Takagi, T.; Naito, Y.; Tsukahara, T.; Inoue, R. Intensive, prolonged exercise seemingly causes gut dysbiosis in female endurance runners. J. Clin. Biochem. Nutr. 2021, 68, 253–258. [Google Scholar] [CrossRef]
- Heydenreich, J.; Kayser, B.; Schutz, Y.; Melzer, K. Total Energy Expenditure, Energy Intake, and Body Composition in Endurance Athletes Across the Training Season: A Systematic Review. Sport. Med.-Open 2017, 3, 1–24. [Google Scholar] [CrossRef] [Green Version]
- Koutedakis, Y. Seasonal Variation in Fitness Parameters in Competitive Athletes. Sport. Med. 2012, 19, 373–392. [Google Scholar] [CrossRef]
- Ruuskanen, O.; Luoto, R.; Valtonen, M.; Heinonen, J.O.; Waris, M. Respiratory Viral Infections in Athletes: Many Unanswered Questions. Sport. Med. 2022 2022, 1–9. [Google Scholar] [CrossRef]
- Jürimäe, J.; Mäestu, J.; Jürimäe, T.; Mangus, B.; Von Duvillard, S.P. Peripheral signals of energy homeostasis as possible markers of training stress in athletes: A review. Metabolism 2011, 60, 335–350. [Google Scholar] [CrossRef]
- Meeusen, R.; Duclos, M.; Foster, C.; Fry, A.; Gleeson, M.; Nieman, D.; Raglin, J.; Rietjens, G.; Steinacker, J.; Urhausen, A. Prevention, diagnosis and treatment of the overtraining syndrome: Joint consensus statement of the European College of Sport Science (ECSS) and the American College of Sports Medicine (ACSM). Eur. J. Sport Sci. 2012, 13, 1–24. [Google Scholar] [CrossRef] [Green Version]
- Schwellnus, M.; Soligard, T.; Alonso, J.; Bahr, R.; Clarsen, B.; Dijkstra, H.P.; Gabbett, T.; Gleeson, M.; Hägglund, M.; Hutchinson, M.R.; et al. How much is too much? (Part 1) International Olympic Committee consensus statement on load in sport and risk of injury. Br. J. Sports Med. 2016, 50, 1030–1041. [Google Scholar] [CrossRef] [Green Version]
- Scheiman, J.; Luber, J.M.; Chavkin, T.A.; MacDonald, T.; Tung, A.; Pham, L.D.; Wibowo, M.C.; Wurth, R.C.; Punthambaker, S.; Tierney, B.T.; et al. Meta-omics analysis of elite athletes identifies a performance-enhancing microbe that functions via lactate metabolism. Nat. Med. 2019, 25, 1104–1109. [Google Scholar] [CrossRef]
- Lin, C.L.; Hsu, Y.J.; Ho, H.H.; Chang, Y.C.; Kuo, Y.W.; Yeh, Y.T.; Tsai, S.Y.; Chen, C.W.; Chen, J.F.; Huang, C.C.; et al. Bifidobacterium longum subsp. longum OLP-01 Supplementation during Endurance Running Training Improves Exercise Performance in Middle- and Long-Distance Runners: A Double-Blind Controlled Trial. Nutrients 2020, 12, 1972. [Google Scholar] [CrossRef]
- Wilmot, E.G.; Edwardson, C.L.; Achana, F.A.; Davies, M.J.; Gorely, T.; Gray, L.J.; Khunti, K.; Yates, T.; Biddle, S.J.H. Sedentary time in adults and the association with diabetes, cardiovascular disease and death: Systematic review and meta-analysis. Diabetologia 2012, 55, 2895–2905. [Google Scholar] [CrossRef]
- Tang, W.H.W.; Hazen, S.L. Microbiome, trimethylamine N-oxide, and cardiometabolic disease. Transl. Res. 2017, 179, 108–115. [Google Scholar] [CrossRef] [Green Version]
- Utzschneider, K.M.; Kratz, M.; Damman, C.J.; Hullarg, M. Mechanisms Linking the Gut Microbiome and Glucose Metabolism. J. Clin. Endocrinol. Metab. 2016, 101, 1445–1454. [Google Scholar] [CrossRef] [Green Version]
- Turnbaugh, P.J.; Ley, R.E.; Mahowald, M.A.; Magrini, V.; Mardis, E.R.; Gordon, J.I. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 2006, 444, 1027–1031. [Google Scholar] [CrossRef]
- Vojinovic, D.; Radjabzadeh, D.; Kurilshikov, A.; Amin, N.; Wijmenga, C.; Franke, L.; Ikram, M.A.; Uitterlinden, A.G.; Zhernakova, A.; Fu, J.; et al. Relationship between gut microbiota and circulating metabolites in population-based cohorts. Nat. Commun. 2019, 10, 1–7. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Goodrich, J.K.; Waters, J.L.; Poole, A.C.; Sutter, J.L.; Koren, O.; Blekhman, R.; Beaumont, M.; Van Treuren, W.; Knight, R.; Bell, J.T.; et al. Human genetics shape the gut microbiome. Cell 2014, 159, 789–799. [Google Scholar] [CrossRef] [Green Version]
- Dhakal, S.; McCormack, L.; Dey, M. Association of the Gut Microbiota with Weight-Loss Response within a Retail Weight-Management Program. Microorganisms 2020, 8, 1246. [Google Scholar] [CrossRef]
- Ottosson, F.; Brunkwall, L.; Ericson, U.; Nilsson, P.M.; Almgren, P.; Fernandez, C.; Melander, O.; Orho-Melander, M. Connection Between BMI-Related Plasma Metabolite Profile and Gut Microbiota. J. Clin. Endocrinol. Metab. 2018, 103, 1491–1501. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Clarke, S.F.; Murphy, E.F.; O’Sullivan, O.; Lucey, A.J.; Humphreys, M.; Hogan, A.; Hayes, P.; O’Reilly, M.; Jeffery, I.B.; Wood-Martin, R.; et al. Exercise and associated dietary extremes impact on gut microbial diversity. Gut 2014, 63, 1913–1920. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fart, F.; Rajan, S.K.; Wall, R.; Rangel, I.; Ganda-Mall, J.P.; Tingö, L.; Brummer, R.J.; Repsilber, D.; Schoultz, I.; Lindqvist, C.M. Differences in Gut Microbiome Composition between Senior Orienteering Athletes and Community-Dwelling Older Adults. Nutrients 2020, 12, 2610. [Google Scholar] [CrossRef] [PubMed]
- Rajilić-Stojanović, M.; de Vos, W.M. The first 1000 cultured species of the human gastrointestinal microbiota. FEMS Microbiol. Rev. 2014, 38, 996–1047. [Google Scholar] [CrossRef]
- Cani, P.D. The gut microbiota manages host metabolism. Nat. Rev. Endocrinol. 2013, 10, 74–76. [Google Scholar] [CrossRef]
- Horton, J.D.; Shimano, H.; Hamilton, R.L.; Brown, M.S.; Goldstein, J.L. Disruption of LDL receptor gene in transgenic SREBP-1a mice unmasks hyperlipidemia resulting from production of lipid-rich VLDL. J. Clin. Invest. 1999, 103, 1067–1076. [Google Scholar] [CrossRef] [Green Version]
- Nielsen, S.; Karpe, F. Determinants of VLDL-triglycerides production. Curr. Opin. Lipidol. 2012, 23, 327. [Google Scholar] [CrossRef]
- Van Herreweghen, F.; De Paepe, K.; Roume, H.; Kerckhof, F.M.; Van de Wiele, T. Mucin degradation niche as a driver of microbiome composition and Akkermansia muciniphila abundance in a dynamic gut model is donor independent. FEMS Microbiol. Ecol. 2018, 94, 186. [Google Scholar] [CrossRef] [Green Version]
- Boesmans, L.; Valles-Colomer, M.; Wang, J.; Eeckhaut, V.; Falony, G.; Ducatelle, R.; Van Immerseel, F.; Raes, J.; Verbeke, K. Butyrate Producers as Potential Next-Generation Probiotics: Safety Assessment of the Administration of Butyricicoccus pullicaecorum to Healthy Volunteers. mSystems 2018, 3. [Google Scholar] [CrossRef] [Green Version]
- Eeckhaut, V.; Machiels, K.; Perrier, C.; Romero, C.; Maes, S.; Flahou, B.; Steppe, M.; Haesebrouck, F.; Sas, B.; Ducatelle, R.; et al. Butyricicoccus pullicaecorum in inflammatory bowel disease. Gut 2013, 62, 1745–1752. [Google Scholar] [CrossRef]
- Sasaki, K.; Sasaki, D.; Hannya, A.; Tsubota, J.; Kondo, A. In vitro human colonic microbiota utilises D-β-hydroxybutyrate to increase butyrogenesis. Sci. Rep. 2020, 10, 1–8. [Google Scholar] [CrossRef]
- Paoli, A.; Mancin, L.; Bianco, A.; Thomas, E.; Mota, J.F.; Piccini, F. Ketogenic Diet and Microbiota: Friends or Enemies? Genes 2019, 10, 534. [Google Scholar] [CrossRef] [Green Version]
- Heinecke, J. HDL and Cardiovascular-Disease Risk—Time for a New Approach? N. Engl. J. Med. 2011, 364, 170–171. [Google Scholar] [CrossRef]
- Ouimet, M.; Barrett, T.J.; Fisher, E.A. HDL and Reverse Cholesterol Transport. Circ. Res. 2019, 124, 1505–1518. [Google Scholar] [CrossRef]
- Karathanasis, S.K.; Freeman, L.A.; Gordon, S.M.; Remaley, A.T. The Changing Face of HDL and the Best Way to Measure It. Clin. Chem. 2017, 63, 196–210. [Google Scholar] [CrossRef] [Green Version]
- Buring, J.E.; O’Connor, G.T.; Goldhaber, S.Z.; Rosner, B.; Herbert, P.N.; Blum, C.B.; Breslow, J.L.; Hennekens, C.H. Decreased HDL2 and HDL3 cholesterol, Apo A-I and Apo A-II, and increased risk of myocardial infarction. Circulation 1992, 85, 22–29. [Google Scholar] [CrossRef] [Green Version]
- Salonen, J.T.; Salonen, R.; Seppänen, K.; Rauramaa, R.; Tuomilehto, J. HDL, HDL2, and HDL3 subfractions, and the risk of acute myocardial infarction. A prospective population study in eastern Finnish men. Circulation 1991, 84, 129–139. [Google Scholar] [CrossRef] [Green Version]
- Deng, X.; Ma, J.; Song, M.; Jin, Y.; Ji, C.; Ge, W.; Guo, C. Effects of products designed to modulate the gut microbiota on hyperlipidaemia. Eur. J. Nutr. 2019, 58, 2713–2729. [Google Scholar] [CrossRef]
- Zheng, M.M.; Wang, R.F.; Li, C.X.; Xu, J.H. Two-step enzymatic synthesis of ursodeoxycholic acid with a new 7β-hydroxysteroid dehydrogenase from Ruminococcus torques. Process Biochem. 2015, 50, 598–604. [Google Scholar] [CrossRef]
- IOC consensus statement on sports nutrition 2010. J. Sports Sci. 2011, 29, S3–S4. [CrossRef]
- Joo, J.; Williamson, S.A.; Vazquez, A.I.; Fernandez, J.R.; Bray, M.S. The influence of 15-week exercise training on dietary patterns among young adults. Int. J. Obes. 2019, 43, 1681–1690. [Google Scholar] [CrossRef]
- Kujala, U.M.; Palviainen, T.; Pesonen, P.; Waller, K.; Sillanpaä&die, E.; Niemelä, M.; Kangas, M.; Vähä-Ypyä, H.; Sievänen, H.; Korpelainen, R.; et al. Polygenic Risk Scores and Physical Activity. Med. Sci. Sport. Exerc. 2020, 52, 1518–1524. [Google Scholar] [CrossRef]
- Sillanpää, E.; Palviainen, T.; Ripatti, S.; Kujala, U.; Kaprio, J. Polygenic Score for Physical Activity Is Associated with Multiple Common Diseases. Med. Sci. Sport. Exerc. 2021, 53, 193–194. [Google Scholar] [CrossRef]
- Grieneisen, L.; Dasari, M.; Gould, T.J.; Björk, J.R.; Grenier, J.-C.; Yotova, V.; Jansen, D.; Gottel, N.; Gordon, J.B.; Learn, N.H.; et al. Gut microbiome heritability is nearly universal but environmentally contingent. Science 2021, 373, 181–186. [Google Scholar] [CrossRef]
- Neuberger-Castillo, L.; Hamot, G.; Marchese, M.; Sanchez, I.; Ammerlaan, W.; Betsou, F. Method Validation for Extraction of DNA from Human Stool Samples for Downstream Microbiome Analysis. Biopreserv. Biobank. 2020, 18, 102–116. [Google Scholar] [CrossRef]
- Rintala, A.; Riikonen, I.; Toivonen, A.; Pietilä, S.; Munukka, E.; Pursiheimo, J.P.; Elo, L.L.; Arikoski, P.; Luopajärvi, K.; Schwab, U.; et al. Early fecal microbiota composition in children who later develop celiac disease and associated autoimmunity. Scand. J. Gastroenterol. 2018, 53, 403–409. [Google Scholar] [CrossRef] [Green Version]
- Bokulich, N.A.; Subramanian, S.; Faith, J.J.; Gevers, D.; Gordon, J.I.; Knight, R.; Mills, D.A.; Caporaso, J.G. Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat. Methods 2013, 10, 57–59. [Google Scholar] [CrossRef]
- Segata, N.; Izard, J.; Waldron, L.; Gevers, D.; Miropolsky, L.; Garrett, W.S.; Huttenhower, C. Metagenomic biomarker discovery and explanation. Genome Biol. 2011, 12, 1–18. [Google Scholar] [CrossRef] [Green Version]
- Soininen, P.; Kangas, A.J.; Würtz, P.; Suna, T.; Ala-Korpela, M. Quantitative Serum Nuclear Magnetic Resonance Metabolomics in Cardiovascular Epidemiology and Genetics. Circ. Cardiovasc. Genet. 2015, 8, 192–206. [Google Scholar] [CrossRef] [Green Version]
- Soininen, P.; Kangas, A.J.; Würtz, P.; Tukiainen, T.; Tynkkynen, T.; Laatikainen, R.; Järvelin, M.-R.; Kähönen, M.; Lehtimäki, T.; Viikari, J.; et al. High-throughput serum NMR metabonomics for cost-effective holistic studies on systemic metabolism. Analyst 2009, 134, 1781–1785. [Google Scholar] [CrossRef]
| Athletes (n = 27) | Controls (n = 27) | |
|---|---|---|
| Age | 27.1 ± 5.1 | 27.4 ± 5.6 |
| Male/Female | 14/13 (52/48%) | 14/13 (52/48%) |
| BMI | 22.05 ± 1.8 | 24.0 ± 3.5 |
| Exercise load, h/week | 15 ± 2 | 4 ± 1 |
| Used oral antibiotics during previous 6 weeks | 2 | 1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hintikka, J.E.; Munukka, E.; Valtonen, M.; Luoto, R.; Ihalainen, J.K.; Kallonen, T.; Waris, M.; Heinonen, O.J.; Ruuskanen, O.; Pekkala, S. Gut Microbiota and Serum Metabolome in Elite Cross-Country Skiers: A Controlled Study. Metabolites 2022, 12, 335. https://doi.org/10.3390/metabo12040335
Hintikka JE, Munukka E, Valtonen M, Luoto R, Ihalainen JK, Kallonen T, Waris M, Heinonen OJ, Ruuskanen O, Pekkala S. Gut Microbiota and Serum Metabolome in Elite Cross-Country Skiers: A Controlled Study. Metabolites. 2022; 12(4):335. https://doi.org/10.3390/metabo12040335
Chicago/Turabian StyleHintikka, Jukka E., Eveliina Munukka, Maarit Valtonen, Raakel Luoto, Johanna K. Ihalainen, Teemu Kallonen, Matti Waris, Olli J. Heinonen, Olli Ruuskanen, and Satu Pekkala. 2022. "Gut Microbiota and Serum Metabolome in Elite Cross-Country Skiers: A Controlled Study" Metabolites 12, no. 4: 335. https://doi.org/10.3390/metabo12040335
APA StyleHintikka, J. E., Munukka, E., Valtonen, M., Luoto, R., Ihalainen, J. K., Kallonen, T., Waris, M., Heinonen, O. J., Ruuskanen, O., & Pekkala, S. (2022). Gut Microbiota and Serum Metabolome in Elite Cross-Country Skiers: A Controlled Study. Metabolites, 12(4), 335. https://doi.org/10.3390/metabo12040335






