The Importance of Microorganisms for Sustainable Agriculture—A Review
Abstract
1. Introduction
2. Rhizosphere and Root Exudates
3. Microbiome and Holobiont
4. Plant Immune System in Plant–PGPM Interactions
5. Mechanisms Underneath PGPM–Plant Interactions
5.1. Plant Antioxidant Defence System
5.2. Phytohormones
5.3. Availability of Micro- and Macronutrients
5.4. Direct Interactions of PGPM with Plant Pathogens
5.5. Induction of the Plant Resistance by Microbial Elicitors
6. The Stringent Response in Plant–Microorganism Interactions
7. Trichoderma–Plant Interaction—A Case Study
Author Contributions
Funding
Conflicts of Interest
References
- Al-Ani, L.K.T. Trichoderma: Beneficial role in sustainable agriculture by plant disease management. In Plant Microbiome: Stress Response; Egamberdieva, D., Ahmad, P., Eds.; Springer: Singapore, 2018; Volume 5, pp. 105–126. [Google Scholar]
- Weltje, L.; Simpson, P.; Gross, M.; Crane, M.; Wheeler, J.R. Comparative acute and chronic sensitivity of fish and amphibians: A critical review of data. Environ. Toxicol. Chem. 2013, 32, 984–994. [Google Scholar] [CrossRef]
- Mukherjee, R.K.; Kumar, V.; Roy, K. Chemometric modeling of plant protection products (PPPs) for the prediction of acute contact toxicity against honey bees (A. mellifera): A 2D-QSAR approach. J. Hazard. Mater. 2022, 423, 127230. [Google Scholar] [CrossRef]
- Bruni, I.; Gentili, R.; de Mattia, F.; Cortis, P.; Rossi, G.; Labra, M. A multi-level analysis to evaluate the extinction risk of and conservation strategy for the aquatic fern Marsilea quadrifolia L. in Europe. Aquat. Bot. 2013, 111, 35–42. [Google Scholar] [CrossRef]
- Matarczyk, J.A.; Willis, A.J.; Vranjic, J.A.; Ash, J.E. Herbicides, weeds and endangered species: Management of bitou bush (Chrysanthemoides monilifera ssp. rotundata) with glyphosate and impacts on the endangered shrub. Pimelea spicata. Biol. Conserv. 2002, 108, 133–141. [Google Scholar]
- Bikrol, A.; Saxena, N.; Singh, K. Response of Glycine max in relation to nitrogen fixation as influenced by fungicide seed treatment. Int. J. Biochem. Biotechnol. 2020, 9, 1–5. [Google Scholar]
- Santos, M.S.; Rodrigues, T.F.; Ferreira, E.; Megias, M.; Nogueira, M.A.; Hungria, M. Method for recovering and counting viable cells from maize seeds inoculated with Azospirillum brasilense. J. Pure Appl. Microbiol. 2020, 14, 195–204. [Google Scholar] [CrossRef]
- Zilli, J.É.; Ribeiro, K.G.; Campo, R.J.; Hungria, M. Influence of fungicide seed treatment on soybean nodulation and grain yield. Rev. Bras. Cienc. Solo 2009, 33, 917–923. [Google Scholar] [CrossRef]
- Streletskii, R.; Astaykina, A.; Krasnov, G.; Gorbatov, V. Changes in bacterial and fungal community of soil under treatment of pesticides. Agronomy 2022, 12, 124. [Google Scholar] [CrossRef]
- Santoyo, G.; Guzmán-Guzmán, P.; Parra-Cota, F.I.; de los Santos-Villalobos, S.; del Carmen Orozco-Mosqueda, M.; Glick, B.R. Plant growth stimulation by microbial consortia. Agronomy 2021, 11, 219. [Google Scholar] [CrossRef]
- Chaudhary, T.; Dixit, M.; Gera, R.; Shukla, A.K.; Prakash, A.; Gupta, G.; Shukla, P. Techniques for improving formulations of bioinoculants. 3 Biotech. 2020, 10, 199. [Google Scholar] [CrossRef]
- Romano, I.; Ventorino, V.; Pepe, O. Effectiveness of plant beneficial microbes: Overview of the methodological approaches for the assessment of root colonization and persistence. Front. Plant Sci. 2020, 11, 6. [Google Scholar] [CrossRef]
- Maitra, S.; Brestic, M.; Bhadra, P.; Shankar, T.; Praharaj, S.; Palai, J.B.; Shah, M.M.R.; Barek, V.; Ondrisik, P.; Skalický, M.; et al. Bioinoculants-natural biological resources for sustainable plant production. Microorganisms 2022, 10, 51. [Google Scholar] [CrossRef]
- Basu, A.; Prasad, P.; Das, S.N.; Kalam, S.; Sayyed, R.Z.; Reddy, M.S.; Enshasy, H. Plant growth promoting rhizobacteria (PGPR) as green bioinoculants: Recent developments, constraints, and prospects. Sustainability 2021, 13, 1140. [Google Scholar] [CrossRef]
- Martínez-Hidalgo, P.; Maymon, M.; Pule-Meulenberg, F.; Hirsch, A.M. Engineering root microbiomes for healthier crops and soils using beneficial, environmentally safe bacteria. Can. J. Microbiol. 2019, 65, 91–104. [Google Scholar] [CrossRef]
- Cai, F.; Chen, W.; Wei, Z.; Pang, G.; Li, R.; Ran, W.; Shen, Q. Colonization of Trichoderma harzianum strain SQR-T037 on tomato roots and its relationship to plant growth, nutrient availability and soil microflora. Plant Soil 2015, 388, 337–350. [Google Scholar] [CrossRef]
- Rojas-Tapias, D.F.; Bonilla, R.; Dussán, J. Effect of inoculation and co-inoculation of Acinetobacter sp. RG30 and Pseudomonas putida GN04 on growth, fitness, and copper accumulation of maize (Zea mays). Water Air Soil Pollut. 2014, 225, 2232. [Google Scholar] [CrossRef]
- Patel, P.; Gajjar, H.; Joshi, B.; Krishnamurthy, R.; Amaresan, N. Inoculation of salt-tolerant Acinetobacter sp. (RSC9) improves the sugarcane (Saccharum sp. hybrids) growth under salinity stress condition. Sugar Tech 2022, 24, 494–501. [Google Scholar] [CrossRef]
- Chihaoui, S.A.; Trabelsi, D.; Jdey, A.; Mhadhbi, H.; Mhamdi, R. Inoculation of Phaseolus vulgaris with the nodule-endophyte Agrobacterium sp. 10C2 affects richness and structure of rhizosphere bacterial communities and enhances nodulation and growth. Arch. Microbiol. 2015, 197, 805–813. [Google Scholar] [CrossRef]
- Sharma, M.; Mishra, V.; Rau, N.; Sharma, R.S. Increased iron-stress resilience of maize through inoculation of siderophore-producing Arthrobacter globiformis from mine. J. Basic Microbiol. 2016, 56, 719–735. [Google Scholar] [CrossRef]
- Fan, P.; Chen, D.; He, Y.; Zhou, Q.; Tian, Y.; Gao, L. Alleviating salt stress in tomato seedlings using Arthrobacter and Bacillus megaterium isolated from the rhizosphere of wild plants grown on saline–alkaline lands. Int. J. Phytoremediation 2016, 18, 1113–1121. [Google Scholar] [CrossRef]
- Parmasi, Z.; Tahmasebi, Z.; Zare, M.J.; Nourollahi, K.; Kanouni, H. Biocontrol of Ascochyta blight by Azospirillum sp. depending on the degree of resistance of chickpea genotypes. J. Phytopathol. 2019, 167, 601–607. [Google Scholar] [CrossRef]
- Mazhar, R.; Ilyas, N.; Saeed, M.; Bibi, F.; Batool, N. Biocontrol and salinity tolerance potential of Azospirillum lipoferum and its inoculation effect in wheat crop. Int. J. Agric. Biol. 2015, 18, 494–500. [Google Scholar] [CrossRef]
- Shirinbayan, S.; Khosravi, H.; Malakouti, M.J. Alleviation of drought stress in maize (Zea mays) by inoculation with Azotobacter strains isolated from semi-arid regions. Appl. Soil Ecol. 2019, 133, 138–145. [Google Scholar] [CrossRef]
- Chen, L.; Liu, Y.; Wu, G.; Njeri, K.V.; Shen, Q.; Zhang, N.; Zhang, R. Induced maize salt tolerance by rhizosphere inoculation of Bacillus amyloliquefaciens SQR9. Physiol. Plant. 2016, 158, 34–44. [Google Scholar] [CrossRef]
- Hafeez, F.Y.; Safdar, M.E.; Chaudhry, A.U.; Malik, K.A. Rhizobial inoculation improves seedling emergence, nutrient uptake and growth of cotton. Aust. J. Exp. Agric. 2004, 44, 617–622. [Google Scholar] [CrossRef]
- Naveed, M.; Hussain, M.B.; Zahir, Z.A.; Mitter, B.; Sessitsch, A. Drought stress amelioration in wheat through inoculation with Burkholderia phytofirmans strain PsJN. Plant Growth Regul. 2014, 73, 121–131. [Google Scholar] [CrossRef]
- Bernabeu, P.R.; Pistorio, M.; Torres-Tejerizo, G.; Estrada-De los Santos, P.; Galar, M.L.; Boiardi, J.L.; Luna, M.F. Colonization and plant growth-promotion of tomato by Burkholderia tropica. Sci. Hortic. 2015, 191, 113–120. [Google Scholar] [CrossRef]
- Tsuda, K.; Kosaka, Y.; Tsuge, S.; Kub, Y.; Horin, O. Evaluation of the endophyte Enterobacter cloacae SM10 isolated from spinach roots for biological control against Fusarium wilt of spinach. J. Gen. Plant Pathol. 2001, 67, 78–84. [Google Scholar] [CrossRef]
- Ngom, M.; Gray, K.; Diagne, N.; Oshone, R.; Fardoux, J.; Gherbi, H.; Hocher, V.; Svistoonoff, S.; Laplaze, L.; Tisa, L.S.; et al. Symbiotic performance of diverse Frankia strains on salt-stressed Casuarina glauca and Casuarina equisetifolia plants. Front. Plant Sci. 2016, 7, 1331. [Google Scholar] [CrossRef]
- Grossi, C.E.M.; Fantino, E.; Serral, F.; Zawoznik, M.S.; Fernandez Do Porto, D.A.; Ulloa, R.M. Methylobacterium sp. 2A is a plant growth-promoting rhizobacteria that has the potential to improve potato crop yield under adverse conditions. Front. Plant Sci. 2020, 11, 71. [Google Scholar] [CrossRef]
- Hernández-Montiel, L.G.; Chiquito-Contreras, C.J.; Murillo-Amador, B.; Vidal-Hernández, L.; Quiñones-Aguilar, E.E.; Chiquito-Contreras, R.G. Efficiency of two inoculation methods of Pseudomonas putida on growth and yield of tomato plants. J. Soil Sci. Plant Nutr. 2017, 17, 1003–1012. [Google Scholar] [CrossRef]
- Fu, Q.; Liu, C.; Ding, N.; Lin, Y.; Guo, B. Ameliorative effects of inoculation with the plant growth-promoting rhizobacterium Pseudomonas sp. DW1 on growth of eggplant (Solanum melongena L.) seedlings under salt stress. Agric. Water Manag. 2010, 97, 1994–2000. [Google Scholar] [CrossRef]
- Szymańska, S.; Dąbrowska, G.B.; Tyburski, J.; Niedojadło, K.; Piernik, A.; Hrynkiewicz, K. Boosting the Brassica napus L. tolerance to salinity by the halotolerant strain Pseudomonas stutzeri ISE12. Environ. Exp. Bot. 2019, 163, 55–68. [Google Scholar] [CrossRef]
- Biswas, J.C.; Ladha, J.K.; Dazzo, F.B. Rhizobia inoculation improves nutrient uptake and growth of lowland rice. Soil Sci. Soc. Am. J. 2000, 64, 1644–1650. [Google Scholar] [CrossRef]
- Biswas, J.C.; Ladha, J.K.; Dazzo, F.B.; Yanni, Y.G.; Rolfe, B.G. Rhizobial inoculation influences seedling vigor and yield of rice. Agron. J. 2000, 92, 880–886. [Google Scholar] [CrossRef]
- Turhan, E.; Kiran, S.; Ates, Ç.; Ates, O.; Kusvuran, S.; Ellialtioglu, S.S. Ameliorative effects of inoculation with Serratia marcescens and grafting on growth of eggplant seedlings under salt stress. J. Plant Nutr. 2020, 43, 594–603. [Google Scholar] [CrossRef]
- Palaniyandi, S.A.; Damodharan, K.; Yang, S.H.; Suh, J.W. Streptomyces sp. strain PGPA39 alleviates salt stress and promotes growth of ‘Micro Tom’ tomato plants. J. Appl. Microbiol. 2014, 117, 766–773. [Google Scholar] [CrossRef]
- Samac, D.A.; Kinkel, L.L. Suppression of the root-lesion nematode (Pratylenchus penetrans) in alfalfa (Medicago sativa) by Streptomyces spp. Plant Soil 2001, 235, 35–44. [Google Scholar] [CrossRef]
- Mauricio-Castillo, J.A.; Salas-Muñoz, S.; Reveles-Torres, L.R.; Salas-Luevano, M.A.; Salazar-Badillo, F.B. Could Alternaria solani IA300 be a plant growth-promoting fungus? Eur. J. Plant Pathol. 2020, 157, 413–419. [Google Scholar] [CrossRef]
- Galeano, R.M.S.; Franco, D.G.; Chaves, P.O.; Giannesi, G.C.; Masui, D.C.; Ruller, R.; Corrêa, B.O.; da Silva Brasil, M.; Zanoelo, F.F. Plant growth promoting potential of endophytic Aspergillus niger 9-p isolated from native forage grass in Pantanal of Nhecolândia region, Brazil. Rhizosphere 2021, 18, 100332. [Google Scholar] [CrossRef]
- Khan, A.L.; Hamayun, M.; Kim, Y.H.; Kang, S.M.; Lee, J.H.; Lee, I.J. Gibberellins producing endophytic Aspergillus fumigatus sp. LH02 influenced endogenous phytohormonal levels, isoflavonoids production and plant growth in salinity stress. Process Biochem. 2011, 46, 440–447. [Google Scholar] [CrossRef]
- Dąbrowska, G.B.; Garstecka, Z.; Trejgell, A.; Dąbrowski, H.P.; Konieczna, W.; Szyp-Borowska, I. The impact of forest fungi on promoting growth and development of Brassica napus L. Agronomy 2021, 11, 2475. [Google Scholar] [CrossRef]
- Velásquez, A.; Vega-Celedón, P.; Fiaschi, G.; Agnolucci, M.; Avio, L.; Giovannetti, M.; D’Onofrio, C.; Seeger, M. Responses of Vitis vinifera cv. Cabernet Sauvignon roots to the arbuscular mycorrhizal fungus Funneliformis mosseae and the plant growth-promoting rhizobacterium Ensifer meliloti include changes in volatile organic compounds. Mycorrhiza 2020, 30, 161–170. [Google Scholar] [CrossRef]
- Saldajeno, M.G.B.; Hyakumachi, M. The plant growth-promoting fungus Fusarium equiseti and the arbuscular mycorrhizal fungus Glomus mosseae stimulate plant growth and reduce severity of anthracnose and damping-off diseases in cucumber (Cucumis sativus) seedlings. Ann. Appl. Biol. 2011, 159, 28–40. [Google Scholar] [CrossRef]
- Radhakrishnan, R.; Khan, A.L.; Kang, S.M.; Lee, I.J. A comparative study of phosphate solubilization and the host plant growth promotion ability of Fusarium verticillioides RK01 and Humicola sp. KNU01 under salt stress. Ann. Microbiol. 2015, 65, 585–593. [Google Scholar] [CrossRef]
- Estaún, V.; Camprubí, A.; Calvet, C.; Pinochet, J. Nursery and field response of olive trees inoculated with two arbuscular mycorrhizal fungi, Glomus intraradices and Glomus mosseae. J. Am. Soc. Hortic. Sci. 2003, 128, 767–775. [Google Scholar] [CrossRef]
- Senthil Kumar, C.M.; Jacob, T.K.; Devasahayam, S.; Thomas, S.; Geethu, C. Multifarious plant growth promotion by an entomopathogenic fungus Lecanicillium psalliotae. Microbiol. Res. 2018, 207, 153–160. [Google Scholar] [CrossRef]
- Ozimek, E.; Jaroszuk-Ściseł, J.; Bohacz, J.; Korniłłowicz-Kowalska, T.; Tyśkiewicz, R.; Słomka, A.; Nowak, A.; Hanaka, A. Synthesis of indoleacetic acid, gibberellic acid and ACC-deaminase by Mortierella strains promote winter wheat seedlings growth under different conditions. Int. J. Mol. Sci. 2018, 19, 3218. [Google Scholar] [CrossRef]
- Rozpądek, P.; Domka, A.; Ważny, R.; Nosek, M.; Jędrzejczyk, R.; Tokarz, K.; Turnau, K. How does the endophytic fungus Mucor sp. improve Arabidopsis arenosa vegetation in the degraded environment of a mine dump? Environ. Exp. Bot. 2018, 147, 31–42. [Google Scholar] [CrossRef]
- Vessey, J.K.; Heisinger, K.G. Effect of Penicillium bilaii inoculation and phosphorus fertilisation on root and shoot parameters of field-grown pea. Can. J. Microbiol. 2011, 81, 361–366. [Google Scholar] [CrossRef]
- Wakelin, S.A.; Gupta, V.V.S.R.; Harvey, P.R.; Ryder, M.H. The effect of Penicillium fungi on plant growth and phosphorus mobilization in neutral to alkaline soils from southern Australia. Can. J. Microbiol. 2007, 53, 106–115. [Google Scholar] [CrossRef]
- Elsharkawy, M.M. Induced systemic resistance against Cucumber mosaic virus by Phoma sp. GS8-2 stimulates transcription of pathogenesis-related genes in Arabidopsis. Pest. Manag. Sci. 2019, 75, 859–866. [Google Scholar] [CrossRef]
- Elsharkawy, M.M.; El-Khateeb, N.M.M. Antifungal activity and resistance induction against Sclerotium cepivorum by plant growth-promoting fungi in onion plants. Egypt. J. Biol. Pest Control 2019, 29, 68. [Google Scholar] [CrossRef]
- Baron, N.C.; de Souza Pollo, A.; Rigobelo, E.C. Purpureocillium lilacinum and Metarhizium marquandii as plant growth-promoting fungi. PeerJ 2020, 2020, e9005. [Google Scholar] [CrossRef]
- Janowska, B.; Andrzejak, R.; Kosiada, T. The influence of fungi of the Trichoderma genus on the flowering of Freesia refracta Klatt ‘Argentea’ in winter. Hortic. Sci. 2020, 47, 203–210. [Google Scholar] [CrossRef]
- Vinayarani, G.; Prakash, H.S. Fungal endophytes of turmeric (Curcuma longa L.) and their biocontrol potential against pathogens Pythium aphanidermatum and Rhizoctonia solani. World J. Microbiol. Biotechnol. 2018, 34, 49. [Google Scholar] [CrossRef]
- Nuangmek, W.; Aiduang, W.; Kumla, J.; Lumyong, S.; Suwannarach, N. Evaluation of a newly identified endophytic fungus, Trichoderma phayaoense for plant growth promotion and biological control of gummy stem blight and wilt of muskmelon. Front. Microbiol. 2021, 12, 410. [Google Scholar] [CrossRef]
- Znajewska, Z.; Dąbrowska, G.; Narbutt, O. Trichoderma viride strains stimulating the growth and development of winter rapeseed (Brassica napus L.). Prog. Plant Prot. 2018, 58, 264–269. [Google Scholar]
- Singh, D.P.; Prabha, R.; Yandigeri, M.S.; Arora, D.K. Cyanobacteria-mediated phenylpropanoids and phytohormones in rice (Oryza sativa) enhance plant growth and stress tolerance. Anton. Leeuw. Int. J. 2011, 100, 557–568. [Google Scholar] [CrossRef]
- Prasanna, R.; Chaudhary, V.; Gupta, V.; Babu, S.; Kumar, A.; Singh, R.; Shivay, Y.S.; Nain, L. Cyanobacteria mediated plant growth promotion and bioprotection against Fusarium wilt in tomato. Eur. J. Plant Pathol. 2013, 136, 337–353. [Google Scholar] [CrossRef]
- Priya, H.; Prasanna, R.; Ramakrishnan, B.; Bidyarani, N.; Babu, S.; Thapa, S.; Renuka, N. Influence of cyanobacterial inoculation on the culturable microbiome and growth of rice. Microbiol. Res. 2015, 171, 78–89. [Google Scholar] [CrossRef]
- Babu, S.; Prasanna, R.; Bidyarani, N.; Singh, R. Analysing the colonisation of inoculated cyanobacteria in wheat plants using biochemical and molecular tools. J. Appl. Phycol. 2015, 27, 327–338. [Google Scholar] [CrossRef]
- Kim, S.J.; Ko, E.J.; Hong, J.K.; Jeun, Y.C. Ultrastructures of Colletotrichum orbiculare in cucumber leaves expressing systemic acquired resistance mediated by Chlorella fusca. Plant Pathol. J. 2018, 34, 113. [Google Scholar] [CrossRef]
- Mohamed Taha, T.; Youssef, M.A. Improvement of growth parameters of Zea mays and properties of soil inoculated with two Chlorella species. Rep. Opin. 2015, 7, 22–27. [Google Scholar]
- Agwa, O.K.; Ogugbue, C.J.; Williams, E.E. Growth promotion of Telfairia occidentalis by application of Chlorella vulgaris (bioinoculant) colonized seeds and soil under tropical field conditions. Am. J. Plant Sci. 2018, 9, 403–415. [Google Scholar] [CrossRef]
- Kuang, X.; Gu, J.D.; Tie, B.Q.; Yao, B.; Shao, J. Interactive effects of cadmium and Microcystis aeruginosa (cyanobacterium) on the growth, antioxidative responses and accumulation of cadmium and microcystins in rice seedlings. Ecotoxicology 2016, 25, 1588–1599. [Google Scholar] [CrossRef]
- Hussain, A.; Hamayun, M.; Shah, S.T. Root colonization and phytostimulation by phytohormones producing entophytic Nostoc sp. AH-12. Curr. Microbiol. 2013, 67, 624–630. [Google Scholar] [CrossRef]
- Maqubela, M.P.; Mnkeni, P.N.S.; Issa, O.M.; Pardo, M.T.; D’Acqui, L.P. Nostoc cyanobacterial inoculation in South African agricultural soils enhances soil structure, fertility, and maize growth. Plant Soil 2009, 315, 79–92. [Google Scholar] [CrossRef]
- Barone, V.; Puglisi, I.; Fragalà, F.; lo Piero, A.R.; Giuffrida, F.; Baglieri, A. Novel bioprocess for the cultivation of microalgae in hydroponic growing system of tomato plants. J. Appl. Phycol. 2019, 31, 465–470. [Google Scholar] [CrossRef]
- Seifikalhor, M.; Hassani, S.B.; Aliniaeifard, S. Seed priming by cyanobacteria (Spirulina platensis) and salep gum enhances tolerance of maize plant against cadmium toxicity. J. Plant Growth Regul. 2020, 39, 1009–1021. [Google Scholar] [CrossRef]
- Rana, A.; Joshi, M.; Prasanna, R.; Shivay, Y.S.; Nain, L. Biofortification of wheat through inoculation of plant growth promoting rhizobacteria and cyanobacteria. Eur. J. Soil Biol. 2012, 50, 118–126. [Google Scholar] [CrossRef]
- Flores, A.C.; Luna, A.A.E.; Portugal, V.O. Yield and quality enhancement of marigold flowers by inoculation with Bacillus subtilis and Glomus fasciculatum. J. Sustain. Agric. 2007, 31, 21–31. [Google Scholar] [CrossRef]
- Nacoon, S.; Jogloy, S.; Riddech, N.; Mongkolthanaruk, W.; Kuyper, T.W.; Boonlue, S. Interaction between phosphate solubilizing bacteria and arbuscular mycorrhizal fungi on growth promotion and tuber inulin content of Helianthus tuberosus L. Sci. Rep. 2020, 10, 4916. [Google Scholar] [CrossRef]
- Oliveira, R.S.; Carvalho, P.; Marques, G.; Ferreira, L.; Nunes, M.; Rocha, I.; Ma, Y.; Carvalho, M.F.; Vosátka, M.; Freitas, H. Increased protein content of chickpea (Cicer arietinum L.) inoculated with arbuscular mycorrhizal fungi and nitrogen-fixing bacteria under water deficit conditions. J. Sci. Food Agric. 2017, 97, 4379–4385. [Google Scholar] [CrossRef]
- Bona, E.; Cantamessa, S.; Massa, N.; Manassero, P.; Marsano, F.; Copetta, A.; Lingua, G.; D’Agostino, G.; Gamalero, E.; Berta, G. Arbuscular mycorrhizal fungi and plant growth-promoting pseudomonads improve yield, quality and nutritional value of tomato: A field study. Mycorrhiza 2017, 27, 1–11. [Google Scholar] [CrossRef]
- Hidri, R.; Barea, J.M.; Mahmoud, O.M.B.; Abdelly, C.; Azcón, R. Impact of microbial inoculation on biomass accumulation by Sulla carnosa provenances, and in regulating nutrition, physiological and antioxidant activities of this species under non-saline and saline conditions. J. Plant Physiol. 2016, 201, 28–41. [Google Scholar] [CrossRef]
- Armada, E.; Probanza, A.; Roldán, A.; Azcón, R. Native plant growth promoting bacteria Bacillus thuringiensis and mixed or individual mycorrhizal species improved drought tolerance and oxidative metabolism in Lavandula dentata plants. J. Plant Physiol. 2016, 192, 1–12. [Google Scholar] [CrossRef]
- Duc, N.H.; Mayer, Z.; Pék, Z.; Helyes, L.; Posta, K. Combined inoculation of arbuscular mycorrhizal fungi, Pseudomonas fluorescens and Trichoderma spp. for enhancing defense enzymes and yield of three pepper cultivars. Appl. Ecol. Environ. Res. 2017, 15, 1815–1829. [Google Scholar] [CrossRef]
- Pandey, A.; Yarzábal, L.A. Bioprospecting cold-adapted plant growth promoting microorganisms from mountain environments. Appl. Microbiol. Biotechnol. 2019, 103, 643–657. [Google Scholar] [CrossRef]
- Van Elsas, J.D.; Chiurazzi, M.; Mallon, C.A.; Elhottovā, D.; Krištůfek, V.; Salles, J.F. Microbial diversity determines the invasion of soil by a bacterial pathogen. PNAS 2012, 109, 1159–1164. [Google Scholar] [CrossRef]
- Goswami, D.; Thakker, J.N.; Dhandhukia, P.C. Portraying mechanics of plant growth promoting rhizobacteria (PGPR): A review. Cogent. Food Agric. 2016, 2, 1127500. [Google Scholar] [CrossRef]
- Middleton, H.; Yergeau, É.; Monard, C.; Combier, J.-P.; el Amrani, A. Rhizospheric plant–microbe interactions: miRNAs as a key mediator. Trends Plant Sci. 2021, 26, 132–141. [Google Scholar] [CrossRef] [PubMed]
- Manfredini, A.; Malusà, E.; Costa, C.; Pallottino, F.; Mocali, S.; Pinzari, F.; Canfora, L. Current methods, common practices, and perspectives in tracking and monitoring bioinoculants in soil. Front. Microbiol. 2021, 12, 698491. [Google Scholar] [CrossRef] [PubMed]
- Shivanna, M.B.; Meera, M.S.; Hyakumachi, M. Sterile fungi from zoysiagrass rhizosphere as plant growth promoters in spring wheat. Can. J. Microbiol. 1994, 40, 637–644. [Google Scholar] [CrossRef]
- Dąbrowska, G.; Hrynkiewicz, K.; Mierek-Adamska, A.; Goc, A. The sensitivity of spring and winter varieties of oilseed rape to heavy metals and rhizobacteria. Oilseed Crop. 2012, 33, 201–220. [Google Scholar]
- Gnanamanickam, S.S.; Immanuel, J.E. Epiphytic bacteria, their ecology and functions. In Plant-Associated Bacteria; Gnanamanickam, S.S., Ed.; Springer: Dordrecht, The Netherlands, 2006; pp. 131–153. [Google Scholar]
- Kucharska, K.; Wachowska, U. The microbiome on the leaves of crop plants. Adv. Microbiol. 2014, 53, 352–359. [Google Scholar]
- Martins, G.; Lauga, B.; Miot-Sertier, C.; Mercier, A.; Lonvaud, A.; Soulas, M.-L.; Soulas, G.; Masneuf-Pomarède, I. Characterization of epiphytic bacterial communities from grapes, leaves, bark and soil of grapevine plants grown, and their relations. PLoS ONE 2013, 8, e73013. [Google Scholar] [CrossRef]
- Whipps, J.M. Microbial interactions and biocontrol in the rhizosphere. J. Exp. Bot. 2001, 52, 487–511. [Google Scholar] [CrossRef]
- Gong, T.; Xin, X. Phyllosphere microbiota: Community dynamics and its interaction with plant hosts. J. Integr. Plant Biol. 2021, 63, 297–304. [Google Scholar] [CrossRef]
- Contreras-Cornejo, H.A.; Macías-Rodríguez, L.; Cortés-Penagos, C.; López-Bucio, J. Trichoderma virens, a plant beneficial fungus, enhances biomass production and promotes lateral root growth through an auxin-dependent mechanism in Arabidopsis. Plant Physiol. 2009, 149, 1579–1592. [Google Scholar] [CrossRef]
- You, Y.-H.; Yoon, H.; Kang, S.-M.; Shin, J.-H.; Choo, Y.-S.; Lee, I.-J.; Lee, J.-M.; Kim, J.-G. Fungal diversity and plant growth promotion of endophytic fungi from six halophytes in Suncheon Bay. J. Microbiol. Biotechnol. 2012, 22, 1549–1556. [Google Scholar] [CrossRef] [PubMed]
- Bergottini, V.M.; Otegui, M.B.; Sosa, D.A.; Zapata, P.D.; Mulot, M.; Rebord, M.; Zopfi, J.; Wiss, F.; Benrey, B.; Junier, P. Bio-inoculation of yerba mate seedlings (Ilex paraguariensis St. Hill.) with native plant growth-promoting rhizobacteria: A sustainable alternative to improve crop yield. Biol. Fertil. Soils 2015, 51, 749–755. [Google Scholar] [CrossRef]
- Poonguzhali, S.; Madhaiyan, M.; Sa, T. Isolation and identification of phosphate solubilizing bacteria from chinese cabbage and their effect on growth and phosphorus utilization of plants. J. Microbiol. Biotechnol. 2008, 18, 773–777. [Google Scholar] [PubMed]
- Zhang, C.; Kong, F. Isolation and identification of potassium-solubilizing bacteria from tobacco rhizospheric soil and their effect on tobacco plants. Appl. Soil Ecol. 2014, 82, 18–25. [Google Scholar] [CrossRef]
- Reed, M.L.E.; Glick, B.R. Growth of canola (Brassica napus) in the presence of plant growth-promoting bacteria and either copper or polycyclic aromatic hydrocarbons. Can. J. Microbiol. 2005, 51, 1061–1069. [Google Scholar] [CrossRef]
- Jousset, A.; Rochat, L.; Lanoue, A.; Bonkowski, M.; Keel, C.; Scheu, S. Plants respond to pathogen infection by enhancing the antifungal gene expression of root-associated bacteria. Mol. Plant-Microbe Interact. 2011, 24, 352–358. [Google Scholar] [CrossRef]
- Schirmböck, M.; Lorito, M.; Wang, Y.L.; Hayes, C.K.; Arisan-Atac, I.; Scala, F.; Harman, G.E.; Kubicek, C.P. Parallel formation and synergism of hydrolytic enzymes and peptaibol antibiotics, molecular mechanisms involved in the antagonistic action of Trichoderma harzianum against phytopathogenic fungi. Appl. Environ. Microbiol. 1994, 60, 4364–4370. [Google Scholar] [CrossRef]
- Crowley, D.E. Microbial siderophores in the plant rhizosphere. In Iron Nutrition in Plants and Rhizospheric Microorganisms; Barton, L.L., Abadia, J., Eds.; Springer: Dordrecht, The Netherlands, 2006; pp. 169–198. [Google Scholar]
- Pinedo, I.; Ledger, T.; Greve, M.; Poupin, M.J. Burkholderia phytofirmans PsJN induces long-term metabolic and transcriptional changes involved in Arabidopsis thaliana salt tolerance. Front. Plant Sci. 2015, 6, 466. [Google Scholar] [CrossRef]
- Coppola, M.; Cascone, P.; di Lelio, I.; Woo, S.L.; Lorito, M.; Rao, R.; Pennacchio, F.; Guerrieri, E.; Digilio, M.C. Trichoderma atroviride P1 colonization of tomato plants enhances both direct and indirect defense barriers against insects. Front. Physiol. 2019, 10, 813. [Google Scholar] [CrossRef]
- Ardanov, P.; Lyastchenko, S.; Karppinen, K.; Häggman, H.; Kozyrovska, N.; Pirttilä, A.M. Effects of Methylobacterium sp. on emergence, yield, and disease prevalence in three cultivars of potato (Solanum tuberosum L.) were associated with the shift in endophytic microbial community. Plant Soil 2016, 405, 299–310. [Google Scholar] [CrossRef]
- Lynch, J.M.; Leij, F. Rhizosphere. In eLS.; John Wiley & Sons: Hoboken, NJ, USA, 2012. [Google Scholar]
- Kawasaki, A.; Donn, S.; Ryan, P.R.; Mathesius, U.; Devilla, R.; Jones, A.; Watt, M. Microbiome and exudates of the root and rhizosphere of Brachypodium distachyon, a model for wheat. PLoS ONE 2016, 11, e0164533. [Google Scholar] [CrossRef] [PubMed]
- Li, X.G.; Zhang, T.L.; Wang, X.X.; Hua, K.; Zhao, L.; Han, Z.M. The composition of root exudates from two different resistant peanut cultivars and their effects on the growth of soil-borne pathogen. Int. J. Biol. Sci. 2013, 9, 164. [Google Scholar] [CrossRef] [PubMed]
- Lucas García, J.A.; Barbas, C.; Probanza, A.; Barrientos, M.L.; Gutierrez Mañero, F.J. Low molecular weight organic acids and fatty acids in root exudates of two Lupinus cultivars at flowering and fruiting stages. Phytochem. Anal. 2001, 12, 305–311. [Google Scholar] [CrossRef]
- Narasimhan, K.; Basheer, C.; Bajic, V.B.; Swarup, S. Enhancement of plant-microbe interactions using a rhizosphere metabolomics-driven approach and its application in the removal of polychlorinated biphenyls. Plant Physiol. 2003, 132, 146–153. [Google Scholar] [CrossRef] [PubMed]
- Nóbrega, F.M.; Santos, I.S.; da Cunha, M.; Carvalho, A.O.; Gomes, V.M. Antimicrobial proteins from cowpea root exudates: Inhibitory activity against Fusarium oxysporum and purification of a chitinase-like protein. Plant Soil 2005, 272, 223–232. [Google Scholar] [CrossRef]
- Schönwitz, R.; Ziegler, H. Exudation of water-soluble vitamins and of some carbohydrates by intact roots of maize seedlings (Zea mays L.) into a mineral nutrient solution. Z. Pflanzenphysiol. 1982, 107, 7–14. [Google Scholar] [CrossRef]
- Wang, J.; Ding, Z.; Bian, J.; Bo, T.; Liu, Y. Chemotaxis response of Meloidogyne incognita to volatiles and organic acids from root exudates. Rhizosphere 2021, 17, 100320. [Google Scholar] [CrossRef]
- Williams, A.; Langridge, H.; Straathof, A.L.; Muhamadali, H.; Hollywood, K.A.; Goodacre, R.; de Vries, F.T. Root functional traits explain root exudation rate and composition across a range of grassland species. J. Ecol. 2022, 110, 21–33. [Google Scholar] [CrossRef]
- Aulakh, M.S.; Wassmann, R.; Bueno, C.; Kreuzwieser, J.; Rennenberg, H. Characterization of root exudates at different growth stages of ten rice (Oryza sativa L.) cultivars. Plant Biol. 2001, 3, 139–148. [Google Scholar] [CrossRef]
- Bekkara, F.; Jay, M.; Viricel, M.R.; Rome, S. Distribution of phenolic compounds within seed and seedlings of two Vicia faba cvs differing in their seed tannin content, and study of their seed and root phenolic exudations. Plant Soil 1998, 203, 27–36. [Google Scholar] [CrossRef]
- Watt, M.; Evans, J.R. Linking development and determinacy with organic acid efflux from proteoid roots of white lupin grown with low phosphorus and ambient or elevated atmospheric CO2 concentration. Plant Physiol. 1999, 120, 705–716. [Google Scholar] [CrossRef] [PubMed]
- Tawaraya, K.; Horie, R.; Saito, A.; Shinano, T.; Wagatsuma, T.; Saito, K.; Oikawa, A. Metabolite profiling of shoot extracts, root extracts, and root exudates of rice plant under phosphorus deficiency. J. Plant Nutr. 2013, 36, 1138–1159. [Google Scholar] [CrossRef]
- Tawaraya, K.; Horie, R.; Shinano, T.; Wagatsuma, T.; Saito, K.; Oikawa, A. Metabolite profiling of soybean root exudates under phosphorus deficiency. Soil Sci. Plant Nutr. 2014, 60, 679–694. [Google Scholar] [CrossRef]
- Neumann, G.; Römheld, V. Root excretion of carboxylic acids and protons in phosphorus-deficient plants. Plant Soil 1999, 211, 121–130. [Google Scholar] [CrossRef]
- Carvalhais, L.C.; Dennis, P.G.; Fedoseyenko, D.; Hajirezaei, M.R.; Borriss, R.; von Wirén, N. Root exudation of sugars, amino acids, and organic acids by maize as affected by nitrogen, phosphorus, potassium, and iron deficiency. J. Plant Nutr. Soil Sci. 2011, 174, 3–11. [Google Scholar] [CrossRef]
- Gargallo-Garriga, A.; Preece, C.; Sardans, J.; Oravec, M.; Urban, O.; Peñuelas, J. Root exudate metabolomes change under drought and show limited capacity for recovery. Sci. Rep. 2018, 8, 12696. [Google Scholar] [CrossRef]
- Henry, A.; Doucette, W.; Norton, J.; Bugbee, B. Changes in crested wheatgrass root exudation caused by flood, drought, and nutrient stress. J. Environ. Qual. 2007, 36, 904–912. [Google Scholar] [CrossRef]
- Wang, E.L.H.; Bergeson, G.B. Biochemical changes in root exudate and xylem sap of tomato plants infected with Meloidogyne incognita. J. Nematol. 1974, 6, 194. [Google Scholar]
- Neumann, G.; Bott, S.; Ohler, M.A.; Mock, H.P.; Lippmann, R.; Grosch, R.; Smalla, K. Root exudation and root development of lettuce (Lactuca sativa l. Cv. Tizian) as affected by different soils. Front. Microbiol. 2014, 5, 2. [Google Scholar] [CrossRef]
- Guttman, D.S.; McHardy, A.C.; Schulze-Lefert, P. Microbial genome-enabled insights into plant–microorganism interactions. Nat. Rev. Genet. 2014, 15, 797–813. [Google Scholar] [CrossRef]
- Mohanram, S.; Kumar, P. Rhizosphere microbiome: Revisiting the synergy of plant-microbe interactions. Ann. Microbiol. 2019, 69, 307–320. [Google Scholar] [CrossRef]
- Virk, A.L.; Lin, B.-J.; Kan, Z.-R.; Qi, J.-Y.; Dang, Y.P.; Lal, R.; Zhao, X.; Zhang, H.-L. Simultaneous effects of legume cultivation on carbon and nitrogen accumulation in soil. In Advances in Agronomy; Elsevier: Amsterdam, The Netherlands, 2022; Volume 171, pp. 75–110. [Google Scholar]
- Stajković, O.; Delić, D.; Jošić, D.; Kuzmanović, D.; Rasulić, N.; Knežević-Vukčević, J. Improvement of common bean growth by co-inoculation with Rhizobium and plant growth-promoting bacteria. Rom. Biotechnol. Lett. 2011, 16, 5919–5926. [Google Scholar]
- Rana, A.; Kabi, S.R.; Verma, S.; Adak, A.; Pal, M.; Shivay, Y.S.; Prasanna, R.; Nain, L. Prospecting plant growth promoting bacteria and cyanobacteria as options for enrichment of macro- and micronutrients in grains in rice–wheat cropping sequence. Cogent Food. Agric. 2015, 1, 1037379. [Google Scholar] [CrossRef]
- Morales, A.; Alvear, M.; Valenzuela, E.; Rubio, R.; Borie, F. Effect of inoculation with Penicillium albidum, a phosphate-solubilizing fungus, on the growth of Trifolium pratense cropped in a volcanic soil. J. Basic Microbiol. 2007, 47, 275–280. [Google Scholar] [CrossRef] [PubMed]
- Hungria, M.; Nogueira, M.A.; Araujo, R.S.; Hungria, M.; Nogueira, M.A.; Araujo, R.S. Soybean seed co-inoculation with Bradyrhizobium spp. and Azospirillum brasilense: A new biotechnological tool to improve yield and sustainability. Am. J. Plant Sci. 2015, 6, 811–817. [Google Scholar] [CrossRef]
- Alkooranee, J.T.; Aledan, T.R.; Ali, A.K.; Lu, G.; Zhang, X.; Wu, J.; Fu, C.; Li, M. Detecting the hormonal pathways in oilseed rape behind induced systemic resistance by Trichoderma harzianum TH12 to Sclerotinia sclerotiorum. PLoS ONE 2017, 12, e0168850. [Google Scholar] [CrossRef]
- Leonetti, P.; Zonno, M.C.; Molinari, S.; Altomare, C. Induction of SA-signaling pathway and ethylene biosynthesis in Trichoderma harzianum-treated tomato plants after infection of the root-knot nematode Meloidogyne incognita. Plant Cell Rep. 2017, 36, 621–631. [Google Scholar] [CrossRef]
- Qin, S.; Zhang, Y.-J.; Yuan, B.; Xu, P.-Y.; Xing, K.; Wang, J.; Jiang, J.-H. Isolation of ACC deaminase-producing habitat-adapted symbiotic bacteria associated with halophyte Limonium sinense (Girard) Kuntze and evaluating their plant growth-promoting activity under salt stress. Plant Soil 2014, 374, 753–766. [Google Scholar] [CrossRef]
- Rafique, H.M.; Khan, M.Y.; Asghar, H.N.; Ahmad Zahir, Z.; Nadeem, S.M.; Sohaib, M.; Alotaibi, F.; Al-Barakah, F.N.I. Converging alfalfa (Medicago sativa L.) and petroleum hydrocarbon acclimated ACC-deaminase containing bacteria for phytoremediation of petroleum hydrocarbon contaminated soil. Int. J. Phytoremediation 2022. [Google Scholar] [CrossRef]
- Vurukonda, S.S.K.P.; Vardharajula, S.; Shrivastava, M.; SkZ, A. Enhancement of drought stress tolerance in crops by plant growth promoting rhizobacteria. Microbiol. Res. 2016, 184, 13–24. [Google Scholar] [CrossRef]
- Glick, B.R.; Gamalero, E. Recent developments in the study of plant microbiomes. Microorganisms 2021, 9, 1533. [Google Scholar] [CrossRef] [PubMed]
- Gobbato, E.; Wang, E.; Higgins, G.; Bano, S.A.; Henry, C.; Schultze, M.; Oldroyd, G.E.D. RAM1 and RAM2 function and expression during arbuscular mycorrhizal symbiosis and Aphanomyces euteiches colonization. Plant Signal. Behav. 2013, 8, e26049. [Google Scholar] [CrossRef] [PubMed]
- Keshavan, N.D.; Chowdhary, P.K.; Haines, D.C.; González, J.E. L-canavanine made by Medicago sativa interferes with quorum sensing in Sinorhizobium meliloti. J. Bacteriol. 2005, 187, 8427–8436. [Google Scholar] [CrossRef] [PubMed]
- Kravchenko, L.V.; Azarova, T.S.; Leonova-Erko, E.I.; Shaposhnikov, A.I.; Makarova, N.M.; Tikhonovich, I.A. Root exudates of tomato plants and their effect on the growth and antifungal activity of Pseudomonas strains. Microbiology 2003, 72, 37–41. [Google Scholar] [CrossRef]
- Rasmann, S.; Köllner, T.G.; Degenhardt, J.; Hiltpold, I.; Toepfer, S.; Kuhlmann, U.; Gershenzon, J.; Turlings, T.C.J. Recruitment of entomopathogenic nematodes by insect-damaged maize roots. Nature 2005, 434, 732–737. [Google Scholar] [CrossRef] [PubMed]
- Zhang, N.; Wang, D.; Liu, Y.; Li, S.; Shen, Q.; Zhang, R. Effects of different plant root exudates and their organic acid components on chemotaxis, biofilm formation and colonization by beneficial rhizosphere-associated bacterial strains. Plant Soil 2014, 374, 689–700. [Google Scholar] [CrossRef]
- Danhorn, T.; Fuqua, C. Biofilm formation by plant-associated bacteria. Annu. Rev. Microbiol. 2007, 61, 401–422. [Google Scholar] [CrossRef]
- Jiménez Bremont, J.; Marina, M.; de la Luz Guerrero-González, M.; Rossi, F.; Sánchez-Rangel, D.; Rodríguez-Kessler, M.; Ruiz, O.; Gárriz, A. Physiological and molecular implications of plant polyamine metabolism during biotic interactions. Front. Plant Sci. 2014, 5, 95. [Google Scholar] [CrossRef]
- Liu, Z.; Beskrovnaya, P.; Melnyk, R.A.; Hossain, S.S.; Khorasani, S.; O’Sullivan, L.R.; Wiesmann, C.L.; Bush, J.; Richard, J.D.; Haney, C.H. A genome-wide screen identifies genes in rhizosphere-associated Pseudomonas required to evade plant defenses. mBio 2018, 9, e00433-18. [Google Scholar] [CrossRef]
- Nelson, M.S.; Sadowsky, M.J. Secretion systems and signal exchange between nitrogen-fixing rhizobia and legumes. Front. Plant Sci. 2015, 6, 491. [Google Scholar] [CrossRef]
- Wang, Q.; Liu, J.; Zhu, H. Genetic and molecular mechanisms underlying symbiotic specificity in legume-rhizobium interactions. Front. Plant Sci. 2018, 9, 313. [Google Scholar] [CrossRef] [PubMed]
- Hugoni, M.; Luis, P.; Guyonnet, J.; el Zahar Haichar, F. Plant host habitat and root exudates shape fungal diversity. Mycorrhiza 2018, 28, 451–463. [Google Scholar] [CrossRef] [PubMed]
- Hu, L.; Robert, C.A.M.; Cadot, S.; Zhang, X.; Ye, M.; Li, B.; Manzo, D.; Chervet, N.; Steinger, T.; van der Heijden, M.G.A.; et al. Root exudate metabolites drive plant-soil feedbacks on growth and defense by shaping the rhizosphere microbiota. Nat. Commun. 2018, 9, 2738. [Google Scholar] [CrossRef] [PubMed]
- Kawasaki, A.; Dennis, P.G.; Forstner, C.; Raghavendra, A.K.H.; Mathesius, U.; Richardson, A.E.; Delhaize, E.; Gilliham, M.; Watt, M.; Ryan, P.R. Manipulating exudate composition from root apices shapes the microbiome throughout the root system. Plant Physiol. 2021, 187, 2279–2295. [Google Scholar] [CrossRef]
- Kudjordjie, E.N.; Sapkota, R.; Nicolaisen, M. Arabidopsis assemble distinct root-associated microbiomes through the synthesis of an array of defense metabolites. PLoS ONE 2021, 16, e0259171. [Google Scholar] [CrossRef]
- Qu, Q.; Li, Y.; Zhang, Z.; Cui, H.; Zhao, Q.; Liu, W.; Lu, T.; Qian, H. Effects of S-metolachlor on wheat (Triticum aestivum L.) seedling root exudates and the rhizosphere microbiome. J. Hazard. Mater. 2021, 411, 125137. [Google Scholar] [CrossRef]
- Weisskopf, L.; Abou-Mansour, E.; Fromin, N.; Tomasi, N.; Santelia, D.; Edelkott, I.; Neumann, G.; Aragno, M.; Tabacchi, R.; Martinoia, E. White lupin has developed a complex strategy to limit microbial degradation of secreted citrate required for phosphate acquisition. Plant Cell Environ. 2006, 29, 919–927. [Google Scholar] [CrossRef]
- EL Zahar Haichar, F.; Marol, C.; Berge, O.; Rangel-Castro, J.I.; Prosser, J.I.; Balesdent, J.; Heulin, T.; Achouak, W. Plant host habitat and root exudates shape soil bacterial community structure. ISME J. 2008, 2, 1221–1230. [Google Scholar] [CrossRef]
- Sasse, J.; Martinoia, E.; Northen, T. Feed your friends: Do plant exudates shape the root microbiome? Trends Plant Sci. 2018, 23, 25–41. [Google Scholar] [CrossRef]
- Cai, T.; Cai, W.; Zhang, J.; Zheng, H.; Tsou, A.M.; Xiao, L.; Zhong, Z.; Zhu, J. Host legume-exuded antimetabolites optimize the symbiotic rhizosphere. Mol. Microbiol. 2009, 73, 507–517. [Google Scholar] [CrossRef]
- Mardani-Korrani, H.; Nakayasu, M.; Yamazaki, S.; Aoki, Y.; Kaida, R.; Motobayashi, T.; Kobayashi, M.; Ohkama-Ohtsu, N.; Oikawa, Y.; Sugiyama, A.; et al. L-canavanine, a root exudate from hairy vetch (Vicia villosa) drastically affecting the soil microbial community and metabolite pathways. Front. Microbiol. 2021, 12, 701796. [Google Scholar] [CrossRef] [PubMed]
- Stringlis, I.A.; Yu, K.; Feussner, K.; de Jonge, R.; van Bentum, S.; van Verk, M.C.; Berendsen, R.L.; Bakker, P.A.H.M.; Feussner, I.; Pieterse, C.M.J. MYB72-dependent coumarin exudation shapes root microbiome assembly to promote plant health. PNAS 2018, 115, E5213–E5222. [Google Scholar] [CrossRef] [PubMed]
- Baune, M.; Kang, K.; Schenkeveld, W.D.C.; Kraemer, S.M.; Hayen, H.; Weber, G. Importance of oxidation products in coumarin-mediated Fe(hydr)oxide mineral dissolution. BioMetals 2020, 33, 305–321. [Google Scholar] [CrossRef] [PubMed]
- Toljander, J.F.; Lindahl, B.D.; Paul, L.R.; Elfstrand, M.; Finlay, R.D. Influence of arbuscular mycorrhizal mycelial exudates on soil bacterial growth and community structure. FEMS Microbiol. Ecol. 2007, 61, 295–304. [Google Scholar] [CrossRef] [PubMed]
- Badri, D.V.; Quintana, N.; el Kassis, E.G.; Kim, H.K.; Choi, Y.H.; Sugiyama, A.; Verpoorte, R.; Martinoia, E.; Manter, D.K.; Vivanco, J.M. An ABC transporter mutation alters root exudation of phytochemicals that provoke an overhaul of natural soil microbiota. Plant Physiol. 2009, 151, 2006–2017. [Google Scholar] [CrossRef] [PubMed]
- Vives-Peris, V.; de Ollas, C.; Gómez-Cadenas, A.; Pérez-Clemente, R.M. Root exudates: From plant to rhizosphere and beyond. Plant Cell Rep. 2020, 39, 3–17. [Google Scholar] [CrossRef]
- Volkov, V.; Schwenke, H. A quest for mechanisms of plant root exudation brings new results and models, 300 years after hales. Plants 2020, 10, 38. [Google Scholar] [CrossRef]
- Kapilan, R.; Vaziri, M.; Zwiazek, J.J. Regulation of aquaporins in plants under stress. Biol. Res. 2018, 51, 4. [Google Scholar] [CrossRef]
- Dietz, S.; von Bülow, J.; Beitz, E.; Nehls, U. The aquaporin gene family of the ectomycorrhizal fungus Laccaria bicolor: Lessons for symbiotic functions. N. Phytol. 2011, 190, 927–940. [Google Scholar] [CrossRef]
- Hwang, J.H.; Ellingson, S.R.; Roberts, D.M. Ammonia permeability of the soybean nodulin 26 channel. FEBS Lett. 2010, 584, 4339–4343. [Google Scholar] [CrossRef]
- Masalkar, P.; Wallace, I.S.; Hwang, J.H.; Roberts, D.M. Interaction of cytosolic glutamine synthetase of soybean root nodules with the C-terminal domain of the symbiosome membrane nodulin 26 aquaglyceroporin. J. Biol. Chem. 2010, 285, 23880–23888. [Google Scholar] [CrossRef] [PubMed]
- Janczak, K.; Dąbrowska, G.; Znajewska, Z.; Hrynkiewicz, K. Effect of bacterial inoculation on the growth of miscanthus and bacterial and fungal density in the polymer-containing soil. Part 2. Non-biodegradable polymers. Chem. Ind. 2014, 93, 2222–2225. [Google Scholar]
- Janczak, K.; Dąbrowska, G.; Znajewska, Z.; Hrynkiewicz, K. Effect of bacterial inoculation on the growth of miscanthus and bacterial and fungal density in the polymer-containing soil. Part 1. Biodegradable polymers. Chem. Ind. 2014, 93, 2218–2221. [Google Scholar]
- Janczak, K.; Dąbrowska, G.B.; Raszkowska-Kaczor, A.; Kaczor, D.; Hrynkiewicz, K.; Richert, A. Biodegradation of the plastics PLA and PET in cultivated soil with the participation of microorganisms and plants. Int. Biodeterior. Biodegrad. 2020, 155, 105087. [Google Scholar] [CrossRef]
- Afzal, M.; Yousaf, S.; Reichenauer, T.G.; Sessitsch, A. The inoculation method affects colonization and performance of bacterial inoculant strains in the phytoremediation of soil contaminated with diesel oil. Int. J. Phytoremediation 2012, 14, 35–47. [Google Scholar] [CrossRef] [PubMed]
- Yousaf, S.; Afzal, M.; Reichenauer, T.G.; Brady, C.L.; Sessitsch, A. Hydrocarbon degradation, plant colonization and gene expression of alkane degradation genes by endophytic Enterobacter ludwigii strains. Environ. Pollut. 2011, 159, 2675–2683. [Google Scholar] [CrossRef] [PubMed]
- Louvel, B.; Cébron, A.; Leyval, C. Root exudates affect phenanthrene biodegradation, bacterial community and functional gene expression in sand microcosms. Int. Biodeterior. Biodegrad. 2011, 65, 947–953. [Google Scholar] [CrossRef]
- Chen, L.; Yang, X.; Raza, W.; Li, J.; Liu, Y.; Qiu, M.; Zhang, F.; Shen, Q. Trichoderma harzianum SQR-T037 rapidly degrades allelochemicals in rhizospheres of continuously cropped cucumbers. Appl. Microbiol. Biotechnol. 2011, 89, 1653–1663. [Google Scholar] [CrossRef]
- Wang, Y.; Li, H.; Feng, G.; Du, L.; Zeng, D. Biodegradation of diuron by an endophytic fungus Neurospora intermedia DP8-1 isolated from sugarcane and its potential for remediating diuron-contaminated soils. PLoS ONE 2017, 12, e0182556. [Google Scholar] [CrossRef]
- Abdelghany, T.M.; El-Ghany Tm, A.; Masmali, I.A. Fungal biodegradation of organophosphorus insecticides and their impact on soil microbial population. J. Plant Pathol. Microbiol. 2016, 7, 1000349. [Google Scholar]
- Guerrero, R.; Margulis, L.; Berlanga, M. Symbiogenesis: The holobiont as a unit of evolution. Int. Microbiol. 2013, 16, 133–143. [Google Scholar] [PubMed]
- Trivedi, P.; Leach, J.E.; Tringe, S.G.; Sa, T.; Singh, B.K. Plant–microbiome interactions: From community assembly to plant health. Nat. Rev. Microbiol. 2020, 18, 607–621. [Google Scholar] [CrossRef] [PubMed]
- Santhanam, R.; Luu, V.T.; Weinhold, A.; Goldberg, J.; Oh, Y.; Baldwin, I.T. Native root-associated bacteria rescue a plant from a sudden-wilt disease that emerged during continuous cropping. Proc. Natl. Acad. Sci. USA 2015, 112, E5013–E5120. [Google Scholar] [CrossRef]
- Garbeva, P.; Silby, M.W.; Raaijmakers, J.M.; Levy, S.B.; de Boer, W. Transcriptional and antagonistic responses of Pseudomonas fluorescens Pf0-1 to phylogenetically different bacterial competitors. ISME J. 2011, 5, 973–985. [Google Scholar] [CrossRef] [PubMed]
- Arendt, K.R.; Hockett, K.L.; Araldi-Brondolo, S.J.; Baltrus, D.A.; Arnold, A.E. Isolation of endohyphal bacteria from foliar Ascomycota and in vitro establishment of their symbiotic associations. Appl. Environ. Microbiol. 2016, 82, 2943–2949. [Google Scholar] [CrossRef]
- Hoffman, M.T.; Gunatilaka, M.K.; Wijeratne, K.; Gunatilaka, L.; Arnold, A.E. Endohyphal bacterium enhances production of indole-3-acetic acid by a foliar fungal endophyte. PLoS ONE 2013, 8, e73132. [Google Scholar] [CrossRef]
- Márquez, L.M.; Redman, R.S.; Rodriguez, R.J.; Roossinck, M.J. A virus in a fungus in a plant: Three-way symbiosis required for thermal tolerance. Science 2007, 315, 513–515. [Google Scholar] [CrossRef]
- Rodriguez, R.J.; Henson, J.; van Volkenburgh, E.; Hoy, M.; Wright, L.; Beckwith, F.; Kim, Y.-O.; Redman, R.S. Stress tolerance in plants via habitat-adapted symbiosis. ISME J. 2008, 2, 404–416. [Google Scholar] [CrossRef]
- Chen, H.; Wu, H.; Yan, B.; Zhao, H.; Liu, F.; Zhang, H.; Sheng, Q.; Miao, F.; Liang, Z. Core microbiome of medicinal plant Salvia miltiorrhiza seed: A rich reservoir of beneficial microbes for secondary metabolism? Int. J. Mol. Sci. 2018, 19, 672. [Google Scholar] [CrossRef]
- Redford, A.J.; Bowers, R.M.; Knight, R.; Linhart, Y.; Fierer, N. The ecology of the phyllosphere: Geographic and phylogenetic variability in the distribution of bacteria on tree leaves: Biogeography of phyllosphere bacterial communities. Environ. Microbiol. 2010, 12, 2885–2893. [Google Scholar] [CrossRef]
- Germida, J.; Siciliano, S. Taxonomic diversity of bacteria associated with the roots of modern, recent and ancient wheat cultivars. Biol. Fertil. Soils 2001, 33, 410–415. [Google Scholar]
- Churchland, C.; Grayston, S.J. Specificity of plant-microbe interactions in the tree mycorrhizosphere biome and consequences for soil C cycling. Front. Microbiol. 2014, 5, 261. [Google Scholar] [CrossRef] [PubMed]
- Grubisha, L.C.; Trappe, J.M.; Molina, R.; Spatafora, J.W. Biology of the ectomycorrhizal genus Rhizopogon. VI. Re-examination of infrageneric relationships inferred from phylogenetic analyses of ITS sequences. Mycologia 2002, 94, 607–619. [Google Scholar] [CrossRef]
- Young, J.P.W.; Johnston, A.W.B. The evolution of specificity in the legume-rhizobium symbiosis. Trends Ecol. Evol. 1989, 4, 341–349. [Google Scholar] [CrossRef]
- Wicaksono, W.A.; Cernava, T.; Berg, C.; Berg, G. Bog ecosystems as a playground for plant–microbe coevolution: Bryophytes and vascular plants harbour functionally adapted bacteria. Microbiome 2021, 9, 170. [Google Scholar] [CrossRef] [PubMed]
- Cobian, G.M.; Egan, C.P.; Amend, A.S. Plant–microbe specificity varies as a function of elevation. ISME J. 2019, 13, 2778–2788. [Google Scholar] [CrossRef]
- Salas-González, I.; Reyt, G.; Flis, P.; Custódio, V.; Gopaulchan, D.; Bakhoum, N.; Dew, T.P.; Suresh, K.; Franke, R.B.; Dangl, J.L.; et al. Coordination between microbiota and root endodermis supports plant mineral nutrient homeostasis. Science 2021, 371, eabd0695. [Google Scholar] [CrossRef]
- Compant, S.; Samad, A.; Faist, H.; Sessitsch, A. A review on the plant microbiome: Ecology, functions, and emerging trends in microbial application. J. Adv. Res. 2019, 19, 29–37. [Google Scholar] [CrossRef]
- Mitter, B.; Pfaffenbichler, N.; Sessitsch, A. Plant–microbe partnerships in 2020. Microb. Biotechnol. 2016, 9, 635–640. [Google Scholar] [CrossRef]
- Abdelfattah, A.; Wisniewski, M.; Schena, L.; Tack, A.J.M. Experimental evidence of microbial inheritance in plants and transmission routes from seed to phyllosphere and root. Environ. Microbiol. 2021, 23, 2199–2214. [Google Scholar] [CrossRef]
- Sánchez-López, A.S.; Pintelon, I.; Stevens, V.; Imperato, V.; Timmermans, J.P.; González-Chávez, C.; Carrillo-González, R.; van hamme, J.; Vangronsveld, J.; Thijs, S. Seed endophyte microbiome of Crotalaria pumila unpeeled: Identification of plant-beneficial methylobacteria. Int. J. Mol. Sci. 2018, 19, 291. [Google Scholar] [CrossRef]
- Coleman-Derr, D.; Desgarennes, D.; Fonseca-Garcia, C.; Gross, S.; Clingenpeel, S.; Woyke, T.; North, G.; Visel, A.; Partida-Martinez, L.P.; Tringe, S.G. Plant compartment and biogeography affect microbiome composition in cultivated and native Agave species. N. Phytol. 2016, 209, 798–811. [Google Scholar] [CrossRef] [PubMed]
- Zarraonaindia, I.; Owens, S.M.; Weisenhorn, P.; West, K.; Hampton-Marcell, J.; Lax, S.; Bokulich, N.A.; Mills, D.A.; Martin, G.; Taghavi, S.; et al. The soil microbiome influences grapevine-associated microbiota. mBio 2015, 6, e02527-14. [Google Scholar] [CrossRef] [PubMed]
- De Souza, R.S.C.; Okura, V.K.; Armanhi, J.S.L.; Jorrín, B.; Lozano, N.; da Silva, M.J.; González-Guerrero, M.; de Araújo, L.M.; Verza, N.C.; Bagheri, H.C.; et al. Unlocking the bacterial and fungal communities assemblages of sugarcane microbiome. Sci. Rep. 2016, 6, 28774. [Google Scholar] [CrossRef] [PubMed]
- Shade, A.; McManus, P.S.; Handelsman, J. Unexpected diversity during community succession in the apple flower microbiome. mBio 2013, 4, e00602-12. [Google Scholar] [CrossRef] [PubMed]
- Wipf, H.M.L.; Bùi, T.N.; Coleman-Derr, D. Distinguishing between the impacts of heat and drought stress on the root microbiome of Sorghum bicolor. Phytobiomes J. 2021, 5, 166–176. [Google Scholar] [CrossRef]
- Nuccio, E.E.; Anderson-Furgeson, J.; Estera, K.Y.; Pett-Ridge, J.; de Valpine, P.; Brodie, E.L.; Firestone, M.K. Climate and edaphic controllers influence rhizosphere community assembly for a wild annual grass. Ecology 2016, 97, 1307–1318. [Google Scholar] [CrossRef]
- Gargouri, M.; Karray, F.; Chebaane, A.; Mhiri, N.; Partida-Martínez, L.P.; Sayadi, S.; Mliki, A. Increasing aridity shapes beta diversity and the network dynamics of the belowground fungal microbiome associated with Opuntia ficus-indica. Sci. Total Environ. 2021, 773, 145008. [Google Scholar] [CrossRef]
- Lau, J.A.; Lennon, J.T. Rapid responses of soil microorganisms improve plant fitness in novel environments. Proc. Natl. Acad. Sci. USA 2012, 109, 14058–14062. [Google Scholar] [CrossRef]
- Zhang, Y.; Jiang, W.; Li, Q.; Xu, W.; Wang, J.; Hu, J.; Zhang, Z. Soil nutrient levels determine the variation of bacterial communities in the rhizosphere of rice under different conditions of climate and genotype. Appl. Soil Ecol. 2021, 167, 104025. [Google Scholar] [CrossRef]
- Pii, Y.; Borruso, L.; Brusetti, L.; Crecchio, C.; Cesco, S.; Mimmo, T. The interaction between iron nutrition, plant species and soil type shapes the rhizosphere microbiome. Plant Physiol. Biochem. 2016, 99, 39–48. [Google Scholar] [CrossRef] [PubMed]
- Rastogi, G.; Sbodio, A.; Tech, J.J.; Suslow, T.V.; Coaker, G.L.; Leveau, J.H.J. Leaf microbiota in an agroecosystem: Spatiotemporal variation in bacterial community composition on field-grown lettuce. ISME J. 2012, 6, 1812–1822. [Google Scholar] [CrossRef] [PubMed]
- Gontia-Mishra, I.; Sapre, S.; Deshmukh, R.; Sikdar, S.; Tiwari, S. Microbe-mediated drought tolerance in plants: Current developments and future challenges. In Plant Microbiomes for Sustainable Agriculture; Yadav, A.N., Singh, J., Rastegari, A.A., Yadav, N., Eds.; Springer: Berlin/Heidelberg, Germany, 2020; pp. 351–379. [Google Scholar]
- Naylor, D.; DeGraaf, S.; Purdom, E.; Coleman-Derr, D. Drought and host selection influence bacterial community dynamics in the grass root microbiome. ISME J. 2017, 11, 2691–2704. [Google Scholar] [CrossRef] [PubMed]
- Santos-Medellín, C.; Edwards, J.; Liechty, Z.; Nguyen, B.; Sundaresan, V. Drought stress results in a compartment-specific restructuring of the rice root-associated microbiomes. mBio 2017, 8, e00764-17. [Google Scholar] [CrossRef] [PubMed]
- Nelson, A.R.; Narrowe, A.B.; Rhoades, C.C.; Fegel, T.S.; Daly, R.A.; Roth, H.K.; Chu, R.K.; Amundson, K.K.; Young, R.B.; Steindorff, A.S.; et al. Wildfire-dependent changes in soil microbiome diversity and function. Nat. Microbiol. 2022, 7, 1419–1430. [Google Scholar] [CrossRef]
- Wei, F.; Feng, H.; Zhang, D.; Feng, Z.; Zhao, L.; Zhang, Y.; Deakin, G.; Peng, J.; Zhu, H.; Xu, X. Composition of rhizosphere microbial communities associated with healthy and Verticillium wilt diseased cotton plants. Front. Microbiol. 2021, 12, 618169. [Google Scholar] [CrossRef]
- Lazcano, C.; Boyd, E.; Holmes, G.; Hewavitharana, S.; Pasulka, A.; Ivors, K. The rhizosphere microbiome plays a role in the resistance to soil-borne pathogens and nutrient uptake of strawberry cultivars under field conditions. Sci. Rep. 2021, 11, 3188. [Google Scholar] [CrossRef]
- Kong, H.G.; Kim, B.K.; Song, G.C.; Lee, S.; Ryu, C.-M. Aboveground whitefly infestation-mediated reshaping of the root microbiota. Front. Microbiol. 2016, 7, 1314. [Google Scholar] [CrossRef]
- Turnbaugh, P.J.; Ley, R.E.; Hamady, M.; Fraser-Liggett, C.M.; Knight, R.; Gordon, J.I. The human microbiome project. Nature 2007, 449, 804–810. [Google Scholar] [CrossRef]
- Neu, A.T.; Allen, E.E.; Roy, K. Defining and quantifying the core microbiome: Challenges and prospects. Proc. Natl. Acad. Sci. USA 2021, 118, e2104429118. [Google Scholar] [CrossRef]
- Lay, C.-Y.; Bell, T.H.; Hamel, C.; Harker, K.N.; Mohr, R.; Greer, C.W.; Yergeau, É.; St-Arnaud, M. Canola root–associated microbiomes in the canadian prairies. Front. Microbiol. 2018, 9, 1188. [Google Scholar] [CrossRef] [PubMed]
- Hassani, M.A.; Durán, P.; Hacquard, S. Microbial interactions within the plant holobiont. Microbiome 2018, 6, 58. [Google Scholar] [CrossRef]
- Agler, M.T.; Ruhe, J.; Kroll, S.; Morhenn, C.; Kim, S.T.; Weigel, D.; Kemen, E.M. Microbial hub taxa link host and abiotic factors to plant microbiome variation. PLoS Biol. 2016, 14, e1002352. [Google Scholar] [CrossRef] [PubMed]
- Niu, B.; Paulson, J.N.; Zheng, X.; Kolter, R. Simplified and representative bacterial community of maize roots. Proc. Natl. Acad. Sci. USA 2017, 114, E2450–E2459. [Google Scholar] [CrossRef] [PubMed]
- Margulis, L.; Fester, R. Symbiosis as a source of evolutionary innovation: Speciation and morphogenesis. In Cambridge Mass; MIT Press: Cambridge, MA, USA, 1991; pp. 1–14. [Google Scholar]
- Teixeira, P.J.P.; Colaianni, N.R.; Fitzpatrick, C.R.; Dangl, J.L. Beyond pathogens: Microbiota interactions with the plant immune system. Curr. Opin. Microbiol. 2019, 49, 7–17. [Google Scholar] [CrossRef] [PubMed]
- Harman, G.E.; Uphoff, N. Symbiotic root-endophytic soil microbes improve crop productivity and provide environmental benefits. Scientifica 2019, 2019, 9106395. [Google Scholar] [CrossRef] [PubMed]
- Zilber-Rosenberg, I.; Rosenberg, E. Role of microorganisms in the evolution of animals and plants: The hologenome theory of evolution. FEMS Microbiol. Rev. 2008, 32, 723–735. [Google Scholar] [CrossRef]
- Rosenberg, E.; Koren, O.; Reshef, L.; Efrony, R.; Zilber-Rosenberg, I. The role of microorganisms in coral health, disease and evolution. Nat. Rev. Microbiol. 2007, 5, 355–362. [Google Scholar] [CrossRef]
- Rosenberg, E.; Zilber-Rosenberg, I. The hologenome concept of evolution after 10 years. Microbiome 2018, 6, 78. [Google Scholar] [CrossRef]
- Romero, F.M.; Marina, M.; Pieckenstain, F.L.; Rossi, F.R.; Gonzalez, M.E.; Vignatti, P.; Gárriz, A. Gaining insight into plant responses to beneficial and pathogenic microorganisms using metabolomic and transcriptomic approaches. In Metabolic Engineering for Bioactive Compounds: Strategies and Processes; Springer: Berlin/Heidelberg, Germany, 2017; pp. 113–140. [Google Scholar]
- Dangl, J.L.; Horvath, D.M.; Staskawicz, B.J. Pivoting the plant immune system from dissection to deployment. Science 2013, 341, 746–751. [Google Scholar] [CrossRef]
- Jubic, L.M.; Saile, S.; Furzer, O.J.; el Kasmi, F.; Dangl, J.L. Help wanted: Helper NLRs and plant immune responses. Curr. Opin. Plant Biol. 2019, 50, 82–94. [Google Scholar] [CrossRef] [PubMed]
- Block, A.; Li, G.; Fu, Z.Q.; Alfano, J.R. Phytopathogen type III effector weaponry and their plant targets. Curr. Opin. Plant Biol. 2008, 11, 396–403. [Google Scholar] [CrossRef] [PubMed]
- Thomma, B.P.H.J.; Nürnberger, T.; Joosten, M.H.A.J. Of PAMPs and effectors: The blurred PTI-ETI dichotomy. Plant Cell 2011, 23, 4–15. [Google Scholar] [CrossRef]
- Yuan, M.; Jiang, Z.; Bi, G.; Nomura, K.; Liu, M.; Wang, Y.; Cai, B.; Zhou, J.M.; He, S.Y.; Xin, X.F. Pattern-recognition receptors are required for NLR-mediated plant immunity. Nature 2021, 592, 105–109. [Google Scholar] [CrossRef]
- Teixeira, P.J.P.L.; Colaianni, N.R.; Law, T.F.; Conway, J.M.; Gilbert, S.; Li, H.; Salas-González, I.; Panda, D.; del Risco, N.M.; Finkel, O.M.; et al. Specific modulation of the root immune system by a community of commensal bacteria. Proc. Natl. Acad. Sci. USA 2021, 118, e2100678118. [Google Scholar] [CrossRef]
- Bai, B.; Liu, W.; Qiu, X.; Zhang, J.; Zhang, J.; Bai, Y. The root microbiome: Community assembly and its contributions to plant fitness. J. Integr. Plant Biol. 2022, 64, 230–243. [Google Scholar] [CrossRef]
- Zhou, F.; Emonet, A.; Dénervaud Tendon, V.; Marhavy, P.; Wu, D.; Lahaye, T.; Geldner, N. Co-incidence of damage and microbial patterns controls localized immune responses in roots. Cell 2020, 180, 440–453.e18. [Google Scholar] [CrossRef]
- Saravanakumar, K.; Fan, L.; Fu, K.; Yu, C.; Wang, M.; Xia, H.; Sun, J.; Li, Y.; Chen, J. Cellulase from Trichoderma harzianum interacts with roots and triggers induced systemic resistance to foliar disease in maize. Sci. Rep. 2016, 6, 35543. [Google Scholar] [CrossRef] [PubMed]
- Zahir, Z.A.; Ghani, U.; Naveed, M.; Nadeem, S.M.; Asghar, H.N. Comparative effectiveness of Pseudomonas and Serratia sp. containing ACC-deaminase for improving growth and yield of wheat (Triticum aestivum L.) under salt-stressed conditions. Arch. Microbiol. 2009, 191, 415–424. [Google Scholar] [CrossRef]
- Adhikari, P.; Pandey, A. Phosphate solubilization potential of endophytic fungi isolated from Taxus wallichiana Zucc. roots. Rhizosphere 2019, 9, 2–9. [Google Scholar] [CrossRef]
- Zhao, L.; Zhang, Y.Q. Effects of phosphate solubilization and phytohormone production of Trichoderma asperellum Q1 on promoting cucumber growth under salt stress. J. Integr. Agric. 2015, 14, 1588–1597. [Google Scholar] [CrossRef]
- Fochi, V.; Chitarra, W.; Kohler, A.; Voyron, S.; Singan, V.R.; Lindquist, E.A.; Barry, K.W.; Girlanda, M.; Grigoriev, I.V.; Martin, F.; et al. Fungal and plant gene expression in the Tulasnella calospora–Serapias vomeracea symbiosis provides clues about nitrogen pathways in orchid mycorrhizas. N. Phytol. 2017, 213, 365–379. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Hu, Y.J.; Hao, Z.P.; Li, H.; Wang, Y.S.; Chen, B.D. First cloning and characterization of two functional aquaporin genes from an arbuscular mycorrhizal fungus Glomus intraradices. N. Phytol. 2013, 197, 617–630. [Google Scholar] [CrossRef] [PubMed]
- Gujar, P.D.; Bhavsar, K.P.; Khire, J.M. Effect of phytase from Aspergillus niger on plant growth and mineral assimilation in wheat (Triticum aestivum Linn.) and its potential for use as a soil amendment. J. Sci. Food Agric. 2013, 93, 2242–2247. [Google Scholar] [CrossRef] [PubMed]
- Viterbo, A.; Landau, U.; Kim, S.; Chernin, L.; Chet, I. Characterization of ACC deaminase from the biocontrol and plant growth-promoting agent Trichoderma asperellum T203. FEMS Microbiol. Lett. 2010, 305, 42–48. [Google Scholar] [CrossRef]
- Viterbo, A.; Chet, I. TasHyd1, a new hydrophobin gene from the biocontrol agent Trichoderma asperellum, is involved in plant root colonization. Mol. Plant Pathol. 2006, 7, 249–258. [Google Scholar] [CrossRef]
- Samolski, I.; Rincón, A.M.; Pinzón, L.M.; Viterbo, A.; Monte, E. The qid74 gene from Trichoderma harzianum has a role in root architecture and plant biofertilization. Microbiology 2012, 158, 129–138. [Google Scholar] [CrossRef]
- Helber, N.; Wippel, K.; Sauer, N.; Schaarschmidt, S.; Hause, B.; Requena, N. A versatile monosaccharide transporter that operates in the arbuscular mycorrhizal fungus Glomus sp is crucial for the symbiotic relationship with plants. Plant Cell 2011, 23, 3812–3823. [Google Scholar] [CrossRef]
- Viterbo, A.; Wiest, A.; Brotman, Y.; Chet, I.; Kenerley, C. The 18mer peptaibols from Trichoderma virens elicit plant defence responses. Mol. Plant Pathol. 2007, 8, 737–746. [Google Scholar] [CrossRef]
- Rubio, M.B.; Hermosa, R.; Reino, J.L.; Collado, I.G.; Monte, E. Thctf1 transcription factor of Trichoderma harzianum is involved in 6-pentyl-2H-pyran-2-one production and antifungal activity. Fungal Genet. Biol. 2009, 46, 17–27. [Google Scholar] [CrossRef]
- Kubicek, C.P.; Herrera-Estrella, A.; Seidl-Seiboth, V.; Martinez, D.A.; Druzhinina, I.S.; Thon, M.; Zeilinger, S.; Casas-Flores, S.; Horwitz, B.A.; Mukherjee, P.K.; et al. Comparative genome sequence analysis underscores mycoparasitism as the ancestral life style of Trichoderma. Genome Biol. 2011, 12, R40. [Google Scholar] [CrossRef] [PubMed]
- Brotman, Y.; Briff, E.; Viterbo, A.; Chet, I. Role of swollenin, an expansin-like protein from Trichoderma, in plant root colonization. Plant Physiol. 2008, 147, 779–789. [Google Scholar] [CrossRef] [PubMed]
- Vargas, W.A.; Djonović, S.; Sukno, S.A.; Kenerley, C.M. Dimerization controls the activity of fungal elicitors that trigger systemic resistance in plants. JBC 2008, 283, 19804–19815. [Google Scholar] [CrossRef] [PubMed]
- Djonović, S.; Vargas, W.A.; Kolomiets, M.V.; Horndeski, M.; Wiest, A.; Kenerley, C.M. A proteinaceous elicitor Sm1 from the beneficial fungus Trichoderma virens is required for induced systemic resistance in maize. Plant Physiol. 2007, 145, 875–889. [Google Scholar] [CrossRef]
- Nelkner, J.; Tejerizo, G.T.; Hassa, J.; Lin, T.W.; Witte, J.; Verwaaijen, B.; Winkler, A.; Bunk, B.; Spröer, C.; Overmann, J.; et al. Genetic potential of the biocontrol agent Pseudomonas brassicacearum (Formerly P. trivialis) 3Re2-7 unraveled by genome sequencing and mining, comparative genomics and transcriptomics. Genes 2019, 10, 601. [Google Scholar] [CrossRef]
- Glick, B.R.; Jacobson, C.B.; Schwarze, M.K.; Pasternak, J.J. 1-Aminocyclopropane-1-carboxylic acid deaminase mutants of the plant growth promoting rhizobacterium Pseudomonas putida GR12-2 do not stimulate canola root elongation. Can. J. Microbiol. 2011, 40, 911–915. [Google Scholar] [CrossRef]
- Blaha, D.; Prigent-Combaret, C.; Mirza, M.S.; Moënne-Loccoz, Y. Phylogeny of the 1-aminocyclopropane-1-carboxylic acid deaminase-encoding gene acdS in phytobeneficial and pathogenic Proteobacteria and relation with strain biogeography. FEMS Microbiol. Ecol. 2006, 56, 455–470. [Google Scholar] [CrossRef]
- Taghavi, S.; van der Lelie, D.; Hoffman, A.; Zhang, Y.B.; Walla, M.D.; Vangronsveld, J.; Newman, L.; Monchy, S. Genome sequence of the plant growth promoting endophytic bacterium Enterobacter sp. 638. PLoS Genet. 2010, 6, e1000943. [Google Scholar] [CrossRef]
- Suryadi, Y.; Susilowati, D.N.; Lestari, P.; Priyatno, T.P.; Samudra, I.M.; Hikmawati, N.; Mubarik, D.N.R.; Mubarik, N.R. Characterization of bacterial isolates producing chitinase and glucanase for biocontrol of plant fungal pathogens. J. Agric. Technol. 2014, 10, 983–999. [Google Scholar]
- Gupta, C.P.; Kumar, B.; Dubey, R.C.; Maheshwari, D.K. Chitinase-mediated destructive antagonistic potential of Pseudomonas aeruginosa GRC1 against Sclerotinia sclerotiorum causing stem rot of peanut. BioControl 2006, 51, 821–835. [Google Scholar] [CrossRef]
- Ramette, A.; Frapolli, M.; Défago, G.; Moënne-Loccoz, Y. Phylogeny of HCN synthase-encoding hcnBC genes in biocontrol fluorescent pseudomonads and its relationship with host plant species and HCN synthesis ability. Mol. Plant Microbe. Interact. 2003, 16, 525–535. [Google Scholar] [CrossRef] [PubMed]
- Zimmer, W.; Hundeshagen, B.; Niederau, E. Demonstration of the indolepyruvate decarboxylase gene homologue in different auxin-producing species of the Enterobacteriaceae. Can. J. Microbiol. 1994, 40, 1072–1076. [Google Scholar] [CrossRef] [PubMed]
- Ryu, R.J.; Patten, C.L. Aromatic amino acid-dependent expression of indole-3-pyruvate decarboxylase is regulated by TyrR in Enterobacter cloacae UW5. J. Bacteriol. 2008, 190, 7200–7208. [Google Scholar] [CrossRef]
- Spaepen, S.; Versées, W.; Gocke, D.; Pohl, M.; Steyaert, J.; Vanderleyden, J. Characterization of phenylpyruvate decarboxylase, involved in auxin production of Azospirillum brasilense. J. Bacteriol. 2007, 189, 7626. [Google Scholar] [CrossRef]
- Poza-Carrión, C.; Jiménez-Vicente, E.; Navarro-Rodríguez, M.; Echavarri-Erasun, C.; Rubio, L.M. Kinetics of nif gene expression in a nitrogen-fixing bacterium. J. Bacteriol. 2014, 196, 595–603. [Google Scholar] [CrossRef] [PubMed]
- Steenhoudt, O.; Vanderleyden, J. Azospirillum, a free-living nitrogen-fixing bacterium closely associated with grasses: Genetic, biochemical and ecological aspects. FEMS Microbiol. Rev. 2000, 24, 487–506. [Google Scholar] [CrossRef]
- Satyanarayana, S.D.V.; Krishna, M.S.R.; Pavan Kumar, P.; Jeereddy, S. In silico structural homology modeling of nif A protein of rhizobial strains in selective legume plants. J. Genet. Eng. Biotechnol. 2018, 16, 731–737. [Google Scholar] [CrossRef]
- Woźniak, M.; Gałązka, A.; Tyśkiewicz, R.; Jaroszuk-ściseł, J. Endophytic bacteria potentially promote plant growth by synthesizing different metabolites and their phenotypic/physiological profiles in the Biolog GEN III MicroPlateTM Test. Int. J. Mol. Sci. 2019, 20, 5283. [Google Scholar] [CrossRef]
- Sherathia, D.; Dey, R.; Thomas, M.; Dalsania, T.; Savsani, K.; Pal, K.K. Biochemical and molecular characterization of DAPG-producing plant growthpromoting rhizobacteria (PGPR) of groundnut (Arachis hypogaea L.). Legum. Res. Int. J. 2016, 39, 614–622. [Google Scholar]
- Lynch, D.; O’Brien, J.; Welch, T.; Clarke, P.; Cuív, P.O.; Crosa, J.H.; O’Connell, M. Genetic organization of the region encoding regulation, biosynthesis, and transport of rhizobactin 1021, a siderophore produced by Sinorhizobium meliloti. J. Bacteriol. 2001, 183, 2576–2585. [Google Scholar] [CrossRef]
- Dong, H.; Beer, S.V. Riboflavin induces disease resistance in plants by activating a novel signal transduction pathway. Phytopathology 2000, 90, 801–811. [Google Scholar] [CrossRef] [PubMed]
- Xie, S.S.; Wu, H.J.; Zang, H.Y.; Wu, L.M.; Zhu, Q.Q.; Gao, X.W. Plant growth promotion by spermidine-producing Bacillus subtilis OKB105. Mol. Plant Microbe Interact. 2014, 27, 655–663. [Google Scholar] [CrossRef] [PubMed]
- Aasfar, A.; Bargaz, A.; Yaakoubi, K.; Hilali, A.; Bennis, I.; Zeroual, Y.; Meftah Kadmiri, I. Nitrogen fixing Azotobacter species as potential soil biological enhancers for crop nutrition and yield stability. Front. Microbiol. 2021, 12, 354. [Google Scholar] [CrossRef]
- Soumare, A.; Diedhiou, A.G.; Thuita, M.; Hafidi, M.; Ouhdouch, Y.; Gopalakrishnan, S.; Kouisni, L. Exploiting biological nitrogen fixation: A route towards a sustainable agriculture. Plants 2020, 9, 1011. [Google Scholar] [CrossRef]
- Ji, J.; Yuan, D.; Jin, C.; Wang, G.; Li, X.; Guan, C. Enhancement of growth and salt tolerance of rice seedlings (Oryza sativa L.) by regulating ethylene production with a novel halotolerant PGPR strain Glutamicibacter sp. YD01 containing ACC deaminase activity. Acta Physiol. Plant. 2020, 42, 42. [Google Scholar] [CrossRef]
- Kumar, S.; Chandra, R.; Keswani, C.; Minkina, T.; Mandzhieva, S.; Voloshina, M.; Meena, M. Trichoderma viride—Mediated modulation of oxidative stress network in potato challenged with Alternaria solani. J. Plant Growth Regul. 2022, 1–18. [Google Scholar] [CrossRef]
- Kumar, A.; Singh, S.; Mukherjee, A.; Rastogi, R.P.; Verma, J.P. Salt-tolerant plant growth-promoting Bacillus pumilus strain JPVS11 to enhance plant growth attributes of rice and improve soil health under salinity stress. Microbiol. Res. 2021, 242, 126616. [Google Scholar] [CrossRef]
- Jain, A.; Singh, A.; Singh, S.; Singh, H.B. Microbial consortium-induced changes in oxidative stress markers in pea plants challenged with Sclerotinia sclerotiorum. J. Plant Growth Regul. 2013, 32, 388–398. [Google Scholar] [CrossRef]
- Singh, S.P.; Gupta, R.; Gaur, R.; Srivastava, A.K. Streptomyces spp. alleviate Rhizoctonia solani-mediated oxidative stress in Solanum lycopersicon. Ann. Appl. Biol. 2016, 168, 232–242. [Google Scholar] [CrossRef]
- Islam, F.; Yasmeen, T.; Arif, M.S.; Ali, S.; Ali, B.; Hameed, S.; Zhou, W. Plant growth promoting bacteria confer salt tolerance in Vigna radiata by up-regulating antioxidant defense and biological soil fertility. Plant Growth Regul. 2016, 80, 23–36. [Google Scholar] [CrossRef]
- Hashem, A.; Alqarawi, A.A.; Radhakrishnan, R.; Al-Arjani, A.-B.F.; Aldehaish, H.A.; Egamberdieva, D.; Abd Allah, E.F. Arbuscular mycorrhizal fungi regulate the oxidative system, hormones and ionic equilibrium to trigger salt stress tolerance in Cucumis sativus L. Saudi J. Biol. Sci. 2018, 25, 1102–1114. [Google Scholar] [CrossRef] [PubMed]
- Mierek-Adamska, A.; Kotowicz, K.; Goc, A.; Boniecka, J.; Berdychowska, J.; Dąbrowska, G.B. Potential involvement of rapeseed (Brassica napus L.) metallothioneins in the hydrogen peroxide-induced regulation of seed vigour. J. Agron. Crop Sci. 2019, 205, 598–607. [Google Scholar] [CrossRef]
- Hassinen, V.H.; Tervahauta, A.I.; Schat, H.; Kärenlampi, S.O. Plant metallothioneins-metal chelators with ROS scavenging activity? Plant Biol. 2011, 13, 225–232. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez-Mañero, F.J.; Ramos-Solano, B.; Probanza, A.; Mehouachi, J.; Tadeo, F.R.; Talon, M. The plant-growth-promoting rhizobacteria Bacillus pumilus and Bacillus licheniformis produce high amounts of physiologically active gibberellins. Physiol. Plant 2001, 111, 206–211. [Google Scholar] [CrossRef]
- Cassán, F.; Perrig, D.; Sgroy, V.; Masciarelli, O.; Penna, C.; Luna, V. Azospirillum brasilense Az39 and Bradyrhizobium japonicum E109, inoculated singly or in combination, promote seed germination and early seedling growth in corn (Zea mays L.) and soybean (Glycine max L.). Eur. J. Soil Biol. 2009, 45, 28–35. [Google Scholar] [CrossRef]
- Perrig, D.; Boiero, M.L.; Masciarelli, O.A.; Penna, C.; Ruiz, O.A.; Cassán, F.D.; Luna, M.V. Plant-growth-promoting compounds produced by two agronomically important strains of Azospirillum brasilense, and implications for inoculant formulation. Appl. Microbiol. Biotechnol. 2007, 75, 1143–1150. [Google Scholar] [CrossRef]
- Timmusk, S.; Nicander, B.; Granhall, U.; Tillberg, E. Cytokinin production by Paenibacillus polymyxa. Soil Biol. Biochem. 1999, 31, 1847–1852. [Google Scholar] [CrossRef]
- García de Salamone, I.E.; Hynes, R.K.; Nelson, L.M. Cytokinin production by plant growth promoting rhizobacteria and selected mutants. Can. J. Microbiol. 2011, 47, 404–411. [Google Scholar] [CrossRef]
- Sorokan, A.; Veselova, S.; Benkovskaya, G.; Maksimov, I. Endophytic strain Bacillus subtilis 26D increases levels of phytohormones and repairs growth of potato plants after Colorado potato beetle damage. Plants 2021, 10, 923. [Google Scholar] [CrossRef]
- Park, Y.-G.; Mun, B.-G.; Kang, S.-M.; Hussain, A.; Shahzad, R.; Seo, C.-W.; Kim, A.-Y.; Lee, S.-U.; Oh, K.Y.; Lee, D.Y.; et al. Bacillus aryabhattai SRB02 tolerates oxidative and nitrosative stress and promotes the growth of soybean by modulating the production of phytohormones. PLoS ONE 2017, 12, e0173203. [Google Scholar] [CrossRef]
- Dixit, R.; Agrawal, L.; Gupta, S.; Kumar, M.; Yadav, S.; Chauhan, P.S.; Nautiyal, C.S. Southern blight disease of tomato control by 1-aminocyclopropane-1-carboxylate (ACC) deaminase producing Paenibacillus lentimorbus B-30488. Plant Signal. Behav. 2016, 11, e1113363. [Google Scholar] [CrossRef] [PubMed]
- Nascimento, F.X.; Vicente, C.S.L.; Barbosa, P.; Espada, M.; Glick, B.R.; Mota, M.; Oliveira, S. Evidence for the involvement of ACC deaminase from Pseudomonas putida UW4 in the biocontrol of pine wilt disease caused by Bursaphelenchus xylophilus. BioControl 2013, 58, 427–433. [Google Scholar] [CrossRef]
- Ravanbakhsh, M.; Sasidharan, R.; Voesenek, L.A.C.J.; Kowalchuk, G.A.; Jousset, A. ACC deaminase-producing rhizosphere bacteria modulate plant responses to flooding. J. Ecol. 2017, 105, 979–986. [Google Scholar] [CrossRef]
- Carlos, M.-H.J.; Stefani, P.-V.Y.; Janette, A.-M.; Melani, M.-S.S.; Gabriela, P.-O. Assessing the effects of heavy metals in ACC deaminase and IAA production on plant growth-promoting bacteria. Microbiol. Res. 2016, 188–189, 53–61. [Google Scholar] [CrossRef] [PubMed]
- Lindström, K.; Mousavi, S.A. Effectiveness of nitrogen fixation in rhizobia. Microb. Biotechnol. 2020, 13, 1314–1335. [Google Scholar] [CrossRef]
- Yang, J.; Kloepper, J.W.; Ryu, C.-M. Rhizosphere bacteria help plants tolerate abiotic stress. Trends Plant. Sci. 2009, 14, 1–4. [Google Scholar] [CrossRef]
- Figueiredo, M.V.B.; Burity, H.A.; Martínez, C.R.; Chanway, C.P. Alleviation of drought stress in the common bean (Phaseolus vulgaris L.) by co-inoculation with Paenibacillus polymyxa and Rhizobium tropici. Appl. Soil Ecol. 2008, 40, 182–188. [Google Scholar] [CrossRef]
- Philippot, L.; Raaijmakers, J.M.; Lemanceau, P.; van der Putten, W.H. Going back to the roots: The microbial ecology of the rhizosphere. Nat. Rev. Microbiol. 2013, 11, 789–799. [Google Scholar] [CrossRef]
- Kafle, A.; Cope, K.; Raths, R.; Krishna Yakha, J.; Subramanian, S.; Bücking, H.; Garcia, K. Harnessing soil microbes to improve plant phosphate efficiency in cropping systems. Agronomy 2019, 9, 127. [Google Scholar] [CrossRef]
- Linu, M.S.; Stephen, J.; Jisha, M.S. Phosphate solubilizing Gluconacetobacter sp., Burkholderia sp. and their potential interaction with cowpea (Vigna unguiculata (L.) Walp.). Int. J. Agric. Res. 2009, 4, 79–87. [Google Scholar] [CrossRef]
- Selvakumar, G.; Mohan, M.; Kundu, S.; Gupta, A.D.; Joshi, P.; Nazim, S.; Gupta, H.S. Cold tolerance and plant growth promotion potential of Serratia marcescens strain SRM (MTCC 8708) isolated from flowers of summer squash (Cucurbita pepo). Lett. Appl. Microbiol. 2008, 46, 171–175. [Google Scholar] [CrossRef] [PubMed]
- Parmar, P.; Sindhu, S.S. Potassium solubilization by rhizosphere bacteria: Influence of nutritional and environmental conditions. J. Microbiol. Res. 2013, 3, 25–31. [Google Scholar]
- Gouda, S.; Kerry, R.G.; Das, G.; Paramithiotis, S.; Shin, H.S.; Patra, J.K. Revitalization of plant growth promoting rhizobacteria for sustainable development in agriculture. Microbiol. Res. 2018, 206, 131–140. [Google Scholar] [CrossRef] [PubMed]
- Ali, A.; Javaid, A.; Shoaib, A.; Khan, I.H. Effect of soil amendment with Chenopodium album dry biomass and two Trichoderma species on growth of chickpea var. Noor 2009 in Sclerotium rolfsii contaminated soil. Egypt. J. Biol. Pest Control 2020, 30, 102. [Google Scholar] [CrossRef]
- Xiao, Y.; Wang, X.; Chen, W.; Huang, Q. Isolation and identification of three potassium-solubilizing bacteria from rape rhizospheric soil and their effects on ryegrass. Geomicrobiol. J. 2017, 34, 873–880. [Google Scholar] [CrossRef]
- Basak, B.B.; Biswas, D.R. Influence of potassium solubilizing microorganism (Bacillus mucilaginosus) and waste mica on potassium uptake dynamics by sudan grass (Sorghum vulgare Pers.) grown under two Alfisols. Plant Soil 2009, 317, 235–255. [Google Scholar] [CrossRef]
- Raji, M.; Thangavelu, M. Isolation and screening of potassium solubilizing bacteria from saxicolous habitat and their impact on tomato growth in different soil types. Arch. Microbiol. 2021, 203, 3147–3161. [Google Scholar] [CrossRef]
- Bityutskii, N.; Yakkonen, K.; Petrova, A.; Nadporozhskaya, M. Xylem sap mineral analyses as a rapid method for estimation plant-availability of Fe, Zn and Mn in carbonate soils: A case study in cucumber. J. Soil Sci. Plant Nutr. 2017, 17, 279–290. [Google Scholar] [CrossRef][Green Version]
- Colombo, C.; Palumbo, G.; He, J.Z.; Pinton, R.; Cesco, S. Review on iron availability in soil: Interaction of Fe minerals, plants, and microbes. J. Soils Sediments 2014, 14, 538–548. [Google Scholar] [CrossRef]
- Rani, N.; Kaur, S.; Nitu, R.; Rajinder, K.; Sukhminderjit, K. Zinc solubilizing bacteria to augment soil fertility—A comprehensive review. Int. Microbiol. 2020, 8, 38–44. [Google Scholar]
- Li, R.-X.; Cai, F.; Pang, G.; Shen, Q.-R.; Li, R.; Chen, W. Solubilisation of phosphate and micronutrients by Trichoderma harzianum and its relationship with the promotion of tomato plant growth. PLoS ONE 2015, 10, e0130081. [Google Scholar] [CrossRef] [PubMed]
- Singh, D.; Geat, N.; Rajawat, M.V.S.; Prasanna, R.; Kar, A.; Singh, A.M.; Saxena, A.K. Prospecting endophytes from different Fe or Zn accumulating wheat genotypes for their influence as inoculants on plant growth, yield, and micronutrient content. Ann. Microbiol. 2018, 68, 815–833. [Google Scholar] [CrossRef]
- Öǧüt, M.; Er, F. Micronutrient composition of field-grown dry bean and wheat inoculated with Azospirillum and Trichoderma. J. Plant Nutr. Soil Sci. 2006, 169, 699–703. [Google Scholar] [CrossRef]
- Durán, P.; Acuña, J.J.; Armada, E.; López-Castillo, O.M.; Cornejo, P.; Mora, M.L.; Azcón, R. Inoculation with selenobacteria and arbuscular mycorrhizal fungi to enhance selenium content in lettuce plants and improve tolerance against drought stress. J. Soil Sci. Plant Nutr. 2016, 16, 211–225. [Google Scholar] [CrossRef]
- De Valença, A.W.; Bake, A.; Brouwer, I.D.; Giller, K.E. Agronomic biofortification of crops to fight hidden hunger in sub-Saharan Africa. Glob. Food Sec. 2017, 12, 8–14. [Google Scholar] [CrossRef]
- Díaz-Gómez, J.; Twyman, R.M.; Zhu, C.; Farré, G.; Serrano, J.C.; Portero-Otin, M.; Muñoz, P.; Sandmann, G.; Capell, T.; Christou, P. Biofortification of crops with nutrients: Factors affecting utilization and storage. Curr. Opin. Biotechnol. 2017, 44, 115–123. [Google Scholar] [CrossRef]
- Khush, G.S.; Lee, S.; Cho, J.I.; Jeon, J.S. Biofortification of crops for reducing malnutrition. Plant Biotechnol. Rep. 2012, 6, 195–202. [Google Scholar] [CrossRef]
- Cabrefiga, J.; Bonaterra, A.; Montesinos, E. Mechanisms of antagonism of Pseudomonas fluorescens EPS62e against Erwinia amylovora, the casual agent of fire blight. Int. Microbiol. 2007, 10, 123132. [Google Scholar]
- Kinsella, K.; Schulthess, C.P.; Morris, T.F.; Stuart, J.D. Rapid quantification of Bacillus subtilis antibiotics in the rhizosphere. Soil Biol. Biochem. 2009, 41, 374–379. [Google Scholar] [CrossRef]
- Bais, H.P.; Fall, R.; Vivanco, J.M. Biocontrol of Bacillus subtilis against infection of Arabidopsis roots by Pseudomonas syringae is facilitated by biofilm formation and surfactin production. Plant Physiol. 2004, 134, 307–319. [Google Scholar] [CrossRef]
- Liu, X.; Bimerew, M.; Ma, Y.; Müller, H.; Ovadis, M.; Eberl, L.; Berg, G.; Chernin, L. Quorum-sensing signaling is required for production of the antibiotic pyrrolnitrin in a rhizospheric biocontrol strain of Serratia plymuthica. FEMS Microbiol. Lett. 2007, 270, 299–305. [Google Scholar] [CrossRef] [PubMed]
- Kurze, S.; Bahl, H.; Dahl, R.; Berg, G. Biological control of fungal strawberry diseases by Serratia plymuthica HRO-C48. Plant Dis. 2001, 85, 529–534. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Wang, J.; Yang, N.; Wen, Z.; Sun, X.; Chai, Y.; Ma, Z. Wheat microbiome bacteria can reduce virulence of a plant pathogenic fungus by altering histone acetylation. Nat. Commun. 2018, 9, 3429. [Google Scholar] [CrossRef] [PubMed]
- Du Jardin, P. Plant biostimulants: Definition, concept, main categories and regulation. Sci. Hortic. 2015, 196, 3–14. [Google Scholar] [CrossRef]
- Mandal, S.; Mitra, A. Reinforcement of cell wall in roots of Lycopersicon esculentum through induction of phenolic compounds and lignin by elicitors. Physiol. Mol. Plant Pathol. 2007, 71, 201–209. [Google Scholar] [CrossRef]
- Egusa, M.; Matsui, H.; Urakami, T.; Okuda, S.; Ifuku, S.; Nakagami, H.; Kaminaka, H. Chitin nanofiber elucidates the elicitor activity of polymeric chitin in plants. Front. Plant Sci. 2015, 6, 1098. [Google Scholar] [CrossRef] [PubMed]
- Proietti, S.; Giangrande, C.; Amoresano, A.; Pucci, P.; Molinaro, A.; Bertini, L.; Caporale, C.; Caruso, C. Xanthomonas campestris lipooligosaccharides trigger innate immunity and oxidative burst in Arabidopsis. Plant Physiol. Biochem. 2014, 85, 51–62. [Google Scholar] [CrossRef][Green Version]
- Nowak, A.; Tyśkiewicz, R.; Wiater, A.; Jaroszuk-ściseł, J. (1→3)-α-D-glucooligosaccharides as elicitors influencing the activity of plant resistance pathways in wheat tissues. Agronomy 2022, 12, 1170. [Google Scholar] [CrossRef]
- Ramkissoon, A.; Francis, J.; Bowrin, V.; Ramjegathesh, R.; Ramsubhag, A.; Jayaraman, J. Bio-efficacy of a chitosan based elicitor on Alternaria solani and Xanthomonas vesicatoria infections in tomato under tropical conditions. Ann. Appl. Biol. 2016, 169, 274–283. [Google Scholar] [CrossRef]
- Qiu, H.; Su, L.; Wang, H.; Zhang, Z. Chitosan elicitation of saponin accumulation in Psammosilene tunicoides hairy roots by modulating antioxidant activity, nitric oxide production and differential gene expression. Plant Physiol. Biochem. 2021, 166, 115–127. [Google Scholar] [CrossRef]
- Aziz, A.; Poinssot, B.; Daire, X.; Adrian, M.; Bézier, A.; Lambert, B.; Joubert, J.M.; Pugin, A. Laminarin elicits defense responses in grapevine and induces protection against Botrytis cinerea and Plasmopara viticola. Mol. Plant Microbe Interact. 2003, 16, 1118–1128. [Google Scholar] [CrossRef] [PubMed]
- Cashel, M.; Gallant, J. Two compounds implicated in the function of the RC gene of Escherichia coli. Nature 1969, 221, 838–841. [Google Scholar] [CrossRef] [PubMed]
- Boniecka, J.; Prusińska, J.; Dąbrowska, G.B.; Goc, A. Within and beyond the stringent response-RSH and (p)ppGpp in plants. Planta 2017, 246, 817–842. [Google Scholar] [CrossRef] [PubMed]
- Dąbrowska, G.; Prusińska, J.; Goc, A. The stringent response—Bacterial mechanism of an adaptive stress response. Adv. Biochem. 2006, 52, 87–93. [Google Scholar]
- Berdychowska, J.; Boniecka, J.; Dąbrowska, G.B. The stringent response and its involvement in the reactions of bacterial cells to stress. Adv. Microbiol. 2019, 58, 127–142. [Google Scholar] [CrossRef]
- Irving, S.E.; Corrigan, R.M.Y.R. Triggering the stringent response: Signals responsible for activating (p)ppGpp synthesis in bacteria. Microbiology 2018, 164, 268–276. [Google Scholar] [CrossRef]
- Irving, S.E.; Choudhury, N.R.; Corrigan, R.M. The stringent response and physiological roles of (pp)pGpp in bacteria. Nat. Rev. Microbiol. 2021, 19, 256–271. [Google Scholar] [CrossRef]
- Sun, D.; Lee, G.; Lee, J.H.; Kim, H.Y.; Rhee, H.W.; Park, S.Y.; Kim, K.J.; Kim, Y.; Kim, B.Y.; Hong, J.I.; et al. A metazoan ortholog of SpoT hydrolyzes ppGpp and functions in starvation responses. Nat. Struct. Mol. Biol. 2010, 17, 1188–1194. [Google Scholar] [CrossRef]
- Sanchez-Vazquez, P.; Dewey, C.N.; Kitten, N.; Ross, W.; Gourse, R.L. Genome-wide effects on Escherichia coli transcription from ppGpp binding to its two sites on RNA polymerase. Proc. Natl. Acad. Sci. USA 2019, 116, 8310–8319. [Google Scholar] [CrossRef]
- Seyfzadeh, M.; Keener, J.; Nomura, M. spoT-dependent accumulation of guanosine tetraphosphate in response to fatty acid starvation in Escherichia coli. Proc. Natl. Acad. Sci. USA 1993, 90, 11004–11008. [Google Scholar] [CrossRef]
- Battesti, A.; Bouveret, E. Acyl carrier protein/SpoT interaction, the switch linking SpoT-dependent stress response to fatty acid metabolism. Mol. Microbiol. 2006, 62, 1048–1063. [Google Scholar] [CrossRef] [PubMed]
- Vinella, D.; Albrecht, C.; Cashel, M.; D’Ari, R. Iron limitation induces SpoT-dependent accumulation of ppGpp in Escherichia coli. Mol. Microbiol. 2005, 56, 958–970. [Google Scholar] [CrossRef] [PubMed]
- Flardh, K.; Axberg, T.; Albertson, N.H.; Kjelleberg, S. Stringent control during carbon starvation of marine Vibrio sp. strain S14: Molecular cloning, nucleotide sequence, and deletion of the relA gene. J. Bacteriol. 1994, 176, 5949–5957. [Google Scholar] [CrossRef] [PubMed]
- Brown, D.R.; Barton, G.; Pan, Z.; Buck, M.; Wigneshweraraj, S. Nitrogen stress response and stringent response are coupled in Escherichia coli. Nat. Commun. 2014, 5, 4115. [Google Scholar] [CrossRef] [PubMed]
- Sivapragasam, S.; Deochand, D.K.; Meariman, J.K.; Grove, A. The stringent response induced by phosphate limitation promotes purine salvage in Agrobacterium fabrum. Biochemistry 2017, 56, 5831–5843. [Google Scholar] [CrossRef] [PubMed]
- English, B.P.; Hauryliuk, V.; Sanamrad, A.; Tankov, S.; Dekker, N.H.; Elf, J. Single-molecule investigations of the stringent response machinery in living bacterial cells. Proc. Natl. Acad. Sci. USA 2011, 108, E365–E373. [Google Scholar] [CrossRef]
- Gallant, J.; Palmer, L.; Pao, C.C. Anomalous synthesis of ppGpp in growing cells. Cell 1977, 11, 181–185. [Google Scholar] [CrossRef]
- Thackray, P.D.; Moir, A. SigM, an extracytoplasmic function sigma factor of Bacillus subtilis, is activated in response to cell wall antibiotics, ethanol, heat, acid, and superoxide stress. J. Bacteriol. 2003, 185, 3491–3498. [Google Scholar] [CrossRef]
- Anderson, K.L.; Roux, C.M.; Olson, M.W.; Luong, T.T.; Lee, C.Y.; Olson, R.; Dunman, P.M. Characterizing the effects of inorganic acid and alkaline shock on the Staphylococcus aureus transcriptome and messenger RNA turnover. FEMS Immunol. Med. Microbiol. 2010, 60, 208–250. [Google Scholar] [CrossRef]
- Heizmann, P.; Howell, S.H. Synthesis of ppGpp and chloroplast ribosomal RNA in Chlamydomonas reinhardi. Biochim. Biophys. Acta 1978, 517, 115–124. [Google Scholar] [CrossRef]
- Van der Biezen, E.A. Arabidopsis RelA/SpoT homologs implicate (p)ppGpp in plant signaling. Proc. Natl. Acad. Sci. USA 2000, 97, 3747–3752. [Google Scholar] [CrossRef] [PubMed]
- Givens, R.M.; Lin, M.-H.; Taylor, D.J.; Mechold, U.; Berry, J.O.; Hernandez, V.J. Inducible expression, enzymatic activity, and origin of higher plant homologues of bacterial RelA/SpoT stress proteins in Nicotiana tabacum. J. Biol. Chem. 2004, 279, 7495–7504. [Google Scholar] [CrossRef] [PubMed]
- Xiong, L.; Lee, M.W.; Qi, M.; Yang, Y. Identification of defense-related rice genes by suppression subtractive hybridization and differential screening. Mol. Plant Microbe Interact. 2001, 14, 685–692. [Google Scholar] [CrossRef] [PubMed]
- Dąbrowska, G.B.; Prusińska, J.; Goc, A. Identification of the RSH gene cDNA (RelA-SpoT homolog) involved in Pharbitis nil response to stress conditions. Zesz. Probl. Postępów Nauk. Rol. 2006, 509, 343–351. [Google Scholar]
- Yamada, A.; Tsutsumi, K.; Tanimoto, S.; Ozeki, Y. Plant RelA/SpoT homolog confers salt tolerance in Escherichia coli and Saccharomyces cerevisiae. Plant Cell Physiol. 2003, 44, 3–9. [Google Scholar] [CrossRef]
- Kim, T.-H.; Ok, S.H.; Kim, D.; Suh, S.-C.; Byun, M.O.; Shin, J.S. Molecular characterization of a biotic and abiotic stress resistance-related gene RelA/SpoT homologue (PepRSH) from pepper. Plant Sci. 2009, 176, 635–642. [Google Scholar] [CrossRef]
- Sato, M.; Takahashi, T.; Ochi, K.; Matsuura, H.; Nabeta, K.; Takahashi, K. Overexpression of RelA/SpoT homologs, PpRSH2a and PpRSH2b, induces the growth suppression of the moss Physcomitrella patens. Biosci. Biotechnol. Biochem. 2015, 79, 36–44. [Google Scholar] [CrossRef]
- Mizusawa, K.; Masuda, S.; Ohta, H. Expression profiling of four RelA/SpoT-like proteins, homologues of bacterial stringent factors, in Arabidopsis thaliana. Planta 2008, 228, 553–562. [Google Scholar] [CrossRef]
- Atkinson, G.C.; Tenson, T.; Hauryliuk, V. The RelA/SpoT Homolog (RSH) superfamily: Distribution and functional evolution of ppGpp synthetases and hydrolases across the tree of life. PLoS ONE 2011, 6, e23479. [Google Scholar] [CrossRef]
- Shin, J.; Singal, B.; Grüber, A.; Wong, D.M.K.; Ragunathan, P.; Grüber, G. Atomic structure of the regulatory TGS domain of Rel protein from Mycobacterium tuberculosis and its interaction with deacylated tRNA. FEBS Lett. 2021, 595, 3006–3018. [Google Scholar] [CrossRef]
- Tozawa, Y.; Nozawa, A.; Kanno, T.; Narisawa, T.; Masuda, S.; Kasai, K.; Nanamiya, H. Calcium-activated (p)ppGpp synthetase in chloroplasts of land plants. J. Biol. Chem. 2007, 282, 35536–35545. [Google Scholar] [CrossRef] [PubMed]
- Ito, K.; Ito, D.; Goto, M.; Suzuki, S.; Masuda, S.; Iba, K.; Kusumi, K. Regulation of ppGpp synthesis and its impact on chloroplast biogenesis during early leaf development in rice. Plant Cell Physiol. 2022, 63, 919–931. [Google Scholar] [CrossRef] [PubMed]
- Yamburenko, M.V.; Zubo, Y.O.; Börner, T. Abscisic acid affects transcription of chloroplast genes via protein phosphatase 2C-dependent activation of nuclear genes: Repression by guanosine-3′-5′-bisdiphosphate and activation by sigma factor 5. Plant J. 2015, 82, 1030–1041. [Google Scholar] [CrossRef] [PubMed]
- Kasai, K.; Kanno, T.; Endo, Y.; Wakasa, K.; Tozawa, Y. Guanosine tetra- and pentaphosphate synthase activity in chloroplasts of a higher plant: Association with 70S ribosomes and inhibition by tetracycline. Nucleic Acids Res. 2004, 32, 5732–5741. [Google Scholar] [CrossRef]
- Imamura, S.; Nomura, Y.; Takemura, T.; Pancha, I.; Taki, K.; Toguchi, K.; Tozawa, Y.; Tanaka, K. The checkpoint kinase TOR (target of rapamycin) regulates expression of a nuclear-encoded chloroplast RelA-SpoT homolog (RSH) and modulates chloroplast ribosomal RNA synthesis in a unicellular red alga. Plant J. 2018, 94, 327–339. [Google Scholar] [CrossRef]
- Sugliani, M.; Abdelkefi, H.; Ke, H.; Bouveret, E.; Robaglia, C.; Caffarri, S.; Field, B. An ancient bacterial signaling pathway regulates chloroplast function to influence growth and development in Arabidopsis. Plant Cell 2016, 28, 661–679. [Google Scholar] [CrossRef]
- Abdelkefi, H.; Sugliani, M.; Ke, H.; Harchouni, S.; Soubigou-Taconnat, L.; Citerne, S.; Mouille, G.; Fakhfakh, H.; Robaglia, C.; Field, B. Guanosine tetraphosphate modulates salicylic acid signalling and the resistance of Arabidopsis thaliana to turnip mosaic virus. Mol. Plant Pathol. 2018, 19, 634–646. [Google Scholar] [CrossRef]
- Kahlau, S.; Bock, R. Plastid transcriptomics and translatomics of tomato fruit development and chloroplast-to-chromoplast differentiation: Chromoplast gene expression largely serves the production of a single protein. Plant Cell 2008, 20, 856–874. [Google Scholar] [CrossRef]
- Maekawa, M.; Honoki, R.; Ihara, Y.; Sato, R.; Oikawa, A.; Kanno, Y.; Ohta, H.; Seo, M.; Saito, K.; Masuda, S. Impact of the plastidial stringent response in plant growth and stress responses. Nat. Plants 2015, 1, 15167. [Google Scholar] [CrossRef]
- Honoki, R.; Ono, S.; Oikawa, A.; Saito, K.; Masuda, S. Significance of accumulation of the alarmone (p)ppGpp in chloroplasts for controlling photosynthesis and metabolite balance during nitrogen starvation in Arabidopsis. Photosynth. Res. 2018, 135, 299–308. [Google Scholar] [CrossRef]
- Romand, S.; Abdelkefi, H.; Lecampion, C.; Belaroussi, M.; Dussenne, M.; Ksas, B.; Citerne, S.; Caius, J.; D’alessandro, S.; Fakhfakh, H.; et al. A guanosine tetraphosphate (ppGpp) mediated brake on photosynthesis is required for acclimation to nitrogen limitation in Arabidopsis. eLife 2022, 11, 75041. [Google Scholar] [CrossRef]
- Li, H.; Nian, J.; Fang, S.; Guo, M.; Huang, X.; Zhang, F.; Wang, Q.; Zhang, J.; Bai, J.; Dong, G.; et al. Regulation of nitrogen starvation responses by the alarmone (p)ppGpp in rice. J. Genet. Genom. 2022, 49, 469–480. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K.; Kasai, K.; Ochi, K. Identification of the bacterial alarmone guanosine 5′-diphosphate 3′-diphosphate (ppGpp) in plants. Proc. Natl. Acad. Sci. USA 2004, 101, 4320–4324. [Google Scholar] [CrossRef] [PubMed]
- Masuda, S.; Mizusawa, K.; Narisawa, T.; Tozawa, Y.; Ohta, H.; Takamiya, K.I. The bacterial stringent response, conserved in chloroplasts, controls plant fertilization. Plant Cell Physiol. 2008, 49, 135–141. [Google Scholar] [CrossRef] [PubMed]
- Ono, S.; Suzuki, S.; Ito, D.; Tagawa, S.; Shiina, T.; Masuda, S. Plastidial (p)ppGpp synthesis by the Ca2+-dependent RelA–SpoT homolog regulates the adaptation of chloroplast gene expression to darkness in Arabidopsis. Plant Cell Physiol. 2021, 61, 2077–2086. [Google Scholar] [CrossRef]
- Nowicki, D.; Rodzik, O.; Herman-Antosiewicz, A.; Szalewska-Pałasz, A. Isothiocyanates as effective agents against enterohemorrhagic Escherichia coli: Insight to the mode of action. Sci. Rep. 2016, 6, 22263. [Google Scholar] [CrossRef]
- Mi, L.; Hood, B.L.; Stewart, N.A.; Xiao, Z.; Govind, S.; Wang, X.; Conrads, T.P.; Veenstra, T.D.; Chung, F.-L. Identification of potential protein targets of isothiocyanates by proteomics. Chem. Res. Toxicol. 2011, 24, 1735–1743. [Google Scholar] [CrossRef]
- Mwita, L.; Chan, W.Y.; Pretorius, T.; Lyantagaye, S.L.; Lapa, S.V.; Avdeeva, L.V.; Reva, O.N. Gene expression regulation in the plant growth promoting Bacillus atrophaeus UCMB-5137 stimulated by maize root exudates. Gene 2016, 590, 18–28. [Google Scholar] [CrossRef]
- Dąbrowska, G.B.; Turkan, S.; Tylman-Mojżeszek, W.; Mierek-Adamska, A. In silico study of the RSH (Rela/spot homologs) gene family and expression analysis in response to PGPR bacteria and salinity in Brassica napus. Int. J. Mol. Sci. 2021, 22, 10666. [Google Scholar] [CrossRef]
- Chapelle, E.; Mendes, R.; Bakker, P.A.H.; Raaijmakers, J.M. Fungal invasion of the rhizosphere microbiome. ISME J. 2016, 10, 265–268. [Google Scholar] [CrossRef]
- Manuel, J.; Berry, C.; Selin, C.; Fernando, W.G.D.; de Kievit, T.R. Repression of the antifungal activity of Pseudomonas sp. strain DF41 by the stringent response. Appl. Environ. Microbiol. 2011, 77, 5635–5642. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Manuel, J.; Selin, C.; Dilantha Fernando, W.G.; de Kievi, T. Stringent response mutants of Pseudomonas chlororaphis PA23 exhibit enhanced antifungal activity against Sclerotinia sclerotiorum in vitro. Microbiology 2012, 158, 207–216. [Google Scholar] [CrossRef] [PubMed]
- Selin, C.; Manuel, J.; Fernando, W.G.D.; de Kievit, T. Expression of the Pseudomonas chlororaphis strain PA23 Rsm system is under control of GacA, RpoS, PsrA, quorum sensing and the stringent response. Biol. Control 2014, 69, 24–33. [Google Scholar] [CrossRef]
- Takeuchi, K.; Yamada, K.; Haas, D. ppGpp controlled by the Gac/Rsm regulatory pathway sustains biocontrol activity in Pseudomonas fluorescens CHA0. Mol. Plant Microbe Interact. 2012, 25, 1440–1449. [Google Scholar] [CrossRef] [PubMed]
- Ochi, K. A rel mutation abolishes the enzyme induction needed for actinomycin synthesis by Streptomyces antibioticus. Agric. Biol. Chem. 2014, 51, 829–835. [Google Scholar]
- Ryu, Y.G.; Wook, J.; Jin, Y.K.; Jae, Y.K.; Sang, H.L.; Kye, J.L. Stringent factor regulates antibiotics production and morphological differentiation of Streptomyces clavuligerus. J. Microbiol. Biotechnol. 2004, 14, 1170–1175. [Google Scholar]
- Brechenmacher, L.; Lei, Z.; Libault, M.; Findley, S.; Sugawara, M.; Sadowsky, M.J.; Sumner, L.W.; Stacey, G. Soybean metabolites regulated in root hairs in response to the symbiotic bacterium Bradyrhizobium japonicum. Plant Physiol. 2010, 153, 1808–1822. [Google Scholar] [CrossRef]
- Murray, J.D. Invasion by invitation: Rhizobial infection in legumes. Mol. Plant Microbe Interact. 2011, 24, 631–639. [Google Scholar] [CrossRef]
- Tsyganova, A.V.; Brewin, N.J.; Tsyganov, V.E. Structure and development of the legume-rhizobial symbiotic interface in infection threads. Cells 2021, 10, 1050. [Google Scholar] [CrossRef]
- Pérez-Giménez, J.; Iturralde, E.T.; Torres Tejerizo, G.; Quelas, J.I.; Krol, E.; Borassi, C.; Becker, A.; Estevez, J.M.; Lodeiro, A.R. A stringent-response-defective Bradyrhizobium diazoefficiens strain does not activate the type 3 secretion system, elicits an early plant defense response, and circumvents NH4NO3 -induced inhibition of nodulation. Appl. Environ. Microbiol. 2021, 87, e02989-20. [Google Scholar] [CrossRef]
- Wippel, K.; Long, S.R. Symbiotic performance of Sinorhizobium meliloti lacking ppGpp depends on the Medicago host species. Mol. Plant Microbe Interact. 2019, 32, 717–728. [Google Scholar] [CrossRef] [PubMed]
- Wells, D.H.; Long, S.R. The Sinorhizobium meliloti stringent response affects multiple aspects of symbiosis. Mol. Microbiol. 2002, 43, 1115–1127. [Google Scholar] [CrossRef] [PubMed]
- Moris, M.; Braeken, K.; Schoeters, E.; Verreth, C.; Beullens, S.; Vanderleyden, J.; Michiels, J. Effective symbiosis between Rhizobium etli and Phaseolus vulgaris requires the alarmone ppGpp. J. Bacteriol. 2005, 187, 5460–5469. [Google Scholar] [CrossRef] [PubMed]
- Calderón-Flores, A.; du Pont, G.; Huerta-Saquero, A.; Merchant-Larios, H.; Servín-González, L.; Durán, S. The stringent response is required for amino acid and nitrate utilization, nod factor regulation, nodulation, and nitrogen fixation in Rhizobium etli. J. Bacteriol. 2005, 187, 5075. [Google Scholar] [CrossRef] [PubMed]
- Qiu, X.; Gao, T.; Yang, J.; Wang, E.; Liu, L.; Yuan, H. Water-soluble humic materials modulating metabolism and triggering stress defense in Sinorhizobium fredii. Microbiol. Spectr. 2021, 9, e00293-21. [Google Scholar] [CrossRef] [PubMed]
- Ancona, V.; Lee, J.H.; Chatnaparat, T.; Oh, J.; Hong, J.I.; Zhao, Y. The bacterial alarmone (p)ppGpp activates the type III secretion system in Erwinia amylovora. J. Bacteriol. 2015, 197, 1433–1443. [Google Scholar] [CrossRef] [PubMed]
- Chatnaparat, T.; Li, Z.; Korban, S.S.; Zhao, Y. The bacterial alarmone (p)ppGpp is required for virulence and controls cell size and survival of Pseudomonas syringae on plants. Environ. Microbiol. 2015, 17, 4253–4270. [Google Scholar] [CrossRef]
- Chatnaparat, T.; Li, Z.; Korban, S.S.; Zhao, Y. The stringent response mediated by (p)ppGpp is required for virulence of Pseudomonas syringae pv. tomato and its survival on tomato. Mol. Plant Microbe Interact. 2015, 28, 776–789. [Google Scholar] [CrossRef]
- Bowden, S.D.; Hale, N.; Chung, J.C.S.; Hodgkinson, J.T.; Spring, D.R.; Welch, M. Surface swarming motility by Pectobacterium atrosepticum is a latent phenotype that requires O antigen and is regulated by quorum sensing. Microbiology 2013, 159, 2375–2385. [Google Scholar] [CrossRef]
- Petrova, O.; Gorshkov, V.; Sergeeva, I.; Daminova, A.; Ageeva, M.; Gogolev, Y. Alternative scenarios of starvation-induced adaptation in Pectobacterium atrosepticum. Res. Microbiol. 2016, 167, 254–261. [Google Scholar] [CrossRef]
- Zhang, Y.; Teper, D.; Xu, J.; Wang, N. Stringent response regulators (p)ppGpp and DksA positively regulate virulence and host adaptation of Xanthomonas citri. Mol. Plant. Pathol. 2019, 20, 1550–1565. [Google Scholar] [CrossRef] [PubMed]
- Harman, G.E.; Howell, C.R.; Viterbo, A.; Chet, I.; Lorito, M. Trichoderma species—Opportunistic, avirulent plant symbionts. Nat. Rev. Microbiol. 2004, 2, 43–56. [Google Scholar] [CrossRef] [PubMed]
- Wojtkowiak-Gębarowska, E. Mechanisms of biological control soil-borne plant pathogen by fungus from genus Trichoderma. Adv. Microbiol. 2006, 45, 261–273. [Google Scholar]
- Bustamante, D.E.; Calderon, M.S.; Leiva, S.; Mendoza, J.E.; Arce, M.; Oliva, M. Three new species of Trichoderma in the Harzianum and Longibrachiatum lineages from Peruvian cacao crop soils based on an integrative approach. Mycologia 2021, 113, 1056–1072. [Google Scholar] [CrossRef]
- Pascale, A.; Vinale, F.; Manganiello, G.; Nigro, M.; Lanzuise, S.; Ruocco, M.; Marra, R.; Lombardi, N.; Woo, S.L.; Lorito, M. Trichoderma and its secondary metabolites improve yield and quality of grapes. Crop Prot. 2017, 92, 176–181. [Google Scholar] [CrossRef]
- Srivastava, S.N.; Singh, V.; Awasthi, S.K. Trichoderma induced improvement in growth, yield and quality of sugarcane. Sugar Tech 2006, 8, 166–169. [Google Scholar] [CrossRef]
- Colla, G.; Rouphael, Y.; di Mattia, E.; El-Nakhel, C.; Cardarelli, M. Co-inoculation of Glomus intraradices and Trichoderma atroviride acts as a biostimulant to promote growth, yield and nutrient uptake of vegetable crops. J. Sci. Food Agric. 2015, 95, 1706–1715. [Google Scholar] [CrossRef]
- Rudresh, D.L.; Shivaprakash, M.K.; Prasad, R.D. Effect of combined application of Rhizobium, phosphate solubilizing bacterium and Trichoderma spp. on growth, nutrient uptake and yield of chickpea (Cicer aritenium L.). Appl. Soil Ecol. 2005, 28, 139–146. [Google Scholar] [CrossRef]
- Khomari, S.; Golshan-Doust, S.; Seyed-Sharifi, R.; Davari, M. Improvement of soybean seedling growth under salinity stress by biopriming of high-vigour seeds with salt-tolerant isolate of Trichoderma harzianum. N. Z. J. Crop Hortic. Sci. 2018, 46, 117–132. [Google Scholar] [CrossRef]
- Asaduzzaman, M.; Alam, M.; Islam, M. Effect of Trichoderma on seed germination and seedling parameters of chili. J. Sci. Found. 2013, 8, 141–150. [Google Scholar] [CrossRef]
- Vargas, W.A.; Mandawe, J.C.; Kenerley, C.M. Plant-derived sucrose is a key element in the symbiotic association between Trichoderma virens and maize plants. Plant Physiol. 2009, 151, 792–808. [Google Scholar] [CrossRef]
- Brotman, Y.; Lisec, J.; Méret, M.; Chet, I.; Willmitzer, L.; Viterbo, A. Transcript and metabolite analysis of the Trichoderma-induced systemic resistance response to Pseudomonas syringae in Arabidopsis thaliana. Microbiology 2012, 158, 139–146. [Google Scholar] [CrossRef] [PubMed]
- Contreras-Cornejo, H.A.; Macías-Rodríguez, L.; Vergara, A.G.; López-Bucio, J. Trichoderma modulates stomatal aperture and leaf transpiration through an abscisic acid-dependent mechanism in Arabidopsis. J. Plant Growth Regul. 2015, 34, 425–432. [Google Scholar] [CrossRef]
- Bae, H.; Sicher, R.C.; Kim, M.S.; Kim, S.-H.; Strem, M.D.; Melnick, R.L.; Bailey, B.A. The beneficial endophyte Trichoderma hamatum isolate DIS 219b promotes growth and delays the onset of the drought response in Theobroma cacao. J. Exp. Bot. 2009, 60, 3279–3295. [Google Scholar] [CrossRef] [PubMed]
- Shoresh, M.; Harman, G.E. The molecular basis of shoot responses of maize seedlings to Trichoderma harzianum T22 inoculation of the root: A proteomic approach. Plant Physiol. 2008, 147, 2147–2163. [Google Scholar] [CrossRef]
- Mercl, F.; García-Sánchez, M.; Kulhánek, M.; Košnář, Z.; Száková, J.; Tlustoš, P. Improved phosphorus fertilisation efficiency of wood ash by fungal strains Penicillium sp. PK112 and Trichoderma harzianum OMG08 on acidic soil. Appl. Soil Ecol. 2020, 147, 103360. [Google Scholar] [CrossRef]
- Fu, J.; Xiao, Y.; Wang, Y.; Liu, Z.; Yang, K. Saline–alkaline stress in growing maize seedlings is alleviated by Trichoderma asperellum through regulation of the soil environment. Sci. Rep. 2021, 11, 11152. [Google Scholar] [CrossRef]
- Zhang, F.; Xu, X.; Wang, G.; Wu, B.; Xiao, Y. Medicago sativa and soil microbiome responses to Trichoderma as a biofertilizer in alkaline-saline soils. Appl. Soil Ecol. 2020, 153, 103573. [Google Scholar] [CrossRef]
- Adams, P.; De-Leij, F.A.A.M.; Lynch, J.M. Trichoderma harzianum Rifai 1295-22 mediates growth promotion of crack willow (Salix fragilis) saplings in both clean and metal-contaminated soil. Microb. Ecol. 2007, 54, 306–313. [Google Scholar] [CrossRef]
- López Errasquín, E.; Vázquez, C. Tolerance and uptake of heavy metals by Trichoderma atroviride isolated from sludge. Chemosphere 2003, 50, 137–143. [Google Scholar] [CrossRef]
- Rajendran, L.; Raja, P.; Jegadeeswari, V.; Santhi, V.P.; Selvaraj, N. Pseudomonas fluorescens and Trichoderma viride enriched bioconsortium for the management of Fusarium wilt in carnation and gerbera under protected cultivation. Indian Phytopathol. 2014, 67, 77–81. [Google Scholar]
- Ambuse, M.G.; Chatage, V.S.; Bhale, U.N. Influence of Trichoderma spp against Alternaria tenuissima incitining leaf spot of Rumex acetosa L. Biosci. Discov. 2012, 3, 259–262. [Google Scholar]
- De Padua, J.C.; de la Cruz, T.E.E. Isolation and characterization of nickel-tolerant Trichoderma strains from marine and terrestrial environments. J. Fungi 2021, 7, 591. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Montesinos, B.; Diánez, F.; Moreno-Gavira, A.; Gea, F.J.; Santos, M. Plant growth promotion and biocontrol of Pythium ultimum by saline tolerant Trichoderma isolates under salinity stress. Int. J. Environ. Res. Public Health 2019, 16, 2053. [Google Scholar] [CrossRef] [PubMed]
- Shi, Z.-Z.; Miao, F.-P.; Fang, S.-T.; Yin, X.-L.; Ji, N.-Y. Sulfurated diketopiperazines from an algicolous isolate of Trichoderma virens. Phytochem. Lett. 2018, 27, 101–104. [Google Scholar] [CrossRef]
- Touati, I.; Ruiz, N.; Thomas, O.; Druzhinina, I.S.; Atanasova, L.; Tabbene, O.; Elkahoui, S.; Benzekri, R.; Bouslama, L.; Pouchus, Y.F.; et al. Hyporientalin A, an anti-Candida peptaibol from a marine Trichoderma orientale. World J. Microbiol. Biotechnol. 2018, 34, 98. [Google Scholar] [CrossRef]
- Zhang, C.; Wang, W.; Hu, Y.; Peng, Z.; Ren, S.; Xue, M.; Liu, Z.; Hou, J.; Xing, M.; Liu, T. A novel salt-tolerant strain Trichoderma atroviride HN082102.1 isolated from marine habitat alleviates salt stress and diminishes cucumber root rot caused by Fusarium oxysporum. BMC Microbiol. 2022, 22, 67. [Google Scholar] [CrossRef]
- Gal-Hemed, I.; Atanasova, L.; Komon-Zelazowska, M.; Druzhinina, I.S.; Viterbo, A.; Yarden, O. Marine isolates of Trichoderma spp. as potential halotolerant agents of biological control for arid-zone agriculture. Appl. Environ. Microbiol. 2011, 77, 5100–5109. [Google Scholar] [CrossRef]
- Nogueira-Lopez, G.; Greenwood, D.R.; Middleditch, M.; Winefield, C.; Eaton, C.; Steyaert, J.M.; Mendoza-Mendoza, A. The apoplastic secretome of Trichoderma virens during interaction with maize roots shows an inhibition of plant defence and scavenging oxidative stress secreted proteins. Front. Plant Sci. 2018, 9, 409. [Google Scholar] [CrossRef]
- Sofo, A.; Scopa, A.; Manfra, M.; de Nisco, M.; Tenore, G.; Troisi, J.; di Fiori, R.; Novellino, E. Trichoderma harzianum strain T-22 induces changes in phytohormone levels in cherry rootstocks (Prunus cerasus × P. canescens). Plant Growth Regul. 2011, 65, 421–425. [Google Scholar] [CrossRef]
- Yedidia, I.; Shoresh, M.; Kerem, Z.; Benhamou, N.; Kapulnik, Y.; Chet, I. Concomitant induction of systemic resistance to Pseudomonas syringae pv. lachrymans in cucumber by Trichoderma asperellum (T-203) and accumulation of phytoalexins. Appl. Environ. Microbiol. 2003, 69, 7343–7353. [Google Scholar] [PubMed]
- Akhter, W.; Bhuiyan, M.K.A.; Sultana, F.; Hossain, M.M. Integrated effect of microbial antagonist, organic amendment and fungicide in controlling seedling mortality (Rhizoctonia solani) and improving yield in pea (Pisum sativum L.). Comptes Rendus. Biol. 2015, 338, 21–28. [Google Scholar] [CrossRef] [PubMed]
- Howell, C.R.; Hanson, L.E.; Stipanovic, R.D.; Puckhaber, L.S. Induction of terpenoid synthesis in cotton roots and control of Rhizoctonia solani by seed treatment with Trichoderma virens. Phytopatology 2000, 90, 248–252. [Google Scholar] [CrossRef] [PubMed]
- Gomes, E.V.; Costa, M.D.N.; de Paula, R.G.; Ricci De Azevedo, R.; da Silva, F.L.; Noronha, E.F.; José Ulhoa, C.; Neves Monteiro, V.; Elena Cardoza, R.; Gutiérrez, S.; et al. The cerato-platanin protein Epl-1 from Trichoderma harzianum is involved in mycoparasitism, plant resistance induction and self cell wall protection. Sci. Rep. 2015, 5, 17998. [Google Scholar] [CrossRef] [PubMed]
- Howell, C.R. Cotton seedling preemergence damping-off incited by Rhizopus oryzae and Pythium spp. and its biological control with Trichoderma spp. Phytopathology 2002, 92, 177–180. [Google Scholar] [CrossRef] [PubMed]
- Hossain, M.M.; Hossain, N.; Sultana, F.; Mohammad, S.; Islam, N.; Khurshed, M.; Bhuiyan, A. Integrated management of Fusarium wilt of chickpea (Cicer arietinum L.) caused by Fusarium oxysporum f. sp. ciceris with microbial antagonist, botanical extract sp. ciceris with microbial antagonist, botanical extract. Afr. J. Biotechnol. 2016, 12, 4699–4706. [Google Scholar]
- Vujanovic, V.; Goh, Y.K. qPCR quantification of sphaerodes mycoparasitica biotrophic mycoparasite interaction with Fusarium graminearum: In vitro and in planta assays. Arch. Microbiol. 2012, 194, 707–717. [Google Scholar] [CrossRef]
- Gveroska, B.; Ziberoski, J. Trichoderma harzianum as a biocontrol agent against Alternaria alternata on tobacco. Appl. Technol. Innov. 2012, 7, 67–76. [Google Scholar] [CrossRef]
- Matroudi, S.; Zamani, M.; Motallebi, M. Antagonistic effects of three species of Trichoderma sp. on Sclerotinia sclerotiorum, the causal agent of canola stem rot. Egypt. Acad. J. Biol. Sci. 2010, 11, 37–44. [Google Scholar]
- De Meyer, G.; Bigirimana, J.; Elad, Y.; Höfte, M. Induced systemic resistance in Trichoderma harzianum T39 biocontrol of Botrytis cinerea. Eur. J. Plant Pathol. 1998, 104, 279–286. [Google Scholar] [CrossRef]
- Herrera-Téllez, V.I.; Cruz-Olmedo, A.K.; Plasencia, J.; Gavilanes-Ruíz, M.; Arce-Cervantes, O.; Hernández-León, S.; Saucedo-García, M. The protective effect of Trichoderma asperellum on tomato plants against Fusarium oxysporum and Botrytis cinerea diseases involves inhibition of reactive oxygen species production. Int. J. Mol. Sci. 2019, 20, 2007. [Google Scholar] [CrossRef] [PubMed]
- Cuervo-Parra, J.A.; Pérez España, V.H.; Zavala-González, E.A.; Peralta-Gil, M.; Aparicio Burgos, J.E.; Romero-Cortes, T. Trichoderma asperellum strains as potential biological control agents against Fusarium verticillioides and Ustilago maydis in maize. Biocontrol. Sci. Technol. 2022, 32, 624–647. [Google Scholar] [CrossRef]
- Daniel, J.F.D.S.; Rodrigues Filho, E. Peptaibols of Trichoderma. Nat. Prod. Rep. 2007, 24, 1128–1141. [Google Scholar] [CrossRef]
- Degenkolb, T.; Berg, A.; Gams, W.; Schlegel, B.; Gräfe, U. The occurrence of peptaibols and structurally related peptaibiotics in fungi and their mass spectrometric identification via diagnostic fragment ions. J. Pept. Sci. 2003, 9, 666–678. [Google Scholar] [CrossRef] [PubMed]
- Shi, M.; Chen, L.; Wang, X.W.; Zhang, T.; Zhao, P.B.; Song, X.Y.; Sun, C.Y.; Chen, X.L.; Zhou, B.C.; Zhang, Y.Z. Antimicrobial peptaibols from Trichoderma pseudokoningii induce programmed cell death in plant fungal pathogens. Microbiology 2012, 158, 166–175. [Google Scholar] [CrossRef]
- Zhang, Y.-Q.; Zhang, S.; Sun, M.-L.; Su, H.-N.; Li, H.-Y.; Liu, K.; Zhang, Y.-Z.; Chen, X.-L.; Cao, H.-Y.; Song, X.-Y. Antibacterial activity of peptaibols from Trichoderma longibrachiatum SMF2 against gram-negative Xanthomonas oryzae pv. oryzae, the causal agent of bacterial leaf blight on rice. Front. Microbiol. 2022, 13, 1034779. [Google Scholar]
- Zhao, P.; Ren, A.; Dong, P.; Sheng, Y.; Li, D. Antimicrobial peptaibols, trichokonins, inhibit mycelial growth and sporulation and induce cell apoptosis in the pathogenic fungus Botrytis cinerea. Appl. Biochem. Microbiol. 2018, 54, 396–403. [Google Scholar] [CrossRef]
- Otto, A.; Laub, A.; Wendt, L.; Porzel, A.; Schmidt, J.; Palfner, G.; Becerra, J.; Krüger, D.; Stadler, M.; Wessjohann, L.; et al. Chilenopeptins A and B, peptaibols from the Chilean Sepedonium aff. chalcipori KSH 883. J. Nat. Prod. 2016, 79, 929–938. [Google Scholar] [CrossRef]
- Tamandegani, P.R.; Marik, T.; Zafari, D.; Balázs, D.; Vágvölgyi, C.; Szekeres, A.; Kredics, L. Changes in peptaibol production of Trichoderma species during in vitro antagonistic interactions with fungal plant pathogens. Biomolecules 2020, 10, 730. [Google Scholar] [CrossRef]
- Luo, Y.; Zhang, D.D.; Dong, X.W.; Zhao, P.B.; Chen, L.L.; Song, X.Y.; Wang, X.J.; Chen, X.L.; Shi, M.; Zhang, Y.Z. Antimicrobial peptaibols induce defense responses and systemic resistance in tobacco against tobacco mosaic virus. FEMS Microbiol. Lett. 2010, 313, 120–126. [Google Scholar] [CrossRef]
- Marik, T.; Tyagi, C.; Balázs, D.; Urbán, P.; Szepesi, Á.; Bakacsy, L.; Endre, G.; Rakk, D.; Szekeres, A.; Andersson, M.A.; et al. Structural diversity and bioactivities of peptaibol compounds from the Longibrachiatum clade of the filamentous fungal genus Trichoderma. Front. Microbiol. 2019, 10, 1434. [Google Scholar] [CrossRef] [PubMed]
- Shoresh, M.; Yedidia, I.; Chet, I. Involvement of jasmonic acid/ethylene signaling pathway in the systemic resistance induced in cucumber by Trichoderma asperellum T203. Phytopathology 2005, 95, 76–84. [Google Scholar] [CrossRef]
- Yedidia, I.; Benhamou, N.; Chet, I. Induction of defense responses in cucumber plants (Cucumis sativus L.) by the biocontrol agent Trichoderma harzianum. Appl. Environ. Microbiol. 1999, 65, 1061–1070. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Chen, L.; Ran, W.; Shen, Q.; Yang, X. Trichoderma harzianum strain SQR-T37 and its bio-organic fertilizer could control Rhizoctonia solani damping-off disease in cucumber seedlings mainly by the mycoparasitism. Appl. Microbiol. Biotechnol. 2011, 91, 741–755. [Google Scholar] [CrossRef] [PubMed]
- Mendoza-Mendoza, A.; Zaid, R.; Lawry, R.; Hermosa, R.; Monte, E.; Horwitz, B.A.; Mukherjee, P.K. Molecular dialogues between Trichoderma and roots: Role of the fungal secretome. Fungal Biol. Rev. 2018, 32, 62–85. [Google Scholar] [CrossRef]
- Brotman, Y.; Landau, U.; Cuadros-Inostroza, Á.; Takayuki, T.; Fernie, A.R.; Chet, I.; Viterbo, A.; Willmitzer, L. Trichoderma-plant root colonization: Escaping early plant defense responses and activation of the antioxidant machinery for saline stress tolerance. PLoS Pathog. 2013, 9, e1003221. [Google Scholar] [CrossRef]
- Engelberth, J.; Koch, T.; Schüler, G.; Bachmann, N.; Rechtenbach, J.; Boland, W. Ion channel-forming alamethicin is a potent elicitor of volatile biosynthesis and tendril coiling. Cross talk between jasmonate and salicylate signaling in lima bean. Plant Physiol. 2001, 125, 369–377. [Google Scholar] [CrossRef]
- Rotblat, B.; Enshell-Seijffers, D.; Gershoni, J.M.; Schuster, S.; Avni, A. Identification of an essential component of the elicitation active site of the EIX protein elicitor. Plant J. 2002, 32, 1049–1055. [Google Scholar] [CrossRef]
- Martinez, C.; Blanc, F.; le Claire, E.; Besnard, O.; Nicole, M.; Baccou, J.-C. Salicylic acid and ethylene pathways are differentially activated in melon cotyledons by active or heat-denatured cellulase from Trichoderma longibrachiatum. Plant Physiol. 2001, 127, 334–344. [Google Scholar] [CrossRef]
- Morán-Diez, E.; Hermosa, R.; Ambrosino, P.; Cardoza, R.E.; Gutiérrez, S.; Lorito, M.; Monte, E. The ThPG1 endopolygalacturonase is required for the Trichoderma harzianum–plant beneficial interaction. Mol. Plant Microbe Interact. 2009, 22, 1021–1031. [Google Scholar] [CrossRef]
- Hermosa, R.; Viterbo, A.; Chet, I.; Monte, E. Plant-beneficial effects of Trichoderma and of its genes. Microbiology 2012, 158, 17–25. [Google Scholar] [CrossRef] [PubMed]
- Alonso-Ramírez, A.; Poveda, J.; Martín, I.; Hermosa, R.; Monte, E.; Nicolás, C. Salicylic acid prevents Trichoderma harzianum from entering the vascular system of roots. Mol. Plant Pathol. 2014, 15, 823–831. [Google Scholar] [CrossRef] [PubMed]
- Ruocco, M.; Lanzuise, S.; Vinale, F.; Marra, R.; Turrà, D.; Woo, S.L.; Lorito, M. Identification of a new biocontrol gene in Trichoderma atroviride: The role of an ABC transporter membrane pump in the interaction with different plant-pathogenic fungi. Mol. Plant Microbe Interact. 2009, 22, 291–301. [Google Scholar] [CrossRef] [PubMed]
- Masunaka, A.; Hyakumachi, M.; Takenaka, S. Plant growth-promoting fungus, Trichoderma koningi suppresses isoflavonoid phytoalexin vestitol production for colonization on/in the roots of Lotus japonicus. Microbes Environ. 2011, 26, 128–134. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Li, J.; Yang, H.; Shin, H.-J. Fungal and mushroom hydrophobins: A review. J. Mushroom 2017, 15, 1–7. [Google Scholar] [CrossRef]
- Dąbrowska, G.B.; Garstecka, Z.; Olewnik-Kruszkowska, E.; Szczepańska, G.; Ostrowski, M.; Mierek-Adamska, A. Comparative study of structural changes of polylactide and poly(ethylene terephthalate) in the presence of Trichoderma viride. Int. J. Mol. Sci. 2021, 22, 3491. [Google Scholar] [CrossRef]
- Viterbo, A.; Harel, M.; Chet, I. Isolation of two aspartyl proteases from Trichoderma asperellum expressed during colonization of cucumber roots. FEMS Microbiol. Lett. 2004, 238, 151–158. [Google Scholar]
- Saloheimo, M.; Paloheimo, M.; Hakola, S.; Pere, J.; Swanson, B.; Nyyssönen, E.; Bhatia, A.; Ward, M.; Penttilä, M. Swollenin, a Trichoderma reesei protein with sequence similarity to the plant expansins, exhibits disruption activity on cellulosic materials. Eur. J. Biochem. 2002, 269, 4202–4211. [Google Scholar] [CrossRef]
- Sánchez-Cruz, R.; Mehta, R.; Atriztán-Hernández, K.; Martínez-Villamil, O.; del Rayo Sánchez-Carbente, M.; Sánchez-Reyes, A.; Lira-Ruan, V.; González-Chávez, C.A.; Tabche-Barrera, M.L.; Bárcenas-Rodríguez, R.C.; et al. Effects on Capsicum annuum plants colonized with Trichoderma atroviride P. Karst strains genetically modified in taswo1, a gene coding for a protein with expansin-like activity. Plants 2021, 10, 1919. [Google Scholar] [CrossRef]
- Macías-Rodríguez, L.; Guzmán-Gómez, A.; García-Juárez, P.; Contreras-Cornejo, H.A. Trichoderma atroviride promotes tomato development and alters the root exudation of carbohydrates, which stimulates fungal growth and the biocontrol of the phytopathogen Phytophthora cinnamomi in a tripartite interaction system. FEMS Microbiol. Ecol. 2018, 94, fiy137. [Google Scholar] [CrossRef]
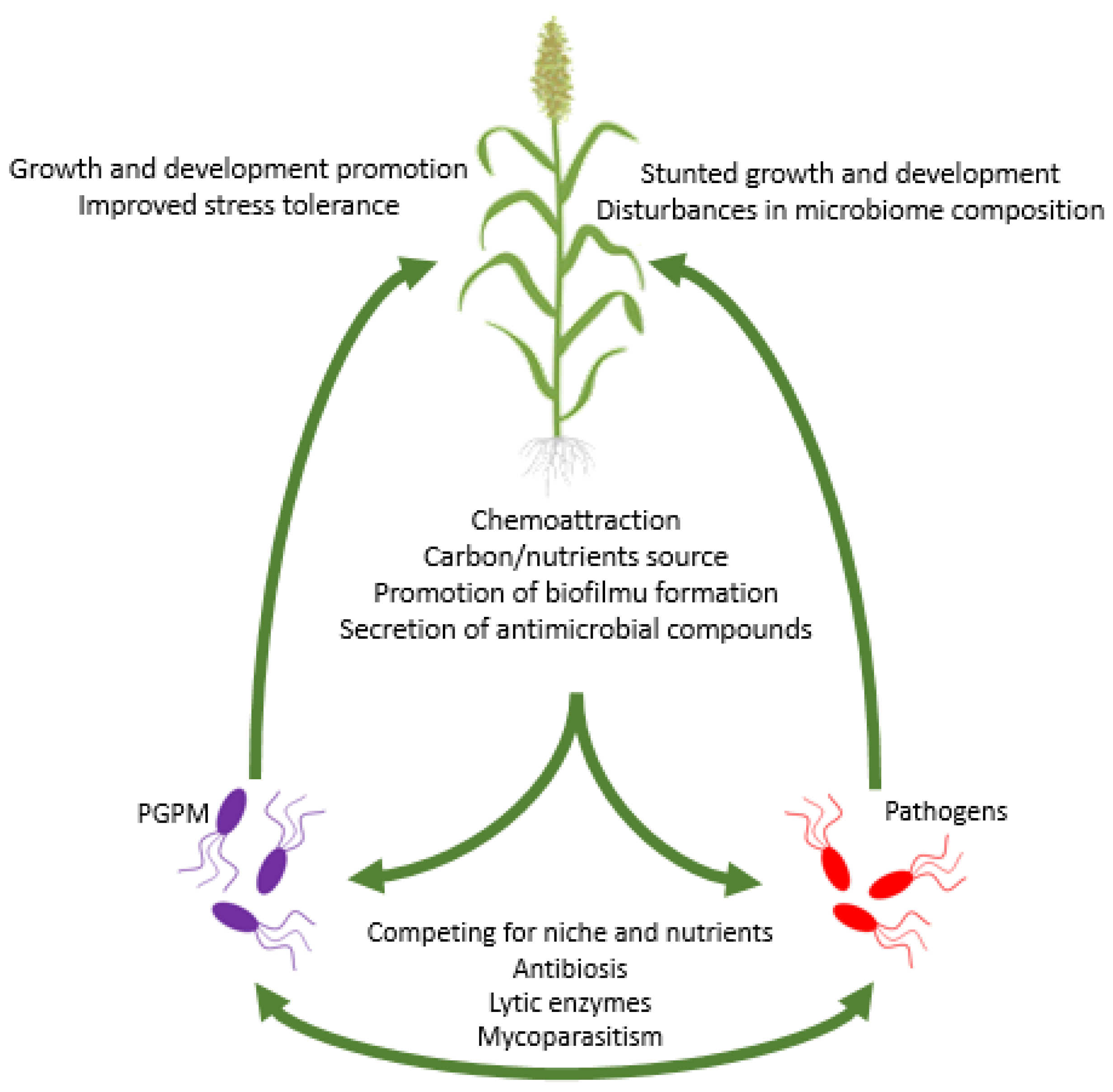
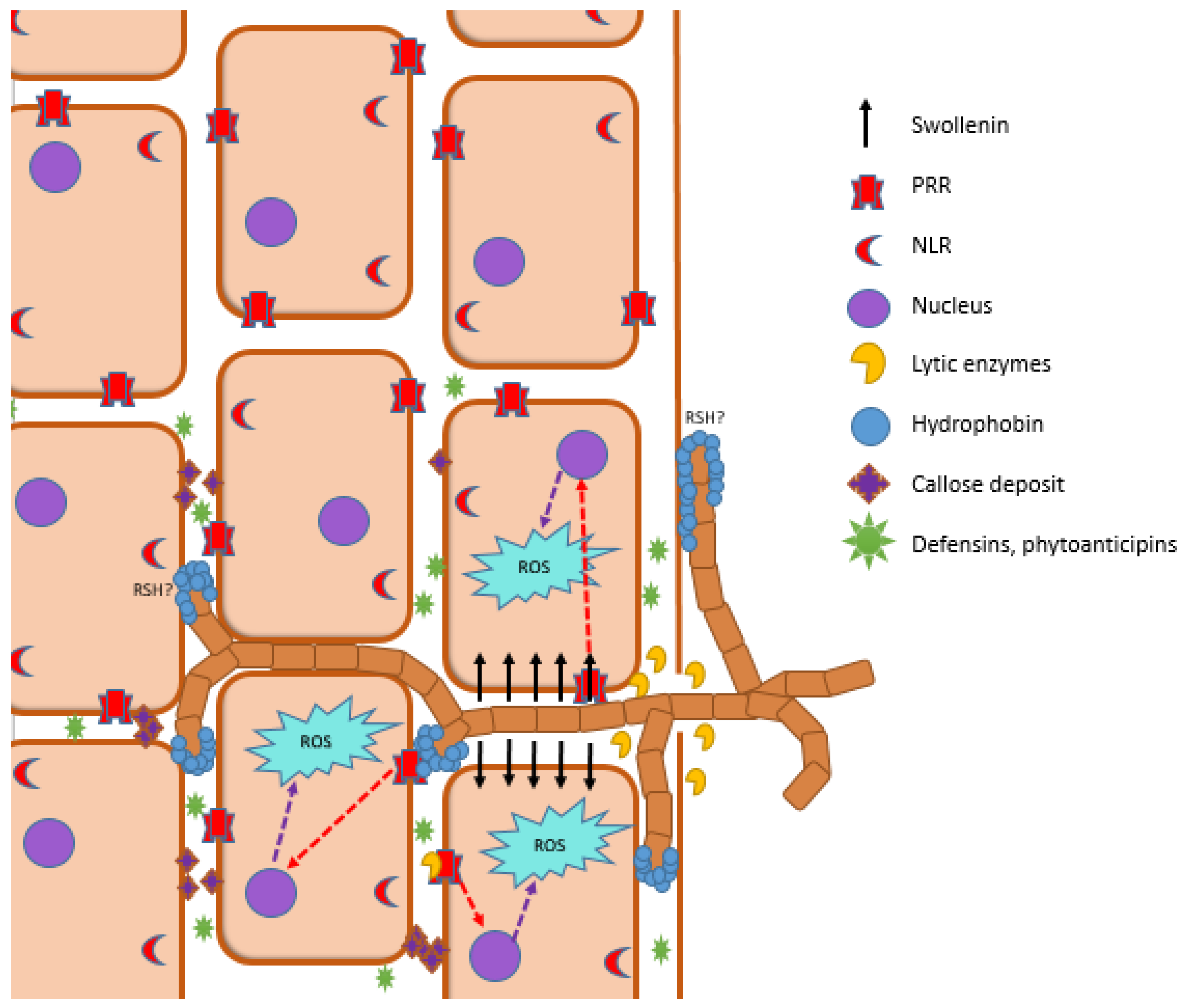
| PGPM | Plant | Remarks | References |
|---|---|---|---|
| Bacteria | |||
| Acinetobacter sp. RG30, Pseudomonas putida GN04 | Zea mays | Plant: - increased tolerance to Cu - enhanced chlorophyll content - increased Cu concentration in tissues Bacteria: - IAA synthesis - production of siderophores - solubilization of Cu and P | [17] |
| Acinetobacter sp. RSC9 | Saccharum sp. | Plant: - under salt stress enhanced number of leaves, fresh, dry weight, and germination ratio Bacteria: - IAA production - P, K, and Zn solubilization - N2 assimilation | [18] |
| Agrobacterium sp. 10C2 | Phaseolus vulgaris | - increased nodule formation - higher plant biomass - enhanced content of P, polyphenols, and flavonoids in grains - changes in the structure of the microbial community | [19] |
| Arthrobacter globiformis | Zea mays, Triticum aestivum | Plant: - enhanced biomass, uptake of Fe and P, and higher chlorophyll content under iron-stress Bacteria: - siderophores production | [20] |
| Arthrobacter sp., Bacillus megaterium | Lycopersicon esculentum | - enhanced seed germination ratio, seedling length, and dry and fresh weight under salt stress | [21] |
| Azospirillum brasilense | Cicer arietinum | - increased resistance to Ascochyta rabiei via induction of plant defense-related genes (Snakin2 and DEF0422) | [22] |
| Azospirillum lipoferum | Triticum aestivum | - improved germination, plant growth, higher chlorophyll content, and improved membrane stability under salt stress - increased production of SOD and osmolytes, i.e., proline, soluble protein, and sugars under salt stress | [23] |
| Azotobacter spp. | Zea mays | Plant: - increased shoot dry weight, chlorophyll content, and N, P, Fe concentration under drought stress Bacteria: - production of siderophores | [24] |
| Bacillus amyloliquefaciens | Zea mays | - increased tolerance to salt stress, enhanced content of chlorophyll, soluble sugars, and glutathione, higher peroxidase/catalase activity | [25] |
| Bradyrhizobium sp., Rhizobium leguminosarum, Azotobacter sp. | Gossypium hirsutum | Plant: - increased rate of seedling emergence, biomass, and N uptake Bacteria: - IAA production | [26] |
| Burkholderia phytofirmans PsJN | Triticum aestivum | - improved water content and CO2 assimilation rate, water use efficiency, chlorophyll content, and higher yield under drought stress - improved ionic balance, antioxidant levels, higher N, P, K, and protein content in grains | [27] |
| Burkholderia tropica | Lycopersicum esculentum | Plant: - increased yield Bacteria: - N-fixation and P solubilization | [28] |
| Enterobacter cloacae | Spinacia oleracea | - protection against Fusarium wilt (Fusarium oxysporum) | [29] |
| Frankia spp. | Casuarina glauca, Casuarina equisetifolia | - salt stress alleviation, higher dry biomass, chlorophyll, and proline content | [30] |
| Methylobacterium sp. 2A | Arabidopsis thaliana, Solanum tuberosum | Plant: - the alleviation of salt stress of A. thaliana, with higher lateral roots density, number of leaves, and larger rosette diameter - reduced necrotic lesions and chlorosis in S. tuberosum infected with P. infestans Bacteria: - production of IAA, P solubilization, biocontrol activity against Phytophtora infestans, Botrytis cinerea, and Fuasrium gramiearum | [31] |
| Pseudomonas putida | Lycopersicum esculentum | Plant: - increased plant height, stem diameter, radical volume, dry biomass, and fruit yield Bacteria: - production of IAA | [32] |
| Pseudomonas sp. DW1 | Solanum melongena | - salt stress ameliorating effect, with increased dry weight, and seed germination - higher SOD activity in leaves | [33] |
| Pseudomonas stutzeri ISE12 | Brassica napus | - enhanced growth under salt stress, with a decrease in non-enzymatic antioxidants accumulation - improved seed germination ratio, number of leaves, chlorophyll content, and dry weight | [34] |
| Rhizobium leguminosarum, Rhizobium sp., Bradyrhizobium sp. | Oryza sativa | Plant: - increased yield and uptake of N, P, K, and Fe - improved seed vigor, dry biomass, and leaf area with faster seedling emergence Bacteria: - production of IAA | [35,36] |
| Serratia marcescens | Solanum melongena | - salt stress alleviation, decreased Na+/Cl- content in leaves, lower lipid peroxidation level, and higher activity of antioxidant enzymes - enhanced biomass, longer stems, and bigger leaf area | [37] |
| Serratia proteamaculans, Pseudomonas putida, Pseudomonas aeruginosa | Triticum aestivum | Plant: - salt stress alleviation with enhanced plant height, root length, and yield, and higher chlorophyll content Bacteria: - ACC deaminase production | [23] |
| Streptomyces sp. | Arabidopsis thaliana, Lycopersicon esculentum | Plant: - salt stress alleviation with increased biomass, chlorophyll content, and decreased proline content Bacteria: - production of IAA, ACC deaminase, P, and NaCl solubilization | [38] |
| Streptomyces sp. | Medicago sativa | - protection against root-lesion nematode (Pratylenchus penetrans) | [39] |
| Fungi | |||
| Alternaria solani IA300 | Capsicum annum | - enhanced number of leaves, flowers, dry, and fresh weight | [40] |
| Apergillus niger 9-p | Phasoleus vulgaris | Plant: - increased biomass Fungus: - production of IAA, ACC deaminase, siderophores, protease, amylase, pectinase, xylanase, and P solubilization | [41] |
| Aspergillus fumigatus | Glycine max | Plant: - salt stress alleviation with enhanced biomass, leaf area, chlorophyll content, and photosynthetic rate - increased isoflavones, proline, SA, and JA content and lower ABA content Fungus: - GAs production (GA4, GA9, GA12) | [42] |
| Collybia tuberosa, Clitocybe sp., Laccaria laccata, Hebeloma mesophaeum, Cyathus olla | Brassica napus | Plant: - enhanced root and shoot growth, number of leaves, and biomass Fungi: - production of IAA | [43] |
| Funneliformis mosseae, Ensifer meliloti | Vitis vinifera | - enhanced plant height and dry weight - higher VOCs content in roots | [44] |
| Fusarium equiseti, Glomus mosseae | Cucumis sativus | - protection against anthracnose (Colletotrichum orbiculare) and damping off (Rhizoctonia solani) - enhanced shoot dry weight | [45] |
| Fusarium verticillioides, Humicola sp. | Glycine max | - salt stress alleviation with increased shoot length, protein content, carotenoid, salicylic acid (SA), and enhanced SOD activity - decreased ABA level and lipid peroxidation level | [46] |
| Glomus intraradices, Glomus mosseae | Olea europaea | - enhanced yield, dry weight, height, stem diameter, and root length | [47] |
| Lecanicillium psalliotae | Elettaria cardamomum | Plant: - enhanced shoot and root length, biomass, and number of leaves - higher chlorophyll content Fungus: - production of IAA, ammonia, siderophores, and cell-wall degrading enzymes - P and Zn solubilization | [48] |
| Mortierella antarctica, Mortierella. Verticillata | Triticum aestivum | Plant: - enhanced fresh weight Fungi: - production of IAA, GA, and ACC deaminase | [49] |
| Mucor sp. | Arabidopsis arenosa | - heavy metal (Zn, Cd, Fe, Pb) stress alleviation with enhanced biomass, root hair growth, improved water, and P content - upregulation of genes involved in nutrient acquisition (HRS1, SPX1, MGD2), and metal homeostasis (MTPA2, ZIP7, IREG2, IRT2) | [50] |
| Penicillium bilaii | Pisum sativum | - increased root dry weight, length, and P content in the shoot | [51] |
| Penicillium sp., Penicillium radicum, Penicillium bilaiae | Medicago lupulina, Lens culinaris, Triticum aestivum | - enhanced shoot growth and dry weight, and increased P uptake | [52] |
| Phoma sp. | Cucumis sativus, Arabidopsis thaliana | - protection against cucumber mosaic virus (CMV) via ISR - higher number of leaves, increased fresh/dry weight, and the yield of cucumber | [53] |
| Phoma spp., Trichoderma asperellum, Fusarium equiseti, Penicillium simplicissmum | Allium cepa | - protection against white rot disease (Sclerotium cepivorum) with enhanced plant height, dry weight, and bulb perimeter - enhanced levels of peroxidase and polyphenol oxidase - upregulation of plant defense genes (PR1, PR2) | [54] |
| Purpureocillium lilacinum, Purpureocillium. lavendulum, Metarhizium marquandii | Zea mays, Phaseolus vulgaris, Glycine max | Plant: - enhanced plant height and biomass and N content in roots (Z. mays) and P in shoots (P. vulgaris) Fungi: - P solubilization and IAA production | [55] |
| Trichoderma hamatum, Trichoderma harzianum, Trichoderma viride | Freesia refracta | - accelerated flowering and enhanced development of lateral inflorescence shoots - increased K, Fe, Mn, and Zn uptake | [56] |
| Trichoderma harzianum | Curcuma longa | Plant: - enhanced plant height and yield Fungi: - biocontrol activity against rhizome rot and leaf blight (Pythium aphanidermatum, Rhizoctonia solani) - production of IAA, HCN, cellulase, and P solubilization | [57] |
| Trichoderma phayaoense | Cucumis melo | Plant: - enhanced plant development, biomass, and fruit yield Fungus: - biocontrol activity against gummy stem blight pathogens (Stagonosporopsis cucurbitacearum, Fusarium equiseti) | [58] |
| Trichoderma viride | Brassica napus | - enhanced biomass, lateral roots development, and germination ratio - changes in microbial composition | [59] |
| Algae | |||
| Anabaena oryzae, Anabaena doliolum, Phormidium fragile, Calothrix geitonos, Hapalosiphon intricatus, Aulosira ferilissima, Tolypothrix tenuis, Oscillatoria acuta, Plectonema boryanum | Oryza sativa | - enhanced shoot and root length and biomass - improved protein, phenolics, flavonoids, and chlorophyll content - a higher activity of enzymes (peroxidase, phenylalanine, and ammonia lyase) - elevated levels of IAA, and IBA | [60] |
| Anabaena variabilis, Anabaena laxa | Lycopersicon esculentum | - protection against Fusarium wilt (F. oxysporum) with significant enhancement of growth, yield, and fruit quality - increased N, P, and Zn concentration - increased activity of defense enzymes (phenylalanine ammonia-lyase, polyphenol oxidase), increased activity of chitosanase, and β-1,3-glucanase | [61] |
| Calothrix elenkinii | Oryza sativa | - enhanced root/shoot length and fresh weight - improved chlorophyll and IAA content - higher nitrogenase and CMCase activity - 10-fold increase in microbiome population abundance | [62] |
| Calothrixsp., A. laxa, Anabaena torulosa, Anabaena azollae, Anabaena oscillarioides | Triticum aestivum | - enhanced biomass - nitrogen-fixing potential - higher endoglucanase activity | [63] |
| Chlorella fusca | Cucumis sativus | - protection against anthracnose (Colletotrichum orbiculare) via the induction of SAR | [64] |
| Chlorella oocystoides, Chlorella minutissima | Zea mays | - enhanced chlorophyll, P, and K content - improved biomass | [65] |
| Chlorella vulgaris | Telfairia occidentalis | - enhanced germination ratio - higher number of leaves and yield - improved chlorophyll, carbohydrates, proteins, and lipid content | [66] |
| Microcystis aeruginsa | Oryza sativa | - heavy metal (Cd) stress alleviation with decreased Cd accumulation, increased translocation of Cd from root to shoot, and enhanced dry weight | [67] |
| Nostoc sp. | Triticum aestivum, Oryza sativa | Plant: - enhanced biomass and shoot/root length Algae: - production of IAA and zeatin | [68] |
| Nostoc sp. | Zea mays | - enhanced dry mass - higher N content - production of exopolysaccharide | [69] |
| Scenedesmus quadricauda, Chlorella vulgaris | Lycopersicon esculentum | - enhanced biomass and root length | [70] |
| Spirulina platensis | Zea mays | - cadmium stress alleviation with improved photosynthetic electron flows and increased non-photochemical quenching - enhanced seed germination, shoot length, root fresh weight, and bigger leaf area - decreased Cd accumulation in shoot | [71] |
| Mixed inoculants | |||
| Anabaena ssp., Calothrix sp., Providencia sp. | Triticum aestivum | - enhanced yield and Fe, Cu, Zn, Mn, and protein content of grains | [72] |
| Glomus fasciculatum, Bacillus subtilis | Tagetes erecta | - enhanced flowering, with improved fresh weight and color of flowers | [73] |
| Klebsiella variicola, Glomus multisubtensum, Rhizophagus intraradices | Helianthus tuberosus | Plant: - enhanced biomass, yield, plant height, and leaf area - increased content of inulin in tubers Microbes: - P solubilization and IAA production | [74] |
| Mesorhizobium mediterraneum, Rhizophagus irregulari | Cicer arietinum | - enhanced yield and protein content of grain under water deficit conditions | [75] |
| Rhizophagus intraradices, Glomus aggregatum, Glomus viscosum, Claroideoglomus etunicatum, Claroideoglomus claroideum, Pseudomonas fluorescens, Linum usitatissimum | Solanum lycopersicum | - enhanced flower and fruit production, with increased lycopene, vitamins, sugars, and citric acid content of the fruits | [76] |
| Rhizophagus intraradices, Pseudomonas sp., Bacillus sp. | Sulla carnosa | Plant: - enhanced biomass, stomatal conductance, photosynthetic pigment content, and photosynthesis rate under salt stress - increased proline content and higher activity of antioxidative enzymes Microbes: - production of IAA | [77] |
| Septoglomus constrictum, Diversispora aunantia, Archaeospora trappei, Glomus versiforme, Paraglomus ocultum, Bacillus thuringiensis | Lavandula dentata | Plant: - increased biomass under drought stress conditions, enhanced activity of the enzymatic antioxidant system, and enhanced nutrient uptake Microbes: - P solubilization, production of IAA, and ACC deaminase | [78] |
| Trichoderma harzianum, Glomus spp., Pseudomonas fluorescens | Capsicum annuum | - enhanced yield, higher activity of antioxidative, and defense enzymes | [79] |
| Gene/Product | Function | Species | Reference |
|---|---|---|---|
| aph/acid phosphatase | - increased P availability via phosphates solubilization | Aspergillus, Trichoderma, Penicillium | [238,239] |
| AMT1; AMT2; AAT9/ammonium transporter; amino acid transporter | - improved N acquisition | Tulasnella calospora | [240] |
| AQPF/aquaporin | - transport of water to the host - enhanced drought stress resistance | Glomus intraradices | [241] |
| Phy/phytase | - increased P availability via solubilization of inositol | Aspergillus, Trichoderma, Penicillium | [238,239,242] |
| acdS/ACC deaminase | - degradation of ethylene precursor and protects against elevated ethylene levels - ameliorates stress effects and promotes root growth | Trichoderma asperellum, Penicillium citrinum, Trichoderma gamsii | [243] |
| Hyd; Qid/hydrophobins | - allows for adhesion of hyphae to the surface of roots and protects hyphae against antifungal compounds - functions as MAMP (microbe-associated molecular pattern) and triggers plant response involved with symbiont recognition | Trichoderma asperellum, Trichoderma harzianum | [244,245] |
| MST2/monosaccharide transporter2 | - development of arbuscules - facilitates root colonization | Glomus sp. | [246] |
| Tex1/non-ribosomal peptided synthase | - synthesis of trichovirin II (peptaibol) which activates the plant immune system | Trichoderma virens | [247] |
| Thctf1/transriptional factor | - regulates the synthesis of 6-pentyl-2H-pyran-2-one (6-PP) (VOC) which exhibits antifungal activity | Trichoderma sp. | [248] |
| Thph1; Thph2/cellulases | - cellulolytic activity - triggers plant immune system | Trichoderma harzianum | [236] |
| sidD/siderophores synthase | - synthesis of siderophores - improved Fe acquisition - defense against pathogens | Trichoderma reesei, Trichoderma virens | [249] |
| Sm1; Sm2; Ep1; Swo /cerato-platanins; swollenin | - fungal elicitors and upregulation of genes involved in JA signaling (modulation of the immune system) - swollenin disrupts the plant cell wall structure and enables penetration of the apoplast | Trichoderma citrinoviride, Trichoderma virens | [250,251,252] |
| Gene/Product | Function | Species | Reference |
|---|---|---|---|
| 2,3-butanediol dehydrogenase | - synthesis of 2,3-butanediol - growth promotion - induction of ISR | Bacillus sp., Aerobacter sp., Serratia sp., Enterobacter sp., Klebsiella sp. | [253] |
| acdS/ACC deaminase | - degradation of ethylene precursor and protects against elevated ethylene levels - ameliorates stress effects | Azospirillum sp. Pseudomonas putida | [254,255] |
| alkaline phosphatase | - increased P availability via phosphates solubilization | Pseudomonas brassicacearum | [253] |
| bud operon | - synthesis of acetoine and 2,3-butanediol - induction of ISR (induced systemic resistance) - increased drought tolerance | Enterobacter sp638 | [256] |
| chitinase; glucanase | - defense against fungal pathogens | Pseudomonas aureginosa, Pseudomonas veronii | [257,258] |
| exoprotease | - N acquisition - protection against pathogens | Pseudomonas brassicacearum | [253] |
| fur/transcription factor | - modulates gene expression encoding Fe transporter - Fe acquisition | Pseudomonas brassicacearum | [253] |
| gcd/pyrroloquinoline quinone (PQQ)-dependant dehydrogenase | - production of gluconic acid - P acquisition | Pseudomonas fluorescens F113, Erwinia herbicola, Enterobacter intermedium | [253] |
| hcnABC/HCN synthase | - protection against pathogens | Pseudomonas fluorescens. Pseudomonas aeruginosa, Pseudomonas chlororaphis | [259] |
| ilvHI; ivlC/acetohydroxyacid synthase; ketol-acid reductoisomerase | - synthesis of secondary metabolites including antibiotics - induction of ISR | Bacillus subtilis | [253] |
| ipd; ppd /indole-3-pyruvate decarboxylase; phenylpyruvate fenylopirogronian decarboxylase | - synthesis of IAA - promotion of root growth | Azospirllum brasilense Sp245, Enterobacter cloacae UW5, Enterobacteriaceae | [260,261,262] |
| nagA/N-acetylglucosamine-6 phosphate deacetylase | - chitinase-like protein and defense against fungal pathogens | Pseudomonas brassicacearum | [253] |
| nif/nitrogenase | - nitrogen assimilation | Azospirllum, Burkoholderia, Rhizobium, Bradyrhizobium, Mesorhizobium, Delftia, Stenotrophomonas, Rhizobium, Brevundimonas, Variovorax, Achromobacter, Novosphingobium, Comamonas | [263,264,265,266] |
| phl operon | - production of antibiotic 2,4-diacetylphloroglucinol - induction of ISR | Pseudomonas fluorescens F113, Pseudomonas protegnes CHA0 | [267] |
| phyC/phytase | - increased P availability via solubilization of inositol | Pseudomonas brassicacearum | [253] |
| rhb; rhtA/siderophore synthase; mebrane Fe-regulated receptor | - production of rhziobactin (siderophore) and Fe acquisition - Fe uptake regulation | Sinorhizobium meliloti | [268] |
| ribC/riboflavin synthase | - growth promotion - defense against pathogens via ISR - upregulation of pathogenesis-related genes | Pseudomonas yamanorum | [269] |
| yecA; speB/polyamine permease; agmatinase | - synthesis and/or secretion of polyamines - lowers ethylene level in root cells - modulation of expansin genes expression - promotion of root growth - increases tolerance to low pH, oxidative and osmotic stress | Bacillus subtilis OKB105 | [270] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Antoszewski, M.; Mierek-Adamska, A.; Dąbrowska, G.B. The Importance of Microorganisms for Sustainable Agriculture—A Review. Metabolites 2022, 12, 1100. https://doi.org/10.3390/metabo12111100
Antoszewski M, Mierek-Adamska A, Dąbrowska GB. The Importance of Microorganisms for Sustainable Agriculture—A Review. Metabolites. 2022; 12(11):1100. https://doi.org/10.3390/metabo12111100
Chicago/Turabian StyleAntoszewski, Marcel, Agnieszka Mierek-Adamska, and Grażyna B. Dąbrowska. 2022. "The Importance of Microorganisms for Sustainable Agriculture—A Review" Metabolites 12, no. 11: 1100. https://doi.org/10.3390/metabo12111100
APA StyleAntoszewski, M., Mierek-Adamska, A., & Dąbrowska, G. B. (2022). The Importance of Microorganisms for Sustainable Agriculture—A Review. Metabolites, 12(11), 1100. https://doi.org/10.3390/metabo12111100






