Stochastic Neighbor Embedding Feature-Based Hyperspectral Image Classification Using 3D Convolutional Neural Network
Abstract
1. Introduction
- We proposed a modified tSNE through the PCA algorithm to solve the visualization, dimensionality, and computational complexity problem. PCA is used to discover the irrelevant feature bands, aiming to increase the performance of tSNE. The tSNE preserves the intra- and inter-band relationship of the HSI, which is the most effective feature of the HSI for classification and visualization.
- We designed a new blended CNN architecture for feature extraction and classification, which is a sequential combination of 3D- and 2D-CNN to incorporate the spatial and spectral information of HSI, where the combined spectral information contains the wavelengths of the bands and the spatial information contains the location information of the band.
- Two benchmark HSI datasets were used to evaluate the proposed system, namely Indian Pines and SalinusA. Finally, the state-of-the-art comparison table proves the superiority of the proposed model over the mentioned systems.
2. Related Work
3. Dataset
3.1. Indian Pines
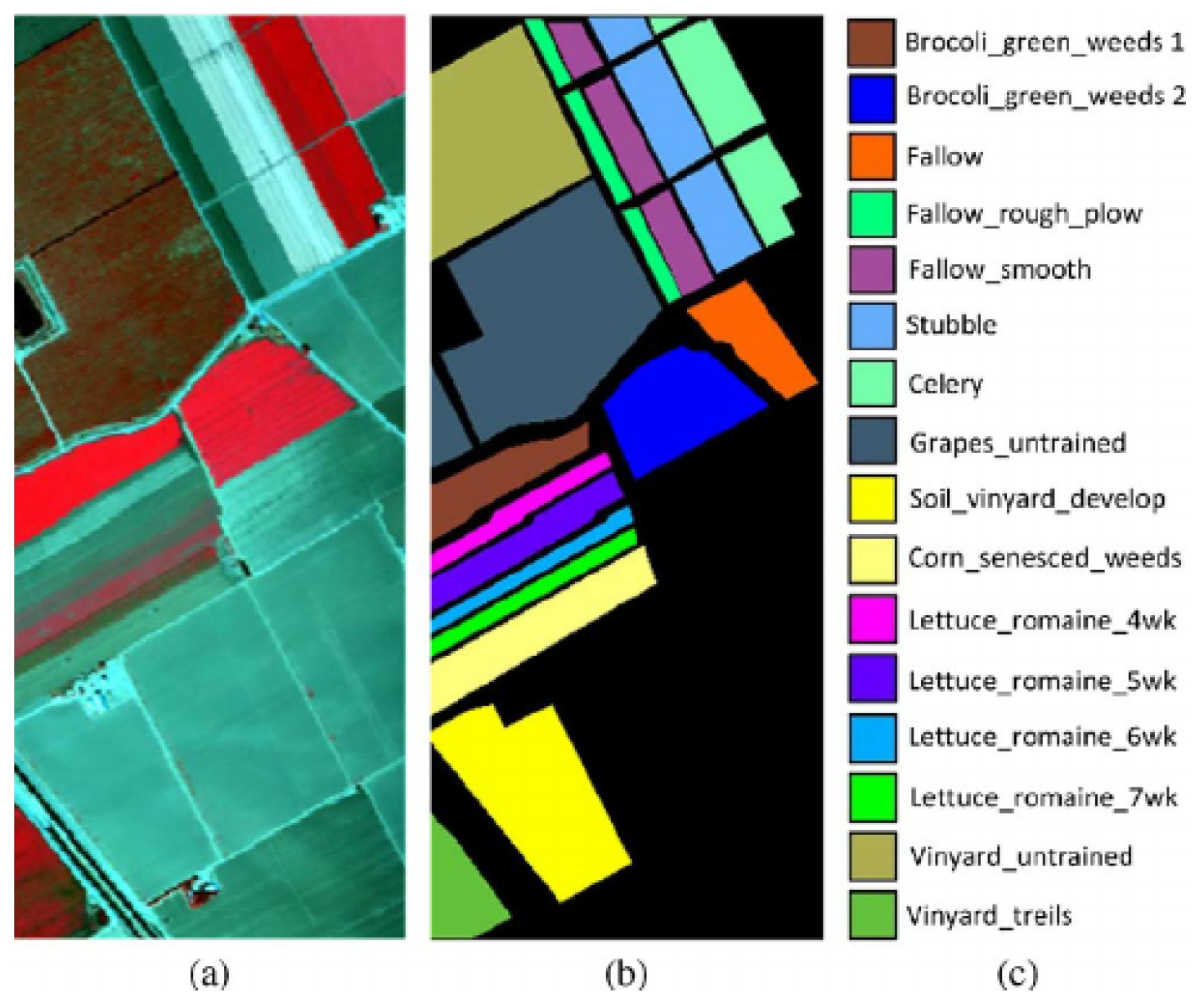
3.2. Salinas
4. Proposed Methodology
| Algorithm 1 Proposed Method Pseudocode |
| Input: Set of Input Dataset with dimension |
| Number of Samples: N, 70% for Training and 30% for Test |
| Output: Set of predicted class |
| define BlendedCNNModel(input=, outputs=ClassificationLayer): |
| return PredictedClass |
| while do |
| // For Training |
| while do |
| // For Testing |
| while do |
4.1. Principle Component Analysis (PCA) for HSI
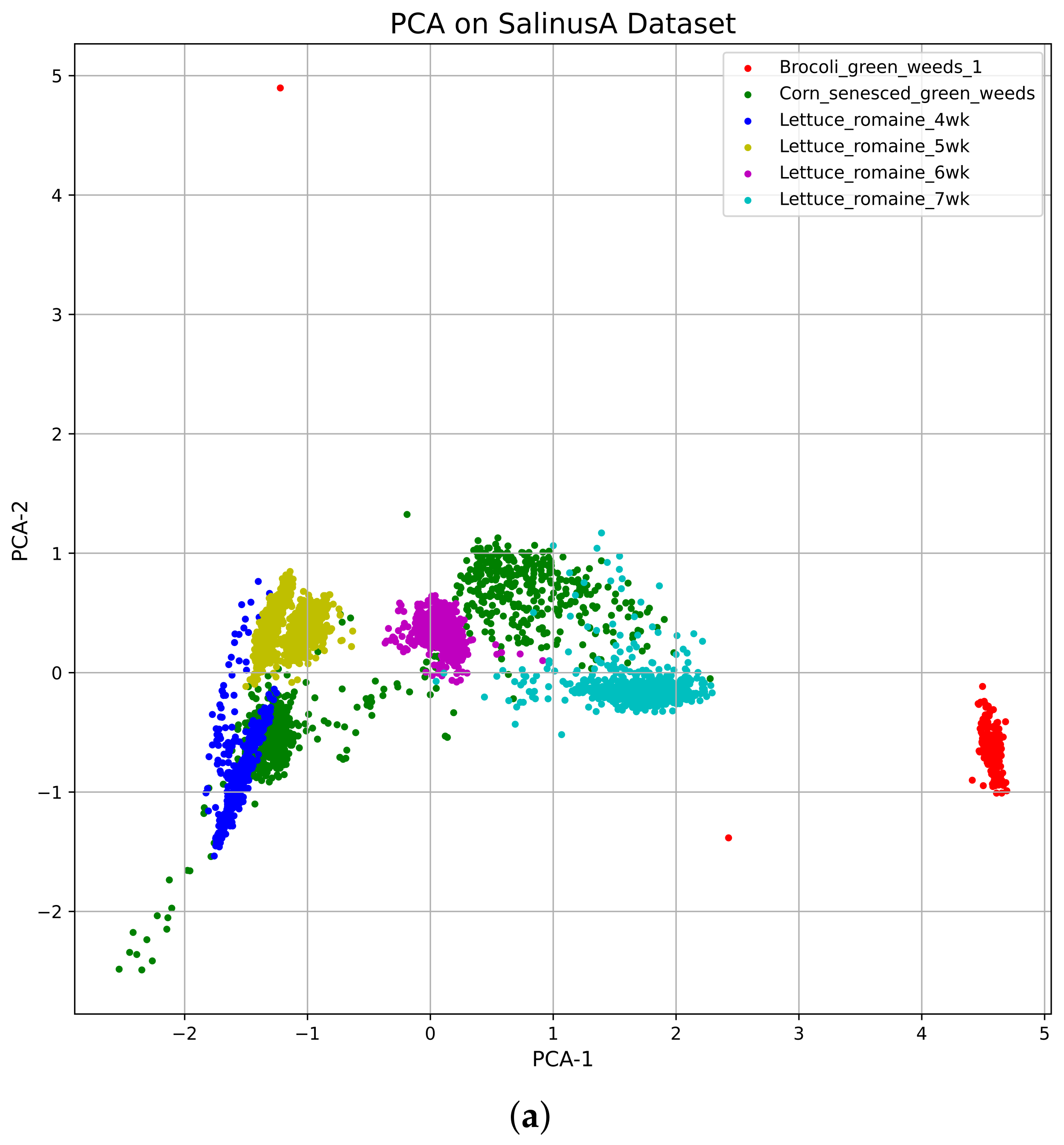
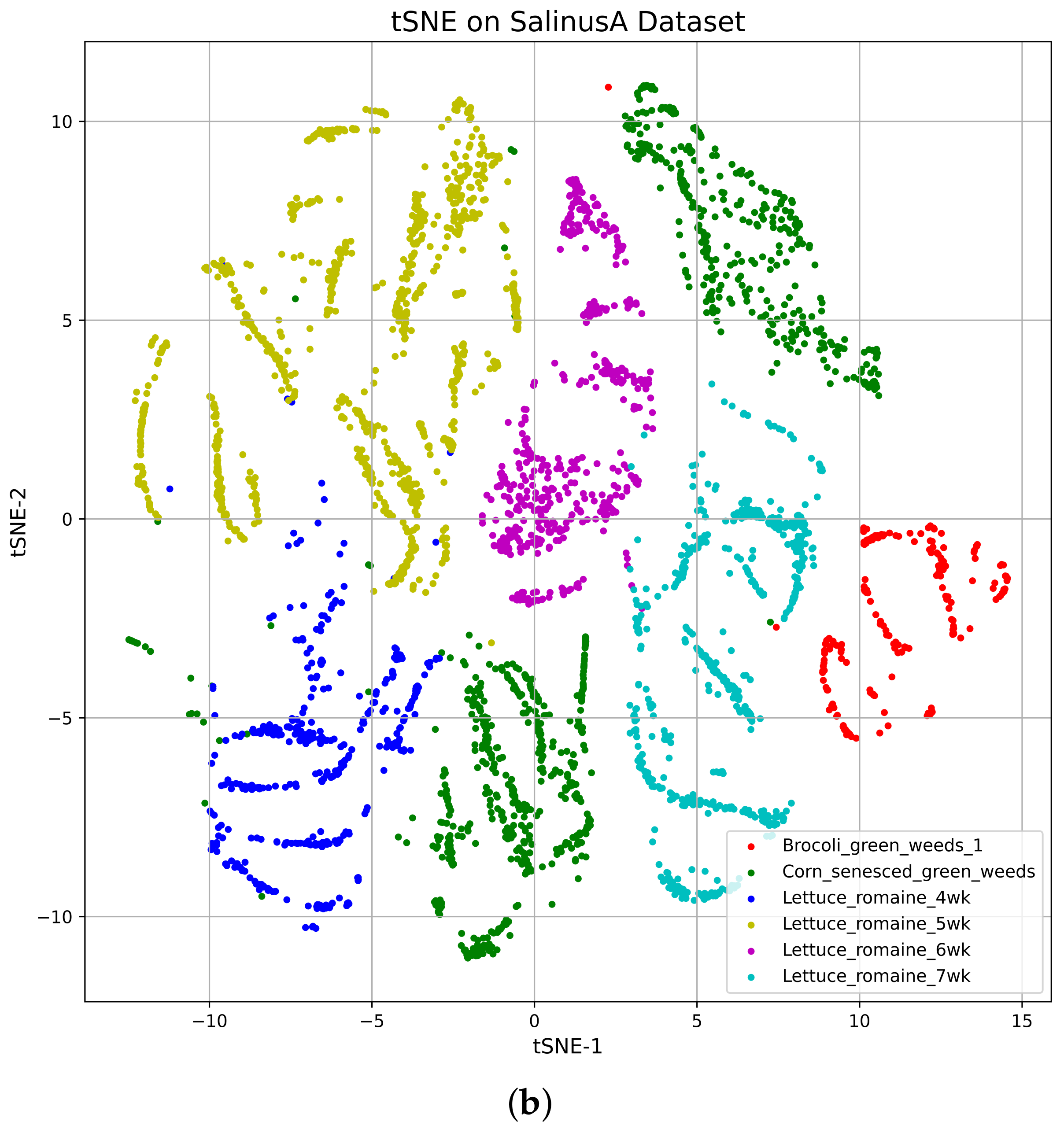
4.2. t-Distributed Stochastic Neighbor Embedding (tSNE)
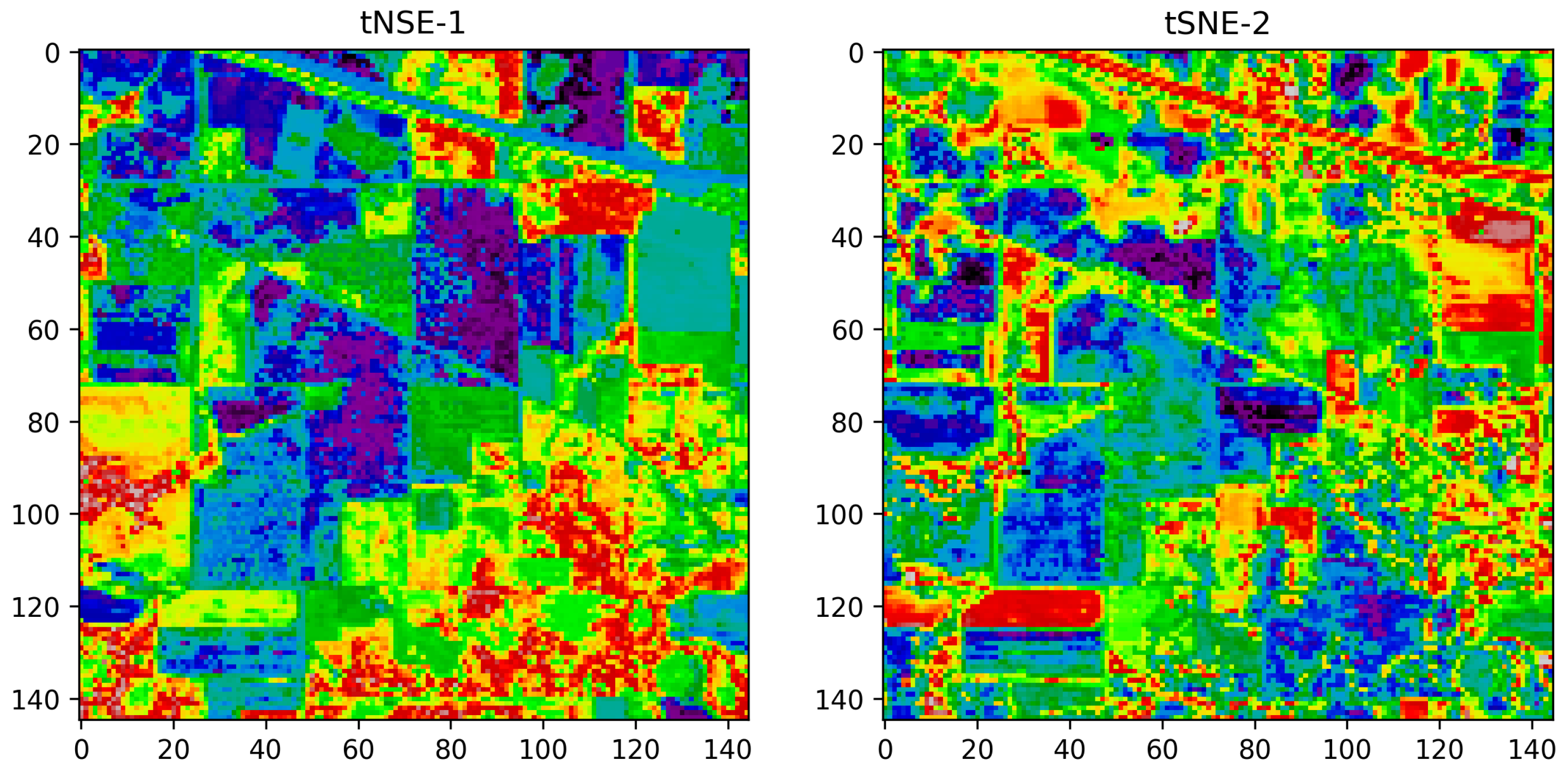
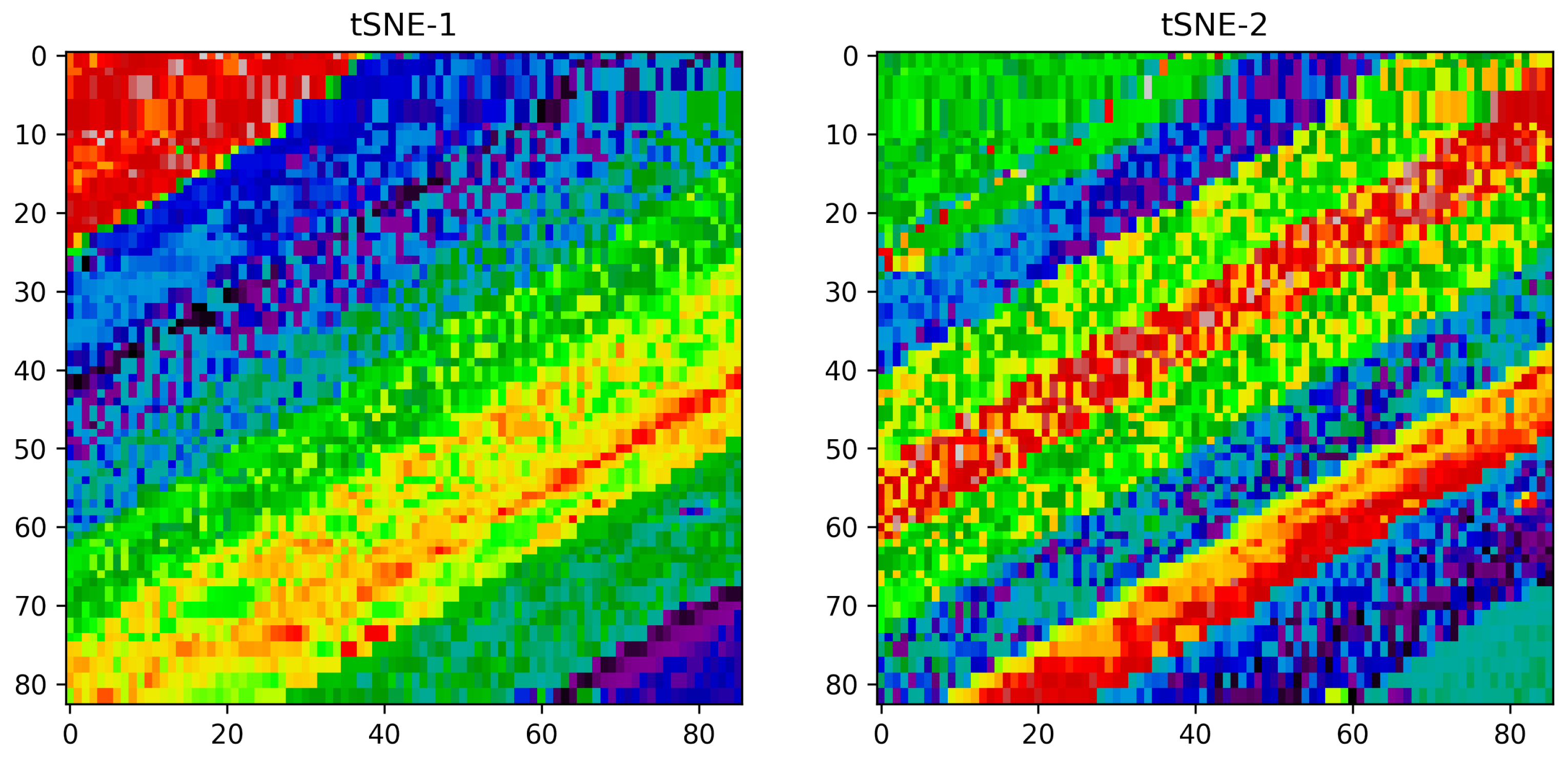
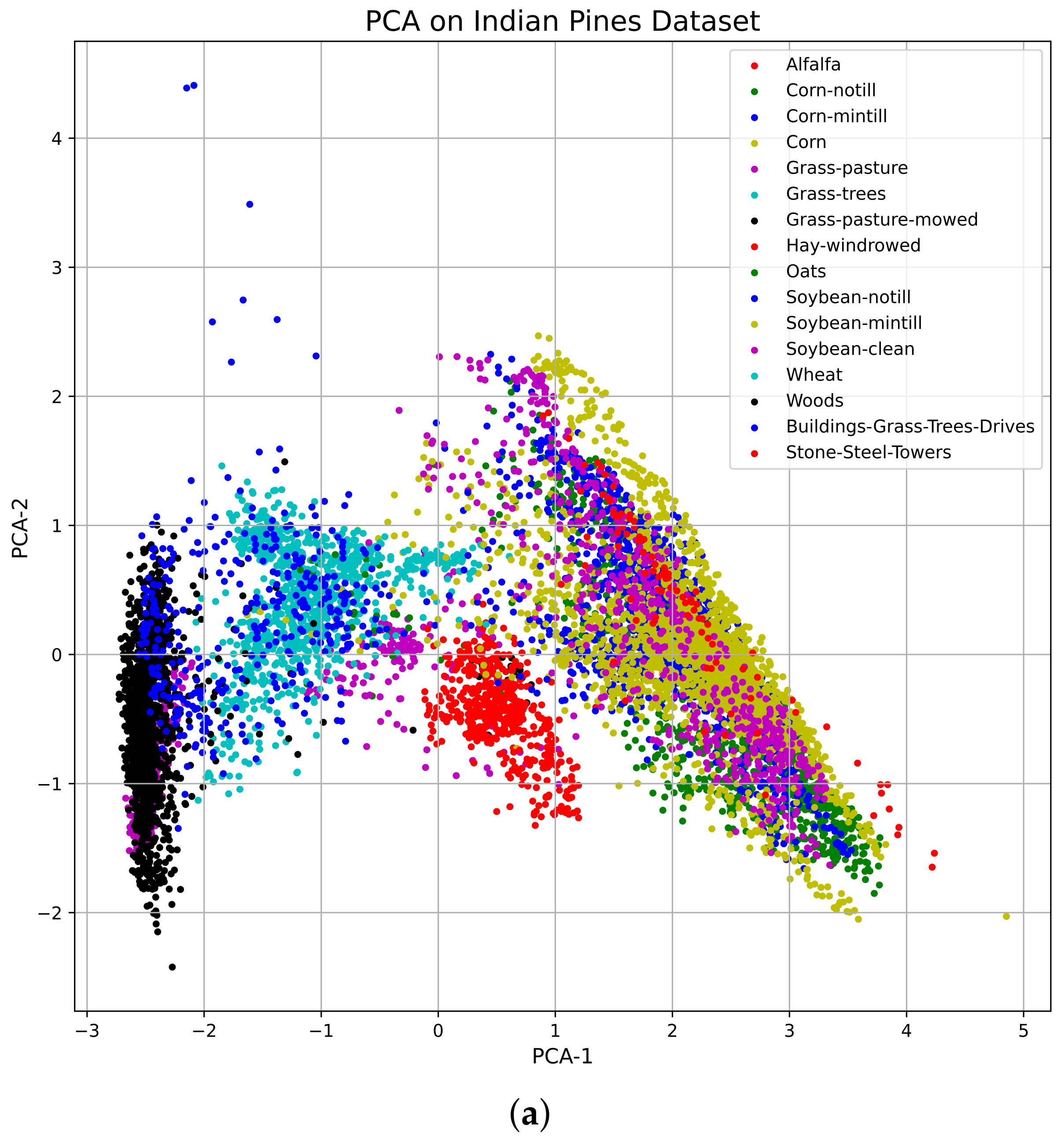
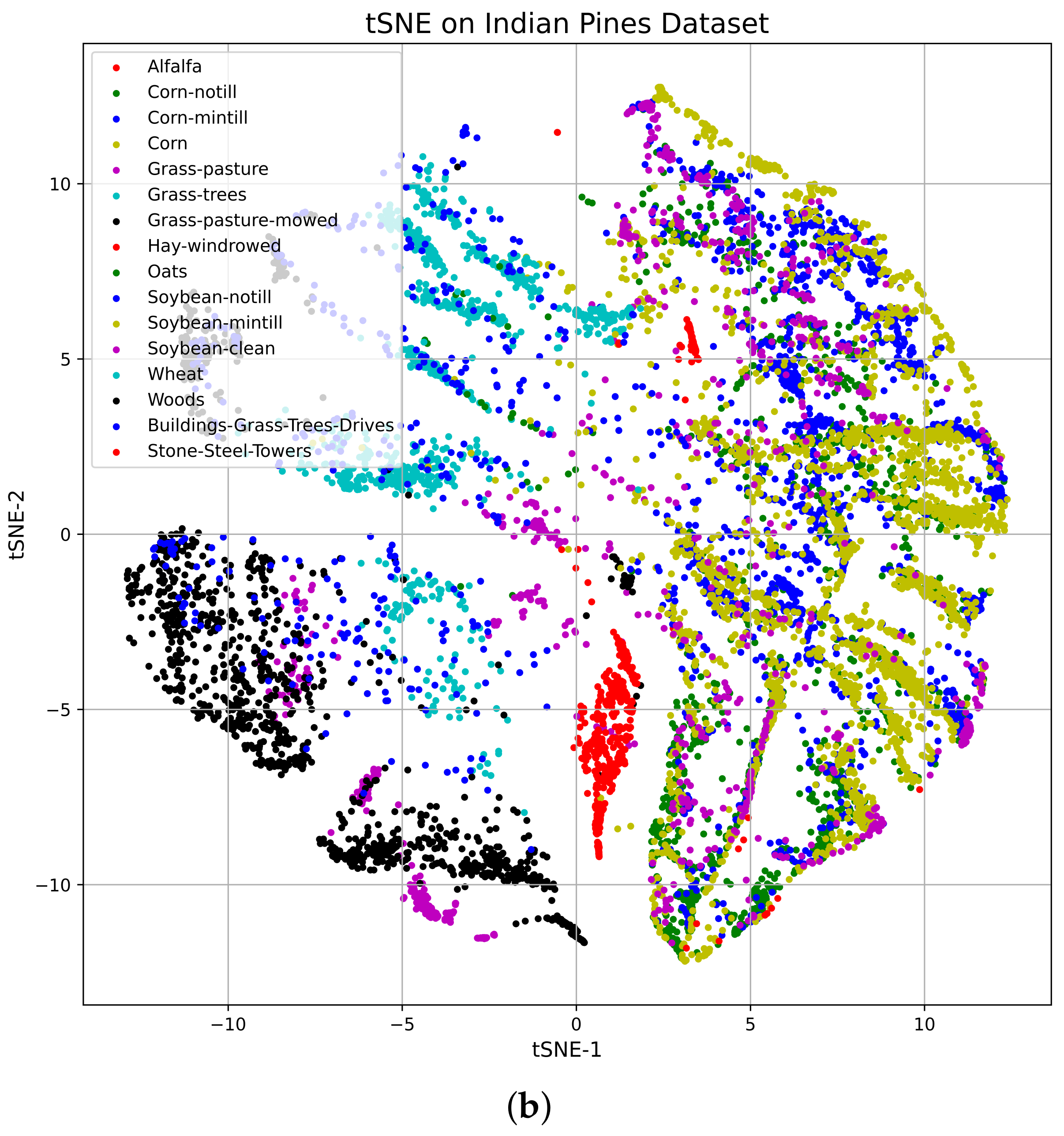
Vizualization of the TSNE
4.3. BlendedCNN Architecture for HSI Classification
5. Experimental Setups and Results
5.1. Experimental Settings
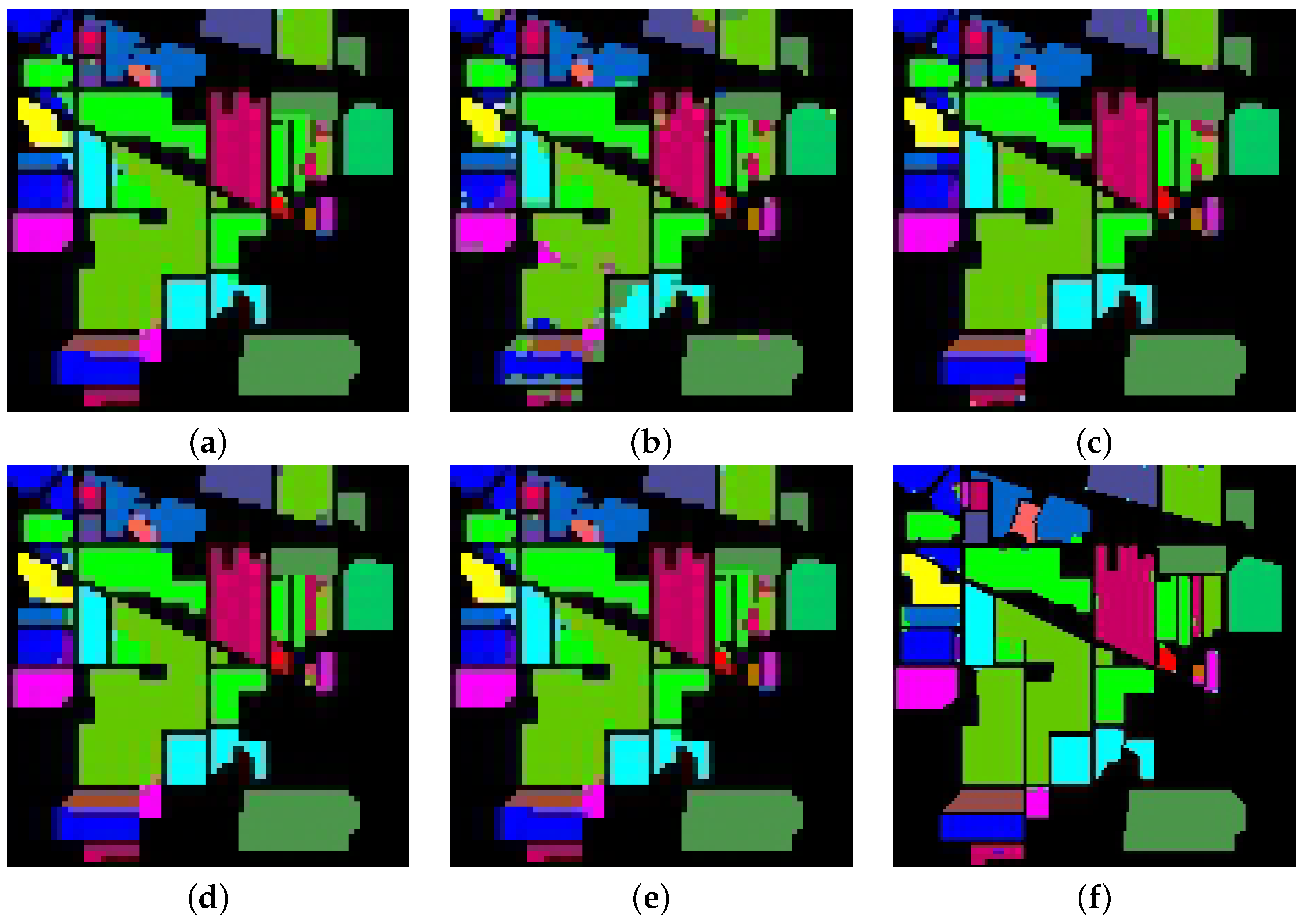
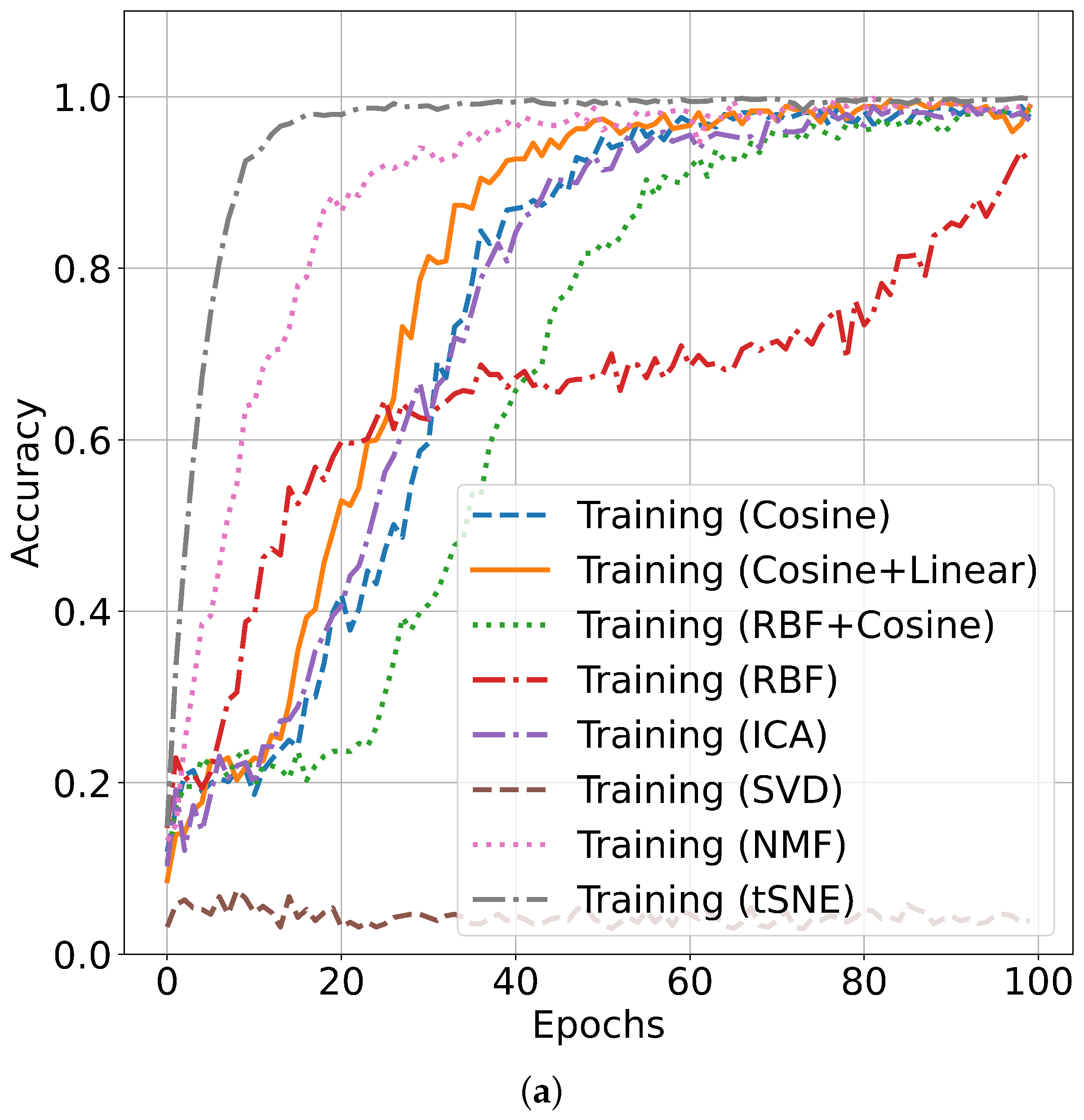
5.2. Performance Accuracy with Indian Pines Dataset
5.3. Performance Accuracy with SalinasA Dataset
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Lebedev, A.; Zavarzin, V.; Gemonov, A. Vegetation Cover Change in Kologrivsky Forest Nature Reserve Detected using Landsat Satellite Image Analysis. IOP Conf. Ser. Earth Environ. Sci. 2020, 507, 012016. [Google Scholar] [CrossRef]
- Fei, B. Chapter 3.6—Hyperspectral imaging in medical applications. In Data Handling in Science and Technology; Amigo, J.M., Ed.; Elsevier: Amsterdam, The Netherlands, 2020; Volume 32, pp. 523–565. [Google Scholar] [CrossRef]
- Contributors. In Hyperspectral Imaging for Food Quality Analysis and Control; Sun, D.W., Ed.; Academic Press: San Diego, CA, USA, 2010; pp. 9–11. [CrossRef]
- Ziemann, A.; Theiler, J. Material Detection in Hyperspectral Imagery in Support of Nuclear Nonproliferation. 2016. Available online: https://public.lanl.gov/jt/Papers/Ziemann_Theiler_ANS2016_v2.pdf (accessed on 1 January 2023).
- Bai, D.; Messinger, D.W.; Howell, D. A pigment analysis tool for hyperspectral images of cultural heritage artifacts. In Proceedings of the Algorithms and Technologies for Multispectral, Hyperspectral, and Ultraspectral Imagery XXIII, Anaheim, CA, USA, 11–13 April 2017; Velez-Reyes, M., Messinger, D.W., Eds.; International Society for Optics and Photonics, SPIE: Bellingham, WA, USA, 2017; Volume 10198, pp. 429–443. [Google Scholar] [CrossRef]
- Edelman, G.; Gaston, E.; van Leeuwen, T.; Cullen, P.; Aalders, M. Hyperspectral Imaging for Non-Contact Analysis of Forensic Traces. Forensic Sci. Int. 2012, 223, 28–39. [Google Scholar] [CrossRef] [PubMed]
- Kibria, K.A.; Chowdhury, A.A.; Miah, A.S.M.; Shahriar, M.R.; Pervin, S.; Shin, J.; Rashid, M.M.; Sarkar, A.R. Bangladeshi Land Cover Change Detection with Satelite Image Using GIS Techniques. In Machine Intelligence and Data Science Applications: Proceedings of MIDAS 2021, Cumilla, Bangladesh, 26–27 December 2021; Springer: Berlin/Heidelberg, Germany, 2022; pp. 125–143. [Google Scholar]
- Mukundan, A.; Tsao, Y.M.; Cheng, W.M.; Lin, F.C.; Wang, H.C. Automatic Counterfeit Currency Detection Using a Novel Snapshot Hyperspectral Imaging Algorithm. Sensors 2023, 23, 2026. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.Y.; Hsiao, Y.P.; Mukundan, A.; Tsao, Y.M.; Chang, W.Y.; Wang, H.C. Classification of Skin Cancer Using Novel Hyperspectral Imaging Engineering via YOLOv5. J. Clin. Med. 2023, 12, 1134. [Google Scholar] [CrossRef] [PubMed]
- Mukundan, A.; Huang, C.C.; Men, T.C.; Lin, F.C.; Wang, H.C. Air Pollution Detection Using a Novel Snap-Shot Hyperspectral Imaging Technique. Sensors 2022, 22, 6231. [Google Scholar] [CrossRef]
- Chang, C. Hyperspectral Data Processing: Algorithm Design and Analysis; John Wiley & Sons: New York, NY, USA, 2013. [Google Scholar] [CrossRef]
- Bu, C.; Zhang, Z. Research on Overfitting Problem and Correction in Machine Learning. J. Phys. Conf. Ser. 2020, 1693, 012100. [Google Scholar] [CrossRef]
- Zhao, W.; Du, S. Spectral–spatial feature extraction for hyperspectral image classification: A dimension reduction and deep learning approach. IEEE Trans. Geosci. Remote Sens. 2016, 54, 4544–4554. [Google Scholar] [CrossRef]
- Momeni, R.; Aplin, P.; Boyd, D. Mapping Complex Urban Land Cover from Spaceborne Imagery: The Influence of Spatial Resolution, Spectral Band Set and Classification Approach. Remote Sens. 2016, 8, 88. [Google Scholar] [CrossRef]
- Ren, J.; Zabalza, J.; Marshall, S.; Zheng, J. Effective Feature Extraction and Data Reduction in Remote Sensing Using Hyperspectral Imaging. Signal Process. Mag. 2014, 31, 149–154. [Google Scholar] [CrossRef]
- Bandos, T.V.; Bruzzone, L.; Camps-Valls, G. Classification of Hyperspectral Images with Regularized Linear Discriminant Analysis. IEEE Trans. Geosci. Remote Sens. 2009, 47, 862–873. [Google Scholar] [CrossRef]
- Menon, V.; Du, Q.; Fowler, J.E. Fast SVD with Random Hadamard Projection for Hyperspectral Dimensionality Reduction. IEEE Geosci. Remote Sens. Lett. 2016, 13, 1275–1279. [Google Scholar] [CrossRef]
- Falco, N.; Benediktsson, J.A.; Bruzzone, L. Spectral and Spatial Classification of Hyperspectral Images Based on ICA and Reduced Morphological Attribute Profiles. IEEE Trans. Geosci. Remote Sens. 2015, 53, 6223–6240. [Google Scholar] [CrossRef]
- Maaten, L.V.D.; Hinton, G.E. Visualizing Data using t-SNE. J. Mach. Learn. Res. 2008, 9, 2579–2605. [Google Scholar]
- Shambulinga, M.; Sadashivappa, G. Hyperspectral Image Classification using Convolutional Neural Networks. Int. J. Adv. Comput. Sci. Appl. 2021, 12, 702–708. [Google Scholar]
- Zhong, S.; Chang, C.I.; Zhang, Y. Iterative support vector machine for hyperspectral image classification. In Proceedings of the 2018 25th IEEE International Conference on Image Processing (ICIP), Athens, Greece, 7–10 October 2018; pp. 3309–3312. [Google Scholar]
- Shambulinga, M.; Sadashivappa, G. Hyperspectral Image Classification using Support Vector Machine with Guided Image Filter. Int. J. Adv. Comput. Sci. Appl. 2019, 10, 271–276. [Google Scholar] [CrossRef]
- Shambulinga, M.; Sadashivappa, G. Supervised Hyperspectral Image Classification using SVM and Linear Discriminant Analysis. Int. J. Adv. Comput. Sci. Appl. 2020, 11, 0111050. [Google Scholar] [CrossRef]
- Miah, A.S.M.; Rahim, M.A.; Shin, J. Motor-Imagery Classification Using Riemannian Geometry with Median Absolute Deviation. Electronics 2020, 9, 1584. [Google Scholar] [CrossRef]
- Miah, A.S.M.; Rashid, M.M.; Arahman, R.M.; Hossain, T.; Sujon, M.M.; Nafisa, N.; Mohammad, H.; Sin, J. Alzheimer’s Disease Detection Using CNN Based on Effective Dimensionality Reduction Approach. In Advances in Intelligent Systems and Computing, Proceedings of the 3rd International Conference on Intelligent Computing and Optimization 2020 (ICO 2020), Huai Khot, Thailand, 8–9 October 2020; Springer: Cham, Switzerland; Volume 1324, pp. 801–811.
- Miah, A.S.M.; Shin, J.; Hasan, M.A.M.; Rahim, M.A. BenSignNet: Bengali Sign Language Alphabet Recognition Using Concatenated Segmentation and Convolutional Neural Network. Appl. Sci. 2022, 12, 3933. [Google Scholar] [CrossRef]
- Miah, A.S.M.; Shin, J.; Hasan, M.A.M.; Rahim, M.A.; Okuyama, Y. Rotation, Translation and Scale Invariant Sign Word Recognition Using Deep Learning. Comput. Syst. Sci. Eng. 2023, 44, 2521–2536. [Google Scholar] [CrossRef]
- Shin, J.; Musa Miah, A.S.; Hasan, M.A.M.; Hirooka, K.; Suzuki, K.; Lee, H.S.; Jang, S.W. Korean Sign Language Recognition Using Transformer-Based Deep Neural Network. Appl. Sci. 2023, 13, 3029. [Google Scholar] [CrossRef]
- Miah, A.S.M.; Hasan, M.A.M.; Shin, J. Dynamic Hand Gesture Recognition using Multi-Branch Attention Based Graph and General Deep Learning Model. IEEE Access 2023, 11, 4703–4716. [Google Scholar] [CrossRef]
- Hasan, M.; Miah, A.S.M.; Hossain, M.M.; Hossain, M.S. LL-PMS8: A time efficient approach to solve planted motif search problem. J. King Saud Univ.-Comput. Inf. Sci. 2022, 34, 3843–3850. [Google Scholar] [CrossRef]
- Roy, S.K.; Krishna, G.; Dubey, S.R.; Chaudhuri, B.B. HybridSN: Exploring 3-D–2-D CNN feature hierarchy for hyperspectral image classification. IEEE Geosci. Remote Sens. Lett. 2019, 17, 277–281. [Google Scholar] [CrossRef]
- Wang, Y.; Ning, D.; Feng, S. A novel capsule network based on wide convolution and multi-scale convolution for fault diagnosis. Appl. Sci. 2020, 10, 3659. [Google Scholar] [CrossRef]
- Aydemir, M.S.; Bilgin, G. Semisupervised hyperspectral image classification using deep features. IEEE J. Sel. Top. Appl. Earth Obs. Remote. Sens. 2019, 12, 3615–3622. [Google Scholar] [CrossRef]
- Kumar, D.; Kumar, D. Hyperspectral Image Classification Using Deep Learning Models: A Review. J. Phys. Conf. Ser. 2021, 1950, 012087. [Google Scholar] [CrossRef]
- Devassy, B.M.; George, S. Dimensionality reduction and visualisation of hyperspectral ink data using t-SNE. Forensic Sci. Int. 2020, 311, 110194. [Google Scholar] [CrossRef]
- Luo, B.; Hussain, A.; Mahmud, M.; Tang, J. Advances in brain-inspired cognitive systems. Cogn. Comput. 2016, 8, 795–796. [Google Scholar]
- Fırat, H.; Asker, M.E.; Hanbay, D. Classification of hyperspectral remote sensing images using different dimension reduction methods with 3D/2D CNN. Remote. Sens. Appl. Soc. Environ. 2022, 25, 100694. [Google Scholar] [CrossRef]
- Ladi, S.K.; Panda, G.; Dash, R.; Ladi, P.K. A Pioneering Approach of Hyperspectral Image Classification Employing the Cooperative Efforts of 3D, 2D and Depthwise Separable-1D Convolutions. In Proceedings of the 2022 IEEE 2nd International Symposium on Sustainable Energy, Signal Processing and Cyber Security (iSSSC), Gunupur, India, 15–17 December 2022; pp. 1–6. [Google Scholar]
- Nayak, O.; Khandare, H.; Parida, N.K.; Giri, R.; Janghel, R.R.; Govil, H. Hyperspectral Image Classification using Hybrid Deep Convolutional Neural Network. Proc. J. Phys. Conf. Ser. 2022, 2273, 012028. [Google Scholar] [CrossRef]
- Butt, M.H.F.; Ayaz, H.; Ahmad, M.; Li, J.P.; Kuleev, R. A Fast and Compact Hybrid CNN for Hyperspectral Imaging-based Bloodstain Classification. In Proceedings of the 2022 IEEE Congress on Evolutionary Computation (CEC), Padua, Italy, 18–23 July 2022; pp. 1–8. [Google Scholar]
- Liu, S.; Shi, Q.; Zhang, L. Few-Shot Hyperspectral Image Classification with Unknown Classes Using Multitask Deep Learning. IEEE Trans. Geosci. Remote Sens. 2021, 59, 5085–5102. [Google Scholar] [CrossRef]
- Yang, X.; Ye, Y.; Li, X.; Lau, R.Y.K.; Zhang, X.; Huang, X. Hyperspectral Image Classification with Deep Learning Models. IEEE Trans. Geosci. Remote Sens. 2018, 56, 5408–5423. [Google Scholar] [CrossRef]
- Hossain, M.M.; Hossain, M.A. Feature Reduction and Classification of Hyperspectral Image Based on Multiple Kernel PCA and Deep Learning. In Proceedings of the 2019 IEEE International Conference on Robotics, Automation, Artificial-intelligence and Internet-of-Things (RAAICON), Dhaka, Bangladesh, 29 November–1 December 2019; pp. 141–144. [Google Scholar] [CrossRef]
- Hossain, M.M.; Hossain, M.A.; Al Mamun, M.; Hossain, M.M. Feature Reduction Based on the Fusion of Spectral and Spatial Transformation for Hyperspectral Image Classification. In Proceedings of the 2020 IEEE Region 10 Symposium (TENSYMP), Dhaka, Bangladesh, 5–7 June 2020; pp. 150–153. [Google Scholar] [CrossRef]
- Hughes, G. On the mean accuracy of statistical pattern recognizers. IEEE Trans. Inf. Theory 1968, 14, 55–63. [Google Scholar] [CrossRef]
- Jia, X.; Kuo, B.; Crawford, M.M. Feature Mining for Hyperspectral Image Classification. Proc. IEEE 2013, 101, 676–697. [Google Scholar] [CrossRef]
- Bartholomew, D.J. Principal Components Analysis. In International Encyclopedia of Education, 3rd ed.; Peterson, P., Baker, E., McGaw, B., Eds.; Elsevier: Oxford, UK, 2010; pp. 374–377. [Google Scholar] [CrossRef]
- Kabir, M.H.; Mahmood, S.; Al Shiam, A.; Musa Miah, A.S.; Shin, J.; Molla, M.K.I. Investigating Feature Selection Techniques to Enhance the Performance of EEG-Based Motor Imagery Tasks Classification. Mathematics 2023, 11, 1921. [Google Scholar] [CrossRef]
- García-Alonso, C.R.; Pérez-Naranjo, L.M.; Fernández-Caballero, J.C. Multiobjective evolutionary algorithms to identify highly autocorrelated areas: The case of spatial distribution in financially compromised farms. Ann. Oper. Res. 2014, 219, 187–202. [Google Scholar] [CrossRef]
- Gisbrecht, A.; Schulz, A.; Hammer, B. Parametric nonlinear dimensionality reduction using kernel t-SNE. Neurocomputing 2015, 147, 71–82. [Google Scholar] [CrossRef]
- Kingman, J.F.C. Information Theory and Statistics. By Solomon Kullback. Pp. 399. 28s. 6d. 1968. (Dover.). Math. Gaz. 1970, 54, 90. [Google Scholar] [CrossRef]
- Li, Y.; Zhang, H.; Shen, Q. Spectral-Spatial Classification of Hyperspectral Imagery with 3D Convolutional Neural Network. Remote Sens. 2017, 9, 67. [Google Scholar] [CrossRef]
- Miao, S.; Wang, J.Z.; Liao, R. Chapter 12-Convolutional Neural Networks for Robust and Real-Time 2-D/3-D Registration. In Deep Learning for Medical Image Analysis; Zhou, S.K., Greenspan, H., Shen, D., Eds.; Academic Press: Cambridge, MA, USA, 2017; pp. 271–296. [Google Scholar] [CrossRef]
- Zhang, Z. Improved Adam Optimizer for Deep Neural Networks. In Proceedings of the 2018 IEEE/ACM 26th International Symposium on Quality of Service (IWQoS), Banff, AB, Canada, 4–6 June 2018; pp. 1–2. [Google Scholar] [CrossRef]
- Abadi, M.; Agarwal, A.; Barham, P.; Brevdo, E.; Chen, Z.; Citro, C.; Corrado, G.S.; Davis, A.; Dean, J.; Devin, M.; et al. Tensorflow: Large-scale machine learning on heterogeneous distributed systems. arXiv 2016, arXiv:1603.04467. [Google Scholar]
- Tock, K. Google CoLaboratory as a platform for Python coding with students. Rtsre Proc. 2019, 2–1, 1–13. [Google Scholar]
- Gollapudi, S. OpenCV with Python. In Learn Computer Vision Using OpenCV; Springer: Berlin/Heidelberg, Germany, 2019; pp. 31–50. [Google Scholar]
- Glorot, X.; Bengio, Y. Understanding the difficulty of training deep feedforward neural networks. In Proceedings of the Thirteenth International Conference on Artificial Intelligence and Statistics, Sardinia, Italy, 13–15 May 2010; pp. 249–256. [Google Scholar]
- Dozat, T. Incorporating Nesterov momentum into Adam. In Proceedings of the Workshop given at International Conference on Learning Representation, San Juan, Puerto Rico, 2–4 May 2016. [Google Scholar]
- Melgani, F.; Bruzzone, L. Classification of hyperspectral remote sensing images with support vector machines. IEEE Trans. Geosci. Remote Sens. 2004, 42, 1778–1790. [Google Scholar] [CrossRef]
- Makantasis, K.; Karantzalos, K.; Doulamis, A.; Doulamis, N. Deep supervised learning for hyperspectral data classification through convolutional neural networks. In Proceedings of the 2015 IEEE International Geoscience and Remote Sensing Symposium (IGARSS), Milan, Italy, 26–31 July 2015; pp. 4959–4962. [Google Scholar]
- Hamida, A.B.; Benoit, A.; Lambert, P.; Amar, C.B. 3-D deep learning approach for remote sensing image classification. IEEE Trans. Geosci. Remote Sens. 2018, 56, 4420–4434. [Google Scholar] [CrossRef]






| Sensors | Organization | No. of Bands | Wavelength Range (m) |
|---|---|---|---|
| AVIRIS | NASA | 224 | 0.40–2.50 |
| AISA | Spectral Imaging Ltd. | 286 | 0.45–0.90 |
| CASI | Itres Research | 288 | 0.43–0.87 |
| PROBE-1 | Earth Search Science Inc. | 128 | 0.40–2.45 |
| Layer (Type) | Output Shape | Parameter # |
|---|---|---|
| Input Layer | (None, 224, 224, 2, 1) | 0 |
| Conv3d Layer | (None, 25, 25, 2, 1) | 0 |
| Conv3d Layer | (None, 23, 23, 1, 8) | 152 |
| Conv3d Layer | (None, 21, 21, 1, 16) | 1168 |
| Conv2D Layer | (None, 19, 19, 64) | 9280 |
| Flatten Layer | (None, 23,104) | 0 |
| FCN Layer | (None, 256) | 5,914,880 |
| Dropout Layer | (None, 256) | 0 |
| Dense Layer | (None, 128) | 32,896 |
| Dropout Layer | (None, 128) | 0 |
| Output Layer | (None, 21, 21, 1, 16) |
| Recall | Precision | F1 Score | Test Loss | TA | KA | OA | AA | Time | |
|---|---|---|---|---|---|---|---|---|---|
| SVD [43] | 6 | 0 | 4 | 98 | 3.9 | 0 | 3.9 | 6.25 | n/a |
| ICA [43] | 85 | 92 | 94 | 27.23 | 94.16 | 93.37 | 94.2 | 85.12 | n/a |
| NMF [43] | 93 | 89 | 94 | 42.62 | 94.25 | 93.44 | 94.25 | 92.89 | n/a |
| KPCA (Cosine) [43] | 94 | 94 | 94 | 37.82 | 93.97 | 93.13 | 93.97 | 90.69 | n/a |
| KPCA (RBF) [43] | 85 | 87 | 85 | 96.82 | 85.49 | 83.35 | 85.49 | 81.36 | n/a |
| MKPCA (Cosine + Linear) [43] | 96 | 96 | 96 | 23.95 | 95.75 | 95.16 | 95.76 | 89.96 | n/a |
| MKPCA (Cosine + RBF) [43] | 95 | 95 | 94 | 23.75 | 94.53 | 93.75 | 94.53 | 90.73 | n/a |
| SVM [60] | n/a | n/a | n/a | n/a | n/a | 85.3 | 83.10 | 79.03 | n/a |
| 2D-CNN [61] | n/a | n/a | n/a | n/a | n/a | 89.48 | 87.96 | 86.14 | 1.3 |
| 3D-CNN [62] | n/a | n/a | n/a | n/a | n/a | 91.10 | 89.98 | 91.58 | 10.6 |
| Proposed | 95 | 99 | 97 | 6.2 | 98.21 | 98.10 | 98.34 | 95.21 | 4.42 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hossain, M.M.; Hossain, M.A.; Musa Miah, A.S.; Okuyama, Y.; Tomioka, Y.; Shin, J. Stochastic Neighbor Embedding Feature-Based Hyperspectral Image Classification Using 3D Convolutional Neural Network. Electronics 2023, 12, 2082. https://doi.org/10.3390/electronics12092082
Hossain MM, Hossain MA, Musa Miah AS, Okuyama Y, Tomioka Y, Shin J. Stochastic Neighbor Embedding Feature-Based Hyperspectral Image Classification Using 3D Convolutional Neural Network. Electronics. 2023; 12(9):2082. https://doi.org/10.3390/electronics12092082
Chicago/Turabian StyleHossain, Md. Moazzem, Md. Ali Hossain, Abu Saleh Musa Miah, Yuichi Okuyama, Yoichi Tomioka, and Jungpil Shin. 2023. "Stochastic Neighbor Embedding Feature-Based Hyperspectral Image Classification Using 3D Convolutional Neural Network" Electronics 12, no. 9: 2082. https://doi.org/10.3390/electronics12092082
APA StyleHossain, M. M., Hossain, M. A., Musa Miah, A. S., Okuyama, Y., Tomioka, Y., & Shin, J. (2023). Stochastic Neighbor Embedding Feature-Based Hyperspectral Image Classification Using 3D Convolutional Neural Network. Electronics, 12(9), 2082. https://doi.org/10.3390/electronics12092082









