Computational Model Informs Effective Control Interventions against Y. enterocolitica Co-Infection
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
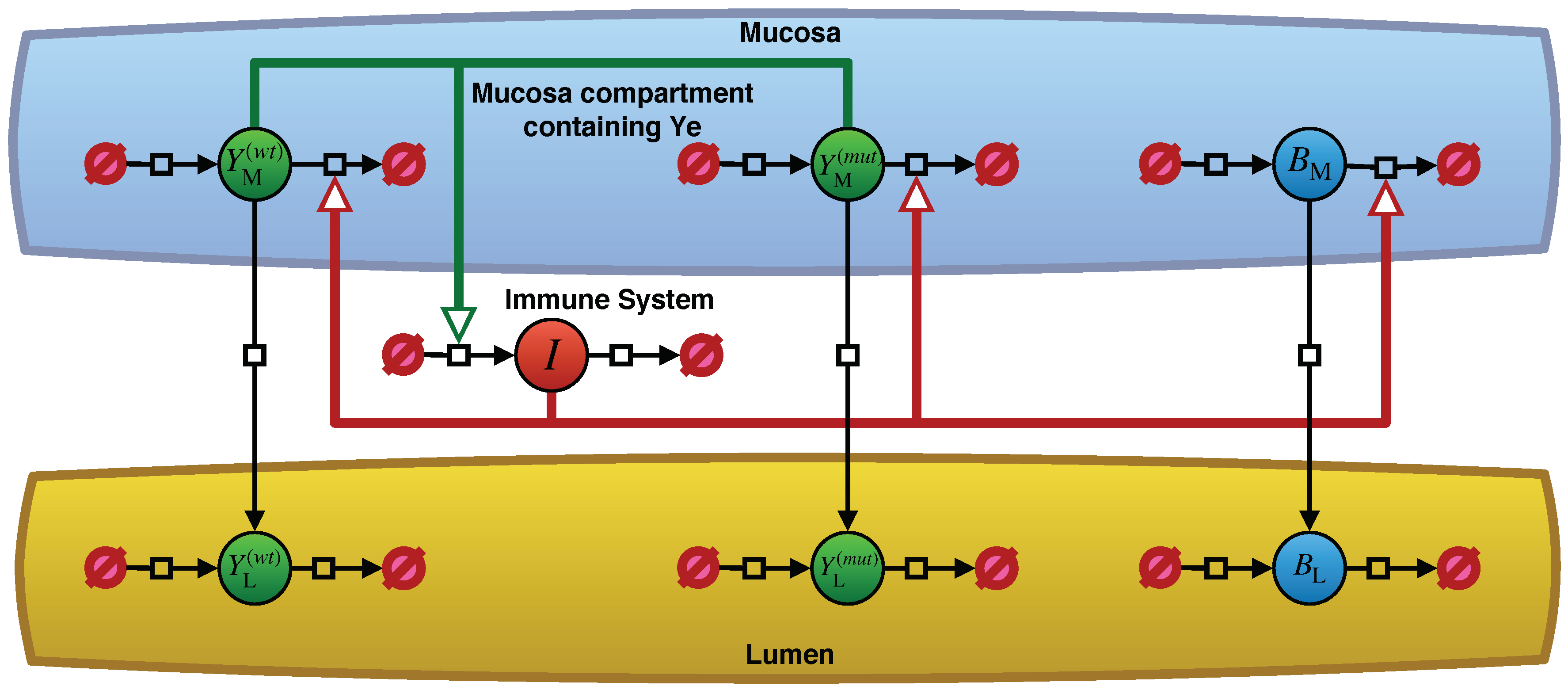
2.1. Model Description
2.2. The Basic Reproduction Number
- A(1)
- (no immigration of individuals into the disease compartments)
- A(2)
- (the new infections will be represented by , so it cannot be negative)
- A(3)
- (if the compartment is empty, it can only have inflow, and the net outflow from the compartment must be negative)
- A(4)
- (sum is net outflow)
- A(5)
- The system has a unique asymptotically stable equilibrium, (all solutions with initial conditions of the form approach a point )
2.3. Center Manifold
- H(1)
- In the balance equations for the infected compartments, nonlinear terms are present only in the rate of the appearance of new infections;
- H(2)
- Nonlinear terms are bilinear;
- H(3)
- There is no linear transfer from infected to uninfected compartments.
3. Results
3.1. Existence of Equilibria
- The trivial equilibrium point is as an origin equilibrium . This solution appears when all populations are extinct. For all parameters, this point never becomes stable due to the positivity of eigenvalues in (A2).
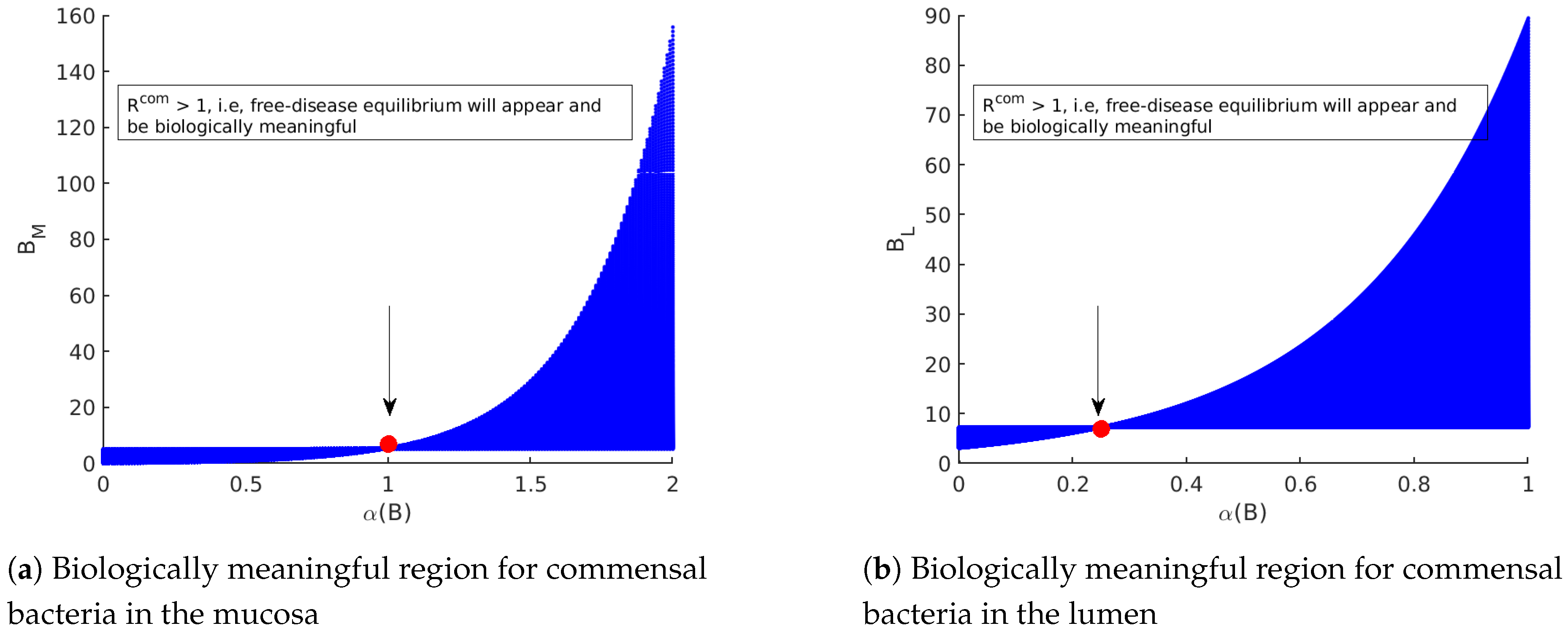
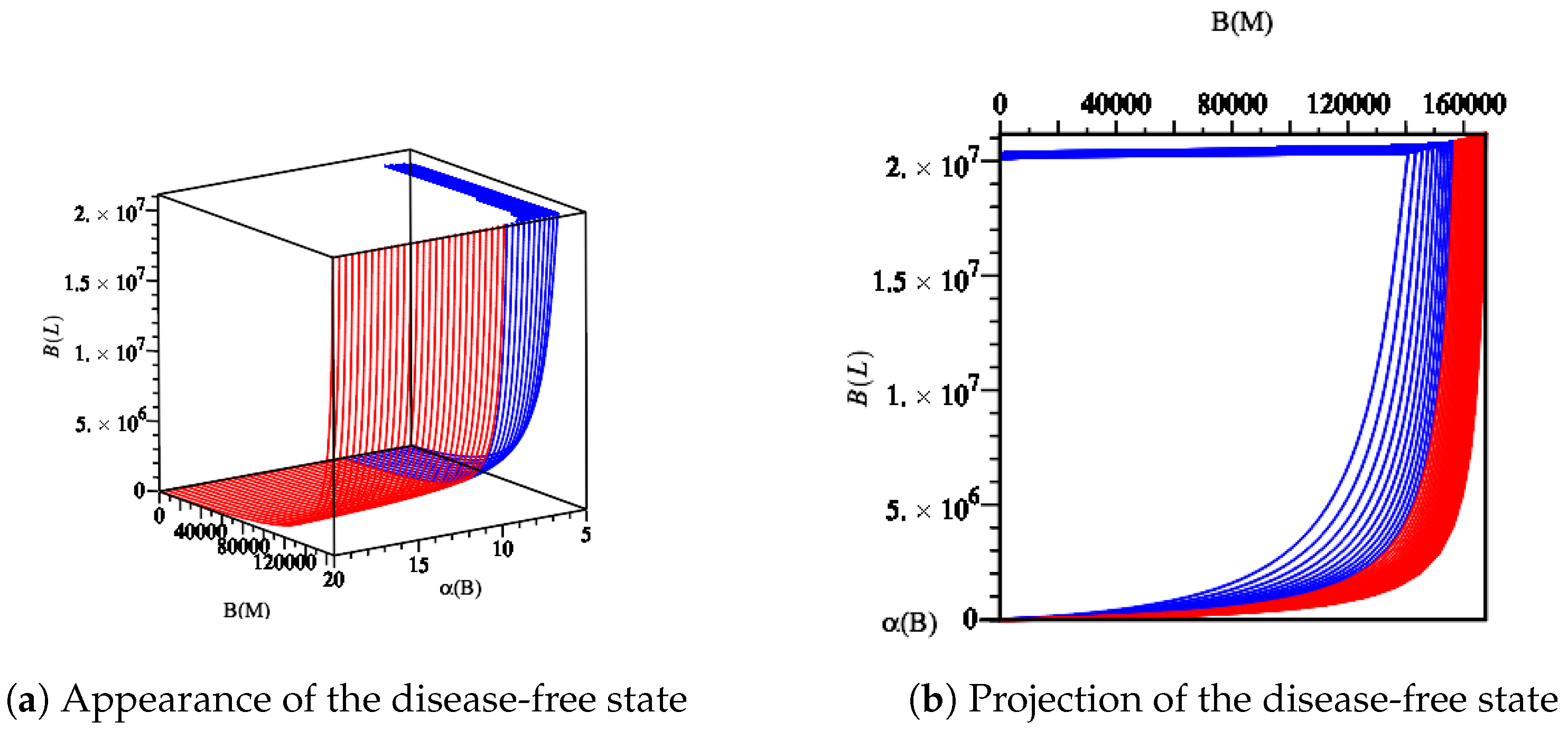
- The first equilibrium point appears in the absence of Yersinia . System (1)–(7) has a disease-free equilibrium, which is given byandIt describes a disease-free state whereby only the commensal bacteria persist. In order for the disease-free state to be biologically meaningful, the conditions and must hold. These conditions correspond to the maximal growth rate of intestinal bacteria exceeding the rate at which intestines are charged and the maximal immunity action, which is not that strong in the absence of Yersinia strains. However, the population of the immune system is at its maximum carrying capacity (in health, not in fighting with any infection).
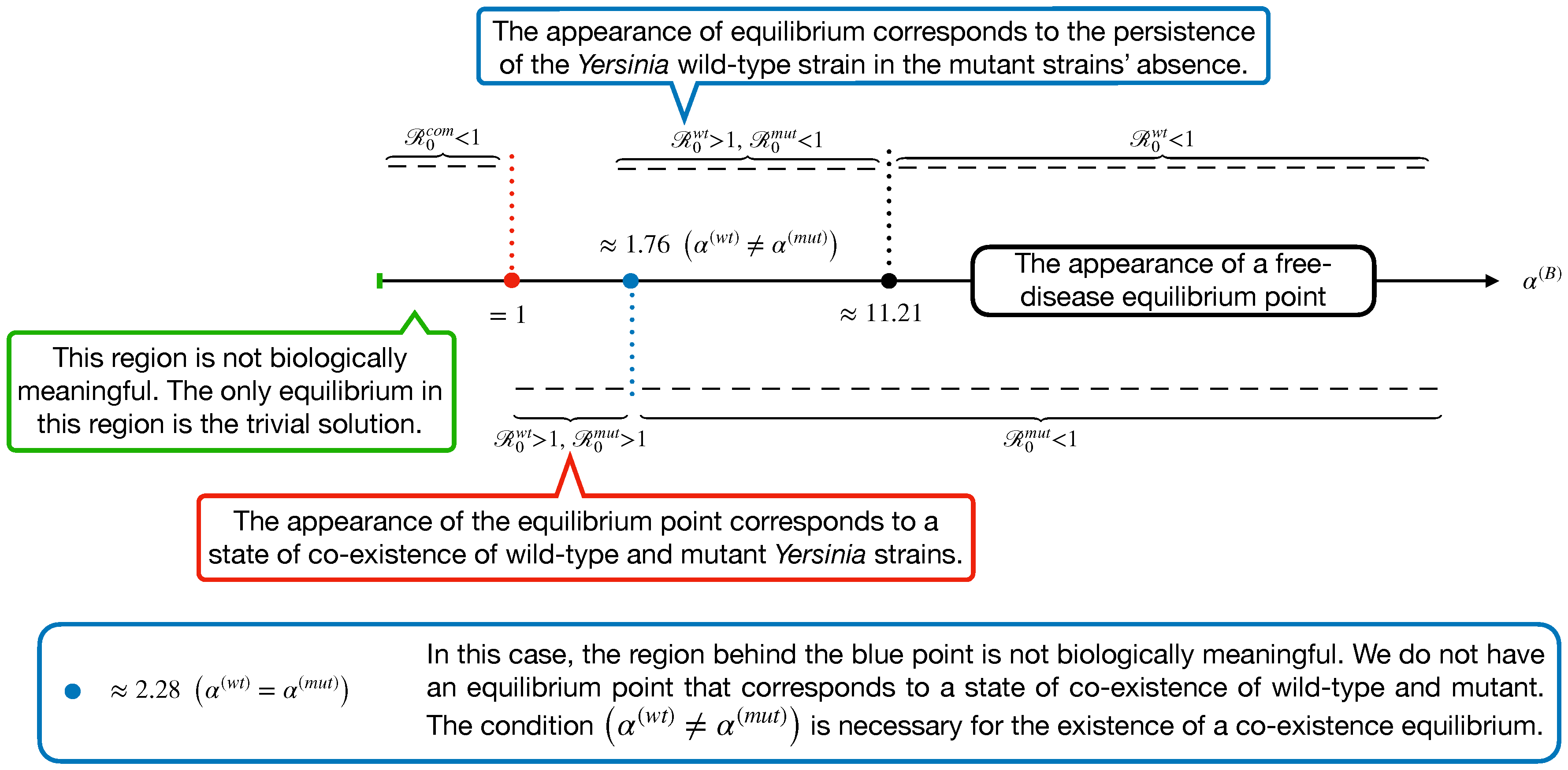
- A second equilibrium corresponds to the commensal bacteria’s persistence and the Yersinia mut strain in the absence of the wt strain. Without loss of generality, the commensal bacteria are supposed to be zero because they are not infective. This point is obtained by setting :with
- The other equilibrium corresponds to the persistence of commensal bacteria and the Yersinia wt strain in the absence of the mut strain. Without loss of generality, the commensal bacteria are supposed to be zero because they are not infective. This point is obtained by setting :with
- Finally, the last equilibrium point corresponds to a state of the co-existence of wt and mut Yersinia strains. This point is achieved by supposing :withwhere Z is defined as to make the equilibrium point biologically meaningful.
3.2. Analysis of the Disease-Free Equilibrium Point
3.3. Computing and Analysis of the System through the Basic Reproduction Number
- : the basic reproductive numbers for wt strain =
- : the basic reproductive numbers for mut strain = .
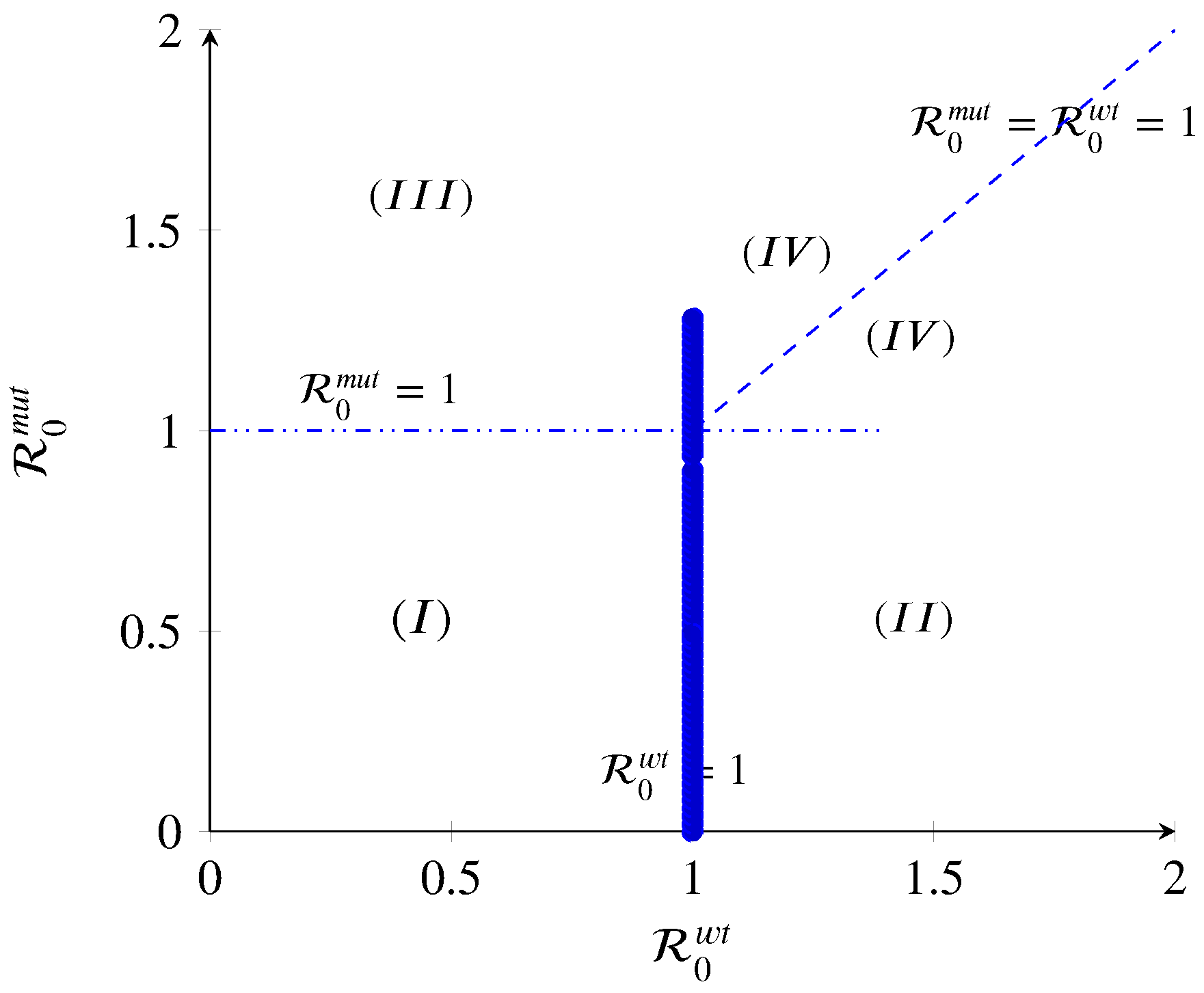
- (I)
- If and , then and . Thus, the intersection of the transcritical curves and results in a triple transcritical bifurcation. As shown in (A9), the Jacobian has a triple zero eigenvalue at this point (). Kuznetsov [34] has proved that such a point would be an indicator of the onset of a non-degenerate or degenerate Bogdanov–Takens bifurcation [34,35]. The disease-free equilibrium loses its stability, and the wt-free and mut-free include one simple zero eigenvalue (), meaning that the dynamics of the model change as the target parameter is within the threshold value.
- (II)
- If , the wt strain equilibrium in region II will persist when . The wt strain will spread and possibly persist within the host population. In general, for a strain to persist, its basic reproduction number has to be strictly greater than one. Therefore, in this region, the disease-free, mut strain, and co-existence state exchange stability: becomes unstable, becomes locally asymptotically stable, and and remain unstable. This means that the immune system could kill one of the strains more efficiently.
- (III)
- If , the mut strain equilibrium in region III will persist when . The mut strain will spread and possibly persist within the host population since its basic reproduction number is greater than one. Therefore, in this region, the disease-free, wt strain, and co-existence state exchange stability: becomes unstable, becomes locally asymptotically stable, and and remain unstable. This means that the immune system could defeat the wt strains. However, the risk of this situation to happen is low because the mut strains are influenced more efficiently than wt strains by immune action.
- (IV)
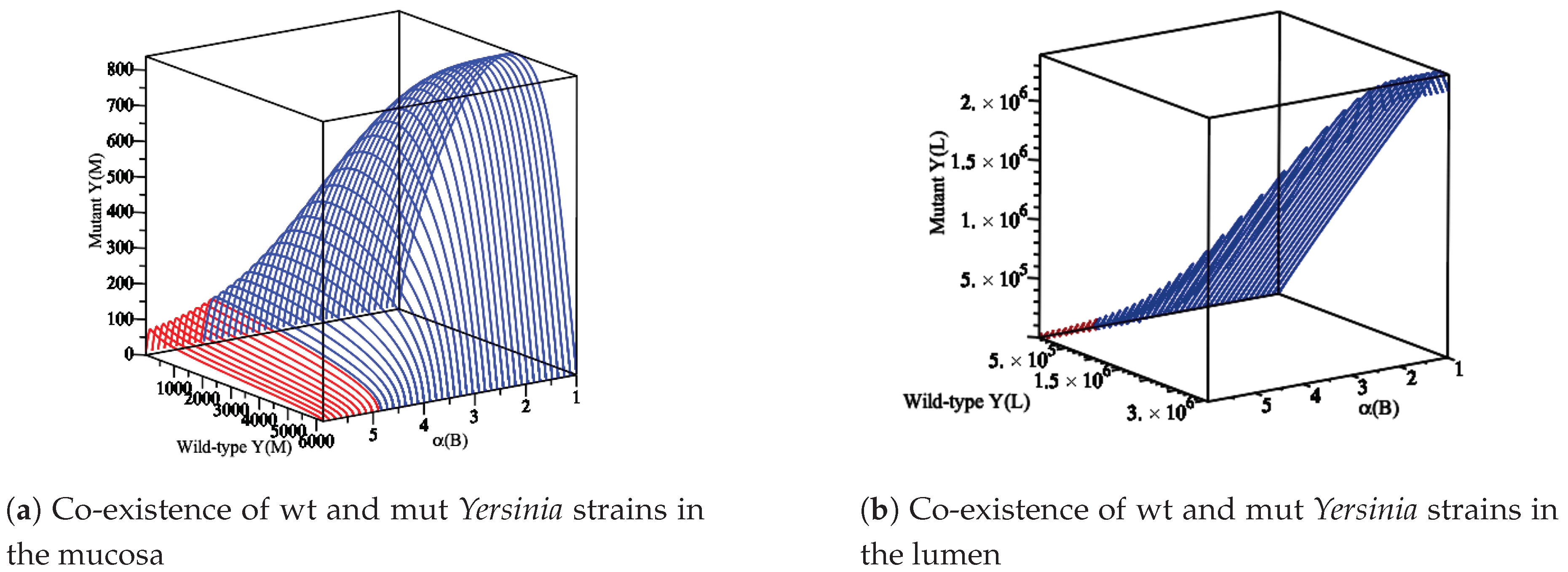
- If and , the co-existence population spreads, and both strains persist. The overall can be defined as . A mut with is thoroughly dominant, while one with is completely recessive; scenarios of incomplete dominance , under-dominance , and over-dominance are possible as well. For instance, a mut could achieve a higher than the wt via a higher growth rate that increases transmission. In a co-infection, the faster-growing mut strain would outcompete the wt and reach its maximum capacity. This situation would change the co-infection to the conditions where a single infection happens. Thus, the overall of the co-infection would be similar to that of the mut by itself, making the mut a dominant one. Furthermore, the effort for having a co-existence equilibrium and analysis of the co-infection model will fail. By contrast, let us assume a mut strain achieves a higher . Nevertheless, the virulence of the wt strain neutralizes the higher value of the mut. This would make the mut a recessive one. In summary, virtually any two-strain co-infection model can be mapped to a set of values for d, allowing scenarios of particular interest to be explored in a context broader than the one possible with typical models.
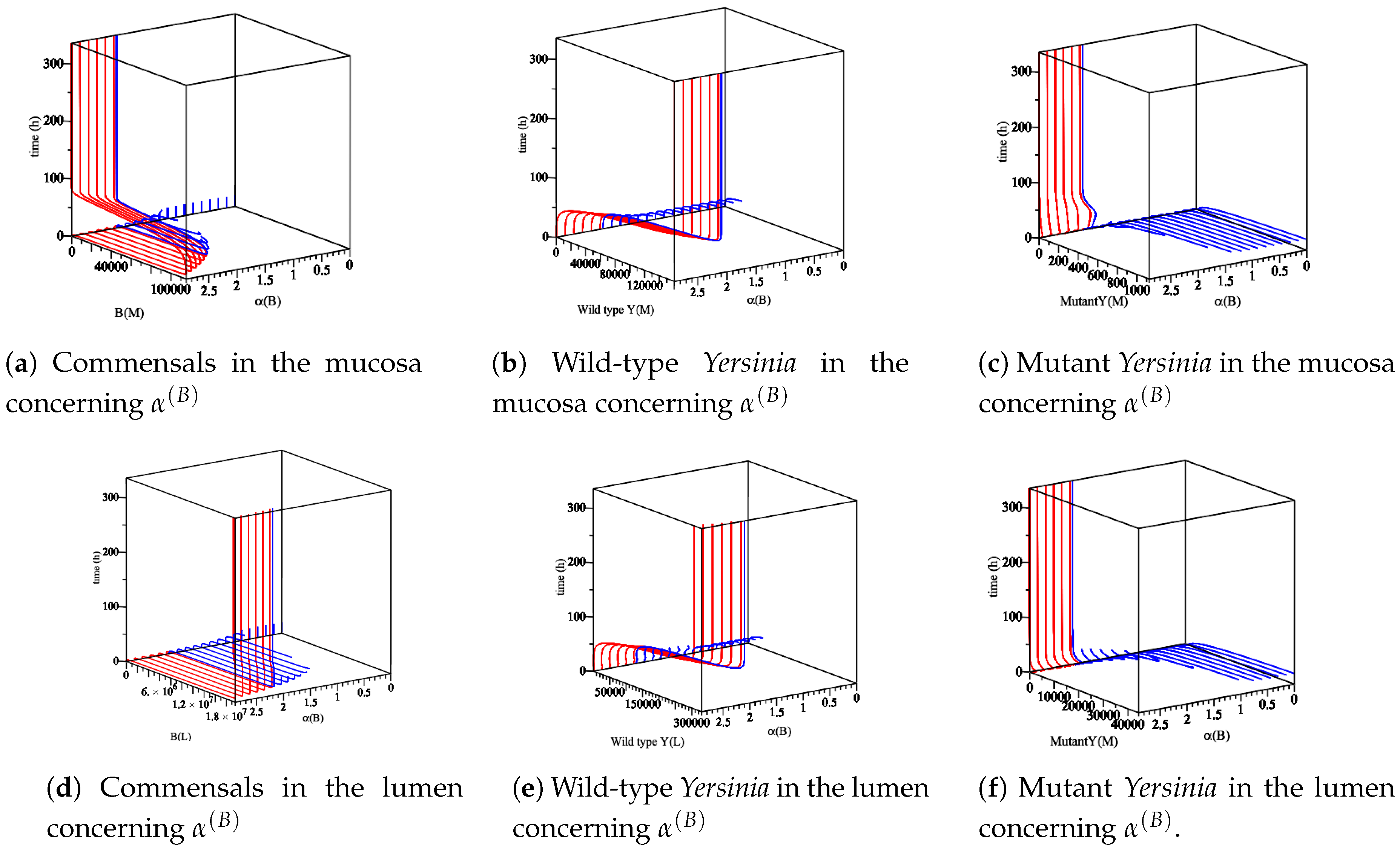
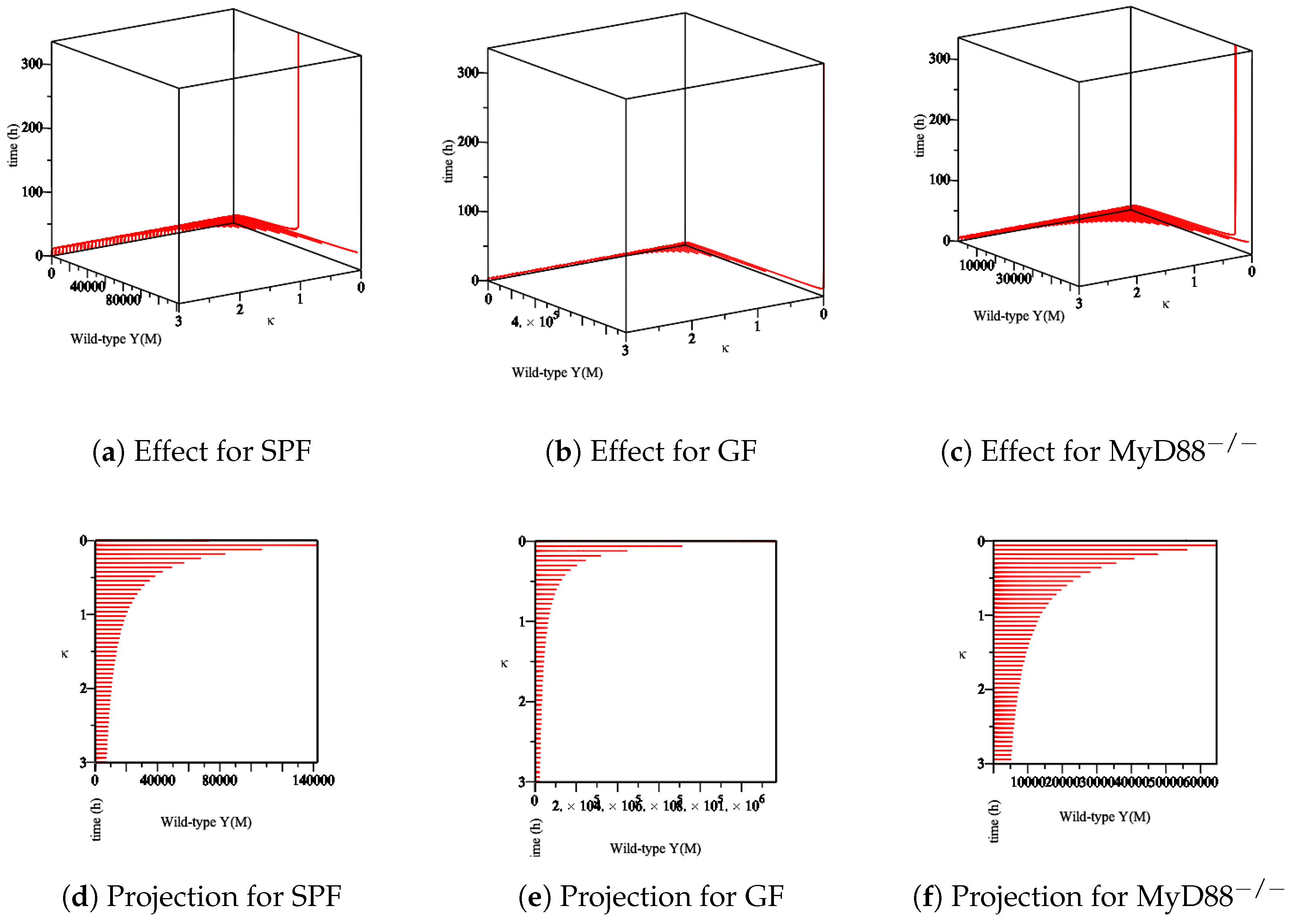
- The effect of the maximum rate of immune growth on wt Yersinia strain in the mucosa, Figure 7;
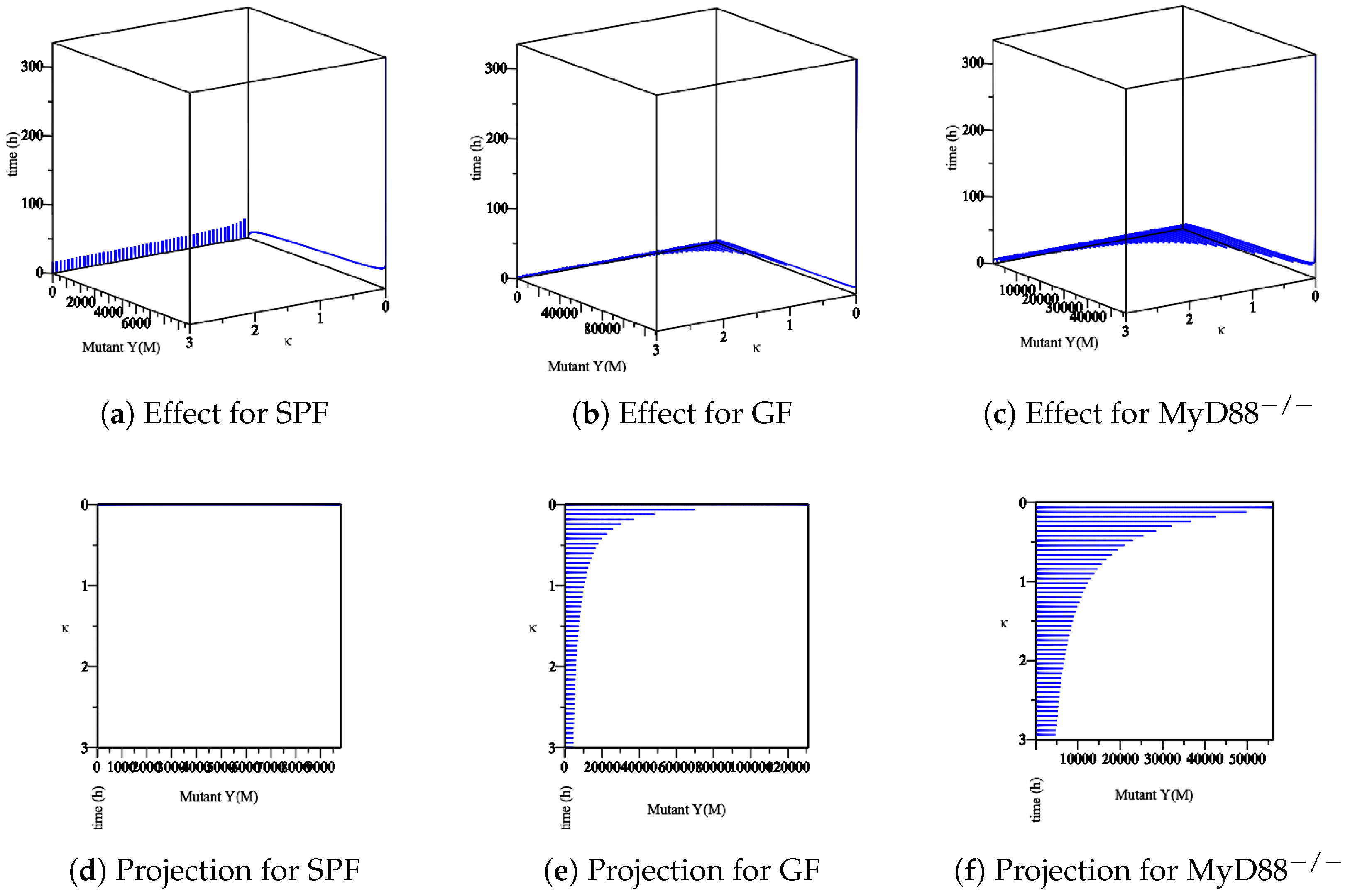
- The effect of the maximum rate of immune growth on mut Yersinia strain in the mucosa, Figure 8;
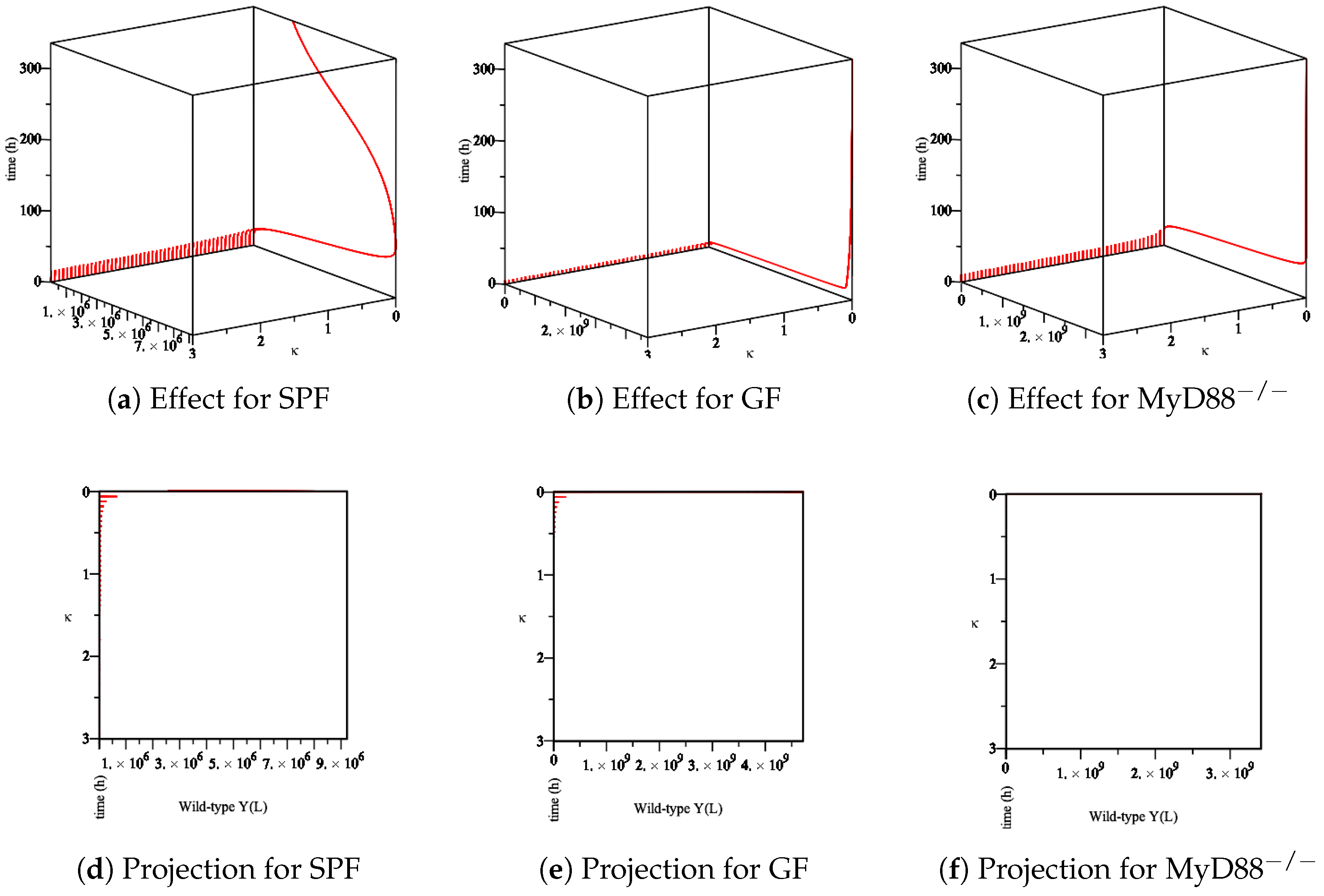
- The effect of the maximum rate of immune growth on wt Yersinia strain in the lumen, Figure 9;
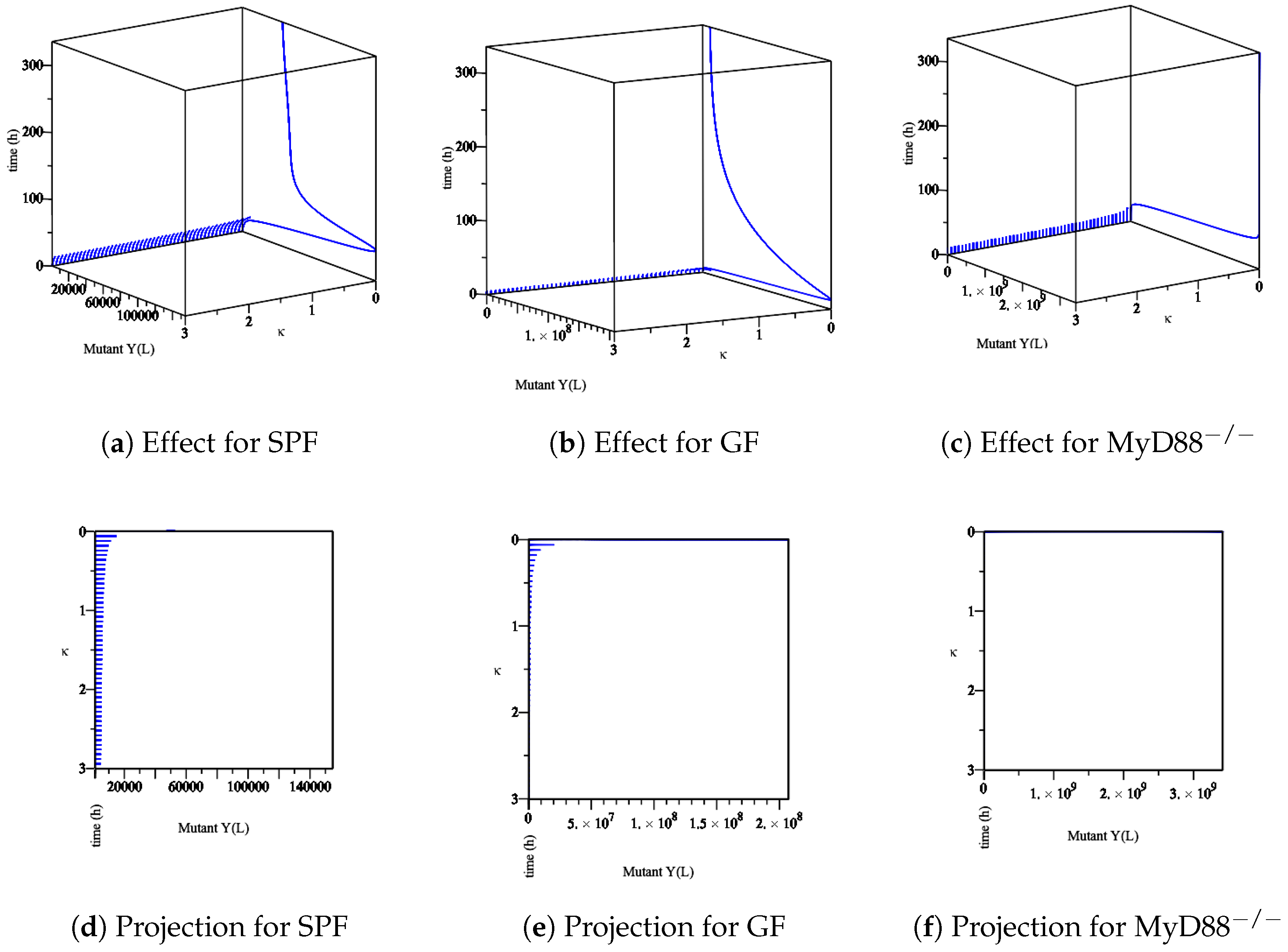
- The effect of the maximum rate of immune growth on mut Yersinia strain in the lumen, Figure 10,
4. Discussion and Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| AMP | antimicrobial peptide |
| BMBF | Federal Ministry of Education and Research |
| BMBF-DZG | Deutsche Zentren der Gesundheitsforschung |
| CFU | colony-forming unit |
| DFE | disease-free equilibrium |
| DFG | Deutsche Forschungsgemeinschaft |
| DZIF | German Center for Infection Research |
| GF | germ-free |
| GIT | gastrointestinal tract |
| mut | mutant |
| MyD88−/− | MyD88-deficient mice |
| ODE | Ordinary Differential Equation |
| PD | Process Description |
| RKI | Robert Koch Institute |
| SBGN | Systems Biology Graphical Notation |
| SI | small intestine |
| SPF | specific-pathogen-free |
| T3SS | Type iii@ secretion system |
| wt | wild-type |
| YadA | Yersinia adhesin A |
| Ye | Yersinia entercolitica |
| Yop | Yersinia outer protein |
Appendix A. Mathematical Calculation
Appendix A.1. Jacobian
Appendix A.2. The Eigenvalues of Trivial Equilibrium Point
Appendix A.3. The Eigenvalues and Eigenvectors of the Disease-Free Equilibrium point
Appendix A.4. The Eigenvalues wt Strain Equilibrium
Appendix A.5. The Eigenvalues mut Strain Equilibrium
Appendix A.6. The Basic Reproduction Numbers
- corresponds to the rate of the appearance of new infections, and corresponds to the rate of transfer assumed asand
- The basic reproduction number is achieved as the spectral radius of the matrix ;as where , , and are as follows:andThen
Appendix B. Data Availability
- The Matlab script
- The Maple workbook
- The described model is available in SBML format [26] (Level 3 Version 2 [27]) from BioModels database [36] under model identifier MODEL2002070001.
References
- El Tahir, Y.; Skurnik, M. YadA, the multifaceted Yersinia Adhes. Int. J. Med. Microbiol. 2001, 291, 209–218. [Google Scholar] [CrossRef] [PubMed]
- Handley, A.J.; Koster, R.; Monsieurs, K.; Perkins, G.D.; Davies, S.; Bossaert, L. European Resuscitation Council Guidelines for Resuscitation 2005: Section 2. Adult basic life support and use of automated external defibrillators. Resuscitation 2005, 67, S7–S23. [Google Scholar] [CrossRef] [PubMed]
- Mühlenkamp, M.; Oberhettinger, P.; Leo, J.C.; Linke, D.; Schütz, M.S. Yersinia Adhes. A (YadA)- Beast. Int. J. Med. Microbiol. 2015, 305, 252–258. [Google Scholar] [CrossRef] [PubMed]
- Young, G.A.; LaVon, G.D.; Taylor, G.W. High Efficiency Absorbent Articles for Incontinence Management. US Patent 5,147,345, 15 September 1992. [Google Scholar]
- Pepe, G.J.; Albrecht, E.D. Actions of placental and fetal adrenal steroid hormones in primate pregnancy. Endocr. Rev. 1995, 16, 608–648. [Google Scholar] [PubMed]
- Cornelis, G.R. Yersinia Type III Secretion: Send Eff. J. Cell Biol. 2002, 158, 401. [Google Scholar] [CrossRef]
- Ruckdeschel, K.; Roggenkamp, A.; Schubert, S.; Heesemann, J. Differential contribution of Yersinia Enterocolitica Virulence Factors Evas. Microbicidal Action Neutrophils. Infect. Immun. 1996, 64, 724–733. [Google Scholar] [CrossRef]
- Lupp, C.; Robertson, M.L.; Wickham, M.E.; Sekirov, I.; Champion, O.L.; Gaynor, E.C.; Finlay, B.B. Host-mediated inflammation disrupts the intestinal microbiota and promotes the overgrowth of Enterobacteriaceae. Cell Host Microbe 2007, 2, 119–129. [Google Scholar] [CrossRef]
- Stecher, B.; Robbiani, R.; Walker, A.W.; Westendorf, A.M.; Barthel, M.; Kremer, M.; Chaffron, S.; Macpherson, A.J.; Buer, J.; Parkhill, J.; et al. Salmonella enterica serovar typhimurium exploits inflammation to compete with the intestinal microbiota. PLoS Biol. 2007, 5, e244. [Google Scholar] [CrossRef]
- Geißert, J.; Bohn, E.; Mostolizadeh, R.; Dräger, A.; Autenrieth, I.; Beier, S.; Deusch, O.; Eichner, M.; Schütz, M. Model-based prediction of bacterial population dynamics in gastrointestinal infection. bioRxiv 2020. [Google Scholar] [CrossRef]
- Nowak, M.; Robert, M. May. Superinfection and the evolution of parasite virulence. Proc. R. Soc. B 1994, 255, 81n89. [Google Scholar]
- Castillo-Chavez, C.; Velasco-Hernández, J.X. On the relationship between evolution of virulence and host demography. J. Theor. Biol. 1998, 192, 437–444. [Google Scholar] [CrossRef] [PubMed]
- Dye, C.; Williams, B.G.; Espinal, M.A.; Raviglione, M.C. Erasing the world’s slow stain: Strategies to beat multidrug-resistant tuberculosis. Science 2002, 295, 2042–2046. [Google Scholar] [CrossRef] [PubMed]
- Boldin, B.; Geritz, S.A.; Kisdi, É. Superinfections and adaptive dynamics of pathogen virulence revisited: A critical function analysis. Evol. Ecol. Res. 2009, 11, 153–175. [Google Scholar]
- Liu, W.m.; Hethcote, H.W.; Levin, S.A. Dynamical behavior of epidemiological models with nonlinear incidence rates. J. Math. Biol. 1987, 25, 359–380. [Google Scholar] [CrossRef] [PubMed]
- Madden, L.V.; Hughes, G.; Van Den Bosch, F. The Study of Plant Disease Epidemics; Am Phytopath Society: St. Paul, MN, USA, 2007. [Google Scholar]
- Gross, K.L.; Porco, T.C.; Grant, R.M. HIV-1 superinfection and viral diversity. Aids 2004, 18, 1513–1520. [Google Scholar] [CrossRef] [PubMed]
- Nurtay, A.; Hennessy, M.G.; Sardanyés, J.; Alsedà, L.; Elena, S.F. Theoretical conditions for the coexistence of viral strains with differences in phenotypic traits: A bifurcation analysis. R. Soc. Open Sci. 2019, 6, 181179. [Google Scholar] [CrossRef]
- Mostolizadeh, R.; Shamsara, E.; Afsharnezhad, Z. Hopf Bifurcation and Chaos in a model for HTLV-I infection of CD4+ T- cells. Ann. Tiberiu Popoviciu Semin. Funct. Equ. Approx. Convexity Ser. B Math. Interdiscip. Res. 2017, 15, 59–79. [Google Scholar]
- Mostolizadeh, R.; Afsharnezhad, Z. Hopf bifurcation in a model for adult T-cell leukemia. Math. Methods Appl. Sci. 2018, 41, 6210–6225. [Google Scholar] [CrossRef]
- Shamsara, E.; Mostolizadeh, R.; Afsharnezhad, Z. Transcritical bifurcation of an immunosuppressive infection model. Iran. J. Numer. Anal. Optim. 2016, 6, 1–16. [Google Scholar]
- Hethcote, H.W. The mathematics of infectious diseases. SIAM Rev. 2000, 42, 599–653. [Google Scholar] [CrossRef]
- Anderson, R.M.; May, R. Infectious Diseases of Humans; Oxford Science Publication: New York, NY, USA, 1991. [Google Scholar]
- Touré, V.; Dräger, A.; Luna, A.; Dogrusoz, U.; Rougny, A. The Systems Biology Graphical Notation: Current Status and Applications in Systems Medicine. In Systems Medicine; Wolkenhauer, O., Ed.; Academic Press: Oxford, UK, 2020; Volume 3, pp. 372–381. [Google Scholar]
- Rougny, A.; Touré, V.; Moodie, S.; Balaur, I.; Czauderna, T.; Borlinghaus, H.; Dogrusoz, U.; Mazein, A.; Dräger, A.; Blinov, M.L.; et al. Systems Biology Graphical Notation: Process Description language Level 1 Version 2.0. J. Integr. Bioinform. 2019, 16. [Google Scholar] [CrossRef] [PubMed]
- Keating, S.M.; Waltemath, D.; König, M.; Zhang, F.; Dräger, A.; Chaouiya, C.; Bergmann, F.T.; Finney, A.; Gillespie, C.S.; Helikar, T.; et al. SBML Level 3: An extensible format for the exchange and reuse of biological models. Mol. Syst. Biol. 2020, 16, e9110. [Google Scholar] [CrossRef] [PubMed]
- Hucka, M.; Bergmann, F.T.; Chaouiya, C.; Dräger, A.; Hoops, S.; Keating, S.M.; König, M.; Le Novère, N.; Myers, C.J.; Olivier, B.G.; et al. Systems Biology Markup Language (SBML) Level 3 Version 2 Core Release 2. J. Integr. Bioinform. 2019, 16, 1. [Google Scholar] [CrossRef] [PubMed]
- Diekmann, O.; Heesterbeek, J.A.P.; Metz, J.A. On the definition and the computation of the basic reproduction ratio R 0 in models for infectious diseases in heterogeneous populations. J. Math. Biol. 1990, 28, 365–382. [Google Scholar] [CrossRef]
- Van den Driessche, P.; Watmough, J. Reproduction numbers and sub-threshold endemic equilibria for compartmental models of disease transmission. Math. Biosci. 2002, 180, 29–48. [Google Scholar] [CrossRef]
- Van den Driessche, P.; Watmough, J. Further notes on the basic reproduction number. In Mathematical Epidemiology; Springer: Berlin/Heidelberg, Germany, 2008; pp. 159–178. [Google Scholar]
- Buonomo, B. A note on the direction of the transcritical bifurcation in epidemic models. Nonlinear Anal. Model Control 2015, 20, 38–55. [Google Scholar] [CrossRef][Green Version]
- Guckenheimer, J.; Holmes, P. Nonlinear Oscillations, Dynamical Systems, and Bifurcations of Vector Fields; Springer Science & Business Media: Berlin/Heidelberg, Germany, 2013; Volume 42. [Google Scholar]
- Wiggins, S. Introduction to Applied Nonlinear Dynamical System and Chaos; Spring: New York, NY, USA, 1990. [Google Scholar]
- Kuznetsov, Y.A. Elements of Applied Bifurcation Theory; Springer Science & Business Media: Berlin/Heidelberg, Germany, 2013; Volume 112. [Google Scholar]
- Kuznetsov, Y.A. Practical computation of normal forms on center manifolds at degenerate Bogdanov–Takens bifurcations. Int. J. Bifurc. Chaos 2005, 15, 3535–3546. [Google Scholar] [CrossRef]
- Malik-Sheriff, R.S.; Glont, M.; Nguyen, T.V.N.; Tiwari, K.; Roberts, M.G.; Xavier, A.; Vu, M.T.; Men, J.; Maire, M.; Kananathan, S.; et al. BioModels—15 years of sharing computational models in life science. Nucleic Acids Res. 2020, 48, D407–D415. [Google Scholar] [CrossRef]
| Variable Symbol | Meaning | Units |
|---|---|---|
| Commensal bacteria in the mucosa | item | |
| wt Yersinia in the mucosa | item | |
| mut Yersinia in the mucosa | item | |
| Commensal bacteria in the lumen | item | |
| wt Yersinia in the lumen | item | |
| mut Yersinia in the lumen | item | |
| I | Strength of immune reaction | dimensionless |
| Parameter | Definition | Unit |
|---|---|---|
| Maximal growth rate of intestinal bacteria | 1/ | |
| Maximal growth rate of wt Yersinia | 1/ | |
| Maximal growth rate of mut Yersinia | 1/ | |
| Immunity adjustment factor for wt Yersinia | dimensionless | |
| Immunity adjustment factor for mut Yersinia | dimensionless | |
| Carrying capacity of the mucosa | item | |
| Carrying capacity of the lumen | item | |
| Carrying capacity of the immune system | item | |
| Maximal immunity action | 1/ | |
| Maximal rate of immune growth | 1/ | |
| Rate at which intestines are discharged | 1/ |
| Parameter | Values in Ye SPF wt/A0 | Values in Ye SPF wt/T3S0 | Values in Ye GF wt/A0 | Values in Ye MyD88−/− wt/A0 |
|---|---|---|---|---|
| 4.89 × 10−1 | 2.00 | 1.99 | 5.40 × 10−1 | |
| 4.44 × 10−1 | 1.86 | 1.60 | 5.78 × 10−1 | |
| 4.44 × 10−1 | 1.86 | 1.60 | 5.78 × 10−1 | |
| 3.96 × 10−1 | 9.48 × 10−3 | 1.10 × 10−1 | 6.23 × 10−2 | |
| 1.95 × 10−1 | 3.73 × 10−1 | 1.19 × 10−1 | 1.28 × 10−1 | |
| 1.76 × 105 | 6.27 × 103 | 1.3 × 106 | 1.28 × 105 | |
| 2.14 × 107 | 6.13 × 106 | 4.99 × 109 | 9.98 × 109 | |
| 1.00 | 1.00 | 1.00 | 1.00 | |
| 1.00 | 1.00 | 9.97 × 10−1 | 1.00 × 10−1 | |
| 7.83 × 10−1 | 4.28 × 10−1 | 6.50 × 10−1 | 4.37 × 10−1 | |
| 2.50 × 10−1 | 2.50 × 10−1 | 8.33 × 10−2 | 1.82 × 10−1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mostolizadeh, R.; Dräger, A. Computational Model Informs Effective Control Interventions against Y. enterocolitica Co-Infection. Biology 2020, 9, 431. https://doi.org/10.3390/biology9120431
Mostolizadeh R, Dräger A. Computational Model Informs Effective Control Interventions against Y. enterocolitica Co-Infection. Biology. 2020; 9(12):431. https://doi.org/10.3390/biology9120431
Chicago/Turabian StyleMostolizadeh, Reihaneh, and Andreas Dräger. 2020. "Computational Model Informs Effective Control Interventions against Y. enterocolitica Co-Infection" Biology 9, no. 12: 431. https://doi.org/10.3390/biology9120431
APA StyleMostolizadeh, R., & Dräger, A. (2020). Computational Model Informs Effective Control Interventions against Y. enterocolitica Co-Infection. Biology, 9(12), 431. https://doi.org/10.3390/biology9120431






