Oncostatin M Mediates Adipocyte Expression and Secretion of Stromal-Derived Factor 1
Abstract
:1. Introduction
2. Materials and Methods
2.1. Animals and Diets
2.2. 3T3-L1 Adipocyte Culture and Treatment
2.3. Transient Transfection of 3T3-L1 Adipocytes
2.4. Gene Expression Analyses
2.5. Immunoblotting
2.6. ELISA and Cytokine Arrays
2.7. Statistical Analyses
3. Results
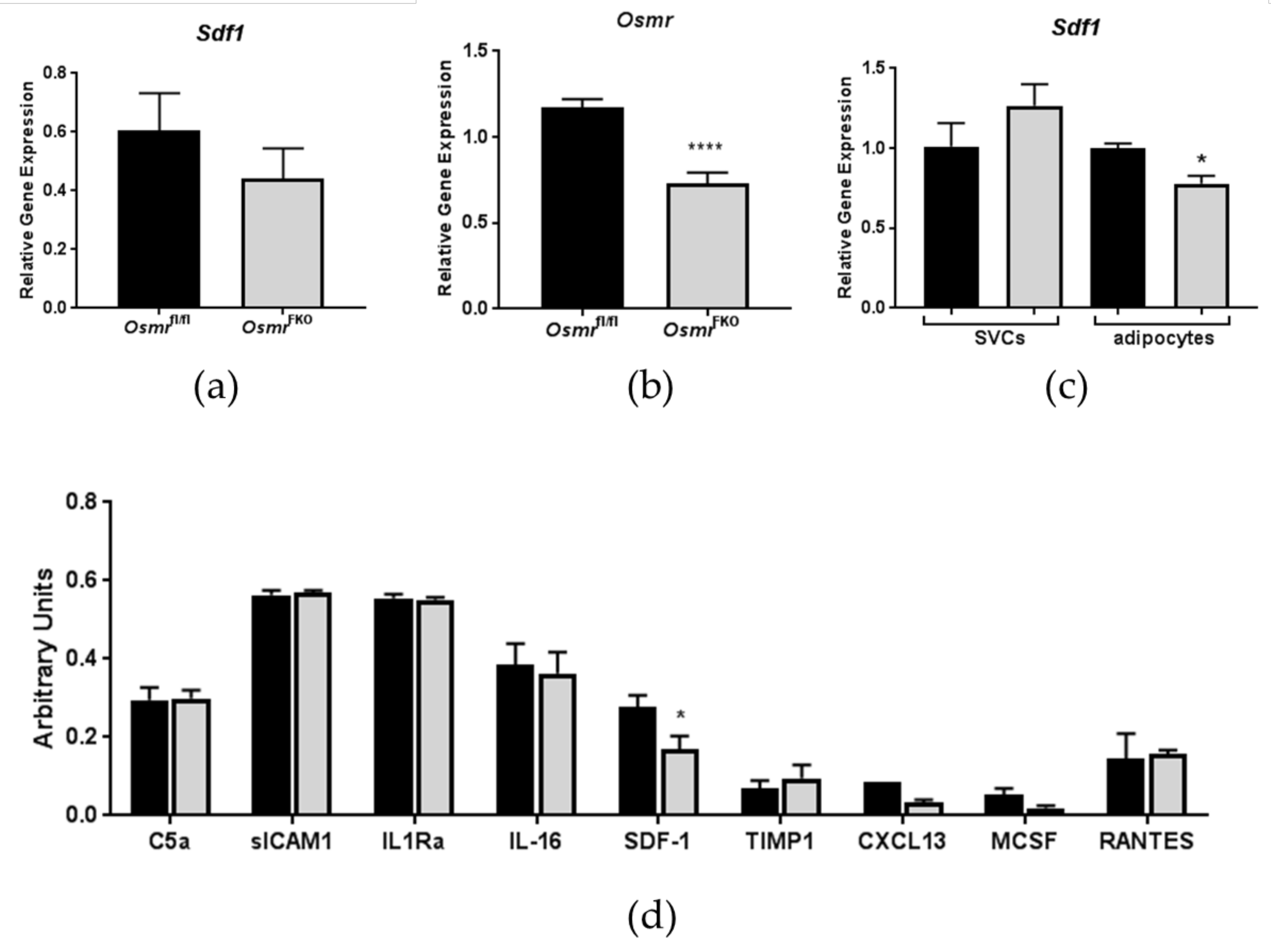
3.1. Effects of In Vivo OSMR Deletion on Adipose Tissue SDF-1 Expression
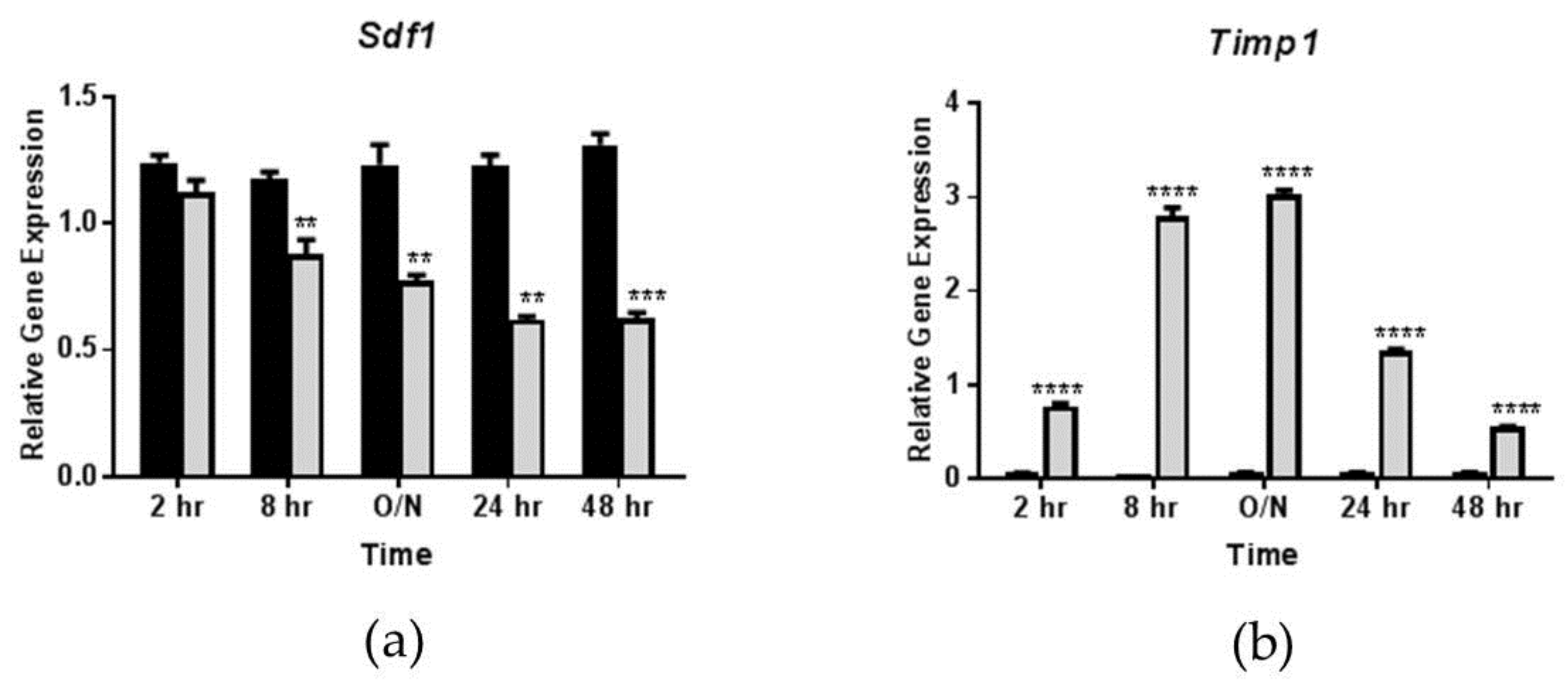
3.2. Effects of OSM Administration on SDF-1 Expression in 3T3-L1 Adipocytes
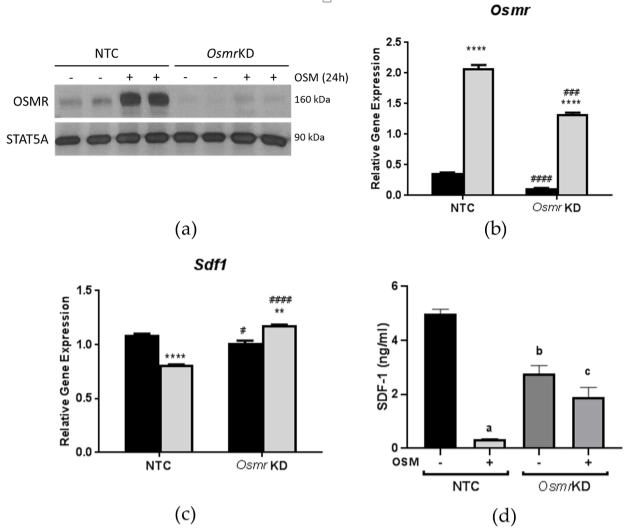
3.3. Effects of OSM Administration on SDF-1 Expression in OSMR-Deficient 3T3-L1 Adipocytes
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Cipolletta, D.; Feuerer, M.; Li, A.; Kamei, N.; Lee, J.; Shoelson, S.E.; Benoist, C.; Mathis, D. PPAR-γ is a major driver of the accumulation and phenotype of adipose tissue Treg cells. Nature 2012, 486, 549. [Google Scholar] [CrossRef] [PubMed]
- Exley, M.A.; Hand, L.; O’Shea, D.; Lynch, L. Interplay between the immune system and adipose tissue in obesity. J. Endocrinol. 2014, 223, R41. [Google Scholar] [CrossRef] [PubMed]
- Grant, R.W.; Dixit, V.D. Adipose tissue as an immunological organ. Obesity 2015, 23, 512–518. [Google Scholar] [CrossRef] [PubMed]
- Macdougall, C.E.; Wood, E.G.; Loschko, J.; Scagliotti, V.; Cassidy, F.C.; Robinson, M.E.; Feldhahn, N.; Castellano, L.; Voisin, M.-B.; Marelli-Berg, F.; et al. Visceral Adipose Tissue Immune Homeostasis Is Regulated by the Crosstalk between Adipocytes and Dendritic Cell Subsets. Cell Metab. 2018, 27, 588–601. [Google Scholar] [CrossRef] [PubMed]
- Mathis, D. Immunological Goings-On in Visceral Adipose Tissue. Cell Metab. 2013, 17, 851–859. [Google Scholar] [CrossRef] [PubMed]
- Maurizi, G.; Della Guardia, L.; Maurizi, A.; Poloni, A. Adipocytes properties and crosstalk with immune system in obesity-related inflammation. J. Cell. Physiol. 2018, 233, 88–97. [Google Scholar] [CrossRef] [PubMed]
- Schipper, H.S.; Prakken, B.; Kalkhoven, E.; Boes, M. Adipose tissue-resident immune cells: Key players in immunometabolism. Trends Endocrinol. Metab. 2012, 23, 407–415. [Google Scholar] [CrossRef]
- Seijkens, T.; Kusters, P.; Chatzigeorgiou, A.; Chavakis, T.; Lutgens, E. Immune Cell Crosstalk in Obesity: A Key Role for Costimulation? Diabetes 2014, 63, 3982–3991. [Google Scholar] [CrossRef] [PubMed]
- Elks, C.M.; Zhao, P.; Grant, R.W.; Hang, H.; Bailey, J.L.; Burk, D.H.; McNulty, M.A.; Mynatt, R.L.; Stephens, J.M. Loss of Oncostatin M Signaling in Adipocytes Induces Insulin Resistance and Adipose Tissue Inflammation in Vivo. J. Biol. Chem. 2016, 291, 17066–17076. [Google Scholar] [CrossRef]
- Stephens, J.M.; Bailey, J.L.; Hang, H.; Rittell, V.; Dietrich, M.A.; Mynatt, R.L.; Elks, C.M. Adipose Tissue Dysfunction Occurs Independently of Obesity in Adipocyte-Specific Oncostatin Receptor Knockout Mice. Obesity 2018, 26, 1439–1447. [Google Scholar] [CrossRef]
- Fasnacht, N.; Müller, W. Conditional gp130 deficient mouse mutants. Semin. Cell Dev. Biol. 2008, 19, 379–384. [Google Scholar] [CrossRef] [PubMed]
- Heinrich, P.C.; Behrmann, I.; Haan, S.; Hermanns, H.M.; Muller-Newen, G.; Schaper, F. Principles of interleukin (IL)-6-type cytokine signalling and its regulation. Biochem. J. 2003, 374, 1–20. [Google Scholar] [CrossRef] [PubMed]
- Wallace, P.M.; MacMaster, J.F.; Rouleau, K.A.; Brown, T.J.; Loy, J.K.; Donaldson, K.L.; Wahl, A.F. Regulation of Inflammatory Responses by Oncostatin M. J. Immunol. 1999, 162, 5547–5555. [Google Scholar] [PubMed]
- Mosley, B.; De Imus, C.; Friend, D.; Boiani, N.; Thoma, B.; Park, L.S.; Cosman, D. Dual oncostatin M (OSM) receptors. Cloning and characterization of an alternative signaling subunit conferring OSM-specific receptor activation. J. Biol. Chem. 1996, 271, 32635–32643. [Google Scholar] [CrossRef] [PubMed]
- Wong, S.; Botelho, F.M.; Rodrigues, R.M.; Richards, C.D. Oncostatin M overexpression induces matrix deposition, STAT3 activation, and SMAD1 Dysregulation in lungs of fibrosis-resistant BALB/c mice. Lab. Investig. 2014, 94, 1003–1016. [Google Scholar] [CrossRef] [PubMed]
- Fritz, D.K.; Kerr, C.; Fattouh, R.; Llop-Guevara, A.; Khan, W.I.; Jordana, M.; Richards, C.D. A Mouse Model of Airway Disease: Oncostatin M-Induced Pulmonary Eosinophilia, Goblet Cell Hyperplasia, and Airway Hyperresponsiveness Are STAT6 Dependent, and Interstitial Pulmonary Fibrosis Is STAT6 Independent. J. Immunol. 2011, 186, 1107–1118. [Google Scholar] [CrossRef] [PubMed]
- Mozaffarian, A.; Brewer, A.W.; Trueblood, E.S.; Luzina, I.G.; Todd, N.W.; Atamas, S.P.; Arnett, H.A. Mechanisms of Oncostatin M-Induced Pulmonary Inflammation and Fibrosis. J. Immunol. 2008, 181, 7243–7253. [Google Scholar] [CrossRef] [PubMed]
- Levy, M.T.; Trojanowska, M.; Reuben, A. Oncostatin M: A cytokine upregulated in human cirrhosis, increases collagen production by human hepatic stellate cells. J. Hepatol. 2000, 32, 218–226. [Google Scholar] [CrossRef]
- Pham Van, T.; Couchie, D.; Martin-Garcia, N.; Laperche, Y.; Zafrani, E.S.; Mavier, P. Expression of matrix metalloproteinase-2 and -9 and of tissue inhibitor of matrix metalloproteinase-1 in liver regeneration from oval cells in rat. Matrix Biol. 2008, 27, 674–681. [Google Scholar] [CrossRef] [PubMed]
- Canady, J.; Arndt, S.; Karrer, S.; Bosserhoff, A.K. Increased KGF Expression Promotes Fibroblast Activation in a Double Paracrine Manner Resulting in Cutaneous Fibrosis. J. Investig. Dermatol. 2013, 133, 647–657. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Infantes, D.; White, U.A.; Elks, C.M.; Morrison, R.F.; Gimble, J.M.; Considine, R.V.; Ferrante, A.W.; Ravussin, E.; Stephens, J.M. Oncostatin M Is Produced in Adipose Tissue and Is Regulated in Conditions of Obesity and Type 2 Diabetes. J. Clin. Endocrinol. Metab. 2014, 99, E217–E225. [Google Scholar] [CrossRef] [PubMed]
- Komori, T.; Tanaka, M.; Senba, E.; Miyajima, A.; Morikawa, Y. Lack of Oncostatin M Receptor β Leads to Adipose Tissue Inflammation and Insulin Resistance by Switching Macrophage Phenotype. J. Biol. Chem. 2013, 288, 21861–21875. [Google Scholar] [CrossRef]
- Komori, T.; Tanaka, M.; Senba, E.; Miyajima, A.; Morikawa, Y. Deficiency of Oncostatin M Receptor β (OSMRβ) Exacerbates High-fat Diet-induced Obesity and Related Metabolic Disorders in Mice. J. Biol. Chem. 2014, 289, 13821–13837. [Google Scholar] [CrossRef] [PubMed]
- Moest, H.; Frei, A.P.; Bhattacharya, I.; Geiger, M.; Wollscheid, B.; Wolfrum, C. Malfunctioning of adipocytes in obesity is linked to quantitative surfaceome changes. Biochim. Biophys. Acta (BBA) Mol. Cell Biol. Lipids 2013, 1831, 1208–1216. [Google Scholar] [CrossRef]
- Kucia, M.; Jankowski, K.; Reca, R.; Wysoczynski, M.; Bandura, L.; Allendorf, D.J.; Zhang, J.; Ratajczak, J.; Ratajczak, M.Z. CXCR4–SDF-1 Signalling, Locomotion, Chemotaxis and Adhesion. J. Mol. Histol. 2004, 35, 233–245. [Google Scholar] [CrossRef] [PubMed]
- Shin, J.; Fukuhara, A.; Onodera, T.; Kita, S.; Yokoyama, C.; Otsuki, M.; Shimomura, I. SDF-1 Is an Autocrine Insulin-Desensitizing Factor in Adipocytes. Diabetes 2018, 67, 1068–1078. [Google Scholar] [CrossRef]
- Wada, T.; Watanabe, E.R.I.; Kotera, Y.; Onogi, Y.; Tsuneki, H.; Sasaoka, T. The SDF1-CXCR4 Signals Regulate Adipose Tissue Expansion by Modulating Angiogenesis in Diet-Induced Obesity in Mice. Diabetes 2018, 67, 2005. [Google Scholar] [CrossRef]
- Maan, Z.N.; Rennert, R.C.; Duscher, D.; Januszyk, M.; Paik, K.; Chung, M.T.; Paik, K.; Fujiwara, T.; Rodrigues, M.; Ho, N.; et al. Abstract 8: SDF-1 Regulates Adipose Niche Homeostasis and Adipose Derived Stromal Cell Function. Plast. Reconstr. Surg. 2014, 133, 15–16. [Google Scholar]
- Albiero, M.; Poncina, N.; Ciciliot, S.; Cappellari, R.; Menegazzo, L.; Ferraro, F.; Bolego, C.; Cignarella, A.; Avogaro, A.; Fadini, G.P. Bone Marrow Macrophages Contribute to Diabetic Stem Cell Mobilopathy by Producing Oncostatin M. Diabetes 2015, 64, 2957. [Google Scholar] [CrossRef] [PubMed]
- Hoermann, G.; Cerny-Reiterer, S.; Sadovnik, I.; Müllauer, L.; Bilban, M.; Gröger, M.; Horny, H.-P.; Reiter, A.; Schmitt-Graeff, A.; Mannhalter, C.; et al. Oncostatin M is a FIP1L1/PDGFRA-dependent mediator of cytokine production in chronic eosinophilic leukemia. Allergy 2013, 68, 713–723. [Google Scholar] [CrossRef]
- Hohensinner, P.J.; Kaun, C.; Rychli, K.; Niessner, A.; Pfaffenberger, S.; Rega, G.; Furnkranz, A.; Uhrin, P.; Zaujec, J.; Afonyushkin, T.; et al. The inflammatory mediator oncostatin M induces stromal derived factor-1 in human adult cardiac cells. FASEB J. 2009, 23, 774–782. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.J.; Song, H.Y.; Kim, M.R.; Sung, S.-M.; Jung, J.S.; Kim, J.H. Oncostatin M stimulates expression of stromal-derived factor-1 in human mesenchymal stem cells. Int. J. Biochem. Cell Biol. 2007, 39, 650–659. [Google Scholar] [CrossRef] [PubMed]
- Minehata, K.-I.; Takeuchi, M.; Hirabayashi, Y.; Inoue, T.; Donovan, P.J.; Tanaka, M.; Miyajima, A. Oncostatin M Maintains the Hematopoietic Microenvironment and Retains Hematopoietic Progenitors in the Bone Marrow. Int. J. Hematol. 2006, 84, 319–327. [Google Scholar] [CrossRef]
- Botelho, F.M.; Rodrigues, R.; Guerette, J.; Wong, S.; Fritz, D.K.; Richards, C.D. Extracellular Matrix and Fibrocyte Accumulation in BALB/c Mouse Lung upon Transient Overexpression of Oncostatin M. Cells 2019, 8, 126. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Kim, J.; Yoon, J.H.; Ghim, J.; Yea, K.; Song, P.; Park, S.; Lee, A.; Hong, C.-P.; Jang, M.S.; et al. CXCL12 secreted from adipose tissue recruits macrophages and induces insulin resistance in mice. Diabetologia 2014, 57, 1456–1465. [Google Scholar] [CrossRef] [PubMed]
- West, N.R.; Murphy, L.C.; Watson, P.H. Oncostatin M suppresses oestrogen receptor-alpha expression and is associated with poor outcome in human breast cancer. Endocr. Relat. Cancer 2012, 19, 181–195. [Google Scholar] [CrossRef] [PubMed]
| Gene Name | Forward | Reverse |
|---|---|---|
| Sdf1/Cxcl12 | GAGCCAACGTCAAGCATCT | CCACTTTAATTTCGGGTCAATGC |
| Timp1 | ACCTGATCCGTCCACAAACA | GGGGTGTGCACAGTGTTTCC |
| Ppia | CCACTGTCGCTTTTCGCCGC | TGCAAACAGCTCGAAGGAGACGC |
| Ubb | CCAGTGGGCAGTGATGG | GCTTACCATGCAACAAAACCT |
| Nono | CATCATCAGCATCACCACCA | TCTTCAGGTCAATAGTCAAGCC |
| Osmr | CGTTCCCCTGTGAGGCCGAG | TCCTCCAAGACTTCGCTTCGGG |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hang, H.; Bailey, J.L.; Elks, C.M. Oncostatin M Mediates Adipocyte Expression and Secretion of Stromal-Derived Factor 1. Biology 2019, 8, 19. https://doi.org/10.3390/biology8010019
Hang H, Bailey JL, Elks CM. Oncostatin M Mediates Adipocyte Expression and Secretion of Stromal-Derived Factor 1. Biology. 2019; 8(1):19. https://doi.org/10.3390/biology8010019
Chicago/Turabian StyleHang, Hardy, Jennifer L. Bailey, and Carrie M. Elks. 2019. "Oncostatin M Mediates Adipocyte Expression and Secretion of Stromal-Derived Factor 1" Biology 8, no. 1: 19. https://doi.org/10.3390/biology8010019
APA StyleHang, H., Bailey, J. L., & Elks, C. M. (2019). Oncostatin M Mediates Adipocyte Expression and Secretion of Stromal-Derived Factor 1. Biology, 8(1), 19. https://doi.org/10.3390/biology8010019






