Along the Axis between Type 1 and Type 2 Immunity; Principles Conserved in Evolution from Fish to Mammals
Abstract
:1. Introduction
General Principles of the i1-i2 Axis as Exemplified by Major Polarizations of Mammalian Helper and Regulatory T Cells
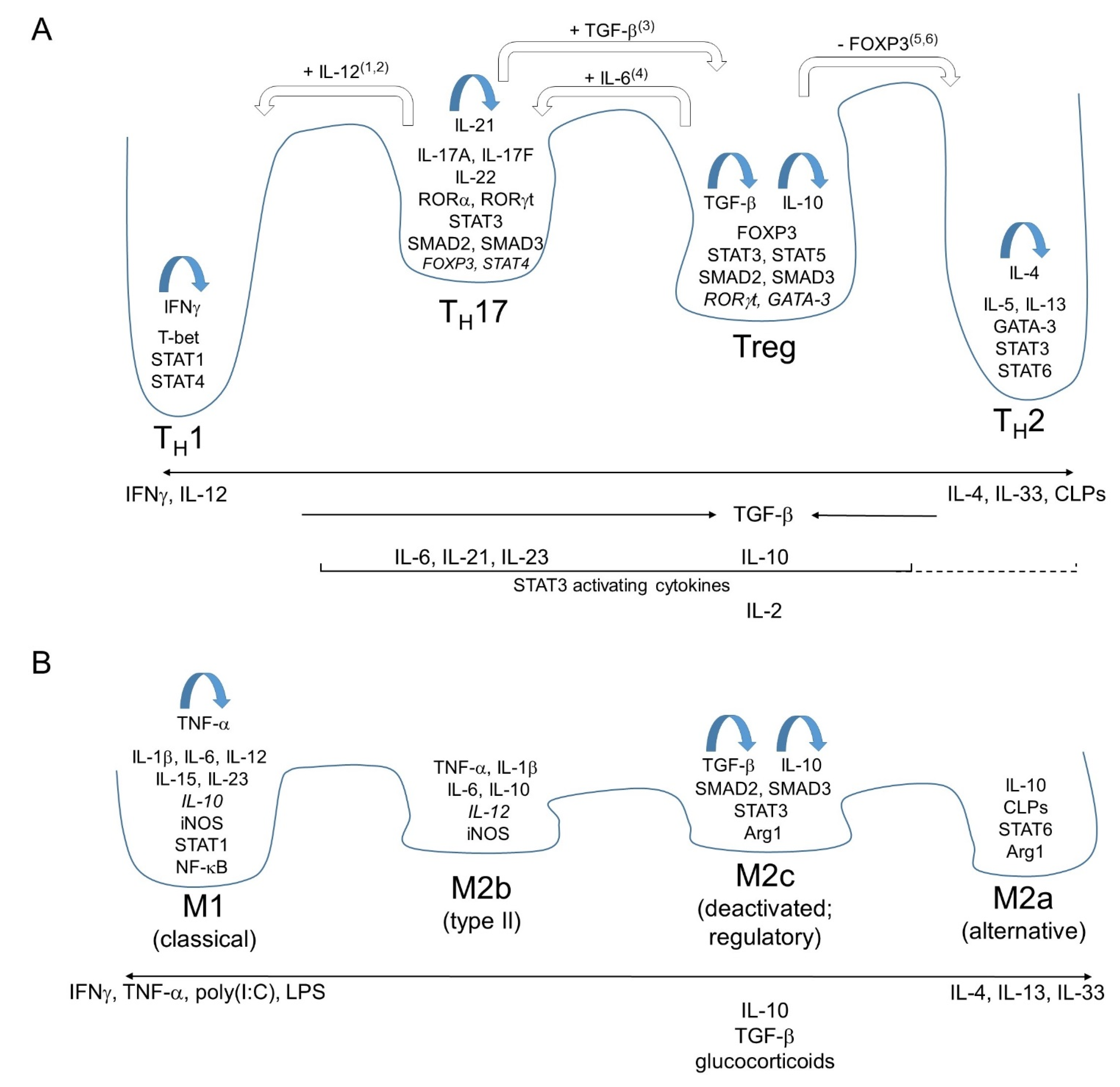
2. Polarizations along the i1-i2 Axis of Mammalian Leukocytes Other than Helper and Regulatory T Cells
3. Fish Orthologues of Mammalian genes for i1-i2 Axis Functions
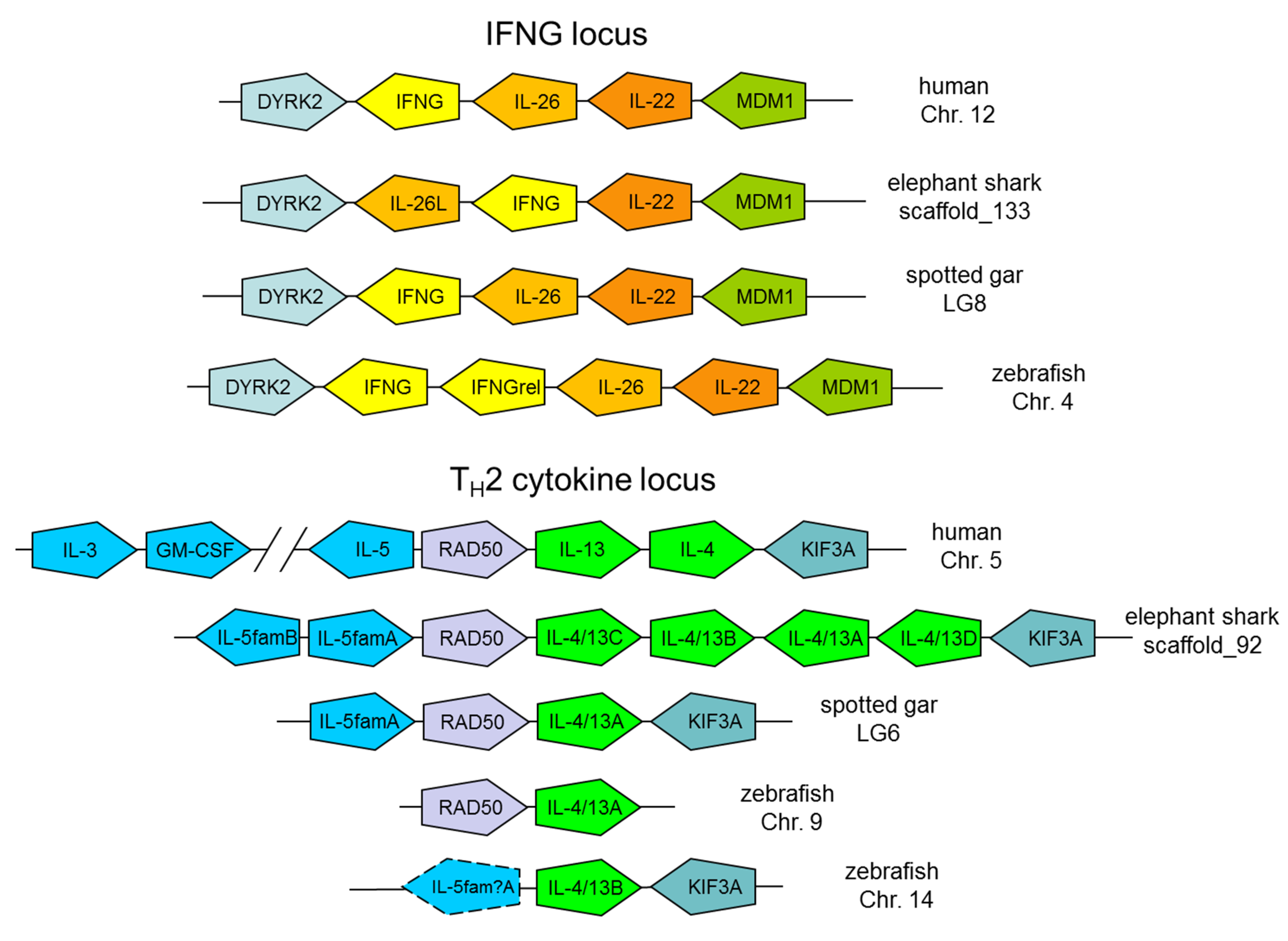
4. Conservation of the IFNG and TH2 Cytokine Loci
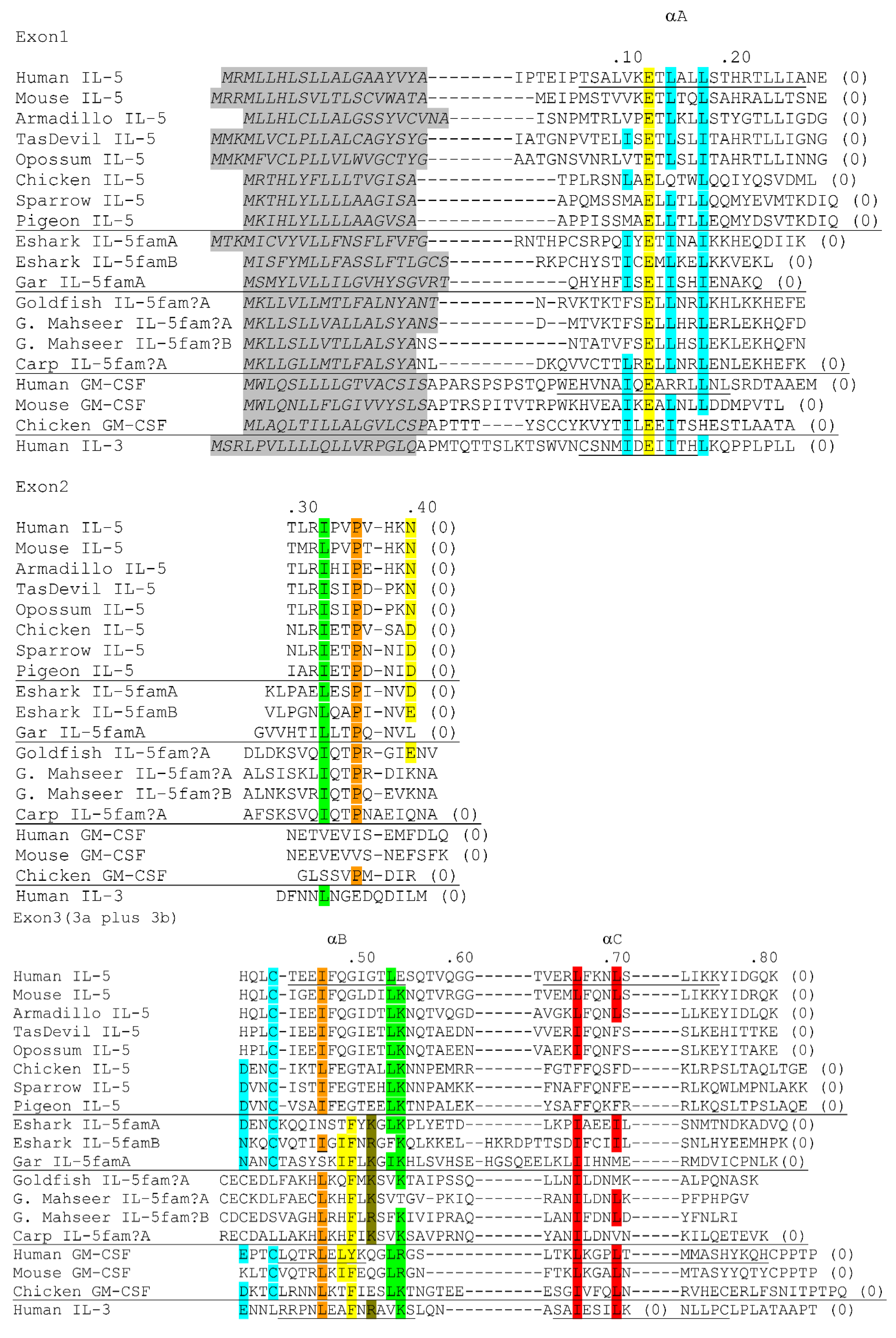
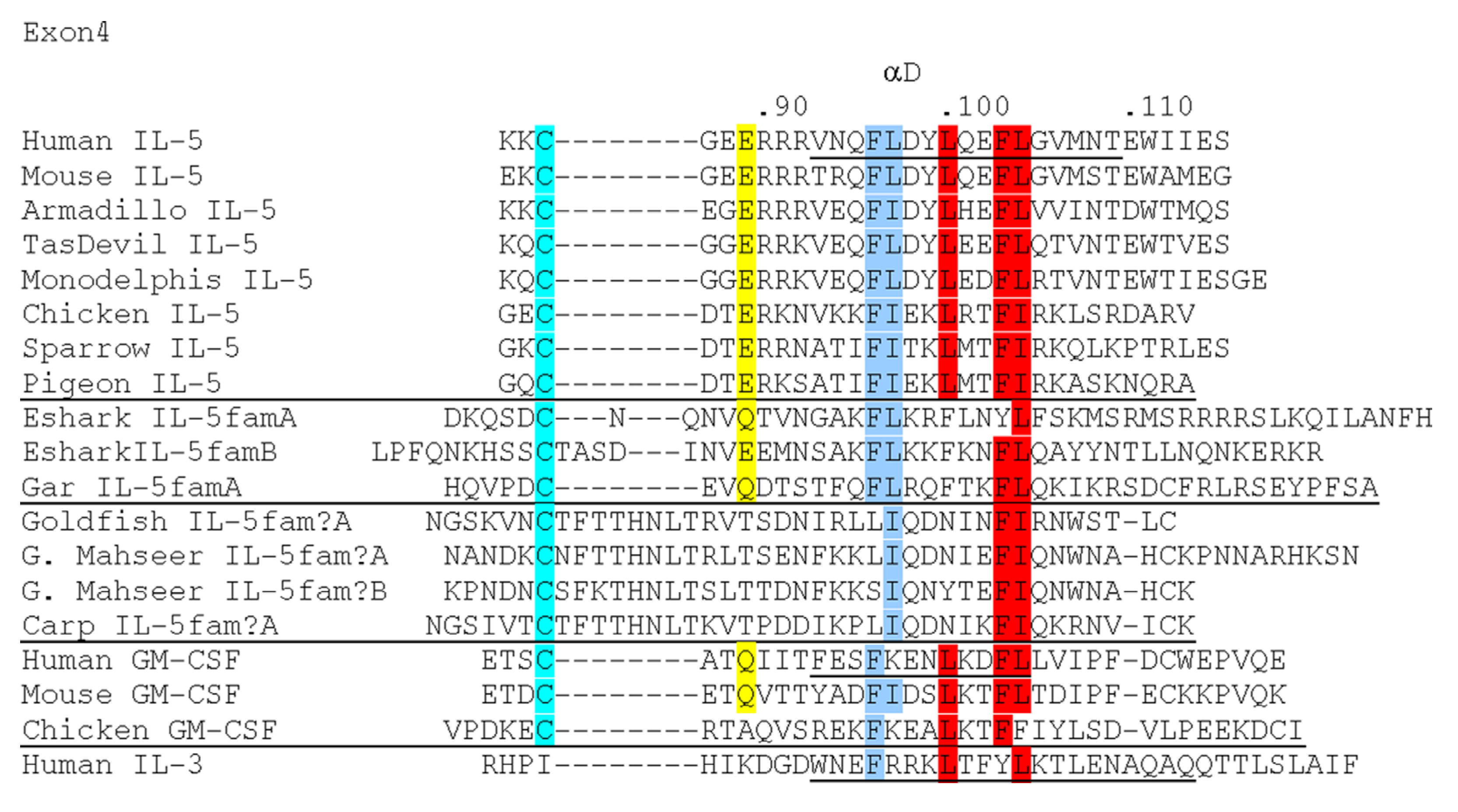
5. Investigation of Tissue-Specific Co-Expressions of TH1, TH17, Treg and TH2 Signature Genes in Fish; Gills Consistently Express High Levels of TH2 Signature Genes
| Elephant Shark (Callorhinchus milii) | |||||
|---|---|---|---|---|---|
| Gill | Kidney | Spleen | Intestine | ||
| TH1-signature | T-bet | 4 | 1 | 121 | 3 |
| STAT1 | 6006 | 701 | 5263 | 2168 | |
| STAT4 | 726 | 131 | 3873 | 738 | |
| IFNγ | 17 | 1 | 78 | 6 | |
| TH17-signature | IL17A/F1 | 1 | 0 | 0 | 7 |
| IL17A/F2 | 21 | 0 | 2 | 66 | |
| IL-21L | 0 | 0 | 3 | 2 | |
| IL-22 | 4 | 0 | 0 | 15 | |
| Treg-signature | Foxp3 | 10 | 1 | 51 | 3 |
| IL-10 | 7 | 4 | 185 | 13 | |
| TGFβ1 | 72 | 36 | 66 | 3 | |
| TH2-signature | GATA3 | 2287 | 92 | 575 | 95 |
| STAT6 | 290 | 41 | 199 | 102 | |
| IL-4/13A | 3 | 0 | 0 | 0 | |
| IL-4/13B | 8 | 0 | 0 | 1 | |
| IL-4/13C | 0 | 1 | 0 | 0 | |
| IL-4/13D | 0 | 0 | 0 | 0 | |
| IL-5A | 0 | 0 | 0 | 0 | |
| IL-5B | 3 | 0 | 0 | 0 | |
| Golden mahseer (Tor putitora) | Northern pike (Esox lucius) | Rainbow trout (Oncorhynchus mykiss) | Atlantic salmon (Salmo salar) | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Gill | Kidney | Spleen | Gill | Kidney | Head Kidney | Spleen | Intestine | Gill | Kidney | Spleen | Intestine | Gill | Kidney | Spleen | Intestine | ||
| TH1-signature | T-bet | 4 | 47 | 54 | 91 | 114 | 663 | 899 | 28 | 12 | 20 | 22 | 11 | 134 | 320 | 1434 | 66 |
| STAT1_1 | 523 | 1174 | 42 | 2589 | 1555 | 3200 | 2579 | 930 | 2041 | 2219 | 5794 | 2422 | 5608 | 5386 | 10317 | 2530 | |
| STAT1_2 | 1990 | 1870 | 3563 | 1878 | |||||||||||||
| STAT4_1 | 1525 | 1714 | 686 | 1866 | 1375 | 3702 | 2708 | 785 | 822 | 964 | 1122 | 679 | 1611 | 729 | 2273 | 1916 | |
| STAT4_2 | 484 | 37 | 62 | 88 | |||||||||||||
| IFNγ_1 | 85 | 47 | 31 | 8 | 0 | 4 | 11 | 1 | 19 | 0 | 189 | 84 | 6 | 6 | 32 | 2 | |
| IFNγ_2 | 17 | 0 | 211 | 99 | |||||||||||||
| TH17-signature | IL-17A/F1a | 14 | 21 | 13 | 13 | 15 | 2 | 0 | 2 | 14 | 5 | 1 | 9 | 0 | 5 | 0 | 0 |
| IL-17A/F1b | 1 | 6 | 0 | 4 | 5 | 11 | 0 | 0 | |||||||||
| IL-17A/F2a | 11 | 6 | 4 | 10 | 0 | 0 | 0 | 0 | 4 | 5 | 0 | 2 | 8 | 39 | 0 | 0 | |
| IL-17A/F2b | 3 | 0 | 0 | 2 | 1 | 0 | 0 | 2 | |||||||||
| IL-17A/F3 | 6 | 10 | 6 | 75 | 7 | 7 | 4 | 1 | 3 | 0 | 2 | 0 | 2 | 1 | 2 | 1 | |
| IL-21 | 8 | 28 | 19 | 53 | 35 | 26 | 14 | 123 | 1 | 0 | 0 | 6 | 0 | 0 | 0 | 0 | |
| IL-22 | 2 | 4 | 1 | 15 | 2 | 0 | 5 | 2 | 13 | 0 | 11 | 5 | 15 | 5 | 7 | 4 | |
| Treg-signature | Foxp3_1 | 11 | 12 | 9 | 100 | 62 | 257 | 138 | 32 | 61 | 11 | 28 | 35 | 230 | 143 | 666 | 149 |
| Foxp3_2 | 115 | 25 | 43 | 63 | |||||||||||||
| IL-10 | 32 | 179 | 130 | 23 | 127 | 111 | 109 | 8 | 0 | 2 | 7 | 1 | 1 | 2 | 2 | 0 | |
| TGFβ1a | 98 | 279 | 59 | 1374 | 891 | 1549 | 1330 | 270 | 177 | 659 | 888 | 211 | 66 | 122 | 125 | 7 | |
| TGFβ1b | 220 | 410 | 910 | 500 | |||||||||||||
| TH2-signature | GATA3 | >20590 | 472 | 113 | 10801 | 409 | 365 | 202 | 141 | 2669 | 98 | 13 | 24 | 16724 | 219 | 334 | 84 |
| STAT6 | 460 | 870 | 17 | 2354 | 2183 | 2650 | 2781 | 875 | 872 | 827 | 1274 | 827 | 1719 | 1313 | 1688 | 788 | |
| IL-4/13A | 1147 | 56 | 36 | 25 | 7 | 4 | 3 | 4 | 29 | 9 | 5 | 5 | 112 | 42 | 33 | 15 | |
| IL-4/13B1 | 13 | 3 | 2 | 3 | 0 | 1 | 0 | 2 | 6 | 0 | 1 | 1 | 31 | 7 | 8 | 14 | |
| IL-4/13B2 | 1 | 6 | 5 | 3 | 8 | 5 | 2 | 2 | 7 | 7 | 3 | 0 | 7 | ||||
| IL-5fam?A | 29 | 13 | 9 | N.A. | N.A. | N.A. | N.A. | N.A. | N.A. | N.A. | N.A. | N.A. | N.A. | N.A. | N.A. | N.A. | |
| IL-5fam?B | 6 | 29 | 12 | N.A. | N.A. | N.A. | N.A. | N.A. | N.A. | N.A. | N.A. | N.A. | N.A. | N.A. | N.A. | N.A. | |
| Elephant shark (Callorhinchus milii) | Golden mahseer (Tor putitora) | Northern pike (Esox lucius) | Rainbow trout (Oncorhynchus mykiss) | Atlantic salmon (Salmo salar) | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| SRA dataset | SRX154852 | SRX154856 | SRX154860 | SRX154855 | SRX768559 | SRX768561 | SRX767362 | SRX514237 | SRX514263 | SRX514240 | SRX514270 | SRX514238 | ERX297522 | ERX297511 | ERX297523 | ERX297509 | SRX608399 | SRX608574 | SRX608599 | SRX608567 |
| Tissue | Gill | Kidney | Spleen | Intestine | Gill | Kidney | Spleen | Gill | Kidney | Head Kidney | Spleen | Intestine | Gill | Kidney | Spleen | Intestine | Gill | Kidney | Spleen | Intestine |
| No. of reads | 71430454 | 118965654 | 83369382 | 147745918 | 41751362 | 34023336 | 51857480 | 58499888 | 60694314 | 61054936 | 61731442 | 60466858 | 39064840 | 32103778 | 41714660 | 40271788 | 59793962 | 61054936 | 60203316 | 59806348 |
6. Evidence Supporting the Existence of TH Cells in Fish
7. TH1-like Responses in Teleost Fish
8. TH17-Like Responses in Teleost Fish
9. Treg-like Responses in Teleost Fish
10. TH2-like Responses in Teleost Fish
11. M1-like vs. M2-Like Macrophage Polarizations in Fish
12. I2-Skewed Tissue Milieus in Healthy Mammals and Fish
For fish this paragraph has an overlap with paragraph 5, but we nevertheless like to dedicate a special paragraph to the comparison between fish and mammals.
13. I2-Skewed Tissue Milieus of Tumors in Mammals and Fish
14. Conclusions and Future Prospects
Supplementary Files
Supplementary File 1Acknowledgments
Author Contributions
Conflicts of Interest
References
- Ceredig, R.; Rolink, A.G.; Brown, G. Models of haematopoiesis: Seeing the wood for the trees. Nat. Rev. Immunol. 2009, 9, 293–300. [Google Scholar] [CrossRef] [PubMed]
- Akirav, E.M.; Alonso-Gonzalez, N.; Truman, L.A.; Ruddle, N.H. Lymphoid Tissues and Organs. In Fundamental Immunology; Paul, W.E., Ed.; Lippincott Williams and Wilkins: Philadelphia, PA, USA, 2012; pp. 47–65. [Google Scholar]
- Ikawa, T. Genetic and epigenetic control of early lymphocyte development. Curr. Top. Microbiol. Immunol. 2014, 381, 1–20. [Google Scholar] [PubMed]
- Fields, P.E.; Lee, G.R.; Kim, S.T.; Bartsevich, V.V.; Flavell, R.A. Th2-specific chromatin remodeling and enhancer activity in the Th2 cytokine locus control region. Immunity 2004, 21, 865–876. [Google Scholar] [CrossRef] [PubMed]
- Ansel, K.M.; Djuretic, I.; Tanasa, B.; Rao, A. Regulation of Th2 differentiation and Il4 locus accessibility. Annu. Rev. Immunol. 2006, 24, 607–656. [Google Scholar] [CrossRef] [PubMed]
- Wilson, C.B.; Rowell, E.; Sekimata, M. Epigenetic control of T-helper-cell differentiation. Nat. Rev. Immunol. 2009, 9, 91–105. [Google Scholar] [CrossRef] [PubMed]
- Mosmann, T.R.; Cherwinski, H.; Bond, M.W.; Giedlin, M.A.; Coffman, R.L. Two types of murine helper T cell clone. I. Definition according to profiles of lymphokine activities and secreted proteins. J. Immunol. 1986, 136, 2348–2357. [Google Scholar] [PubMed]
- Mosmann, T.R.; Coffman, R.L. TH1 and TH2 cells: Different patterns of lymphokine secretion lead to different functional properties. Annu. Rev. Immunol. 1989, 7, 145–173. [Google Scholar] [CrossRef] [PubMed]
- Heinzel, F.P.; Sadick, M.D.; Holaday, J.; Coffman, R.L.; Locksley, R.M. Reciprocal expression of interferon gamma or interleukin 4 during the resolution or progression of murine leishmaniasis. Evidence for expansion of distinct helper T cell subsets. J. Exp. Med. 1989, 169, 59–72. [Google Scholar] [CrossRef] [PubMed]
- Murphy, E.; Shibuya, K.; Hosken, N.; Openshaw, P.; Maino, V.; Davis, K.; Murphy, K.; O’Garra, A. Reversibility of T helper 1 and 2 populations is lost after long-term stimulation. J. Exp. Med. 1996, 183, 901–913. [Google Scholar] [CrossRef] [PubMed]
- Bird, J.J.; Brown, D.R.; Mullen, A.C.; Moskowitz, N.H.; Mahowald, M.A.; Sider, J.R.; Gajewski, T.F.; Wang, C.R.; Reiner, S.L. Helper T cell differentiation is controlled by the cell cycle. Immunity 1998, 9, 229–237. [Google Scholar] [CrossRef]
- McGhee, J.R. The world of TH1/TH2 subsets: First proof. J. Immunol. 2005, 175, 3–4. [Google Scholar] [CrossRef] [PubMed]
- Lighvani, A.A.; Frucht, D.M.; Jankovic, D.; Yamane, H.; Aliberti, J.; Hissong, B.D.; Nguyen, B.V.; Gadina, M.; Sher, A.; Paul, W.E.; et al. T-bet is rapidly induced by interferon-γ in lymphoid and myeloid cells. Proc. Natl. Acad. Sci. USA 2001, 98, 15137–15142. [Google Scholar] [CrossRef] [PubMed]
- Afkarian, M.; Sedy, J.R.; Yang, J.; Jacobson, N.G.; Cereb, N.; Yang, S.Y.; Murphy, T.L.; Murphy, K.M. T-bet is a STAT1-induced regulator of IL-12R expression in naïve CD4+ T cells. Nat. Immunol. 2002, 3, 549–557. [Google Scholar] [CrossRef] [PubMed]
- Kaplan, M.H.; Schindler, U.; Smiley, S.T.; Grusby, M.J. Stat6 is required for mediating responses to IL-4 and for development of Th2 cells. Immunity 1996, 4, 313–319. [Google Scholar] [CrossRef]
- Ouyang, W.; Ranganath, S.H.; Weindel, K.; Bhattacharya, D.; Murphy, T.L.; Sha, W.C.; Murphy, K.M. Inhibition of Th1 development mediated by GATA-3 through an IL-4-independent mechanism. Immunity 1998, 9, 745–755. [Google Scholar] [CrossRef]
- Ouyang, W.; Löhning, M.; Gao, Z.; Assenmacher, M.; Ranganath, S.; Radbruch, A.; Murphy, K.M. Stat6-independent GATA-3 autoactivation directs IL-4-independent Th2 development and commitment. Immunity 2000, 12, 27–37. [Google Scholar] [CrossRef]
- Takemoto, N.; Kamogawa, Y.; Jun Lee, H.; Kurata, H.; Arai, K.I.; O’Garra, A.; Arai, N.; Miyatake, S. Cutting edge: Chromatin remodeling at the IL-4/IL-13 intergenic regulatory region for Th2-specific cytokine gene cluster. J. Immunol. 2000, 165, 6687–6691. [Google Scholar] [CrossRef] [PubMed]
- Lee, G.R.; Kim, S.T.; Spilianakis, C.G.; Fields, P.E.; Flavell, R.A. T helper cell differentiation: Regulation by cis elements and epigenetics. Immunity 2006, 24, 369–379. [Google Scholar] [CrossRef] [PubMed]
- Kidd, P. Th1/Th2 balance: The hypothesis, its limitations, and implications for health and disease. Altern. Med. Rev. 2003, 8, 223–246. [Google Scholar] [PubMed]
- Langston, H.P.; Ke, Y.; Gewirtz, A.T.; Dombrowski, K.E.; Kapp, J.A. Secretion of IL-2 and IFN-γ, but not IL-4, by antigen-specific T cells requires extracellular ATP. J. Immunol. 2003, 170, 2962–2970. [Google Scholar] [CrossRef] [PubMed]
- Pulendran, B.; Artis, D. New paradigms in type 2 immunity. Science 2012, 337, 431–435. [Google Scholar] [CrossRef] [PubMed]
- Gause, W.C.; Wynn, T.A.; Allen, J.E. Type 2 immunity and wound healing: Evolutionary refinement of adaptive immunity by helminths. Nat. Rev. Immunol. 2013, 13, 607–614. [Google Scholar] [CrossRef] [PubMed]
- Koscsó, B.; Csóka, B.; Kókai, E.; Németh, Z.H.; Pacher, P.; Virág, L.; Leibovich, S.J.; Haskó, G. Adenosine augments IL-10-induced STAT3 signaling in M2c macrophages. J. Leukoc Biol. 2013, 94, 1309–1315. [Google Scholar] [CrossRef] [PubMed]
- Steinman, L. A rush to judgment on Th17. J. Exp. Med. 2008, 205, 1517–1522. [Google Scholar] [CrossRef] [PubMed]
- Annunziato, F.; Romagnani, C.; Romagnani, S. The 3 major types of innate and adaptive cell-mediated effector immunity. J. Allergy Clin. Immunol. 2015, 135, 626–635. [Google Scholar] [CrossRef] [PubMed]
- Ouyang, W.; Kolls, J.K.; Zheng, Y. The biological functions of T helper 17 cell effector cytokines in inflammation. Immunity 2008, 28, 454–467. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.O.; Pappu, B.P.; Nurieva, R.; Akimzhanov, A.; Kang, H.S.; Chung, Y.; Ma, L.; Shah, B.; Panopoulos, A.D.; Schluns, K.S.; et al. T helper 17 lineage differentiation is programmed by orphan nuclear receptors ROR α and RORγ. Immunity 2008, 28, 29–39. [Google Scholar] [CrossRef] [PubMed]
- Maloy, K.J.; Kullberg, M.C. IL-23 and Th17 cytokines in intestinal homeostasis. Mucosal Immunol. 2008, 1, 339–349. [Google Scholar] [CrossRef] [PubMed]
- Denning, T.L.; Norris, B.A.; Medina-Contreras, O.; Manicassamy, S.; Geem, D.; Madan, R.; Karp, C.L.; Pulendran, B. Functional specializations of intestinal dendritic cell and macrophage subsets that control Th17 and regulatory T cell responses are dependent on the T cell/APC ratio, source of mouse strain, and regional localization. J. Immunol. 2011, 187, 733–747. [Google Scholar] [CrossRef] [PubMed]
- Goto, Y.; Panea, C.; Nakato, G.; Cebula, A.; Lee, C.; Diez, M.G.; Laufer, T.M.; Ignatowicz, L.; Ivanov, I.I. Segmented filamentous bacteria antigens presented by intestinal dendritic cells drive mucosal Th17 cell differentiation. Immunity 2014, 40, 594–607. [Google Scholar] [CrossRef] [PubMed]
- Cohen, C.J.; Crome, S.Q.; MacDonald, K.G.; Dai, E.L.; Mager, D.L.; Levings, M.K. Human Th1 and Th17 cells exhibit epigenetic stability at signature cytokine and transcription factor loci. J. Immunol. 2011, 187, 5615–5626. [Google Scholar] [CrossRef] [PubMed]
- Adkins, B.; Bu, Y.; Guevara, P. The generation of Th memory in neonates versus adults: Prolonged primary Th2 effector function and impaired development of Th1 memory effector function in murine neonates. J. Immunol. 2001, 166, 918–925. [Google Scholar] [CrossRef] [PubMed]
- Rose, S.; Lichtenheld, M.; Foote, M.R.; Adkins, B. Murine neonatal CD4+ cells are poised for rapid Th2 effector-like function. J. Immunol. 2007, 178, 2667–2678. [Google Scholar] [CrossRef] [PubMed]
- Allen, J.E.; Sutherland, T.E. Host protective roles of type 2 immunity: Parasite killing and tissue repair, flip sides of the same coin. Semin Immunol. 2014, 26, 329–340. [Google Scholar] [CrossRef] [PubMed]
- Foster, P.S.; Hogan, S.P.; Ramsay, A.J.; Matthaei, K.I.; Young, I.G. Interleukin 5 deficiency abolishes eosinophilia, airways hyperreactivity, and lung damage in a mouse asthma model. J. Exp. Med. 1996, 183, 195–201. [Google Scholar] [CrossRef] [PubMed]
- Wynn, T.A. Fibrotic disease and the TH1/TH2 paradigm. Nat. Rev. Immunol. 2004, 4, 583–594. [Google Scholar] [CrossRef] [PubMed]
- Wynn, T.A. Type 2 cytokines: Mechanisms and therapeutic strategies. Nat. Rev. Immunol. 2015, 15, 271–282. [Google Scholar] [CrossRef] [PubMed]
- Arora, M.; Chen, L.; Paglia, M.; Gallagher, I.; Allen, J.E.; Vyas, Y.M.; Ray, A.; Ray, P. Simvastatin promotes Th2-type responses through the induction of the chitinase family member Ym1 in dendritic cells. Proc. Natl. Acad. Sci. USA 2006, 103, 7777–7782. [Google Scholar] [CrossRef] [PubMed]
- Loke, P.; Gallagher, I.; Nair, M.G.; Zang, X.; Brombacher, F.; Mohrs, M.; Allison, J.P.; Allen, J.E. Alternative activation is an innate response to injury that requires CD4+ T cells to be sustained during chronic infection. J. Immunol. 2007, 179, 3926–3936. [Google Scholar] [CrossRef] [PubMed]
- Muallem, G.; Hunter, C.A. ParadYm shift: Ym1 and Ym2 as innate immunological regulators of IL-17. Nat. Immunol. 2014, 15, 1099–1100. [Google Scholar] [CrossRef] [PubMed]
- Sutherland, T.E.; Logan, N.; Rückerl, D.; Humbles, A.A.; Allan, S.M.; Papayannopoulos, V.; Stockinger, B.; Maizels, R.M.; Allen, J.E. Chitinase-like proteins promote IL-17-mediated neutrophilia in a tradeoff between nematode killing and host damage. Nat. Immunol. 2014, 15, 1116–1125. [Google Scholar] [CrossRef] [PubMed]
- Sica, A.; Saccani, A.; Bottazzi, B.; Polentarutti, N.; Vecchi, A.; van Damme, J.; Mantovani, A. Autocrine production of IL-10 mediates defective IL-12 production and NF-kappa B activation in tumor-associated macrophages. J. Immunol. 2000, 164, 762–767. [Google Scholar] [CrossRef] [PubMed]
- Marie, J.C.; Letterio, J.J.; Gavin, M.; Rudensky, A.Y. TGF-β1 maintains suppressor function and Foxp3 expression in CD4+CD25+ regulatory T cells. J. Exp. Med. 2005, 201, 1061–1067. [Google Scholar] [CrossRef] [PubMed]
- Deenick, E.K.; Tangye, S.G. Autoimmunity: IL-21: A new player in Th17-cell differentiation. Immunol. Cell Biol. 2007, 85, 503–505. [Google Scholar] [CrossRef] [PubMed]
- Chaudhry, A.; Samstein, R.M.; Treuting, P.; Liang, Y.; Pils, M.C.; Heinrich, J.M.; Jack, R.S.; Wunderlich, F.T.; Brüning, J.C.; Müller, W.; et al. Interleukin-10 signaling in regulatory T cells is required for suppression of Th17 cell-mediated inflammation. Immunity 2011, 34, 566–578. [Google Scholar] [CrossRef] [PubMed]
- Wan, Y.Y.; Flavell, R.A. Regulatory T-cell functions are subverted and converted owing to attenuated Foxp3 expression. Nature 2007, 445, 766–770. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.O.; Nurieva, R.; Martinez, G.J.; Kang, H.S.; Chung, Y.; Pappu, B.P.; Shah, B.; Chang, S.H.; Schluns, K.S.; Watowich, S.S.; et al. Molecular antagonism and plasticity of regulatory and inflammatory T cell programs. Immunity 2008, 29, 44–56. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.K.; Turner, H.; Maynard, C.L.; Oliver, J.R.; Chen, D.; Elson, C.O.; Weaver, C.T. Late developmental plasticity in the T helper 17 lineage. Immunity 2009, 30, 92–107. [Google Scholar] [CrossRef] [PubMed]
- Mukasa, R.; Balasubramani, A.; Lee, Y.K.; Whitley, S.K.; Weaver, B.T.; Shibata, Y.; Crawford, G.E.; Hatton, R.D.; Weaver, C.T. Epigenetic instability of cytokine and transcription factor gene loci underlies plasticity of the T helper 17 cell lineage. Immunity 2010, 32, 616–627. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Souabni, A.; Flavell, R.A.; Wan, Y.Y. An intrinsic mechanism predisposes Foxp3-expressing regulatory T cells to Th2 conversion in vivo. J. Immunol. 2010, 185, 5983–5992. [Google Scholar] [CrossRef] [PubMed]
- Muranski, P.; Restifo, N.P. Essentials of Th17 cell commitment and plasticity. Blood 2013, 121, 2402–2414. [Google Scholar] [CrossRef] [PubMed]
- Obermajer, N.; Popp, F.C.; Soeder, Y.; Haarer, J.; Geissler, E.K.; Schlitt, H.J.; Dahlke, M.H. Conversion of Th17 into IL-17Aneg regulatory T cells: A novel mechanism in prolonged allograft survival promoted by mesenchymal stem cell-supported minimized immunosuppressive therapy. J. Immunol. 2014, 193, 4988–4999. [Google Scholar] [CrossRef] [PubMed]
- Panzer, M.; Sitte, S.; Wirth, S.; Drexler, I.; Sparwasser, T.; Voehringer, D. Rapid in vivo conversion of effector T cells into Th2 cells during helminth infection. J. Immunol. 2012, 188, 615–623. [Google Scholar] [CrossRef] [PubMed]
- Sawant, D.V.; Vignali, D.A. Once a Treg, always a Treg? Immunol. Rev. 2014, 259, 173–191. [Google Scholar] [CrossRef] [PubMed]
- Sakaguchi, S.; Vignali, D.A.; Rudensky, A.Y.; Niec, R.E.; Waldmann, H. The plasticity and stability of regulatory T cells. Nat. Rev. Immunol. 2013, 13, 461–467. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Su, M.A.; Wan, Y.Y. An essential role of the transcription factor GATA-3 for the function of regulatory T cells. Immunity 2011, 35, 337–348. [Google Scholar] [CrossRef] [PubMed]
- Hatton, R.D.; Weaver, C.T. Duality in the Th17-Treg developmental decision. F1000 Biol. Rep. 2009. [Google Scholar] [CrossRef] [PubMed]
- Mathur, A.N.; Chang, H.C.; Zisoulis, D.G.; Stritesky, G.L.; Yu, Q.; O’Malley, J.T.; Kapur, R.; Levy, D.E.; Kansas, G.S.; Kaplan, M.H. Stat3 and Stat4 direct development of IL-17-secreting Th cells. J. Immunol. 2007, 178, 4901–4907. [Google Scholar] [CrossRef] [PubMed]
- Mantovani, A.; Sica, A.; Sozzani, S.; Allavena, P.; Vecchi, A.; Locati, M. The chemokine system in diverse forms of macrophage activation and polarization. Trends Immunol. 2004, 25, 677–686. [Google Scholar] [CrossRef] [PubMed]
- Lombardo, E.; Alvarez-Barrientos, A.; Maroto, B.; Boscá, L.; Knaus, U.G. TLR4-mediated survival of macrophages is MyD88 dependent and requires TNF-α autocrine signalling. J. Immunol. 2007, 178, 3731–3739. [Google Scholar] [CrossRef] [PubMed]
- Yarilina, A.; Park-Min, K.H.; Antoniv, T.; Hu, X.; Ivashkiv, L.B. TNF activates an IRF1-dependent autocrine loop leading to sustained expression of chemokines and STAT1-dependent type I interferon-response genes. Nat. Immunol. 2008, 9, 378–387. [Google Scholar] [CrossRef] [PubMed]
- Zizzo, G.; Hilliard, B.A.; Monestier, M.; Cohen, P.L. Efficient clearance of early apoptotic cells by human macrophages requires M2c polarization and MerTK induction. J. Immunol. 2012, 189, 3508–3520. [Google Scholar] [CrossRef] [PubMed]
- Fadok, V.A.; Bratton, D.L.; Konowal, A.; Freed, P.W.; Westcott, J.Y.; Henson, P.M. Macrophages that have ingested apoptotic cells in vitro inhibit proinflammatory cytokine production through autocrine/paracrine mechanisms involving TGF-β, PGE2, and PAF. J. Clin. Invest. 1998, 101, 890–898. [Google Scholar] [CrossRef] [PubMed]
- Waddington, C.H. The Strategy of the Genes; A Discussion of Some Aspects of Theoretical Biology; Allen & Unwin: London, UK, 1957. [Google Scholar]
- Rebhahn, J.A.; Deng, N.; Sharma, G.; Livingstone, A.M.; Huang, S.; Mosmann, T.R. An animated landscape representation of CD4+ T-cell differentiation, variability, and plasticity: Insights into the behavior of populations versus cells. Eur. J. Immunol. 2014, 44, 2216–2229. [Google Scholar] [CrossRef] [PubMed]
- Kaplan, M.H.; Hufford, M.M.; Olson, M.R. The development and in vivo function of T helper 9 cells. Nat. Rev. Immunol. 2015, 15, 295–307. [Google Scholar] [CrossRef] [PubMed]
- Glatman Zaretsky, A.; Taylor, J.J.; King, I.L.; Marshall, F.A.; Mohrs, M.; Pearce, E.J. T follicular helper cells differentiate from Th2 cells in response to helminth antigens. J. Exp. Med. 2009, 206, 991–999. [Google Scholar] [CrossRef] [PubMed]
- Ueno, H.; Banchereau, J.; Vinuesa, C.G. Pathophysiology of T follicular helper cells in humans and mice. Nat. Immunol. 2015, 16, 142–152. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Lopes, J.E.; Chong, M.M.; Ivanov, I.I.; Min, R.; Victora, G.D.; Shen, Y.; Du, J.; Rubtsov, Y.P.; Rudensky, A.Y.; et al. TGF-β-induced Foxp3 inhibits TH17 cell differentiation by antagonizing RORgammat function. Nature 2008, 453, 236–240. [Google Scholar] [CrossRef] [PubMed]
- Hatton, R.D. TGF-β in Th17 cell development: The truth is out there. Immunity 2011, 34, 288–290. [Google Scholar] [CrossRef] [PubMed]
- Gorissen, M.; de Vrieze, E.; Flik, G.; Huising, M.O. STAT genes display differential evolutionary rates that correlate with their roles in the endocrine and immune system. J. Endocrinol. 2011, 209, 175–184. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yu, H.; Pardoll, D.; Jove, R. STATs in cancer inflammation and immunity: A leading role for STAT3. Nat. Rev. Cancer 2009, 9, 798–809. [Google Scholar] [CrossRef] [PubMed]
- Stritesky, G.L.; Muthukrishnan, R.; Sehra, S.; Goswami, R.; Pham, D.; Travers, J.; Nguyen, E.T.; Levy, D.E.; Kaplan, M.H. The transcription factor STAT3 is required for T helper 2 cell development. Immunity 2011, 34, 39–49. [Google Scholar] [CrossRef] [PubMed]
- Heltemes-Harris, L.M.; Farrar, M.A. The role of STAT5 in lymphocyte development and transformation. Curr. Opin. Immunol. 2012, 24, 146–152. [Google Scholar] [CrossRef] [PubMed]
- Laurence, A.; Tato, C.M.; Davidson, T.S.; Kanno, Y.; Chen, Z.; Yao, Z.; Blank, R.B.; Meylan, F.; Siegel, R.; Hennighausen, L.; Shevach, E.M.; O’shea, J.J. Interleukin-2 signaling via STAT5 constrains T helper 17 cell generation. Immunity 2007, 26, 371–381. [Google Scholar] [CrossRef] [PubMed]
- Waldmann, T.A. The biology of interleukin-2 and interleukin-15: Implications for cancer therapy and vaccine design. Nat. Rev. Immunol. 2006, 6, 595–601. [Google Scholar] [CrossRef] [PubMed]
- Hazenberg, M.D.; Spits, H. Human innate lymphoid cells. Blood 2014, 124, 700–709. [Google Scholar] [CrossRef] [PubMed]
- Moro, K.; Koyasu, S. Innate lymphoid cells, possible interaction with microbiota. Semin. Immunopathol. 2015, 37, 27–37. [Google Scholar] [CrossRef] [PubMed]
- Bernink, J.H.; Peters, C.P.; Munneke, M.; te Velde, A.A.; Meijer, S.L.; Weijer, K.; Hreggvidsdottir, H.S.; Heinsbroek, S.E.; Legrand, N.; Buskens, C.J.; et al. Human type 1 innate lymphoid cells accumulate in inflamed mucosal tissues. Nat. Immunol. 2013, 14, 221–229. [Google Scholar] [CrossRef] [PubMed]
- Turner, J.E.; Morrison, P.J.; Wilhelm, C.; Wilson, M.; Ahlfors, H.; Renauld, J.C.; Panzer, U.; Helmby, H.; Stockinger, B. IL-9-mediated survival of type 2 innate lymphoid cells promotes damage control in helminth-induced lung inflammation. J. Exp. Med. 2013, 210, 2951–2965. [Google Scholar] [CrossRef] [PubMed]
- Bernink, J.H.; Germar, K.; Spits, H. The role of ILC2 in pathology of type 2 inflammatory diseases. Curr. Opin. Immunol. 2014, 31, 115–120. [Google Scholar] [CrossRef] [PubMed]
- Hepworth, M.R.; Fung, T.C.; Masur, S.H.; Kelsen, J.R.; McConnell, F.M.; Dubrot, J.; Withers, D.R.; Hugues, S.; Farrar, M.A.; Reith, W.; et al. Group 3 innate lymphoid cells mediate intestinal selection of commensal bacteria-specific CD4+ T cells. Science 2015, 348, 1031–1035. [Google Scholar] [CrossRef] [PubMed]
- Sad, S.; Marcotte, R.; Mosmann, T.R. Cytokine-induced differentiation of precursor mouse CD8+ T cells into cytotoxic CD8+ T cells secreting Th1 or Th2 cytokines. Immunity 1995, 2, 271–279. [Google Scholar] [CrossRef]
- Harris, D.P.; Haynes, L.; Sayles, P.C.; Duso, D.K.; Eaton, S.M.; Lepak, N.M.; Johnson, L.L.; Swain, S.L.; Lund, F.E. Reciprocal regulation of polarized cytokine production by effector B and T cells. Nat. Immunol. 2000, 1, 475–482. [Google Scholar] [CrossRef] [PubMed]
- Fridlender, Z.G.; Sun, J.; Kim, S.; Kapoor, V.; Cheng, G.; Ling, L.; Worthen, G.S.; Albelda, S.M. Polarization of tumor-associated neutrophil phenotype by TGF-β: “N1” versus “N2” TAN. Cancer Cell. 2009, 16, 183–194. [Google Scholar] [CrossRef] [PubMed]
- Feili-Hariri, M.; Falkner, D.H.; Morel, P.A. Polarization of naive T cells into Th1 or Th2 by distinct cytokine-driven murine dendritic cell populations: Implications for immunotherapy. J. Leukoc Biol. 2005, 78, 656–664. [Google Scholar] [CrossRef] [PubMed]
- Qin, H.; Holdbrooks, A.T.; Liu, Y.; Reynolds, S.L.; Yanagisawa, L.L.; Benveniste, E.N. SOCS3 deficiency promotes M1 macrophage polarization and inflammation. J. Immunol. 2012, 189, 3439–3448. [Google Scholar] [CrossRef] [PubMed]
- Biswas, S.K.; Mantovani, A. Macrophage plasticity and interaction with lymphocyte subsets: Cancer as a paradigm. Nat. Immunol. 2010, 11, 889–896. [Google Scholar] [CrossRef] [PubMed]
- Messi, M.; Giacchetto, I.; Nagata, K.; Lanzavecchia, A.; Natoli, G.; Sallusto, F. Memory and flexibility of cytokine gene expression as separable properties of human TH1 and TH2 lymphocytes. Nat. Immunol. 2003, 4, 78–86. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.Y.; Yang, H.Y.; Monie, A.; Ma, B.; Tsai, H.H.; Wu, T.C.; Hung, C.F. Intraperitoneal administration of poly(I:C) with polyethylenimine leads to significant antitumor immunity against murine ovarian tumors. Cancer Immunol. Immunother. 2011, 60, 1085–1096. [Google Scholar] [CrossRef] [PubMed]
- Shime, H.; Matsumoto, M.; Oshiumi, H.; Tanaka, S.; Nakane, A.; Iwakura, Y.; Tahara, H.; Inoue, N.; Seya, T. Toll-like receptor 3 signaling converts tumor-supporting myeloid cells to tumoricidal effectors. Proc. Natl. Acad. Sci. USA 2012, 109, 2066–2071. [Google Scholar] [CrossRef] [PubMed]
- Doherty, T.M.; Seder, R.A.; Sher, A. Induction and regulation of IL-15 expression in murine macrophages. J. Immunol. 1996, 156, 735–741. [Google Scholar] [PubMed]
- Ka, M.B.; Daumas, A.; Textoris, J.; Mege, J.L. Phenotypic diversity and emerging new tools to study macrophage activation in bacterial infectious diseases. Front Immunol. 2014. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kennedy, M.K.; Glaccum, M.; Brown, S.N.; Butz, E.A.; Viney, J.L.; Embers, M.; Matsuki, N.; Charrier, K.; Sedger, L.; Willis, C.R.; et al. Reversible defects in natural killer and memory CD8 T cell lineages in interleukin 15-deficient mice. J. Exp. Med. 2000, 191, 771–780. [Google Scholar] [CrossRef] [PubMed]
- Edwards, J.P.; Zhang, X.; Frauwirth, K.A.; Mosser, D.M. Biochemical and functional characterization of three activated macrophage populations. J. Leukoc. Biol. 2006, 80, 1298–1307. [Google Scholar] [CrossRef] [PubMed]
- Gerber, J.S.; Mosser, D.M. Reversing lipopolysaccharide toxicity by ligating the macrophage Fcγ receptors. J. Immunol. 2001, 166, 6861–6868. [Google Scholar] [CrossRef] [PubMed]
- Anderson, C.F.; Mosser, D.M. Cutting edge: Biasing immune responses by directing antigen to macrophage Fcγ receptors. J. Immunol. 2002, 168, 3697–3701. [Google Scholar] [CrossRef] [PubMed]
- Modolell, M.; Corraliza, I.M.; Link, F.; Soler, G.; Eichmann, K. Reciprocal regulation of the nitric oxide synthase/arginase balance in mouse bone marrow-derived macrophages by TH1 and TH2 cytokines. Eur. J. Immunol. 1995, 25, 1101–1104. [Google Scholar] [CrossRef] [PubMed]
- Munder, M. Arginase: An emerging key player in the mammalian immune system. Br. J. Pharmacol. 2009, 158, 638–651. [Google Scholar] [CrossRef] [PubMed]
- Whyte, C.S.; Bishop, E.T.; Rückerl, D.; Gaspar-Pereira, S.; Barker, R.N.; Allen, J.E.; Rees, A.J.; Wilson, H.M. Suppressor of cytokine signaling (SOCS)1 is a key determinant of differential macrophage activation and function. J. Leukoc Biol. 2011, 90, 845–854. [Google Scholar] [CrossRef] [PubMed]
- Stempin, C.C.; Dulgerian, L.R.; Garrido, V.V.; Cerban, F.M. Arginase in parasitic infections: Macrophage activation, immunosuppression, and intracellular signals. J. Biomed. Biotechnol. 2010. [Google Scholar] [CrossRef] [PubMed]
- Miron, V.E.; Boyd, A.; Zhao, J.W.; Yuen, T.J.; Ruckh, J.M.; Shadrach, J.L.; van Wijngaarden, P.; Wagers, A.J.; Williams, A.; Franklin, R.J.; et al. M2 microglia and macrophages drive oligodendrocyte differentiation during CNS remyelination. Nat. Neurosci. 2013, 16, 1211–1218. [Google Scholar] [CrossRef] [PubMed]
- Kurowska-Stolarska, M.; Stolarski, B.; Kewin, P.; Murphy, G.; Corrigan, C.J.; Ying, S.; Pitman, N.; Mirchandani, A.; Rana, B.; van Rooijen, N.; et al. IL-33 amplifies the polarization of alternatively activated macrophages that contribute to airway inflammation. J. Immunol. 2009, 183, 6469–6477. [Google Scholar] [CrossRef] [PubMed]
- Nelson, M.P.; Christmann, B.S.; Werner, J.L.; Metz, A.E.; Trevor, J.L.; Lowell, C.A.; Steele, C. IL-33 and M2a alveolar macrophages promote lung defense against the atypical fungal pathogen Pneumocystis murina. J. Immunol. 2011, 186, 2372–2381. [Google Scholar] [CrossRef] [PubMed]
- Korenaga, H.; Kono, T.; Sakai, M. Isolation of seven IL-17 family genes from the Japanese pufferfish Takifugu rubripes. Fish Shellfish Immunol. 2010, 28, 809–818. [Google Scholar] [CrossRef] [PubMed]
- Venkatesh, B.; Lee, A.P.; Ravi, V.; Maurya, A.K.; Lian, M.M.; Swann, J.B.; Ohta, Y.; Flajnik, M.F.; Sutoh, Y.; Kasahara, M.; et al. Elephant shark genome provides unique insights into gnathostome evolution. Nature 2014, 505, 174–179. [Google Scholar] [CrossRef] [PubMed]
- Welch, J.S.; Escoubet-Lozach, L.; Sykes, D.B.; Liddiard, K.; Greaves, D.R.; Glass, C.K. TH2 cytokines and allergic challenge induce Ym1 expression in macrophages by a STAT6-dependent mechanism. J. Biol. Chem. 2002, 277, 42821–42829. [Google Scholar] [CrossRef] [PubMed]
- Pepe, G.; Calderazzi, G.; de Maglie, M.; Villa, A.M.; Vegeto, E. Heterogeneous induction of microglia M2a phenotype by central administration of interleukin-4. J. Neuroinflammation. 2014. [Google Scholar] [CrossRef] [PubMed]
- Ehrchen, J.; Helming, L.; Varga, G.; Pasche, B.; Loser, K.; Gunzer, M.; Sunderkötter, C.; Sorg, C.; Roth, J.; Lengeling, A. Vitamin D receptor signaling contributes to susceptibility to infection with Leishmania major. FASEB J. 2007, 21, 3208–3218. [Google Scholar] [CrossRef] [PubMed]
- Hao, N.B.; Lü, M.H.; Fan, Y.H.; Cao, Y.L.; Zhang, Z.R.; Yang, S.M. Macrophages in tumor microenvironments and the progression of tumors. Clin. Dev. Immunol. 2012. [Google Scholar] [CrossRef] [PubMed]
- Lawrence, T.; Natoli, G. Transcriptional regulation of macrophage polarization: Enabling diversity with identity. Nat. Rev. Immunol. 2011, 11, 750–761. [Google Scholar] [CrossRef] [PubMed]
- Thomas, A.C.; Mattila, J.T. “Of mice and men”: Arginine metabolism in macrophages. Front Immunol. 2014. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sica, A.; Schioppa, T.; Mantovani, A.; Allavena, P. Tumour-associated macrophages are a distinct M2 polarised population promoting tumour progression: Potential targets of anti-cancer therapy. Eur. J. Cancer 2006, 42, 717–727. [Google Scholar] [CrossRef] [PubMed]
- Wynn, T.A.; Chawla, A.; Pollard, J.W. Macrophage biology in development, homeostasis and disease. Nature 2013, 496, 445–455. [Google Scholar] [CrossRef] [PubMed]
- Fridlender, Z.G.; Jassar, A.; Mishalian, I.; Wang, L.C.; Kapoor, V.; Cheng, G.; Sun, J.; Singhal, S.; Levy, L.; Albelda, S.M. Using macrophage activation to augment immunotherapy of established tumours. Br. J. Cancer 2013, 108, 1288–1297. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Knolhoff, B.L.; Meyer, M.A.; Nywening, T.M.; West, B.L.; Luo, J.; Wang-Gillam, A.; Goedegebuure, S.P.; Linehan, D.C.; DeNardo, D.G. CSF1/CSF1R blockade reprograms tumor-infiltrating macrophages and improves response to T-cell checkpoint immunotherapy in pancreatic cancer models. Cancer Res. 2014, 74, 5057–5069. [Google Scholar] [CrossRef] [PubMed]
- Elephant Shark Genome Project. Available online: http://esharkgenome.imcb.a-star.edu.sg/ (accessed on 10 November 2015).
- Ensembl Genome Browser. Available online: http://www.ensembl.org/index.html (accessed on 10 November 2015).
- Zou, J.; Yoshiura, Y.; Dijkstra, J.M.; Sakai, M.; Ototake, M.; Secombes, C. Identification of an interferon gamma homologue in Fugu, Takifugu rubripes. Fish Shellfish Immunol. 2004, 17, 403–409. [Google Scholar] [CrossRef] [PubMed]
- Li, J.H.; Shao, J.Z.; Xiang, L.X.; Wen, Y. Cloning, characterization and expression analysis of pufferfish interleukin-4 cDNA: The first evidence of Th2-type cytokine in fish. Mol. Immunol. 2007, 44, 2078–2086. [Google Scholar] [CrossRef] [PubMed]
- Ohtani, M.; Hayashi, N.; Hashimoto, K.; Nakanishi, T.; Dijkstra, J.M. Comprehensive clarification of two paralogous interleukin 4/13 loci in teleost fish. Immunogenetics 2008, 60, 383–397. [Google Scholar] [CrossRef] [PubMed]
- Igawa, D.; Sakai, M.; Savan, R. An unexpected discovery of two interferon γ-like genes along with interleukin (IL)-22 and -26 from teleost: IL-22 and -26 genes have been described for the first time outside mammals. Mol. Immunol. 2006, 43, 999–1009. [Google Scholar] [CrossRef] [PubMed]
- Dijkstra, J.M. TH2 and Treg candidate genes in elephant shark. Nature 2014, 511, E7–E9. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Secombes, C.J. The evolution of IL-4 and IL-13 and their receptor subunits. Cytokine 2015, 75, 8–13. [Google Scholar] [CrossRef] [PubMed]
- Jaillon, O.; Aury, J.M.; Brunet, F.; Petit, J.L.; Stange-Thomann, N.; Mauceli, E.; Bouneau, L.; Fischer, C.; Ozouf-Costaz, C.; Bernot, A.; et al. Genome duplication in the teleost fish Tetraodon nigroviridis reveals the early vertebrate proto-karyotype. Nature 2004, 431, 946–957. [Google Scholar] [PubMed]
- Berthelot, C.; Brunet, F.; Chalopin, D.; Juanchich, A.; Bernard, M.; Noël, B.; Bento, P.; da Silva, C.; Labadie, K.; Alberti, A.; et al. The rainbow trout genome provides novel insights into evolution after whole-genome duplication in vertebrates. Nat. Commun. 2014. [Google Scholar] [CrossRef] [PubMed]
- Yoshiura, Y.; Kiryu, I.; Fujiwara, A.; Suetake, H.; Suzuki, Y.; Nakanishi, T.; Ototake, M. Identification and characterization of Fugu orthologues of mammalian interleukin-12 subunits. Immunogenetics 2003, 55, 296–306. [Google Scholar] [PubMed]
- Bird, S.; Zou, J.; Kono, T.; Sakai, M.; Dijkstra, J.M.; Secombes, C. Characterisation and expression analysis of interleukin 2 (IL-2) and IL-21 homologues in the Japanese pufferfish, Fugu rubripes, following their discovery by synteny. Immunogenetics 2005, 56, 909–923. [Google Scholar] [CrossRef] [PubMed]
- Mitra, S.; Alnabulsi, A.; Secombes, C.J.; Bird, S. Identification and characterization of the transcription factors involved in T-cell development, t-bet, stat6 and foxp3, within the zebrafish, Danio rerio. FEBS J. 2010, 277, 128–147. [Google Scholar] [CrossRef] [PubMed]
- Liongue, C.; Ward, A.C. Evolution of class I cytokine receptors. BMC Evol. Biol. 2007. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Gorgoglione, B.; Maehr, T.; Holland, J.W.; Vecino, J.L.; Wadsworth, S.; Secombes, C.J. Fish suppressors of cytokine signaling (SOCS): Gene discovery, modulation of expression and function. J. Signal Transduct. 2011. [Google Scholar] [CrossRef] [PubMed]
- Skjesol, A.; Liebe, T.; Iliev, D.B.; Thomassen, E.I.; Tollersrud, L.G.; Sobhkhez, M.; Secombes, C.J.; Joensen, L.L.; Jørgensen, J.B. Functional conservation of suppressors of cytokine signaling proteins between teleosts and mammals: Atlantic salmon SOCS1 binds to JAK/STAT family members and suppresses type I and II IFN signaling. Dev. Comp. Immunol. 2014, 45, 177–189. [Google Scholar] [CrossRef] [PubMed]
- Stolte, E.H.; van Kemenade, B.M.; Savelkoul, H.F.; Flik, G. Evolution of glucocorticoid receptors with different glucocorticoid sensitivity. J. Endocrinol. 2006, 190, 17–28. [Google Scholar] [CrossRef] [PubMed]
- Hussain, M.; Wilson, J.B. New paralogues and revised time line in the expansion of the vertebrate GH18 family. J. Mol. Evol. 2013, 76, 240–260. [Google Scholar] [CrossRef] [PubMed]
- Huising, M.O.; Stet, R.J.M.; Savelkoul, H.F.J.; Verburg-van Kemenade, B.M.L. The molecular evolution of the interleukin-1 family of cytokines; IL-18 in teleost fish. Dev. Comp. Immunol. 2004, 28, 395–413. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Bird, S.; Koussounadis, A.; Holland, J.W.; Carrington, A.; Zou, J.; Secombes, C.J. Identification of a novel IL-1 cytokine family member in teleost fish. J. Immunol. 2009, 183, 962–974. [Google Scholar] [CrossRef] [PubMed]
- Stansberg, C.; Subramaniam, S.; Olsen, L.; Secombes, C.J.; Cunningham, C. Cloning and characterisation of a putative ST2L homologue from Atlantic salmon (Salmo salar). Fish Shellfish Immunol. 2003, 15, 211–224. [Google Scholar] [CrossRef]
- Köbis, J.M.; Rebl, A.; Kühn, C.; Goldammer, T. Comparison of splenic transcriptome activity of two rainbow trout strains differing in robustness under regional aquaculture conditions. Mol. Biol. Rep. 2013, 40, 1955–1966. [Google Scholar] [CrossRef] [PubMed]
- Gibson, M.S.; Kaiser, P.; Fife, M. The chicken IL-1 family: Evolution in the context of the studied vertebrate lineage. Immunogenetics 2014, 66, 427–438. [Google Scholar] [CrossRef] [PubMed]
- Dijkstra, J.M.; Grimholt, U.; Leong, J.; Koop, B.F.; Hashimoto, K. Comprehensive analysis of MHC class II genes in teleost fish genomes reveals dispensability of the peptide-loading DM system in a large part of vertebrates. BMC Evol. Biol. 2013. [Google Scholar] [CrossRef] [PubMed]
- Grimholt, U.; Tsukamoto, K.; Azuma, T.; Leong, J.; Koop, B.F.; Dijkstra, J.M. A comprehensive analysis of teleost MHC class I sequences. BMC Evol. Biol. 2015. [Google Scholar] [CrossRef] [PubMed]
- Nomiyama, H.; Hieshima, K.; Osada, N.; Kato-Unoki, Y.; Otsuka-Ono, K.; Takegawa, S.; Izawa, T.; Yoshizawa, A.; Kikuchi, Y.; Tanase, S.; et al. Extensive expansion and diversification of the chemokine gene family in zebrafish: Identification of a novel chemokine subfamily CX. BMC Genomics 2008. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Yang, L.; Liu, W. Distinct evolution process among type I interferon in mammals. Protein Cell. 2013, 4, 383–392. [Google Scholar] [CrossRef] [PubMed]
- Cannon, J.P.; Haire, R.N.; Magis, A.T.; Eason, D.D.; Winfrey, K.N.; Prada, J.A.H.; Bailey, K.M.; Jakoncic, J.; Litman, G.W.; Ostrov, D.A. A bony fish immunological receptor of the NITR multigene family mediates allogeneic recognition. Immunity 2008, 29, 228–237. [Google Scholar] [CrossRef] [PubMed]
- Amemiya, C.T.; Saha, N.R.; Zapata, A. Evolution and development of immunological structures in the lamprey. Curr. Opin. Immunol. 2007, 19, 535–541. [Google Scholar] [CrossRef] [PubMed]
- Flajnik, M.F.; du Pasquier, L. Evolution of the immune system. In Fundamental Immunology; Paul, W.E., Ed.; Lippincott Williams and Wilkins: Philadelphia, PA, USA, 2012; pp. 67–128. [Google Scholar]
- Kasahara, M.; Sutoh, Y. Two forms of adaptive immunity in vertebrates: Similarities and differences. Adv. Immunol. 2014, 122, 59–90. [Google Scholar] [PubMed]
- Seumois, G.; Chavez, L.; Gerasimova, A.; Lienhard, M.; Omran, N.; Kalinke, L.; Vedanayagam, M.; Ganesan, A.P.; Chawla, A.; Djukanović, R.; et al. Epigenomic analysis of primary human T cells reveals enhancers associated with TH2 memory cell differentiation and asthma susceptibility. Nat. Immunol. 2014, 15, 777–788. [Google Scholar] [CrossRef] [PubMed]
- Loots, G.G.; Locksley, R.M.; Blankespoor, C.M.; Wang, Z.E.; Miller, W.; Rubin, E.M.; Frazer, K.A. Identification of a coordinate regulator of interleukins 4, 13, and 5 by cross-species sequence comparisons. Science 2000, 288, 136–140. [Google Scholar] [CrossRef] [PubMed]
- Santangelo, S.; Cousins, D.J.; Winkelmann, N.E.; Staynov, D.Z. DNA methylation changes at human Th2 cytokine genes coincide with DNase I hypersensitive site formation during CD4+ T cell differentiation. J. Immunol. 2002, 169, 1893–1903. [Google Scholar] [CrossRef] [PubMed]
- Sallusto, F.; Reiner, S.L. Sliding doors in the immune response. Nat. Immunol. 2005, 6, 10–12. [Google Scholar] [CrossRef] [PubMed]
- Zeng, W.P. “All things considered”: Transcriptional regulation of T helper type 2 cell differentiation from precursor to effector activation. Immunology 2013, 140, 31–38. [Google Scholar] [CrossRef] [PubMed]
- PHYRE2 Protein Fold Recognition Server. Available online: www.sbg.bio.ic.ac.uk/phyre2/ (accessed on 10 November 2015).
- Huising, M.O.; Kruiswijk, C.P.; Flik, G. Phylogeny and evolution of class-I helical cytokines. J. Endocrinol. 2006, 189, 1–25. [Google Scholar] [CrossRef] [PubMed]
- Hurley, I.A.; Mueller, R.L.; Dunn, K.A.; Schmidt, E.J.; Friedman, M.; Ho, R.K.; Prince, V.E.; Yang, Z.; Thomas, M.G.; Coates, M.I. A new time-scale for ray-finned fish evolution. Proc. Biol. Sci. 2007, 274, 489–498. [Google Scholar] [CrossRef] [PubMed]
- Wells, T.N.; Graber, P.; Proudfoot, A.E.; Arod, C.Y.; Jordan, S.R.; Lambert, M.H.; Hassel, A.M.; Milburn, M.V. The three-dimensional structure of human interleukin-5 at 2.4-angstroms resolution: Implication for the structures of other cytokines. Ann. NY Acad. Sci. 1994, 725, 118–127. [Google Scholar] [CrossRef] [PubMed]
- Rozwarski, D.A.; Diederichs, K.; Hecht, R.; Boone, T.; Karplus, P.A. Refined crystal structure and mutagenesis of human granulocyte-macrophage colony-stimulating factor. Proteins 1996, 26, 304–313. [Google Scholar] [CrossRef]
- Feng, Y.; Klein, B.K.; McWherter, C.A. Three-dimensional solution structure and backbone dynamics of a variant of human interleukin-3. J. Mol. Biol. 1996, 259, 524–541. [Google Scholar] [CrossRef] [PubMed]
- Rozwarski, D.A.; Gronenborn, A.M.; Clore, G.M.; Bazan, J.F.; Bohm, A.; Wlodawer, A.; Hatada, M.; Karplus, P.A. Structural comparisons among the short-chain helical cytokines. Structure 1994, 2, 159–173. [Google Scholar] [CrossRef]
- Dijkstra, J.M.; Takizawa, F.; Fischer, U.; Friedrich, M.; Soto-Lampe, V.; Lefèvre, C.; Lenk, M.; Karger, A.; Matsui, T.; Hashimoto, K. Identification of a gene for an ancient cytokine, interleukin 15-like, in mammals; interleukins 2 and 15 co-evolved with this third family member, all sharing binding motifs for IL-15Rα. Immunogenetics 2014, 66, 93–103. [Google Scholar] [CrossRef] [PubMed]
- Barry, S.C.; Bagley, C.J.; Phillips, J.; Dottore, M.; Cambareri, B.; Moretti, P.; D’Andrea, R.; Goodall, G.J.; Shannon, M.F.; Vadas, M.A.; et al. Two contiguous residues in human interleukin-3, Asp21 and Glu22, selectively interact with the alpha- and beta-chains of its receptor and participate in function. J. Biol. Chem. 1994, 269, 8488–8492. [Google Scholar] [PubMed]
- Hercus, T.R.; Bagley, C.J.; Cambareri, B.; Dottore, M.; Woodcock, J.M.; Vadas, M.A.; Shannon, M.F.; Lopez, A.F. Specific human granulocyte-macrophage colony-stimulating factor antagonists. Proc. Natl. Acad. Sci. USA 1994, 91, 5838–5842. [Google Scholar] [CrossRef] [PubMed]
- McKinnon, M.; Page, K.; Uings, I.J.; Banks, M.; Fattah, D.; Proudfoot, A.E.; Graber, P.; Arod, C.; Fish, R.; Wells, T.N.; et al. An interleukin 5 mutant distinguishes between two functional responses in human eosinophils. J. Exp. Med. 1997, 186, 121–129. [Google Scholar] [CrossRef] [PubMed]
- Hansen, G.; Hercus, T.R.; McClure, B.J.; Stomski, F.C.; Dottore, M.; Powell, J.; Ramshaw, H.; Woodcock, J.M.; Xu, Y.; Guthridge, M.; et al. The structure of the GM-CSF receptor complex reveals a distinct mode of cytokine receptor activation. Cell 2008, 134, 496–507. [Google Scholar] [CrossRef] [PubMed]
- Takizawa, F.; Koppang, E.O.; Ohtani, M.; Nakanishi, T.; Hashimoto, K.; Fischer, U.; Dijkstra, J.M. Constitutive high expression of interleukin-4/13A and GATA-3 in gill and skin of salmonid fishes suggests that these tissues form Th2-skewed immune environments. Mol. Immunol. 2011, 48, 1360–1368. [Google Scholar] [CrossRef] [PubMed]
- Kumari, J.; Bogwald, J.; Dalmo, R.A. Transcription factor GATA-3 in Atlantic salmon (Salmo salar): Molecular characterization, promoter activity and expression analysis. Mol. Immunol. 2009, 46, 3099–3107. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Holland, J.W.; Martin, S.A.; Secombes, C.J. Sequence and expression analysis of two T helper master transcription factors, T-bet and GATA3, in rainbow trout Oncorhynchus mykiss and analysis of their expression during bacterial and parasitic infection. Fish Shellfish Immunol. 2010, 29, 705–715. [Google Scholar] [CrossRef] [PubMed]
- Koppang, E.O.; Fischer, U.; Moore, L.; Tranulis, M.A.; Dijkstra, J.M.; Köllner, B.; Aune, L.; Jirillo, E.; Hordvik, I. Salmonid T cells assemble in the thymus, spleen and in novel interbranchial lymphoid tissue. J. Anat. 2010, 217, 728–739. [Google Scholar] [CrossRef] [PubMed]
- Costa, M.M.; Pereiro, P.; Wang, T.; Secombes, C.J.; Figueras, A.; Novoa, B. Characterization and gene expression analysis of the two main Th17 cytokines (IL-17A/F and IL-22) in turbot, Scophthalmus maximus. Dev. Comp. Immunol. 2012, 38, 505–516. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Flajnik, M.F. Re-evaluation of the immunological Big Bang. Curr Biol. 2014, 24, R1060–R1065. [Google Scholar] [CrossRef] [PubMed]
- Star, B.; Nederbragt, A.J.; Jentoft, S.; Grimholt, U.; Malmstrøm, M.; Gregers, T.F.; Rounge, T.B.; Paulsen, J.; Solbakken, M.H.; Sharma, A.; et al. The genome sequence of Atlantic cod reveals a unique immune system. Nature 2011, 477, 207–210. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hashimoto, K.; Nakanishi, T.; Kurosawa, Y. Isolation of carp genes encoding major histocompatibility complex antigens. Proc. Natl. Acad. Sci. USA 1990, 87, 6863–6867. [Google Scholar] [CrossRef] [PubMed]
- Wittamer, V.; Bertrand, J.Y.; Gutschow, P.W.; Traver, D. Characterization of the mononuclear phagocyte system in zebrafish. Blood 2011, 117, 7126–7135. [Google Scholar] [CrossRef] [PubMed]
- Dijkstra, J.M.; Kiryu, I.; Köllner, B.; Yoshiura, Y.; Ototake, M. MHC class II invariant chain homologues in rainbow trout (Oncorhynchus mykiss). Fish Shellfish Immunol. 2003, 15, 91–105. [Google Scholar] [CrossRef]
- Lewis, K.L.; del Cid, N.; Traver, D. Perspectives on antigen presenting cells in zebrafish. Dev. Comp. Immunol. 2014, 46, 63–73. [Google Scholar] [CrossRef] [PubMed]
- Rast, J.P.; Litman, G.W. T-cell receptor gene homologs are present in the most primitive jawed vertebrates. Proc. Natl. Acad. Sci. USA 1994, 91, 9248–9252. [Google Scholar] [CrossRef] [PubMed]
- Boudinot, P.; Boubekeur, S.; Benmansour, A. Rhabdovirus infection induces public and private T cell responses in teleost fish. J. Immunol. 2001, 167, 6202–6209. [Google Scholar] [CrossRef] [PubMed]
- Suetake, H.; Araki, K.; Suzuki, Y. Cloning, expression, and characterization of fugu CD4, the first ectothermic animal CD4. Immunogenetics 2004, 56, 368–374. [Google Scholar] [CrossRef] [PubMed]
- Dijkstra, J.M.; Somamoto, T.; Moore, L.; Hordvik, I.; Ototake, M.; Fischer, U. Identification and characterization of a second CD4-like gene in teleost fish. Mol. Immunol. 2006, 43, 410–419. [Google Scholar] [CrossRef] [PubMed]
- Laing, K.J.; Zou, J.J.; Purcell, M.K.; Phillips, R.; Secombes, C.J.; Hansen, J.D. Evolution of the CD4 family: Teleost fish possess two divergent forms of CD4 in addition to lymphocyte activation gene-3. J. Immunol. 2006, 177, 3939–3951. [Google Scholar] [CrossRef] [PubMed]
- Taylor, E.B.; Wilson, M.; Bengten, E. The Src tyrosine kinase Lck binds to CD2, CD4–1, and CD4–2 T cell co-receptors in channel catfish, Ictalurus punctatus. Mol. Immunol. 2015, 66, 126–138. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Moore, L.; Koppang, E.O.; Hordvik, I. Characterization of the CD3zeta, CD3gammadelta and CD3epsilon subunits of the T cell receptor complex in Atlantic salmon. Dev. Comp. Immunol. 2008, 32, 26–35. [Google Scholar] [CrossRef] [PubMed]
- Øvergård, A.C.; Nerland, A.H.; Patel, S. Cloning, characterization, and expression pattern of Atlantic halibut (Hippoglossus hippoglossus L.) CD4-2, Lck, and ZAP-70. Fish Shellfish Immunol. 2010, 29, 987–997. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fischer, U.; Dijkstra, J.M.; Köllner, B.; Kiryu, I.; Koppang, E.O.; Hordvik, I.; Sawamoto, Y.; Ototake, M. The ontogeny of MHC class I expression in rainbow trout (Oncorhynchus mykiss). Fish Shellfish Immunol. 2005, 18, 49–60. [Google Scholar] [CrossRef] [PubMed]
- Takizawa, F.; Dijkstra, J.M.; Kotterba, P.; Korytář, T.; Kock, H.; Köllner, B.; Jaureguiberry, B.; Nakanishi, T.; Fischer, U. The expression of CD8α discriminates distinct T cell subsets in teleost fish. Dev. Comp. Immunol. 2011, 35, 752–763. [Google Scholar] [CrossRef] [PubMed]
- Toda, H.; Saito, Y.; Koike, T.; Takizawa, F.; Araki, K.; Yabu, T.; Somamoto, T.; Suetake, H.; Suzuki, Y.; Ototake, M.; et al. Conservation of characteristics and functions of CD4 positive lymphocytes in a teleost fish. Dev. Comp. Immunol. 2011, 35, 650–660. [Google Scholar] [CrossRef] [PubMed]
- Picchietti, S.; Guerra, L.; Buonocore, F.; Randelli, E.; Fausto, A.M.; Abelli, L. Lymphocyte differentiation in sea bass thymus: CD4 and CD8-α gene expression studies. Fish Shellfish Immunol. 2009, 27, 50–56. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Picchietti, S.; Abelli, L.; Guerra, L.; Randelli, E.; Serafini, F.P.; Belardinelli, M.C.; Buonocore, F.; Bernini, C.; Fausto, A.M.; Scapigliati, G. MHC II-β chain gene expression studies define the regional organization of the thymus in the developing bony fish Dicentrarchus labrax (L.). Fish Shellfish Immunol. 2015, 42, 483–493. [Google Scholar] [CrossRef] [PubMed]
- Nakanishi, T. Effects of X-irradiation and thymectomy on the immune response of the marine teleost, Sebastiscus marmoratus. Dev. Comp. Immunol. 1986, 10, 519–527. [Google Scholar] [CrossRef]
- Yocum, D.; Cuchens, M.; Clem, L.W. The hapten-carrier effect in teleost fish. J. Immunol. 1975, 114, 925–927. [Google Scholar] [PubMed]
- Miller, N.W.; Sizemore, R.C.; Clem, L.W. Phylogeny of lymphocyte heterogeneity: The cellular requirements for in vitro antibody responses of channel catfish leukocytes. J. Immunol. 1985, 134, 2884–2888. [Google Scholar] [PubMed]
- Miller, N.; Wilson, M.; Bengtén, E.; Stuge, T.; Warr, G.; Clem, W. Functional and molecular characterization of teleost leukocytes. Immunol Rev. 1998, 166, 187–197. [Google Scholar] [CrossRef] [PubMed]
- Somamoto, T.; Kondo, M.; Nakanishi, T.; Nakao, M. Helper function of CD4+ lymphocytes in antiviral immunity in ginbuna crucian carp, Carassius auratus langsdorfii. Dev. Comp. Immunol. 2014, 44, 111–115. [Google Scholar] [CrossRef] [PubMed]
- Yoon, S.; Mitra, S.; Wyse, C.; Alnabulsi, A.; Zou, J.; Weerdenburg, E.M.; van der Sar, A.; Wang, D.; Secombes, C.J.; Bird, S. First demonstration of antigen induced cytokine expression by CD4-1+ lymphocytes in a poikilotherm: Studies in zebrafish (Danio rerio). PLoS ONE 2015, 10, e0126378. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Edholm, E.S.; Stafford, J.L.; Quiniou, S.M.; Waldbieser, G.; Miller, N.W.; Bengtén, E.; Wilson, M. Channel catfish, Ictalurus punctatus, CD4-like molecules. Dev. Comp. Immunol. 2007, 31, 172–187. [Google Scholar] [CrossRef] [PubMed]
- Takizawa, F.; Xu, Z.; Gómez, D.; Parra, D.; Sunyer, J.O. Novel T cell subpopulations expressing CD4-1 and CD4-2 molecules in rainbow trout. Fish Shellfish Immunol. 2013. [Google Scholar] [CrossRef]
- Kato, G.; Goto, K.; Akune, I.; Aoka, S.; Kondo, H.; Hirono, I. CD4 and CD8 homologues in Japanese flounder, Paralichthys olivaceus: Differences in the expressions and localizations of CD4-1, CD4-2, CD8α and CD8β. Dev. Comp. Immunol. 2013, 39, 293–301. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, T.; Katakura, F.; Someya, K.; Dijkstra, J.M.; Moritomo, T.; Nakanishi, T. Clonal growth of carp (Cyprinus carpio) T cells in vitro: Long-term proliferation of Th2-like cells. Fish Shellfish Immunol. 2013, 34, 433–442. [Google Scholar] [CrossRef] [PubMed]
- Fischer, U.; Koppang, E.O.; Nakanishi, T. Teleost T and NK cell immunity. Fish Shellfish Immunol. 2013, 35, 197–206. [Google Scholar] [CrossRef] [PubMed]
- Somamoto, T.; Koppang, E.O.; Fischer, U. Antiviral functions of CD8+ cytotoxic T cells in teleost fish. Dev. Comp. Immunol. 2014, 43, 197–204. [Google Scholar] [CrossRef] [PubMed]
- Kono, T.; Korenaga, H. Cytokine gene expression in CD4 positive cells of the Japanese pufferfish, Takifugu rubripes. PLoS ONE 2013, 8, e66364. [Google Scholar] [CrossRef] [PubMed]
- Jung, C.Y.; Hikima, J.; Ohtani, M.; Jang, H.B.; del Castillo, C.S.; Nho, S.W.; Cha, I.S.; Park, S.B.; Aoki, T.; Jung, T.S. Recombinant interferon-γ activates immune responses against Edwardsiella tarda infection in the olive flounder, Paralichthys olivaceus. Fish Shellfish Immunol. 2012, 33, 197–203. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Holland, J.W.; Carrington, A.; Zou, J.; Secombes, C.J. Molecular and functional characterization of IL-15 in rainbow trout Oncorhynchus mykiss: A potent inducer of IFN-γ expression in spleen leukocytes. J. Immunol. 2007, 179, 1475–1488. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Husain, M.; Hong, S.; Holland, J.W. Differential expression, modulation and bioactivity of distinct fish IL-12 isoforms: Implication towards the evolution of Th1-like immune responses. Eur. J. Immunol. 2014, 44, 1541–1551. [Google Scholar] [CrossRef] [PubMed]
- Berg, R.E.; Cordes, C.J.; Forman, J. Contribution of CD8+ T cells to innate immunity: IFN-γ secretion induced by IL-12 and IL-18. Eur. J. Immunol. 2002, 32, 2807–2816. [Google Scholar] [CrossRef]
- Fauriat, C.; Long, E.O.; Ljunggren, H.G.; Bryceson, Y.T. Regulation of human NK-cell cytokine and chemokine production by target cell recognition. Blood 2010, 115, 2167–2176. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Jiang, Y.; Wang, A.; Husain, M.; Xu, Q.; Secombes, C.J. Identification of the salmonid IL-17A/F1a/b, IL-17A/F2b, IL-17A/F3 and IL-17N genes and analysis of their expression following in vitro stimulation and infection. Immunogenetics 2015, 67, 395–412. [Google Scholar] [CrossRef] [PubMed]
- Victoratos, P.; Yiangou, M.; Avramidis, N.; Hadjipetrou, L. Regulation of cytokine gene expression by adjuvants in vivo. Clin. Exp. Immunol. 1997, 109, 569–578. [Google Scholar] [CrossRef] [PubMed]
- Yabu, T.; Toda, H.; Shibasaki, Y.; Araki, K.; Yamashita, M.; Anzai, H.; Mano, N.; Masuhiro, Y.; Hanazawa, S.; Shiba, H.; et al. Antiviral protection mechanisms mediated by ginbuna crucian carp interferon gamma isoforms 1 and 2 through two distinct interferon gamma-receptors. J. Biochem. 2011, 150, 635–648. [Google Scholar] [CrossRef] [PubMed]
- Korn, T.; Bettelli, E.; Oukka, M.; Kuchroo, V.K. IL-17 and Th17 Cells. Annu. Rev. Immunol. 2009, 27, 485–517. [Google Scholar] [CrossRef] [PubMed]
- Kumar, V.; Sharma, A. Neutrophils: Cinderella of innate immune system. Int. Immunopharmacol. 2010, 10, 1325–1334. [Google Scholar] [CrossRef] [PubMed]
- Verburg-van Kemenade, B.M.; Daly, J.G.; Groeneveld, A.; Wiegertjes, G.F. Multiple regulation of carp (Cyprinus carpio L.) macrophages and neutrophilic granulocytes by serum factors: Influence of infection with atypical Aeromonas salmonicida. Vet. Immunol. Immunopathol. 1996, 51, 189–200. [Google Scholar] [CrossRef]
- Afonso, A.; Lousada, S.; Silva, J.; Ellis, A.E.; Silva, M.T. Neutrophil and macrophage responses to inflammation in the peritoneal cavity of rainbow trout Oncorhynchus mykiss. A light and electron microscopic cytochemical study. Dis. Aquat. Organ. 1998, 34, 27–37. [Google Scholar] [CrossRef] [PubMed]
- Lieschke, G.J.; Oates, A.C.; Crowhurst, M.O.; Ward, A.C.; Layton, J.E. Morphologic and functional characterization of granulocytes and macrophages in embryonic and adult zebrafish. Blood 2001, 98, 3087–3096. [Google Scholar] [CrossRef] [PubMed]
- Palić, D.; Ostojić, J.; Andreasen, C.B.; Roth, J.A. Fish cast NETs: Neutrophil extracellular traps are released from fish neutrophils. Dev. Comp. Immunol. 2007, 31, 805–816. [Google Scholar] [CrossRef] [PubMed]
- Gunimaladevi, I.; Savan, R.; Sakai, M. Identification, cloning and characterization of interleukin-17 and its family from zebrafish. Fish Shellfish Immunol. 2006, 21, 393–403. [Google Scholar] [CrossRef] [PubMed]
- Du, L.; Feng, S.; Yin, L.; Wang, X.; Zhang, A.; Yang, K.; Zhou, H. Identification and functional characterization of grass carp IL-17A/F1: An evaluation of the immunoregulatory role of teleost IL-17A/F1. Dev. Comp. Immunol. 2015, 51, 202–211. [Google Scholar] [CrossRef] [PubMed]
- De Oliveira, S.; Reyes-Aldasoro, C.C.; Candel, S.; Renshaw, S.A.; Mulero, V.; Calado, A. Cxcl8 (IL-8) mediates neutrophil recruitment and behavior in the zebrafish inflammatory response. J. Immunol. 2013, 190, 4349–4359. [Google Scholar] [CrossRef] [PubMed]
- Brugman, S.; Witte, M.; Scholman, R.C.; Klein, M.R.; Boes, M.; Nieuwenhuis, E.E. T lymphocyte-dependent and -independent regulation of Cxcl8 expression in zebrafish intestines. J. Immunol. 2014, 192, 484–491. [Google Scholar] [CrossRef] [PubMed]
- Ribeiro, C.M.; Pontes, M.J.; Bird, S.; Chadzinska, M.; Scheer, M.; Verburg-van Kemenade, B.M.; Savelkoul, H.F.; Wiegertjes, G.F. Trypanosomiasis-induced Th17-like immune responses in carp. PLoS ONE 2010, 5, e13012. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Diaz-Rosales, P.; Costa, M.M.; Campbell, S.; Snow, M.; Collet, B.; Martin, S.A.; Secombes, C.J. Functional characterization of a nonmammalian IL-21: Rainbow trout Oncorhynchus mykiss IL-21 upregulates the expression of the Th cell signature cytokines IFN-γ, IL-10, and IL-22. J. Immunol. 2011, 186, 708–721. [Google Scholar] [CrossRef] [PubMed]
- Corripio-Miyar, Y.; Zou, J.; Richmond, H.; Secombes, C.J. Identification of interleukin-22 in gadoids and examination of its expression level in vaccinated fish. Mol. Immunol. 2009, 46, 2098–2106. [Google Scholar] [CrossRef] [PubMed]
- Monte, M.M.; Zou, J.; Wang, T.; Carrington, A.; Secombes, C.J. Cloning, expression analysis and bioactivity studies of rainbow trout (Oncorhynchus mykiss) interleukin-22. Cytokine 2011, 55, 62–73. [Google Scholar] [CrossRef] [PubMed]
- Costa, M.M.; Saraceni, P.R.; Forn-Cuní, G.; Dios, S.; Romero, A.; Figueras, A.; Novoa, B. IL-22 is a key player in the regulation of inflammation in fish and involves innate immune cells and PI3K signaling. Dev. Comp. Immunol. 2013, 41, 746–755. [Google Scholar] [CrossRef] [PubMed]
- Qi, Z.; Zhang, Q.; Wang, Z.; Zhao, W.; Chen, S.; Gao, Q. Molecular cloning, expression analysis and functional characterization of interleukin-22 in So-iny mullet, Liza haematocheila. Mol. Immunol. 2015, 63, 245–252. [Google Scholar] [CrossRef] [PubMed]
- Mughal, M.S.; Farley-Ewens, E.K.; Manning, M.J. Effects of direct immersion in antigen on immunological memory in young carp, Cyprinus carpio. Vet. Immunol. Immunopathol. 1986, 12, 181–192. [Google Scholar] [CrossRef]
- Joosten, P.H.; Engelsma, M.Y.; van der Zee, M.D.; Rombout, J.H. Induction of oral tolerance in carp (Cyprinus carpio L.) after feeding protein antigens. Vet. Immunol. Immunopathol. 1997, 60, 187–196. [Google Scholar] [CrossRef]
- Hori, S.; Nomura, T.; Sakaguchi, S. Control of regulatory T cell development by the transcription factor Foxp3. Science 2003, 299, 1057–1061. [Google Scholar] [CrossRef] [PubMed]
- Horwitz, D.A.; Zheng, S.G.; Gray, J.D. Natural and TGF-β-induced Foxp3+CD4+CD25+ regulatory T cells are not mirror images of each other. Trends Immunol. 2008, 29, 429–435. [Google Scholar] [CrossRef] [PubMed]
- Wen, Y.; Fang, W.; Xiang, L.X.; Pan, R.L.; Shao, J.Z. Identification of Treg-like cells in Tetraodon: Insight into the origin of regulatory T subsets during early vertebrate evolution. Cell Mol. Life Sci. 2011, 68, 2615–2626. [Google Scholar] [CrossRef] [PubMed]
- Sadlack, B.; Löhler, J.; Schorle, H.; Klebb, G.; Haber, H.; Sickel, E.; Noelle, R.J.; Horak, I. Generalized autoimmune disease in interleukin-2-deficient mice is triggered by an uncontrolled activation and proliferation of CD4+ T cells. Eur. J. Immunol. 1995, 25, 3053–3059. [Google Scholar] [CrossRef] [PubMed]
- Almeida, A.R.; Legrand, N.; Papiernik, M.; Freitas, A.A. Homeostasis of peripheral CD4+ T cells: IL-2R α and IL-2 shape a population of regulatory cells that controls CD4+ T cell numbers. J. Immunol. 2002, 169, 4850–4860. [Google Scholar] [CrossRef] [PubMed]
- Busse, D.; de la Rosa, M.; Hobiger, K.; Thurley, K.; Flossdorf, M.; Scheffold, A.; Höfer, T. Competing feedback loops shape IL-2 signaling between helper and regulatory T lymphocytes in cellular microenvironments. Proc. Natl. Acad. Sci. USA 2010, 107, 3058–3063. [Google Scholar] [CrossRef] [PubMed]
- Quintana, F.J.; Iglesias, A.H.; Farez, M.F.; Caccamo, M.; Burns, E.J.; Kassam, N.; Oukka, M.; Weiner, H.L. Adaptive autoimmunity and Foxp3-based immunoregulation in zebrafish. PLoS ONE 2010, 5, e9478. [Google Scholar] [CrossRef] [PubMed]
- Grayfer, L.; Hodgkinson, J.W.; Hitchen, S.J.; Belosevic, M. Characterization and functional analysis of goldfish (Carassius auratus L.) interleukin-10. Mol. Immunol. 2011, 48, 563–571. [Google Scholar] [CrossRef] [PubMed]
- Piazzon, M.C.; Savelkoul, H.S.; Pietretti, D.; Wiegertjes, G.F.; Forlenza, M. Carp II10 has anti-inflammatory activities on phagocytes, promotes proliferation of memory T cells, and regulates B cell differentiation and antibody secretion. J. Immunol. 2015, 194, 187–199. [Google Scholar] [CrossRef] [PubMed]
- Haddad, G.; Hanington, P.C.; Wilson, E.C.; Grayfer, L.; Belosevic, M. Molecular and functional characterization of goldfish (Carassius auratus L.) transforming growth factor beta. Dev. Comp. Immunol. 2008, 32, 654–663. [Google Scholar] [CrossRef] [PubMed]
- Balla, K.M.; Lugo-Villarino, G.; Spitsbergen, J.M.; Stachura, D.L.; Hu, Y.; Bañuelos, K.; Romo-Fewell, O.; Aroian, R.V.; Traver, D. Eosinophils in the zebrafish: Prospective isolation, characterization, and eosinophilia induction by helminth determinants. Blood 2010, 116, 3944–3954. [Google Scholar] [CrossRef] [PubMed]
- Fast, M.D. Fish immune responses to parasitic copepod (namely sea lice) infection. Dev. Comp. Immunol. 2014, 43, 300–312. [Google Scholar] [CrossRef] [PubMed]
- Mulero, I.; Sepulcre, M.P.; Meseguer, J.; García-Ayala, A.; Mulero, V. Histamine is stored in mast cells of most evolutionarily advanced fish and regulates the fish inflammatory response. Proc. Natl. Acad. Sci. USA 2007, 104, 19434–19439. [Google Scholar] [CrossRef] [PubMed]
- Dezfuli, B.S.; Giari, L. Mast cells in the gills and intestines of naturally infected fish: Evidence of migration and degranulation. J. Fish Dis. 2008, 31, 845–852. [Google Scholar] [CrossRef] [PubMed]
- Sfacteria, A.; Brines, M.; Blank, U. The mast cell plays a central role in the immune system of teleost fish. Mol. Immunol. 2015, 63, 3–8. [Google Scholar] [CrossRef] [PubMed]
- Prykhozhij, S.V.; Berman, J.N. The progress and promise of zebrafish as a model to study mast cells. Dev. Comp. Immunol. 2014, 46, 74–83. [Google Scholar] [CrossRef] [PubMed]
- Chettri, J.K.; Kuhn, J.A.; Jaafar, R.M.; Kania, P.W.; Møller, O.S.; Buchmann, K. Epidermal response of rainbow trout to Ichthyobodo necator: Immunohistochemical and gene expression studies indicate a Th1-/Th2-like switch. J. Fish Dis. 2014, 37, 771–783. [Google Scholar] [CrossRef] [PubMed]
- Benedicenti, O.; Collins, C.; Wang, T.; McCarthy, U.; Secombes, C.J. Which Th pathway is involved during late stage amoebic gill disease? Fish Shellfish Immunol. 2015, 46, 417–425. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.Y.; Pan, P.P.; Fang, W.; Shao, J.Z.; Xiang, L.X. Essential role of IL-4 and IL-4Rα interaction in adaptive immunity of zebrafish: Insight into the origin of Th2-like regulatory mechanism in ancient vertebrates. J. Immunol. 2012, 188, 5571–5584. [Google Scholar] [CrossRef] [PubMed]
- Lin, A.F.; Xiang, L.X.; Wang, Q.L.; Dong, W.R.; Gong, Y.F.; Shao, J.Z. The DC-SIGN of zebrafish: Insights into the existence of a CD209 homologue in a lower vertebrate and its involvement in adaptive immunity. J. Immunol. 2009, 183, 7398–7410. [Google Scholar] [CrossRef] [PubMed]
- Farrar, J.J.; Howard, M.; Fuller-Farrar, J.; Paul, W.E. Biochemical and physicochemical characterization of mouse B cell growth factor: A lymphokine distinct from interleukin 2. J. Immunol. 1983, 131, 1838–1842. [Google Scholar] [PubMed]
- Grayfer, L.; Hodgkinson, J.W.; Belosevic, M. Antimicrobial responses of teleost phagocytes and innate immune evasion strategies of intracellular bacteria. Dev. Comp. Immunol. 2014, 43, 223–242. [Google Scholar] [CrossRef] [PubMed]
- Richardson, R.; Slanchev, K.; Kraus, C.; Knyphausen, P.; Eming, S.; Hammerschmidt, M. Adult zebrafish as a model system for cutaneous wound-healing research. J. Investig. Dermatol. 2013, 133, 1655–1665. [Google Scholar] [CrossRef] [PubMed]
- Keightley, M.C.; Wang, C.H.; Pazhakh, V.; Lieschke, G.J. Delineating the roles of neutrophils and macrophages in zebrafish regeneration models. Int. J. Biochem. Cell Biol. 2014, 56, 92–106. [Google Scholar] [CrossRef] [PubMed]
- Nguyen-Chi, M.; Laplace-Builhe, B.; Travnickova, J.; Luz-Crawford, P.; Tejedor, G.; Phan, Q.T.; Duroux-Richard, I.; Levraud, J.P.; Kissa, K.; Lutfalla, G.; et al. Identification of polarized macrophage subsets in zebrafish. Elife 2015. [Google Scholar] [CrossRef] [PubMed]
- Petrie, T.A.; Strand, N.S.; Yang, C.T.; Rabinowitz, J.S.; Moon, R.T. Macrophages modulate adult zebrafish tail fin regeneration. Development 2014, 141, 2581–2591. [Google Scholar] [CrossRef] [PubMed]
- Forlenza, M.; Fink, I.R.; Raes, G.; Wiegertjes, G.F. Heterogeneity of macrophage activation in fish. Dev. Comp. Immunol. 2011, 35, 1246–1255. [Google Scholar] [CrossRef] [PubMed]
- Arts, J.A.; Tijhaar, E.J.; Chadzinska, M.; Savelkoul, H.F.; Verburg-van Kemenade, B.M. Functional analysis of carp interferon-γ: Evolutionary conservation of classical phagocyte activation. Fish Shellfish Immunol. 2010, 29, 793–802. [Google Scholar] [CrossRef] [PubMed]
- Grayfer, L.; Belosevic, M. Molecular characterization, expression and functional analysis of goldfish (Carassius aurutus L.) interferon gamma. Dev. Comp Immunol. 2009, 33, 235–246. [Google Scholar] [CrossRef] [PubMed]
- Grayfer, L.; Garcia, E.G.; Belosevic, M. Comparison of macrophage antimicrobial responses induced by type II interferons of the goldfish (Carassius auratus L.). J. Biol. Chem. 2010, 285, 23537–23547. [Google Scholar] [CrossRef] [PubMed]
- Grayfer, L.; Walsh, J.G.; Belosevic, M. Characterization and functional analysis of goldfish (Carassius auratus L.) tumor necrosis factor-alpha. Dev. Comp. Immunol. 2008, 32, 532–543. [Google Scholar] [CrossRef] [PubMed]
- Joerink, M.; Ribeiro, C.M.; Stet, R.J.; Hermsen, T.; Savelkoul, H.F.; Wiegertjes, G.F. Head kidney-derived macrophages of common carp (Cyprinus carpio L.) show plasticity and functional polarization upon differential stimulation. J. Immunol. 2006, 177, 61–69. [Google Scholar] [CrossRef] [PubMed]
- Weigert, A.; Tzieply, N.; von Knethen, A.; Johann, A.M.; Schmidt, H.; Geisslinger, G.; Brüne, B. Tumor cell apoptosis polarizes macrophages role of sphingosine-1-phosphate. Mol. Biol. Cell. 2007, 18, 3810–3819. [Google Scholar] [CrossRef] [PubMed]
- Soehnlein, O.; Lindbom, L. Phagocyte partnership during the onset and resolution of inflammation. Nat. Rev. Immunol. 2010, 10, 427–439. [Google Scholar] [CrossRef] [PubMed]
- Brown, G.D.; Taylor, P.R.; Reid, D.M.; Willment, J.A.; Williams, D.L.; Martinez-Pomares, L.; Wong, S.Y.; Gordon, S. Dectin-1 is a major beta-glucan receptor on macrophages. J. Exp. Med. 2002, 196, 407–412. [Google Scholar] [CrossRef] [PubMed]
- Municio, C.; Alvarez, Y.; Montero, O.; Hugo, E.; Rodríguez, M.; Domingo, E.; Alonso, S.; Fernández, N.; Crespo, M.S. The response of human macrophages to β-glucans depends on the inflammatory milieu. PLoS ONE 2013, 8, e62016. [Google Scholar] [CrossRef] [PubMed]
- Rieger, A.M.; Konowalchuk, J.D.; Grayfer, L.; Katzenback, B.A.; Havixbeck, J.J.; Kiemele, M.D.; Belosevic, M.; Barreda, D.R. Fish and mammalian phagocytes differentially regulate pro-inflammatory and homeostatic responses in vivo. PLoS ONE 2012, 7, e47070. [Google Scholar] [CrossRef] [PubMed]
- Szulzewsky, F.; Pelz, A.; Feng, X.; Synowitz, M.; Markovic, D.; Langmann, T.; Holtman, I.R.; Wang, X.; Eggen, B.J.; Boddeke, H.W.; et al. Glioma-associated microglia/macrophages display an expression profile different from M1 and M2 polarization and highly express Gpnmb and Spp1. PLoS ONE 2015, 10, e0116644. [Google Scholar] [CrossRef] [PubMed]
- Hendricks, D.W.; Fink, P.J. Recent thymic emigrants are biased against the T-helper type 1 and toward the T-helper type 2 effector lineage. Blood 2011, 117, 1239–1249. [Google Scholar] [CrossRef] [PubMed]
- Jørgensen, S.M.; Hetland, D.L.; Press, C.M.; Grimholt, U.; Gjøen, T. Effect of early infectious salmon anaemia virus (ISAV) infection on expression of MHC pathway genes and type I and II interferon in Atlantic salmon (Salmo salar L.) tissues. Fish Shellfish Immunol. 2007, 23, 576–588. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Shen, B.; Wu, H.; Gao, L.; Liu, Q.; Wang, Q.; Xiao, J.; Zhang, Y. Th17-like immune response in fish mucosal tissues after administration of live attenuated Vibrio anguillarum via different vaccination routes. Fish Shellfish Immunol. 2014, 37, 229–238. [Google Scholar] [CrossRef] [PubMed]
- Wegmann, T.G.; Lin, H.; Guilbert, L.; Mosmann, T.R. Bidirectional cytokine interactions in the maternal-fetal relationship: Is successful pregnancy a TH2 phenomenon? Immunol. Today 1993, 14, 353–356. [Google Scholar] [CrossRef]
- Adkins, B. Peripheral CD4+ lymphocytes derived from fetal versus adult thymic precursors differ phenotypically and functionally. J. Immunol. 2003, 171, 5157–5164. [Google Scholar] [CrossRef] [PubMed]
- Chaouat, G. The Th1/Th2 paradigm: Still important in pregnancy? Semin. Immunopathol. 2007, 29, 95–113. [Google Scholar] [CrossRef] [PubMed]
- Sykes, L.; MacIntyre, D.A.; Yap, X.J.; Teoh, T.G.; Bennett, P.R. The Th1:th2 dichotomy of pregnancy and preterm labour. Mediators Inflamm. 2012, 2012, e967629. [Google Scholar] [CrossRef] [PubMed]
- Morein, B.; Blomqvist, G.; Hu, K. Immune responsiveness in the neonatal period. J. Comp. Pathol. 2007, 137, S27–S31. [Google Scholar] [CrossRef] [PubMed]
- Halonen, M.; Lohman, I.C.; Stern, D.A.; Spangenberg, A.; Anderson, D.; Mobley, S.; Ciano, K.; Peck, M.; Wright, A.L. Th1/Th2 patterns and balance in cytokine production in the parents and infants of a large birth cohort. J. Immunol. 2009, 182, 3285–3293. [Google Scholar] [CrossRef] [PubMed]
- Spencer, S.P.; Wilhelm, C.; Yang, Q.; Hall, J.A.; Bouladoux, N.; Boyd, A.; Nutman, T.B.; Urban, J.F., Jr.; Wang, J.; Ramalingam, T.R.; et al. Adaptation of innate lymphoid cells to a micronutrient deficiency promotes type 2 barrier immunity. Science 2014, 343, 432–437. [Google Scholar] [CrossRef] [PubMed]
- Couzin-Frankel, J. Breakthrough of the year 2013. Cancer immunotherapy. Science. 2013, 342, 1432–1433. [Google Scholar] [CrossRef] [PubMed]
- Fukuda, A.; Wang, S.C.; Morris, J.P., IV; Folias, A.E.; Liou, A.; Kim, G.E.; Akira, S.; Boucher, K.M.; Firpo, M.A.; Mulvihill, S.J.; et al. Stat3 and MMP7 contribute to pancreatic ductal adenocarcinoma initiation and progression. Cancer Cell. 2011, 19, 441–455. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Yan, W.; Collins, M.A.; Bednar, F.; Rakshit, S.; Zetter, B.R.; Stanger, B.Z.; Chung, I.; Rhim, A.D.; di Magliano, M.P. Interleukin-6 is required for pancreatic cancer progression by promoting MAPK signaling activation and oxidative stress resistance. Cancer Res. 2013, 73, 6359–6374. [Google Scholar] [CrossRef] [PubMed]
- Zambirinis, C.P.; Pushalkar, S.; Saxena, D.; Miller, G. Pancreatic cancer, inflammation, and microbiome. Cancer J. 2014, 20, 195–202. [Google Scholar] [CrossRef] [PubMed]
- Sanford, D.E.; Belt, B.A.; Panni, R.Z.; Mayer, A.; Deshpande, A.D.; Carpenter, D.; Mitchem, J.B.; Plambeck-Suess, S.M.; Worley, L.A.; Goetz, B.D.; et al. Inflammatory monocyte mobilization decreases patient survival in pancreatic cancer: A role for targeting the CCL2/CCR2 axis. Clin. Cancer Res. 2013, 19, 3404–3415. [Google Scholar] [CrossRef] [PubMed]
- Zheng, L.; Xue, J.; Jaffee, E.M.; Habtezion, A. Role of immune cells and immune-based therapies in pancreatitis and pancreatic ductal adenocarcinoma. Gastroenterology 2013, 144, 1230–1240. [Google Scholar] [CrossRef] [PubMed]
- Clark, C.E.; Hingorani, S.R.; Mick, R.; Combs, C.; Tuveson, D.A.; Vonderheide, R.H. Dynamics of the immune reaction to pancreatic cancer from inception to invasion. Cancer Res. 2007, 67, 9518–9527. [Google Scholar] [CrossRef] [PubMed]
- Gardian, K.; Janczewska, S.; Olszewski, W.L.; Durlik, M. Analysis of pancreatic cancer microenvironment: Role of macrophage infiltrates and growth factors expression. J. Cancer 2012, 3, 285–291. [Google Scholar] [CrossRef] [PubMed]
- Yabushita, S.; Fukamachi, K.; Tanaka, H.; Fukuda, T.; Sumida, K.; Deguchi, Y.; Mikata, K.; Nishioka, K.; Kawamura, S.; Uwagawa, S.; et al. Metabolomic and transcriptomic profiling of human K-ras oncogene transgenic rats with pancreatic ductal adenocarcinomas. Carcinogenesis 2013, 34, 1251–1259. [Google Scholar] [CrossRef] [PubMed]
- Partecke, L.I.; Günther, C.; Hagemann, S.; Jacobi, C.; Merkel, M.; Sendler, M.; van Rooijen, N.; Käding, A.; Trung, D.N.; Lorenz, E.; et al. Induction of M2-macrophages by tumour cells and tumour growth promotion by M2-macrophages: A quid pro quo in pancreatic cancer. Pancreatology 2013, 13, 508–516. [Google Scholar] [CrossRef] [PubMed]
- Yan, C.; Huo, X.; Wang, S.; Feng, Y.; Gong, Z. Stimulation of hepatocarcinogenesis by neutrophils upon induction of oncogenic kras expression in transgenic zebrafish. J. Hepatol. 2015, 63, 420–428. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yamaguchi, T.; Takizawa, F.; Fischer, U.; Dijkstra, J.M. Along the Axis between Type 1 and Type 2 Immunity; Principles Conserved in Evolution from Fish to Mammals. Biology 2015, 4, 814-859. https://doi.org/10.3390/biology4040814
Yamaguchi T, Takizawa F, Fischer U, Dijkstra JM. Along the Axis between Type 1 and Type 2 Immunity; Principles Conserved in Evolution from Fish to Mammals. Biology. 2015; 4(4):814-859. https://doi.org/10.3390/biology4040814
Chicago/Turabian StyleYamaguchi, Takuya, Fumio Takizawa, Uwe Fischer, and Johannes M. Dijkstra. 2015. "Along the Axis between Type 1 and Type 2 Immunity; Principles Conserved in Evolution from Fish to Mammals" Biology 4, no. 4: 814-859. https://doi.org/10.3390/biology4040814
APA StyleYamaguchi, T., Takizawa, F., Fischer, U., & Dijkstra, J. M. (2015). Along the Axis between Type 1 and Type 2 Immunity; Principles Conserved in Evolution from Fish to Mammals. Biology, 4(4), 814-859. https://doi.org/10.3390/biology4040814






