Biological Sequence Representation Methods and Recent Advances: A Review
Simple Summary
Abstract
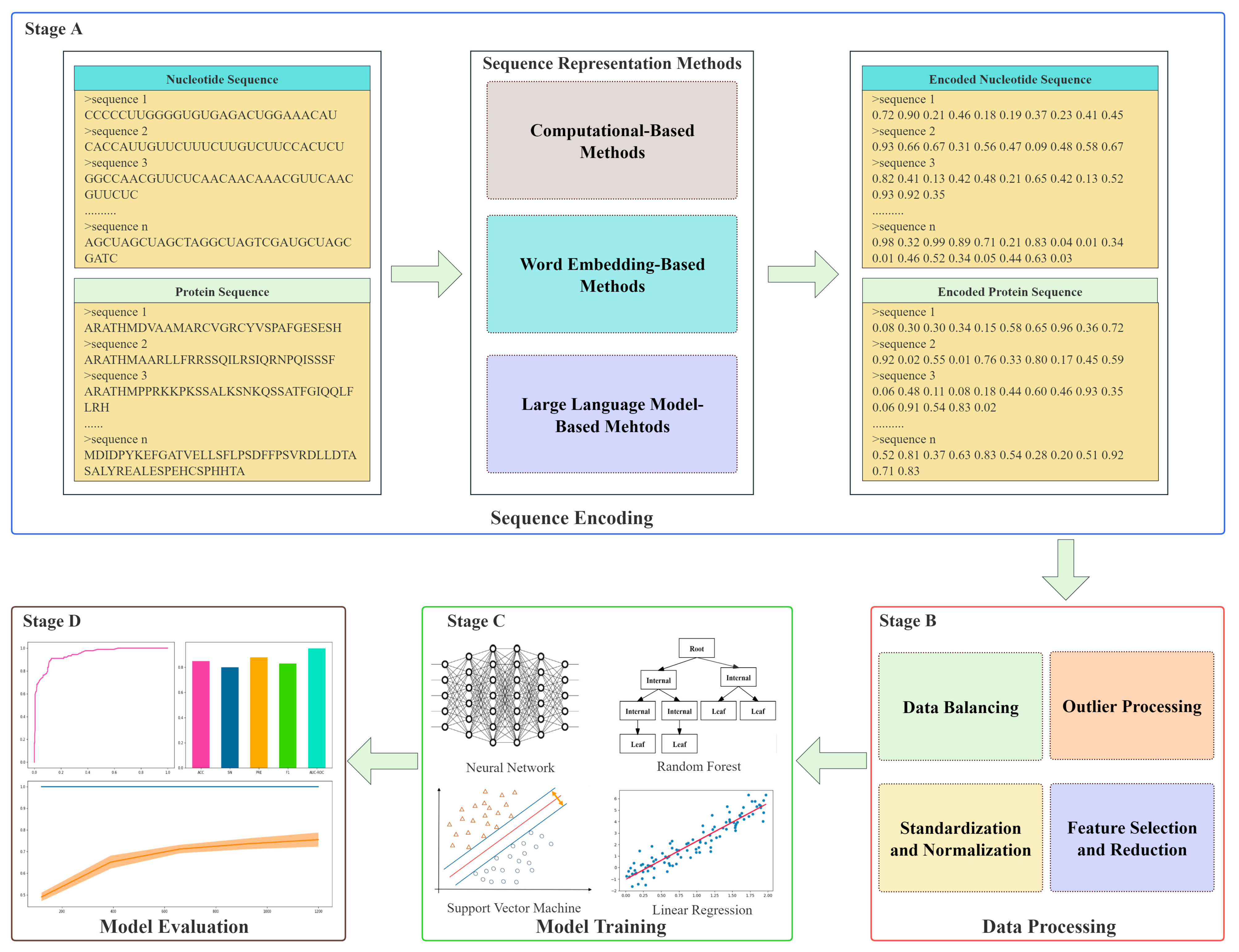
1. Introduction
2. Computational-Based Methods
2.1. K-Mer-Based Methods
2.1.1. Overview
2.1.2. Applications
2.1.3. Advantages and Limitations
2.2. Group-Based Methods
2.2.1. Overview
2.2.2. Applications
2.2.3. Advantages and Limitations
2.3. Correlation-Based Methods
2.3.1. Overview
2.3.2. Applications
2.3.3. Advantages and Limitations
2.4. PSSM-Based Methods
2.4.1. Overview
2.4.2. Applications
2.4.3. Advantages and Limitations
2.5. Structure-Based Methods
2.5.1. Overview
2.5.2. Applications
2.5.3. Advantages and Limitations
3. Word Embedding-Based Methods
3.1. Local Feature Embedding-Based Methods
3.1.1. Overview
3.1.2. Applications
3.1.3. Advantages and Limitations
3.2. Global Feature Embedding-Based Methods
3.2.1. Overview
3.2.2. Applications
3.2.3. Advantages and Limitations
4. Large Language Model-Based Methods
4.1. Self-Supervised Learning Methods
4.1.1. Overview
4.1.2. Applications
4.1.3. Advantages and Limitations
4.2. Multi-Task Learning Methods
4.2.1. Overview
4.2.2. Applications
4.2.3. Advantages and Limitations
5. Challenges and Future Directions
6. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| ML | Machine Learning |
| SVM | Support Vector Machine |
| RF | Random Forest |
| NN | Neural Network |
| LR | Linear Regression |
| AUC-ROC | Area Under the Receiver Operating Characteristic Curve |
| CNN | Convolutional Neural Network |
| LSTM | Long Short-Term Memory |
| LLM | Large Language Model |
| PPI | Protein–Protein Interaction |
| XGBOOST | eXtreme Gradient Boosting |
| PCA | Principal Component Analysis |
| PSSM | Position Specific Scoring Matrix |
| MNC | Mononucleotide Composition |
| DNC | Dinucleotide Composition |
| TNC | Trinucleotide Composition |
| AAC | Amino Acid Composition |
| DPC | Dipeptide Composition |
| TPC | Tripeptide Composition |
| CTD | Composition, Transition, and Distribution |
| CT | Conjoint Triad |
| AC | Auto-Covariance |
| CC | Cross-Covariance |
| DAC | Dinucleotide Auto-Covariance |
| TAC | Trinucleotide Auto-Covariance |
| DCC | Dinucleotide Cross-Covariance |
| TCC | Trinucleotide Cross-Covariance |
| DACC | Dinucleotide Auto- and Cross-Covariance |
| Pse-PSSM | Pseudo Position Specific Scoring Matrix |
| SPSSC | Split Protein Secondary Structure Composition |
| TS | Triplet Structure |
| MLM | Masked Language Modeling |
| PLM | Protein Language Model |
| GO | Gene Ontology |
| TAPE | Tasks Assessing Protein Embeddings |
| SHAP | SHapley Additive exPlanations |
| LIME | Local Interpretable Model-agnostic Explanations |
References
- Olson, M.V. The human genome project. Proc. Natl. Acad. Sci. USA 1993, 90, 4338–4344. [Google Scholar] [CrossRef] [PubMed]
- Lander, E.S.; Linton, L.M.; Birren, B.; Nusbaum, C.; Zody, M.C.; Baldwin, J.; Morris, W.; Meldrim, J.; Devon, K.; Santos, R.; et al. Initial sequencing and analysis of the hum1an genome. Nature 2001, 409, 860–921. [Google Scholar] [PubMed]
- Venter, J.C.; Adams, M.D.; Myers, E.W.; Li, P.W.; Mural, R.J.; Sutton, G.G.; Smith, H.O.; Yandell, M.; Evans, C.A.; Holt, R.A.; et al. The sequence of the human genome. Science 2001, 291, 1304–1351. [Google Scholar] [CrossRef]
- Moraes, F.; Góes, A. A decade of human genome project conclusion: Scientific diffusion about our genome knowledge. Biochem. Mol. Biol. Educ. 2016, 44, 215–223. [Google Scholar] [CrossRef] [PubMed]
- Aganezov, S.; Yan, S.M.; Soto, D.C.; Kirsche, M.; Zarate, S.; Avdeyev, P.; Taylor, D.J.; Shafin, K.; Shumate, A.; Xiao, C.; et al. A complete reference genome improves analysis of human genetic variation. Science 2022, 376, eabl3533. [Google Scholar] [CrossRef]
- Nurk, S.; Koren, S.; Rhie, A.; Rautiainen, M.; Bzikadze, A.V.; Mikheenko, A.; Vollger, M.R.; Altemose, N.; Uralsky, L.; Gershman, A.; et al. The complete sequence of a human genome. Science 2022, 376, 44–53. [Google Scholar] [CrossRef]
- Altemose, N.; Logsdon, G.A.; Bzikadze, A.V.; Sidhwani, P.; Langley, S.A.; Caldas, G.V.; Hoyt, S.J.; Uralsky, L.; Ryabov, F.D.; Shew, C.J.; et al. Complete genomic and epigenetic maps of human centromeres. Science 2022, 376, eabl4178. [Google Scholar] [CrossRef]
- Gershman, A.; Sauria, M.E.; Guitart, X.; Vollger, M.R.; Hook, P.W.; Hoyt, S.J.; Jain, M.; Shumate, A.; Razaghi, R.; Koren, S.; et al. Epigenetic patterns in a complete human genome. Science 2022, 376, eabj5089. [Google Scholar] [CrossRef]
- Hoyt, S.J.; Storer, J.M.; Hartley, G.A.; Grady, P.G.; Gershman, A.; de Lima, L.G.; Limouse, C.; Halabian, R.; Wojenski, L.; Rodriguez, M.; et al. From telomere to telomere: The transcriptional and epigenetic state of human repeat elements. Science 2022, 376, eabk3112. [Google Scholar] [CrossRef]
- Vollger, M.R.; Guitart, X.; Dishuck, P.C.; Mercuri, L.; Harvey, W.T.; Gershman, A.; Diekhans, M.; Sulovari, A.; Munson, K.M.; Lewis, A.P.; et al. Segmental duplications and their variation in a complete human genome. Science 2022, 376, eabj6965. [Google Scholar] [CrossRef]
- Sravani, C.; Pavani, P.; Vybhavi, G.Y.; Ramesh, G.; Farman, A. Decoding the HumanGenome: Machine Learning Techniques for DNA Sequencing Analysis. In Proceedings of the E3S Web of Conferences, Newcastle, UK, 5–8 September 2023; EDP Sciences. Volume 430, p. 01067. [Google Scholar]
- Cortes, C.; Vapnik, V. Support-vector networks. Mach. Learn. 1995, 20, 273–297. [Google Scholar] [CrossRef]
- Breiman, L. Random forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Formenti, G.; Rhie, A.; Walenz, B.P.; Thibaud-Nissen, F.; Shafin, K.; Koren, S.; Myers, E.W.; Jarvis, E.D.; Phillippy, A.M. Merfin: Improved variant filtering, assembly evaluation and polishing via k-mer validation. Nat. Methods 2022, 19, 696–704. [Google Scholar] [CrossRef]
- Xu, R.; Zhou, J.; Wang, H.; He, Y.; Wang, X.; Liu, B. Identifying DNA-binding proteins by combining support vector machine and PSSM distance transformation. BMC Syst. Biol. 2015, 9 (Suppl. 1), S10. [Google Scholar] [CrossRef]
- Zahiri, J.; Yaghoubi, O.; Mohammad-Noori, M.; Ebrahimpour, R.; Masoudi-Nejad, A. PPIevo: Protein–protein interaction prediction from PSSM based evolutionary information. Genomics 2013, 102, 237–242. [Google Scholar] [CrossRef]
- LeCun, Y.; Bottou, L.; Bengio, Y.; Haffner, P. Gradient-based learning applied to document recognition. Proc. IEEE 1998, 86, 2278–2324. [Google Scholar] [CrossRef]
- Hochreiter, S.; Schmidhuber, J. Long short-term memory. Neural Comput. 1997, 9, 1735–1780. [Google Scholar] [CrossRef] [PubMed]
- Yilmaz, S.; Toklu, S. A deep learning analysis on question classification task using Word2vec representations. Neural Comput. Appl. 2020, 32, 2909–2928. [Google Scholar] [CrossRef]
- Qiu, J.; Bernhofer, M.; Heinzinger, M.; Kemper, S.; Norambuena, T.; Melo, F.; Rost, B. ProNA2020 predicts protein–DNA, protein–RNA, and protein–protein binding proteins and residues from sequence. J. Mol. Biol. 2020, 432, 2428–2443. [Google Scholar] [CrossRef] [PubMed]
- Bahdanau, D.; Cho, K.; Bengio, Y. Neural machine translation by jointly learning to align and translate. arXiv 2014, arXiv:1409.0473. [Google Scholar]
- Abramson, J.; Adler, J.; Dunger, J.; Evans, R.; Green, T.; Pritzel, A.; Ronneberger, O.; Willmore, L.; Ballard, A.J.; Bambrick, J.; et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 2024, 630, 493–500. [Google Scholar] [CrossRef]
- Krishna, R.; Wang, J.; Ahern, W.; Sturmfels, P.; Venkatesh, P.; Kalvet, I.; Lee, G.R.; Morey-Brrows, F.S.; Anishchenko, I.; Humphreys, I.R.; et al. Generalized biomolecular modeling and design with RoseTTAFold All-Atom. Science 2024, 384, eadl2528. [Google Scholar] [CrossRef]
- Burge, C.; Campbell, A.M.; Karlin, S. Over-and under-representation of short oligonucleotides in DNA sequences. Proc. Natl. Acad. Sci. USA 1992, 89, 1358–1362. [Google Scholar] [CrossRef]
- Karlin, S.; Ladunga, I. Comparisons of eukaryotic genomic sequences. Proc. Natl. Acad. Sci. USA 1994, 91, 12832–12836. [Google Scholar] [CrossRef]
- Ghandi, M.; Lee, D.; Mohammad-Noori, M.; Beer, M.A. Enhanced regulatory sequence prediction using gapped k-mer features. PLoS Comput. Biol. 2014, 10, e1003711. [Google Scholar] [CrossRef]
- Zielezinski, A.; Vinga, S.; Almeida, J.; Karlowski, W.M. Alignment-free sequence comparison: Benefits, applications, and tools. Genome Biol. 2017, 18, 186. [Google Scholar] [CrossRef] [PubMed]
- Moeckel, C.; Mareboina, M.; Konnaris, M.A.; Chan, C.S.; Mouratidis, I.; Montgomery, A.; Chantzi, N.; Pavlopoulos, G.A.; Georgakopoulos-Soares, I. A survey of k-mer methods and applications in bioinformatics. Comput. Struct. Biotechnol. J. 2024, 23, 2289–2303. [Google Scholar] [CrossRef]
- Ghandi, M.; Mohammad-Noori, M.; Ghareghani, N.; Lee, D.; Garraway, L.; Beer, M.A. gkmSVM: An R package for gapped-kmer SVM. Bioinformatics 2016, 32, 2205–2207. [Google Scholar] [CrossRef] [PubMed]
- Wood, D.E.; Salzberg, S.L. Kraken: Ultrafast metagenomic sequence classification using exact alignments. Genome Biol. 2014, 15, R46. [Google Scholar] [CrossRef] [PubMed]
- Yan, J.; Qiu, Y.; Ribeiro dos Santos, A.M.; Yin, Y.; Li, Y.E.; Vinckier, N.; Nariai, N.; Benaglio, P.; Raman, A.; Li, X.; et al. Systematic analysis of binding of transcription factors to noncoding variants. Nature 2021, 591, 147–151. [Google Scholar] [CrossRef]
- Zhou, J.; Troyanskaya, O.G. Predicting effects of noncoding variants with deep learning–based sequence model. Nat. Methods 2015, 12, 931–934. [Google Scholar] [CrossRef]
- Yao, Y.H.; Lv, Y.P.; Li, L.; Xu, H.M.; Ji, B.B.; Chen, J.; Li, C.; Liao, B.; Nan, X.Y. Protein sequence information extraction and subcellular localization prediction with gapped k-Mer method. BMC Bioinform. 2019, 20 (Suppl. 22), 719. [Google Scholar] [CrossRef]
- Ahmad, A.; Akbar, S.; Hayat, M.; Ali, F.; Khan, S.; Sohail, M. Identification of antioxidant proteins using a discriminative intelligent model of k-space amino acid pairs based descriptors incorporating with ensemble feature selection. Biocybern. Biomed. Eng. 2022, 42, 727–735. [Google Scholar] [CrossRef]
- Bae, D.; Kim, M.; Seo, J.; Nam, H. AI-guided discovery and optimization of antimicrobial peptides through species-aware language model. Brief. Bioinform. 2025, 26, bbaf343. [Google Scholar] [CrossRef]
- Shen, J.; Zhang, J.; Luo, X.; Zhu, W.; Yu, K.; Chen, K.; Li, Y.; Jiang, H. Predicting protein–protein interactions based only on sequences information. Proc. Natl. Acad. Sci. USA 2007, 104, 4337–4341. [Google Scholar] [CrossRef] [PubMed]
- Dubchak, I.; Muchnik, I.; Mayor, C.; Dralyuk, I.; Kim, S.H. Recognition of a protein fold in the context of the SCOP classification. Proteins Struct. Funct. Bioinform. 1999, 35, 401–407. [Google Scholar] [CrossRef]
- Chothia, C.; Finkelstein, A.V. The classification and origins of protein folding patterns. Annu. Rev. Biochem. 1990, 59, 1007–1035. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Yan, W.; Yang, Y. DeepTP: A deep learning model for thermophilic protein prediction. Int. J. Mol. Sci. 2023, 24, 2217. [Google Scholar] [CrossRef]
- Yan, K.; Lv, H.; Wen, J.; Guo, Y.; Xu, Y.; Liu, B. PreTP-Stack: Prediction of therapeutic peptide based on the stacked ensemble learning. IEEE/ACM Trans. Comput. Biol. Bioinform. 2022, 20, 1337–1344. [Google Scholar] [CrossRef]
- Sureyya Rifaioglu, A.; Doğan, T.; Jesus Martin, M.; Cetin-Atalay, R.; Atalay, V. DEEPred: Automated protein function prediction with multi-task feed-forward deep neural networks. Sci. Rep. 2019, 9, 7344. [Google Scholar] [CrossRef]
- Wold, S.; Jonsson, J.; Sjörström, M.; Sandberg, M.; Rännar, S. DNA and peptide sequences and chemical processes multivariately modelled by principal component analysis and partial least-squares projections to latent structures. Anal. Chim. Acta 1993, 277, 239–253. [Google Scholar] [CrossRef]
- Guo, Y.; Yu, L.; Wen, Z.; Li, M. Using support vector machine combined with auto covariance to predict protein–protein interactions from protein sequences. Nucleic Acids Res. 2008, 36, 3025–3030. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Xiao, X.; Yu, D.J.; Jia, J.; Qiu, W.R.; Chou, K.C. pRNAm-PC: Predicting N6-methyladenosine sites in RNA sequences via physical–chemical properties. Anal. Biochem. 2016, 497, 60–67. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Liu, F.; Fang, L.; Wang, X.; Chou, K.C. repDNA: A Python package to generate various modes of feature vectors for DNA sequences by incorporating user-defined physicochemical properties and sequence-order effects. Bioinformatics 2015, 31, 1307–1309. [Google Scholar] [CrossRef]
- Dong, Q.; Zhou, S.; Guan, J. A new taxonomy-based protein fold recognition approach based on autocross-covariance transformation. Bioinformatics 2009, 25, 2655–2662. [Google Scholar] [CrossRef]
- Wang, X.; Liu, Y.; Li, J.; Wang, G. StackCirRNAPred: Computational classification of long circRNA from other lncRNA based on stacking strategy. BMC Bioinform. 2022, 23, 563. [Google Scholar] [CrossRef]
- Khan, S.; Uddin, I.; Noor, S.; AlQahtani, S.A.; Ahmad, N. N6-methyladenine identification using deep learning and discriminative feature integration. BMC Med. Genom. 2025, 18, 58. [Google Scholar] [CrossRef]
- Uddin, I.; Awan, H.H.; Khalid, M.; Khan, S.; Akbar, S.; Sarker, M.R.; Abdolrasol, M.G.; Alghamdi, T.A. A hybrid residue based sequential encoding mechanism with XGBoost improved ensemble model for identifying 5-hydroxymethylcytosine modifications. Sci. Rep. 2024, 14, 20819. [Google Scholar] [CrossRef]
- An, H.E.; Mun, M.H.; Malik, A.; Kim, C.B. Development of a two-layer machine learning model for the forensic application of legal and illegal poppy classification based on sequence data. Forensic Sci. Int. Genet. 2024, 71, 103061. [Google Scholar] [CrossRef] [PubMed]
- Gribskov, M.; McLachlan, A.D.; Eisenberg, D. Profile analysis: Detection of distantly related proteins. Proc. Natl. Acad. Sci. USA 1987, 84, 4355–4358. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef]
- Chou, K.C.; Shen, H.B. MemType-2L: A web server for predicting membrane proteins and their types by incorporating evolution information through Pse-PSSM. Biochem. Biophys. Res. Commun. 2007, 360, 339–345. [Google Scholar] [CrossRef]
- Liu, T.; Zheng, X.; Wang, J. Prediction of protein structural class for low-similarity sequences using support vector machine and PSI-BLAST profile. Biochimie 2010, 92, 1330–1334. [Google Scholar] [CrossRef]
- Réau, M.; Renaud, N.; Xue, L.C.; Bonvin, A.M. DeepRank-GNN: A graph neural network framework to learn patterns in protein–protein interfaces. Bioinformatics 2023, 39, btac759. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, C. PLM-ATG: Identification of Autophagy Proteins by Integrating Protein Language Model Embeddings with PSSM-Based Features. Molecules 2025, 30, 1704. [Google Scholar] [CrossRef]
- Zhang, Y.; Yu, L.; Xue, L.; Liu, F.; Jing, R.; Luo, J. Optimizing lipocalin sequence classification with ensemble deep learning models. PLoS ONE 2025, 20, e0319329. [Google Scholar] [CrossRef]
- Zhang, S. Accurate prediction of protein structural classes by incorporating PSSS and PSSM into Chou’s general PseAAC. Chemom. Intell. Lab. Syst. 2015, 142, 28–35. [Google Scholar] [CrossRef]
- Shen, B.; Zhang, H.; Li, C.; Zhao, T.; Liu, Y. Deep Learning Method for RNA Secondary Structure Prediction with Pseudoknots Based on Large-Scale Data. J. Healthc. Eng. 2021, 2021, 1–9. [Google Scholar] [CrossRef]
- Hofacker, I.L.; Fontana, W.; Stadler, P.F.; Bonhoeffer, L.S.; Tacker, M.; Schuster, P. Fast folding and comparison of RNA secondary structures. Monatshefte Chem. Chem. Mon. 1994, 125, 167–188. [Google Scholar] [CrossRef]
- Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 2003, 31, 3406–3415. [Google Scholar] [CrossRef] [PubMed]
- Zuker, M.; Stiegler, P. Optimal computer folding of large RNA sequences using thermodynamics and auxiliary information. Nucleic Acids Res. 1981, 9, 133–148. [Google Scholar] [CrossRef]
- Zuker, M.; Jaeger, J.A.; Turner, D.H. A comparison of optimal and suboptimal RNA secondary structures predicted by free energy minimization with structures determined by phylogenetic comparison. Nucleic Acids Res. 1991, 19, 2707–2714. [Google Scholar] [CrossRef]
- Xue, C.; Li, F.; He, T.; Liu, G.P.; Li, Y.; Zhang, X. Classification of real and pseudo microRNA precursors using local structure-sequence features and support vector machine. BMC Bioinform. 2005, 6, 310. [Google Scholar] [CrossRef] [PubMed]
- Pengcheng, S. Prediction of membrane protein types based on fusion features and voting ensemble learning. Hans J. Comput. Biol. 2022, 11, 49. [Google Scholar]
- Cuff, J.A.; Barton, G.J. Evaluation and improvement of multiple sequence methods for protein secondary structure prediction. Proteins Struct. Funct. Bioinform. 1999, 34, 508–519. [Google Scholar] [CrossRef]
- Liu, B.; Fang, L.; Liu, F.; Wang, X.; Chen, J.; Chou, K.C. Identification of real microRNA precursors with a pseudo structure status composition approach. PLoS ONE 2015, 10, e0121501. [Google Scholar] [CrossRef]
- Zhang, Y.; Yu, L.; Jing, R.; Han, B.; Luo, J. Fast and efficient design of deep neural networks for predicting n7-methylguanosine sites using autobioseqpy. ACS Omega 2023, 8, 19728–19740. [Google Scholar] [CrossRef]
- Min, H.; Xin, X.H.; Gao, C.Q.; Wang, L.; Du, P.F. XGEM: Predicting essential miRNAs by the ensembles of various sequence-based classifiers with XGBoost algorithm. Front. Genet. 2022, 13, 877409. [Google Scholar] [CrossRef]
- Yu, B.; Wang, X.; Zhang, Y.; Gao, H.; Wang, Y.; Liu, Y.; Gao, X. RPI-MDLStack: Predicting RNA–protein interactions through deep learning with stacking strategy and LASSO. Appl. Soft Comput. 2022, 120, 108676. [Google Scholar] [CrossRef]
- Iuchi, H.; Matsutani, T.; Yamada, K.; Iwano, N.; Sumi, S.; Hosoda, S.; Zhao, S.; Fukunaga, T.; Hamada, M. Representation learning applications in biological sequence analysis. Comput. Struct. Biotechnol. J. 2021, 19, 3198–3208. [Google Scholar] [CrossRef]
- Hamid, M.N.; Friedberg, I. Identifying antimicrobial peptides using word embedding with deep recurrent neural networks. Bioinformatics 2019, 35, 2009–2016. [Google Scholar] [CrossRef]
- Mikolov, T.; Sutskever, I.; Chen, K.; Corrado, G.S.; Dean, J. Distributed representations of words and phrases and their compositionality. Adv. Neural Inf. Process. Syst. 2013, 26, 1–9. [Google Scholar]
- Asgari, E.; Mofrad, M.R. Continuous distributed representation of biological sequences for deep proteomics and genomics. PLoS ONE 2015, 10, e0141287. [Google Scholar] [CrossRef]
- Ng, P. dna2vec: Consistent vector representations of variable-length k-mers. arXiv 2017, arXiv:1701.06279. [Google Scholar]
- Zulfiqar, H.; Guo, Z.; Ahmad, R.M.; Ahmed, Z.; Cai, P.; Chen, X.; Zhang, Y.; Lin, H.; Shi, Z. Deep-STP: A deep learning-based approach to predict snake toxin proteins by using word embeddings. Front. Med. 2024, 10, 1291352. [Google Scholar] [CrossRef]
- Li, H.; Zhang, S.; Chen, L.; Pan, X.; Li, Z.; Huang, T.; Cai, Y.D. Identifying functions of proteins in mice with functional embedding features. Front. Genet. 2022, 13, 909040. [Google Scholar] [CrossRef]
- Cao, L.; Liu, P.; Chen, J.; Deng, L. Prediction of transcription factor binding sites using a combined deep learning approach. Front. Oncol. 2022, 12, 893520. [Google Scholar] [CrossRef]
- Le, Q.; Mikolov, T. Distributed representations of sentences and documents. In Proceedings of the International Conference on Machine Learning, Beijing, China, 18 June 2014; pp. 1188–1196. [Google Scholar]
- Pennington, J.; Socher, R.; Manning, C.D. Glove: Global vectors for word representation. In Proceedings of the 2014 Conference on Empirical Methods in Natural Language Processing (EMNLP), Doha, Qatar, 25–29 October 2014; pp. 1532–1543. [Google Scholar]
- Yang, X.; Wuchty, S.; Liang, Z.; Ji, L.; Wang, B.; Zhu, J.; Zhang, Z.; Dong, Y. Multi-modal features-based human-herpesvirus protein–protein interaction prediction by using LightGBM. Brief. Bioinform. 2024, 25, 1–13. [Google Scholar] [CrossRef]
- Tran, H.N.; Nguyen, P.X.; Guo, F.; Wang, J. Prediction of protein–protein interactions based on integrating deep learning and feature fusion. Int. J. Mol. Sci. 2024, 25, 5820. [Google Scholar] [CrossRef]
- Yang, Y.; Hou, Z.; Ma, Z.; Li, X.; Wong, K.C. iCircRBP-DHN: Identification of circRNA-RBP interaction sites using deep hierarchical network. Brief. Bioinform. 2021, 22, bbaa274. [Google Scholar] [CrossRef]
- Ji, Y.; Sun, J.; Xie, J.; Wu, W.; Shuai, S.C.; Zhao, Q.; Chen, W. m5UMCB: Prediction of RNA 5-methyluridine sites using multi-scale convolutional neural network with BiLSTM. Comput. Biol. Med. 2024, 168, 107793. [Google Scholar] [CrossRef]
- Mahmud, S.H.; Goh, K.O.; Hosen, M.F.; Nandi, D.; Shoombuatong, W. Deep-WET: A deep learning-based approach for predicting DNA-binding proteins using word embedding techniques with weighted features. Sci. Rep. 2024, 14, 2961. [Google Scholar] [CrossRef]
- Ruffolo, J.A.; Madani, A. Designing proteins with language models. Nat. Biotechnol. 2024, 42, 200–202. [Google Scholar] [CrossRef] [PubMed]
- Vaswani, A.; Shazeer, N.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A.N.; Kaiser, Ł.; Polosukhin, I. Attention is all you need. Adv. Neural Inf. Process. Syst. 2017, 30, 5998–6008. [Google Scholar]
- Rives, A.; Meier, J.; Sercu, T.; Goyal, S.; Lin, Z.; Liu, J.; Guo, D.; Ott, M.; Zitnick, C.L.; Ma, J.; et al. Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences. Proc. Natl. Acad. Sci. USA 2021, 118, e2016239118. [Google Scholar] [CrossRef]
- Elnaggar, A.; Heinzinger, M.; Dallago, C.; Rehawi, G.; Wang, Y.; Jones, L.; Gibbs, T.; Feher, T.; Angerer, C.; Steinegger, M.; et al. Prottrans: Toward understanding the language of life through self-supervised learning. IEEE Trans. Pattern Anal. Mach. Intell. 2021, 44, 7112–7127. [Google Scholar] [CrossRef] [PubMed]
- Shen, T.; Hu, Z.; Sun, S.; Liu, D.; Wong, F.; Wang, J.; Chen, J.; Wang, Y.; Hong, L.; Xiao, J.; et al. Accurate RNA 3D structure prediction using a language model-based deep learning approach. Nat. Methods 2024, 21, 2287–2298. [Google Scholar] [CrossRef]
- Lv, L.; Lin, Z.; Li, H.; Liu, Y.; Cui, J.; Chen, C.Y.; Yuan, L.; Tian, Y. Prollama: A protein large language model for multi-task protein language processing. IEEE Trans. Artif. Intell. Early Access 2025, 1–12. [Google Scholar] [CrossRef]
- Jang, Y.J.; Qin, Q.Q.; Huang, S.Y.; Peter, A.T.; Ding, X.M.; Kornmann, B. Accurate prediction of protein function using statistics-informed graph networks. Nat. Commun. 2024, 15, 6601. [Google Scholar] [CrossRef]
- Rao, R.; Bhattacharya, N.; Thomas, N.; Duan, Y.; Chen, P.; Canny, J.; Abbeel, P.; Song, Y. Evaluating protein transfer learning with TAPE. Adv. Neural Inf. Process. Syst. 2019, 32, 9689–9701. [Google Scholar]
- Zhang, N.; Bi, Z.; Liang, X.; Cheng, S.; Hong, H.; Deng, S.; Lian, J.; Zhang, Q.; Chen, H. Ontoprotein: Protein pretraining with gene ontology embedding. arXiv 2022, arXiv:2201.11147. [Google Scholar] [CrossRef]
- Wang, N.; Bian, J.; Li, Y.; Li, X.; Mumtaz, S.; Kong, L.; Xiong, H. Multi-purpose RNA language modelling with motif-aware pretraining and type-guided fine-tuning. Nat. Mach. Intell. 2024, 6, 548–557. [Google Scholar] [CrossRef]
- Hayes, T.; Rao, R.; Akin, H.; Sofroniew, N.J.; Oktay, D.; Lin, Z.; Verkuil, R.; Tran, V.Q.; Deaton, J.; Wiggert, M.; et al. Simulating 500 million years of evolution with a language model. Science 2025, 387, 850–858. [Google Scholar] [CrossRef]
- Lam, H.Y.; Ong, X.E.; Mutwil, M. Large language models in plant biology. Trends Plant Sci. 2024, 29, 1145–1155. [Google Scholar] [CrossRef]
- Li, W.; Mao, Z.; Xiao, Z.; Liao, X.; Koffas, M.; Chen, Y.; Tang, Y.J. Large language model for knowledge synthesis and AI-enhanced biomanufacturing. Trends Biotechnol. 2025, 43, 1864–1875. [Google Scholar] [CrossRef]
- Hu, E.J.; Shen, Y.; Wallis, P.; Allen-Zhu, Z.; Li, Y.; Wang, S.; Wang, L.; Chen, W. Lora: Low-rank adaptation of large language models. ICLR 2022, 1, 3. [Google Scholar]
- Suzuki, S.; Horie, K.; Amagasa, T.; Fukuda, N. Genomic language models with k-mer tokenization strategies for plant genome annotation and regulatory element strength prediction. Plant Mol. Biol. 2025, 115, 100. [Google Scholar] [CrossRef]
- Silva, J.C.; Schuster, L.; Sexson, N.; Erdem, M.; Hulk, R.; Kirst, M.; Resende, M.F.; Dias, R. InteracTor: Feature Engineering and Explainable AI for Profiling Protein Structure-Interaction-Function Relationships. bioRxiv 2025. [Google Scholar] [CrossRef]
- Dickinson, Q.; Meyer, J.G. Positional SHAP (PoSHAP) for Interpretation of machine learning models trained from biological sequences. PLoS Comput. Biol. 2022, 18, e1009736. [Google Scholar] [CrossRef]
- Clarke, D.J.; Marino, G.B.; Deng, E.Z.; Xie, Z.; Evangelista, J.E.; Ma’ayan, A. Rummagene: Massive mining of gene sets from supporting materials of biomedical research publications. Commun. Biol. 2024, 7, 482. [Google Scholar] [CrossRef]
- Song, B.; Li, Z.; Lin, X.; Wang, J.; Wang, T.; Fu, X. Pretraining model for biological sequence data. Brief. Funct. Genom. 2021, 20, 181–195. [Google Scholar] [CrossRef]
- Stock, M.; Van Criekinge, W.; Boeckaerts, D.; Taelman, S.; Van Haeverbeke, M.; Dewulf, P.; De Baets, B. Hyperdimensional computing: A fast, robust, and interpretable paradigm for biological data. PLoS Comput. Biol. 2024, 20, e1012426. [Google Scholar] [CrossRef] [PubMed]
- Tam, B.; Qin, Z.; Zhao, B.; Sinha, S.; Lei, C.L.; Wang, S.M. Classification of MLH1 Missense VUS Using Protein Structure-Based Deep Learning-Ramachandran Plot-Molecular Dynamics Simulations Method. Int. J. Mol. Sci. 2024, 25, 850. [Google Scholar] [CrossRef] [PubMed]
- Tang, Z.; Huang, J.; Chen, G.; Chen, C.Y. Comprehensive View Embedding Learning for Single-Cell Multimodal Integration. In Proceedings of the AAAI Conference on Artificial Intelligence, Vancouver, BC, Canada, 24 March 2024; Volume 38, pp. 15292–15300. [Google Scholar]
| Methods | Core Applications | Advantages | Limitations |
|---|---|---|---|
| k-mer-based | Genome assembly, motif discovery, sequence classification | Computationally efficient, captures local patterns | High dimensionality, limited long-range dependency capture |
| Group-based | Protein function prediction, Protein annotation, protein–protein interaction prediction | Encodes physicochemical properties, biologically interpretable | Sparsity in long sequences, parameter optimization needed |
| Correlation-based | RNA classification, epigenetic modification prediction | Models complex dependencies, robust for multi-property interactions | High computational cost, limited for RNA trinucleotide correlations |
| PSSM-based | Protein structure/function prediction, PPI prediction | Leverages evolutionary conservation, robust feature extraction | Dependent on alignment quality, computationally intensive |
| Structure-based | RNA modification prediction, protein function prediction | Captures local structural motifs, biologically meaningful | Relies on accurate structural predictions, limited global context |
| Methods | Core Applications | Advantages | Limitations |
|---|---|---|---|
| Local feature embedding-based | Protein sequence classification, transcription factor binding prediction, gene annotation | Captures short-range patterns, robust to sequence length variability, computationally efficient | Limited to local dependencies, requires large training datasets, lacks direct biological interpretability |
| Global feature embedding-based | Protein function prediction, RNA methylation site prediction, regulatory RNA identification | Models sequence-wide context, adaptable to diverse tasks, robust to variable sequence lengths | Misses fine-grained local patterns, computationally intensive, sensitive to dataset quality |
| Methods | Core Applications | Advantages | Limitations |
|---|---|---|---|
| Self-supervised learning | RNA/protein structure prediction, binding site identification, functional annotation | Models complex sequence relationships, generalizable with unlabeled data, high predictive accuracy | High computational cost, requires large datasets |
| Multi-task learning | RNA type/structure classification, protein function/structure prediction, cross-modal analysis | Integrates sequence, structure, and function data, adaptable to diverse tasks, enhanced by multi-modal learning | Complex training process, resource-intensive, sensitive to annotated dataset quality |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, H.; Shi, Y.; Wang, Y.; Yang, X.; Li, K.; Im, S.-K.; Han, Y. Biological Sequence Representation Methods and Recent Advances: A Review. Biology 2025, 14, 1137. https://doi.org/10.3390/biology14091137
Zhang H, Shi Y, Wang Y, Yang X, Li K, Im S-K, Han Y. Biological Sequence Representation Methods and Recent Advances: A Review. Biology. 2025; 14(9):1137. https://doi.org/10.3390/biology14091137
Chicago/Turabian StyleZhang, Hongwei, Yan Shi, Yapeng Wang, Xu Yang, Kefeng Li, Sio-Kei Im, and Yu Han. 2025. "Biological Sequence Representation Methods and Recent Advances: A Review" Biology 14, no. 9: 1137. https://doi.org/10.3390/biology14091137
APA StyleZhang, H., Shi, Y., Wang, Y., Yang, X., Li, K., Im, S.-K., & Han, Y. (2025). Biological Sequence Representation Methods and Recent Advances: A Review. Biology, 14(9), 1137. https://doi.org/10.3390/biology14091137






