Dynamic Species Distribution Modeling Reveals the Pivotal Role of Human-Mediated Long-Distance Dispersal in Plant Invasion
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Ecological Process Model
2.1.1. General Structure
- -
- -
- if (within-cell dispersal),
- -
- determines the proportion of seeds from i’ that reach i if the latter is adjacent (short-distance dispersal), noted
- -
- , where is the proportion of urban habitat area in cell i’, determines the proportion of seeds from i’ transported via long-distance dispersal to i, i.e., when i is not adjacent to i’, noted . In other words, long-distance dispersal diffuses a portion of seeds homogeneously and instantaneously across the domain.
- -
- The proportion of seeds from i participating in local recruitment is set as the reference in this parametrization.
2.1.2. Age-Structured Fecundity
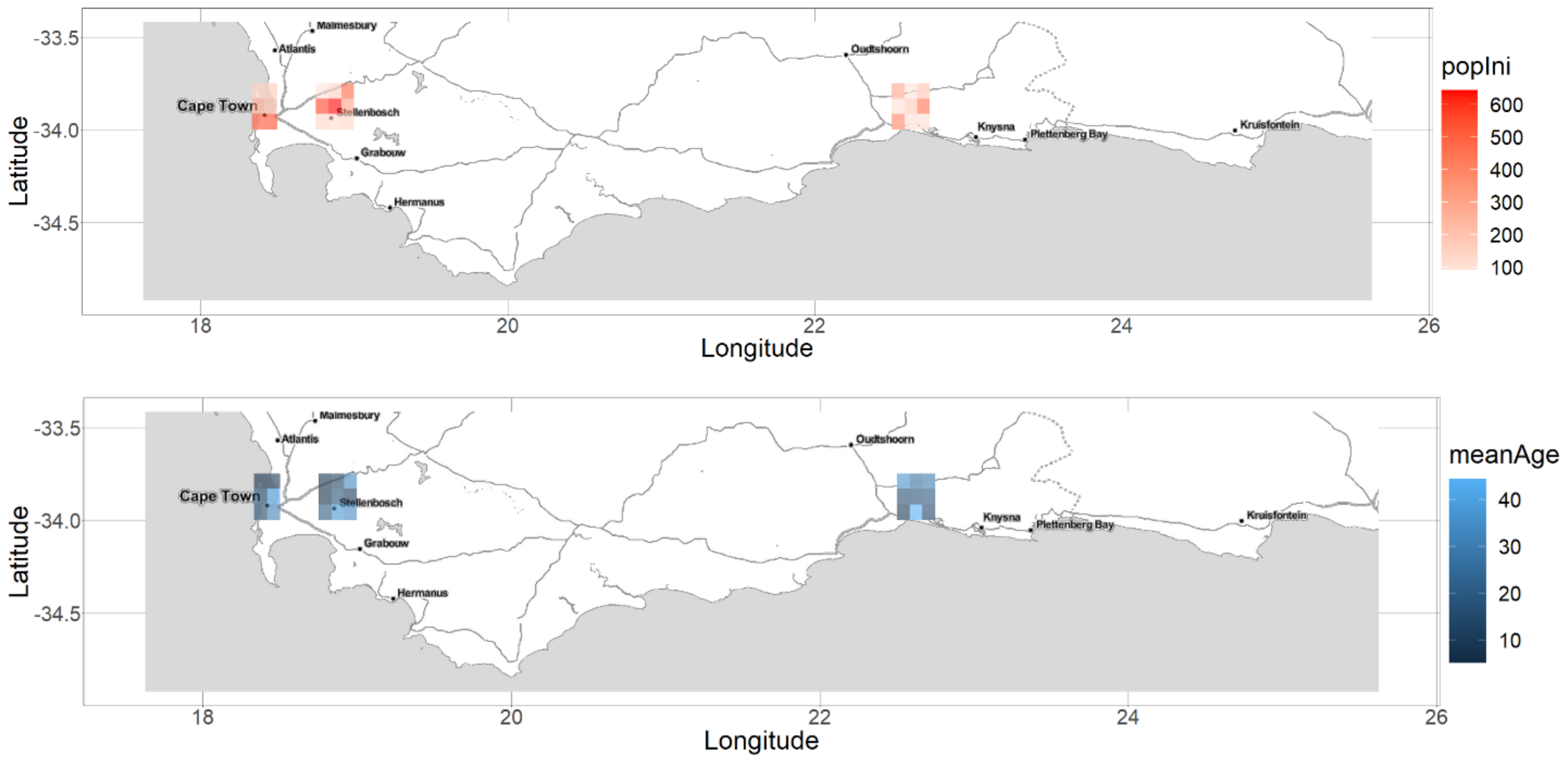
2.1.3. Initial Populations
2.2. Sampling Process Model
2.3. Data
2.3.1. Temporal and Spatial Extent
2.3.2. Occurrence Data
2.3.3. Environmental Variables
2.4. Model Fitting, Convergence Assessment and Posterior Samples
2.5. Output Representations
3. Results
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
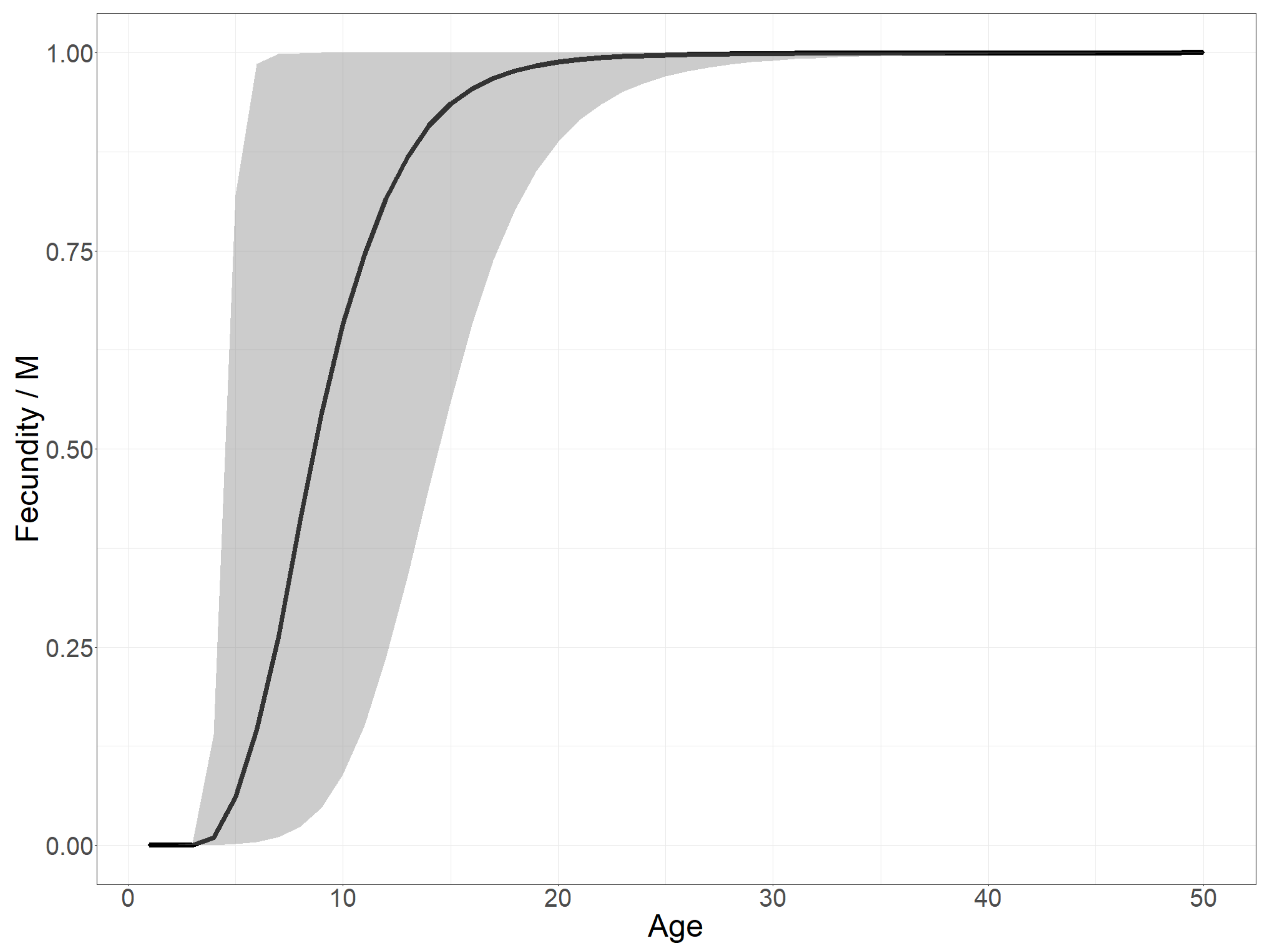
Appendix A.1. Illustration of Fecundity Function
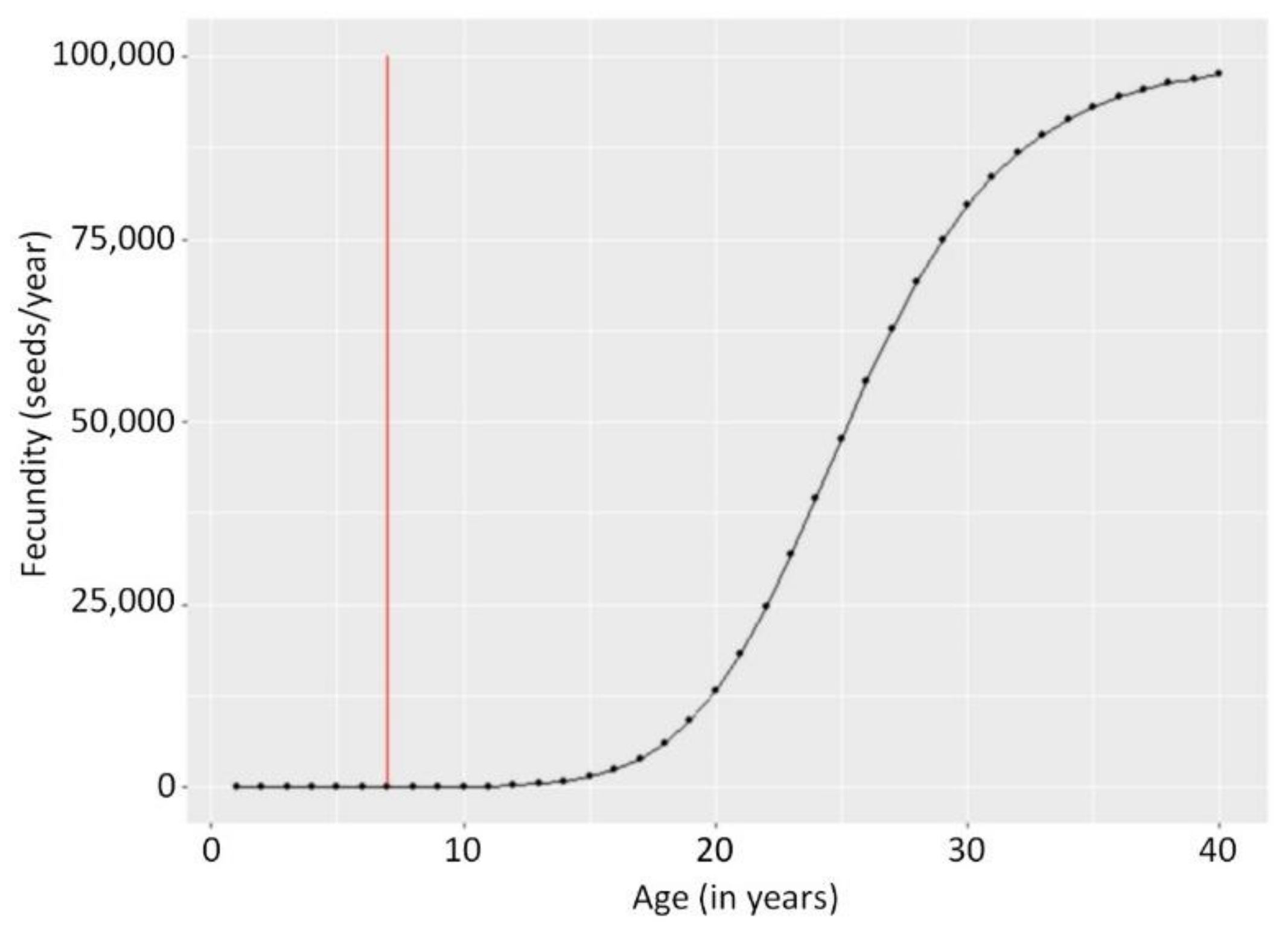
Appendix A.2. Parameter Prior Distributions
- -
- -
- -
- , which in practice does not constrain the potential functional shapes of the fecundity as an increase of this factor makes the shape converge quickly towards a step function.
- -
- -
- : the expectation of is then approximately 200,000 seeds/plant/year, and its standard deviation is 50,000, which is coherent with prior knowledge on woody plant fecundity while acknowledging a large uncertainty.
- -
- : the expectation of is 9.5 million plants/cell (of 10 × 10 km). This number is very close to the one taken for Acacia longifolia from [16]. The standard deviation is set so that the standard deviation of is of approximately 8 million plants/cell, acknowledging a large uncertainty around this number.
- -
- : the lower bound is defined such that all plants of an approximately saturated cell population will certainly die before reaching the maximal age, set as 50 in our application.
- -
- -
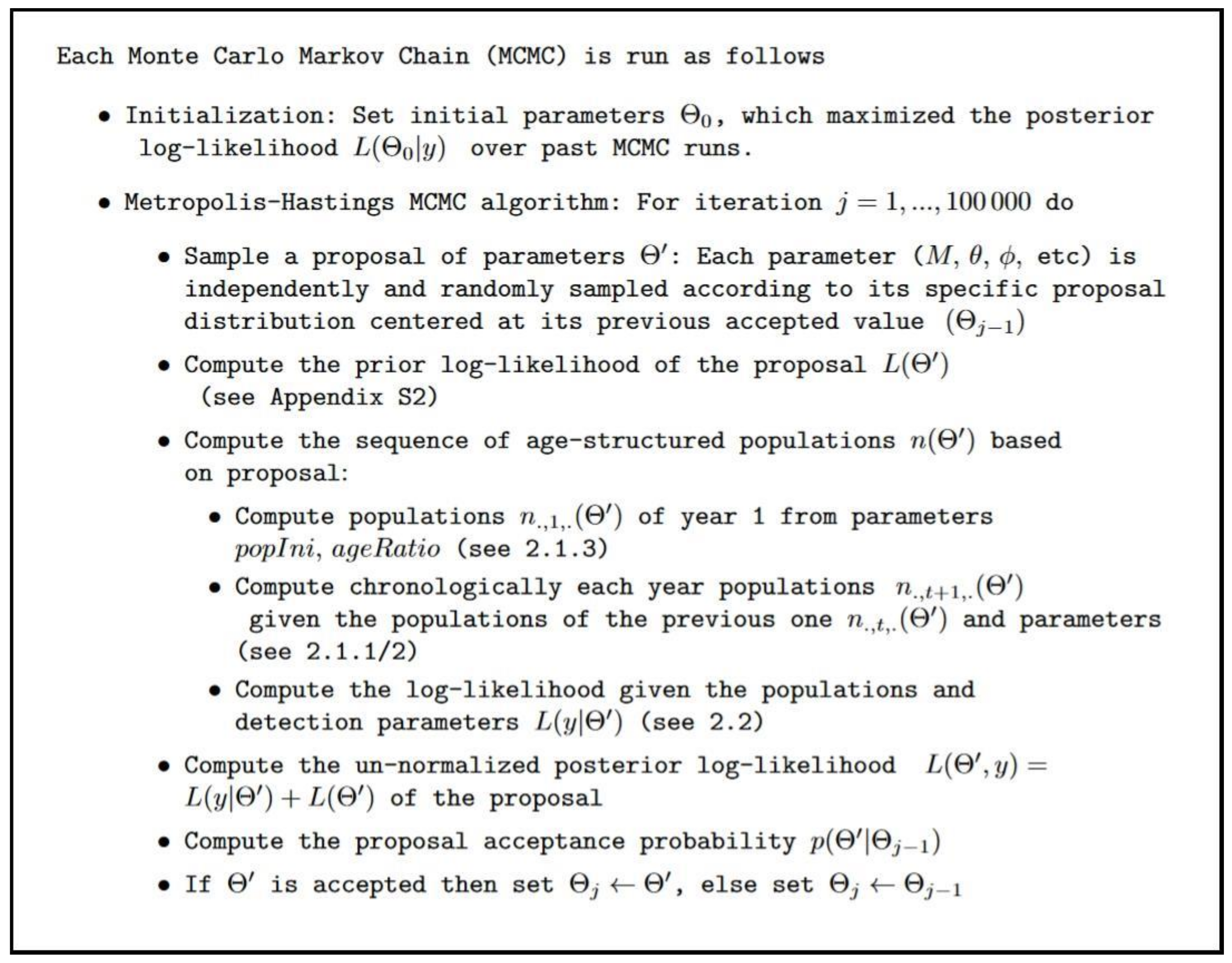
Appendix A.3. MCMC Procedure and Convergence Tests


| Parameter | PSRF’s Point Estimate | PSRF’s Upper Confidence Interval Bound |
|---|---|---|
| 1.99 | 3.37 | |
| 1.54 | 4.63 | |
| 1.75 | 2.82 | |
| 1.50 | 2.24 | |
| 2.1 | 3.50 | |
| 1.43 | 1.87 | |
| 1.08 | 1.17 | |
| 1.36 | 1.82 | |
| 1.18 | 1.39 | |
| 1.07 | 1.15 | |
| 1.61 | 2.41 | |
| 1.40 | 1.84 | |
| 1.48 | 2.15 | |
| 1.42 | 1.86 | |
| 1.53 | 2.07 | |
| 1.37 | 1.76 |
Appendix A.4. Parameter Estimability


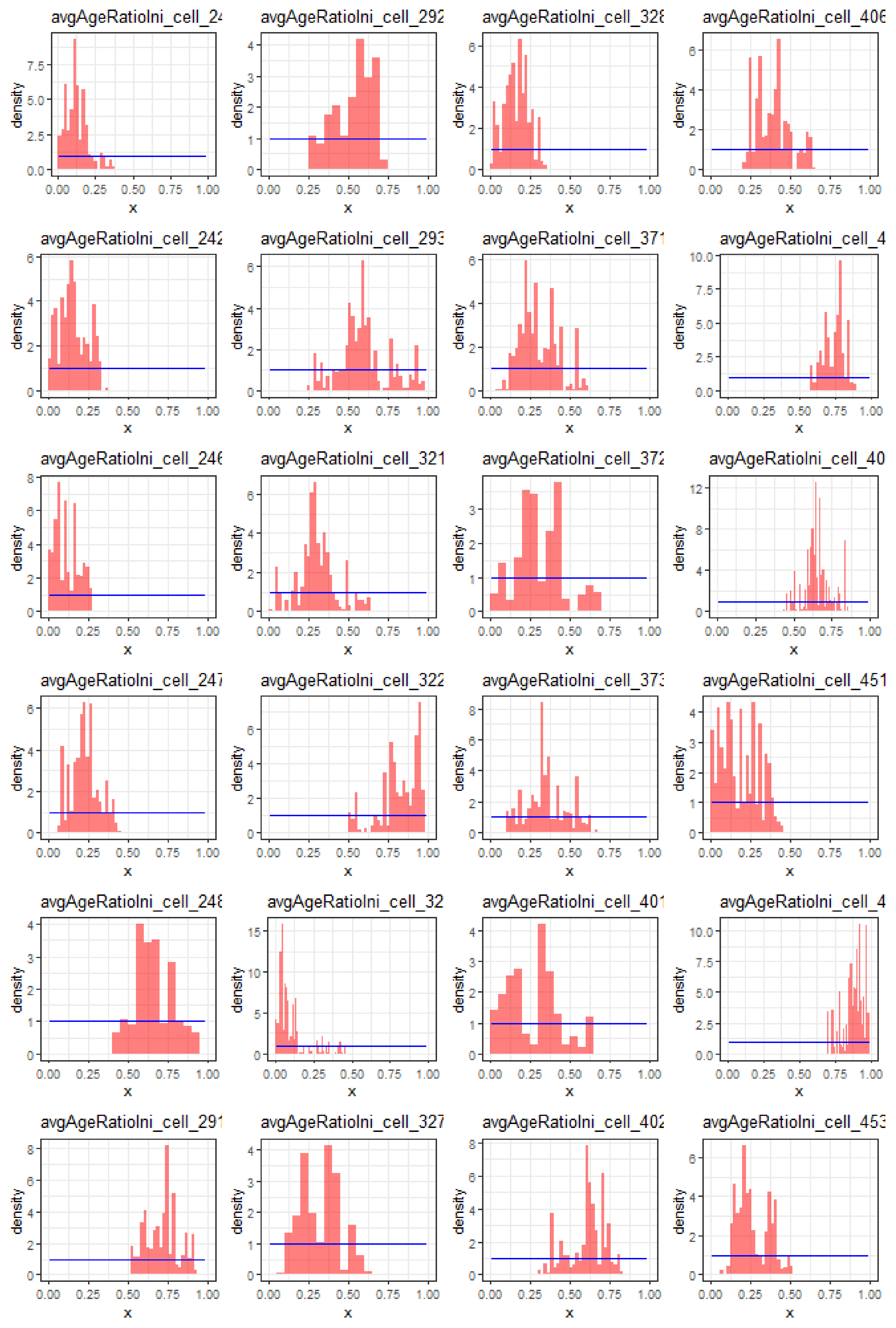
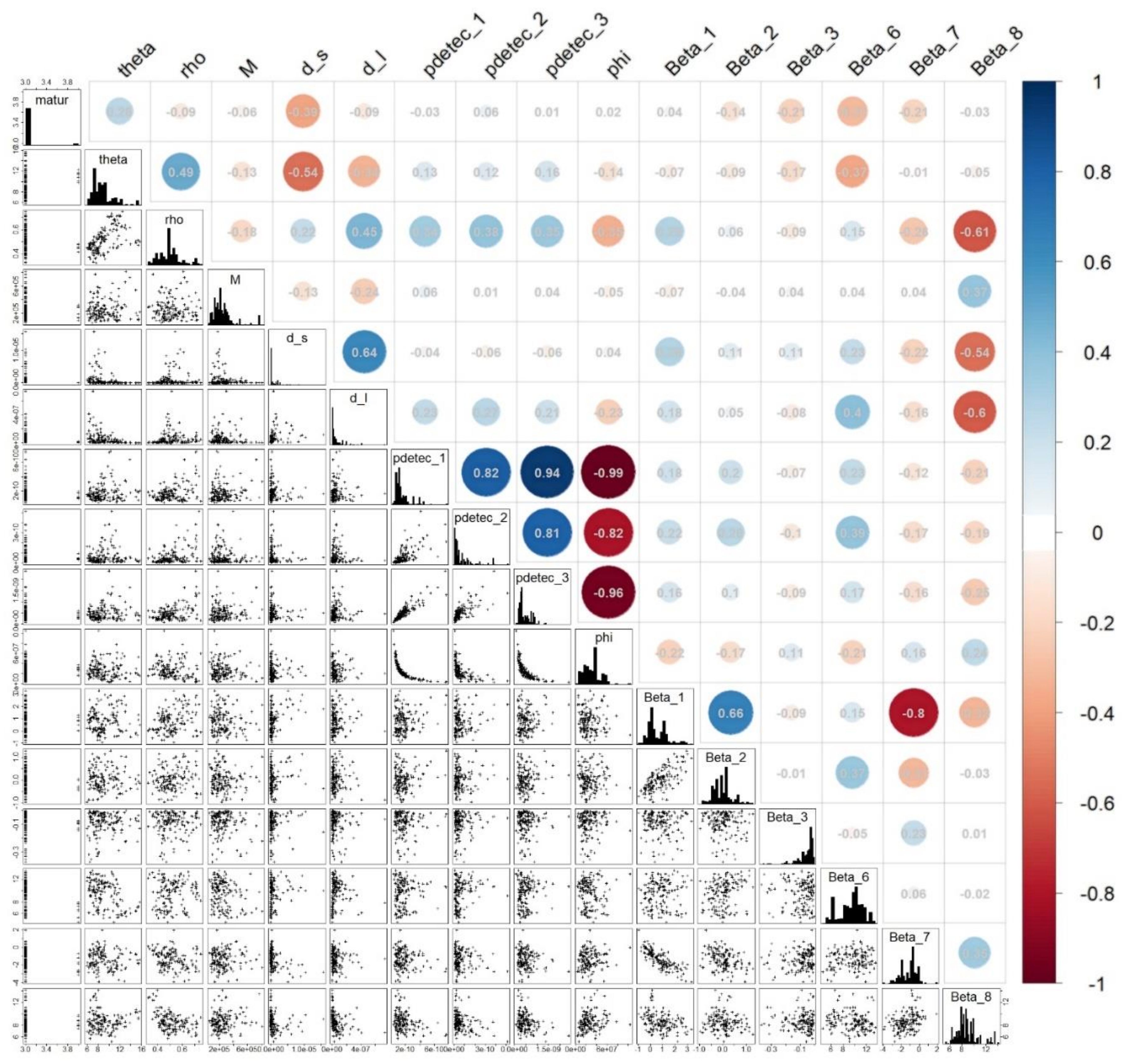
Appendix A.5. Environmental Suitability Estimates and Interpretations
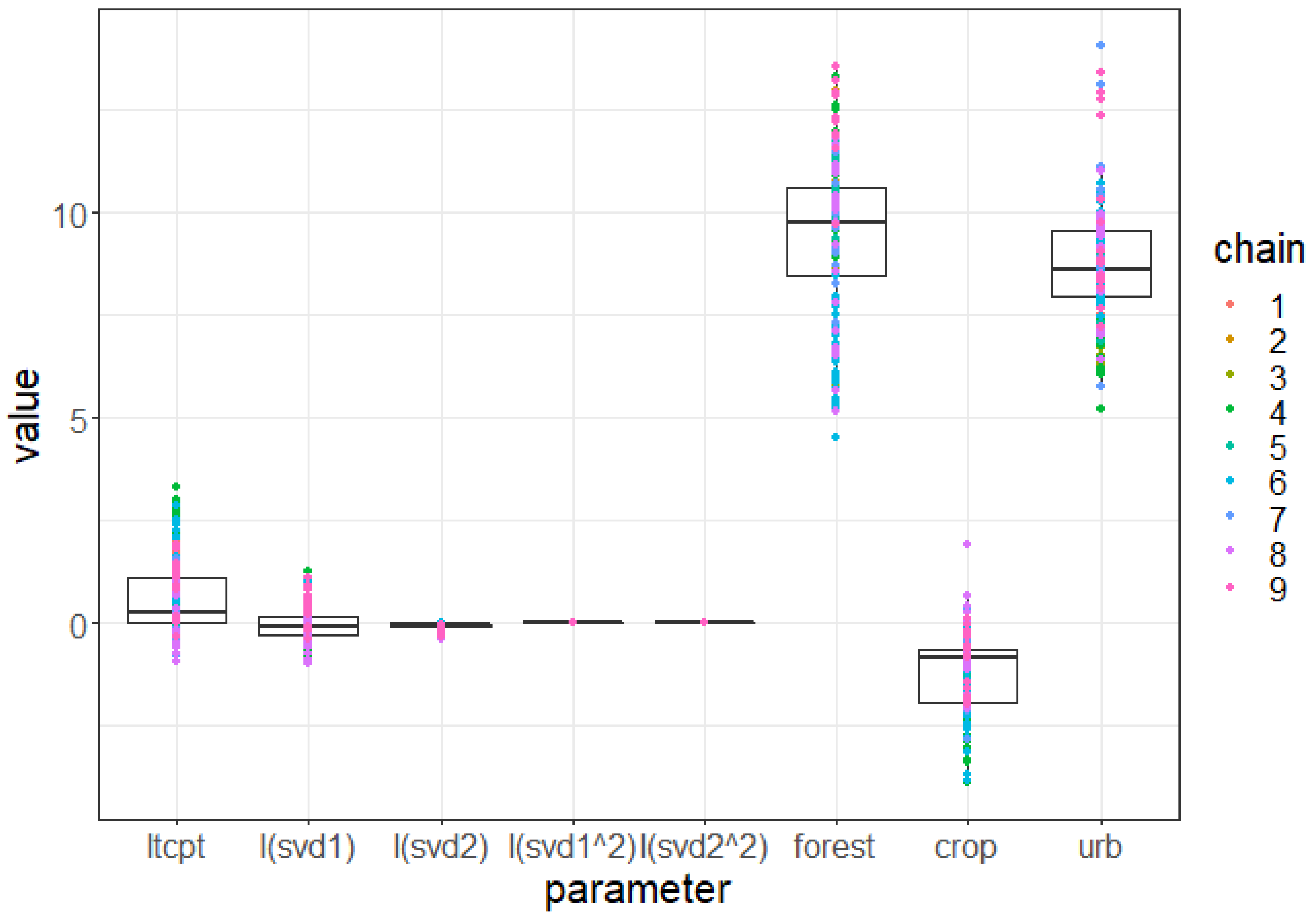
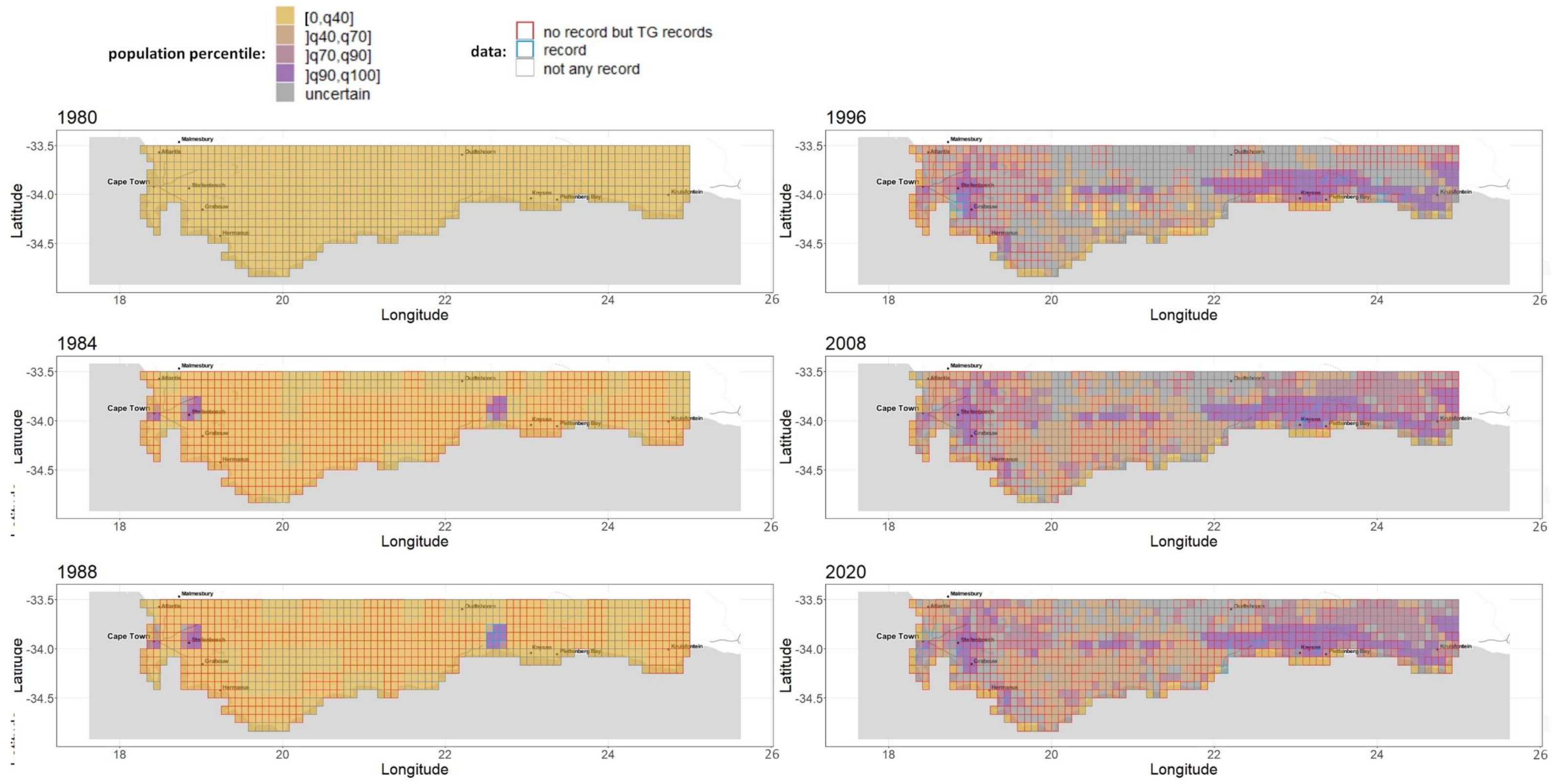
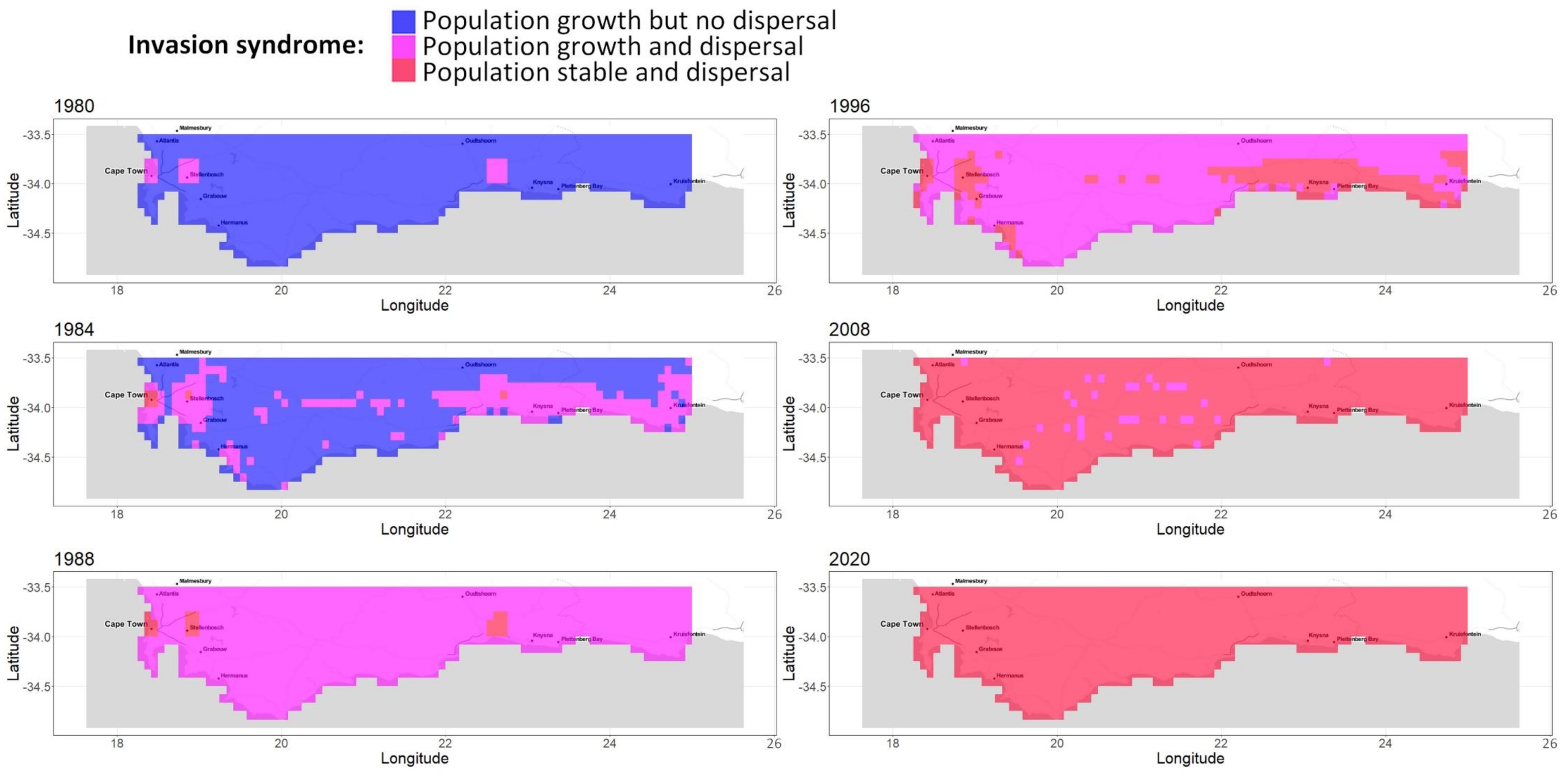
Appendix A.6. Complete Sequence of Growth and Population Maps
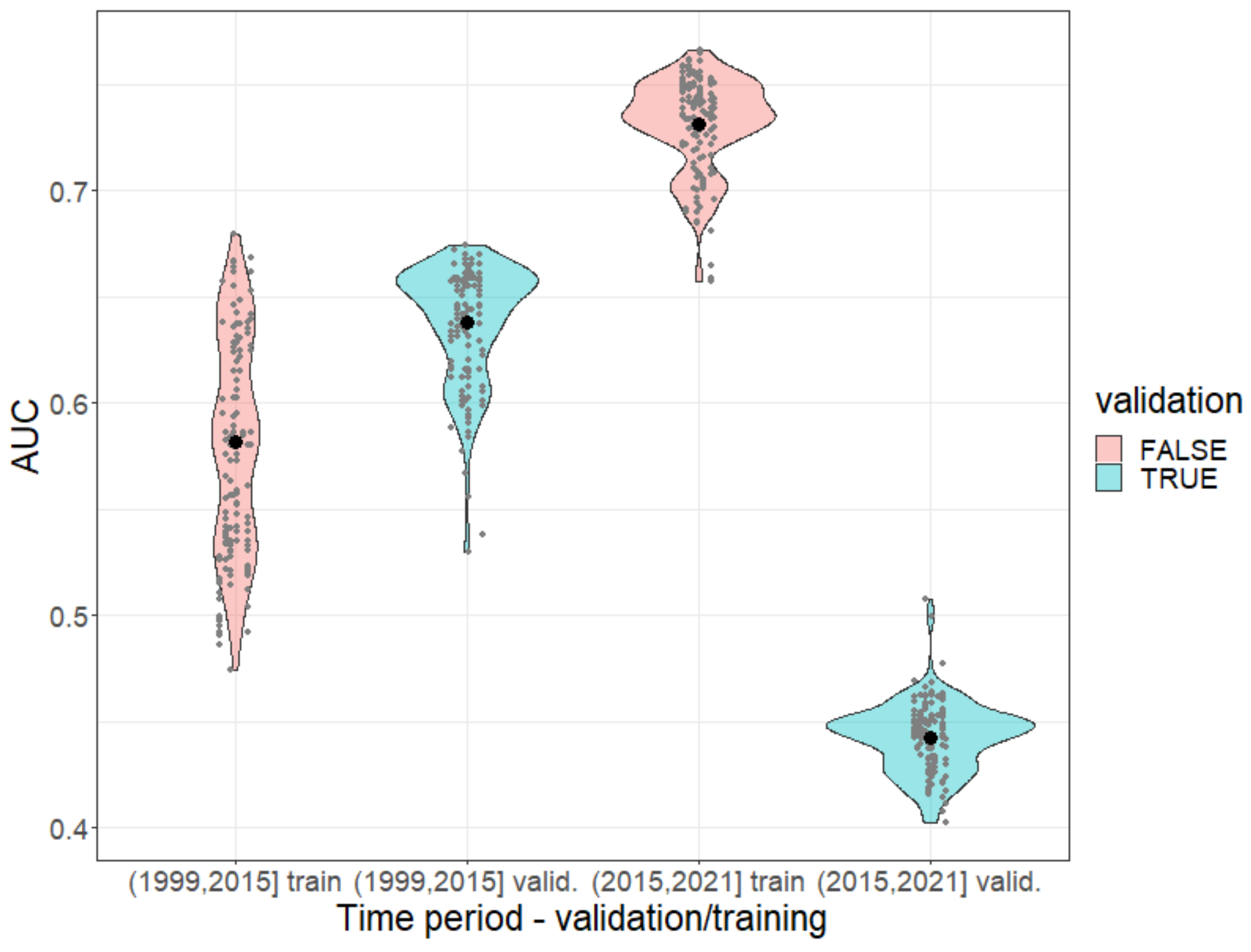
Appendix A.7. Model Validation

References
- Pyšek, P.; Křivánek, M.; Jarošík, V. Planting intensity, residence time, and species traits determine invasion success of alien woody species. Ecology 2009, 90, 2734–2744. [Google Scholar] [CrossRef] [PubMed]
- Haubrock, P.J.; Cuthbert, R.N.; Tricarico, E.; Diagne, C.; Courchamp, F.; Gozlan, R.E. The recorded economic costs of alien invasive species in Italy. NeoBiota 2021, 67, 247. [Google Scholar] [CrossRef]
- Renault, D.; Manfrini, E.; Leroy, B.; Diagne, C.; Ballesteros-Mejia, L.; Angulo, E.; Courchamp, F. Biological invasions in France: Alarming costs and even more alarming knowledge gaps. NeoBiota 2021, 67, 191. [Google Scholar] [CrossRef]
- Cuthbert, R.N.; Bartlett, A.C.; Turbelin, A.J.; Haubrock, P.J.; Diagne, C.; Pattison, Z.; Catford, J.A. Economic costs of biological invasions in the United Kingdom. NeoBiota 2021, 67, 299–328. [Google Scholar] [CrossRef]
- Haubrock, P.J.; Turbelin, A.J.; Cuthbert, R.N.; Novoa, A.; Taylor, N.G.; Angulo, E.; Courchamp, F. Economic costs of invasive alien species across Europe. NeoBiota 2021, 67, 153–190. [Google Scholar] [CrossRef]
- Seebens, H.; Blackburn, T.M.; Dyer, E.E.; Genovesi, P.; Hulme, P.E.; Jeschke, J.M.; Essl, F. No saturation in the accumulation of alien species worldwide. Nat. Commun. 2017, 8, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Rouget, M.; Robertson, M.P.; Wilson, J.R.; Hui, C.; Essl, F.; Renteria, J.L.; Richardson, D.M. Invasion debt–quantifying future biological invasions. Divers. Distrib. 2016, 22, 445–456. [Google Scholar] [CrossRef]
- Kowarik, I. Time lags in biological invasions with regard to the success and failure of alien species. Plant Invasions Gen. Asp. Spec. Probl. 1995, 15–38. [Google Scholar]
- Wilson, J.R.; Panetta, F.D.; Lindgren, C. Detecting and Responding to Alien Plant Incursions; Cambridge University Press: Cambridge, UK, 2016. [Google Scholar]
- Encarnação, J.; Teodósio, M.A.; Morais, P. Citizen science and biological invasions: A review. Front. Environ. Sci. 2021, 8, 303. [Google Scholar] [CrossRef]
- Hui, C.; Richardson, D.M. Invasion Dynamics; Oxford University Press: Oxford, UK, 2017. [Google Scholar]
- Elith, J. Predicting distributions of invasive species. Invasive Species Risk Assess. Manag. 2017, 10, 93–129. [Google Scholar]
- Thomas, C.D. Climate, climate change and range boundaries. Divers. Distrib. 2010, 16, 488–495. [Google Scholar] [CrossRef]
- Häkkinen, H.; Hodgson, D.; Early, R. Plant naturalizations are constrained by temperature but released by precipitation. Glob. Ecol. Biogeogr. 2021, 31, 504–521. [Google Scholar] [CrossRef]
- Roura-Pascual, N.; Bas, J.M.; Thuiller, W.; Hui, C.; Krug, R.M.; Brotons, L. From introduction to equilibrium: Reconstructing the invasive pathways of the Argentine ant in a Mediterranean region. Glob. Change Biol. 2009, 15, 2101–2115. [Google Scholar] [CrossRef]
- Donaldson, J.E.; Hui, C.; Richardson, D.M.; Robertson, M.P.; Webber, B.L.; Wilson, J.R. Invasion trajectory of alien trees: The role of introduction pathway and planting history. Glob. Change Biol. 2014, 20, 1527–1537. [Google Scholar] [CrossRef] [PubMed]
- Melbourne, B.A.; Hastings, A. Highly variable spread rates in replicated biological invasions: Fundamental limits to predictability. Science 2009, 325, 1536–1539. [Google Scholar] [CrossRef]
- Drenovsky, R.E.; Grewell, B.J.; D’antonio, C.M.; Funk, J.L.; James, J.J.; Molinari, N.; Richards, C.L. A functional trait perspective on plant invasion. Ann. Bot. 2012, 110, 141–153. [Google Scholar] [CrossRef]
- Daehler, C.C. Variation in self-fertility and the reproductive advantage of self-fertility for an invading plant (Spartina alterniflora). Evol. Ecol. 1998, 12, 553–568. [Google Scholar] [CrossRef]
- Pyšek, P. Is there a taxonomic pattern to plant invasions? Oikos 1998, 82, 282–294. [Google Scholar] [CrossRef]
- Schurr, F.M.; Pagel, J.; Cabral, J.S.; Groeneveld, J.; Bykova, O.; O’Hara, R.B.; Zimmermann, N.E. How to understand species’ niches and range dynamics: A demographic research agenda for biogeography. J. Biogeogr. 2012, 39, 2146–2162. [Google Scholar] [CrossRef]
- Louvrier, J.; Papaïx, J.; Duchamp, C.; Gimenez, O. A mechanistic–statistical species distribution model to explain and forecast wolf (Canis lupus) colonization in South-Eastern France. Spat. Stat. 2020, 36, 100428. [Google Scholar] [CrossRef]
- Roques, L.; Desbiez, C.; Berthier, K.; Soubeyrand, S.; Walker, E.; Klein, E.K.; Papaïx, J. Emerging strains of watermelon mosaic virus in Southeastern France: Model-based estimation of the dates and places of introduction. Sci. Rep. 2021, 11, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Rejmánek, M.; Richardson, D.M. What attributes make some plant species more invasive? Ecology 1996, 77, 1655–1661. [Google Scholar] [CrossRef]
- Higgins, S.I.; Richardson, D.M. Predicting plant migration rates in a changing world: The role of long-distance dispersal. Am. Nat. 1999, 153, 464–475. [Google Scholar] [CrossRef] [PubMed]
- Caswell, H.; Lensink, R.; Neubert, M.G. Demography and dispersal: Life table response experiments for invasion speed. Ecology 2003, 84, 1968–1978. [Google Scholar] [CrossRef]
- Pemberton, R.W.; Liu, H. Marketing time predicts naturalization of horticultural plants. Ecology 2009, 90, 69–80. [Google Scholar] [CrossRef]
- Castro-Díez, P.; Godoy, O.; Saldaña, A.; Richardson, D.M. Predicting invasiveness of Australian acacias on the basis of their native climatic affinities, life history traits and human use. Divers. Distrib. 2011, 17, 934–945. [Google Scholar]
- Caswell, H. Matrix Population Models; Sinauer: Sunderland, MA, USA, 2000; Volume 1. [Google Scholar]
- Stott, I.; Townley, S.; Hodgson, D.J. A framework for studying transient dynamics of population projection matrix models. Ecol. Lett. 2011, 14, 959–970. [Google Scholar] [CrossRef]
- Qiu, T.; Aravena, M.C.; Andrus, R.; Ascoli, D.; Bergeron, Y.; Berretti, R.; Clark, J.S. Is there tree senescence? The fecundity evidence. Proc. Natl. Acad. Sci. USA 2021, 118, e2106130118. [Google Scholar] [CrossRef]
- Wilson, J.R.U.; Richardson, D.M.; Rouget, M.; Procheş, Ş.; Amis, M.A.; Henderson, L.; Thuiller, W. Residence time and potential range: Crucial considerations in modelling plant invasions. Divers. Distrib. 2007, 13, 11–22. [Google Scholar] [CrossRef]
- Caley, P.; Groves, R.H.; Barker, R. Estimating the invasion success of introduced plants. Divers. Distrib. 2008, 14, 196–203. [Google Scholar] [CrossRef]
- Williamson, M.; Dehnen-Schmutz, K.; Kühn, I.; Hill, M.; Klotz, S.; Milbau, A.; Pyšek, P. The distribution of range sizes of native and alien plants in four European countries and the effects of residence time. Divers. Distrib. 2009, 15, 158–166. [Google Scholar] [CrossRef]
- Cook, A.; Marion, G.; Butler, A.; Gibson, G. Bayesian inference for the spatio-temporal invasion of alien species. Bull. Math. Biol. 2007, 69, 2005–2025. [Google Scholar] [CrossRef] [PubMed]
- Clark, J.S.; Scher, C.L.; Swift, M. The emergent interactions that govern biodiversity change. Proc. Natl. Acad. Sci. USA 2020, 117, 17074–17083. [Google Scholar] [CrossRef] [PubMed]
- West, M.; Harrison, P.J. Bayesian Forecasting and Dynamic Models, 2nd ed.; Springer: New York, NY, USA, 1997. [Google Scholar]
- Miller, D.A.; Pacifici, K.; Sanderlin, J.S.; Reich, B.J. The recent past and promising future for data integration methods to estimate species’ distributions. Methods Ecol. Evol. 2019, 10, 22–37. [Google Scholar] [CrossRef]
- Hastie, T.; Fithian, W. Inference from presence-only data; the ongoing controversy. Ecography 2013, 36, 864–867. [Google Scholar] [CrossRef] [PubMed]
- Alasbahi, R.H.; Melzig, M.F. Plectranthus barbatus: A review of phytochemistry, ethnobotanical uses and pharmacology-Part 1. Planta Med. 2010, 76, 653–661. [Google Scholar] [CrossRef]
- Phillips, L.A.; Greer, C.W.; Farrell, R.E.; Germida, J.J. Field-scale assessment of weathered hydrocarbon degradation by mixed and single plant treatments. Appl. Soil Ecol. 2009, 42, 9–17. [Google Scholar] [CrossRef]
- Botella, C.; Joly, A.; Monestiez, P.; Bonnet, P.; Munoz, F. Bias in presence-only niche models related to sampling effort and species niches: Lessons for background point selection. PLoS ONE 2020, 15, e0232078. [Google Scholar] [CrossRef]
- Loveland, T.R.; Reed, B.C.; Brown, J.F.; Ohlen, D.O.; Zhu, Z.; Yang, L.W.M.J.; Merchant, J.W. Development of a global land cover characteristics database and IGBP DISCover from 1 km AVHRR data. Int. J. Remote Sens. 2000, 21, 1303–1330. [Google Scholar] [CrossRef]
- Harris, I.; Jones, P.D.; Osborn, T.J.; Lister, D.H. Updated high-resolution grids of monthly climatic observations—The CRU TS3.10 Dataset. Int. J. Climatol. 2014, 34, 623–642. [Google Scholar] [CrossRef]
- Fick, S.E.; Hijmans, R.J. WorldClim 2: New 1km spatial resolution climate surfaces for global land areas. Int. J. Climatol. 2017, 37, 4302–4315. [Google Scholar] [CrossRef]
- Bellard, C.; Thuiller, W.; Leroy, B.; Genovesi, P.; Bakkenes, M.; Courchamp, F. Will climate change promote future invasions? Glob. Chang. Biol. 2013, 19, 3740–3748. [Google Scholar] [CrossRef] [PubMed]
- Araújo, M.B.; Pearson, R.G.; Thuiller, W.; Erhard, M. Validation of species–climate impact models under climate change. Glob. Chang. Biol. 2005, 11, 1504–1513. [Google Scholar] [CrossRef] [Green Version]
- Chalmandrier, L.; Hartig, F.; Laughlin, D.C.; Lischke, H.; Pichler, M.; Stouffer, D.B.; Pellissier, L. Linking functional traits and demography to model species-rich communities. Nat. Commun. 2021, 12, 1–9. [Google Scholar] [CrossRef]
- McLean, P.; Gallien, L.; Wilson, J.R.U.; Gaertner, M.; Richardson, D.M. Small urban centres as launching sites for plant invasions in natural areas: Insights from South Africa. Biol. Invasions 2017, 19, 3541–3555. [Google Scholar] [CrossRef]
- Potgieter, L.J.; Douwes, E.; Gaertner, M.; Measey, G.J.; Paap, T.; Richardson, D.M. Biological invasions in South Africa’s urban ecosystems: Patterns, processes, impacts and management. In Biological Invasions in South Africa; Van Wilgen, B.W., Measey, J., Richardson, D.M., Wilson, J.R.U., Zengeya, T.A., Eds.; Springer: Berlin/Heidelberg, Germany, 2020; pp. 275–309. [Google Scholar]
- Pages, M.; Fischer, A.; van der Wal, R.; Lambin, X. Empowered communities or “cheap labour”? Engaging volunteers in the rationalised management of invasive alien species in Great Britain. J. Environ. Manag. 2019, 229, 102–111. [Google Scholar] [CrossRef]
- Catterall, S.; Cook, A.R.; Marion, G.; Butler, A.; Hulme, P.E. Accounting for uncertainty in colonisation times: A novel approach to modelling the spatio-temporal dynamics of alien invasions using distribution data. Ecography 2012, 35, 901–911. [Google Scholar] [CrossRef]
- Groom, Q.; Adriaens, T.; Bertolino, S.; Poelen, J.H.; Reeder, D.; Richardson, D.M.; Simmons, N. Holistic understanding of contemporary ecosystems requires integration of data on domesticated, captive, and cultivated organisms. Biodivers. Data J. 2021, 9, e65371. Available online: https://bdj.pensoft.net/article/65371/ (accessed on 29 August 2022). [CrossRef]
- Li, E.; Parker, S.S.; Pauly, G.B.; Randall, J.M.; Brown, B.V.; Cohen, B.S. An urban biodiversity assessment framework that combines an urban habitat classification scheme and citizen science data. Front. Ecol. Evol. 2019, 7, 277. [Google Scholar] [CrossRef]
- Aikio, S.; Duncan, R.P.; Hulme, P.E. Lag-phases in alien plant invasions: Separating the facts from the artefacts. Oikos 2010, 119, 370–378. [Google Scholar] [CrossRef]
- Nelson, G.; Ellis, S. The history and impact of digitization and digital data mobilization on biodiversity research. Philos. Trans. R. Soc. B 2019, 374, 20170391. [Google Scholar] [CrossRef] [PubMed]
- Randin, C.F.; Dirnböck, T.; Dullinger, S.; Zimmermann, N.E.; Zappa, M.; Guisan, A. Are niche-based species distribution models transferable in space? J. Biogeogr. 2006, 33, 1689–1703. [Google Scholar] [CrossRef]
- Elliott-Graves, A. The problem of prediction in invasion biology. Biol. Philos. 2016, 31, 373–393. [Google Scholar] [CrossRef]
- Cole, D.J. Bayesian Identifiability. In Parameter Redundancy and Identifiability; Chapman and Hall: London, UK, 2020; pp. 101–153. [Google Scholar]
- MacKenzie, D.I.; Nichols, J.D.; Lachman, G.B.; Droege, S.; Royle, J.A.; Langtimm, C.A. Estimating site occupancy rates when detection probabilities are less than one. Ecology 2002, 83, 2248–2255. [Google Scholar] [CrossRef]
- Rosenthal, J.S. Optimal proposal distributions and adaptive MCMC. Handb. Markov Chain. Monte Carlo 2011, 4, 93–112. [Google Scholar]
- Hartig, F. BayesianTools: General-Purpose MCMC and SMC Samplers and Tools for Bayesian Statistics, R package version 0.1-7; 2017; CRAN; Available online: https://cran.r-project.org/web/packages/BayesianTools/index.html (accessed on 29 August 2022).
- Rosenthal, M.; Glew, R. Medical Biochemistry, 1st ed.; Wiley: Hoboken, NJ, USA, 2011; Original work published 2011; Available online: https://www.perlego.com/book/1008615/medical-biochemistry-pdf (accessed on 29 August 2022).
- Brooks, S.P.; Gelman, A. General methods for monitoring convergence of iterative simulations. J. Comput. Graph. Stat. 1998, 7, 434–455. [Google Scholar]
- Jiménez-Valverde, A. Insights into the area under the receiver operating characteristic curve (AUC) as a discrimination measure in species distribution modelling. Glob. Ecol. Biogeogr. 2012, 21, 498–507. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Botella, C.; Bonnet, P.; Hui, C.; Joly, A.; Richardson, D.M. Dynamic Species Distribution Modeling Reveals the Pivotal Role of Human-Mediated Long-Distance Dispersal in Plant Invasion. Biology 2022, 11, 1293. https://doi.org/10.3390/biology11091293
Botella C, Bonnet P, Hui C, Joly A, Richardson DM. Dynamic Species Distribution Modeling Reveals the Pivotal Role of Human-Mediated Long-Distance Dispersal in Plant Invasion. Biology. 2022; 11(9):1293. https://doi.org/10.3390/biology11091293
Chicago/Turabian StyleBotella, Christophe, Pierre Bonnet, Cang Hui, Alexis Joly, and David M. Richardson. 2022. "Dynamic Species Distribution Modeling Reveals the Pivotal Role of Human-Mediated Long-Distance Dispersal in Plant Invasion" Biology 11, no. 9: 1293. https://doi.org/10.3390/biology11091293
APA StyleBotella, C., Bonnet, P., Hui, C., Joly, A., & Richardson, D. M. (2022). Dynamic Species Distribution Modeling Reveals the Pivotal Role of Human-Mediated Long-Distance Dispersal in Plant Invasion. Biology, 11(9), 1293. https://doi.org/10.3390/biology11091293






