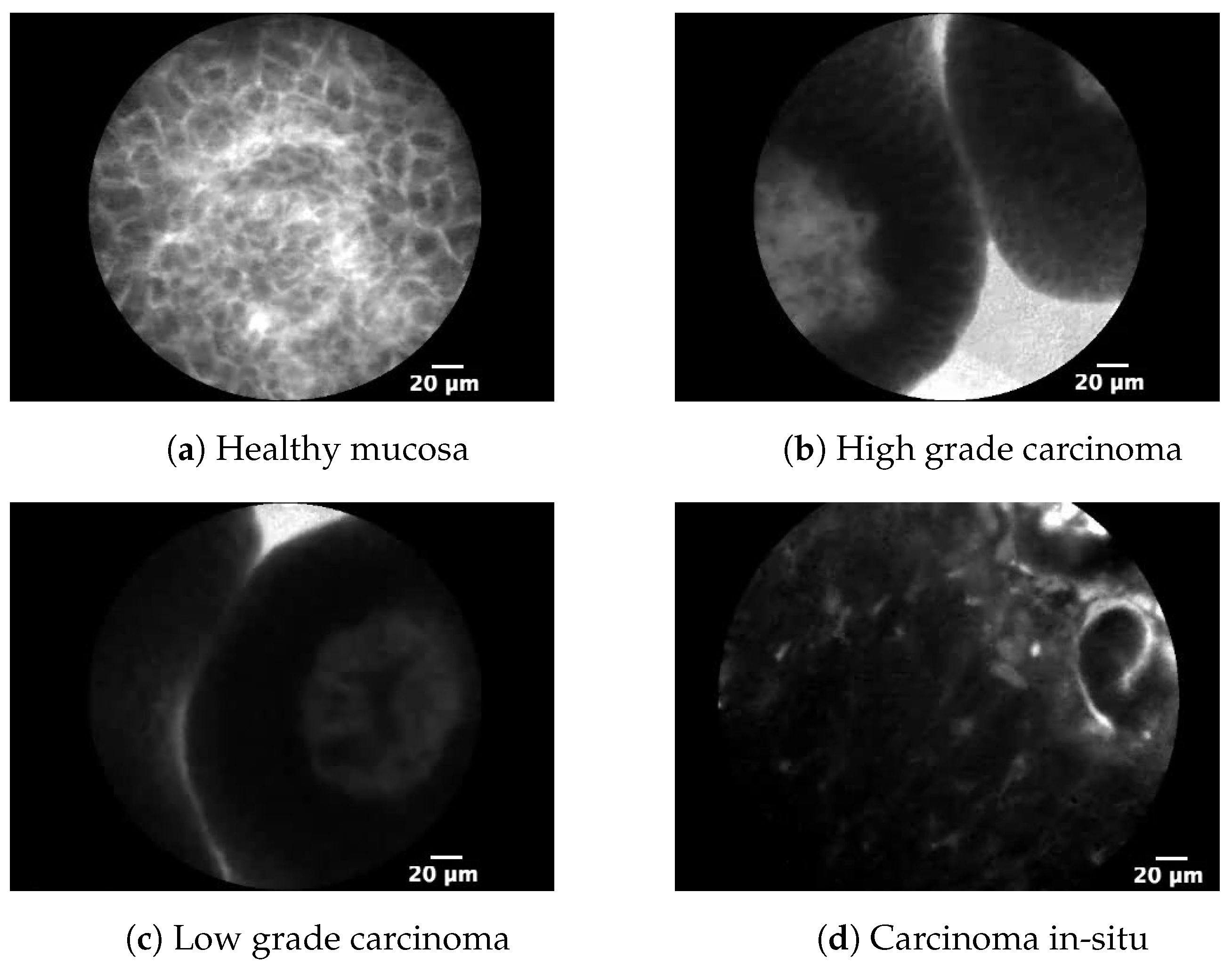
1. Introduction
Urinary bladder cancer, as one of the most common malignant diseases of the urinary tract, is a consequence of a mutation in the bladder’s mucosa cells which causes their uncontrolled growth. Such growth shows a high tendency to spread to the rest of mucosa, but also to other parts of the human body. Above stated facts, urinary bladder cancer is showing elevated recurrence rates ranging from 61% in the first year, to 78% in the first five years. Such recurrence rates are among the highest compared to other malignant diseases. For this reason, diagnosis, treatment and follow-up are extremely challenging [
1]. There is a multiplicity of urinary bladder cancer types, among which the most common are:
urothelial carcinoma [
2,
3],
squamous cell carcinoma [
4,
5],
small cell carcinoma [
8,
9] and
Most urinary bladder cancers are urothelial carcinomas or transitional cell carcinomas (TCC). These carcinomas are, in their papillary form, characterized by low-grade metastatic potential [
12]. On the other hand, high-grade cancer and carcinoma in-situ (CIS) are characterized by higher metastatic potential. Unlike TCC, CIS lesion of urinary bladder mucosa is dominantly flat, making it harder to differentiate from benign growths.
The dominant medical examination for the diagnosis of urinary bladder cancer is cystoscopy, an endoscopic method where a probe called a cystoscope is inserted into the urinary bladder via ureter. Modern cystoscopes are equipped with confocal laser endomicroscopes (CLE) that allow in-vivo evaluation of the mucosa, without the need for biopsy and patho-histological examination. Such a method called an optical biopsy, shows high results from the point of view of detecting papillary lesions. On the other hand, the same method has failed in the detection of CIS, and its accuracy does not exceed 75%. To increase the likelihood of a positive outcome, the goal is an accurate and timely diagnosis of CIS, which, as already mentioned, is characterized by its high metastatic potential [
13,
14].
With aim of increasing accuracy, machine learning (ML) algorithms are introduced and integrated with CLE-based cystoscopy [
15]. Such integration can be of particular importance especially in the case of CIS recognition. There were several research works based on ML utilization for cystoscopic image classification. Research presented in [
16] proposed the utilization of a multilayer perceptron (MLP) whose initial weights were pre-determined with a genetic algorithm. The authors of the research presented in [
17] have proposed a Convolutional Neural Network-based (CNN) approach that achieved
of
. CNN-based approach is also presented in [
18], where authors have achieved accuracies up to
. A similar approach is presented in [
19], where authors have proposed utilization of pre-defined networks in order to achieve high classification performances. The best results have been achieved with Xception architecture (
). A hybrid approach is presented in [
20] where authors have proposed a combination of Laplacian edge detector and MLP that follows CNN methodology. By using this approach,
values up to
were reached. From the presented overview, it can be noticed that CNN-based architectures are playing a lead role in ML-based algorithms for the classification of cystoscopic images. However, regardless of the encouraging results in this area, there is always room for improvement, both from CNN and data preprocessing standpoint.
The collection of data for Artificial Intelligence (AI) studies in the field of medicine, especially in the classification of rarely appearing diseases—such as specific types of carcinoma—can be complex. Not only can the data collection process be hard due to a low number of patients on which the testing procedures need to be performed, ethical questions can arise. Not all patients may consent to the data collected during procedures performed with them as the subject being shared with non-medical researchers. Additionally, it can be hard to guarantee near equal numbers across various classes, as, naturally, the larger number of patients on who the testing procedure is performed are positive, due to the test usually being performed on symptomatic subjects. Due to this, there might be a larger number of positive test findings in the input, in comparison to the negative (healthy) patients [
20,
21].
Because of the above-listed issues, data augmentation is commonly used. Data augmentation refers to the process of generating new inputs for Artificial Neural Network (ANN) training. This process is commonly performed on datasets that consist of pictures as inputs. A common practice is the use of deterministic modifications on the images of the input dataset, where standard transformations are used [
22,
23]. These transformations include vertical and horizontal mirroring, rotating, and scaling of different image sections [
24]. While these transformations can provide additional robustness to the model, they are still tightly connected to the existing data and may not provide needed variance to the input dataset.
The answer to this issue lies in the use of Generative Adversarial Networks (GAN) [
25]. Through the use of GAN new images can be generated, which can then be used as inputs, and the resulting models can be tested using previously established metrics, to test the precision of models created using generated data.
The aim of this research is to examine the influence of GAN-based image data augmentation of CNNs for urinary bladder cancer diagnosis. This research is focused on increasing the performance of classifiers by applying dataset pre-processing, while using established CNN architectures that have achieved high classification performance in solving various problems in practice. From facts stated above, several questions could be asked:
Is it possible to utilize GAN fo urinary bladder cancer image data augmentation,
Are classifier performance higher if an augmented set is used and
How does the share of data generated in the training set affect the performance of the classifier?
3. Use of DCGAN Networks in Generation of Medical Data
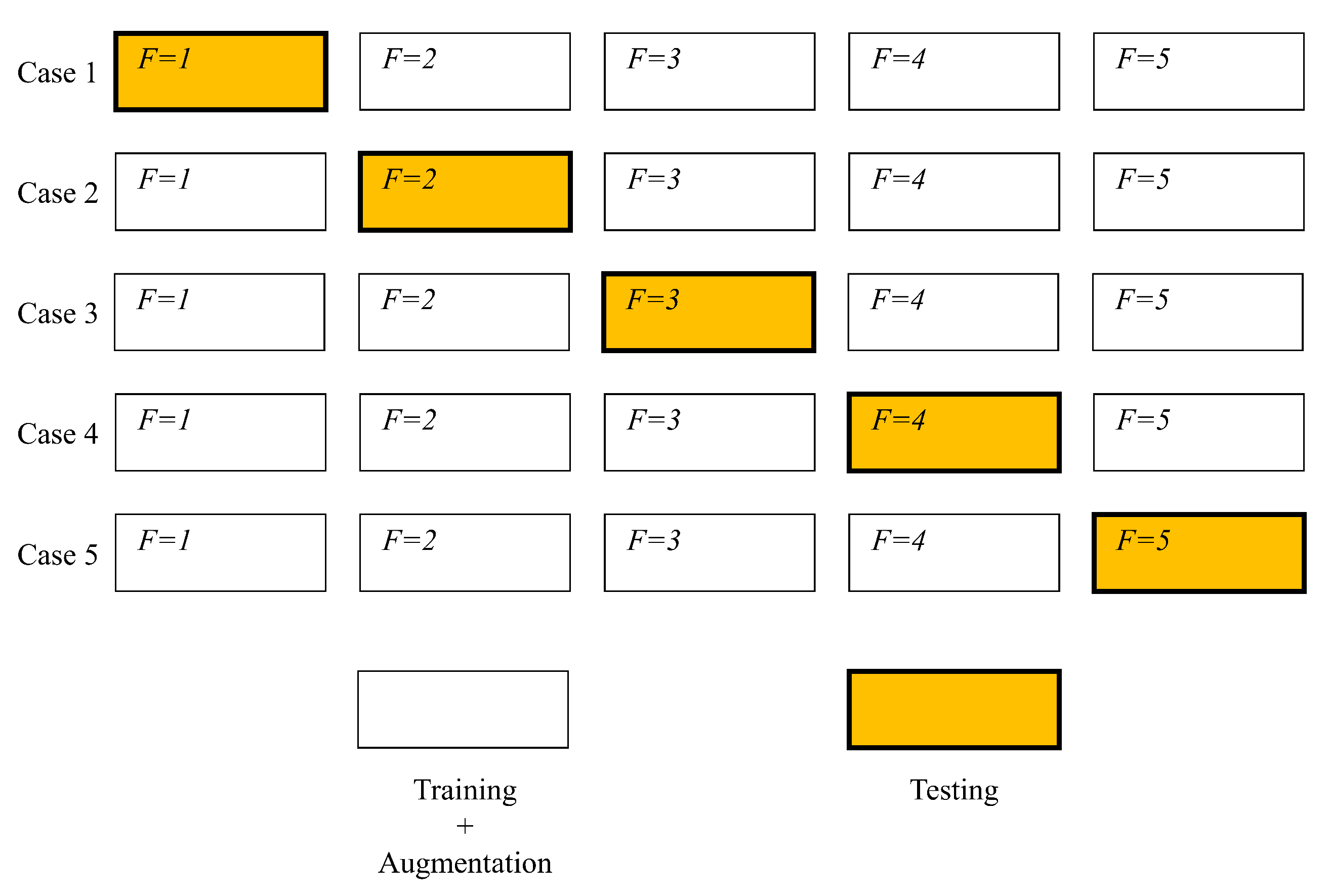
In this section, a brief overview of a GAN-based image generating process will be presented. The overview of the process is relatively simple. Dataset is split into two parts—training and testing set, as is the standard ANN training practice [
27,
28]. The testing part of the dataset is set aside, while the training set is fed into the two parts of GAN—discriminator and generator [
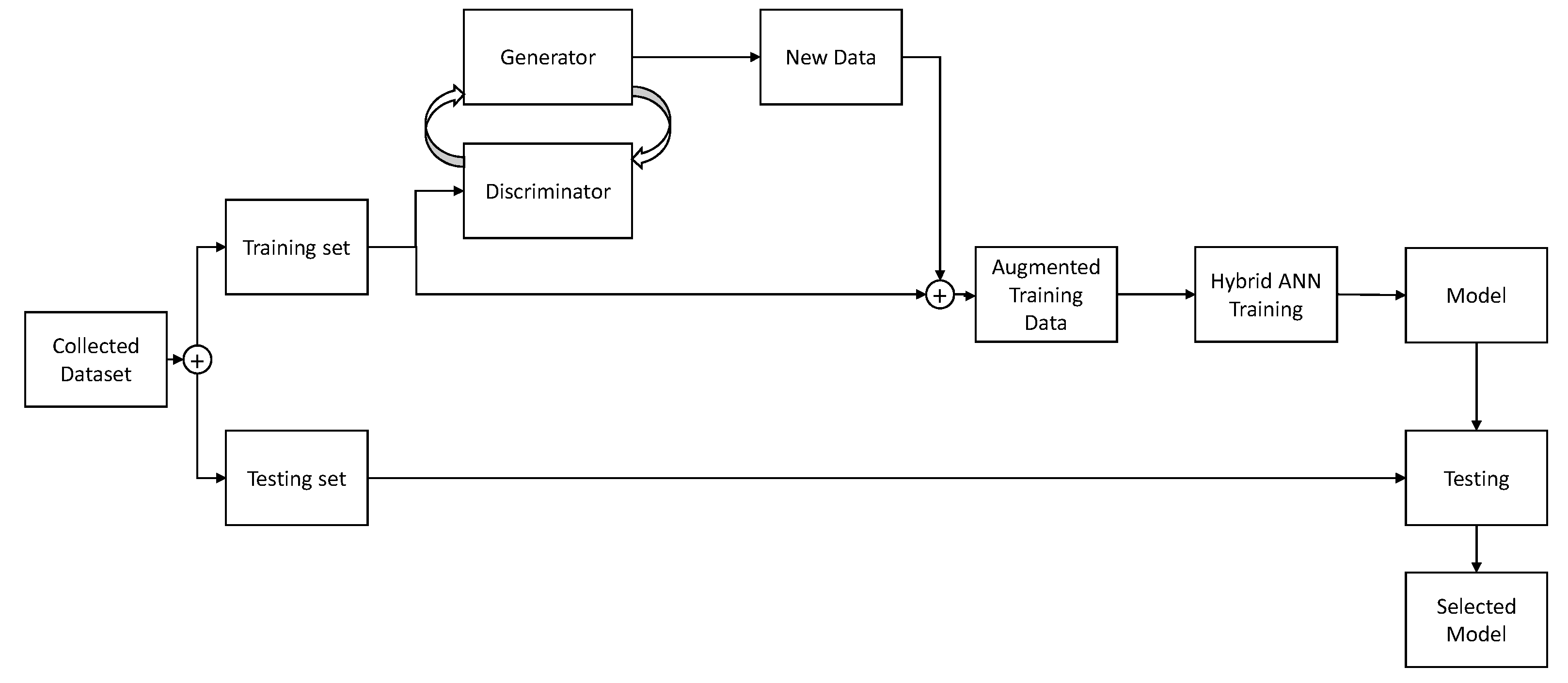
29]. A discriminator is trained to distinguish real and fake images, while Generator is trained to generate images. The generator adjusts its’ parameters in an attempt of generating images that are realistic enough that the discriminator does not detect them as false. Once this process is completed, the generated images are mixed with the original training set, providing an augmented training set. This augmented training set is then used to train the CNN and created models which are evaluated on images of the testing set. It is important to note that the testing set does not contain images generated via the described process. If generated images were of low-quality due to improper settings of the GAN, scores could still be high if they were used in the evaluation. By avoiding using the generated images within the test set, if they are of low-quality CNN model will be trained using inappropriate data—where inappropriate data means generated images which are massively inconsistent in comparison to real data. If the model is trained with such data, and then evaluated using real data the scores will be low. In this way, the data testing process evaluates the CNN and GAN quality as well, as both ANNs will have to achieve good results for the scores to be acceptable. This process is shown in
Figure 3.
The ANN setup used consists of two separate ANNs—generator and discriminator. The generator and discriminator used in this research are both deep convolutional neural networks, which makes GAN used in this research a Deep Convolutional Generative Adversarial Network (DCGAN) [
30].
Convolution is a mathematical operation defined on two functions (
x and
h) which produces a third function that expresses how the function
h changes the shape of function
x [
31]. If
n and
k refer to indices of the discreete signals, it can be defined with:
Equation (
1) defines the one-dimensional, discrete convolution. Considering the fact that the convolution is performed on images, a two-dimensional definition is necessary, given by [
31,
32]:
or:
Convolutional neural networks work by applying convolution onto the input of the layer [
28,
33]. They filter the image, capturing the spatial and temporal dependencies contained within. The role of convolutional networks is transforming images into smaller, easier to process forms, without losing the important features [
34]. The convolution layer is defined using four parameters [
35]:
number of filters—which define the dimensionality of the output,
the size of kernel (h)—which specify the dimensions of convolution window,
strides—which specify convolutional strides along the height and width, and
padding—which defines whether padding will be applied to the output of convolution or should the size remain the same.
Assuming the input image of dimensions
pixels, where the first number represents the height of the image in pixels, the second represents the width, and final is the number of color channels—in this case one, as a gray-scale image is assumed. The size of the image is defined, for both its height and width using the following [
36]:
where:
is the output dimension (width or height) of the image,
L is the dimension (width or height) size of the image,
F is the dimension of the kernel in the same direction (width or height) as the image dimension being transformed,
P is the size of padding applied to the image, and
S is the total number of strides.
The “same” padding is used. This padding will assure that the input image gets fully covered by the filter and specified stride. As this is the setting used, the Equation (
4) can be simplified to [
36]:
Transposed convolution allows the input to be expanded instead of compressed as it is when using convolution. It is commonly used in upscaling, but its role in DCGAN is the generation of an image from randomized input data using learned weights stored within the kernel
h. Transposed convolution is performed by taking the original image
x and kernel
h. Then zero-padding equal to the dimensions of
h, minus 1, is added to the image
x. The convolution is then performed and the calculation for each element in the new matrix
is performed using Equation (
3), and, to differentiate it, commonly marked using:
The size of the resulting matrix is defined with [
32,
36]:
which can, when using “same” padding be simplified similar to Equation (
5) [
32,
36]:
The networks in question are referred to as “adversarial”, as they are working “against each other” [
37]. The discriminator is trained as a classifier—using the original collected data—with the goal of differentiating between real and fake images [
38]. The generator is created to generate images, with the goal of tricking the discriminator [
30,
38]. It takes a randomized input, which is in this case a uniformly distributed random vector of 100 elements in a range between 0.0 and 1.0. This randomized input is fed into the generator which, using two-dimensional transposed convolution, generates the image. The generated image is then fed into the discriminator which classifies it as belonging to the original dataset or not. In other words; it discriminates between false (generated) images and real ones [
38]. If the discriminator determines the image generated is fake, the parameters of the generator are adjusted, and the process repeats [
39]. In this way, the generator continually adjusts its parameters, until the images produced by it are indistinguishable from the original dataset images to the discriminator [
40].
The input to the generator shaped (1, 100) is fed into a densely connected input layer shaped (
) where
represents the expected width of the generated image, and
represents the expected height of the generated image. This input passes through batch normalization, which normalizes the output of the previous layer through the subtraction of the image batch mean and dividing by the image batch standard deviation. Such an approach provides higher stability to the network as it avoids too large of a value appearing in the layer output [
41]. It also provides a more independent learning environment to the layers surrounding it [
33,
41]. The output is then passed through LeakyReLU activation function. LeakyReLU comes from Rectified Linear Unit activation function, which maps the positive input directly to the output, while eliminating the negative outputs—defined with
. LeakyReLU functions in a similar manner, but instead of completely eliminating negative values it just lowers their value by the factor of
[
42]. LeakyReLU can be defined using [
43]:
The generator then uses two two-dimensional transposed convolution layers to further expand the image, going to shapes of (
) to (
) and finally to (
). The output of each transposed convolution layer, except the last, is processed through batch normalization and LeakyReLU activation layers. It can be seen, that the changes to the image dimensions follow Equation (
8). For example, generating process of 28 × 28 image starts with the aforementioned (1, 100) uniformly randomized vector, which is mapped using a densely connected layer into the shape of (1, 12,544). Such a vector is reshaped into the three-dimensional layer shaped (7, 7, 256). Through the process of transposed convolution, this is transformed into (14, 14, 64), and finally (28, 28, 1). This form represents the desired output shape. The third dimension is a value defined via the arbitrary number of filters contained in the two-dimensional transposed convolution layer.
The discriminator is also defined as CNN. It consists of two two-dimensional convolution layers, followed by leakyReLU activation and a Dropout Layer. The dropout layer applies dropout regularization to the output of a previous layer. Regularization is important in avoiding the over-fitting of ANN to certain prominent features [
27]. Dropout achieves this by setting a fraction of inputs to the layer to 0. In this case, the rates for dropout are
(
) for both dropout layers. Finally, a flatten layer is used to flatten the vector of
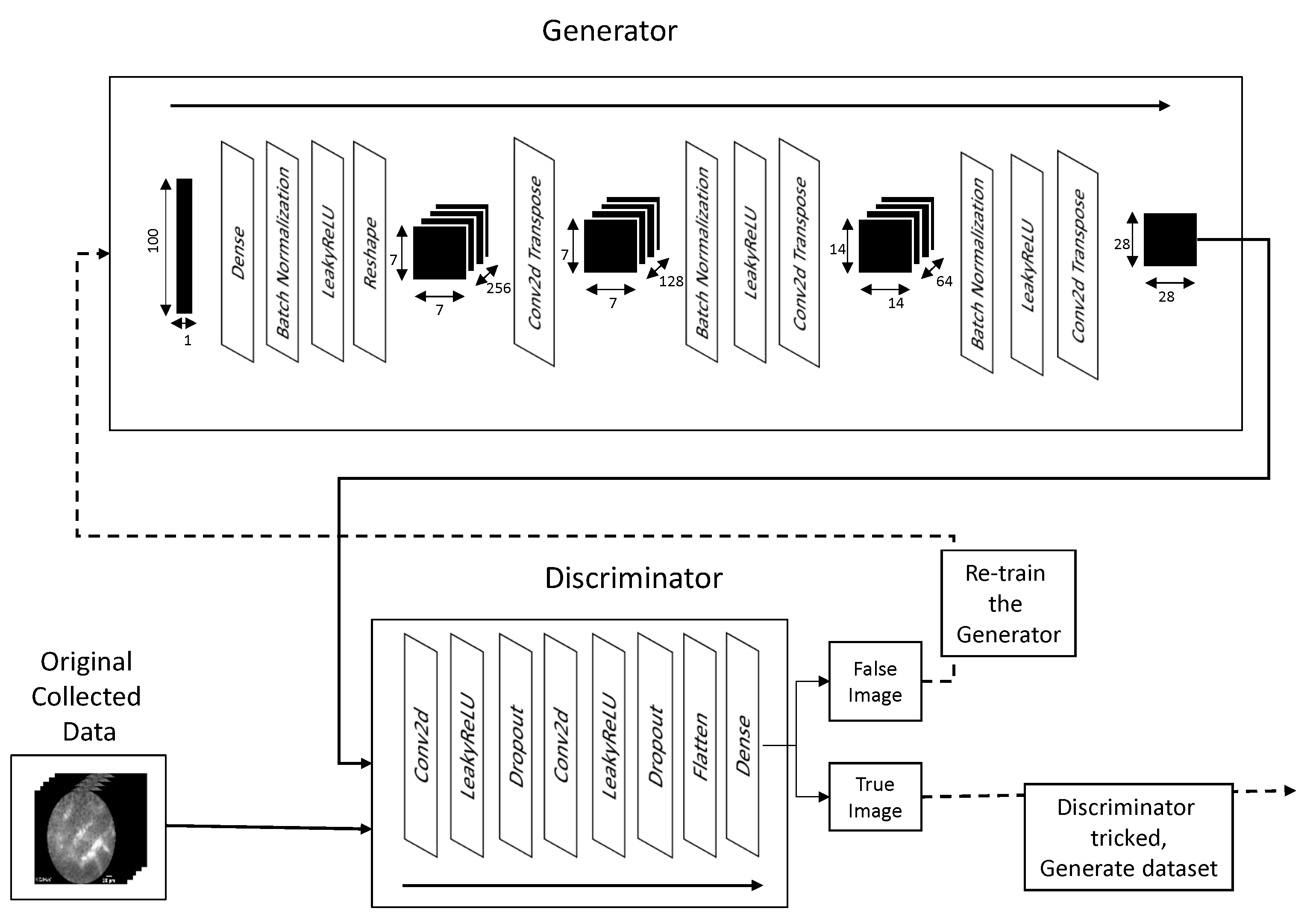
n-dimensional matrices into a vector which can be interpreted by the final layer of the CNN—a dense layer containing a single output neuron. The value of the neuron in the final layer defines the output of the discriminator. The architecture and process of training are shown in
Figure 4. The transformation of image sizes within the network follows the Equation (
5). For example, if the input image has the dimensions of (28, 28, 1)—which is the expected image size—the first two-dimensional convolution layer transforms this into (14, 14, 64)—where the first two dimensions follow the Equation (
5), and the third being arbitrarily defined number of filters. The second convolutional layer transforms this into (7, 7, 128), by following the same rules. Finally, this is flattened using the flatten layer into a vector of dimensions (1, 6272), where the number of 6272 elements is obtained by multiplying all dimension in the output tensor of the previous layer (
), in order for both of these layer outputs to have the same number of elements. Finally, this is transformed into a single output neuron using a densely connected layer with the same number of connections as the outputs of the flatten layer—6272. It is important to note that in GAN networks the output of the generator needs to fit not only the expected generated image size, but also the input to the discriminator. When working with DCGANs this means that convolution and transposed convolution layers need to be adjusted in such a manner that this can be achieved.
Loss
L is a measure of difference between the expected and actual input of the neural network. In other words, it defines the error of the neural network. By lowering the value of loss, a more precise ANN is achieved, and this is the goal of training stage of ANNs (
) [
33]. In this research, cross-entropy loss is used. Cross-entropy measures the difference between two probability distributions for a random variable [
27,
44]. Entropy can be defined as the number of bits needed to represent the information of an event occurring. Lower probability events will have higher entropy, as more information needs to be conveyed, while higher probability events will have low entropy. For a random variable
z, which can be in a set of discrete states
Z, with the number of these states being
, with the probability of each of these states occurring being defined as
, entropy
can be defined as [
45]:
Cross-entropy on the other hand calculates the number of bits required to represent event
z from distribution
in comparison to distribution
. In other words—how many bits are necessary to represent an event using distribution
, instead of distribution
. When used as a loss function the discrete states
Z represent the possible classes. Cross-entropy is derived from the Equation (
10), being given as [
44]:
The classes of discriminator output need to be defined. The discriminator is a binary classifier, meaning its’ output will be one of the two classes “0” or “1”. In the presented case the discriminator will return “1” if it classifies the images as “real” and “0” if it classifies the images as fake. To calculate the cross-entropy of generator the output is compared using cross-entropy to an array of ones, because if the generator is performing good, the output of discriminator will be ones for images—as it will classify them as real For the discriminator the real cross-entropy is calculated using discriminator output of real images in comparison to image array of ones (where a value of “1” denotes the class of real image), and fake cross-entropy is calculated in comparison of discriminator output of fake images (expected “0”) to an array of zeros. Following this, cross-entropy of discriminator
is calculated as the sum of cross entropies of the real (collected) output
and fake (generated) output
:
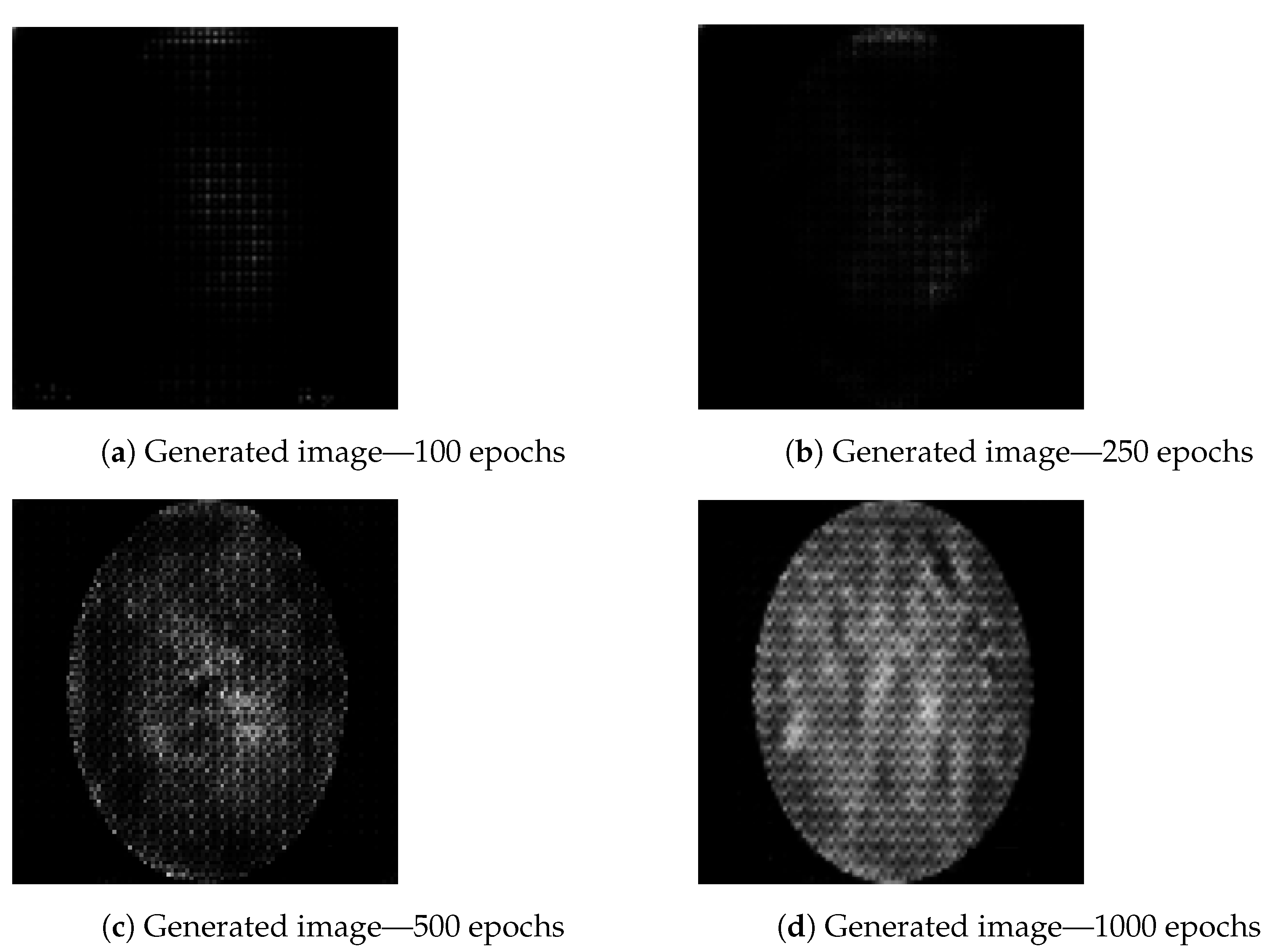
The images are generated using a generator and discriminator with the above-described hyperparameters, with the only variation being the number of epochs (iterations) for which the generator is trained to generate new images. Three different epoch ranges are used: 100 epochs, 250 epochs, 500 epochs and 1000 epochs. The number of epochs is an important hyperparameter for GAN performance tuning. Having a number set too low will cause the generator network to be undertrained and images will not be of satisfactory quality. Setting the number of epochs too high will have a negative effect on training times [
46]. Still, under training can generate weak results, as it can be seen in
Figure 5, where image generated using 100 epochs (
Figure 5a) contains lower amount of data. The same figure shows an increase in details from the image obtained while training with 500 epochs (
Figure 5c) to image obtained from training the generator over 1000 epochs (
Figure 5d). The artifacts apparent in
Figure 5c,d are to be expected as a result of previously described transposed convolution process, known as “chekerboard” artifacts [
47]. Still, while the effect of number of epochs is visible to the naked eye, the way the number of epochs affects actual data contained within the image in terms of classification scoring cannot be determined in this way, and training and testing need to be performed using predefined CNN architectures, results of which are presented in the following section. It can be possible that images which, to a human observer, appear to be weaker could contain enough data to better the classifier performance, while the images which appear better could be a result of overfitting and fail to do the same [
27].
7. Results and Discussion
In order to compare the achieved results and to determine the influence of GAN-based augmentation on the proposed classifiers, first the performances achieved by classifiers trained with the original dataset must be measured. In this case, it can be noticed that higher results (up to 0.97) are achieved if VGG-16 is used. Furthermore, it can be noticed that slightly lower classification performances are achieved when AlexNet is used for classification, as presented in
Table 5.
It can be noticed that the AlexNet architecture that has achieved the highest classification performances is characterized with higher and . For these reasons it can be concluded that AlexNet has lower generalization ability in comparison with VGG-16. Furthermore, it can be concluded that VGG-16 deals better with the dataset diversity. This property can be attributed to its deeper architecture.
When the influence of GAN-parameters and share of generated images in the training dataset on the performances of the top-ranked AlexNet models are observed, it is interesting to notice that the highest performances are achieved by increasing the number of GAN epochs to the particular level. It is interesting to notice that increasing the share of generated images in the training dataset does not increase classification performances, rather it decreases them. It can be noticed that the lowest classification performances are achieved when the set augmented with images generated by GAN in 100 epochs. However, these classification performances are still higher than classification performances achieved without data augmentation, pointing to the conclusion that GAN-based augmentation has, in general, a positive impact on the image classification performed by using AlexNet. All statement could be supported by data presented in
Table 6, where the first column corresponds to nomenclature presented in
Table 4.
When results achieved with VGG-16 are compared, it can be noticed that in all cases,
and
up to 0.99 are achieved. As in the case of AlexNet, lower classification performances are achieved if the original training dataset is augmented with images generated by GAN in 1000 epochs. Furthermore, a slight performance decay can be seen in the case when 18180 generated images are used for augmenting the original set. For these reasons, it can be concluded that a higher share of augmented images reduces the performances of top-ranked VGG-16 architectures. All presented facts could be seen in
Table 7.
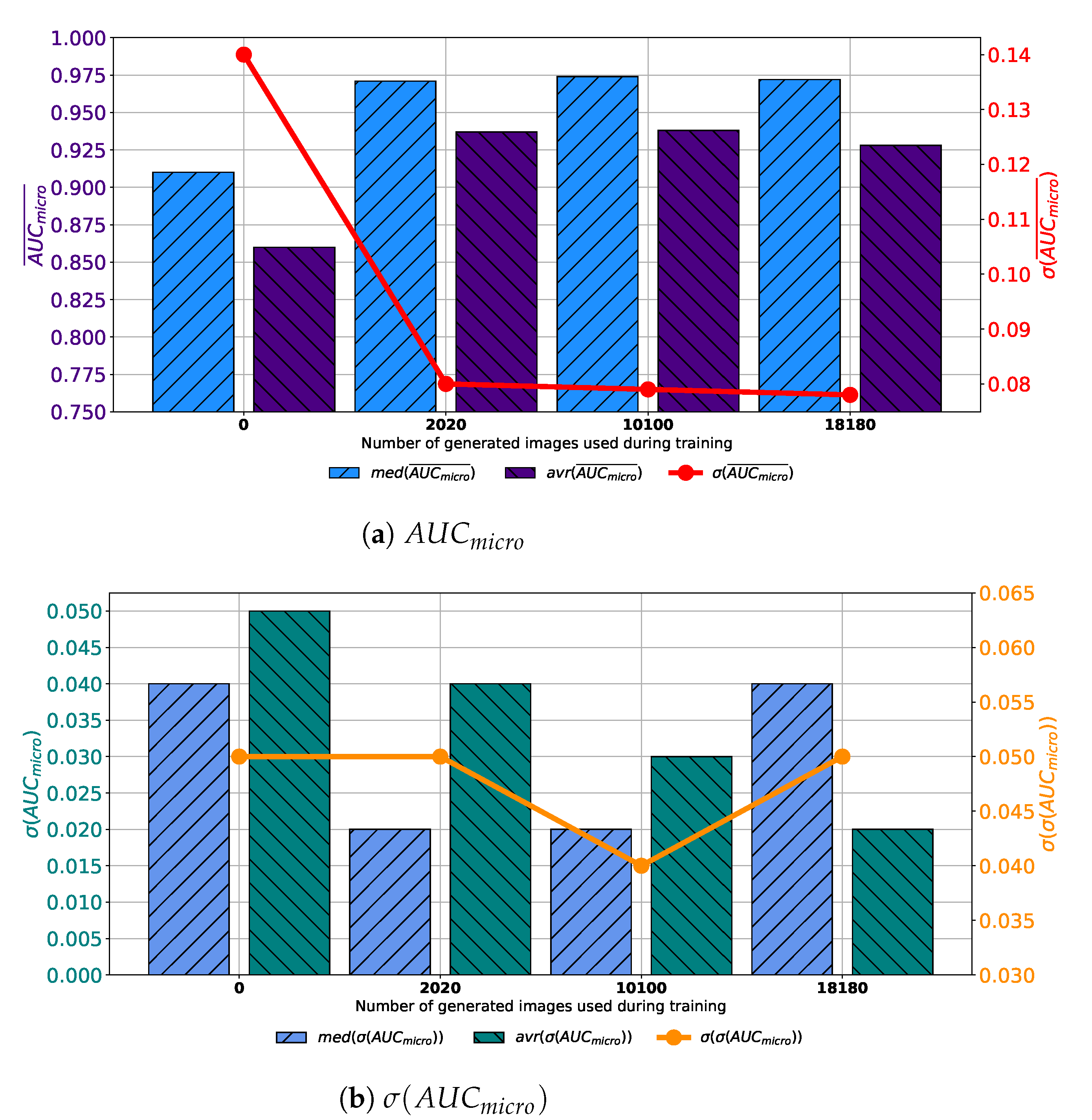
If results achieved with each variation of AlexNet trained with images generated by GAN in 100 consecutive epochs are compared, it can be noticed that the highest median and average
is achieved if the training dataset is augmented with 10,100 GAN-generated images. Furthermore, it can be noticed that the standard deviation of
is showing a significant decreasing trend suggesting the more stable network behavior, as presented in
Figure 7a. When
is observed, it can be noticed that lower median and average values were achieved if augmented datasets are used for training. It can be noticed that the lowest median and
values are achieved if 10,100 generated images are added to the training set. The standard deviation of
is also the lowest in the same case, as presented in
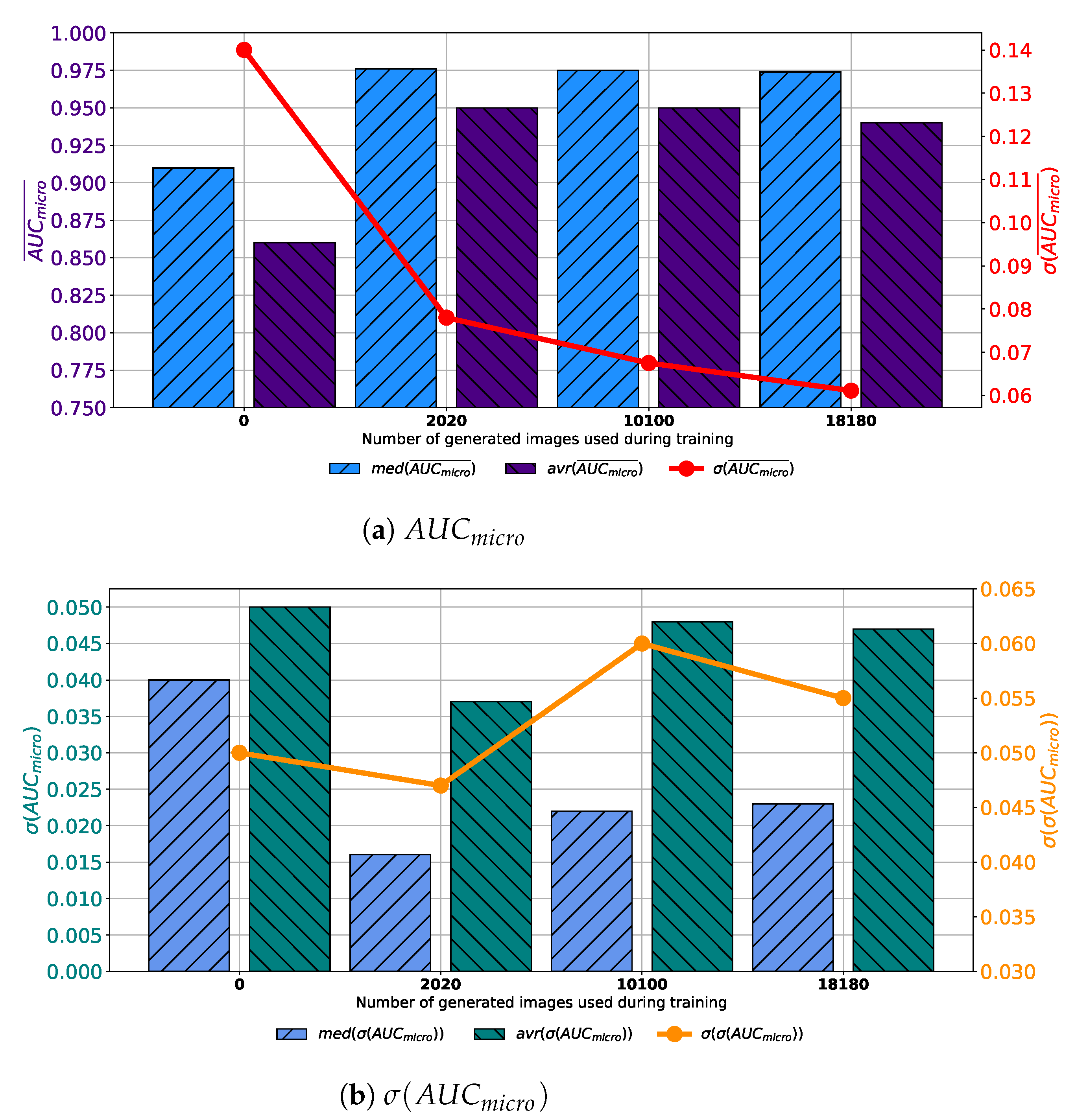
Figure 7b. Presented results are suggesting improvement from a standpoint of generalization when an augmented training dataset is used. When presented results are summed up, it can be concluded that, in the case of 100 consecutive GAN epochs, the highest performances are achieved if 10,100 images are added to the training dataset.
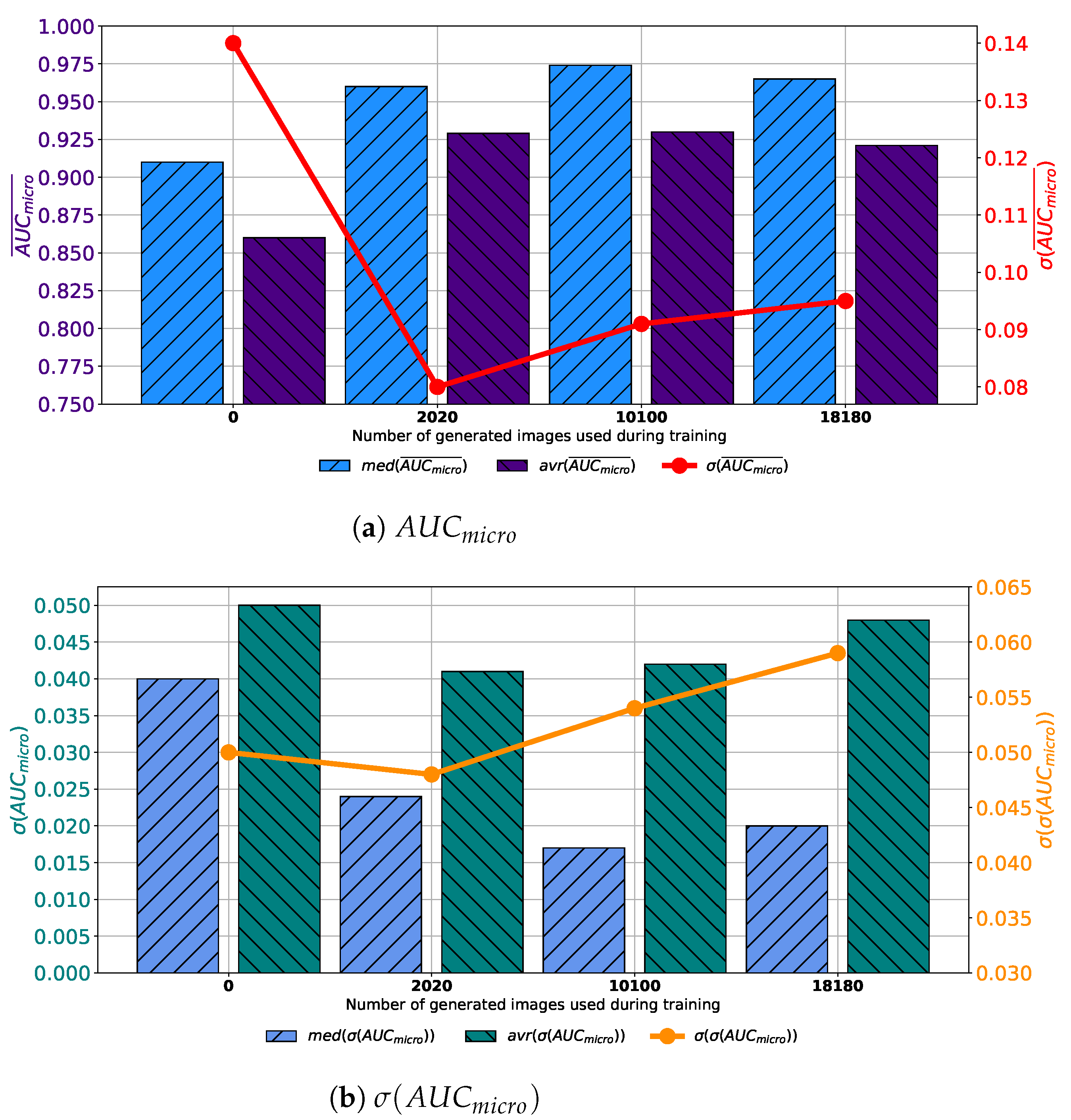
Similar results are achieved when images are generated with GAN trained for 250 epochs. In this case, a set assembled with original images combined with 10,100 generated images again achieves the highest median and average values of
. The difference lays in the fact that the standard deviation of
is the lowest when 2020 generated images are used for training dataset construction, as presented in
Figure 8a. When
is observed, it can be noticed that the lowest average value is achieved in the case of 2020 images, while the lowest median value is achieved in the case of 10,100 images. The standard deviation of
achieves the minimum value of 0.026 when Case 2 is observed. By combining results from both
and
standpoint it can be concluded that, again, the best results will be achieved if 10,100 generated images are added to the training dataset. This conclusion could be drawn from the fact that average and median
values are significantly lower in Case 2, as seen in
Figure 8b. This fact, due to classification capability, plays the dominant role in variation rankings, regardless of better generalization performances of the configuration presented in Case 2.
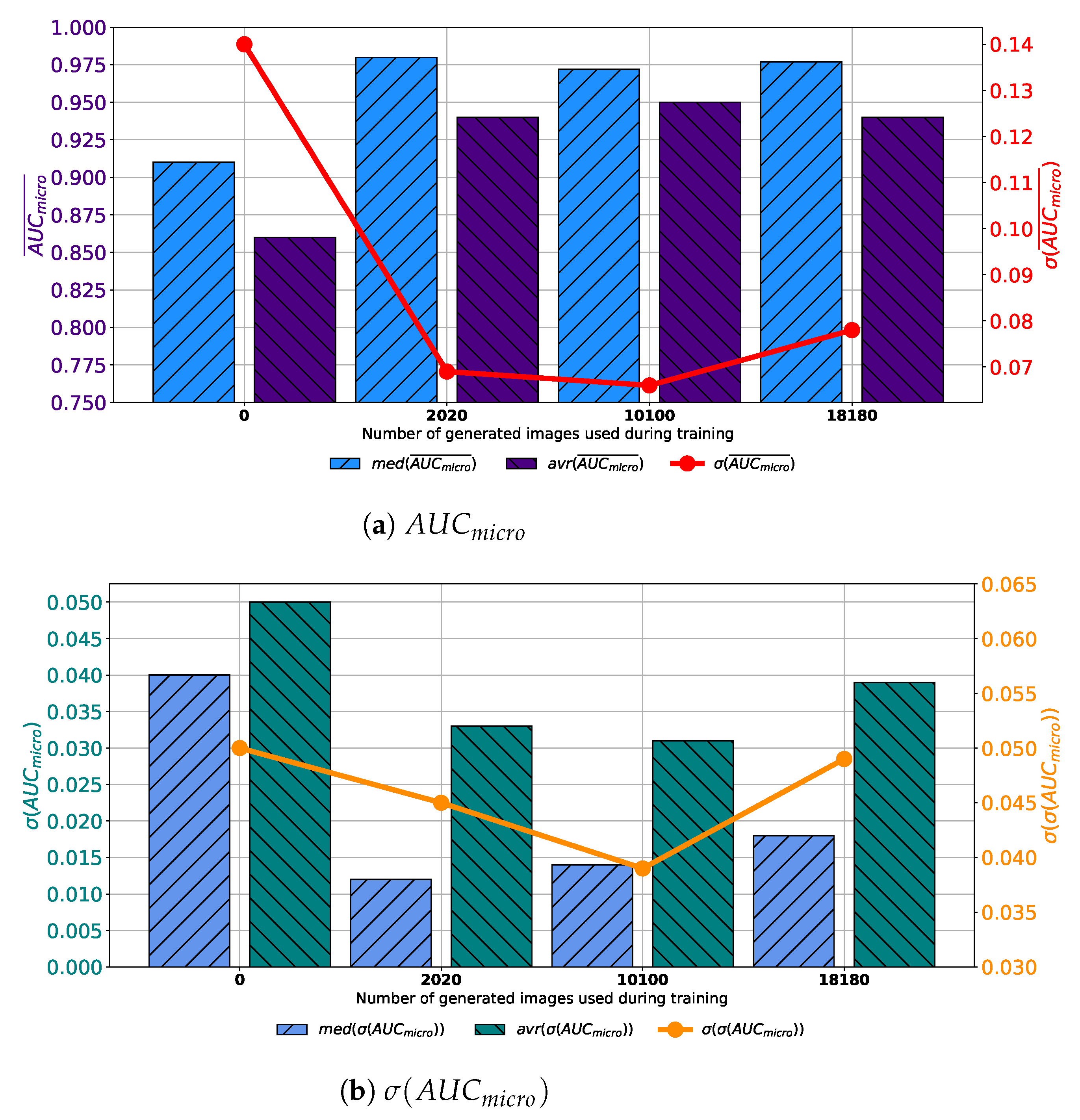
When images generated by using GAN executed for 500 consecutive epochs are added to the training dataset, similar results are achieved. In this case, average
values higher up to 0.95 are achieved. Presented value is achieved when configuration marked as Case 3 is used. Is it interesting to notice that median values are higher in Case 2 and Case 4 than in Case 3, as presented in
Figure 9a. When the standard deviation of
is observed, it can be seen that the lowest value is achieved in Case 3. For these reasons, it can be concluded that the presented configuration has the best consistency of classification performances, regardless of batch size, the number of training epochs and solver used. When generalization performances are observed, it can be noticed that the lowest average
is achieved in Case 3. Furthermore, it is interesting to notice that the median value is lower in Case 2. When the standard deviation of all achieved
values is observed, it can be noticed that the lowest standard deviation is also achieved in the case when 10,100 generated images are added to the training set, as presented in
Figure 8b. From presented results, it can be concluded that the best performances are achieved when Case 3 is used for the construction of the training dataset. This property is valid from both classification and generalization standpoints.
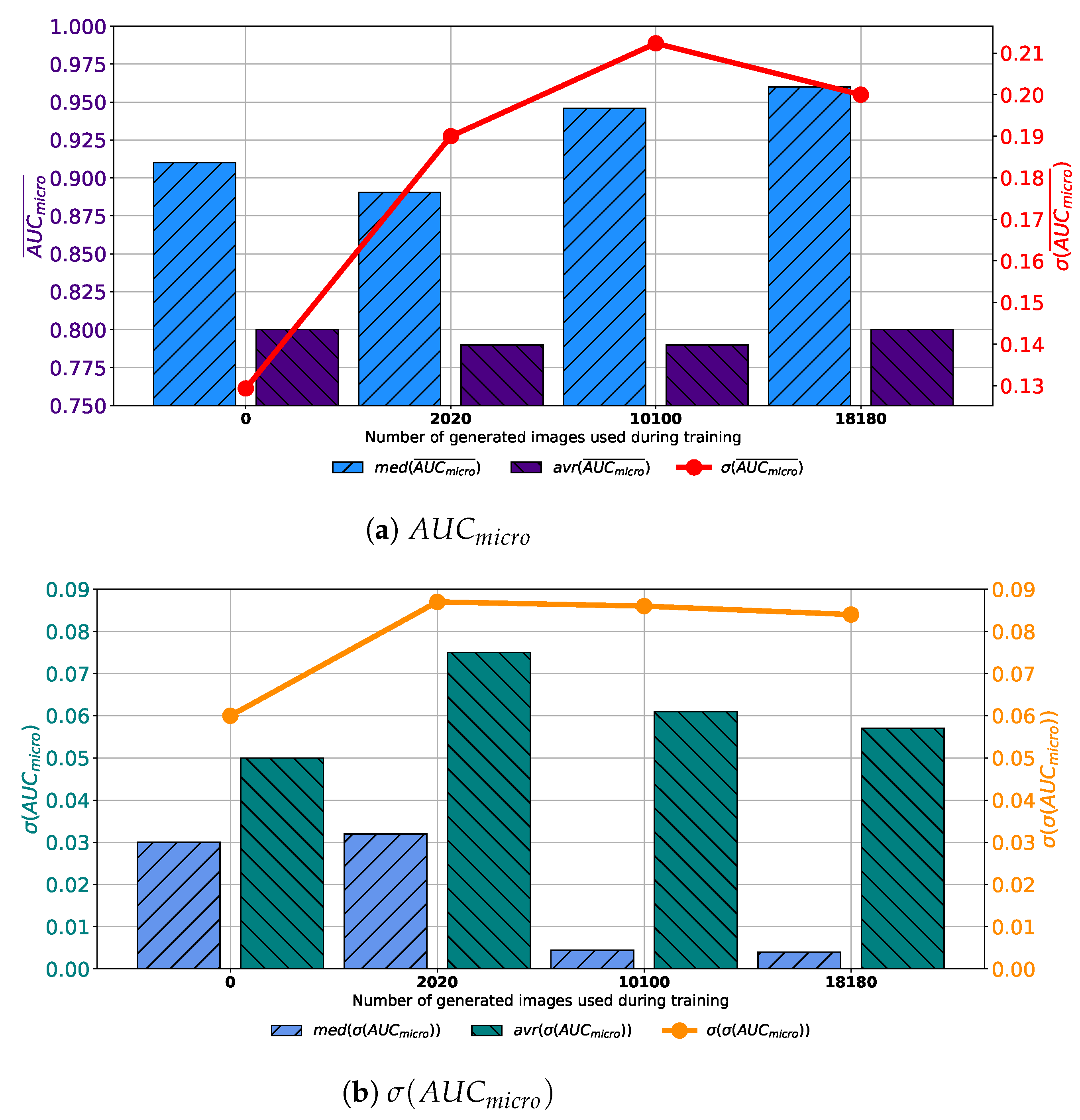
When the training dataset is augmented with images generated by GAN in 1000 consecutive epochs, slightly different conclusions could be drawn. In this case, a decay of median and average values of
can be noticed, as presented in
Figure 10a. It can be noticed that by increasing the share of generated data in the training dataset, no significant change in median and average value can be noticed. Furthermore, a significant fall of standard deviation can be noticed, pointing towards the conclusion that by using a lager share of generated images, a significantly lower change in classification performances is achieved. When generalization performances are observed, it can be noticed that the lowest median and average values of
are achieved in the case when 2020 generated images are used for testing dataset augmentation. Furthermore, it can be noticed that the lowest standard deviation is also achieved in this case, as presented in
Figure 10b. From presented results, it can be noticed that the best results are, in this case, achieved when 2020 generated images are used for data augmentation. When results obtained for all four GAN configurations are summed up, it can be concluded that the dominantly highest performances, both from classification and generalization standpoints, are achieved when images are generated by using GAN for 500 consecutive epochs.
When the results achieved with VGG-16 architecture trained with augmented set generated by GAN executed for 100 executive epochs are observed, it can be noticed that a significant improvement of classification performances can be seen only if 10,100 generated images are added to the training dataset. In all other cases, a significant increase can be seen only when median values are observed, as presented in
Figure 11a. Furthermore, a significant increase in the standard deviation of
can be noticed, suggesting more unstable behavior of networks trained with augmented datasets. When generalization performances are observed, it can be noticed that if augmented training sets are used, higher average values of
are achieved. On the other hand, a significant decrease in median values can be noticed for Case 3 and Case 4. The standard deviation of
does not show a significant difference, regardless of the share of generated images, as presented in
Figure 11b. Presented results are suggesting that there is no significant positive impact on generalization performances of VGG-16 when it is trained by using the training dataset augmented with images generated by GAN in 100 epochs.
When images added to the training dataset are generated by GAN in 250 epochs, similar trends could be noticed. In this case, a slight increase in VGG-16 performances could be noticed. It is interesting to notice that higher median values are achieved when the training dataset is augmented by using 10,100 and 18,180 augmented images. On the other hand, there is no significant change of average
values, as presented in
Figure 12a. By observing the change of standard deviation of
. When generalization performances are observed, it can be noticed a significant decrease of median
can be noticed, as presented in
Figure 12b. On the other hand, there significant decrease of average
and its standard deviation, which is even higher. These results are pointing towards the conclusion that there is no increase in generalization performances.
When classification performances of VGG-16 trained with augmented dataset constructed with images generated by GAN in 500 epochs are compared, a slight increase of average and median value of
can be noticed. In the same time, due to higher standard deviations, a conclusion that defines a more unstable behavior of VGG-16 networks trained with augmented dataset can be drawn. All presented properties are shown in
Figure 13a. When generalization performances are observed, it can be noticed that there is no significant decrease of median
. On the other hand, a decrease in median value can be noticed, as presented in
Figure 13b. As it is in the case of all presented variations of training dataset used for training of VGG-16, there is no significant change in generalization capabilities of such a network. On the other hand, only limited improvements are achieved in classification performances.
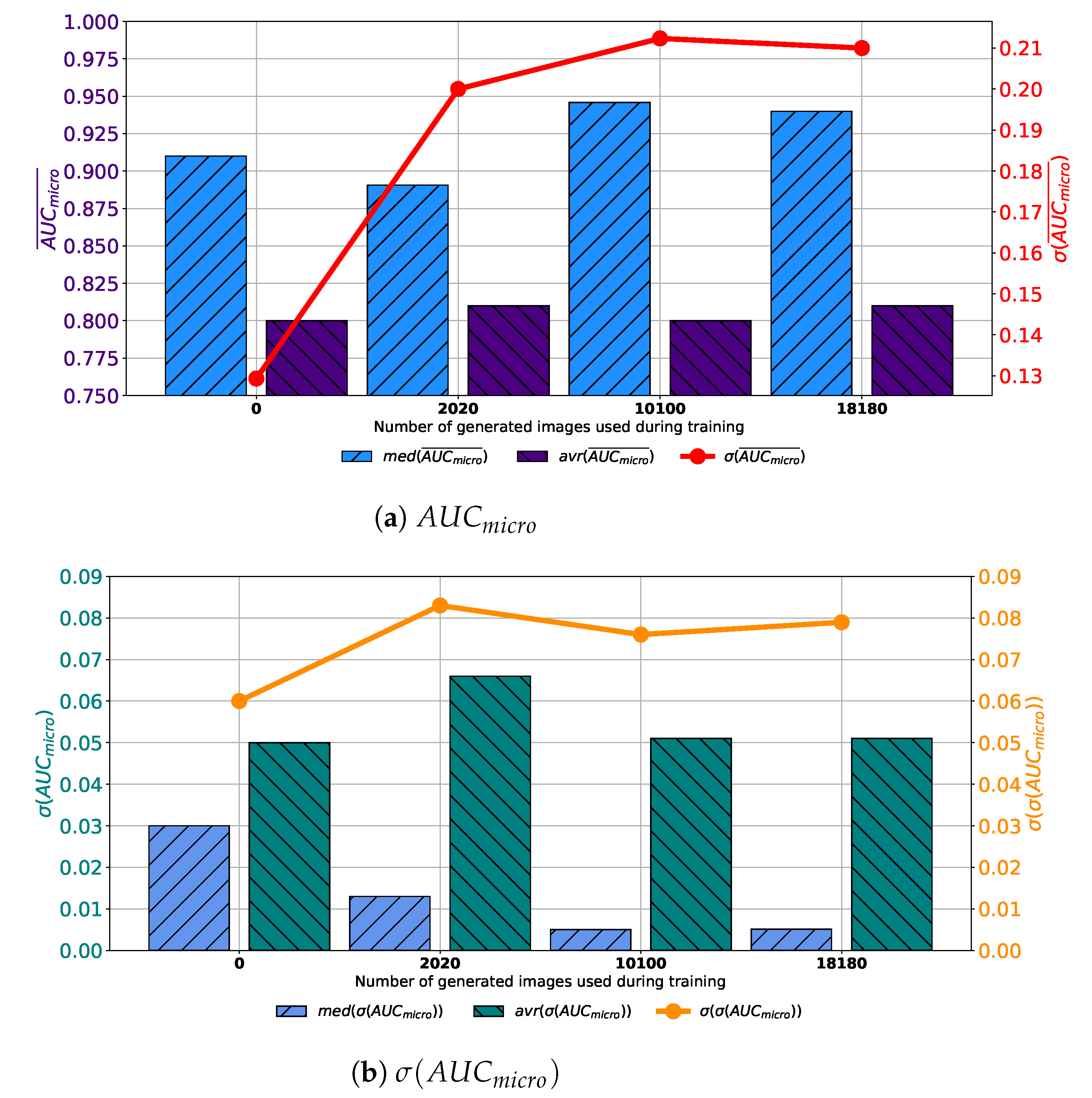
As the last case, VGG-16 trained with a dataset augmented with images generated by GAN in 1000 consecutive epochs. When classification performances are compared, a significant increase of the median value of
can be noticed in Case 2 and Case 3. On the other hand, there is no significant increase in average value, as presented in
Figure 14a. Furthermore, it can be noticed that the standard deviation of
is significantly larger if augmented datasets are used, suggesting the less stable behavior. When generalization performances are observed, it can be noticed that by increasing the share of GAN-generated images, lover median values of
are achieved. This property is valid only for Cases 3 and 4. On the other hand, there is no decrease in average value. In other words, average
is lower when the original dataset is used for the training of VGG-16. When the standard deviation of
is observed, no significant change can be noticed, as presented in
Figure 14b. From presented facts, it can be concluded that there is an increase in generalization performances of VGG-16 when images generated in 1000 epochs are used for training dataset augmentation.
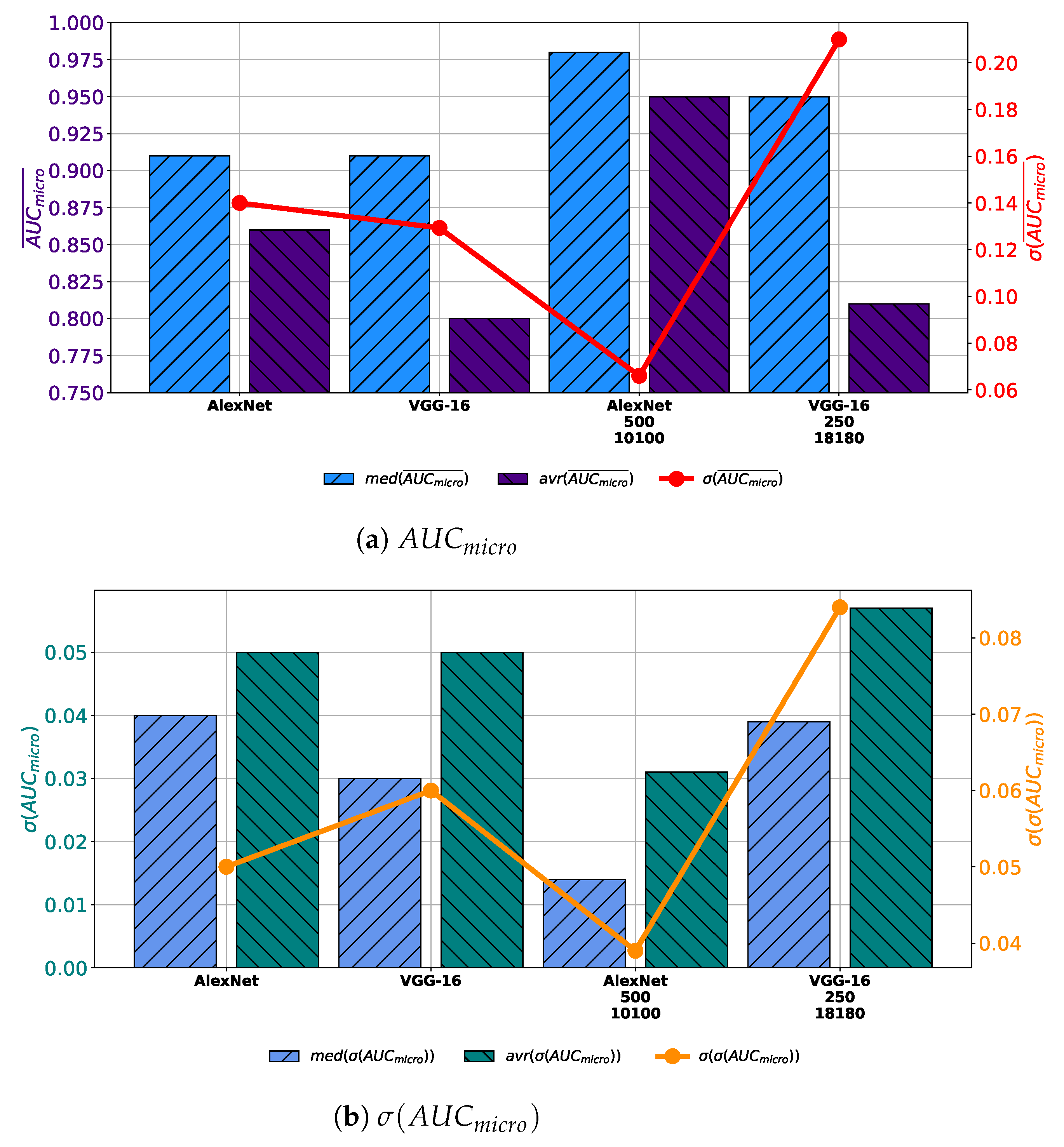
When all obtained results are compared, it can be noticed that by using AlexNet trained with augmented dataset constructed with 10,100 images generated with 500 GAN epochs significantly larger average and median values of
are achieved. Furthermore, the most stable performance is also achieved by using this configuration, judging by the lowest value of the standard deviation. Furthermore, if VGG-16 architecture trained with augmented dataset constructed with 18180 images generated trough 250 epochs is used, no significant improvement in regard to the average and median value of
is achieved. Presented values are only slightly higher than values achieved with VGG-16 architecture trained with the original dataset, as presented in
Figure 15a. When generalization performances are compared, it can be noticed that by using AlexNet architecture lower average and median values of
, as well as its standard deviation. These results are pointing to the conclusion that AlexNet trained with an augmented dataset has achieved higher generalization performances, in comparison with networks trained with the original dataset. On the other hand, it can be noticed that VGG-16 architecture trained with augmented dataset achieves higher average, median and standard deviation of
, as presented in
Figure 15b. These results are pointing to the conclusion that VGG-16 architecture, trained with augmented dataset performs poorer than networks trained with original dataset.