Flavonoids as Potential anti-MRSA Agents through Modulation of PBP2a: A Computational and Experimental Study
Abstract
:1. Introduction
2. Results
2.1. In-Silico Screening
2.1.1. Target Identification Using Inverse Virtual Screening
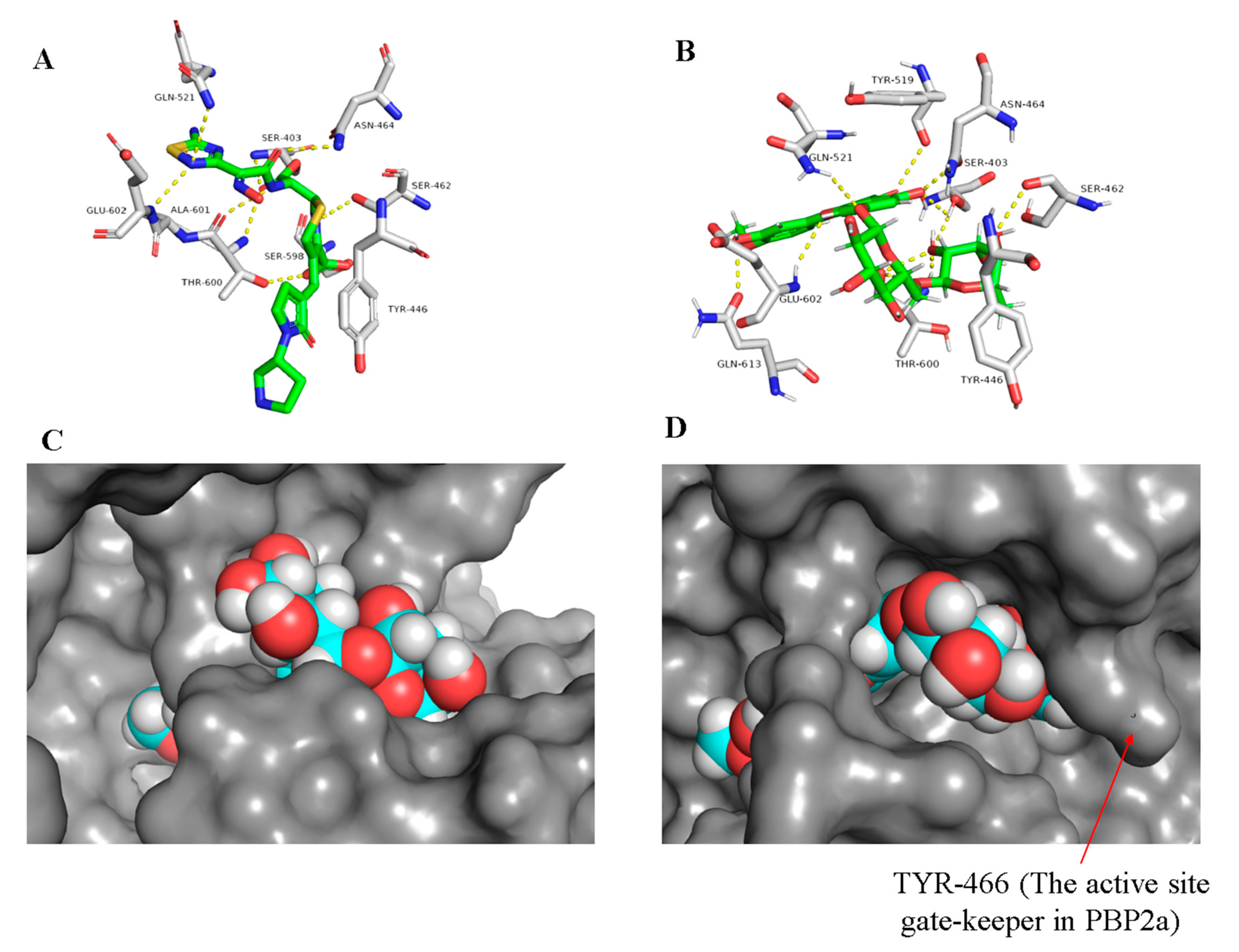
2.1.2. Docking Studies and Molecular Dynamic Simulation Refinement
2.2. In Vitro Inhibitory Activity
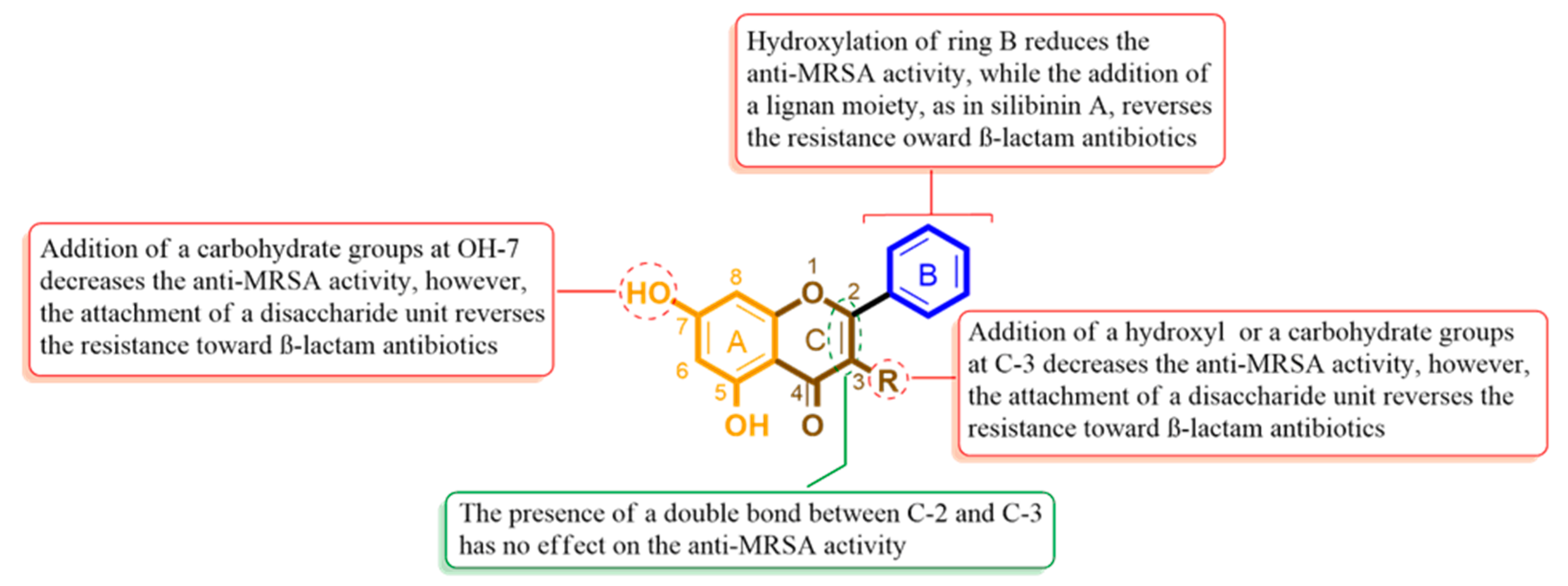
2.3. Structure–Activity Relationship
3. Discussion
4. Materials and Methods
4.1. Library Construction
4.2. Bacterial Strains
4.3. Minimum Inhibitory Concentration (MIC) Assay
4.4. Checkerboard Microdilution Assay
4.5. Inverse Virtual Screening
4.6. Molecular Docking
4.7. Molecular Dynamic Simulation
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Antibiotic Resistance. Available online: https://www.who.int/news-room/fact-sheets/detail/antibiotic-resistance (accessed on 1 August 2020).
- About Antibiotic Resistance. Available online: https://www.cdc.gov/drugresistance/about.htm (accessed on 1 August 2020).
- Kokoska, L.; Kloucek, P.; Leuner, O.; Novy, P. Plant-derived products as antibacterial and antifungal agents in human health care. Curr. Med. Chem. 2019, 26, 5501–5541. [Google Scholar] [CrossRef] [PubMed]
- Lovering, A.L.; Gretes, M.C.; Safadi, S.S.; Danel, F.; De Castro, L.; Page, M.G.; Strynadka, N.C. Structural insights into the anti-methicillin-resistant Staphylococcus aureus (MRSA) activity of ceftobiprole. J. Biol. Chem. 2012, 287, 32096–32102. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sayed, A.M.; Alhadrami, H.A.; El-Hawary, S.S.; Mohammed, R.; Hassan, H.M.; Rateb, M.E.; Bakeer, W. Discovery of two brominated oxindole alkaloids as Staphylococcal DNA gyrase and pyruvate kinase inhibitors via inverse virtual screening. Microorganisms 2020, 8, 293. [Google Scholar] [CrossRef] [Green Version]
- Wang, J.C.; Chu, P.Y.; Chen, C.M.; Lin, J.H. idTarget: A web server for identifying protein targets of small chemical molecules with robust scoring functions and a divide-and-conquer docking approach. Nuc. Acids Res. 2012, 40, W393–W399. [Google Scholar] [CrossRef] [Green Version]
- Bernard, F.X.; Sable, S.; Cameron, B.; Provost, J.; Desnottes, J.F.; Crouzet, J.; Blanche, F. Glycosylated flavones as selective inhibitors of topoisomerase IV. Antimicrob. Agents Chemotherap. 1997, 41, 992–998. [Google Scholar] [CrossRef] [Green Version]
- Wu, D.; Kong, Y.; Han, C.; Chen, J.; Hu, L.; Jiang, H.; Shen, X. D-Alanine: D-alanine ligase as a new target for the flavonoids quercetin and apigenin. Int. J. Antimicrob. Agents 2008, 32, 421–426. [Google Scholar] [CrossRef]
- Gaillard, T. Evaluation of AutoDock and AutoDock Vina on the CASF-2013 benchmark. J. Chem. Inform. Model. 2018, 58, 1697–1706. [Google Scholar] [CrossRef] [PubMed]
- Ramírez, D.; Caballero, J. Is it reliable to take the molecular docking top scoring position as the best solution without considering available structural data? Molecules 2018, 23, 1038. [Google Scholar] [CrossRef] [Green Version]
- Sayed, A.M.; Alhadrami, H.A.; El-Gendy, A.O.; Shamikh, Y.I.; Belbahri, L.; Hassan, H.M.; Abdelmohsen, U.R.; Rateb, M.E. Microbial Natural Products as Potential Inhibitors of SARS-CoV-2 Main Protease (Mpro). Microorganisms 2020, 8, 970. [Google Scholar] [CrossRef]
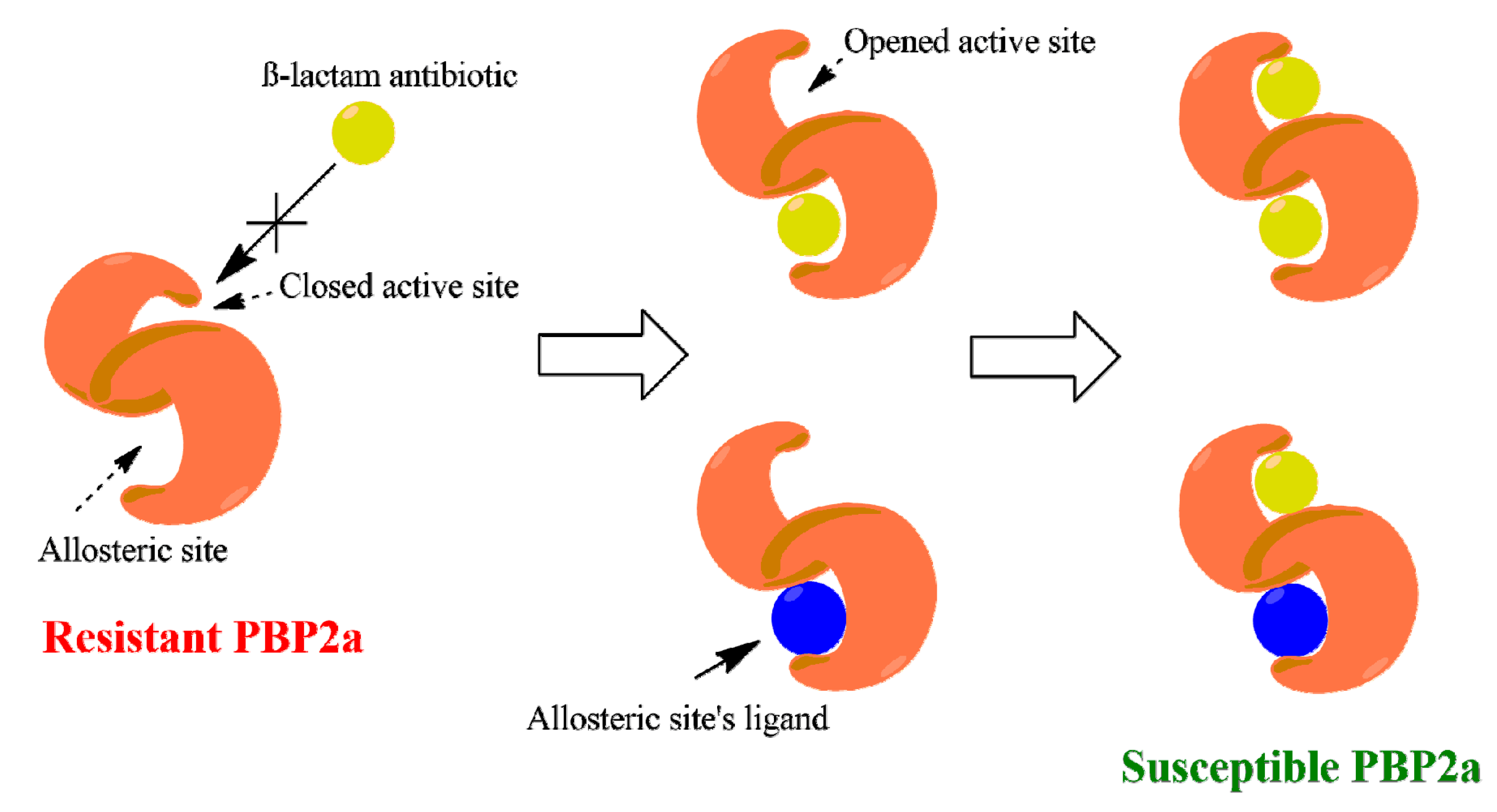
- Otero, L.H.; Rojas-Altuve, A.; Llarrull, L.I.; Carrasco-López, C.; Kumarasiri, M.; Lastochkin, E.; Johnson, J.W. How allosteric control of Staphylococcus aureus penicillin binding protein 2a enables methicillin resistance and physiological function. Proc. Natl. Acad. Sci. USA 2013, 110, 16808–16813. [Google Scholar] [CrossRef] [Green Version]
- Shalaby, M.A.W.; Dokla, E.M.; Serya, R.A.; Abouzid, K.A. Penicillin binding protein 2a: An overview and a medicinal chemistry perspective. Eur. J. Med. Chem. 2020, 112312. [Google Scholar] [CrossRef] [PubMed]
- Chassagne, F.; Cabanac, G.; Hubert, G.; David, B.; Marti, G. The landscape of natural product diversity and their pharmacological relevance from a focus on the Dictionary of Natural Products®. Phytochem. Rev. 2019, 18, 601–622. [Google Scholar] [CrossRef] [Green Version]
- Dixon, R.A. Natural products and plant disease resistance. Nature 2001, 411, 843–847. [Google Scholar] [CrossRef] [PubMed]
- Gibbons, S. Anti-staphylococcal plant natural products. Nat. Prod. Rep. 2004, 21, 263–277. [Google Scholar] [CrossRef]
- Wishart, D.S.; Tzur, D.; Knox, C.; Eisner, R.; Guo, A.C.; Young, N. HMDB: The human metabolome database. Nuc. Acids Res. 2007, 35, D521–D526. [Google Scholar] [CrossRef]
- Rani, N.; Vijayakumar, S.; PTV, L.; Arunachalam, A. Allosteric site-mediated active site inhibition of PBP2a using Quercetin 3-O-rutinoside and its combination. J. Biomol. Struc. Dynam. 2016, 34, 1778–1796. [Google Scholar] [CrossRef]
- Usman Amin, M.; Khurram, M.; Khan, T.A.; Faidah, H.S.; Ullah Shah, Z.; Ur Rahman, S.; Iriti, M. Effects of luteolin and quercetin in combination with some conventional antibiotics against methicillin-resistant Staphylococcus aureus. Int. J. Mol. Sci. 2016, 17, 1947. [Google Scholar] [CrossRef] [Green Version]
- Shuai-cheng, W.; Qian, L.; Zhi-Qiang, Y.; Fei, L.; Wen-Jing, P.; Xiang-Bin, S.; JIan-Zhong, S. Antibacterial effect and mode of action of flavonoids from licorice against methicillin-resistant Staphylococcus aureus. Front. Microbiol. 2019, 10, 2489. [Google Scholar] [CrossRef] [Green Version]
- Hatano, T.; Kusuda, M.; Inada, K.; Ogawa, T.O.; Shiota, S.; Tsuchiya, T.; Yoshida, T. Effects of tannins and related polyphenols on methicillin-resistant Staphylococcus Aureus. Phytochem. 2005, 66, 2047–2055. [Google Scholar] [CrossRef]
- Mun, S.H.; Lee, Y.S.; Han, S.H.; Lee, S.W.; Cha, S.W.; Kim, S.B. In vitro potential effect of morin in the combination with β-lactam antibiotics against methicillin-resistant Staphylococcus aureus. Foodborn Pathogen. Dis. 2015, 12, 545–550. [Google Scholar] [CrossRef]
- Pushkaran, A.C.; Vinod, V.; Vanuopadath, M.; Nair, S.S.; Nair, S.V.; Vasudevan, A.K. Combination of repurposed drug diosmin with amoxicillin-clavulanic acid causes synergistic inhibition of mycobacterial growth. Sci. Rep. 2019, 9, 1–14. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tong, Y.; Zhou, X.M.; Wang, S.J.; Yang, Y.; Cao, Y.L. Analgesic activity of myricetin isolated from Myrica Rubra Sieb. Zucc. Leaves. Arch. Pharm. Res. 2009, 32, 527–533. [Google Scholar] [CrossRef] [PubMed]
- Fang, X.K.; Gao, J.; Zhu, D.N. Kaempferol and quercetin isolated from Euonymus alatus improve glucose uptake of 3T3-L1 cells without adipogenesis activity. Life Sci. 2008, 82, 615–622. [Google Scholar] [CrossRef]
- Wolfman, C.; Viola, H.; Paladini, A.; Dajas, F.; Medina, J.H. Possible anxiolytic effects of chrysin, a central benzodiazepine receptor ligand isolated from Passiflora coerulea. Pharmacol. Biochem. Behav. 1994, 47, 1–4. [Google Scholar] [CrossRef]
- Londoño-Londoño, J.; De Lima, V.R.; Jaramillo, C.; Creczynski-pasa, T. Hesperidin & hesperetin membrane interaction: Understanding role of 7-O-glycoside moiety in flavonoids. Arch. Biochem. Biophys. 2010, 499, 6–16. [Google Scholar] [PubMed]
- El-Hawary, S.S.; Sayed, A.M.; Mohammed, R.; Hassan, H.M.; Zaki, M.A.; Rateb, M.E.; Abdelmohsen, U.R. Epigenetic modifiers induce bioactive phenolic metabolites in the marine-derived fungus Penicillium brevicompactum. Mar. Drugs. 2018, 16, 253. [Google Scholar] [CrossRef] [Green Version]
- Jeong, S.I.; Han, W.S.; Yun, Y.H.; Kim, K.J. Continentalic acid from Aralia continentalis shows activity against methicillin-resistant Staphylococcus Aureus. Phytother. Res. 2006, 20, 511–514. [Google Scholar] [CrossRef]
- Lorian, V. Antibiotics in Laboratory Medicine; Lippincott Williams & Wilkins: New York, NY, USA, 2005. [Google Scholar]
- Fratini, F.; Mancini, S.; Turchi, B.; Friscia, E.; Pistelli, L.; Giusti, G.; Cerri, D. A novel interpretation of the fractional inhibitory concentration index: The case Origanum vulgare L. and Leptospermum scoparium JR et G. Forst essential oils against Staphylococcus aureus strains. Microbiol. Res. 2017, 195, 11–17. [Google Scholar] [CrossRef]
- Cozzini, P. From Medicinal Chemistry to Food Science: A Transfer of In Silico Methods Applications; Nova Science Pub Inc.: Hauppauge, NY, USA, 2015; UK ed. Edition. [Google Scholar]
- Seeliger, D.; de Groot, B.L. Ligand docking and binding site analysis with PyMOL and Autodock/Vina. J. Comput. Mol. Des. 2010, 24, 417–422. [Google Scholar] [CrossRef] [Green Version]
- Lill, M.A.; Danielson, M.L. Computer-aided drug design platform using PyMOL. J. Comput. Mol. Des. 2011, 25, 13–19. [Google Scholar] [CrossRef]
- Phillips, J.C.; Braun, R.; Wang, W.; Gumbart, J.; Tajkhorshid, E.; Villa, E.; Schulten, K. Scalable molecular dynamics with NAMD. J. Comput. Chem. 2005, 26, 1781–1802. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- MacKerell, A.D., Jr.; Bashford, D.; Bellott, M.L.; Dunbrack, R.L., Jr.; Evanseck, J.D.; Field, M.J.; Fischer, S.; Gao, J.; Guo, H.; Ha, S.; et al. All-atom empirical potential for molecular modeling and dynamics studies of proteins. J. Phys. Chem. B 1998, 102, 3586–3616. [Google Scholar] [CrossRef] [PubMed]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD: Visual molecular dynamics. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef]
- Jo, S.; Kim, T.; Iyer, V.G.; Im, W. CHARMM-GUI: A web-based graphical user interface for CHARMM. J. Comput. Chem. 2008, 29, 1859–1865. [Google Scholar] [CrossRef] [PubMed]
- Jiménez, J.; Skalic, M.; Martinez-Rosell, G.; De Fabritiis, G. K deep: Protein–ligand absolute binding affinity prediction via 3d-convolutional neural networks. J. Chem. Informat. Mod. 2018, 58, 287–296. [Google Scholar] [CrossRef]





| Ligand | ∆GVina (Active and Allosteric Sites) | ∆G * KDEEP (Active and Allosteric Sites) | Hydrogen Bonding Interactions | Hydrophobic Interactions | ||
|---|---|---|---|---|---|---|
| Active Site | Allosteric Site | Active Site | Allosteric Site | |||
| Myricetin (1) | >−7.0, −7.0 | >−7.0, −6.8 | ASN-464, TYR-519, GLN-521 | TYR-105, ASN-146 | TYR-446 | TYR-297 |
| Quercetin (2) | −7.2, −7.2 | <−7.0, −7.1 | ASN-464, TYR-519, GLN-521 | TYR-105,TYR-297 | TYR-446 | - |
| Kaempferol (3) | −7.6, <−7.0 | <−7.0, <−7.0 | SER-403 GLN-521 | ASN-146, ILE-144 | TYR-446 | - |
| Apigenin (4) | −8.0, <−7.0 | −7.8, <−7.0 | SER-403, ASN-464, TYR-519, GLN-521 | ARG-298, GLY-296 | TYR-446 | - |
| Chrysin (5) | −7.9, <−7.0 | −8.0, <−7.0 | SER-403, ASN-464, TYR-519, GLN-521, THR-600 | GLU-145, ASP-295 | TYR-446 | TYR-297 |
| Hesperetin (6) | −7.8, −7.2 | −8.0, <−7.0 | SER-403, ASN-464, TYR-519, THR-600 | ILE-144, ASP-295 | TYR-446 | TYR-297 |
| Astragalin (7) | −9.0, −8.1 | −8.7, −8.5 | SER-403, ASN-464, TYR-519, THR-600, GLN-613 | ILE-144, GLU-145, ASN-146, GLY-296 | - | TYR-105 |
| Kaempferol 7-O-glucoside (8) | −9.1, −7.9 | −8.7, −8.4 | SER-403, SER-462, ASN-464, TYR-519 | TYR-105, ILE-144, GLU-145, ASN-146 | - | TYR-297 |
| Quercitrin (9) | −9.0, −8.1 | −8.8, −8.3 | SER-403, ASN-464, TYR-519, THR-600, GLN-613 | ILE-144, GLU-145, ASN-146, GLY-296 | - | TYR-105 |
| Rutin (10) | −9.4, −8.8 | −9.0, −8,7 | SER-403, ASN-464, TYR-519, THR-600, GLN-613 | TYR-105, ILE-144, GLU-145, ASN-146, GLY-296 | - | - |
| Diosmin (11) | −9.6, −9.8 | −9.9, −9.2 | SER-403, SER-462, ASN-464, TYR-519, THR-600, GLN-613 | TYR-105, ILE-144, GLU-145, ASN-146, GLY-296 | - | - |
| Hesperidin (12) | −9.5, −10.3 | −9.6, −9.9 | SER-403, SER-462, ASN-464, TYR-519, THR-600, GLN-613 | TYR-105, ILE-144, GLU-145, ASN-146, GLY-296, ARG-298, SER-306 | - | TYR-297 |
| Silibinin A (13) | −8.8, −9.5 | −8.9, −9.3 | SER-403, SER-462, ASN-464, TYR-519, THR-600 | TYR-105, ILE-144, GLU-145, ASN-146, ASP-295, GLY-296, ARG-298, SER-306 | TYR-446 | TYR-105, TYR-297 |
| Ampicillin (23) | >−7.0, >−7.0 | >−7.0, −7.1 | SER-462, ASN-464, TYR-519 | ILE-144, GLU-145, ASN-146 | - | TYR-297 |
| Co-crystalized ligands (24 and 25) | −8.9 a, −9.1 b | −8.5 a, −9.0 b | SER-403 #, SER-462, ASN-464, GLN-521, SER-598, AlA-601 GLU-602, THR-600 | TYR-105, ILE-144, ASN-146, ARG-298 | TYR-446 | TYR-297 |
| Tested Compound | MIC MSSA (µg/mL) | MIC MRSA (µg/mL) | FICI * | Inference |
|---|---|---|---|---|
| Myricetin (1) | 31.25 | 125 | 2 | Indifferent |
| Quercetin (2) | 31.25 | 62.5 | 1.5 | Indifferent |
| Kaempferol (3) | 31.25 | 62.5 | 1.5 | Indifferent |
| Apigenin (4) | 15.62 | 31.25 | 1 | Additive |
| Chrysin (5) | 15.62 | 15.62 | 1 | Additive |
| Hesperetin (6) | 15.62 | 31.25 | 1 | Additive |
| Astragalin (7) | 31.25 | 250 | 1.5 | Indifferent |
| kaempferol 7-O-glucoside (8) | 31.25 | 250 | 1.5 | Indifferent |
| Quercitrin (9) | 31.25 | 250 | 1.5 | Indifferent |
| Rutin (10) | 125 | 250 | 0.625 | Synergistic |
| Diosmin (11) | 31.25 | 250 | 0.31 | Synergistic |
| Hesperidin (12) | 31.25 | 250 | 0.31 | Synergistic |
| Silibinin A (13) | 62.5 | 250 | 0.31 | Synergistic |
| Resveratrol (14) | 125 | 250 | 1.5 | Indifferent |
| Caffeic acid (15) | >250 | 250 | 1.5 | Indifferent |
| Sinapic acid (16) | 125 | >250 | 2 | Indifferent |
| Rosmarinic acid (17) | >250 | >250 | 2 | Indifferent |
| Gallic acid (18) | >250 | >250 | 2 | Indifferent |
| Syringic acid (19) | >250 | >250 | 2 | Indifferent |
| Trimethoxy benzoic acid (20) | >250 | >250 | 2 | Indifferent |
| Gentisic acid (21) | >250 | >250 | 2 | Indifferent |
| Benzyle anisate (22) | >250 | >250 | 2 | Indifferent |
| Ampicillin (23) | 0.25 | 125 | - | - |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Alhadrami, H.A.; Hamed, A.A.; Hassan, H.M.; Belbahri, L.; Rateb, M.E.; Sayed, A.M. Flavonoids as Potential anti-MRSA Agents through Modulation of PBP2a: A Computational and Experimental Study. Antibiotics 2020, 9, 562. https://doi.org/10.3390/antibiotics9090562
Alhadrami HA, Hamed AA, Hassan HM, Belbahri L, Rateb ME, Sayed AM. Flavonoids as Potential anti-MRSA Agents through Modulation of PBP2a: A Computational and Experimental Study. Antibiotics. 2020; 9(9):562. https://doi.org/10.3390/antibiotics9090562
Chicago/Turabian StyleAlhadrami, Hani A., Ahmed A. Hamed, Hossam M. Hassan, Lassaad Belbahri, Mostafa E. Rateb, and Ahmed M. Sayed. 2020. "Flavonoids as Potential anti-MRSA Agents through Modulation of PBP2a: A Computational and Experimental Study" Antibiotics 9, no. 9: 562. https://doi.org/10.3390/antibiotics9090562









