Characterization of Fosfomycin and Nitrofurantoin Resistance Mechanisms in Escherichia coli Isolated in Clinical Urine Samples
Abstract
:1. Introduction
2. Methods
2.1. Bacterial Isolates
2.2. Carbohydrate Utilization Test
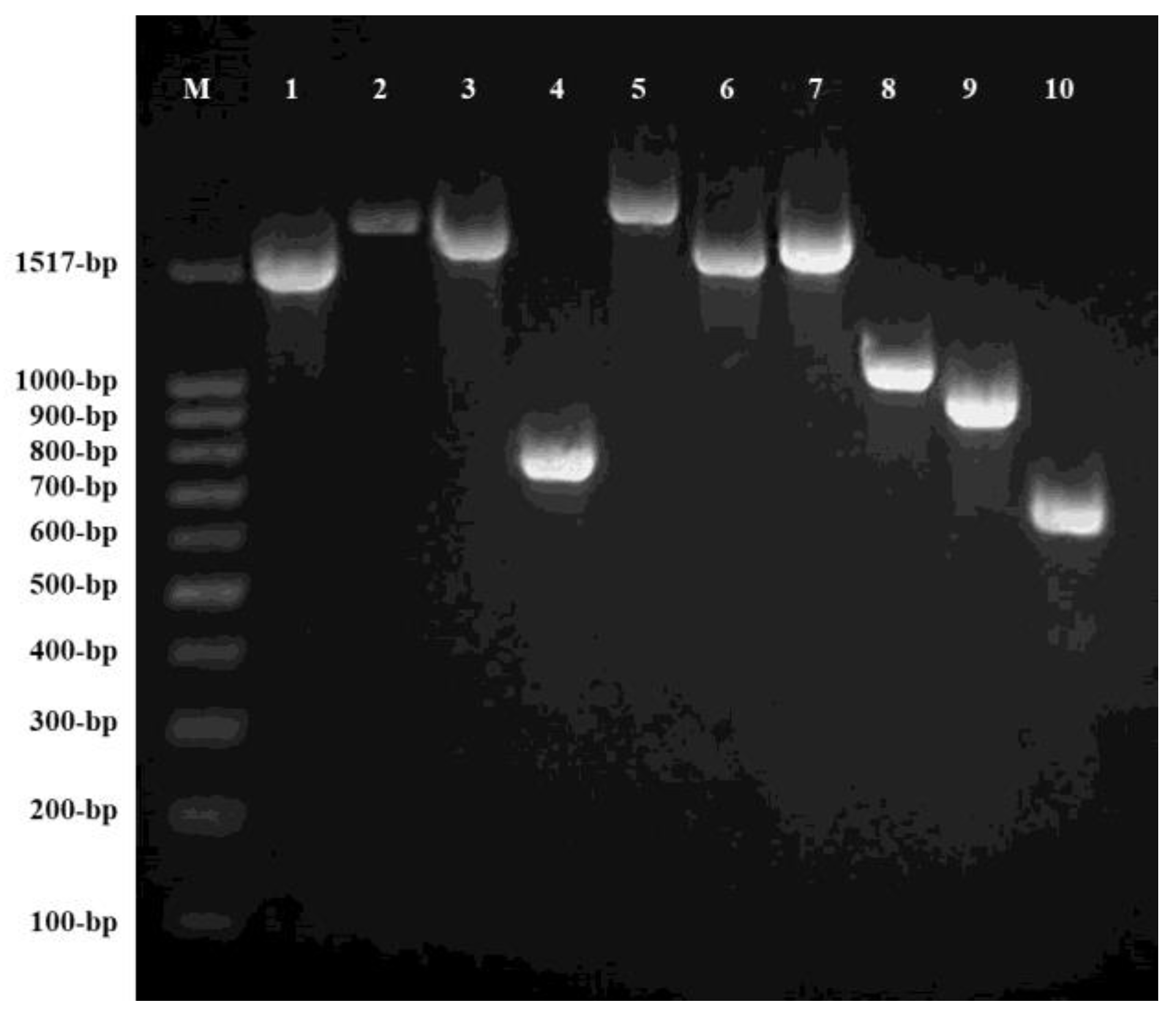
2.3. PCR Amplification
2.4. Nucleotide Sequencing
3. Results
3.1. Fosfomycin Resistance
3.2. Nitrofurantoin Resistance
4. Discussion
4.1. Mechanisms of Resistance to Fosfomycin in E. coli
4.1.1. Modification or Overexpression of the Target (MurA)
4.1.2. Permeability Reduction
4.1.3. Enzymatic Modification of Fosfomycin
4.2. Mechanisms of Resistance to Nitrofurantoin in E. coli
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Sorlozano, A.; Jimenez-Pacheco, A.; de Dios Luna Del Castillo, J.; Sampedro, A.; Martinez-Brocal, A.; Miranda-Casas, C.; Navarro-Marí, J.M.; Gutiérrez-Fernández, J. Evolution of the resistance to antibiotics of bacteria involved in urinary tract infections: A 7-year surveillance study. Am. J. Infect. Control 2014, 42, 1033–1038. [Google Scholar] [CrossRef] [PubMed]
- Vardakas, K.Z.; Legakis, N.J.; Triarides, N.; Falagas, M.E. Susceptibility of contemporary isolates to fosfomycin: A systematic review of the literature. Int. J. Antimicrob. Agents. 2016, 47, 269–285. [Google Scholar] [CrossRef] [PubMed]
- Falagas, M.E.; Kastoris, A.C.; Kapaskelis, A.M.; Karageorgopoulos, D.E. Fosfomycin for the treatment of multidrug-resistant, including extended-spectrum beta-lactamase producing, Enterobacteriaceae infections: A systematic review. Lancet Infect. Dis. 2010, 10, 43–50. [Google Scholar] [CrossRef]
- Sánchez-García, J.M.; Sorlózano-Puerto, A.; Navarro-Marí, J.M.; Gutiérrez Fernández, J. Evolution of the antibiotic-resistance of microorganisms causing urinary tract infections: A 4-year epidemiological surveillance study in a hospital population. Rev. Clin. Esp. 2019, 219, 116–123. [Google Scholar] [CrossRef]
- Sandegren, L.; Lindqvist, A.; Kahlmeter, G.; Andersson, D.I. Nitrofurantoin resistance mechanism and fitness cost in Escherichia coli. J. Antimicrob. Chemother. 2008, 62, 495–503. [Google Scholar] [CrossRef] [Green Version]
- Giske, C.G. Contemporary resistance trends and mechanisms for the old antibiotics colistin, temocillin, fosfomycin, mecillinam and nitrofurantoin. Clin. Microbiol. Infect. 2015, 21, 899–905. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- CLSI. Performance Standards for Antimicrobial Susceptibility Testing, 27th ed.; CLSI supplement M100; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2017. [Google Scholar]
- Nakamura, G.; Wachino, J.; Sato, N.; Kimura, K.; Yamada, K.; Jin, W.; Shibayama, K.; Yagi, T.; Kawamura, K.; Arakawa, Y. Practical agar-based disk potentiation test for detection of fosfomycin-nonsusceptible Escherichia coli clinical isolates producing glutathione S-transferases. J. Clin. Microbiol. 2014, 52, 3175–3179. [Google Scholar] [CrossRef] [Green Version]
- Takahata, S.; Ida, T.; Hiraishi, T.; Sakakibara, S.; Maebashi, K.; Terada, S.; Muratani, T.; Matsumoto, T.; Nakahama, C.; Tomono, K. Molecular mechanisms of fosfomycin resistance in clinical isolates of Escherichia coli. Int. J. Antimicrob. Agents. 2010, 35, 333–337. [Google Scholar] [CrossRef]
- Hou, J.; Huang, X.; Deng, Y.; He, L.; Yang, T.; Zeng, Z.; Chen, Z.; Liu, J.H. Dissemination of the fosfomycin resistance gene fosA3 with CTX-M β-lactamase genes and rmtB carried on IncFII plasmids among Escherichia coli isolates from pets in China. Antimicrob. Agents Chemother. 2012, 56, 2135–2138. [Google Scholar] [CrossRef] [Green Version]
- Benzerara, Y.; Gallah, S.; Hommeril, B.; Genel, N.; Decré, D.; Rottman, M.; Arlet, G. Emergence of plasmid-mediated fosfomycin-resistance genes among Escherichia coli isolates, France. Emerg. Infect. Dis. 2017, 23, 1564–1567. [Google Scholar] [CrossRef] [Green Version]
- Chen, X.; Zhang, W.; Pan, W.; Yin, J.; Pan, Z.; Gao, S.; Jiao, X. Prevalence of qnr, aac(6=)-Ib-cr, qepA, and oqxAB in Escherichia coli isolates from humans, animals, and the environment. Antimicrob. Agents Chemother. 2012, 56, 3423–3427. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Choi, Y.; Chan, A.P. PROVEAN web server: A tool to predict the functional effect of amino acid substitutions and indels. Bioinformatics 2015, 31, 2745–2747. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Castañeda-García, A.; Blázquez, J.; Rodríguez-Rojas, A. Molecular mechanisms and clinical impact of acquired and intrinsic fosfomycin resistance. Antibiotics 2013, 2, 217–236. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, D.H.; Lees, W.J.; Kempsell, K.E.; Lane, W.S.; Duncan, K.; Walsh, C.T. Characterization of a Cys115 to Asp substitution in the Escherichia coli cell wall biosynthetic enzyme UDP-GlcNAc enolpyruvyl transferase (MurA) that confers resistance to inactivation by the antibiotic fosfomycin. Biochemistry 1996, 35, 4923–4928. [Google Scholar] [CrossRef] [PubMed]
- Herring, C.D.; Blattner, F.R. Conditional lethal amber mutations in essential Escherichia coli genes. J. Bacteriol. 2004, 186, 2673–2681. [Google Scholar] [CrossRef] [Green Version]
- Falagas, M.E.; Vouloumanou, E.K.; Samonis, G.; Vardakas, K.Z. Fosfomycin. Clin. Microbiol. Rev. 2016, 29, 321–347. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lucas, A.E.; Ito, R.; Mustapha, M.M.; McElheny, C.L.; Mettus, R.T.; Bowler, S.L.; Kantz, S.F.; Pacey, M.P.; Pasculle, A.W.; Cooper, V.S.; et al. Frequency and mechanisms of spontaneous fosfomycin nonsusceptibility observed upon disk diffusion testing of Escherichia coli. J. Clin. Microbiol. 2017, 56, e01368-17. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tseng, S.P.; Wang, S.F.; Kuo, C.Y.; Huang, J.W.; Hung, W.C.; Ke, G.M.; Lu, P.L. Characterization of fosfomycin resistant extended-spectrum β-lactamase-producing Escherichia coli isolates from human and pig in Taiwan. PLoS ONE 2015, 10, e0135864. [Google Scholar] [CrossRef] [PubMed]
- Wachino, J.; Yamane, K.; Suzuki, S.; Kimura, K.; Arakawa, Y. Prevalence of fosfomycin resistance among CTX-M-producing Escherichia coli clinical isolates in Japan and identification of novel plasmid-mediated fosfomycin-modifying enzymes. Antimicrob. Agents Chemother. 2010, 54, 3061–3064. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, Y.; Zheng, B.; Li, Y.; Zhu, S.; Xue, F.; Liu, J. Antimicrobial susceptibility and molecular mechanisms of fosfomycin resistance in clinical Escherichia coli isolates in Mainland China. PLoS ONE 2015, 10, e0135269. [Google Scholar] [CrossRef] [Green Version]
- Nilsson, A.I.; Berg, O.G.; Aspevall, O.; Kahlmeter, G.; Andersson, D.I. Biological costs and mechanisms of fosfomycin resistance in Escherichia coli. Antimicrob. Agents Chemother. 2003, 47, 2850–2858. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Karageorgopoulos, D.E.; Wang, R.; Yu, X.H.; Falagas, M.E. Fosfomycin: Evaluation of the published evidence on the emergence of antimicrobial resistance in Gram-negative pathogens. J. Antimicrob. Chemother. 2012, 67, 255–268. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vervoort, J.; Xavier, B.B.; Stewardson, A.; Coenen, S.; Godycki-Cwirko, M.; Adriaenssens, N.; Kowalczyk, A.; Lammens, C.; Harbarth, S.; Goossens, H.; et al. An in vitro deletion in ribE encoding lumazine synthase contributes to nitrofurantoin resistance in Escherichia coli. Antimicrob. Agents Chemother. 2014, 58, 7225–7233. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, X.; Zhang, Y.; Wang, F.; Wang, C.; Chen, L.; Liu, H.; Lu, H.; Wen, H.; Zhou, T. Unravelling mechanisms of nitrofurantoin resistance and epidemiological characteristics among Escherichia coli clinical isolates. Int. J. Antimicrob. Agents. 2018, 52, 226–232. [Google Scholar] [CrossRef]
- Whiteway, J.; Koziarz, P.; Veall, J.; Sandhu, N.; Kumar, P.; Hoecher, B.; Lambert, I.B. Oxygen-insensitive nitroreductases: Analysis of the roles of nfsA and nfsB in development of resistance to 5-nitrofuran derivatives in Escherichia coli. J. Bacteriol. 1998, 180, 5529–5539. [Google Scholar] [CrossRef] [Green Version]
- Ho, P.L.; Ng, K.Y.; Lo, W.U.; Law, P.Y.; Lai, E.L.; Wang, Y.; Chow, K.H. Plasmid-mediated oqxAB is an important mechanism for nitrofurantoin resistance in Escherichia coli. Antimicrob. Agents Chemother. 2015, 60, 537–543. [Google Scholar] [CrossRef] [Green Version]
- Campos, J.; Mourão, J.; Marçal, S.; Machado, J.; Novais, C.; Peixe, L.; Antunes, P. Clinical Salmonella Typhimurium ST34 with metal tolerance genes and an IncHI2 plasmid carrying oqxAB-aac(6′)-Ib-cr from Europe. J. Antimicrob. Chemother. 2016, 71, 843–845. [Google Scholar] [CrossRef] [Green Version]
- Dotto, G.; Giacomelli, M.; Grilli, G.; Ferrazzi, V.; Carattoli, A.; Fortini, D.; Piccirillo, A. High prevalence of oqxAB in Escherichia coli isolates from domestic and wild lagomorphs in Italy. Microb. Drug Resist. 2014, 20, 118–123. [Google Scholar] [CrossRef]
- Rodríguez-Martínez, J.M.; Díaz de Alba, P.; Briales, A.; Machuca, J.; Lossa, M.; Fernández-Cuenca, F.; Rodríguez Baño, J.; Martínez-Martínez, L.; Pascual, Á. Contribution of OqxAB efflux pumps to quinolone resistance in extended-spectrum-β-lactamase-producing Klebsiella pneumoniae. J. Antimicrob. Chemother. 2013, 68, 68–73. [Google Scholar] [CrossRef] [Green Version]

| Gene | Forward Primer | Reverse Primer | Amplicon Size (bp) | Reference |
|---|---|---|---|---|
| murA | 5′-AAACAGCAGACGGTCTATGG-3′ | 5′-CCATGAGTTTATCGACAGAACG-3′ | 1542 | [9] |
| glpT | 5′-GCGAGTCGCGAGTTTTCATTG-3′ | 5′-GGCAAATATCCACTGGCACC-3′ | 1785 | |
| uhpT | 5′-TTTTTGAACGCCCAGACACC-3′ | 5′-AGTCAGGGGCTATTTGATGG-3′ | 1667 | |
| uhpA | 5′-GATCGCGGTGTTTTTTCAG-3′ | 5′-GATACTCCACAGGCAAAACC-3′ | 771 | |
| ptsI | 5′-GAAAGCGGTTGAACATCTGG-3′ | 5′-TCCTTCTTGTCGTCGGAAAC-3′ | 1908 | |
| cyaA1 | 5′-AACCAGGCGCGAAAAGTGG-3′ | 5′-TGATGGCTGATGATCGACTC-3′ | 1559 | [9] This study |
| cyaA2 | 5′-AAAGCTCAGCCGTGAACGC-3′ | 5′-ACCTTCTGGGATTTGCTGG-3′ | 1648 | |
| nfsA | 5′-ATTTTCTCGGCCAGAAGTGC-3′ | 5′-AGAATTTCAACCAGGTGACC-3′ | 1036 | [5] |
| nfsB | 5′-CTTCGCGATCTGATCAACG-3′ | 5′-CAACAGCAGCCTATGATGAC-3′ | 923 | |
| ribE | 5′-AAGGGAAGCAGCGCACGAA-3′ | 5′-GGACAACTGCCAGGAGTAGA-3′ | 634 | This study |
| fosA3 | 5′-GCGTCAAGCCTGGCATTT-3′ | 5′-GCCGTCAGGGTCGAGAAA-3′ | 282 | [10] |
| fosA4 | 5′-CTGGCGTTTTATCAGCGGTT-3′ | 5′-CTTCGCTGCGGTTGTCTTT-3′ | 230 | [11] |
| fosA5 | 5′-TATTAGCGAAGCCGATTTTGCT-3′ | 5′-CCCCTTATACGGCTGCTCG-3′ | 177 | |
| fosA6 | 5′-GCTACGGTTCAGCTTCCAGA-3′ | 5′-CGAGCGTGGCGTTTTATCAG-3′ | 242 | This study |
| fosC2 | 5′-CGTTCCGTGGAGTTCTATAC-3′ | 5′-CTTGATAGGGTTTAGACTTC-3′ | 334 | [8] |
| oqxA | 5′-GACAGCGTCGCACAGAATG-3′ | 5′-GGAGACGAGGTTGGTATGGA-3′ | 339 | [12] |
| oqxB | 5′-CGAAGAAAGACCTCCCTACCC-3′ | 5′-CGCCGCCAATGAGATACA-3′ | 240 |
| Strain | Fosfomycin Disk 1 | Clinical Category 2 | Fosfomycin Disk Plus PPF 3 | G3P 4 | G6P 5 | Amino Acid Substitutions in | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| MurA | GlpT | UhpT | UhpA | PtsI | CyaA | ||||||
| 11 | 6 | R | 6 | + | − | None | Leu297Phe Glu443Gln Gln444Glu Glu448Lys | Not detected | Not detected | Arg367Lys | Asn142Ser Ala349Glu Ser352Thr Ser356Leu Gly359Glu Glu362Asp |
| 17 | 6 | R | 6 | + | − | None | Glu448Lys | Glu350Gln | Not detected | Val25Ile Arg367Lys | Asn142Ser Asp837Glu Thr840Ala |
| 26 | 12 | R | 12 | + | − a | None | Gly84Asp Glu448Lys | Not detected | None | Arg367Lys | Asn142Ser Ala349Glu Ser356Leu Gly359Glu Glu362Asp Ala363Ser Asp837Glu Thr840Ala |
| 66 | 6 | R | 13 | + | − | None | Glu448Lys | Glu350Gln | Not detected | Val25Ile Arg367Lys | Asn142Ser Asp837Glu Thr840Ala |
| 302 | 29 | S | 30 | + | + | None | Glu448Lys | None | None | Arg367Lys | Asn142Ser Gly222Ser |
| 334 | 31 | S | 32 | + | + | None | Glu448Lys | None | None | Arg367Lys | Asn142Ser |
| 381 | 6 | R | 6 | + | − | None | Glu448Lys | Glu350Gln | Not detected | Val25Ile Arg367Lys | Asn142Ser Asp837Glu Thr840Ala |
| 387 | 6 | R | 6 | + | − | None | Glu448Lys | Glu350Gln | Not detected | Val25Ile Arg367Lys | Asn142Ser Asp837Glu Thr840Ala |
| 462 | 12 | R | 13 | + | − | None | Glu448Lys | Glu350Gln | Not detected | Val25Ile Arg367Lys | Asn142Ser Asp837Glu Thr840Ala |
| 632 | 6 | R | 6 | + | − | None | Leu297Phe Glu443Gln Gln444Glu Glu448Lys | Glu350Gln | Not detected | Arg367Lys | Asn142Ser Ala349Glu Ser352Thr Ser356Leu Gly359Glu Glu362Asp Ala363Gly |
| 751 | 30 | S | 32 | + | + | None | Ala16Thr Glu448Lys | None | Arg46Cys | Ala306Thr Arg367Lys | Asn142Ser Ala349Glu Ser356Leu Gly359Glu Glu362Asp Asp837Glu Thr840Ala |
| 752 | 11 | R | 11 | + | − | None | Pro212Leu Glu448Lys | Glu350Gln | Not detected | Val25Ile Arg367Lys | Asn142Ser Asp837Glu Thr840Ala |
| 757 | 6 | R | 6 | + | − a | None | Leu297Phe Glu443Gln Gln444Glu Glu448Lys | None | Not detected | Arg367Lys | Asn142Ser Ala349Glu Ser352Thr Ser356Leu Gly359Glu Glu362Asp |
| 776 | 12 | R | 12 | + | − a | None | Glu448Lys | Glu350Gln | Not detected | Val25Ile Arg367Lys | Asn142Ser Asp837Glu Thr840Ala |
| 789 | 6 | R | 6 | + | − | Leu370Ile | Leu297Phe Glu443Gln Gln444Glu Glu448Lys | Glu350Gln | Not detected | Arg367Lys | Asn142Ser Ala349Glu Ser352Thr Ser356Leu Gly359Glu Glu362Asp |
| 792 | 6 | R | 6 | + | − | None | Glu448Lys | Glu350Gln | Not detected | Val25Ile Arg367Lys | Asn142Ser Asp837Glu Thr840Ala |
| 795 | 6 | R | 6 | + | − | None | Leu297Phe Glu443Gln Gln444Glu Glu448Lys | Glu350Gln | Not detected | Arg367Lys | Asn142Ser Ala349Glu Ser352Thr Ser356Leu Gly359Glu Glu362Asp |
| 797 | 20 | S | 21 | + | + | None | Ala16Thr Leu373Arg Glu448Lys | None | Arg46Cys | Ala306Thr Arg367Lys | Asn142Ser Ala349Glu Ser356Leu Gly359Glu Glu362Asp Asp837Glu Thr840Ala |
| 799 | 6 | R | 7 | + | − | None | Leu297Phe Glu443Gln Gln444Glu Glu448Lys | None | Not detected | Arg367Lys | Asn142Ser Ala349Glu Ser352Thr Ser356Leu Gly359Glu Glu362Asp |
| 802 | 30 | S | 30 | + | + | None | Phe133Cys Gly135Trp Ala197Val Glu448Lys | None | None | Arg367Lys | Asn142Ser |
| 809 | 6 | R | 6 | + | − | Leu370Ile | Leu297Phe Glu443Gln Gln444Glu Glu448Lys | Glu350Gln | Not detected | Arg367Lys | Asn142Ser Ala349Glu Ser352Thr Ser356Leu Gly359Glu Glu362Asp |
| 853 | 6 | R | 8 | + | − | Leu370Ile | Leu297Phe Glu443Gln Gln444Glu Glu448Lys | Glu350Gln | Not detected | Arg367Lys | Asn142Ser Ala349Glu Ser352Thr Ser356Leu Gly359Glu Glu362Asp |
| 854 | 6 | R | 6 | + | − | None | Glu448Lys | None | Not detected | Arg367Lys | Asn142Ser |
| 860 | 6 | R | 7 | + | − | None | Met52Leu Leu297Phe Glu443Gln Gln444Glu Glu448Lys | None | Not detected | Arg367Lys | Asn142Ser Ala349Glu Ser352Thr Ser356Leu Gly359Glu Glu362Asp |
| 871 | 6 | R | 14 | + | − a | None | Leu297Phe Glu443Gln Gln444Glu Glu448Lys | Glu350Gln | Not detected | Arg367Lys | Asn142Ser Ala349Glu Ser352Thr Ser356Leu Gly359Glu Glu362Asp Ala363Gly |
| 872 | 35 | S | 35 | + | + | None | Glu448Lys | None | None | Arg367Lys | Asn142Ser Asp837Glu Thr840Ala |
| 883 | 11 | R | 12 | + | − | None | Glu448Lys | Glu350Gln | Not detected | Val25Ile Arg367Lys | Asn142Ser Asp837Glu Thr840Ala |
| 891 | 11 | R | 11 | + | − | None | Glu448Lys | Glu350Gln | Not detected | Val25Ile Arg367Lys | Asn142Ser Asp837Glu Thr840Ala |
| 892 | 21 | S | 21 | + | + | None | Glu448Lys | None | None | Arg367Lys | Asn142Ser |
| Strain | Nitrofurantoin Disk 1 | Clinical Category 2 | Amino Acid Substitutions in | ||
|---|---|---|---|---|---|
| NfsA | NfsB | RibE | |||
| 11 | 11 | R | Ile117Thr Gly126Arg Lys141Glu Gln147Arg Gly187Asp | Truncated at Glu54 | None |
| 17 | 27 | S | Ile117Thr Lys141Glu Gly187Asp | Gly66Asp Val93Ala Ala174Glu | None |
| 26 | 24 | S | Ile117Thr Lys141Glu Gly187Asp | Gly66Asp Val93Ala Ala174Glu | None |
| 66 | 14 | R | Ile117Thr Lys141Glu Gly187Asp | Gly66Asp Val93Ala Ala174Glu Arg207His | None |
| 302 | 14 | R | None | Gly66Asp Met75Ile Val93Ala Ala174Glu Arg207His | None |
| 334 | 15 | I | Gln67Leu | Gly66Asp Met75Ile Val93Ala Arg107Cys | Pro55His |
| 381 | 25 | S | Ile117Thr Lys141Glu Gly187Asp | Gly66Asp Val93Ala Ala174Glu | None |
| 387 | 21 | S | Ile117Thr Lys141Glu Gly187Asp | Gly66Asp Val93Ala Ala174Glu | None |
| 462 | 14 | R | Cys80Arg Ile117Thr Lys141Glu Gly187Asp | Gly66Asp Val93Ala Ala174Glu Gly192Ser | None |
| 632 | 20 | S | Glu58Asp Ile117Thr Lys141Glu Gln147Arg Gly187Asp | Val93Ala | None |
| 751 | 14 | R | Ile117Thr Lys141Glu Gly187Asp Arg203Cys | Gly66Asp Val93Ala Ala174Glu | Val51Ile |
| 752 | 22 | S | Ile117Thr Lys141Glu Gly187Asp | Gly66Asp Val93Ala Ala174Glu | None |
| 757 | 16 | I | Glu58Asp Truncated at Gln67 | Val93Ala Lys122Arg | None |
| 776 | 19 | S | Ile117Thr Lys141Glu Gly187Asp | Gly66Asp Val93Ala Ala174Glu | None |
| 789 | 24 | S | Glu58Asp Ile117Thr Lys141Glu Gln147Arg Gly187Asp | Val93Ala | None |
| 792 | 25 | S | Ile117Thr Lys141Glu Gly187Asp | Gly66Asp Val93Ala Ala174Glu | None |
| 795 | 16 | I | His11Tyr Glu58Asp Ile117Thr Lys141Glu Gln147Arg Gly187Asp | Val93Ala | None |
| 797 | 12 | R | Ile117Thr Lys141Glu Gly154Glu Gly187Asp | Leu22Ile Gly66Asp Val93Ala Ala174Glu | Val51Ile |
| 799 | 13 | R | Ile117Thr Lys141Glu Gln147Arg Gly187Asp | Val93Ala | None |
| 802 | 12 | R | Truncated at Gln147 | Met75Ile Val93Ala | None |
| 809 | 28 | S | Glu58Asp Ile117Thr Lys141Glu Gln147Arg Gly187Asp | Val93Ala | None |
| 853 | 20 | S | Glu58Asp Ile117Thr Lys141Glu Gln147Arg Gly187Asp | Val93Ala | None |
| 854 | 16 | I | None | Gly66Asp Met75Ile Val93Ala | None |
| 860 | 20 | S | Ile117Thr Lys141Glu Gln147Arg Gly187Asp | Val93Ala | None |
| 871 | 24 | S | Glu58Asp Ile117Thr Lys141Glu Gln147Arg Gly187Asp | Val93Ala | None |
| 872 | 15 | I | Asp19Asn Ser33Arg Ile117Thr Lys141Glu Gly187Asp | Gln44His Gly66Asp Val93Ala Ala174Glu | None |
| 883 | 13 | R | Ile117Thr Lys141Glu Gly187Asp | Gly66Asp Phe84Ser Val93Ala Ala174Glu | None |
| 891 | 23 | S | Ile117Thr Lys141Glu Gly187Asp | Gly66Asp Val93Ala Ala174Glu | None |
| 892 | 10 | R | None | Gly66Asp Met75Ile Val93Ala | None |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sorlozano-Puerto, A.; Lopez-Machado, I.; Albertuz-Crespo, M.; Martinez-Gonzalez, L.J.; Gutierrez-Fernandez, J. Characterization of Fosfomycin and Nitrofurantoin Resistance Mechanisms in Escherichia coli Isolated in Clinical Urine Samples. Antibiotics 2020, 9, 534. https://doi.org/10.3390/antibiotics9090534
Sorlozano-Puerto A, Lopez-Machado I, Albertuz-Crespo M, Martinez-Gonzalez LJ, Gutierrez-Fernandez J. Characterization of Fosfomycin and Nitrofurantoin Resistance Mechanisms in Escherichia coli Isolated in Clinical Urine Samples. Antibiotics. 2020; 9(9):534. https://doi.org/10.3390/antibiotics9090534
Chicago/Turabian StyleSorlozano-Puerto, Antonio, Isaac Lopez-Machado, Maria Albertuz-Crespo, Luis Javier Martinez-Gonzalez, and Jose Gutierrez-Fernandez. 2020. "Characterization of Fosfomycin and Nitrofurantoin Resistance Mechanisms in Escherichia coli Isolated in Clinical Urine Samples" Antibiotics 9, no. 9: 534. https://doi.org/10.3390/antibiotics9090534
APA StyleSorlozano-Puerto, A., Lopez-Machado, I., Albertuz-Crespo, M., Martinez-Gonzalez, L. J., & Gutierrez-Fernandez, J. (2020). Characterization of Fosfomycin and Nitrofurantoin Resistance Mechanisms in Escherichia coli Isolated in Clinical Urine Samples. Antibiotics, 9(9), 534. https://doi.org/10.3390/antibiotics9090534







