Abstract
Oceans cover seventy percent of the planet’s surface and besides being an immense reservoir of biological life, they serve as vital sources for human sustenance, tourism, transport and commerce. Yet, it is estimated by the National Oceanic and Atmospheric Administration (NOAA) that eighty percent of the oceans remain unexplored. The untapped biological resources present in oceans may be fundamental in solving several of the world’s public health crises of the 21st century, which span from the rise of antibiotic resistance in bacteria, pathogenic fungi and parasites, to the rise of cancer incidence and viral infection outbreaks. In this review, health risks as well as how marine bacterial derived natural products may be tools to fight them will be discussed. Moreover, an overview will be made of the research pipeline of novel molecules, from identification of bioactive bacterial crude extracts to the isolation and chemical characterization of the molecules within the framework of the One Health approach. This review highlights information that has been published since 2014, showing the current relevance of marine bacteria for the discovery of novel natural products.
1. Health Challenges Faced Today
1.1. The Health and Economic Burden of Antibiotic Resistance
With the increase of global travel and the incorrect and excessive use of antibiotics, namely for animal production, a drastic rise in antibiotic resistance is being observed in bacterial populations throughout the world [1]. Infections with resistant organisms not only entail higher morbidity and mortality, but are also more expensive to treat and result in longer hospital stays, which places a greater burden on healthcare systems [2]. These facts make the rise in antibiotic resistance one of the greatest health challenges of the century. Based on World Health Organization (WHO) reports on the global rise of resistance in pathogenic bacteria, the Global Antimicrobial Resistance Surveillance System (GLASS) report [3,4] forecasts an alarming future for humankind. Patients with third-generation cephalosporin-resistant Escherichia coli infections have experienced a two-fold increase in mortality attributable to bacterial infections and a significant mortality increase is also observed in patients with methicillin-resistant Staphylococcus aureus infections. Facts like these point to the imperative need for the discovery of novel antibiotics to fight these super bacteria. Moreover, the global priority list of antibiotic-resistant bacteria to guide research, discovery, and development of new antibiotics also released by the WHO, specifies the top priorities for research and development of novel antibiotics. Critical bacteria comprise carbapenem-resistant Enterobacteriaceae (e.g., Klebsiella pneumoniae, E. coli, Enterobacter sp., Serratia sp., Proteus sp., Providencia sp. and Morganella sp.) and Acinetobacter baumannii, Pseudomonas aeruginosa, as well as 3rd generation cephalosporin-resistant Enterobacteriaceae. High priority bacteria comprise vancomycin-resistant Enterococcus faecium, methicillin-resistant and/or vancomycin-resistant S. aureus, clarithromycin-resistant Helicobacter pylori, fluoroquinolone-resistant Campylobacter sp. and fluoroquinolone-resistant Salmonella sp. and Neisseria gonorrhoeae, as well as 3rd generation cephalosporin-resistant N. gonorrhoeae. Medium priority bacteria include fluoroquinolone-resistant Shigella sp., ampicillin-resistant Haemophilus influenzae and penicillin-non-susceptible Streptococcus pneumoniae [5].
Regarding fungi, infections by resistant Candida spp. and Aspergillus spp. are increasing. The Centers for Disease Control and Prevention (CDC) recently published data showing that 7% of all bloodstream Candida isolates are resistant to fluconazole [6]. Moreover, the CDC also reports that about 90% of Candida auris isolates are resistant to fluconazole and 33% to amphotericin B [7]. Aspergillus infections, although only common in immunosuppressed individuals, are also becoming harder to fight, as azole resistance is also being observed [8,9,10].
In addition, the spread of antimicrobial resistance entails a vast economic loss. It is estimated that the economic impact of these diseases will reach up to US$3.5 billion in losses per year due to deaths caused by resistant microorganisms in the next 30 years just in Europe, North America and Australia [11].
Likewise, there is a significant rise of resistance in protozoal parasites, belonging to the genera Plasmodium, Trypanosoma and Leishmania [12,13,14]. These parasites are responsible for malaria, African and American trypanosomiasis and leishmaniasis, respectively, which disproportionally affect the poorest and least prepared nations. This is aggravated by the fact that few drugs are effective to combat these diseases. For example, in the 1960s, Plasmodium developed resistance to chloroquine, a first line treatment drug against malaria [15] and only a few families of drugs remain capable of treating malaria. However, in Southeast Asia, the infectious agents of malaria already show signs of resistance to artemisinin [16], which was discovered in 1975 and is currently the main treatment for malaria [17].
1.2. Impacts of Viral Infectious Outbreaks
Viruses are infectious agents that can only reproduce inside living cells. In their most basic form, they consist of only genetic material, such as DNA or RNA (retroviruses) and a protein coat, the capsid. This basic plan can then be supplemented with other components, like an outer envelope of lipid coating or, for retroviruses, the reverse transcriptase enzyme [18].
Viruses have a great capacity to spread causing viral infections in humans. Due to horizontal transmission, the most common mechanism of virus spreading in populations, we have witnessed several viral pandemics that have led to health and economic crises. Since December 2019, the world is in the grips of another pandemic outbreak, caused by the SARS-CoV-2 virus, which has already caused thousands of deaths and disrupted the global economy [19]. However, this is just the most recent of several pandemic and epidemic outbreaks caused by viruses, like those of Zika in 2015 [20], Ebola which flared up in West Africa in 2013 [21], the 2009 influenza pandemic caused by H1N1/09 and the severe acute respiratory syndrome (SARS), caused by SARS-CoV-1 in 2002. This last mentioned epidemic, like SARS-CoV-2, affected China, causing 774 deaths and 100 billion dollars in losses [22]. Moreover, humanity is still struggling with past pandemics, such as the HIV/AIDS pandemic, which still infects 37 million people around the world [23]. Furthermore, as novel viral diseases keep appearing, and antivirals are usually specifically targeted, research of new therapeutics is continuously needed [24].
1.3. “One Health” Framework
Many of these novel microbial threats are zoonotic in nature [25]. Zoonotic diseases are found to be spreading due to pressures on the environment like, for example, encroachment of human populations on animal territory. Moreover, animals are known reservoirs of endemic diseases also [26,27]. The “One Health” framework thus looks for combat diseases by a three-pronged strategy of communication, coordination and collaboration between human, animal, environmental health, and other relevant experts [28,29]. This integrated, interdisciplinary work is vital to understand and tackle emergent and re-emergent threats of infectious diseases, as it looks for environmental clues for discerning infectious threats and generating plans for confronting these threats.
1.4. Cancer Incidence and Mortality
As one of the most important public health problems, cancer is currently the second most frequent cause of death worldwide, being responsible for an estimated 9.6 million deaths in 2018 [30]. Between 2005 and 2015, it was the cause of one-third of all deaths worldwide [31]. For 2018, the GLOBOCAN database detailed that the major cancer types are of the lung, female breast and colorectum, which account for one-third of the cancer incidence and mortality [32].
Cancer is the generic name for a collection of more than 100 diseases due to uncontrolled division of cells that have developed over time. In cancer, the tightly controlled processes of cell division are deregulated by either the activation of proto-oncogenes or the inactivation of tumour suppressor genes. However, in order for a cell to develop into a cancerous tumour, other processes, such as apoptosis and senescence, cannot occur in the tumour cells. Cells must also have access to oxygen and nutrients [33]. Genetic factors can play an important part in the appearance of cancerous formations. This is the case of breast cancer and the tumour suppressing gene, BRCA1, as carriers are predisposed to life-time risks of up to 80% [34]. Furthermore, environmental factors, such as tobacco smoking, increase in urban pollution, increased intake of high caloric foods and longer lifespans all contribute to increasing the risk of cancer diagnosis [35]. Cancer is treated through the management of the risk factors and the use of antitumoral chemotherapeutic agents, surgery, radiotherapy or immunotherapy.
Antitumoral chemotherapeutics of microbial origin, many of which are produced by bacteria of the genus Streptomyces, are widely used in clinical practice and comprise mostly cytotoxic compounds that include the anthracycline daunomycin and its derivative doxorubicin, the polyketides aclarubicin and mithramycin, the glycopeptidic bleomycins A2 and B2, the chromopeptide actinomycin D and mitomycin C [36]. Nevertheless, the discovery of novel anticancer drugs remains a highly challenging endeavour. Not only must molecules have a specific effect, usually inhibitory which leads to cytotoxic effect on a target (or multiple targets), but toxic side effects need to be manageable [37]. This implies that many molecules must be extensively screened for side effects before their possible application. Natural products can reliably increase the diversity of bioactive molecular structures and, thus, increase the likelihood of developing novel therapeutics.
After pointing out the fundamental health risks faced by humankind, this review explores the contribution of marine bacterial derived natural products in the fight against cancer, antimicrobial resistance and viruses, with an update highlighting information from the literature published since 2014. Moreover, the technicalities needed for discoveries will be detailed.
2. Natural Products
2.1. The History of Marine Natural Products
Nature has long been the most important source of therapeutics. The use of poultices and mixtures of plant material to treat infections goes back to the early bronze age civilizations [38]. Building on the prior knowledge acquired, early medicine, pharmacology and chemistry started to develop therapeutics by studying nature. For example, extracts of willow bark (genus Salix), containing salicylic acid, which was identified to be the bioactive molecule present in the bark in 1828, were already used by Sumerians and Egyptians to treat inflammation and pain. In 1852 acetylsalicylic acid was first synthesised and in 1899, Bayer patented it as aspirin [39]. Ever since then, many terrestrial organisms and in particular plants have been sought for use as natural products. It was only around the 1970s, that attention was first given to the ocean as a source of useful natural products. As the oceans cover most of the Earth’s surface, they are home to a substantial portion of the world’s biodiversity [40] which lives in distinctive and varied conditions and has evolved through a long period of metabolic adaptations. When exploration of the ocean’s biodiversity and metabolic richness began, it resulted in the discovery of thousands of structurally unique bioactive marine natural products [41].
Initially, the exploitation of marine wildlife for natural bioactive products focused on a small number of organisms which included sponges, molluscs, tunicates and macroalgae [42]. These were shown to produce a very diverse range of unique molecular structures, like halogenated terpenes, polyketides and prostaglandins [43,44,45,46]. This diversity of bioactive structures is considered to be part of the defence, survival and predatorial strategies employed by these organisms, such as, for example, sponges, which are sessile, soft-bodied organisms, lacking morphological defences like biological armature or spines [47]. Thus, these organisms appeared to be a great source for the discovery of novel bioactive molecules. However, the bioactive molecules produced by these organisms can be present in quite small amounts. For example, halichondrin B (1) (Figure 1), a macrolide first isolated from Halichondria okadai that has potent anticancer activity, impeding mitotic division by targeting tubulin [48], is present in concentrations as low as 400 µg per kg wet weight of tissue of Lissodendoryx sp. [49]. Due to their low concentrations of bioactive molecules the use of these organisms poses environmental problems, because high quantities of organisms would be needed to produce enough molecules to even begin preclinical trials [50]. Yet, its structure inspired a synthetic analogue, eribulin mesylate, which is now used in breast cancer and lipocarcinoma treatment [51]. As such, sponges, molluscs, tunicates and macroalgae still remain relevant sources of new marine natural products [52].
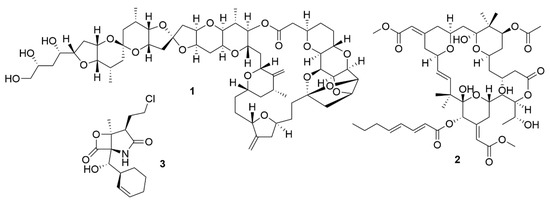
Figure 1.
Bioactive metabolites isolated from marine organisms. Halichondrin B (1), a macrolide first isolated from H. okadai but also present in other sponges. Bryostatin 1 (2), which belongs to a family of polyketide macrolides first identified in the marine bryozoan B. neritina. Salinosporamide A (3), is a proteasome inhibitor isolated from bacteria from the genus Salinospora and is in phase III clinical trials for the treatment of multiple myeloma.
The exploitation of other sources of bioactive marine organisms, mainly microorganisms, has also led to the discovery of new promising leads. Indeed, some of the molecules associated with macroorganisms such as sponges, may have their origin in associated microorganisms [53]. This may be the case of bryostatins, found in the marine bryozoan Bugula neritina [54]. The bryostatins, exemplified by bryostatin 1 (2) are polyketide macrolactones with neurological and anticancer properties that work by modulating the activity of the protein kinase C family and they were first isolated from this bryozoan in 1982 [55]. Yet, these macrolactones appear to be synthesised by a group of PKS genes of bacterial origin, indicating that bryostatins are produced by a bacterium [56].
Marine microorganisms, like members of Actinobacteria, Proteobacteria, Firmicutes, Cyanobacteria, fungi and dinoflagellates, have shown to be great reservoirs of bioactive molecules [52]. Yet, even though initial predictions pointed to an immediate increase in the number of natural products discovered with the shift in focus from macroorganisms to microorganisms, this did not occur [57]. However, advances in the isolation of novel taxa, provided a boost in the discovery of novel bioactive molecules. Analysis of the literature reveals an increase in the number of new molecules discovered in all microorganisms from 2014 to 2018 [52], with a special emphasis on fungi and bacteria. Members of the genus Salinospora (Actinobacteria) are examples of bacteria that lead to the discovery of salinosporamide A (3) [58]. This molecule has a potent cytotoxic activity due to a unique functionalisation of the core-fused γ-lactam-β-lactone bicyclic ring, which contributes significantly to its activity [59]. Salinosporamide A is now in phase III clinical trials for the treatment of multiple myeloma under the brand name Marizomib [60].
2.2. Marine Bacterial Natural Products
Although salinosporamide A is the best example of the potential present in bacteria, many novel molecular structures are discovered each year. In fact, in 2016, 179 new natural products of marine bacterial origin were discovered [61], in 2017, the number rose to 242 [62] and in 2018 a total of 240 new molecules were reported [52]. The upwards trend seen in the number of discovered molecules and the remarkable chemical diversity displayed, which ranges from peptides, siderophores and polyketides to esters, macrolactones, quinones and terpenes, shows the bacterial potential for the discovery of novel active principles.
2.2.1. Antimicrobial Marine Bacterial Natural Products
Bacteria are promising sources for novel antimicrobial natural product discovery. This is primarily due to two factors. One is their variable and malleable metabolism [63] and the other is their competitive pressure for resources against other microbes. Several recent examples of natural products from marine bacteria are provided below. The novel molecule bacicyclin (4) (Figure 2), which is a cyclic peptide, was isolated from a Bacillus sp. strain BC028 isolated from the common mussel (Mytilus edulis) [64]. It displays antibacterial activity against E. faecalis and S. aureus with minimal inhibitory concentration (MIC) values of 8 and 12 µM, respectively, and can help in the design of analogues with increased antibiotic efficacy [64]. Anthracimycin B (5), a polyketide with powerful anti-Gram-positive bacteria activity that was obtained from a marine-derived Streptomyces cyaneofuscatus M-169, has expanded the knowledge of how the methyl group at C-2 of anthracimycins plays a role in its antibacterial effect [65]. Taromycin B (6), a lipodepsipeptide with potent activity against methicillin-resistant S. aureus and vancomycin-resistant E. faecium which was isolated from the marine actinomycete Saccharomonospora sp. CNQ-490, provides a promising start for the development of novel antibacterial scaffoldings [66]. Janthinopolyenemycin A (7) and B (8) are also polyketides and the first examples of molecules of their structural type. Janthinopolyenemycins were isolated from the proteobacterium Janthinobacterium spp., strains ZZ145 and ZZ148, and have activity against Candida albicans [67]. Streptoseomycin (9), a macrolactone isolated from the actinobacterium Streptomyces seoulensis A01, has specific activity against microaerophilic bacteria, specially the pathogen H. pylori [68]. This restricted activity makes streptoseomycin a good starting point for the discovery of antibiotics for the treatment of H. pylori infections. The polyketides ansalactams B (10), C (11) and D (12) are highly modified ansamycins that show weak and mild anti-methicillin-resistant S. aureus and were identified in cultures of Streptomyces sp. CNH189, isolated from marine sediments. Ansalactams B and D are cyclic polyketides with similarities to ansalactam A. However, ansalactam D shows evidence of an uncommon oxetane ring. Ansalactam C is an open polyketide chain resulting from a Baeyer–Villiger-type oxidation [69]. Micromonohalimanes A (13) and B (14) are rare halimane-type diterpenoids isolated from the actinobacterium Micromonospora sp. WMMC-218. Micromonohalimane A displays a very weak inhibitory effect on methicillin-resistant S. aureus while micromonohalimane B displays moderate bacteriostatic activity against it. Xestostreptin (15) is a modified diketopiperazine isolated from Streptomyces sp. S.4, resulting from the condensation of the aminoacids threonine and alanine [70]. Xestostreptin shows weak activity against the malarial agent P. falciparum. Two macrolides, branimycins B (16) and C (17) were identified from a fermentation of the actinobacterium Pseudonocardia carboxydivorans M-227, isolated from deep sea water [71]. Branimycin B shows moderate antibacterial activity against Gram-positive bacteria, while branimycin C displayed moderate antibacterial activity against Gram-negative bacteria.
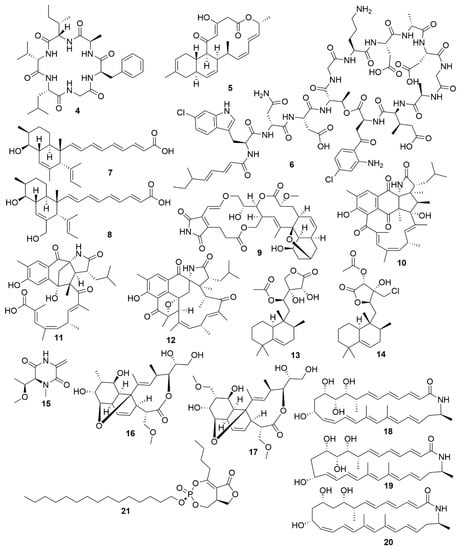
Figure 2.
Examples of recently isolated antimicrobial natural products from marine bacteria. Bacicyclin (4), a cyclic peptide isolated from a Bacillus sp. BC028. Anthramicin B (5), a polyketide isolated from S. cyaneofuscatus M-169. Taromycin B (6), a lipopeptide from Saccharomonospora sp. CNQ-490. Janthinopolyenemycin A (7) and Janthinopolyenemycin B (8). The janthinopolyenemycins are polyketides isolated from two strains of the genus Janthinobacterium. Streptoseomycin (9), a macrolactone isolated from S. seoulensis. Ansalactam B (10), a pentacyclic polyketide. Ansalactam C (11), an open polyketide unlike ansalactam B. Ansalactam D (12), a hexacyclic polyketide. Ansalactams B, C and D were isolated from Streptomyces sp. CNH189. Micromonohalimane A (13). Micromonohalimane B (14). Micromonohalimanes A and B are terpenes isolated from Micromonospora sp. WMMC-218. Xestostreptin (15), a diketopiperazine isolated from Streptomyces sp. S.4. Branimycin B (16) and Branimycin C (17) C are macrolides isolated from the deep-sea bacterium P. carboxydivorans M-227. Lobosamide A (18), Lobosamide B (19) and Lobosamide C (20) are macrolactams isolated from Micromonospora sp. RL09-050-HVF-A. Salinipostin A (21), a bicyclic phosphotriester isolated from Salinispora sp. RLUS08-036-SPS-B.
Utilizing a genome-assisted discovery strategy, three macrolactams, lobosamides A (18), B (19) and C (20), were isolated from Micromonospora sp. RL09-050-HVF-A [72]. Lobosamides A and B showed bioactivity against the microbial agent of African trypanosomiasis, Trypanosoma brucei in low concentrations. However, lobosamide C was not bioactive. Schulze and colleagues [73] also identified salinipostins A-K bicyclic phosphotriesters isolated from the actinobacterium Salinispora sp. RLUS08-036-SPS-B with potent and selective activity against P. falciparum. The salinipostin scaffold considerably differs from any of the known antimalarial compounds, representing a novel lead structure in the development of therapeutics for malaria. Experiments with salinipostin A (21), the most bioactive of the 11 salinipostins, indicate that it exhibits growth stage-specific effects and no apparent resistance could be identified in parasite populations. A hybrid peptide-polyketide, mollemycin A (22) (Figure 3) was isolated from the marine bacterium Streptomyces sp. CMB-M0244 and shows potent antimalarial and broad antibacterial activities [74]. Actinosporin A (23) is a glycosilated polyketide which shows antiparasitic activity against T. brucei and was isolated from a marine sponge associated Actinokineospora sp. EG49 [75]. Likewise, actinosporin B, was also isolated but showed no bioactivity, suggesting that actinosporin A is acting selectively against the parasite. The linear lipopetides, gageopeptides A-D (24–27), gageotetrins A–C (28–30) and gageostatins A–C (31–33) were isolated from the marine Bacillus subtilis 109GGC020. These lipopetides showed a range of different antimicrobial bioactivities, with gageostatins A, B and C all showing good antimicrobial activity and moderate cytotoxic activity to lung cancer cell line NCI-H23 [76]. Gageotetrins A, B and C showed potent antimicrobial bioactivities but not cytotoxic effect on human myeloid leukaemia K-562 [77]. Furthermore, gageopeptides A, B, C and D all showed good antifungal and moderate broad antibacterial activity, while not showing cytotoxicity to human myeloid leukaemia K-562 and mouse leukemic macrophage RAW 264.7 cell lines [78].
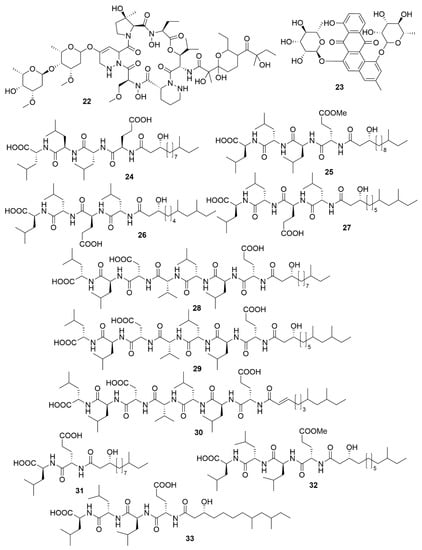
Figure 3.
Examples of recently isolated antimicrobial natural products from marine bacteria. Mollemycin A (22) is a hybrid peptide-polyketide isolated from Streptomyces sp. CMB-M0244. Actinosporin A (23), a polyketide isolated from Actinokineospora sp. EG49. Gageopeptide A (24). Gageopeptide B (25). Gageopeptide C (26), Gageopeptide D (27), Gageotetrin A (28). Gageotetrin B (29). Gageotetrin C (30). Gageostatin A (31). Gageostatin B (32). Gageostatin C (33). The gageopeptides, gageotetrins and gageostatins are linear lipopeptides isolated from B. subtilis 109GGC020.
2.2.2. Antiviral Marine Bacterial Natural Products
It is estimated that as many as 1031 viruses inhabit the oceans [79], with concentrations ranging from 3 × 106 viruses mL−1 in deep sea waters to 108 viruses mL−1 in coastal waters [80], many of which are bacteriophages. Consequently, marine bacteria are subjected to evolutionary pressure to develop defences against viral attacks. As such, marine bacteria may be great reservoirs of antiviral leads.
A number of examples of marine bacterial natural products with antiviral bioactivities have been recently reported. Three novel abyssomicin monomers, neoabyssomicins D (34) (Figure 4), E and A2 and two dimers—neoabyssomicins F (35) and G (36)—were isolated from the marine Streptomyces koyangensis SCSIO 5802, with neoabyssomicin D showing moderate anti-herpes simplex virus activity, and neoabyssomicins F and G showing low activity against vesicular stomatitis virus [81]. Streptomyces sp. OUCMDZ-3434, isolated from the marine alga Enteromorpha prolifera, was shown to produce five new phenolic polyketides [82]. Of these molecules, wailupemycin J (37) and (R)-wailupemycin K (38) proved to be bioactive against the influenza A virus (H1N1). The indolosesquiterpenoids xiamycins C (39), D (40) and E (41) were isolated from the marine-derived Streptomyces sp. #HK18, and showed strong inhibitory effect against the coronavirus porcine epidemic diarrhoea virus [83]. As such, xiamycins may provide useful leads in the development of antivirals with broader spectrum activity against other coronaviruses.
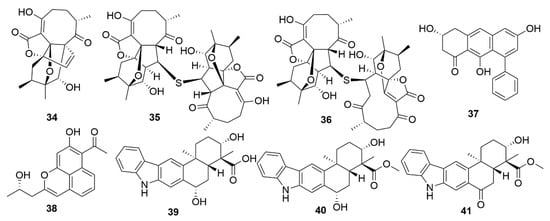
Figure 4.
Examples of recently isolated natural products from marine bacteria with antiviral activity. Neoabyssomicin D (34). Neoabyssomicin F (35) and Neoabyssomicin E (36) are polycyclic polyketides isolated from S. koyangensis SCSIO 5802. Wailupemycin J (37) and (R)-wailupemycin K (38) are phenolic polyketides isolated from Streptomyces sp. OUCMDZ-3434. Xiamycin C (39), Xiamycin D (40) and Xiamycin E (41) were isolated from Streptomyces sp. #HK18.
2.2.3. Anticancer Marine Bacterial Natural Products
As with antimicrobials, it is ascertained that marine bacteria are great reservoirs for cytotoxic natural products. While salinosporamide A, already mentioned above, is a great example of marine cytotoxic drug discovery [58], every year, novel anticancer bioactive structures are discovered. Actinobacteria, especially those of the genera Streptomyces and Micromonospora, have been a very prolific source of cytotoxic compounds. There are several recent examples of structures isolated from these bacteria. Dentigerumycin E (42) (Figure 5), is a cyclic hexapeptide bearing three piperazic acids and a pyran-bearing polyketide acyl chain, isolated from the marine actinoabcterium Streptomyces albogriseolus JB5 [84]. It showed moderate cytotoxicity against lung carcinoma A549, colorectal cancer HCT116, breast cancer MDA-MB-231, liver cancer SK-HEP-1 and stomach cancer SNU638 cell lines, while not being cytotoxic to the normal human breast epithelial cell line MCF-10A [84]. Likewise, neothioviridamide (43) is a polythioamide cyclic peptide with strong cytotoxicity against human ovarian adenocarcinoma (SKOV-3), malignant pleural mesothelioma (Meso-1) and immortalized human T lymphocyte (Jurkat) cell lines [85]. It was isolated after discovery of a novel biosynthetic cluster (thioviridamide-like biosynthetic gene) in Streptomyces sp. MSB090213SC12 by genome mining and heterologous expression of a bacterial artificial chromosome in Streptomyces avermitilis SUKA. Three cyclic depsipeptides, rakicidins G-I (44–46), isolated from the marine actinobacterium Micromonospora chalcea FIM 02–523, have potent cytotoxic activity against the human pancreatic cancer cell line PANC-1 and human colon carcinoma cell line HCT-8 [86]. Rakicidins G-I differ in the length of their β-hydroxy fatty acid moiety. The 26-membered polyene macrolactam, FW05328-1 (47), isolated from Micromonospora sp. FIM05328, has potent bioactivity against three cells lines of human oesophageal squamous cell carcinoma (KYSE30, KYSE180 and EC109) [86]. The integration of genomic data in association with nuclear magnetic resonance (NMR) analysis allowed the determination of the stereostructure of neaumycin B, a cytovaricin-ossamycin-oligomycin macrolide, that was isolated from Micromonospora sp. CNY-010 [87]. Neaumycin B (48) has shown potent anti-human glioblastoma cell line U87 activity but it is unstable. Through genetic manipulation of promoters, six new polyketides, pactamides A–F, were isolated from Streptomyces pactum SCSIO 02999. Pactamides B-F showed low to moderate cytotoxic activity against human glioblastoma cell line (SF-268), human breast cancer cell (MCF-7), human large-cell lung carcinoma (NCI-H460) and human liver cancer cell (HepG2) while pactamide A (49) showed potent activity [88]. Two cyclodepsipeptides, streptodepsipeptides P11A (50) and P11B (51), were isolated from Streptomyces sp. P11-23B and displayed potent anti-proliferative bioactivity in four cell lines of human glioblastoma (U251, U87-MG, SHG-44 and C6) [89]. Research with Streptomyces sp. strain THS-55 yielded four new antimycin alkaloids, antimycins E-H (52–55), which showed potent cytotoxic effect on HPV-transformed human cervix adenocarcinoma (HeLa) cells and moderate anti-proliferative activity in human cervical cancer cell SiHa, human myelogenous leukaemia cell line K562, and human leukaemia cell line HL-60 [90]. However, cytotoxic effects were shown in healthy human embryonic kidney cells 293T. In a knockout mutant of Streptomyces sp. CHQ-64, two new alkaloids, geranylpyrrol A and piericidin F (56), were discovered [91]. Of these, piericidin F showed potent anti-proliferative activity against several cancer cell lines, including HeLa, human acute promyelocytic leukaemia (NB4) and human lung carcinoma (A549 and H1975).
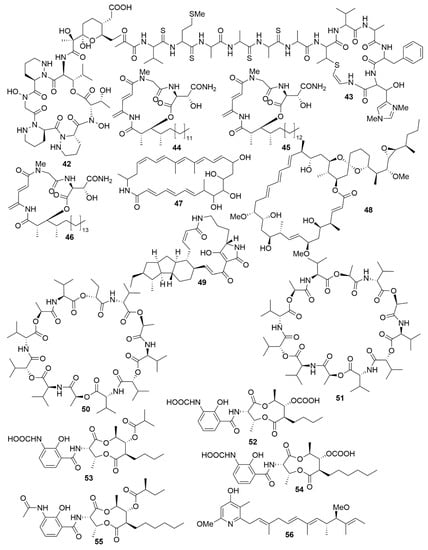
Figure 5.
Examples of recently isolated cytotoxic natural products from marine bacteria. Dentigerumycin E (42), a cyclic hexapeptide isolated from S. albogriseolus JB5. Neothioviridamide (43), a cyclic peptide from Streptomyces sp. MSB090213SC12. Rakicidin G (44), Rakicidin H (45) and Rakicidin I (46) are cyclic depsipeptides isolated from M. chalcea FIM 02–523. FW05328-1 (47), a polyene macrolactam isolated from Micromonospora sp FIM05328. Neaumycin B (48), a macrolide from Micromonospora sp. CNY-010. Pactamide A (49), a polyketide isolated from S. pactum SCSIO 02999. Streptodepsipeptide P11A (50) and Streptodepsipeptide P11B (51) are cyclodepsipeptides from Streptomyces sp. P11-23B. Antimycin E (52), Antimycin F (53), Antimycin G (54) and Antimycin H (55) are alkaloids isolated from Streptomyces sp. THS-55. Piericidin F (56) is an alkaloid isolated Streptomyces sp. CHQ-64.
Two new chromodepsipeptides, neo-actinomycin A (57) (Figure 6) and neo-actinomycin B (58) were isolated from a marine-derived Streptomyces sp. IMB094 [92]. They showed strong cytotoxic effect on human colorectal carcinoma cell line HCT116 and human lung carcinoma cell line A549.
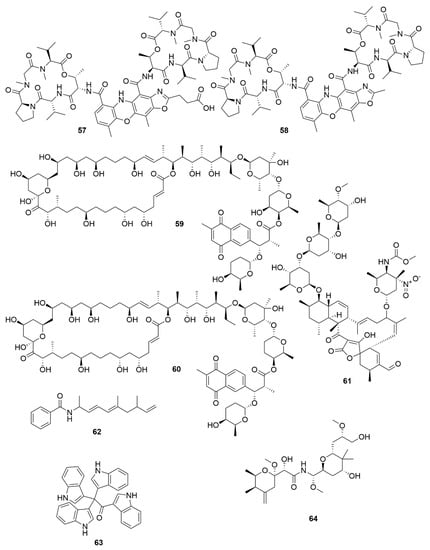
Figure 6.
Examples of recently isolated cytotoxic natural products from marine bacteria. Neo-actinomycin A (57) and Neo-actinomycin B (58) are chromopeptides from Streptomyces sp. IMB094. PM100117 (59) and PM100118 (60) are macrolides isolated from S. caniferus GUA-06-05-006A. Lobophorin I (61), a spirotetronate isolated from Streptomyces sp. 1053U.I.1a.3b. Haliamide (62), a hybrid of a polyketide synthase from H. ochraceum SMP-2. Tetra(indol-3-yl)ethenone (63), an indole isolated from P. denitrificans BBCC725. O-Demethylpederin (64), a polyketide with a tetrahydropyran-core from Labrenzia sp. PHM005.
Two new macrolides, PM100117 (59) and PM100118 (60) were isolated from a marine Streptomyces caniferus GUA-06-05-006A [93]. Both PM100117 and PM100118 show potent cytotoxic effect on human breast adenocarcinoma (MDA-MB-231), human lung carcinoma (A549) and human colorectal carcinoma (HT-29) cell lines. The study of a symbiotic Streptomyces sp. (strain 1053U.I.1a.3b) of cone snails lead to the isolation of two lobophorins, H and I [94]. Lobophorins are a large family of spirotetronates with antimicrobial and cytotoxic bioactivities. Of these, lobophorin I (61) showed potent cytotoxic activity against human T-cell leukaemia cell line CEM-TART.
Besides Actinobacteria, other marine bacterial phyla are also proving to be relevant for the isolation of novel bioactive molecules. As a result of a hybrid polyketide synthase (PKS) and non-ribosomal peptide synthetase (NRPS) biosynthesis, haliamide (62) was isolated from a marine myxobacterium, Haliangium ochraceum SMP-2, and shows moderate cytotoxicity towards the HeLa cell line [95]. A novel cytotoxic indole, tetra(indol-3-yl)ethenone (63) was isolated from the marine proteobacterium Pseudovibrio denitrificans BBCC725 [96]. Tetra(indol-3-yl)ethenone has moderate cytotoxicity to human lung carcinoma cell line A549 and the mouse fibroblasts cell line L929. Another marine proteobacterium, Labrenzia sp. PHM005 produced a new tetrahydropyran-core polyketide and analogue of pederin [97]. This novel pederin, 18-O-demethylpederin (64) shows potent anti-proliferative activity against four cell lines: human lung carcinoma cell line A549, human colon adenocarcinoma cell line HT-29, human breast adenocarcinoma cell line MDA-MB-231 and human pancreas adenocarcinoma cell line PSN-1.
All these examples show the great potential displayed by marine bacteria which reveals the extraordinary chemical diversity of their metabolism. When analysing the bioactive bacteria and their taxonomic groups, Actinobacteria proved to be the most prolific and diverse producers (Table 1, Table 2 and Table 3). However, ignoring other less studied phyla denies access to valuable chemical diversity, which is essential in the drug discovery process. Moreover, marine bacterial metabolites have shown to have potential as treatment in both human and animal pathologies. Thus, marine bacterial natural products have great significance under the “One Health” framework.

Table 1.
Recently isolated bioactive molecules from marine bacteria with antimicrobial properties.

Table 2.
Recently isolated bioactive molecules from marine bacteria with antiviral properties.

Table 3.
Recently isolated bioactive molecules from marine bacteria with cytotoxic properties.
3. Screening Methodologies for the Detection of Bioactive Marine Bacteria
3.1. Conventional Screening for Antimicrobials
Traditional screening methodologies for antimicrobials are based on agar diffusion assays, with the Kirby-Bauer diffusion assay being the most commonly used. This assay was standardized in the 1960s by Kirby and colleagues [98] and this and other variations of agar diffusion assays are, in fact, the models currently used by the Clinical Laboratory Standards Institute (CLSI) for antimicrobial susceptibility. The diffusion assays consist of the inoculation of specific agar media, usually Mueller-Hinton or Mueller-Hinton II agar, with the pathogenic strains under test. Paper filters impregnated with the substance to be tested are then placed on top of the medium surface and the culture is incubated usually at 37 °C, for 4 to 10 h. After this period, the medium is inspected for the formation of circular spots without bacterial growth, designated inhibition halos, and their radiuses are measured and compared to reference values of other antimicrobials, as instructed by the CLSI [99]. The Kirby-Bauer diffusion assay is very simple and relatively low-cost assay, and it allows for a rapid testing of a considerable number of pathogens and bioactive antimicrobial principles. However, it offers no information on the potential bactericidal or bacteriostatic effect produced by the molecule being tested. Moreover, diffusion in the agar medium is size-dependent, meaning that for large molecules [100], antimicrobial susceptibility may be underestimated or that false negatives can be obtained. Likewise, although there are reports of the successful use of these assays in fungi [101], not all microorganisms can be tested, as it is the case for fastidious, slow-growing and microaerophiles bacteria. Furthermore, the impossibility of its automation also hampers use of this assay in large-scale drug discovery [102].
Regarding antiparasitics, as parasites are usually obligate intracellular, they cannot be tested in agar diffusion assays. Therefore, the discovery of these drugs relies on the use of either cultures of parasites or in vivo treatments. Direct treatment of sick patients can be a useful tool, as it allows several different interactions to occur, like host related immunity or metabolism or the possibility of re-infection. However, direct treatment come with critical disadvantages, ranging from limited testing capacity to possible dangerous side effects for the patients. However, most antiparasitic therapeutics come from the veterinary world, as this saves human patients from contact with potentially toxic substances, while maintaining all the advantages of the direct treatment. Nevertheless, it can still be a costly endeavour and may yield results which do not prove to be suitable for human treatment [103]. In vitro culturing of parasites can be done with free living or infectious parasites. Free living parasites can be cultivated in axenic cultures [104] while infectious parasites come from intracellular infection of a host cell and which are more physiologically relevant [105]. By isolating the parasites from the host, the interferences caused in in vivo cultures are excluded. Yet, inactive drug precursors that must be activated by the host or drugs which activity require the synergistic effect of the host immunity cannot be tested. Moreover, in vitro testing is more resource intensive, making it hard to apply in the regions that are most affected by these diseases [103].
3.2. Conventional Screening for Antiviral Molecules
Vaccines, as a prophylactic approach, are the primary strategy for controlling viral outbreaks and even successfully contributing to total eradication of the agent responsible [106]. However, post-infection treatments are also needed, mostly due to the high pathogenicity that current pandemic-responsible virus displays, as stated previously [20,21,22,23]. The urgency for new and innovative antiviral therapeutics is also reinforced by the quick mutation rate of viral strains, which often leads to the creation of unique and drug-resistant viruses [107].
Successful infection by a virus is dependent on various key steps, such as the kind of entry in the host cell and the genome replication [108]. Traditional screenings for evaluating promising antiviral compounds were firstly based on non-specific approaches like the plaque inhibition assay [109], which still plays an important role in the determination of viral inhibitory concentrations. The general premise is simple, as it is based on the incubation of viral plaque-forming units in plates with overlaid layers containing media, the host cells, and the drug to be tested. The formed plaques are then stained, normally with neutral red, and counted [110]. Several modified versions of the assay have already been developed [111,112,113]. Nevertheless, antiviral screenings have progressed to more specific approaches, by focusing on the obtainment of bioactive molecules with high specificity to affect particular steps in the virus cell cycle [24]. This approach, which focuses on the obtainment of highly specific molecules, is seen in a variety of in vitro cell based and biochemical assays [108]. Reporter proteins such as luciferase or β-galactosidase can be incorporated in the assay [114,115,116].However, to do so, a virus containing a copy of a related luciferase or a-galactosidase reporter gene in its genome is needed.
As viruses will always need to be cultivated in host cells, complementary cytotoxic studies of the potential antiviral drug (detailed in the next section) are also necessary to guarantee its non-cytotoxic effect against the cell utilized in the screening.
3.3. Conventional Screening for Cytotoxic Molecules
The first anti-cancer therapeutics came about from in vivo experimentations, as was the case of, for example, methotrexate [117]. As already stated, these assays have disadvantages, such as toxic side effects on patients or the low number of individuals and molecules tested. The introduction of the first cultivable immortalized cancer cell lines [118] lead to a great increase in the number of discovered therapeutics, by allowing more bioactive molecules to be safely tested. In vitro cytotoxicity testing is usually performed utilizing six techniques for the evaluation of molecule bioactivity. One is staining for viable cells, as in the sulphorhodamine assay [119] and another is dye exclusion, where only dead or damaged cells are stained, as is the case of Trypan Blue [120]. Cytotoxic activity can also be assessed through metabolic activity, or a lack thereof, viability of protease activity, clonogenic capability and DNA synthesis. Methods based on metabolic activity usually rely on reduction reactions, namely the reduction of tetrazolium salts [121] and the resazurin reduction assay [122], or on the measurement of adenosine triphosphate levels, that drop as cells die [123]. In protease viability assays, cell viability is assessed by measuring the intracellular or extracellular protease activity. In intracellular protease assays, viability is positively correlated with protease activity and are performed with fluorogenic protease substrates that can penetrate cell membranes. However, in extracellular protease assays, the viability is negatively correlated with protease activity and are performed with fluorogenic protease substrates that cannot penetrate cell membranes [124]. The clonogenic capability of a cell is its ability to proliferate indefinitely, and the loss of this capacity can be related to the antitumor capacity of compounds [125]. When cells proliferate, DNA must be synthesised. In this case, quantifying the de novo synthesis of DNA is one of the most reliable and accurate proliferation assays. This can be done by incubation of cells with radioactive labels like 3H or specific thymidine analogues that can be detected though immunoblotting [126].
3.4. Genetic Analysis of Bioactive Potential
As mentioned earlier, the described bacterial bioactive secondary metabolites demonstrate a high degree of chemical and structural diversity and complexity. As these molecules do not appear to participate directly in the growth and development of the organism but are often important for other actions like stress response, competition or defence, they are considered secondary metabolites. Hence their potential for having biotechnologically interesting features [127]. Polyketide synthases (PKS) and non-ribosomal peptide synthetases (NRPS) are examples of families of complex enzymatic machineries that have been proven to be responsible for the production of most bacterial bioactive molecules [128]. The search for genes related to these complexes can be made using simple and low-cost molecular techniques such as DNA extraction and amplification using specific primers [129,130]. Early-on, genetic screenings can thus complement preliminary bioactivity screenings and help identify the most promising strains for use in further molecule isolation studies. In fact, this is important not only to help reduce the number of strains to work with, but also to discriminate promising strains belonging to the same species, as even phylogenetically close strains can have different genomic content regarding the presence of biosynthetic gene clusters [131]. Moreover, this approach has proven to be successful in helping select good candidates which ultimately showed interesting bioactive profiles, including the potential production of novel molecules [131,132,133]. Furthermore, with the increase in available data on sequenced bacterial genomes, genome mining techniques have rapidly evolved for detecting secondary metabolite gene clusters [134]. Several online platforms dedicated to the search for genes encoding core-biosynthetic enzymes are currently available, such as BAGEL [135], NaPDos and antiSMASH [136], amongst others. BAGEL can detect putative bacteriocins and ribosomally synthesized and post-translationally modified peptides (RiPPs) of gene clusters in bacterial genomes whereas NaPDos extracts C- and KS- domains to identify secondary metabolite genes in general. AntiSMASH is the most complete platform, as it uses a library of enzymes/protein domains that are normally seen in secondary metabolite biosynthetic pathways to identify possible hits with 44 different types of gene clusters [136]. These include, not only various types of PKS, NRPS and PKS/NRPS hybrids, but also other important secondary metabolite gene clusters for biosynthesis of compounds like RiPPs, terpenoids/isoprenoids, and ectoines, among others. This approach is already successfully assisting in discovering and characterize new natural products, as is the case for the macrolactams lobosamide A, B and C [72], which were characterized with the help of antiSMASH and the polythioamide neothioviridamide [85], whose biosynthetic cluster belongs to a cryptic RiPP and was heterogeneously expressed in S. avermitilis (discussed in Section 2.2.1 and Section 2.2.3). Genome mining for secondary metabolite genes can also be applied in metagenomic data of still uncultured organisms, as well as to help unveil the biotechnological potential of unique and still under-explored bacterial groups, such as, for example, the planctomycetes [137,138,139].
3.5. High-Throughput Screening Methodologies
Because of the drawbacks imposed by traditional screening methodologies, biotechnological pharmaceutical companies have largely abandoned these assay models in favour of high-throughput screening (HTS) methodologies. HTS methodologies are characterized by rapid, high-volume testing of compounds, in the range of 105 to 106 molecules per day, and are inserted in a whole drug discovery strategy directed towards novel therapeutic leads [140]. HTS are easily miniaturized in 96-, 384- or 1,536-microwell plate formats and are generally based on cell or biochemical targets in which the response is measured as either an increase or a decrease of a signal. The signal measured is usually based on absorbance methods, e.g., fluorescence [141]. Non-conventional methodologies, genetic tools and HTS methodologies, are less labour demanding, improve the speed and capacity of testing, and discovering of novel bioactive molecules. They potentially allow the decrease of the needed time for obtainment of new pharmaceuticals, which is a fundamental aspect in the lengthy process of drug discovery.
4. Strategies for Avoiding Reisolation and Recharacterization of Known Bioactive Compounds
The great chemical diversity present in natural molecules has been the impetus behind the discovery of novel therapeutics [142]. Yet, the isolation and characterization of novel natural molecules is a laborious, time consuming endeavour that requires particular expertise and competences [143]. This is mainly due to the presence of known and redundant compounds in crude extracts [144]. Thus, a strategy for recognizing and eliminating already known bioactive natural molecules in the early stage of the screening process is crucial. This process is called dereplication [145].
The dereplication process analyses previously identified active samples in preliminary biological screenings through a combination of analytical and spectroscopic methods to putatively identify any possible known bioactive molecules present. In this section, current strategies for the dereplication of bioactive extracts will be explained.
4.1. Analytical Separation Techniques
Crude bacterial extracts are complex mixtures of organic molecules [146], usually containing nutrients, primary and secondary metabolites [147]. Due to this complexity, in the dereplication process, crude extracts must first be fractionated prior to any spectroscopic analysis [145]. While many developed separation techniques exist, like thin-layer chromatography, gas chromatography and capillary electrophoresis to name but a few, the most commonly used techniques in the dereplication process pivot between high-performance liquid chromatography (HPLC) and ultra-high-performance liquid chromatography (UHPLC) [148]. The extensive use of HPLC systems in the isolation of natural products has allowed it to become a powerful and versatile tool, as HPLC systems can be easily hyphenated with detection techniques (like ultraviolet/visible (UV/Vis) detectors, mass (MS) and NMR spectrometers, see below) and copious amounts of data available, makes HPLC the de facto standard chromatographic technique for dereplication [142]. Still, while UHPLC systems are less used, they can operate at very high pressures with very small particle size columns, which grants them great sensitivity while diminishing sample and eluent volumes used and the analysis time [149]. However, both techniques allow for the use of compatible column types and chemical phases. Because of the chemical nature of most natural products, small molecules with molecular weights bellow 1000 Da and low or no polarity [147], the best method for their separation is revered-phase chromatography which allows a longer retention time in the column during the polar and elution in the non-polar phase [150,151]. The time that the metabolite is retained in the column, also called retention time, gives an important chromatographic property intrinsic to each molecule that can be used in its identification [145,148,152].
4.2. Detection Methods
In the dereplication process, molecules are identified due to intrinsic properties, like absorption peaks or molecular mass, which can be measured by a number of detection techniques. The most common techniques include, UV/Vis diode array detectors (DAD), mass spectrometers and NMR spectrometers [142].
In UV/Vis spectroscopy, UV/vis spectra can be immediately acquired as part of the chromatographic separation of the extracts [147]. UV/vis spectra are normally obtained through the use of DAD, which can scan a range of wavelengths and produce the absorption spectrum. UV/Vis spectroscopy gives useful information on the molecules, such as chromophores present and the max absorption wavelength (λmax), giving important structural information for the identification of yet unknown molecules [153]. Yet, but for the dereplication process, the full spectrum is a superior and more definitive approach, as it contains more information than just the λmax can give [148]. However, in the dereplication process, UV/vis spectra are normally used only as a secondary criterion. Currently, the principal criterion for identification is based on high resolution-MS (HRMS) [142]. MS is a destructive technique that provides the mass-to-charge ratio (m/z) of the ionized molecule or ionized fragments of the molecule. MS data is presented as spectrum of m/z and relative intensity. The m/z can be used to estimate the molecular mass of the molecule and thus, its molecular formula [154]. Additionally, as the ionization of the molecule may fragment it in smaller pieces, this fragmentation pattern can also be used in the identification of the molecule. Moreover, if it is a novel molecule, the fragmentation pattern can give useful information for structural elucidation of the molecule [155]. Initial dereplication efforts were based around low resolution-MS (LRMS) [156]. However, LRMS is ineffective in resolving between two very closely weight molecules, as LRMS can only confidently detect up to the integer of the mass. Thus, HRMS, which can give extremely accurate molecular masses [154], was adopted. Nevertheless, an LC-DAD-MS dereplication only putatively identifies the compounds present in the extract, because of the possible existence of regioisomers or stereoisomers [148]. Therefore, full confirmation of the identity is only possible through NMR spectroscopy. NMR spectroscopy of natural products is normally performed on protons (1H-NMR) and carbons (13C-NMR) where the chemical shifts measured correlate with the position and type of each atom in the molecule. NMR has various advantages compared to MS, as it can analyse all kinds of small metabolites and samples can be recovered for further analyses. It is considered the universal detector for structure elucidation, has high analytical reproducibility and simplicity of sample preparation [142] and can be used for checking the purity of the sample, a problem that can arise from the common purification steps undertaken in most natural product isolation [145]. An example of this are lipids, problematic impurities that are undetectable in LC-DAD-MS analysis because they have low UV absorption, have difficulty of ionizing and high lipophilicity, which, 1H-NMR can easily recognize by the presence of the large peak of the lipidic methylene chain [157]. Yet, for NMR spectroscopy, sample size is a limiting factor, as it may require up to milligrams of the sample [142]. In contrast, MS can work with attomolar concentrations of sample [158]. This limitation in NMR spectroscopy is especially important in the dereplication process, as crude extracts may only contain minute quantities of molecules.
4.3. Natural Product Databases
Databases of natural products are critical to successfully complete the dereplication process [142]. Databases contain crucial information on natural products, including molecular weight, exact mass and fragmentation pattern, UV/vis λmax absorption or full spectra and NMR spectra or chemical shifts [148]. Databases of natural products should also contain relevant taxonomic information of the producer organism and known biological activity. For the purpose of identification of natural products, public domain and commercial databases are available. The most important commercial databases include Chemical Abstracts Service’s Registry File (CAS) [159], NaprAlert [160], the Chapman & Hall Dictionary of Natural Products (DNP), AntiBase [161], MarinLit [162], the open-access databases Global Natural Products Social Networking (GNPS) [163,164], The Natural Products Atlas (NPAtlas) [165,166] and Small Molecule Accurate Recognition Technology (SMART) [167,168]. The CAS database comprises the largest online repository of chemical structures, including natural products. NaprAlert has significant resources for dereplicating the terrestrial sources. MarinLit is a database dedicated to marine natural products established by Professors John Blunt and Murray Munro. Records contain bibliographic information and the database can be searched by querying substructure, 13C- and 1H-NMR shift data, exact mass, chemical formula, or UV λmax absorption. AntiBase focuses on molecules from microorganisms and higher fungi. AntiBase incorporates molecular formula and mass, melting point, optical rotation, UV λmax absorption, 13C and 1H-NMR shift data and mass spectra. AntiBase also contains bioactivity data and information on origin, isolation and literature sources.
The DNP is considered the most complete natural product database, containing relevant bibliographic references [148]. The DNP contains information on exact mass and molecular formula, UV λmax absorption, biological sources and activity. Of the commercial databases, for marine bacterial natural products, MarinLit, AntiBase and the DNP and its subset on marine natural products (DMNP) are considered the most relevant databases for marine bacterial natural products dereplication. GNPS is an open-access database hosted at University of California, San Diego of LC and MS/MS spectra below m/z 2000 of natural products with data of over 18,000 compounds. GNPS also can detect sets of spectra from related molecules, even if the spectra used is not matched to any known compounds [142]. It allows the use of experimental MS/MS spectra as input in searches, allowing the direct dereplication of molecules present in extracts or chromatographic fractions without the need for determining the molecular formulae of their components. Additionally, the algorithm behind the platform allows networks of molecules based on their cosine similarity to be established, a process that has proved to be very useful in the identification of new members of known families of natural products. Bioactivity data can also be added to the process, allowing the direct correlation between the presence of certain components in samples and their bioactivity, using the so-called “bioactive molecular networking” (BMN) approach [169].
GNPS molecular networking has been successfully applied to the discovery of several bioactive natural products, including the antimicrobial angucycline-derived polycyclic aromatic polyketide lugdunomycin (65) (Figure 7) [170], and the aromatic polyketide accramycin A (66) [171], both isolated from Streptomyces sp., as well as the antibacterial chlorinated cyclic hexapeptides noursamycins A-F (67-72), obtained from cultures of Streptomyces noursei NTR-SR4 [172]. NPAtlas is a database which includes over 25,500 compounds from microbial sources. It contains referenced data for structure, compound names, source organisms, isolation references, total syntheses, and instances of structural reassignment and is integrated with other natural product databases, like GNPS [165]. SMART is an NMR database [in particular, non-uniform sampling heteronuclear single quantum coherence (NUS HSQC)] assisted by a deep convolutional neural network (dCNN) to not only identify compounds, but for novel unidentified ones, placing them in natural product families with similar HSQC spectra, thus allowing for faster structural elucidation. As SMART is assisted by dCNN, as the size of the training set increases for a given compound class, so does its accuracy and capability of prediction of a new compound’s structure improve [167]. Additionally, with further training, SMART has the potential to enable novel experimentation at the chemistry-biology interface, like predictions of biological activity and or target. The SMART technology in combination with GNPS was successfully applied recently to the discovery of symplocolide A (73), a new chimeric swinholide-like macrolide with cytotoxic properties against the NCI-H460 human lung cancer cell line obtained from extracts of the filamentous marine cyanobacterium Symploca sp. [173].
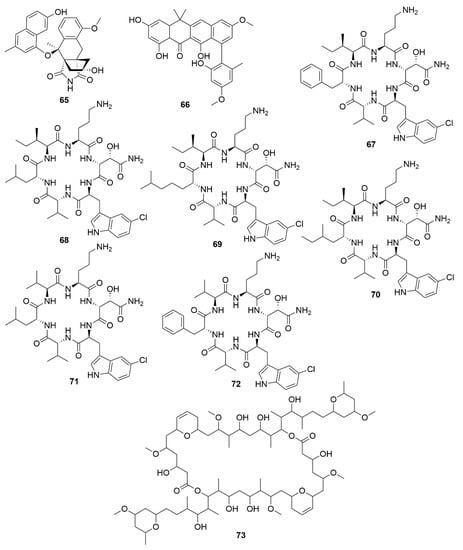
Figure 7.
Examples of recently isolated bacterial natural products employing the GNPS and SMART technologies. Lugdunomycin (65), a novel angucycline-derived polycyclic aromatic polyketide with antimicrobial properties, isolated from Streptomyces sp. QL37 utilising a metabolomic combined with MS-based molecular networking analysis. Accramycin A (66), a naphthacene-type aromatic polyketide isolated from Streptomyces sp. MA37, which characterization was aided by MS/MS molecular networking. Noursamycin A (67), Noursamycin B (68), Noursamycin C1 (69), Noursamycin C2 (70), Noursamycin D (71) and Noursamycin E (72) are chlorinated cyclic hexapeptides isolated from S. noursei with the aid of MS/MS molecular networking. Symplocolide A (73), a swinholide-like macrolide with cytotoxic properties obtained from extracts of the filamentous marine cyanobacterium Symploca sp. by employing both SMART technologies and MS/MS molecular networking.
5. Conclusions
It is evident that great heath challenges are still in need of novel pharmaceutical responses. Resistance of bacteria and fungi to antibiotics, of parasites like malaria to the available treatments, unavailability of medicines to treat new viruses and efficient drugs for cancer treatment, require additional research and development focus in discovering novel bioactive molecules. Oceans, home to a substantial portion of the world’s biodiversity, are still underexplored, and are an important source of drugs and drug leads. Since 2014, several novel marine bacterial natural products, obtained mainly from Actinobacteria, Firmicutes and Proteobacteria, were described and reported in this review. In supplement to conventional methods, high-throughput analysis, like genome and metagenomic analysis and HTS, and the use of new dereplication and identification tools, will foster the discovery of new leads. This review shows that marine bacteria are key to the development of new pharmaceuticals, especially if combined with a rational, high-throughput approach.
Author Contributions
Conceptualization and investigation by J.D.S., writing—original draft preparation by J.D.S., I.V., review and editing by J.D.S., I.V., F.R.; F.V. and O.M.L. All authors have read and agreed to the published version of the manuscript.
Funding
This research was partially supported by the Strategic Funding UID/Multi/04423/2019 through national funds provided by FCT—Foundation for Science and Technology and European Regional Development Fund (ERDF), in the framework of the programme PT2020. José Diogo Santos is supported by an FCT doctoral grant (SFRH/BD/145576/2019). Inês Vitorino is supported by an FCT doctoral grant (SFRH/BD/145577/2019).
Conflicts of Interest
J.D.S., I.V. and O.M. declare no conflict of interest. F.R. and F.V. disclosed the receipt of financial support from Fundación MEDINA, a public–private partnership of Merck Sharp & Dohme de España S.A./Universidad de Granada/Junta de Andalucía.
References
- Van Boeckel, T.P.; Pires, J.; Silvester, R.; Zhao, C.; Song, J.; Criscuolo, N.G.; Gilbert, M.; Bonhoeffer, S.; Laxminarayan, R. Global trends in antimicrobial resistance in animals in low- and middle-income countries. Science 2019, 365, eaaw1944. [Google Scholar] [CrossRef]
- Frost, I.; Van Boeckel, T.P.; Pires, J.; Craig, J.; Laxminarayan, R. Global geographic trends in antimicrobial resistance: The role of international travel. J. Travel Med. 2019, 26, taz036. [Google Scholar] [CrossRef]
- WHO. Global Antimicrobial Resistance Surveillance System (GLASS) Report: Early Implementation 2017–2018; World Health Organization: Geneva, Switzerland, 2019. [Google Scholar]
- WHO. Antimicrobial Resistance: Global Report on Surveillance; World Health Organization: Geneva, Switzerland, 2014; p. xxii. 232p. [Google Scholar]
- WHO. Global Priority List of Antibiotic-Resistant Bacteria to Guide Research, Discovery, and Development of New Antibiotics; World Health Organization: Geneva, Switzerland, 2017; p. 7. [Google Scholar]
- Toda, M.; Williams, S.R.; Berkow, E.L.; Farley, M.M.; Harrison, L.H.; Bonner, L.; Marceaux, K.M.; Hollick, R.; Zhang, A.Y.; Schaffner, W.; et al. Population-Based Active Surveillance for Culture-Confirmed Candidemia—Four Sites, United States, 2012–2016. MMWR Surveill. Summ. 2019, 68, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Lockhart, S.R.; Etienne, K.A.; Vallabhaneni, S.; Farooqi, J.; Chowdhary, A.; Govender, N.P.; Colombo, A.L.; Calvo, B.; Cuomo, C.A.; Desjardins, C.A.; et al. Simultaneous Emergence of Multidrug-Resistant Candida auris on 3 Continents Confirmed by Whole-Genome Sequencing and Epidemiological Analyses. Clin. Infect. Dis. 2017, 64, 134–140. [Google Scholar] [CrossRef]
- Pfaller, M.A. Antifungal drug resistance: Mechanisms, epidemiology, and consequences for treatment. Am. J. Med. 2012, 125, S3–S13. [Google Scholar] [CrossRef] [PubMed]
- Bueid, A.; Howard, S.J.; Moore, C.B.; Richardson, M.D.; Harrison, E.; Bowyer, P.; Denning, D.W. Azole antifungal resistance in Aspergillus fumigatus: 2008 and 2009. J. Antimicrob. Chemother. 2010, 65, 2116–2118. [Google Scholar] [CrossRef] [PubMed]
- Howard, S.J.; Cerar, D.; Anderson, M.J.; Albarrag, A.; Fisher, M.C.; Pasqualotto, A.C.; Laverdiere, M.; Arendrup, M.C.; Perlin, D.S.; Denning, D.W. Frequency and evolution of Azole resistance in Aspergillus fumigatus associated with treatment failure. Emerg. Infect. Dis. 2009, 15, 1068–1076. [Google Scholar] [CrossRef]
- Hofer, U. The cost of antimicrobial resistance. Nat. Rev. Microbiol. 2019, 17, 3. [Google Scholar] [CrossRef]
- Tjitra, E.; Anstey, N.M.; Sugiarto, P.; Warikar, N.; Kenangalem, E.; Karyana, M.; Lampah, D.A.; Price, R.N. Multidrug-resistant Plasmodium vivax associated with severe and fatal malaria: A prospective study in Papua, Indonesia. PLoS Med. 2008, 5, e128. [Google Scholar] [CrossRef]
- Kennedy, P.G. Clinical features, diagnosis, and treatment of human African trypanosomiasis (sleeping sickness). Lancet Neurol. 2013, 12, 186–194. [Google Scholar] [CrossRef]
- Barrett, M.P.; Croft, S.L. Management of trypanosomiasis and leishmaniasis. Br. Med. Bull. 2012, 104, 175–196. [Google Scholar] [CrossRef] [PubMed]
- Ridley, R.G. Medical need, scientific opportunity and the drive for antimalarial drugs. Nature 2002, 415, 686–693. [Google Scholar] [CrossRef]
- Leang, R.; Taylor, W.R.; Bouth, D.M.; Song, L.; Tarning, J.; Char, M.C.; Kim, S.; Witkowski, B.; Duru, V.; Domergue, A.; et al. Evidence of Plasmodium falciparum Malaria Multidrug Resistance to Artemisinin and Piperaquine in Western Cambodia: Dihydroartemisinin-Piperaquine Open-Label Multicenter Clinical Assessment. Antimicrob. Agents Chemother. 2015, 59, 4719–4726. [Google Scholar] [CrossRef] [PubMed]
- Li, Y. Qinghaosu (artemisinin): Chemistry and pharmacology. Acta Pharmacol. Sin. 2012, 33, 1141–1146. [Google Scholar] [CrossRef] [PubMed]
- Lostroh, P. Molecular and Cellular Biology of Viruses, 1st ed.; CRC Press: Boca Raton, FL, USA, 2019. [Google Scholar]
- Paules, C.I.; Marston, H.D.; Fauci, A.S. Coronavirus Infections-More Than Just the Common Cold. JAMA 2020. [Google Scholar] [CrossRef]
- Zanotto, P.M.A.; Leite, L.C.C. The Challenges Imposed by Dengue, Zika, and Chikungunya to Brazil. Front. Immunol. 2018, 9, 1964. [Google Scholar] [CrossRef]
- Gostin, L.O.; Lucey, D.; Phelan, A. The Ebola Epidemic: A Global Health Emergency. JAMA 2014, 312, 1095–1096. [Google Scholar] [CrossRef]
- de Wilde, A.H.; Snijder, E.J.; Kikkert, M.; van Hemert, M.J. Host Factors in Coronavirus Replication. Curr. Top Microbiol. Immunol. 2018, 419, 1–42. [Google Scholar] [CrossRef]
- Vega, L.E.; Espinoza, L.R. Human immunodeficiency virus infection (HIV)-associated rheumatic manifestations in thepre- and post-HAART eras. Clin. Rheumatol. 2020. [Google Scholar] [CrossRef]
- Morens, D.M.; Fauci, A.S. Emerging infectious diseases: Threats to human health and global stability. PLoS Pathog. 2013, 9, e1003467. [Google Scholar] [CrossRef]
- Destoumieux-Garzon, D.; Mavingui, P.; Boetsch, G.; Boissier, J.; Darriet, F.; Duboz, P.; Fritsch, C.; Giraudoux, P.; Le Roux, F.; Morand, S.; et al. The One Health Concept: 10 Years Old and a Long Road Ahead. Front. Vet. Sci. 2018, 5, 14. [Google Scholar] [CrossRef] [PubMed]
- Robinson, T.P.; Bu, D.P.; Carrique-Mas, J.; Fevre, E.M.; Gilbert, M.; Grace, D.; Hay, S.I.; Jiwakanon, J.; Kakkar, M.; Kariuki, S.; et al. Antibiotic resistance is the quintessential One Health issue. Trans. R. Soc. Trop. Med. Hyg. 2016, 110, 377–380. [Google Scholar] [CrossRef] [PubMed]
- Chang, Q.; Wang, W.; Regev-Yochay, G.; Lipsitch, M.; Hanage, W.P. Antibiotics in agriculture and the risk to human health: How worried should we be? Evol. Appl. 2015, 8, 240–247. [Google Scholar] [CrossRef] [PubMed]
- CDC. One Health Basics. Available online: https://www.cdc.gov/onehealth/basics/index.html (accessed on 23 June 2020).
- WHO. One Health. Available online: https://www.who.int/news-room/q-a-detail/one-health (accessed on 23 June 2020).
- Global Burden of Disease Cancer Collaboration. Global, Regional, and National Cancer Incidence, Mortality, Years of Life Lost, Years Lived With Disability, and Disability-Adjusted Life-years for 32 Cancer Groups, 1990 to 2015: A Systematic Analysis for the Global Burden of Disease Study. JAMA Oncol. 2017, 3, 524–548. [Google Scholar] [CrossRef]
- GBD 2015 Mortality; Causes of Death Collaborators. Global, regional, and national life expectancy, all-cause mortality, and cause-specific mortality for 249 causes of death, 1980–2015: A systematic analysis for the Global Burden of Disease Study 2015. Lancet 2016, 388, 1459–1544. [Google Scholar] [CrossRef]
- Ferlay, J.; Colombet, M.; Soerjomataram, I.; Mathers, C.; Parkin, D.M.; Pineros, M.; Znaor, A.; Bray, F. Estimating the global cancer incidence and mortality in 2018: GLOBOCAN sources and methods. Int. J. Cancer 2019, 144, 1941–1953. [Google Scholar] [CrossRef]
- Bertram, J.S. The molecular biology of cancer. Mol. Asp. Med. 2000, 21, 167–223. [Google Scholar] [CrossRef]
- Semmler, L.; Reiter-Brennan, C.; Klein, A. BRCA1 and Breast Cancer: A Review of the Underlying Mechanisms Resulting in the Tissue-Specific Tumorigenesis in Mutation Carriers. J. Breast Cancer 2019, 22, 1–14. [Google Scholar] [CrossRef]
- Global Burden of Disease Cancer, C.; Fitzmaurice, C.; Dicker, D.; Pain, A.; Hamavid, H.; Moradi-Lakeh, M.; MacIntyre, M.F.; Allen, C.; Hansen, G.; Woodbrook, R.; et al. The Global Burden of Cancer 2013. JAMA Oncol. 2015, 1, 505–527. [Google Scholar] [CrossRef]
- Newman, D.J.; Cragg, G.M. Microbial antitumor drugs: Natural products of microbial origin as anticancer agents. Curr. Opin. Investig. Drugs 2009, 10, 1280–1296. [Google Scholar]
- Kamb, A.; Wee, S.; Lengauer, C. Why is cancer drug discovery so difficult? Nat. Rev. Drug Discov. 2007, 6, 115–120. [Google Scholar] [CrossRef]
- Forrest, R.D. Early history of wound treatment. J. R. Soc. Med. 1982, 75, 198–205. [Google Scholar] [PubMed]
- Montinari, M.R.; Minelli, S.; De Caterina, R. The first 3500 years of aspirin history from its roots—A concise summary. Vasc. Pharmacol. 2019, 113, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Haefner, B. Drugs from the deep: Marine natural products as drug candidates. Drug Discov. Today 2003, 8, 536–544. [Google Scholar] [CrossRef]
- Dias, D.A.; Urban, S.; Roessner, U. A historical overview of natural products in drug discovery. Metabolites 2012, 2, 303–336. [Google Scholar] [CrossRef]
- Gerwick, W.H.; Moore, B.S. Lessons from the past and charting the future of marine natural products drug discovery and chemical biology. Chem. Biol. 2012, 19, 85–98. [Google Scholar] [CrossRef] [PubMed]
- Weinheimer, A.J.; Spraggins, R.L. The occurrence of two new prostaglandin derivatives (15-epi-PGA2 and its acetate, methyl ester) in the gorgonian Plexaura homomalla chemistry of coelenterates. XV. Tetrahedron Lett. 1969, 10, 5185–5188. [Google Scholar] [CrossRef]
- Masuda, M.; Abe, T.; Suzuki, T.; Suzuki, M. Morphological and chemotaxonomic studies on Laurencia composita and L. okamurae (Ceramiales, Rhodophyta). Phycologia 1996, 35, 550–562. [Google Scholar] [CrossRef]
- Taylor, R.E. Tedanolide and the evolution of polyketide inhibitors of eukaryotic protein synthesis. Nat. Prod. Rep. 2008, 25, 854–861. [Google Scholar] [CrossRef]
- Selvin, J.; Ninawe, A.S.; Kiran, G.; Arya, S.; Priyadharshini, S. Bioactive Marine Natural Products: Insights into Marine Microbes, Seaweeds, and Marine Sponges as Potential Sources of Drug Discovery. In Marine Microorganisms Extraction and Analysis of Bioactive Compounds, 1st ed.; Nollet, L.M.L., Ed.; CRC Press: Boca Raton, FL, USA, 2016. [Google Scholar]
- Laport, M.S.; Santos, O.C.; Muricy, G. Marine sponges: Potential sources of new antimicrobial drugs. Curr. Pharm. Biotechnol. 2009, 10, 86–105. [Google Scholar] [CrossRef]
- Bai, R.L.; Paull, K.D.; Herald, C.L.; Malspeis, L.; Pettit, G.R.; Hamel, E. Halichondrin B and homohalichondrin B, marine natural products binding in the vinca domain of tubulin. Discovery of tubulin-based mechanism of action by analysis of differential cytotoxicity data. J. Biol. Chem. 1991, 266, 15882–15889. [Google Scholar] [PubMed]
- Sipkema, D.; Osinga, R.; Schatton, W.; Mendola, D.; Tramper, J.; Wijffels, R.H. Large-scale production of pharmaceuticals by marine sponges: Sea, cell, or synthesis? Biotechnol. Bioeng. 2005, 90, 201–222. [Google Scholar] [CrossRef]
- Munro, M.H.; Blunt, J.W.; Dumdei, E.J.; Hickford, S.J.; Lill, R.E.; Li, S.; Battershill, C.N.; Duckworth, A.R. The discovery and development of marine compounds with pharmaceutical potential. J. Biotechnol. 1999, 70, 15–25. [Google Scholar] [CrossRef]
- Shetty, N.; Gupta, S. Eribulin drug review. South Asian J. Cancer 2014, 3, 57–59. [Google Scholar] [CrossRef] [PubMed]
- Carroll, A.R.; Copp, B.R.; Davis, R.A.; Keyzers, R.A.; Prinsep, M.R. Marine natural products. Nat. Prod. Rep. 2020, 37, 175–223. [Google Scholar] [CrossRef] [PubMed]
- Haygood, M.G.; Schmidt, E.W.; Davidson, S.K.; Faulkner, D.J. Microbial symbionts of marine invertebrates: Opportunities for microbial biotechnology. J. Mol. Microbiol. Biotechnol. 1999, 1, 33–43. [Google Scholar] [PubMed]
- Pettit, G.R. The bryostatins. Fortschr. Chem. Org. Naturst. 1991, 57, 153–195. [Google Scholar] [CrossRef]
- Pettit, G.R.; Herald, C.L.; Doubek, D.L.; Herald, D.L.; Arnold, W.; Clardy, J. Isolation and structure of bryostatin 1. J. Am. Chem. Soc. 1982, 104, 6846–6848. [Google Scholar] [CrossRef]
- Sudek, S.; Lopanik, N.B.; Waggoner, L.E.; Hildebrand, M.; Anderson, C.; Liu, H.; Patel, A.; Sherman, D.H.; Haygood, M.G. Identification of the putative bryostatin polyketide synthase gene cluster from “Candidatus Endobugula sertula”, the uncultivated microbial symbiont of the marine bryozoan Bugula neritina. J. Nat. Prod. 2007, 70, 67–74. [Google Scholar] [CrossRef]
- John Faulkner, D. Marine natural products. Nat. Prod. Rep. 2000, 17, 7–55. [Google Scholar] [CrossRef]
- Feling, R.H.; Buchanan, G.O.; Mincer, T.J.; Kauffman, C.A.; Jensen, P.R.; Fenical, W. Salinosporamide A: A highly cytotoxic proteasome inhibitor from a novel microbial source, a marine bacterium of the new genus salinospora. Angew. Chem. Int. Ed. Engl. 2003, 42, 355–357. [Google Scholar] [CrossRef] [PubMed]
- Blunt, J.W.; Copp, B.R.; Munro, M.H.G.; Northcote, P.T.; Prinsep, M.R. Marine natural products. Nat. Prod. Rep. 2005, 22, 15–61. [Google Scholar] [CrossRef] [PubMed]
- Dyshlovoy, S.A.; Honecker, F. Marine Compounds and Cancer: The First Two Decades of XXI Century. Mar. Drugs 2019, 18, 20. [Google Scholar] [CrossRef] [PubMed]
- Blunt, J.W.; Carroll, A.R.; Copp, B.R.; Davis, R.A.; Keyzers, R.A.; Prinsep, M.R. Marine natural products. Nat. Prod. Rep. 2018, 35, 8–53. [Google Scholar] [CrossRef] [PubMed]
- Carroll, A.R.; Copp, B.R.; Davis, R.A.; Keyzers, R.A.; Prinsep, M.R. Marine natural products. Nat. Prod. Rep. 2019, 36, 122–173. [Google Scholar] [CrossRef]
- Medema, M.H.; Fischbach, M.A. Computational approaches to natural product discovery. Nat. Chem. Biol. 2015, 11, 639–648. [Google Scholar] [CrossRef]
- Wiese, J.; Abdelmohsen, U.R.; Motiei, A.; Humeida, U.H.; Imhoff, J.F. Bacicyclin, a new antibacterial cyclic hexapeptide from Bacillus sp. strain BC028 isolated from Mytilus edulis. Bioorg. Med. Chem. Lett. 2018, 28, 558–561. [Google Scholar] [CrossRef]
- Rodriguez, V.; Martin, J.; Sarmiento-Vizcaino, A.; de la Cruz, M.; Garcia, L.A.; Blanco, G.; Reyes, F. Anthracimycin B, a Potent Antibiotic against Gram-Positive Bacteria Isolated from Cultures of the Deep-Sea Actinomycete Streptomyces cyaneofuscatus M-169. Mar. Drugs 2018, 16, 406. [Google Scholar] [CrossRef]
- Reynolds, K.A.; Luhavaya, H.; Li, J.; Dahesh, S.; Nizet, V.; Yamanaka, K.; Moore, B.S. Isolation and structure elucidation of lipopeptide antibiotic taromycin B from the activated taromycin biosynthetic gene cluster. J. Antibiot. 2018, 71, 333–338. [Google Scholar] [CrossRef]
- Anjum, K.; Sadiq, I.; Chen, L.; Kaleem, S.; Li, X.-C.; Zhang, Z.; Lian, X.-Y. Novel antifungal janthinopolyenemycins A and B from a co-culture of marine-associated Janthinobacterium spp. ZZ145 and ZZ148. Tetrahedron Lett. 2018, 59, 3490–3494. [Google Scholar] [CrossRef]
- Zhang, B.; Wang, K.B.; Wang, W.; Bi, S.F.; Mei, Y.N.; Deng, X.Z.; Jiao, R.H.; Tan, R.X.; Ge, H.M. Discovery, Biosynthesis, and Heterologous Production of Streptoseomycin, an Anti-Microaerophilic Bacteria Macrodilactone. Org. Lett. 2018, 20, 2967–2971. [Google Scholar] [CrossRef] [PubMed]
- Le, T.C.; Yang, I.; Yoon, Y.J.; Nam, S.J.; Fenical, W. Ansalactams B-D Illustrate Further Biosynthetic Plasticity within the Ansamycin Pathway. Org. Lett. 2016, 18, 2256–2259. [Google Scholar] [CrossRef] [PubMed]
- Harinantenaina Rakotondraibe, L.; Rasolomampianina, R.; Park, H.Y.; Li, J.; Slebodnik, C.; Brodie, P.J.; Blasiak, L.C.; Hill, R.; TenDyke, K.; Shen, Y.; et al. Antiproliferative and antiplasmodial compounds from selected Streptomyces species. Bioorg. Med. Chem. Lett. 2015, 25, 5646–5649. [Google Scholar] [CrossRef] [PubMed]
- Braña, A.F.; Sarmiento-Vizcaino, A.; Perez-Victoria, I.; Otero, L.; Fernandez, J.; Palacios, J.J.; Martin, J.; de la Cruz, M.; Diaz, C.; Vicente, F.; et al. Branimycins B and C, Antibiotics Produced by the Abyssal Actinobacterium Pseudonocardia carboxydivorans M-227. J. Nat. Prod. 2017, 80, 569–573. [Google Scholar] [CrossRef]
- Schulze, C.J.; Donia, M.S.; Siqueira-Neto, J.L.; Ray, D.; Raskatov, J.A.; Green, R.E.; McKerrow, J.H.; Fischbach, M.A.; Linington, R.G. Genome-Directed Lead Discovery: Biosynthesis, Structure Elucidation, and Biological Evaluation of Two Families of Polyene Macrolactams against Trypanosoma brucei. ACS Chem. Biol. 2015, 10, 2373–2381. [Google Scholar] [CrossRef]
- Schulze, C.J.; Navarro, G.; Ebert, D.; DeRisi, J.; Linington, R.G. Salinipostins A-K, long-chain bicyclic phosphotriesters as a potent and selective antimalarial chemotype. J. Org. Chem. 2015, 80, 1312–1320. [Google Scholar] [CrossRef]
- Raju, R.; Khalil, Z.G.; Piggott, A.M.; Blumenthal, A.; Gardiner, D.L.; Skinner-Adams, T.S.; Capon, R.J. Mollemycin A: An antimalarial and antibacterial glyco-hexadepsipeptide-polyketide from an Australian marine-derived Streptomyces sp. (CMB-M0244). Org. Lett. 2014, 16, 1716–1719. [Google Scholar] [CrossRef]
- Abdelmohsen, U.R.; Cheng, C.; Viegelmann, C.; Zhang, T.; Grkovic, T.; Ahmed, S.; Quinn, R.J.; Hentschel, U.; Edrada-Ebel, R. Dereplication strategies for targeted isolation of new antitrypanosomal actinosporins A and B from a marine sponge associated-Actinokineospora sp. EG49. Mar. Drugs 2014, 12, 1220–1244. [Google Scholar] [CrossRef]
- Tareq, F.S.; Lee, M.A.; Lee, H.S.; Lee, J.S.; Lee, Y.J.; Shin, H.J. Gageostatins A-C, antimicrobial linear lipopeptides from a marine Bacillus subtilis. Mar. Drugs 2014, 12, 871–885. [Google Scholar] [CrossRef]
- Tareq, F.S.; Lee, M.A.; Lee, H.S.; Lee, Y.J.; Lee, J.S.; Hasan, C.M.; Islam, M.T.; Shin, H.J. Gageotetrins A-C, noncytotoxic antimicrobial linear lipopeptides from a marine bacterium Bacillus subtilis. Org. Lett. 2014, 16, 928–931. [Google Scholar] [CrossRef]
- Tareq, F.S.; Lee, M.A.; Lee, H.S.; Lee, Y.J.; Lee, J.S.; Hasan, C.M.; Islam, M.T.; Shin, H.J. Non-cytotoxic antifungal agents: Isolation and structures of gageopeptides A-D from a Bacillus strain 109GGC020. J. Agric. Food Chem. 2014, 62, 5565–5572. [Google Scholar] [CrossRef]
- Breitbart, M.; Rohwer, F. Here a virus, there a virus, everywhere the same virus? Trends Microbiol. 2005, 13, 278–284. [Google Scholar] [CrossRef] [PubMed]
- Suttle, C.A. Viruses in the sea. Nature 2005, 437, 356–361. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Song, Y.; Li, X.; Wang, X.; Ling, C.; Qin, X.; Zhou, Z.; Li, Q.; Wei, X.; Ju, J. Abyssomicin Monomers and Dimers from the Marine-Derived Streptomyces koyangensis SCSIO 5802. J. Nat. Prod. 2018, 81, 1892–1898. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Chen, Z.; Zhu, G.; Wang, L.; Du, Y.; Wang, Y.; Zhu, W. Phenolic polyketides from the marine alga-derived Streptomyces sp. OUCMDZ-3434. Tetrahedron 2017, 73, 5451–5455. [Google Scholar] [CrossRef]
- Kim, S.H.; Ha, T.K.; Oh, W.K.; Shin, J.; Oh, D.C. Antiviral Indolosesquiterpenoid Xiamycins C-E from a Halophilic Actinomycete. J. Nat. Prod. 2016, 79, 51–58. [Google Scholar] [CrossRef]
- Shin, D.; Byun, W.S.; Moon, K.; Kwon, Y.; Bae, M.; Um, S.; Lee, S.K.; Oh, D.C. Coculture of Marine Streptomyces sp. With Bacillus sp. Produces a New Piperazic Acid-Bearing Cyclic Peptide. Front. Chem. 2018, 6, 498. [Google Scholar] [CrossRef]
- Kawahara, T.; Izumikawa, M.; Kozone, I.; Hashimoto, J.; Kagaya, N.; Koiwai, H.; Komatsu, M.; Fujie, M.; Sato, N.; Ikeda, H.; et al. Neothioviridamide, a Polythioamide Compound Produced by Heterologous Expression of a Streptomyces sp. Cryptic RiPP Biosynthetic Gene Cluster. J. Nat. Prod. 2018, 81, 264–269. [Google Scholar] [CrossRef]
- Nie, Y.L.; Wu, Y.D.; Wang, C.X.; Lin, R.; Xie, Y.; Fang, D.S.; Jiang, H.; Lian, Y.Y. Structure elucidation and antitumour activity of a new macrolactam produced by marine-derived actinomycete Micromonospora sp. FIM05328. Nat. Prod. Res. 2018, 32, 2133–2138. [Google Scholar] [CrossRef]
- Kim, M.C.; Machado, H.; Jang, K.H.; Trzoss, L.; Jensen, P.R.; Fenical, W. Integration of Genomic Data with NMR Analysis Enables Assignment of the Full Stereostructure of Neaumycin B, a Potent Inhibitor of Glioblastoma from a Marine-Derived Micromonospora. J. Am. Chem. Soc. 2018, 140, 10775–10784. [Google Scholar] [CrossRef]
- Saha, S.; Zhang, W.; Zhang, G.; Zhu, Y.; Chen, Y.; Liu, W.; Yuan, C.; Zhang, Q.; Zhang, H.; Zhang, L.; et al. Activation and characterization of a cryptic gene cluster reveals a cyclization cascade for polycyclic tetramate macrolactams. Chem. Sci. 2017, 8, 1607–1612. [Google Scholar] [CrossRef] [PubMed]
- Ye, X.; Anjum, K.; Song, T.; Wang, W.; Liang, Y.; Chen, M.; Huang, H.; Lian, X.Y.; Zhang, Z. Antiproliferative cyclodepsipeptides from the marine actinomycete Streptomyces sp. P11-23B downregulating the tumor metabolic enzymes of glycolysis, glutaminolysis, and lipogenesis. Phytochemistry 2017, 135, 151–159. [Google Scholar] [CrossRef]
- Zhang, W.; Che, Q.; Tan, H.; Qi, X.; Li, J.; Li, D.; Gu, Q.; Zhu, T.; Liu, M. Marine Streptomyces sp. derived antimycin analogues suppress HeLa cells via depletion HPV E6/E7 mediated by ROS-dependent ubiquitin-proteasome system. Sci. Rep. 2017, 7, 42180. [Google Scholar] [CrossRef] [PubMed]
- Han, X.; Liu, Z.; Zhang, Z.; Zhang, X.; Zhu, T.; Gu, Q.; Li, W.; Che, Q.; Li, D. Geranylpyrrol A and Piericidin F from Streptomyces sp. CHQ-64 DeltardmF. J. Nat. Prod. 2017, 80, 1684–1687. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Zhang, Y.; Wang, M.; Tan, Y.; Hu, X.; He, H.; Xiao, C.; You, X.; Wang, Y.; Gan, M. Neo-actinomycins A and B, natural actinomycins bearing the 5H-oxazolo [4–b] phenoxazine chromophore, from the marine-derived Streptomyces sp. IMB094. Sci. Rep. 2017, 7, 3591. [Google Scholar] [CrossRef]
- Perez, M.; Schleissner, C.; Fernandez, R.; Rodriguez, P.; Reyes, F.; Zuniga, P.; de la Calle, F.; Cuevas, C. PM100117 and PM100118, new antitumor macrolides produced by a marine Streptomyces caniferus GUA-06-05-006A. J. Antibiot. 2016, 69, 388–394. [Google Scholar] [CrossRef]
- Lin, Z.; Koch, M.; Pond, C.D.; Mabeza, G.; Seronay, R.A.; Concepcion, G.P.; Barrows, L.R.; Olivera, B.M.; Schmidt, E.W. Structure and activity of lobophorins from a turrid mollusk-associated Streptomyces sp. J. Antibiot. 2014, 67, 121–126. [Google Scholar] [CrossRef]
- Sun, Y.; Tomura, T.; Sato, J.; Iizuka, T.; Fudou, R.; Ojika, M. Isolation and Biosynthetic Analysis of Haliamide, a New PKS-NRPS Hybrid Metabolite from the Marine Myxobacterium Haliangium ochraceum. Molecules 2016, 21, 59. [Google Scholar] [CrossRef]
- Rodrigues, A.M.S.; Rohée, C.; Fabre, T.; Batailler, N.; Sautel, F.; Carletti, I.; Nogues, S.; Suzuki, M.T.; Stien, D. Cytotoxic indole alkaloids from Pseudovibrio denitrificans BBCC725. Tetrahedron Lett. 2017, 58, 3172–3173. [Google Scholar] [CrossRef]
- Schleissner, C.; Canedo, L.M.; Rodriguez, P.; Crespo, C.; Zuniga, P.; Penalver, A.; de la Calle, F.; Cuevas, C. Bacterial Production of a Pederin Analogue by a Free-Living Marine Alphaproteobacterium. J. Nat. Prod. 2017, 80, 2170–2173. [Google Scholar] [CrossRef]
- Bauer, A.W.; Perry, D.M.; Kirby, W.M. Single-disk antibiotic-sensitivity testing of staphylococci; an analysis of technique and results. AMA Arch. Intern. Med. 1959, 104, 208–216. [Google Scholar] [CrossRef]
- Jorgensen, J.H.; Turnidge, J.D. Susceptibility test methods: Dilution and disk diffusion methods. Man. Clin. Microbiol. 2015, 1253–1273. [Google Scholar]
- Bauer, A.W.; Kirby, W.M.; Sherris, J.C.; Turck, M. Antibiotic susceptibility testing by a standardized single disk method. Am. J. Clin. Pathol. 1966, 45, 493–496. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Oviedo, E.; Aller, A.I.; Martin, C.; Castro, C.; Ramirez, M.; Peman, J.M.; Canton, E.; Almeida, C.; Martin-Mazuelos, E. Evaluation of disk diffusion method for determining posaconazole susceptibility of filamentous fungi: Comparison with CLSI broth microdilution method. Antimicrob. Agents Chemother. 2006, 50, 1108–1111. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Reller, L.B.; Weinstein, M.; Jorgensen, J.H.; Ferraro, M.J. Antimicrobial Susceptibility Testing: A Review of General Principles and Contemporary Practices. Clin. Infect. Dis. 2009, 49, 1749–1755. [Google Scholar] [CrossRef]
- Le Bras, J.; Clain, J.R.; Secor, W.E. Susceptibility Test Methods: Parasites. In Manual of Clinical Microbiology, 11th ed.; American Society of Microbiology: Washington, DC, USA, 2015. [Google Scholar] [CrossRef]
- Gupta, S.; Nishi. Visceral leishmaniasis: Experimental models for drug discovery. Indian J. Med. Res. 2011, 133, 27–39. [Google Scholar] [PubMed]
- Sayed, A.A.; Simeonov, A.; Thomas, C.J.; Inglese, J.; Austin, C.P.; Williams, D.L. Identification of oxadiazoles as new drug leads for the control of schistosomiasis. Nat. Med. 2008, 14, 407–412. [Google Scholar] [CrossRef]
- Henderson, D.A. The eradication of smallpox--an overview of the past, present, and future. Vaccine 2011, 29 (Suppl. 4), D7–D9. [Google Scholar] [CrossRef]
- Burke, J.D.; Fish, E.N. Antiviral Strategies: The Present and Beyond. Curr. Mol. Pharmacol. 2009, 2, 32–39. [Google Scholar] [CrossRef]
- Rumlova, M.; Ruml, T. In vitro methods for testing antiviral drugs. Biotechnol. Adv. 2018, 36, 557–576. [Google Scholar] [CrossRef]
- Baer, A.; Kehn-Hall, K. Viral concentration determination through plaque assays: Using traditional and novel overlay systems. J. Vis. Exp. 2014, e52065. [Google Scholar] [CrossRef] [PubMed]
- Hayden, F.G.; Cote, K.M.; Douglas, R.G., Jr. Plaque inhibition assay for drug susceptibility testing of influenza viruses. Antimicrob. Agents Chemother. 1980, 17, 865–870. [Google Scholar] [CrossRef] [PubMed]
- Yin, Y.; Xu, Y.; Ou, Z.; Yang, X.; Liu, H. An antiviral drug screening system for enterovirus 71 based on an improved plaque assay: A potential high-throughput method. J. Med. Virol. 2019, 91, 1440–1447. [Google Scholar] [CrossRef] [PubMed]
- Landry, M.L.; Stanat, S.; Biron, K.; Brambilla, D.; Britt, W.; Jokela, J.; Chou, S.; Drew, W.L.; Erice, A.; Gilliam, B.; et al. A standardized plaque reduction assay for determination of drug susceptibilities of cytomegalovirus clinical isolates. Antimicrob. Agents Chemother. 2000, 44, 688–692. [Google Scholar] [CrossRef] [PubMed]
- Fang, J.; Sun, L.; Peng, G.; Xu, J.; Zhou, R.; Cao, S.; Chen, H.; Song, Y. Identification of three antiviral inhibitors against Japanese encephalitis virus from library of pharmacologically active compounds 1280. PLoS ONE 2013, 8, e78425. [Google Scholar] [CrossRef] [PubMed]
- Rumschlag-Booms, E.; Zhang, H.; Soejarto, D.D.; Fong, H.H.; Rong, L. Development of an Antiviral Screening Protocol: One-Stone-Two-Birds. J. Antivir. Antiretrovir. 2011, 3, 8–10. [Google Scholar] [CrossRef]
- Naarding, M.A.; Fernandez, N.; Kappes, J.C.; Hayes, P.; Ahmed, T.; Icyuz, M.; Edmonds, T.G.; Bergin, P.; Anzala, O.; Hanke, T.; et al. Development of a luciferase based viral inhibition assay to evaluate vaccine induced CD8 T-cell responses. J. Immunol. Methods 2014, 409, 161–173. [Google Scholar] [CrossRef]
- Matza-Porges, S.; Eisen, K.; Ibrahim, H.; Haberman, A.; Fridlender, B.; Joseph, G. A new antiviral screening method that simultaneously detects viral replication, cell viability, and cell toxicity. J. Virol. Methods 2014, 208, 138–143. [Google Scholar] [CrossRef]
- Wright, J.C.; Prigot, A.; Wright, B.; Weintraub, S.; Wright, L.T. An evaluation of folic acid antagonists in adults with neoplastic diseases: A study of 93 patients with incurable neoplasms. J. Natl. Med. Assoc. 1951, 43, 211–240. [Google Scholar]
- Gillet, J.P.; Varma, S.; Gottesman, M.M. The clinical relevance of cancer cell lines. J. Natl. Cancer Inst. 2013, 105, 452–458. [Google Scholar] [CrossRef]
- Skehan, P.; Storeng, R.; Scudiero, D.; Monks, A.; McMahon, J.; Vistica, D.; Warren, J.T.; Bokesch, H.; Kenney, S.; Boyd, M.R. New Colorimetric Cytotoxicity Assay for Anticancer-Drug Screening. JNCI 1990, 82, 1107–1112. [Google Scholar] [CrossRef] [PubMed]
- Strober, W. Trypan blue exclusion test of cell viability. Curr. Protoc. Immunol. 2001. [Google Scholar] [CrossRef]
- Stockert, J.C.; Horobin, R.W.; Colombo, L.L.; Blazquez-Castro, A. Tetrazolium salts and formazan products in Cell Biology: Viability assessment, fluorescence imaging, and labeling perspectives. Acta Histochem. 2018, 120, 159–167. [Google Scholar] [CrossRef] [PubMed]
- Shum, D.; Radu, C.; Kim, E.; Cajuste, M.; Shao, Y.; Seshan, V.E.; Djaballah, H. A high density assay format for the detection of novel cytotoxic agents in large chemical libraries. J. Enzyme. Inhib. Med. Chem. 2008, 23, 931–945. [Google Scholar] [CrossRef]
- Riss, T.; Moravec, R.; Beck, M.; Hannah, R.; Wilson, K.; Swanson, R. CellTiter-Glo Luminescent Cell Viability Assay: Fast, Sensitive and Flexible. Promega Notes Mag. 2002, 81, 2–5. [Google Scholar]
- Aslantürk, O.S. In Vitro Cytotoxicity and Cell Viability Assays: Principles, Advantages, and Disadvantages, Genotoxicity. In Genotoxicity—A Predictable Risk to Our Actual World; Larramendy, M.L., Soloneski, S., Eds.; IntechOpen: London, UK, 2017. [Google Scholar] [CrossRef]
- Munshi, A.; Hobbs, M.; Meyn, R.E. Clonogenic cell survival assay. Methods Mol. Med. 2005, 110, 21–28. [Google Scholar] [CrossRef] [PubMed]
- Madhavan, H. Simple Laboratory methods to measure cell proliferation using DNA synthesis property. J. Stem Cells Regen. Med. 2007, 3, 12–14. [Google Scholar]
- Demain, A.L.; Fang, A. The natural functions of secondary metabolites. Adv. Biochem. Eng. Biotechnol. 2000, 69, 1–39. [Google Scholar] [CrossRef]
- Donadio, S.; Monciardini, P.; Sosio, M. Polyketide synthases and nonribosomal peptide synthetases: The emerging view from bacterial genomics. Nat. Prod. Rep. 2007, 24, 1073–1109. [Google Scholar] [CrossRef]
- Kim, T.K.; Garson, M.J.; Fuerst, J.A. Marine actinomycetes related to the “Salinospora” group from the Great Barrier Reef sponge Pseudoceratina clavata. Environ. Microbiol. 2005, 7, 509–518. [Google Scholar] [CrossRef]
- Neilan, B.A.; Dittmann, E.; Rouhiainen, L.; Bass, R.A.; Schaub, V.; Sivonen, K.; Borner, T. Nonribosomal peptide synthesis and toxigenicity of cyanobacteria. J. Bacteriol. 1999, 181, 4089–4097. [Google Scholar] [CrossRef] [PubMed]
- Graça, A.P.; Calisto, R.; Lage, O.M. Planctomycetes as Novel Source of Bioactive Molecules. Front. Microbiol. 2016, 7, 1241. [Google Scholar] [CrossRef]
- Santos, J.D.; Vitorino, I.; de la Cruz, M.; Diaz, C.; Cautain, B.; Annang, F.; Perez-Moreno, G.; Gonzalez, I.; Tormo, J.R.; Martin, J.; et al. Diketopiperazines and other bioactive compounds from bacterial symbionts of marine sponges. Antonie Van Leeuwenhoek 2020. [Google Scholar] [CrossRef] [PubMed]
- Santos, J.D.; Vitorino, I.; De la Cruz, M.; Diaz, C.; Cautain, B.; Annang, F.; Perez-Moreno, G.; Gonzalez Martinez, I.; Tormo, J.R.; Martin, J.M.; et al. Bioactivities and Extract Dereplication of Actinomycetales Isolated From Marine Sponges. Front. Microbiol. 2019, 10, 727. [Google Scholar] [CrossRef]
- Ziemert, N.; Alanjary, M.; Weber, T. The evolution of genome mining in microbes—A review. Nat. Prod. Rep. 2016, 33, 988–1005. [Google Scholar] [CrossRef] [PubMed]
- van Heel, A.J.; de Jong, A.; Song, C.; Viel, J.H.; Kok, J.; Kuipers, O.P. BAGEL4: A user-friendly web server to thoroughly mine RiPPs and bacteriocins. Nucleic Acids Res. 2018, 46, W278–W281. [Google Scholar] [CrossRef]
- Blin, K.; Shaw, S.; Steinke, K.; Villebro, R.; Ziemert, N.; Lee, S.Y.; Medema, M.H.; Weber, T. antiSMASH 5.0: Updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res. 2019, 47, W81–W87. [Google Scholar] [CrossRef]
- Wiegand, S.; Jogler, M.; Jogler, C. On the maverick Planctomycetes. FEMS Microbiol. Rev. 2018, 42, 739–760. [Google Scholar] [CrossRef]
- Wiegand, S.; Jogler, M.; Boedeker, C.; Pinto, D.; Vollmers, J.; Rivas-Marin, E.; Kohn, T.; Peeters, S.H.; Heuer, A.; Rast, P.; et al. Cultivation and functional characterization of 79 planctomycetes uncovers their unique biology. Nat. Microbiol. 2020, 5, 126–140. [Google Scholar] [CrossRef]
- Jeske, O.; Jogler, M.; Petersen, J.; Sikorski, J.; Jogler, C. From genome mining to phenotypic microarrays: Planctomycetes as source for novel bioactive molecules. Antonie Van Leeuwenhoek 2013, 104, 551–567. [Google Scholar] [CrossRef]
- Inglese, J.; Johnson, R.L.; Simeonov, A.; Xia, M.; Zheng, W.; Austin, C.P.; Auld, D.S. High-throughput screening assays for the identification of chemical probes. Nat. Chem. Biol. 2007, 3, 466–479. [Google Scholar] [CrossRef] [PubMed]
- Macarrón, R.; Hertzberg, R.P. Design and Implementation of High Throughput Screening Assays. Mol. Biotechnol. 2011, 47, 270–285. [Google Scholar] [CrossRef] [PubMed]
- Gaudencio, S.P.; Pereira, F. Dereplication: Racing to speed up the natural products discovery process. Nat. Prod. Rep. 2015, 32, 779–810. [Google Scholar] [CrossRef] [PubMed]
- Berdy, J. Thoughts and facts about antibiotics: Where we are now and where we are heading. J. Antibiot. 2012, 65, 385–395. [Google Scholar] [CrossRef]
- Koehn, F.E.; Carter, G.T. The evolving role of natural products in drug discovery. Nat. Rev. Drug Discov. 2005, 4, 206–220. [Google Scholar] [CrossRef]
- Ito, T.; Masubuchi, M. Dereplication of microbial extracts and related analytical technologies. J. Antibiot. 2014, 67, 353–360. [Google Scholar] [CrossRef]
- Takagi, M.; Shin-Ya, K. Construction of a natural product library containing secondary metabolites produced by actinomycetes. J. Antibiot. 2012, 65, 443–447. [Google Scholar] [CrossRef]
- Wolfender, J.L.; Marti, G.; Thomas, A.; Bertrand, S. Current approaches and challenges for the metabolite profiling of complex natural extracts. J. Chromatogr. A 2015, 1382, 136–164. [Google Scholar] [CrossRef]
- Perez-Victoria, I.; Martin, J.; Reyes, F. Combined LC/UV/MS and NMR Strategies for the Dereplication of Marine Natural Products. Planta. Med. 2016, 82, 857–871. [Google Scholar] [CrossRef]
- Eugster, P.J.; Guillarme, D.; Rudaz, S.; Veuthey, J.L.; Carrupt, P.A.; Wolfender, J.L. Ultra high pressure liquid chromatography for crude plant extract profiling. J. AOAC Int. 2011, 94, 51–70. [Google Scholar] [CrossRef]
- Ganzera, M.; Sturm, S. Recent advances on HPLC/MS in medicinal plant analysis—An update covering 2011–2016. J. Pharm. Biomed. Anal. 2018, 147, 211–233. [Google Scholar] [CrossRef]
- Steinmann, D.; Ganzera, M. Recent advances on HPLC/MS in medicinal plant analysis. J. Pharm. Biomed. Anal. 2011, 55, 744–757. [Google Scholar] [CrossRef]
- Boswell, P.G.; Schellenberg, J.R.; Carr, P.W.; Cohen, J.D.; Hegeman, A.D. A study on retention “projection” as a supplementary means for compound identification by liquid chromatography-mass spectrometry capable of predicting retention with different gradients, flow rates, and instruments. J. Chromatogr. A 2011, 1218, 6732–6741. [Google Scholar] [CrossRef]
- Nielsen, K.F.; Mansson, M.; Rank, C.; Frisvad, J.C.; Larsen, T.O. Dereplication of microbial natural products by LC-DAD-TOFMS. J. Nat. Prod. 2011, 74, 2338–2348. [Google Scholar] [CrossRef]
- Yates III, J.R. A century of mass spectrometry: From atoms to proteomes. Nat. Methods 2011, 8, 633–637. [Google Scholar] [CrossRef]
- Nguyen, D.D.; Wu, C.H.; Moree, W.J.; Lamsa, A.; Medema, M.H.; Zhao, X.; Gavilan, R.G.; Aparicio, M.; Atencio, L.; Jackson, C.; et al. MS/MS networking guided analysis of molecule and gene cluster families. Proc. Natl. Acad. Sci. USA 2013, 110, E2611–E2620. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, K.F.; Smedsgaard, J. Fungal metabolite screening: Database of 474 mycotoxins and fungal metabolites for dereplication by standardised liquid chromatography-UV-mass spectrometry methodology. J. Chromatogr. A 2003, 1002, 111–136. [Google Scholar] [CrossRef]
- Lang, G.; Mayhudin, N.A.; Mitova, M.I.; Sun, L.; van der Sar, S.; Blunt, J.W.; Cole, A.L.J.; Ellis, G.; Laatsch, H.; Munro, M.H.G. Evolving Trends in the Dereplication of Natural Product Extracts: New Methodology for Rapid, Small-Scale Investigation of Natural Product Extracts. J. Nat. Prod. 2008, 71, 1595–1599. [Google Scholar] [CrossRef] [PubMed]
- Cifani, P.; Kentsis, A. High Sensitivity Quantitative Proteomics Using Automated Multidimensional Nano-flow Chromatography and Accumulated Ion Monitoring on Quadrupole-Orbitrap-Linear Ion Trap Mass Spectrometer. Mol. Cell Proteom. 2017, 16, 2006–2016. [Google Scholar] [CrossRef] [PubMed]
- Scifinder. Chemical Abstracts Service’s Registry File. Available online: http://www.cas.org/products/scifinder (accessed on 13 June 2020).
- Farnsworth, N.R. NAPRALERT. Available online: https://www.napralert.org/ (accessed on 13 June 2020).
- Laatsch, H. AntiBase: The Natural Compound Identifier. Available online: https://www.wiley.com/en-us/AntiBase%3A+The+Natural+Compound+Identifier-p-9783527343591 (accessed on 13 June 2020).
- RSC. MarinLit. A Database of the Marine Natural Products Literature. Available online: http://pubs.rsc.org/marinlit/ (accessed on 13 June 2020).
- Wang, M.; Carver, J.J.; Phelan, V.V.; Sanchez, L.M.; Garg, N.; Peng, Y.; Nguyen, D.D.; Watrous, J.; Kapono, C.A.; Luzzatto-Knaan, T.; et al. Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking. Nat. Biotechnol. 2016, 34, 828–837. [Google Scholar] [CrossRef]
- Dorrestein, P.C. Global Natural Products Social Networking (GNPS). Available online: https://gnps.ucsd.edu/ProteoSAFe/static/gnps-splash.jsp (accessed on 16 June 2020).
- van Santen, J.A.; Jacob, G.; Singh, A.L.; Aniebok, V.; Balunas, M.J.; Bunsko, D.; Neto, F.C.; Castano-Espriu, L.; Chang, C.; Clark, T.N.; et al. The Natural Products Atlas: An Open Access Knowledge Base for Microbial Natural Products Discovery. ACS Cent. Sci. 2019, 5, 1824–1833. [Google Scholar] [CrossRef] [PubMed]
- Linington, R.G. The Natural Products Atlas. Available online: https://www.npatlas.org/joomla/ (accessed on 13 June 2020).
- Zhang, C.; Idelbayev, Y.; Roberts, N.; Tao, Y.; Nannapaneni, Y.; Duggan, B.M.; Min, J.; Lin, E.C.; Gerwick, E.C.; Cottrell, G.W.; et al. Small Molecule Accurate Recognition Technology (SMART) to Enhance Natural Products Research. Sci. Rep. 2017, 7, 14243. [Google Scholar] [CrossRef] [PubMed]
- Gerwick, W.H. Small Molecule Accurate Recognition Technology (SMART). Available online: https://smart.ucsd.edu/ (accessed on 13 June 2020).
- Nothias, L.F.; Nothias-Esposito, M.; da Silva, R.; Wang, M.; Protsyuk, I.; Zhang, Z.; Sarvepalli, A.; Leyssen, P.; Touboul, D.; Costa, J.; et al. Bioactivity-Based Molecular Networking for the Discovery of Drug Leads in Natural Product Bioassay-Guided Fractionation. J. Nat. Prod. 2018, 81, 758–767. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.; van der Heul, H.U.; Melnik, A.V.; Lubben, J.; Dorrestein, P.C.; Minnaard, A.J.; Choi, Y.H.; van Wezel, G.P. Lugdunomycin, an Angucycline-Derived Molecule with Unprecedented Chemical Architecture. Angew. Chem. Int. Ed. Engl. 2019, 58, 2809–2814. [Google Scholar] [CrossRef] [PubMed]
- Maglangit, F.; Fang, Q.; Leman, V.; Soldatou, S.; Ebel, R.; Kyeremeh, K.; Deng, H. Accramycin A, a New Aromatic Polyketide, from the Soil Bacterium, Streptomyces sp. MA37. Molecules 2019, 24, 3384. [Google Scholar] [CrossRef] [PubMed]
- Mudalungu, C.M.; von Torne, W.J.; Voigt, K.; Ruckert, C.; Schmitz, S.; Sekurova, O.N.; Zotchev, S.B.; Sussmuth, R.D. Noursamycins, Chlorinated Cyclohexapeptides Identified from Molecular Networking of Streptomyces noursei NTR-SR4. J. Nat. Prod. 2019, 82, 1478–1486. [Google Scholar] [CrossRef]
- Reher, R.; Kim, H.W.; Zhang, C.; Mao, H.H.; Wang, M.; Nothias, L.F.; Caraballo-Rodriguez, A.M.; Glukhov, E.; Teke, B.; Leao, T.; et al. A Convolutional Neural Network-Based Approach for the Rapid Annotation of Molecularly Diverse Natural Products. J. Am. Chem. Soc. 2020, 142, 4114–4120. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).